Abstract
Usher syndrome is a rare disorder causing retinitis pigmentosa, together with sensorineural hearing loss. Due to the phenotypic and genetic heterogeneity of this disease, the best method to screen the causative mutations is by high-throughput sequencing. In this study, we tested a semiconductor chip based sequencing approach with 77 unrelated patients, as a molecular diagnosis routine. In addition, Multiplex Ligation-dependent Probe Amplification and microarray-based Comparative Genomic Hybridization techniques were applied to detect large rearrangements, and minigene assays were performed to confirm the mRNA processing aberrations caused by splice-site mutations. The designed panel included all the USH causative genes (MYO7A, USH1C, CDH23, PCDH15, USH1G, CIB2, USH2A, ADGRV1, WHRN and CLRN1) as well as four uncertainly associated genes (HARS, PDZD7, CEP250 and C2orf71). The outcome showed an overall mutation detection ratio of 82.8% and allowed the identification of 42 novel putatively pathogenic mutations. Furthermore, we detected two novel nonsense mutations in CEP250 in a patient with a disease mimicking Usher syndrome that associates visual impairment due to cone-rod dystrophy and progressive hearing loss. Therefore, this approach proved reliable results for the molecular diagnosis of the disease and also allowed the consolidation of the CEP250 gene as disease causative for an Usher-like phenotype.
Similar content being viewed by others
Introduction
Usher syndrome (USH) is a rare autosomal recessive disease that associates retinitis pigmentosa (RP), sensorineural hearing loss (SNHL) and, in some cases, vestibular dysfunction. It is the most common form of hereditary disease combining hearing and vision impairment, with a prevalence ranging from 3 to 6.2 per 100,0001,2. Three types of USH are distinguished depending on the severity and progression of the pathology: Type 1 (USH I) is typically characterized by a severe-profound congenital hearing loss, onset of RP usually within the first decade of life, and vestibular dysfunction. Type 2 (USH II) patients present with a moderate-severe congenital hearing impairment, a pubertal onset of RP and normal vestibular function. Type 3 (USH III) is defined by progressive hearing loss starting after post-lingual phase and an age-variable onset of RP, whereas the vestibular dysfunction is variable3. Despite the three major divisions of the disorder, some patients display a clinical profile not matching any of these categories, being classified as atypical USH.
As well as clinically, USH is genetically heterogeneous. To date, 13 genes have been associated with the disease and these do not explain all the reported cases, suggesting other still unknown genes may be responsible for the disorder4.
USH I is commonly caused by mutations in six genes: MYO7A, USH1C, CDH23, PCDH15, USH1G and CIB2. On the other hand, USH2A, ADGRV1 and WHRN are the three genes usually responsible for USH II, whilst the CLRN1 gene is the only one currently associated to USH III cases.
In addition, other genes have been related to the disease. The PDZD7 gene has been reported to behave as a modifier of retinal disease with USH2A and a contributor to digenic inheritance with ADGRV15. Recently, HARS was proposed as a novel causative gene of USH III, based on a mutation found in two patients6 and CEP250 has been reported as responsible for an atypical Usher syndrome with SNHL and a relatively mild RP7.
Most of the USH-causing mutations are private and most of the involved genes are of a large size. These issues can be overcome with the use of high-throughput sequencing (HTS) tools, which enable a rapid, feasible method for the genetic diagnosis of the disease, and they are being increasingly employed8,9,10,11,12. The main objective of the present study is the molecular diagnosis of a large cohort of USH patients by means of a HTS screening. Thus, we developed a custom targeted exome design, including the ten disease causative genes and four additional candidates, for its use in Ion Torrent platforms.
Methods
Patients
A cohort of 77 USH patients was selected for this study. The probands were classified into the different USH subtypes according to their clinical records. The data (when feasible) consisted of the patient’s ophthalmological studies, including best-corrected visual acuity measurements (BCVA), fundus ophthalmoscopy, visual field examination and electrophysiological examination; and audiological tests13,14,15. Hearing loss severity was established as mild (between >25 and ≤40 dB), moderate (between >40 and ≤70 dB) or severe/profound (>70 dB). Patients presenting a bilateral severe congenital hearing loss (>70 dB), early RP onset and altered vestibular function were diagnosed as USH I. Patients suffering from bilateral congenital moderate-severe hearing loss (40–70 dB) and adolescent-to-adult onset of RP were categorized as USH II. If the patients displayed progressive hearing loss, with or without vestibular dysfunction, and late onset RP, were recognized as USH III. Patients with a profile not quite matching any of these three categories were diagnosed as atypical USH cases. When the clinical data was insufficient, the type was stated as general USH. For case RP1973, further ophthalmological examinations were performed, which included measurements of fundus autofluorescence (FAF), optical coherence tomography (OCT) (acquired with a Heidelberg Spectralis OCT Bluepeak) and visual fields 30-2 and 120-2 strategies by the Humphrey Visual Field Analyzer. Full-field electroretinography was performed according to the International Society for Clinical Electrophysiology of Vision Standards16.
From all the patients included, 19 were assigned to a test group in order to evaluate the sequencing platform performance. Eight cases out of these already had a complete molecular diagnosis (at least two USH causative mutations) and 11 where partially solved with only one previously known disease causing mutation. The test group comprised a total of 4 Copy Number Variations (CNVs) and 22 point mutations, represented by variants of different nature (Table 1). Finally, a cohort of 58 previously unscreened USH patients of Spanish origin were recruited for this study in order to determine their genetic diagnosis. Among these to be characterized, 15 were USH I, 31 USH II, and 12 undetermined USH.
Segregation analysis was performed by conventional Sanger sequencing when DNA samples of family members were available.
Ethics Statement
This study was approved by the Hospital La Fe Ethics Committee and authorizations from all the patients and the participating relatives were obtained by signing an informed consent form. All research was performed in accordance with the relevant guidelines and regulations.
Samples
Genomic DNA (gDNA) from the probands was obtained and purified using standard procedures. The concentration of the resulting DNA samples was determined with Nanodrop and Qubit fluorometer (Thermo Fisher Scientific).
Targeted USH exome sequencing design
A customized AmpliSeq panel was designed using Ion AmpliSeq Designer tool from Thermo Fisher Scientific (www.ampliseq.com) to generate the targeted library. The designed targeted exome (Table 2) included all exons contemplated in all isoforms of 14 genes: the 10 USH causative genes (MYO7A, USH1C, CDH23, PCDH15, USH1G, CIB2, USH2A, ADGRV1, WHRN and CLRN1), the additional locus comprising the c.7595 − 2144A > G intronic mutation in USH2A17, and 4 USH associated genes (HARS, PDZD7, CEP250 and C2orf71).
Sequence enrichment and HTS
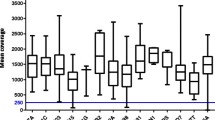
The amplification of the targets was performed according to the Ion AmpliSeq Library Kit 2.0 protocol (Thermo Fisher Scientific) for Ion Torrent sequencing. The sequencing was carried out with a theoretical minimum coverage of 500x either on the PGM (Ion 318 chip, 500 flows) or Proton system (Ion PI chip, 520 flows).
Variant filtering and analysis
The resulting sequencing data were analyzed with Ion Reporter Software tool (https://ionreporter.thermofisher.com) in regard to the human assembly GRCh37/hg19. The annotated variants were filtered according to a Minor Allele Frequency (MAF) value ≤0.01, the frequency of the variants was explored in the Exome Aggregation Consortium (ExAC) database, their annotation in the dbSNP (www.ncbi.nlm.nih.gov/SNP/), their description in the Usher syndrome mutation database (https://grenada.lumc.nl/LOVD2/Usher_montpellier/) and the mutation type.
In order to determine the pathogenicity of novel missense or splice-site mutations, the variants were analyzed using several in silico prediction tools according to the nature of the mutation. Aminoacid change effects were examinated using the SIFT18, PolyPhen-219 and PROVEAN20 programs and the additional tools ATGpr21, NetStart22 and TIS Miner23 were applied when concerning translation start loss variants. Putative variants affecting the splicing pattern were investigated with the Human Splicing Finder 3.124, MaxEnt25 and NNSplice26 algorithms.
All the putative pathogenic variants were validated through conventional Sanger sequencing. All the poorly or null covered regions were screened by conventional Sanger for all the cases with only one or none putative disease causing mutations detected through HTS. Additionally, the same patients were screened by Sanger sequencing for recently identified deep intronic mutations that were published after the start of this study and could therefore not be included in the panel design: four in USH2A, namely c.14134 − 3169A > G27, c.5573 − 843A > G, c.8845 + 628C > T, c.9959 − 4159A > G28; and variant c.254–649T > G of CLRN129.
Copy Number Variation analysis
The screening for large rearrangements was performed in all patients where either none or only one mutation was detected with the panel, using either multiplex ligation-dependent probe amplification (MLPA) or a custom microarray-based Comparative Genomic Hybridisation (aCGH).
The MLPA technique (MRC-Holland) allows the identification of large rearrangements for the USH2A (probemixes P361 and P362) and PCDH15 (probemix P292) genes.
In order to screen possible CNVs in the remaining genes, aCGH was designed covering all the genes included in this study. The resulting custom 60 K microarray (Agilent Technologies, AMADID-082310) contained 62976 probes. Three DNA samples with known CNVs from the test group (RP1895, RP1522, RP1034) were used as positive controls in the first batch of the analysis, in order to validate the custom design of the array.The gDNAs were prepared according to the manufacturer’s protocol, as described before30.
Splicing Effect Analysis by Minigenes
Minigene assay was performed for all novel intronic mutations found in this study in order to confirm the splicing alterations, adopting a procedure previously described31 and using HEK293 cells. All experiments were performed in duplicate.
Extended exome screening
Whole exome sequencing (WES) using SureSelect Human All Exon V6 kit (Agilent Techologies) for Illumina platform was performed for sample RP1973 to discard mutations in other genes, in view of the results obtained for this case.
Results
Test group
The sequencing results allowed us to identify 20 of the 22 point mutations from the positive controls, while none of the CNVs was detected. When forcing a general parameters relaxation, the two previously undetected changes consisting of frameshift duplications were then recognized. These results and detailed information are summarized in Table 1.
In this study, the second disease causing mutation was detected in 7 out of the 11 patients included in the test group in whom only one of the pathological alleles was previously registered and, therefore, their genetic diagnosis was fulfilled (Table 3). These included 6 novel mutations: two nonsense mutations, two CNVs, 1 frameshift and 1 splice-site mutation.
Cohort of previously unscreened USH patients
We identified both pathogenic variants in 45 out of the 58 the analyzed cases and in 6 patients only one mutation was detected. No likely pathogenic mutations were found in other 7 probands.
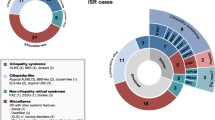
This work allowed to detect mutations of different nature, from which 42 were novel variants (Table 3). Among these novel changes, we were able to detect 8 missense, 14 nonsense, 11 frameshift, 4 splice-site, 1 start loss variant and 4 large rearrangements (Tables 3 and 4). Regarding all the detected mutations, USH2A shows the highest prevalence, accounting for 48% of the causative variants (Fig. 1).
Recurrence of mutated genes included in the design of this study and distribution of the type of mutations. The data includes all the disease causative variants from the previously unscreened cohort and from the seven ultimately solved patients of the test group, which at the beginning of the study had only one causing mutation identified and the second was finally detected with the technology used in this work. Abbreviations: PE, Pseudo-exon; InDel, Insertion/Deletion; CNV, Copy Number Variation.
Custom aCGH unveiled three CNVs in the ADGRV1 gene: one large heterozygous duplication involving exons 79–83 in patient RP580, the heterozygous deletion of exon 85 in patient RP1936, and one large homozygous deletion comprising exons 28–33 in proband RP2011. The latter was also suspected beforehand by the null coverage of that region on the HTS sequencing results.
Minigene splice assay analysis
We detected 3 canonical splice-site mutations (c.1691 − 1G > A, MYO7A; c.5776 + 1G > A, USH2A; c.12295 − 1G > A, USH2A) and one variant with dubious consequences (c.5314 − 5T > A, ADGRV1) (Table 4). These variant candidates were tested through minigenes and for all of them the exon skipping was proved, confirming therefore the pathogenicity of the mutations (Fig. 2).
Minigene assay results for the four splicing mutations. The gel electrophoresis displays the splicing outcome of the minigene transcription for the WT and mutant alleles. In vitro experiments were performed in duplicate and therefore the results show both repetitions. Sanger sequencing of the results confirm the splicing processes by evidencing the transcript joints. SD6 and SA2 are the exons included in the pSPL3 exon trapping vector used in the assay. (a) c.5314 − 5T > A (ADGRV1). Band A corresponds to the correct transcript of exon 25. Band B from the mutant construction denotes the skipping of the same exon. If the transcript harboring the mutation were translated, the newly generated protein would of 1,772 aminoacids in length, p.Asn1772*. (b) c.5776 + 1G > A (USH2A). Band A is the correct transcript corresponding to the exon 28 and Band B is the skipping of the same exon. If the aberrant transcript were translated, it would generate a new truncated protein of 5,134 aminoacids in length, p.Gly1858_Thr1925del. (c) c.1691 − 1G > A (MYO7A). Band A corresponds to the correct transcript of exon 25. Band B is the aberrant splicing process due to the new site generated by the lack of a guanine at the acceptor site, entailing therefore a frameshift effect. The fragment C corresponds to de exon skipping of exon 15. If the transcript with the mutation were translated, it would generate the two proteins p. Gly564Alafs*58 and p.Phe565Argfs*11. (d) c.12295 − 1G > A (USH2A). Band A corresponds to the correct transcript of exon 63 and band B, from the mutant allele, evidences the skipping of the exon. The displayed images of the gels have been cropped to improve the clarity of the presentation, and the full-length gels are presented in Supplementary Fig. S2.
For the c.1691 − 1G > A MYO7A mutation, the minigene assay was of particular interest. Besides the skipping of exon 15, the mutant allele displayed an additional aberrant band (Fig. 2c). This additional fragment corresponds to the recognition of the first guanine of the exon as the acceptor site (the mutation is a transition of G > A), resulting in a frameshift effect starting at the first base of the exon. The in silico algorithms had predicted the same consequence (Table 4).
Clinical description of RP1973
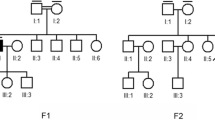
A remarkable case was RP1973, which was found to be a compound heterozygous for two nonsense mutations in CEP250. Both nonsense mutations segregate with the family, which is composed of both parents and an unaffected sibling (Fig. 3a). Patient RP1973 suffered from bilateral moderate-severe progressive hearing loss manifested at 13 years old (Fig. 3b) and late-onset progressive diminution of vision in both eyes with photophobia (first ophthalmologic examination at 44 years old). There is no history of any similar condition in any other family member. The BCVA was 0.6 in the right eye and 0.5 in the left eye (Snellen). The anterior segment findings were within normal limits, but fundus examination revealed migration of pigment in a bone-spicule pattern within a mid-peripheral annular zone of both eyes and narrowing of the peripheral retinal blood vessels (Fig. 3c). The left eye showed a glistening reflex of the inner retinal surface secondary to an epiretinal membrane. The macula of the right eye was relatively normal. Humphrey perimetry revealed peripheral visual field constriction with relative defects in the paracentral region in both eyes that has remained stable for the last five years. Macular autofluorescence images were normal in both eyes. The OCT shows a normal macular thickness with discontinuity of the outer segment layer of the photoreceptors around the foveal center in both eyes (Fig. 3c). Full-field electroretinography showed only mild alterations in the scotopic flash electroretinography (ERG), as the amplitudes of the b-wave were reduced in the right eye and absent in the left eye. Macular ERG showed an absence of response in both eyes, and Visual Evoked Potentials (VEP) were altered (Supplemental Fig. S1). Due to the symptoms, the nature of the variants leading to a premature stop codon and the co-segregation analysis we consider both mutations as disease causing for an USH-like phenotype.
Clinical and molecular data of patient RP1973 harboring the nonsense mutations in CEP250. (a) Family pedigree with the Sanger sequencing results revealing the segregation pattern of the mutations. (b) Audiometric results evincing the progression of the bilateral hearing loss. (c) Ocular phenotype. Upper images correspond to the right eye, bottom images are from the left eye. Fundus pictures showing pigment clumps (c1, c4) and thinning of the peripheral arterioles (c2, c5). OCT images of the foveal region showing loss and discontinuity of the retinal pigment epithelium layer (c3, c6). Abbreviations: yo, years old; dB, decibel; Hz, hertz.
The analysis of the targeted panel and WES results showed no additional putative pathogenic mutations, except for one heterozygous missense variant in USH2A (rs773526991: c.4561C > T/p.Arg1521Cys), presenting an allele frequency of 0.00002 in ExAC and for which the in silico tools implied a deleterious effect. Nevertheless, no other potential mutation was identified in USH2A.
Discussion
Usher syndrome is genetically heterogeneous mostly due to the high number of genes involved and their large size. The genetic etiopathogeny relies on all kinds of mutations among these genes and additionally, most of the variants are private. For that reason, it is very difficult to perform molecular diagnosis by conventional genotyping or direct gene sequencing. In our study, we were able to detect biallelic mutations in an USH gene in 45 out of the 58 previously unscreened patients (77.6%) and we identified 96 out of the 116 expected mutated alleles (82.8% detection ratio). That percentage difference is due to the fact that 6 cases were carriers of only one pathogenic variant (Table 3). The remaining undiscovered second mutation, as well as both variants of unresolved cases, may be located either in other USH responsible still unknown genes or in other non-coding regions that were not incorporated in our design. The pathogenic deep intronic mutation c.7595 − 2144A > G in USH2A was included in the study, but five other have been recently designated to be pathogenic27,28,29, which proves that still many deep intronic mutations may remain unveiled and further whole gene screen studies are of great interest.
The output of the analysis of the raw data is very dependent on the algorithm used for the mapping and variant calling. Two control variants, consisting of frameshift duplications, were detected only when relaxing the software quality parameters, suggesting a possible hindrance for the Ion Torrent mapping algorithm to align homopolymers. Indeed, other studies have reported these homopolymer-associated errors and even over and under-calling errors in non-homopolymer regions32,33. Additional factors for this technology suggested by other authors, such as the biases produced by the GC content or the underestimation of the quality scores32,34, probably contribute to the false negative calling errors.
The platform and panel design provided a 91% reliability based on the point mutation detection rate of the test group, but it reached a 100% of accuracy when thresholds of the mapping and annotation variables were decreased. However, no additional causative variants were found in the group of unresolved cases after applying the same procedure. Nevertheless, the failure to detect these variants could also fall on the HTS system used for the study, escaping variant detection independently on the resulting data management.
Among the study, two patients from the previously unscreened group presented mutations in genes that were not consistent with their clinical diagnosis, being these genes usually responsible for another USH subtype. One USH I patient (RP1748) carried biallelic mutations in USH2A and an USH II case in the MYO7A gene (RP1567). Still, the event of a molecular diagnosis not quite matching the clinical phenotype is not unusual and has been previously reported in other studies8,35. Indeed, this supports the further investigation of USH patients by HTS to establish better genotype – phenotype associations.
Many previous studies have evidenced the presence of large rearrangements among different USH populations, establishing them as a significant genetic alteration causing the disease. PCDH15 and USH2A are the most common genes displaying such CNVs30,36, but also large rearrangements have been found in MYO7A, CDH23 and ADGRV137,38.
A CNVs survey based on the coverage of the sequencing results was not possible due to the wide deviation of the target enrichment technique by loci amplification. However, large homozygous deletions could be inferred from null covered regions corresponding to several adjacent probes, when observed in punctual cases. The supplemental analysis by MLPA or aCGH allowed us to detect a total of five large rearrangements among the test group and the previously unscreened cohort, four of these rearrangements being novel. Concerning our series of patients without prior genetic diagnosis, the CNVs account for 5.2% of the total identified pathogenic alleles.
Four of the novel mutations were intronic variants located in splicing regions that, though all but one were set on canonical ±1 loci, a sort of functional analysis would provide further support of their pathogenicity. Certainly, the minigene assays proved that all these four mutations cause an aberrant splicing.
Regarding the compound heterozygous case for the two nonsense mutations in CEP250, our study provides sufficient data for the gene to be classified as USH-like causative. The association was firstly introduced in a study of a consanguineous family of Iranian Jewish origin characterized by early onset hearing loss and mild RP7 and, very recently, Kubota et al.39 presented a Japanese family carrying compound heterozygous nonsense mutations in CEP250, with a clinical phenotype of cone-rod dystrophy and sensorineural hearing loss. Our patient with the CEP250 mutations (RP1973) presented with progressive hearing loss and mild macular affectation with lowering of the visual acuity and photophobia, which are similar symptoms to those of the latter work, thus consolidating its role as a gene responsible for mimicking Usher syndrome. It has to be remarked that RP1973 shows a clearly progressive SNHL, yet the aforementioned studies do not give any details about the deafness evolution and, thus, a full comparative analysis is not feasible. There is another study correlating CEP250 with non-syndromic RP (nsRP) due to a detected homozygous missense mutation40. These findings are in agreement with ours and other authors observations that different diseases can be caused by the same gene depending on specific mutations, such as USH2A that can cause either nsRP or USH, or the USH genes MYO7A, USH1C, CDH23, PCDH15, USH1G, WHRN or CIB2 that can cause non syndromic hearing loss or USH41,42,43,44,45,46,47,48. In view of the different but closely related phenotypes associated to CEP250, thorough clinical examinations of the cases should be performed to better understand the consequences of mutations in this gene, particularly those regarding cone affectation.
In the last years, the USH molecular diagnosis through HTS approaches have replaced the traditional techniques based on Sanger sequencing35,49. The more recent next generation sequencing approaches enable a detection ratio between 50–100%, depending on the cohort and design of study8,9,12,50,51,52,53. Here, we provide a HTS method based on targeted exome library generation by amplification and the subsequent ion sensing-based sequencing that allows an average allele detection ratio compared to the other mentioned studies. It is, though, unfair to compare these varying efficiencies, since they do not only rely on the sequencing system, but also on the cohort selection criteria of the samples. For instance, the group analyzed by Qu et al.53, Besnard et al.50 and Eandi et al.9 consisted of only five, thirteen and seventeen probands respectively, a rather scarce number of samples that might bias the efficiency outcome. Additionally, those and other studies such as Aparisi et al.8 and Bonnet et al.51 involved only well USH characterized patients. Our study not only included a larger number of samples, but also some unclassified USH cases. Therefore, another partial reason for the unsolved cases could be the misdiagnosis of some patients as USH, who could harbor several mutations in other genes that together may mimic the syndrome. From the seven unresolved patients without genetic diagnosis, three lack in detailed clinical notes that clearly support the cases as USH. The remaining four patients do not fully harmonize with the syndrome, since they present a late-onset hearing impairment. If we were not to take these samples into account, the detection ratio would increase from the 82.8% up to 94.1%. Even displaying such a solid outcome, this HTS approach falls short of CNV detection, yet it allows the use of only 10 ng of starting DNA (admitting as well some degradation). All these features shall be taken in consideration, depending on the requirements and resources of each center and the group of study.
Our updated custom design for USH targeted exome sequencing is a reliable tool for molecular diagnosis of the disease, and its implementation in the health care system would lead to a great profit for the patients. Furthermore, CEP250 should be officially recognized as a gene causative of Usher-like syndrome.
Data Availability Statement
The datasets generated and analysed during the current study are available from the corresponding author on reasonable request.
References
Boughman, J. A., Vernon, M. & Shaver, K. A. Usher syndrome: definition and estimate of prevalence from two high-risk populations. J. Chronic Dis. 36, 595–603 (1983).
Cohen, M., Bitner-Glindzicz, M. & Luxon, L. The changing face of Usher syndrome: clinical implications. Int. J. Audiol. 46, 82–93 (2007).
Millán, J. M. et al. An Update on the Genetics of Usher Syndrome. J. Ophthalmol. 2011 (2011).
Mathur, P. & Yang, J. Usher syndrome: Hearing loss, retinal degeneration and associated abnormalities. Biochim. Biophys. Acta 1852, 406–420 (2015).
Ebermann, I. et al. PDZD7 is a modifier of retinal disease and a contributor to digenic Usher syndrome. J. Clin. Invest. 120, 1812–1823 (2010).
Puffenberger, E. G. et al. Genetic mapping and exome sequencing identify variants associated with five novel diseases. PloS One 7, e28936 (2012).
Khateb, S. et al. A homozygous nonsense CEP250 mutation combined with a heterozygous nonsense C2orf71 mutation is associated with atypical Usher syndrome. J. Med. Genet. 51, 460–469 (2014).
Aparisi, M. J. et al. Targeted next generation sequencing for molecular diagnosis of Usher syndrome. Orphanet J. Rare Dis. 9, 168 (2014).
Eandi, C. M. et al. Targeted next generation sequencing in Italian patients with Usher syndrome: phenotype-genotype correlations. Sci. Rep. 7 (2017).
Kooshavar, D., Razipour, M., Movasat, M. & Keramatipour, M. Targeted next generation sequencing identified a novel mutation in MYO7A causing Usher syndrome type 1 in an Iranian consanguineous pedigree. Int. J. Pediatr. Otorhinolaryngol. 104, 10–13 (2018).
Neuhaus, C. et al. Next-generation sequencing reveals the mutational landscape of clinically diagnosed Usher syndrome: copy number variations, phenocopies, a predominant target for translational read-through, andPEX26mutated in Heimler syndrome. Mol. Genet. Genomic Med. 5, 531–552 (2017).
Jiang, L. et al. Comprehensive molecular diagnosis of 67 Chinese Usher syndrome probands: high rate of ethnicity specific mutations in Chinese USH patients. Orphanet J. Rare Dis. 10, 110 (2015).
Kumar, A., Fishman, G. & Torok, N. Vestibular and auditory function in Usher’s syndrome. Ann. Otol. Rhinol. Laryngol. 93, 600–608 (1984).
Möller, C. G. et al. Usher syndrome: an otoneurologic study. The Laryngoscope 99, 73–79 (1989).
Seeliger, M. et al. Comparative study of visual, auditory, and olfactory function in Usher syndrome. Graefes Arch. Clin. Exp. Ophthalmol. Albrecht Von Graefes Arch. Klin. Exp. Ophthalmol. 237, 301–307 (1999).
McCulloch, D. L. et al. ISCEV Standard for full-field clinical electroretinography (2015update). Doc. Ophthalmol. Adv. Ophthalmol. 130, 1–12 (2015).
Vaché, C. et al. Usher syndrome type 2 caused by activation of an USH2A pseudoexon: implications for diagnosis and therapy. Hum. Mutat. 33, 104–108 (2012).
Kumar, P., Henikoff, S. & Ng, P. C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Choi, Y. & Chan, A. P. PROVEAN web server: a tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 31, 2745–2747 (2015).
Salamov, A. A., Nishikawa, T. & Swindells, M. B. Assessing protein coding region integrity in cDNA sequencing projects. Bioinforma. Oxf. Engl. 14, 384–390 (1998).
Pedersen, A. G. & Nielsen, H. Neural network prediction of translation initiation sites in eukaryotes: perspectives for EST and genome analysis. Proc. Int. Conf. Intell. Syst. Mol. Biol. 5, 226–233 (1997).
Liu, H., Han, H., Li, J. & Wong, L. DNAFSMiner: a web-based software toolbox to recognize two types of functional sites in DNA sequences. Bioinforma. Oxf. Engl. 21, 671–673 (2005).
Desmet, F.-O. et al. Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 37, e67 (2009).
Yeo, G. & Burge, C. B. Maximum entropy modeling of short sequence motifs with applications to RNA splicing signals. J. Comput. Biol. J. Comput. Mol. Cell Biol. 11, 377–394 (2004).
Reese, M. G., Eeckman, F. H., Kulp, D. & Haussler, D. Improved splice site detection in Genie. J. Comput. Biol. J. Comput. Mol. Cell Biol. 4, 311–323 (1997).
Baux, D. et al. Combined genetic approaches yield a 48% diagnostic rate in a large cohort of French hearing-impaired patients. Sci. Rep. 7, (2017).
Liquori, A. et al. Whole USH2A Gene Sequencing Identifies Several New Deep Intronic Mutations. Hum. Mutat. 37, 184–193 (2016).
Khan, A. O. et al. A deep intronic CLRN1 (USH3A) founder mutation generates an aberrant exon and underlies severe Usher syndrome on the Arabian Peninsula. Sci. Rep. 7, 1411 (2017).
Aller, E. et al. Identification of large rearrangements of the PCDH15 gene by combined MLPA and a CGH: large duplications are responsible for Usher syndrome. Invest. Ophthalmol. Vis. Sci. 51, 5480–5485 (2010).
Aparisi, M. J. et al. Study of USH1 Splicing Variants through Minigenes and Transcript Analysis from Nasal Epithelial Cells. Plos One 8 (2013).
Bragg, L. M., Stone, G., Butler, M. K., Hugenholtz, P. & Tyson, G. W. Shining a Light on Dark Sequencing: Characterising Errors in Ion Torrent PGM Data. PLoS Comput. Biol. 9 (2013).
Song, L. et al. Comparison of error correction algorithms for Ion Torrent PGM data: application to hepatitis B virus. Sci. Rep. 7 (2017).
Loman, N. J. et al. Performance comparison of benchtop high-throughput sequencing platforms. Nat. Biotechnol. 30, 434–439 (2012).
Bonnet, C. et al. Complete exon sequencing of all known Usher syndrome genes greatly improves molecular diagnosis. Orphanet J. Rare Dis. 6, 21 (2011).
García-García, G. et al. Novel deletions involving the USH2A gene in patients with Usher syndrome and retinitis pigmentosa. Mol. Vis. 20, 1398–1410 (2014).
Roux, A.-F. et al. Four-year follow-up of diagnostic service in USH1 patients. Invest. Ophthalmol. Vis. Sci. 52, 4063–4071 (2011).
Besnard, T. et al. Non-USH2A mutations in USH2 patients. Hum. Mutat. 33, 504–510 (2012).
Kubota, D. et al. CEP250 mutations associated with mild cone-rod dystrophy and sensorineural hearing loss in a Japanese family. Ophthalmic Genet. 1–8, https://doi.org/10.1080/13816810.2018.1466338 (2018).
de Castro-Miró, M. et al. Novel Candidate Genes and a Wide Spectrum of Structural and Point Mutations Responsible for Inherited Retinal Dystrophies Revealed by Exome Sequencing. PLoS One 11 (2016).
Weil, D. et al. The autosomal recessive isolated deafness, DFNB2, and the Usher 1B syndrome are allelic defects of the myosin-VIIA gene. Nat. Genet. 16, 191–193 (1997).
Ouyang, X. M. et al. Mutations in the alternatively spliced exons of USH1C cause non-syndromic recessive deafness. Hum. Genet. 111, 26–30 (2002).
Ahmed, Z. M. et al. Nonsyndromic recessive deafness DFNB18 and Usher syndrome type IC are allelic mutations of USHIC. Hum. Genet. 110, 527–531 (2002).
Bork, J. M. et al. Usher syndrome 1D and nonsyndromic autosomal recessive deafness DFNB12 are caused by allelic mutations of the novel cadherin-like gene CDH23. Am. J. Hum. Genet. 68, 26–37 (2001).
Ahmed, Z. M. et al. PCDH15 is expressed in the neurosensory epithelium of the eye and ear and mutant alleles are responsible for both USH1F and DFNB23. Hum. Mol. Genet. 12, 3215–3223 (2003).
Maria Oonk, A. M. et al. Nonsyndromic hearing loss caused by USH1G mutations: widening the USH1G disease spectrum. Ear Hear. 36, 205–211 (2015).
Mburu, P. et al. Defects in whirlin, a PDZ domain molecule involved in stereocilia elongation, cause deafness in the whirler mouse and families with DFNB31. Nat. Genet. 34, 421–428 (2003).
Riazuddin, S. et al. Alterations of the CIB2 calcium- and integrin-binding protein cause Usher syndrome type 1J and nonsyndromic deafness DFNB48. Nat. Genet. 44, 1265–1271 (2012).
Le Quesne Stabej, P. et al. Comprehensive sequence analysis of nine Usher syndrome genes in the UK National Collaborative Usher Study. J. Med. Genet. 49, 27–36 (2012).
Besnard, T. et al. Experience of targeted Usher exome sequencing as a clinical test. Mol. Genet. Genomic Med. 2, 30–43 (2014).
Bonnet, C. et al. An innovative strategy for the molecular diagnosis of Usher syndrome identifies causal biallelic mutations in 93% of European patients. Eur. J. Hum. Genet. EJHG 24, 1730–1738 (2016).
Oishi, M. et al. Comprehensive molecular diagnosis of a large cohort of Japanese retinitis pigmentosa and Usher syndrome patients by next-generation sequencing. Invest. Ophthalmol. Vis. Sci. 55, 7369–7375 (2014).
Qu, L.-H., Jin, X., Xu, H.-W., Li, S.-Y. & Yin, Z.-Q. Detecting novel genetic mutations in Chinese Usher syndrome families using next-generation sequencing technology. Mol. Genet. Genomics MGG 290, 353–363 (2015).
Aller, E. et al. Identification of 14 novel mutations in the long isoform of USH2A in Spanish patients with Usher syndrome type II. J. Med. Genet. 43, e55 (2006).
Jaijo, T. et al. Microarray-based mutation analysis of 183 Spanish families with Usher syndrome. Invest. Ophthalmol. Vis. Sci. 51, 1311–1317 (2010).
Oshima, A. et al. Mutation profile of the CDH23 gene in 56 probands with Usher syndrome type I. Hum. Mutat. 29, E37–46 (2008).
Ben-Yosef, T. et al. A mutation of PCDH15 among Ashkenazi Jews with the type 1 Usher syndrome. N. Engl. J. Med. 348, 1664–1670 (2003).
Jaijo, T. et al. Mutation screening of the PCDH15 gene in Spanish patients with Usher syndrome type I. Mol. Vis. 18, 1719–1726 (2012).
Liu, X. Z. et al. A mutation (2314delG) in the Usher syndrome type IIA gene: high prevalence and phenotypic variation. Am. J. Hum. Genet. 64, 1221–1225 (1999).
Dreyer, B. et al. Identification of novel USH2A mutations: implications for the structure of USH2A protein. Eur. J. Hum. Genet. EJHG 8, 500–506 (2000).
Garcia-Garcia, G. et al. Mutational screening of the USH2A gene in Spanish USH patients reveals 23 novel pathogenic mutations. Orphanet J. Rare Dis. 6, 65 (2011).
Jaijo, T. et al. MYO7A mutation screening in Usher syndrome type I patients from diverse origins. J. Med. Genet. 44, e71 (2007).
Bharadwaj, A. K., Kasztejna, J. P., Huq, S., Berson, E. L. & Dryja, T. P. Evaluation of the myosin VIIA gene and visual function in patients with Usher syndrome type I. Exp. Eye Res. 71, 173–181 (2000).
Ahmed, Z. M. et al. Mutations of the protocadherin gene PCDH15 cause Usher syndrome type 1F. Am. J. Hum. Genet. 69, 25–34 (2001).
Bolz, H. et al. Mutation of CDH23, encoding a new member of the cadherin gene family, causes Usher syndrome type 1D. Nat. Genet. 27, 108–112 (2001).
Astuto, L. M. et al. CDH23 mutation and phenotype heterogeneity: a profile of 107 diverse families with Usher syndrome and nonsyndromic deafness. Am. J. Hum. Genet. 71, 262–275 (2002).
Adato, A. et al. Mutation profile of all 49 exons of the human myosin VIIA gene, and haplotype analysis, in Usher 1B families from diverse origins. Am. J. Hum. Genet. 61, 813–821 (1997).
Roux, A.-F. et al. Survey of the frequency of USH1 gene mutations in a cohort of Usher patients shows the importance of cadherin 23 and protocadherin 15 genes and establishes a detection rate of above 90%. J. Med. Genet. 43, 763–768 (2006).
Neveling, K. et al. Next-generation genetic testing for retinitis pigmentosa. Hum. Mutat. 33, 963–972 (2012).
Baux, D. et al. Enrichment of LOVD-USHbases with 152 USH2A genotypes defines an extensive mutational spectrum and highlights missense hotspots. Hum. Mutat. 35, 1179–1186 (2014).
Jacobson, S. G. et al. Disease boundaries in the retina of patients with Usher syndrome caused by MYO7A gene mutations. Invest. Ophthalmol. Vis. Sci. 50, 1886–1894 (2009).
Baux, D. et al. Molecular and in silico analyses of the full-length isoform of usherin identify new pathogenic alleles in Usher type II patients. Hum. Mutat. 28, 781–789 (2007).
Dreyer, B. et al. Spectrum of USH2A mutations in Scandinavian patients with Usher syndrome type II. Hum. Mutat. 29, 451 (2008).
von Brederlow, B. et al. Identification and in vitro expression of novel CDH23 mutations of patients with Usher syndrome type 1D. Hum. Mutat. 19, 268–273 (2002).
Bernal, S. et al. Clinical and genetic studies in Spanish patients with Usher syndrome type II: description of new mutations and evidence for a lack of genotype–phenotype correlation. Clin. Genet. 68, 204–214 (2005).
van Wijk, E. et al. Identification of 51 novel exons of the Usher syndrome type 2A (USH2A) gene that encode multiple conserved functional domains and that are mutated in patients with Usher syndrome type II. Am. J. Hum. Genet. 74, 738–744 (2004).
Pennings, R. J. E. et al. Evaluation of visual impairment in Usher syndrome 1b and Usher syndrome 2a. Acta Ophthalmol. Scand. 82, 131–139 (2004).
Nakanishi, H. et al. Identification of 11 novel mutations in USH2A among Japanese patients with Usher syndrome type 2. Clin. Genet. 76, 383–391 (2009).
Ouyang, X. M. et al. Characterization of Usher syndrome type I gene mutations in an Usher syndrome patient population. Hum. Genet. 116, 292–299 (2005).
Cuevas, J. M. et al. Identification of three novel mutations in the MYO7A gene. Hum. Mutat. 14, 181 (1999).
Acknowledgements
This work was supported by the Institute of Health Carlos III (ISCIII) and the European Development Regional Funds (grants PI13/00638, PI16/00425, PI16/00539), and by a grant from Fundación ONCE (grant 2015/0398). C.F.G. is a recipient of a fellowship from the ISCIII and the European Social Fund (IFI14/00021). G.G.G. is recipient of a senior postdoctoral contract from CIBERER. Ion Proton and PGM were acquired thanks to a grant (PIE13/00046) bestowed by the ISCIII.
Author information
Authors and Affiliations
Contributions
C.F.G., E.A. and J.M.M. conceived and designed the study. C.F.G., G.G.G., T.J., N.F., E.A. and J.M.M. performed the molecular experiments and analysed the data. M.F.B. and A.S.D.M. provided the RP1973 sample and carried out the clinical evaluation of the patient. C.F.G. drafted the manuscript. J.M.M. and C.A. obtained the funding and provided the samples. C.F.G., G.G.G., T.J., E.A. and J.M.M. reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fuster-García, C., García-García, G., Jaijo, T. et al. High-throughput sequencing for the molecular diagnosis of Usher syndrome reveals 42 novel mutations and consolidates CEP250 as Usher-like disease causative. Sci Rep 8, 17113 (2018). https://doi.org/10.1038/s41598-018-35085-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-35085-0
- Springer Nature Limited
Keywords
This article is cited by
-
Genetics, pathogenesis and therapeutic developments for Usher syndrome type 2
Human Genetics (2022)
-
The genetic and phenotypic landscapes of Usher syndrome: from disease mechanisms to a new classification
Human Genetics (2022)
-
Novel Usher syndrome pathogenic variants identified in cases with hearing and vision loss
BMC Medical Genetics (2019)