Abstract
Smokeless tobacco associated Gingivobuccal squamous cell carcinoma (GB-SCC) is a major public health problem but available oral cancer cell lines are mostly from smoking associated tongue SCC raising the need for pertinent GB-SCC cell line models. As part of the International Cancer Genome Consortium (ICGC) Project, 4 novel cell lines, namely, Indian Tata Memorial Centre Oral Cancer (ITOC) −01 to −04 were established and characterized with conventional methods, karyotyping, ultrastructure, in vivo tumourigenicity, Whole exome sequencing (WES) and RNA sequencing. These hyperploid cell lines form xenografts in mice and show metabolically active and necrotic areas on fluorodeoxyglucose-positron emission tomography (FDG-PET) imaging. WES of ITOC cell lines recapitulate the genomic tumor profile of ICGC GB-SCC database. We further identified smokeless tobacco associated genetic alterations (PCLO, FAT3 and SYNE2) and oncogenic PIK3CA mutation in GB-SCC cell lines. Transcriptome profiling identified deregulation of pathways commonly altered in cancer and down-regulation of arachidonic acid metabolism pathway, implying its possible role in GB-SCC. Clinical application of high throughput sequencing data depends on relevant cell line models to validate potential targets. Extensively characterized, these oral SCC cell lines are particularly suited for mechanistic studies and pre-clinical drug development for smokeless tobacco associated oral cancer.
Similar content being viewed by others
Introduction
Advanced stage oral cancer with poor clinical outcome is a major public health problem in south Asian and east Asian countries where smokeless tobacco use is prevalent1,2. Smokeless tobacco is consumed in various forms with areca nut, lime or betel leaves as quid, pan, masheri or gutka3,4. Most oral cancers are squamous cell carcinomas (SCC) and arise from the lip, tongue, gingivobuccal mucosa, retromolar trigone, floor of mouth or the hard palate. The form of tobacco use not only influences the site of cancer within the oral cavity but also its biology and mutational profile. In western countries where smoking is the predominant form of tobacco use, tongue cancer is the most common oral cancer while in south Asian and east Asian countries where tobacco chewing is more prevalent, Gingivobuccal SCC (GB-SCC) is the most common oral cancer. GB-SCC differs from the tongue SCC in its clinical behaviour5 as well as molecular6,7,8 and immunoproteome profile9, yet the treatment modalities are identical. Previous studies have revealed that as compared to tobacco smoking, the smokeless tobacco associated cancers have predominance of specific transversions or transitions in TP53 and the genome10,11,12.
Comprehensive molecular genetic characterization using whole genome, exome and transcriptome analysis has been reported in over 400 Head and Neck SCC (HNSCC) tumours. These include GB-SCC by our group as part of the International Cancer Genome Consortium (ICGC) India Project6 and The Cancer Genome Atlas (TCGA) project in USA for diverse HNSCC13. Large number of novel and recurrently mutated genes and deregulated pathways in HNSCC, with some being unique to GB-SCC were identified. Mechanistic studies on the role of an increasing number of genetic variants being identified in different cancers and their translational exploitation requires cell lines from same organ and histology with full molecular genetic characterization.
Primary cultures and immortalized cell lines are indispensible source for conducting translational cancer research. Tumor-derived cell lines are widely used in vitro models due to the ease of performing large scale experiments, genetic manipulation and reproducibility14. Cell lines maintained using appropriate conditions retain parent tumor features and phenotype, making them preferred tumor representative models in laboratory studies15,16. The dearth of commercial cell lines for tumours which are uncommon in large parts of the world necessitates the need to establish and characterize novel cell lines from primary tumor17.
Of the several hundred HNSCC cell lines reported worldwide, very few are from GB-SCC in patients with smokeless tobacco use18,19,20 and none from the Indian subcontinent. As part of our systematic efforts to establish relevant cell lines in the ICGC India oral cancer project, we have established and performed comprehensive molecular genetic characterization of 4 oral squamous cell carcinoma (OSCC) cell lines from tumour tissues obtained from tobacco chewing Indian patients. These cell lines are named ITOC-01, 02, 03 and 04, after the ethnic origin, institute and cancer site - Indian Tata Memorial Centre Oral Cancer (ITOC). These cell line models established from smokeless tobacco consuming patients provide a platform to carry out wide spectrum of assays spanning from functional genomics to drug response studies. To the best of our knowledge this is the first study from Indian subcontinent characterizing smokeless tobacco GB-SCC derived cell lines.
Results
Cell line establishment and characterization
Details of the patients from whom these cell lines were derived are provided in Table 1. All the 4 cell lines are stably self immortalized and have under gone at least 70 passages. They have been maintained for over 4 years and cryopreserved at regular intervals.
Morphology and ultrastructure analysis
Classical epithelial morphology and characteristic SCC features were seen in all four parent tumors (Fig. 1A–D). Detailed histological evaluation of OSCC tumors is tabulated in Supplementary Table S1. ITOC cells were mononuclear and appeared flat and polygonal in shape. ITOC-01 cells adhered to each other forming clumps while ITOC-02, ITOC-03 and ITOC-04 cells grew in a monolayer (Fig. 1E–H). ITOC cells morphology is indicative of squamous epithelial cells, hence suggesting the squamous origin of the parent tumor being retained in all the 4 OSCC cell lines. Transmission electron microscopy (TEM) revealed studded desmosomes on the adjacent cell membranes, characteristic of epithelial morphology, condensed nuclear chromatin, irregular nuclear envelope and other cytoplasmic organelles (Fig. 1I–L). Bundles of intermediate filament in cytoplasm and microvilli on the cell surface were seen. Membranous staining for epithelial marker cytokeratin 8 confirmed epithelial origin of all 4 cell lines (Fig. 2A–D).
Morphology of OSCC cells in culture. (A–D) Histological sections of the primary oral tumors showing squamous cell carcinoma. (E–H) Phase contrast photomicrographs of cells in culture at 10X magnification. (I–L) TEM images of all the four cell lines exhibiting desmosomes (white arrow) and intracellular spaces (black arrow).
DNA profiles
Genotyping confirmed the human origin and unique genotype of each cell line (Supplementary Table S2). Comparison with Children’s Oncology Group (COG) and American Type Tissue Culture (ATCC) short tandem repeat (STR) databases showed no significant relatedness with any of the established cell lines affirming unique origin of each cell line. Apparent gender mismatch was observed in ITOC-04 cell line established from a male patient. The Y allele was not detected in the Amelogenin marker in the cell line but both X and Y alleles were intact in the normal and tumor tissue of this patient. Loss of Y allele during cell passaging is a known phenomenon21.
Chromosome analyses
Chromosomal analyses of ITOC-01, ITOC-02, ITOC-03 and ITOC-04 cell lines showed complex near triploid, hypertriploid, hypotetraploid, and near tetraploid karyotype respectively. Human male karyotype was observed in ITOC-01 and -03 cell line whilst ITOC-02 cell line showed human female karyotype. Loss of Y allele was observed in ITOC-04 cell line as seen in STR marker analysis. Aneuploidy was observed in all cell lines wherein the chromosome number ranged between 63–93 chromosomes (Fig. 3A–D).
Chromosomal analysis of OSCC cell lines. (A) Composite karyotype of ITOC-01 cell line: 63 ∼ 70,XY, +der(1)(p36), +der(2)t(2;6)(p24;q13), +der(3)del(3)(p10)x2,4, +5, +6,dup(7)(q11.2q22), +der(7)t(4;7)(q21;p22),del(7)(q32), +9, +der(9)t(9;?13)(p23;?q10), +10, +10, +der(11)t(4;11)(pterq26;p15),add(12)(p13), +13, +14, +14,der(14)t(13;14)(q10;q10), +15, +der(15)add(15)(p11.5), +16, +17,der(18)dup(18)(q21.1;q23), +20, +20,der(21)t(21;?)(q21;?),add(21)(q21)[cp20]. (B) Composite karyotype of ITOC-02 cell line: 72 ∼ 82,XX, +1, +3, +5, +8, +8, +9, +9,i(9)(q10;q10), +11, +11, +der(11)dup(11)(q13q23), +der(11)?t(11;?)(q;?),der(13)t(13;14)(q10;q10)x2, +der(13)t(13;?)(q10;?), +14, +14, +15, +der(15)t(14;15)(q10;q10)x3, +16, +16, +16, +17, +18, +18, +der(18)t(11;18)(q23;q23),19, +20, +20, +, +der(21)t(17;21)(q21;q22),der(21)t(21;21)(q10;q22)x3, +mar[cp20]. (C) Composite karyotype of ITOC-03 cell line: 80 ∼ 89,XY, +X, +1, +2, +3, +3, +4, +5, +der(6)t(6;?)(q10;?), +7, +7, +8, +9, +10, +11, +11, +12, +12, +der(13),t(13;14)(q10;q10)x2, +der(13)t(13;13)(q10;q10)x3, +14, +, +14, +der(14)add(14)(q11.2), +15, +15, +der(15)t(15;?)(q11.2;?)x2, +16, +16, +17, +17, +18, +19, +20, +20, +mar x3[cp20]. (D) Composite karyotype of ITOC-04 cell line: 84 ∼ 93,X,der(1)dup(1)(q22p33); +der(2)add(2)(p22p23), +der(3)t(3;?)(p21;?)x2, +4, +5, +5, +6, +7, +7, +8, +9, +9,i(9)(q10;q10), +der(10)t(10;?)(p10;?),der(11)dup(11)(q23)x3, +12, +12, +13, +15, +15,der(15)dup(15)(p10), +16, +17,18, +20, +20, +20, +21, +21, +21, +21, +der(21)t(21;21)(q10;q10), +22, +22, +mar1, +mar2x2, +mar3[cp20].
Growth characteristics and DNA content
Analysis of cell proliferation was carried out using growth curve. Population doubling time of ITOC-01, 02, 03 and 04 cell line was 47.31, 51.15, 67.82 and 47.75 hours respectively (Supplementary Fig. S1A–D). DNA histogram analysis of ITOC-01, ITOC-02, ITOC-03 revealed hyperploidy with DNA indices of 1.6, 1.8, 1.2, respectively, while ITOC-04 revealed tetraploidy, with DNA index of 2.0 (Supplementary Fig. S1E–H).
Transfection efficiency
Transfection efficiency of OSCC cell lines was evaluated by transfecting enhanced green fluorescent protein expression plasmid (pEGFP-C1) with GeneJuice transfection reagent. EGFP expression observed after 48 hours of transfection revealed low transfection efficiency in ITOC-01 (1%) and ITOC-02 (6%) but high efficiency in ITOC-03 (30%) and ITOC-04 cell lines (20%) (Supplementary Fig. S2).
Scratch assay and anchorage independent growth
At 24 hours, cell migration calculated using ImageJ software showed 24%, 37%, 51% and 100% wound healing for ITOC-01, ITOC-02, ITOC-03 and ITOC-04 cell lines respectively. One-way ANOVA (analysis of variance) and Bonferroni post-tests showed high significance between control and time points of 24 and 48 hours (Supplementary Fig. S3A). At 21 days, anchorage independent growth in ITOC-01 and ITOC-02 showed large colonies while ITOC-03 and ITOC-04 showed small colonies (Supplementary Fig. S3 B–E).
In vivo tumourigenicity and fluorodeoxyglucose - positron emission tomography (FDG-PET) imaging
ITOC-01 cell line developed subcutaneous tumor in the dorsal flank region in NUDE mice in 6 weeks, while ITOC-02, ITOC-03 and ITOC-04 cell line developed subcutaneous tumor in nonobese diabetic – severe combined immunodeficiency (NOD-SCID) mice in 3, 2 and 2 weeks respectively (Fig. 4A–D). Grossly, the xenograft tumors were firm, grey-white and partially necrotic with whitish thick exudate. Histological analysis was done on the sections taken from grey-white firm areas revealing characteristic squamous cell carcinoma features. Distant metastasis was not observed on FDG-PET scan (Fig. 4E–H) or on histology done after subcutaneous injection in any of the animals (Fig. 4I–L, Supplementary Table S1). Histopathological evaluation of primary and respective xenograft tumors revealed squamous cell carcinoma of varying grades. Detailed morphological analysis of the tumors confirmed characteristic features of the parent tumor being retained in the xenograft tumors (Supplementary Table S1).
Tumourigenicity of OSCC cell lines. (A–D) Subcutaneous tumor formation in nude (ITOC-01) and NOD SCID mice (ITOC-02, ITOC-03, ITOC-04) after inoculation of OSCC cell lines. (E–H) Micro PET images of 18-FDG uptake in coronal planes for cell line xenografted mice. Tumor is indicated with white broken line with areas of necrosis depicted with *. Blue arrow indicates the viable base tumor in PET scan. (I–L) (H&E) stained microphotographs from tumor xenografts captured at 10X magnification.
Exome analysis
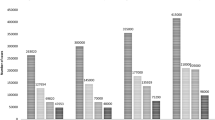
Whole exome sequencing (WES) was done at a mean depth of 155x, 112x, 128x and 113x for ITOC-01, -02, -03 and -04 cell line respectively with mean mapping quality of 58.12. A total of 24150, 21921, 22576 and 22018 exonic variants were identified in the ITOC-01, 02, 03 and 04 cell lines respectively, of which, 369, 265, 343 and 312 somatic variants were considered deleterious. Each cell line had a mutational profile matching with the GB-SCC or HNSCC mutational profile reported in the ICGC GB-SCC and TCGA HNSCC database (Supplementary Table S3, S4). Of the 15 most recurrently mutated genes in ≥10% of HNSCC or GB-SCC tumours as reported in the TCGA and ICGC database (TP53, FAT1, CASP8, NOTCH1, KMT2B, PCLO, UNC13C, SMG1, FAT3, EP300, KMT2D, SYNE2, TRPM3, PIK3CA and NSD1), 12 were mutated in one or more of the ITOC cell lines (Fig. 5A,B and Supplementary Table S5). All 4 cell lines had TP53 mutation while 3 cell lines had NOTCH1 deleterious mutations. The somatic variants mainly belonged to the apoptosis (TP53, BMP3, BMP4, TRAF5, AVEN) and inflammatory pathways (IKBKB, MMP10, CYP2A6, CYP4B1), implying their role in cancer progression22,23,24,25,26,27,28,29. Mutational signature in the 4 cell lines were similar with C > T transitions (44%) being most common, followed by T > C (20%), C > A (21%) and C > G (15%) transversions (Supplementary Fig. S4).
Mutational profile and validation of RNA-seq data. (A) Somatic nonsynonymous variants in genes recurrently mutated in ≥10% ICGC GB-SCC or TCGA HNSCC samples. (B) Somatic truncating and splice site mutations and missense variants predicted to be deleterious by ≥5 functional prediction tools. *Variants classified as deleterious by 4 prediction tools. (C–E) Validation of RNA-seq data by qPCR. 30 genes identified to be differentially expressed across OSCC cell lines were validated by Real time qRT-PCR. Comparison of gene expression values of RNA-seq and qPCR showed concordance in the data.
Comparative genomic analyses of parent tumor, cell lines at early passage (ITOC-01: P-10, ITOC-02: P-8, ITOC-03: P- 12, ITOC-04: P-9), cell lines at late passage (ITOC-01: P-51, ITOC-02: P-48, ITOC-03: P-58, ITOC-04: P-57) and xenograft tumors was performed by evaluating TP53 mutational status, the most frequently mutated gene in HNSCC and which is also found to be mutated in all 4 OSCC cell lines (Supplementary Fig. S5). We identified R196X nonsense mutation and Y126H missense mutation in ITOC-01, R306X nonsense mutation in ITOC-02, IVS2 + 38 C > G mutation in ITOC-03 and ITOC-04 and IVS3 + 36_52del16 and R342X in ITOC-04 parent tumor, cell line at early and late passage and xenograft tumors. However, in ITOC-02, the R306X mutation was seen in the cell line and primary tumour but not in the xenograft, possibly due to the heterogeneity in the xenograft.
Transcriptome analyses
RNA-seq of three normal buccal mucosa tissues and four OSCC cell lines generated ~65 million reads each. Compared to the 3 normal buccal mucosa tissues, 661, 811, 885, 907 genes were upregulated and 1627, 1997, 2343, 1950 genes down-regulated in ITOC-01, -02, -03 and -04 cell lines respectively (Supplementary Tables S6, S7). Gene Set Enrichment Analysis (GSEA) with Kyoto Encyclopedia of Genes and Genomes (KEGG) gene sets identified Notch signaling, p53 signaling, arachidonic acid metabolism (AAM), mitogen activated protein kinase (MAPK), chemokine, focal adhesion, extracellular matrix (ECM) receptor interaction, cell cycle, DNA replication pathway as significantly deregulated pathways in one or more of these 4 OSCC cell lines (Supplementary Fig. S6). Analysis limited to the 3 GB-SCC cell lines (ITOC-01, 03 and 04) identified similar deregulated pathways (Supplementary Fig. S7).
RNA-seq data was validated by quantitative polymerase chain reaction (qPCR) for 30 genes which are known to be deregulated in cancer and also in these cell lines (Fig. 5C). Gene expression of individual cell lines was normalized to normal buccal mucosa tissue using β-actin endogenous control. Up-regulation of keratins (KRT8,18) and down-regulation of Vimentin validates the epithelial origin of these cell line and also corroborates with the earlier result of expression of keratin 8 in OSCC cell lines (Fig. 2A–D). The AAM pathway genes which are mutated in GB-SCC (PLA2G3, PLA2G4E, PLA2G4F, PTGIS, TBXAS1, CYP2U1, CYP2C19, GPX7) were down-regulated in ITOC cell lines.
Discussion
Human cancer cell lines are indispensable to functionally characterize new or previously uncharacterized genetic alterations identified by large scale genomic studies. In the precision medicine era, the utility of a cell line for identifying clinically relevant targets requires these cell lines to closely recapitulate the molecular genetic profile of specific human cancers. Despite the common SCC histology, prior tobacco exposure and several common genetic alterations, the carcinomas arising at different primary sites within the Head and Neck region have several distinct features7,8,18. The Expert protein analysis system (ExPASy) – Cellosaurus database has reported over 150 SCC cell lines derived from tongue cancer but very few from buccal mucosa30,31,32 or gingiva33,34,35. Surprisingly, no GB-SCC cell lines are reported in literature or databases from the Indian subcontinent where it is the most common oral cancer. One possible reason may be the difficulty in culturing and establishing cell lines from heavily infected locally advanced gingivobuccal cancers.
We have successfully established and characterized 4 OSCC cell lines including 3 from GB-SCC derived from tobacco chewers. We demonstrate these cell lines to be of human origin with characteristic features of SCC. Identification of identical TP53 mutations across parent tumor, cell line at early and late passage and xenograft tumors affirms the genomic characteristics of primary tumor being preserved throughout the cell line cultivation and further in the xenograft model as well. The genomic and histopathological analyses assure the OSCC cell lines are representative models of the parent tumors. Through exome and transcriptome analysis we confirm that ITOC cell lines recapitulate the mutational and gene expression profile of GB-SCC and HNSCC in the ICGC and TCGA studies6,13. Of the 15 most recurrently mutated HNSCC or GB-SCC genes, 8 genes (PCLO, SYNE2, FAT3, EP300, KMT2B, SMG1, TRPM3 and UNC13C) were mutated in >10% of smokeless tobacco associated GB-SCC tumours of ICGC6 but not in the cigarette smoking associated HNSCC of TCGA study13 and may be considered as smokeless tobacco associated genes (Fig. 5A,B). Three of these 8 smokeless tobacco associated genes (PCLO, SYNE2 and FAT3) had deleterious mutation in GB-SCC ITOC cell lines (ITOC-01, 04) but not in the Tongue-SCC ITOC-02 cell line. This highlights the need for establishing, characterizing and using cell lines from tumours of the same subsite within HNSCC that have developed after similar form of tobacco exposure. Interestingly, the tongue SCC ITOC-02 cell line had truncating mutations in a few genes which are rarely associated with some cancers, PYROXD2, PCMTD1 reported to be mutated in hepatocellular cancer36,37 and PABCP1 reported to be upregulated in gastric carcinoma and targeting miR-34c38.
Moreover, ITOC-01 is the only GB-SCC cell line harbouring an oncogenic PIK3CA mutation (H1047R). It would be particularly useful to study PIK3CA inhibitors like Alpelisib (BYL719) which has been evaluated in only 2 PIK3CA mutated HNSCC cell lines39. Encouraging results from first in human use of Alpelesib in diverse tumours with oncogenic PIK3CA mutations has been reported40 and merits further evaluation in HNSCC including GB-SCC.
Earlier studies have proposed genetic signature 29 with abundance of C > A transversion for smokeless tobacco associated cancers10, however, this was not observed in any of the 4 ITOC cell lines. Similar to our finding, Fadlullah et al.20 had also not observed any correlation between the patient habits and mutation signatures. It is important to note that smokeless tobacco is used in many forms with or without lime, areca nut and other potentially carcinogenic ingredients. Hence there is a need to re-examine signature 29 and its association with smokeless tobacco and forms of smokeless tobacco use.
A common limitation of genetic profiling studies on cell lines is the absence of paired normal germline and gene expression data20,41. For the ITOC cell lines we used stringent bioinformatic methods to deduce somatic mutations. This seems to be a valid approach since the known mutational profile of HNSCC and GB-SCC is recapitulated in the ITOC cell lines. Since gene expression is tissue dependent, to interpret deregulated pathways in specific cancers, it is important to use representative normal tissues. In the TCGA HNSCC study with 16 GB-SCC tumours, their head neck region normal tissue dataset did not have any normal buccal mucosa sample for gene expression analysis. Similarly, the oral cancer cell line panel study used cultured keratinocytes in lieu of normal oral mucosa tissue20. Pure normal cell population in culture may be used as a convenient surrogate for normal tissues. However it is well known that isolation of specific cell populations, immortalization of normal or tumour cells, and the culturing conditions could all result in major alterations in gene expression as compared to their counterpart normal tissue or primary tumor42. This may result in misidentification of altered genes and pathways leading to false discovery. Possibly as a result of using normal buccal mucosa for determining differential gene expression, the deregulated genes in ITOC cell lines show excellent correlation with the qPCR data (Fig. 5C).
The ITOC cell lines show up-regulation of inducers of epithelial to mesenchymal transition (VEGFA)43, epigenetic markers (AURKA, AURKB, HDAC9); ECM receptor interaction molecules (laminin - LAMC2) and IL23A. A recent study has shown up-regulation of LAMC2 as a marker of cancer invasion in oral leukoplakia44. We propose a similar role could be observed in OSCC-GB. ITOC cell lines also show up-regulation of ECM receptor interaction and focal adhesion pathway, similar to previous large scale transcriptomic analysis of 326 oral cancer tissues45. Previous report by ICGC India team has shown loss-of-function somatic mutations in AAM pathway leading to better patient survival46. Considering the identification of somatic mutations and down-regulation of gene expression in AAM pathway components of these novel cell lines it would be interesting to further elucidate the role of AAM pathway in OSCC.
This is the first report of comprehensive molecular genetic characterization of 4 OSCC cell lines established from Indian patients. The ITOC cell lines recapitulate the molecular genetic profile of smokeless tobacco associated GB-SCC and tongue SCC as identified by the ICGC and TCGA studies. Moreover, ITOC-01 is the only GB-SCC cell line with oncogenic PIK3CA mutation. Tumorigenic potential along with the genomic characterization make the ITOC cell lines particularly useful for mechanistic and translational studies and screening novel agents in a disease with high recurrence rates despite best multimodality treatment.
Methods
Tumor specimen collection and establishment of cell lines
This study was approved by Tata Memorial Centre (TMC) – Advanced Centre for Treatment, Research & Education in Cancer (ACTREC) Institutional Ethics Committee III (Project No. 3). This study was conducted at TMC, ACTREC and all experiments involving human tissue were performed in accordance with relevant guidelines and regulations. Tumor tissues were collected at the time of surgery after obtaining informed consent from patients with smokeless tobacco associated oral SCC. Tumor tissues were transported in 5 mL Plain Iscove’s Modified Dulbecco’s Medium (IMDM) (Gibco) with double strength antibiotic mixture of Penicillin 400 U/mL, Streptomycin 40 µg/mL, Gentamycin 50 µg/mL and Mycostatin 10 U/mL (PSGM). Tissues were processed within half an hour of resection and were rinsed twice with sterile 1X phosphate buffered saline (PBS) followed by 10% Povidone-iodine (Wockhardt) and PBS wash. Tumor tissues were minced finely to small pieces of approximately 1–2 mm3 in size and incubated for 48 hours in 4 mL of complete IMDM [90% plain IMDM and 10% fetal bovine serum (FBS) (Gibco) supplemented with antibiotic] at 37 °C in a humidified atmosphere containing 5% CO2. Culture medium was replaced every 48 hours and the epithelial pool of cells was enriched by differential trypsinization and subsequent removal of fibroblasts. Cells at 80% confluency were split in the ratio of 1:2 twice a week. Four cell lines were established – 3 from GB-SCC (ITOC-01, 03 and 04) and 1 from tongue SCC (ITOC-02).
Phase contrast and electron microscopy
Morphological characteristics of the cell lines were studied using Olympus IX51 phase contrast microscope and ultra structure analysis was performed with Transmission Electron Microscope (JEM 1010, JEOL, Tokyo, Japan) at 120 kV (Supplementary Materials & Methods).
Chromosome preparation and giemsa banding
Cells were harvested and arrested with colcemide (0.05 µg/mL) for 1 hour at 37 °C. Mitotic cells were centrifuged at 2000 rpm for 5 minutes followed by hypotonic KCl treatment for 30 minutes at 37 °C. Cells were fixed in fresh methanol: glacial acetic acid (3:1, v/v) for 15 minutes at 4 °C. G-banding was performed by treating the slides with trypsin followed by Giemsa staining (GTG). Metaphases were captured using Olympus BX61 microscope (Olympus, USA) and analyzed using Applied Spectral Imaging (ASI) GenASIs software version 7.0 (ASI, Israel).
DNA profiling
Genomic DNA of cell lines was used for STR analysis of 8 autosomal markers (CSF1PO, D5S818, D7S820, D13S317, D16S539, THO1, TPOX, vWA) and 1 gender marker, Amelogenin. PCR products were analyzed with 3500 Genetic Analyzer (Applied Biosystems) and compared with COG STR database (http://strdb.cogcell.org) and ATCC STR database (http://atcc.org/STR_Database) of cell lines using match threshold of ≥80%.
Cell proliferation assay and doubling time
Cells were seeded at a density of 5 × 103 cells/well in 150 μL medium. 3-(4,5-dimethylthiazolyl-2)-2,5-diphenyltetrazolium bromide (MTT) solution was added at a final concentration of 0.5 mg/mL and incubated for 4 hours at 37 °C, 5% CO2. Purple formazan crystals were dissolved by addition of 10% SDS. Absorbance was measured at 540 nm with a spectrophotometer (BMG Labtech). Cell proliferation was assayed at different time intervals - 0, 24, 48, 72 and 96 hours. MTT readings were recorded in triplicates and the overall mean plotted against time. Standard error of the mean (SEM) were calculated for each day and the exponential growth curve plotted. Doubling time calculated using the equation ln(2)/K, where K is the growth rate constant.
DNA measurement
Lymphocytes isolated from fresh blood of a healthy donor were used as control for ploidy studies. Cells fixed in 70% ethanol for 60 minutes were incubated with 100 µg/mL RNase A (Sigma-Aldrich) and stained with 50 µL propidium iodide (Sigma-Aldrich) for 30 minutes at 37 °C. Acquisitions were performed on BD FACSCalibur (Becton Dickinson) and data analyzed using Modfit software (version 2.0).
Immunofluorescence staining
Cells were grown on sterile coverslips for 48 hours, fixed in ice-cold methanol at −20 °C for 10 minutes. Coverslips were rinsed with PBS and permeabilized using 0.3% Triton X and ice-cold methanol at room temperature (RT). Blocking was carried out using 3% bovine serum albumin (BSA) in PBS for 15 minutes at RT. Cells were incubated with primary antibodies for 1 hour at RT; p53 (Mouse monoclonal ab26, Abcam: 1:50), Cytokeratin 8 (Mouse monoclonal, Abcam, 1:100) followed by 1 hour incubation with fluorescein isothiocyanate (FITC) secondary antibody (Anti-Mouse IgG-FITC, Sigma), Alexa Fluor 568 goat anti-mouse IgG (H + L), Molecular Probes). Coverslips were mounted using Vectashield (Vector Labs) mounting medium with 4′,6-diamidino-2-phenylindole (DAPI) and viewed under confocal microscope (Carl Zeiss, LSM780).
Transfection efficiency assay
Cells seeded at a density of 3 × 105 in 35 mm cell culture dish were transfected with 1 µg of pEGFP-C1 plasmid using GeneJuice (Merck) transfection reagent as recommended by the manufacturer’s instructions. Cells were incubated at 37 °C for 48 hours and transfection efficiency was evaluated by counting 200 cells from five different fields under a fluorescent microscope.
Cell migration assay
Cells were seeded at a density of 2 × 106 in 6-well plate and on reaching 100% confluency a wound was inflicted with a pipette tip and images were captured every 24 hours on Olympus IX51 microscope and analyzed using ImageJ software. The relative cell free area at respective time points was calculated by considering the inflicted wound at 0 hours as 100%. One-way analysis of variance (ANOVA) and Bonferroni post-tests were performed to analyze effect of wound healing with respect to time.
Anchorage independent growth
1 × 105 cells/mL were suspended in 0.2% agar in IMDM with 10% FBS and overlaid onto 0.5% basal agar coat in 35 mm plates. The plates were incubated at 37 °C in a humidified atmosphere containing 5% CO2 for 21 days and colonies formed were visualized under an inverted microscope. Images were captured using Axiovert 200 inverted microscope (Carl Zeiss).
In vivo tumourigenicity and PET
For animal study approved by Institutional Animal Ethics Committee (Project no. 05/2012), 1 × 107 cells were injected subcutaneously in nude (ITOC-01) and NOD-SCID mice (ITOC-02, ITOC-03, ITOC-04). All experiments were performed in accordance with relevant guidelines and regulations of Institutional Animal Ethics Committee. The mice were housed in sterile pathogen-free environment and observed daily for appearance of tumors. FDG PET studies were performed for the xenografts established from these cell lines in nude and NOD-SCID mice (Supplementary Materials and Methods). Tumors from sacrificed mice were stained with hematoxylin and eosin (H&E) and images captured using Axio Imager.Z1 upright microscope (Carl Zeiss).
Whole exome sequencing
Genomic DNA libraries were prepared using Nextera® Rapid Capture Enrichment Kit (Illumina) as per the manufacturer’s instructions and paired-end reads were generated using Illumina HiSeq 1500 platform. Approximately 37 Mb region of human genome comprising of 214,405 exons was captured. Detailed analysis provided in Supplementary Materials and Methods.
TP53 mutational analysis
Genomic DNA was isolated from cells (Qiagen, 51104), fresh frozen tissue (PreAnalytiX, 767134) and formalin-fixed paraffin-embedded (FFPE) tissue (Qiagen, 56404) as per the manufacturer’s instructions. Primer sequences used for amplification of Exon 1–11 of TP53 gene were as described earlier47. Mutational analysis was carried out by bidirectional Sanger sequencing of PCR products on ABI 3730 capillary sequencer (Applied Biosystems).
RNA sequencing
Total RNA was isolated from the 4 tumor cell lines and 3 normal buccal mucosa tissues using miRNeasy mini kit (Qiagen). RNA libraries were prepared using TruSeq RNA sample preparation kit (Illumina) as per the manufacturer’s instructions and paired-end reads were generated using Illumina HiSeq 2500 platform. Please refer to Supplementary Materials & Methods for detailed RNA-seq analysis. RNA-seq data was validated by qPCR (Supplementary Materials & Methods). Primer sequences are specified in Supplementary Table S8.
Data Availability
Data is available on reasonable request according to data protection policy in the ethics agreement.
References
Warnakulasuriya, S. Global epidemiology of oral and oropharyngeal cancer. Oral Oncol. 45, 309–316 (2009).
Walvekar, R. et al. Squamous cell carcinoma of the gingivobuccal complex: Predictors of locoregional failure in stage III–IV cancers. Oral Oncol. 45, 135–140 (2009).
Jemal, A. et al. Global Cancer Statistics. CA Cancer J Clin. 61, 69–90 (2011).
Shield, K. et al. The Global Incidence of Lip, Oral Cavity, and Pharyngeal Cancers by Subsite in 2012. CA Cancer J Clin. 67, 51–64 (2017).
Nair, S. et al. Squamous cell carcinoma of tongue and buccal mucosa: clinico-pathologically different entities. Eur Arch OtoRhinoLaryngol. 273, 3921–8 (2016).
India Project Team of ICGC. Mutational landscape of gingivo-buccal oral squamous cell carcinoma reveals new recurrently-mutated genes and molecular subgroups. Nat Commun. 4, 2873 (2013).
Liao, C. T. et al. Tongue and buccal mucosa carcinoma: is there a difference in outcome? Ann Surg Oncol. 17, 2984–91 (2010).
Sathyan, K. M. et al. Carcinoma of tongue and the buccal mucosa represent different biological subentities of the oral carcinoma. J Cancer Res Clin Oncol. 132, 601–9 (2006).
Shukla, S. et al. Immunoproteomics reveals that cancer of the tongue and the gingivobuccal complex exhibit differential autoantibody response. Cancer Biomarkers. 5, 127–35 (2009).
Alexandrov, L. Understanding the origins of human cancer. Science. 350, 1175 (2015).
Pfeifer, G. & Hainaut, P. On the origin of G-> T transversions in lung cancer. Mutat Res. 526, 39–43 (2003).
Kannan, K. et al. Low incidence of p53 mutations in betel quid and tobacco chewing-associated oral squamous carcinoma from India. Int J Oncol. 15, 1133–6 (1999).
Cancer Genome Atlas Network. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature. 517, 576–582 (2015).
Van Staveren, W. C. G. et al. Human cancer cell lines: Experimental models for cancer cells in situ? For cancer stem cells? Biochim Biophys Acta Rev Cancer. 1795, 92–103 (2009).
Masters, J. R. Human cancer cell lines: fact and fantasy. Nat. Rev. Mol. Cell Biol. 1, 233 (2000).
Lohberger, B. et al. Liegl-Atzwanger, Establishment of a novel cellular model for myxofibrosarcoma heterogeneity. Sci. Rep. 7, 44700 (2017).
Miserocchi, G. et al. Management and potentialities of primary cancer cultures in preclinical and translational studies. J Transl Med. 15, 229 (2017).
Lin, C. et al. Head and neck squamous cell carcinoma cell lines: established models and rationale for selection. Head Neck. 29, 163–88 (2007).
Hamid, S. et al. Establishment and characterization of Asian oral cancer cell lines as in vitro models to study a disease prevalent in Asia. Int J Mol Med. 19, 453–460 (2007).
Fadlullah, M. et al. Genetically-defined novel oral squamous cell carcinoma cell lines for the development of molecular therapies. Oncotarget. 7, 27802 (2016).
Shah, K., McCormack, C. & Bradbury, N. Do you know the sex of your cells? Am J Physiol Cell Physiol. 306, C3–C18 (2014).
Yoshino, Y. & Ishioka, C. Inhibition of glycogen synthase kinase-3 beta induces apoptosis and mitotic catastrophe by disrupting centrosome regulation in cancer cells. Sci Rep. 5, 13249 (2015).
Beurel, E. & Jope, R. The paradoxical pro-and anti-apoptotic actions of GSK3 in the intrinsic and extrinsic apoptosis signaling pathways. Prog Neurobiol. 79, 173–89 (2006).
Fridman, J. & Lowe, S. Control of apoptosis by p53. Oncogene. 22, 9030–40 (2003).
Olsson, M. & Zhivotovsky, B. Caspases and cancer. Cell Death Differ. 18, 1441–9 (2011).
Singh, A. & Morris, R. The Yin and Yang of bone morphogenetic proteins in cancer. Cytokine Growth Factor Rev. 21, 299–313 (2010).
Eissmann, M. et al. Overexpression of the anti-apoptotic protein AVEN contributes to increased malignancy in hematopoietic neoplasms. Oncogene. 32, 2586–91 (2013).
Shao, Z., Zhang, W., Chen, X. & Shang, Z. Expression of EphA2 and VEGF in squamous cell carcinoma of the tongue: correlation with the angiogenesis and clinical outcome. Oral Oncol. 44, 1110–7 (2008).
Del Prete, A. et al. Molecular pathways in cancer-related inflammation. Biochem Med (Zagreb). 21, 264–75 (2011).
Lee, E. et al. Characterization of newly established oral cancer cell lines derived from six squamous cell carcinoma and two mucoepidermoid carcinoma cells. Exp Mol Med. 37, 379–90 (2005).
Fujisawa, A., Ikeda, H., Tobita, T. & Inokuchi, T. Establishment and characterization of human oral squamous cell carcinoma cell lines. Oral Med Pathol. 9, 133–139 (2004).
Kok, S. et al. Establishment and characterization of a tumorigenic cell line from areca quid and tobacco smoke-associated buccal carcinoma. Oral Oncol. 43, 639–47 (2007).
Wong, D., Chang, K., Chen, C. & Chang, R. Characterization of Two New Cell Lines Derived From Oral Cavity Human Squamous Cell Carcinomas-OC1 and OC2. J Oral Maxillofac Surg. 48, 385–390 (1990).
Zhao, M. et al. Assembly and initial characterization of a panel of 85 genomically validated cell lines from diverse head and neck tumor sites. Clin Cancer Res. 17, 7248–64 (2011).
Lee, G. et al. Characterization of novel cell lines established from three human oral squamous cell carcinomas. Int J Oncol. 20, 1151–9 (2002).
Liu, H. et al. Myeloid zinc finger 1 protein is a key transcription stimulating factor of PYROXD2 promoter. Oncol Rep. 38, 3245–3253 (2017).
Cleary, S. et al. Identification of driver genes in hepatocellular carcinoma by exome sequencing. Hepatology. 58, 1693–702 (2013).
Zhu, J., Ding, H., Wang, X. & Lu, Q. PABPC1 exerts carcinogenesis in gastric carcinoma by targeting miR-34c. Int J Clin Exp Pathol. 8, 3794–802 (2015).
Keam, B. et al. In vitro anticancer activity of PI3K alpha selective inhibitor BYL719 in head and neck cancer. Anticancer Res. 35, 175–82 (2015).
Juric, D. et al. Phosphatidylinositol 3-Kinase α–Selective Inhibition With Alpelisib (BYL719) in PIK3CA-Altered Solid Tumors: Results From the First-in-Human Study. J Clin Oncol. 36, 1291–1299 (2018).
Reinhold, W. et al. NCI-60 whole exome sequencing and pharmacological CellMiner analyses. PLoS One. 9, e101670 (2014).
Ertel, A., Verghese, A., Byers, S., Ochs, M. & Tozeren, A. Pathway-specific differences between tumor cell lines and normal and tumor tissue cells. Mol Cancer. 5, 55 (2006).
Kim, M. et al. VEGFA links self-renewal and metastasis by inducing Sox2 to repress miR-452, driving Slug. Oncogene. 36, 5199–5211 (2017).
Nguyen, C. et al. LAMC2 is a predictive marker for the malignant progression of leukoplakia. J Oral Pathol & Med. 46, 223–31 (2017).
He, Y. et al. Large scale transcriptomics analysis suggests over-expression of BGH3, MMP9 and PDIA3 in oral squamous cell carcinoma. PLoS One. 11, e0146530 (2016).
Biswas, N. K., Das, S., Maitra, A., Sarin, R. & Majumder, P. P. Somatic mutations in arachidonic acid metabolism pathway genes enhance oral cancer post-treatment disease-free survival. Nature Commun. 5, 5835 (2014).
Liu, Y. & Bodmer, W. F. Analysis of P53 mutations and their expression in 56 colorectal cancer cell lines. PNAS. 13, 976–981 (2006).
Acknowledgements
We thank Ms Vineeta Sawant and Ms Siddhi Redkar from Electron microscopy facility; Dr. Pradip Chaudhari and Mr Bhabani Mohanty from Small Animal Imaging facility; Mr Naresh Mahida and Ms Sharda Haralkar from Genomics facility, Dr. Dhanalaxmi Shetty from Department of Cancer Cytogenetics, Mr Nikhil Gadewal from Bioinformatics, Mr Madan Ludbe and Ms Nikita Sharma from Biorepository, Ms Pallavi Pandharipande and Ms Neha Gupta from Sarin lab at ACTREC for their active support in the conduct of various experiments and analysis. This work was supported by the grant from Department of Biotechnology, Govt. of India (Grant Number (BT/MB/01/ICGC/2009) under the International Cancer Genome Consortium project.
Author information
Authors and Affiliations
Contributions
Conception and design: R.S., K.A. and P.K. Development of methodology: K.P., S.K., P.D. and R.P. Analysis and interpretation of data: K.P., N.G. and P.G. Writing the manuscript: K.P. and R.S.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pansare, K., Gardi, N., Kamat, S. et al. Establishment and genomic characterization of gingivobuccal carcinoma cell lines with smokeless tobacco associated genetic alterations and oncogenic PIK3CA mutation. Sci Rep 9, 8272 (2019). https://doi.org/10.1038/s41598-019-44143-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44143-0
- Springer Nature Limited
This article is cited by
-
ARID2 suppression promotes tumor progression and upregulates cytokeratin 8, 18 and β-4 integrin expression in TP53-mutated tobacco-related oral cancer and has prognostic implications
Cancer Gene Therapy (2022)
-
Effect of Cold Atmospheric Plasma Jet and Gamma Radiation Treatments on Gingivobuccal Squamous Cell Carcinoma and Breast Adenocarcinoma Cells
Plasma Chemistry and Plasma Processing (2022)
-
Evidence of Association of CYP1A1 Expression in Blood Lymphocytes and Clinicopathological Variables in Oral Cancer
Indian Journal of Clinical Biochemistry (2022)
-
Funding for cancer research by an Indian funding agency, DBT
Journal of Biosciences (2021)