Abstract
The fungus, Fusarium oxysporum f. sp. cubense (Foc), is the causal agent of Fusarium wilt disease, which is the most serious disease affecting the whole banana industry. Although extensive studies have characterized many Foc-responsive genes in banana, the molecular mechanisms on microRNA level underlying both banana defense and Foc pathogenesis are not yet fully understood. In this study, we aimed to reveal the role of miRNA during banana-Foc TR4 interactions. Illumina sequencing was used to reveal the changes in small RNAome profiles in roots of Foc TR4-inoculated ‘Tianbaojiao’ banana (Musa acuminata cv. Tianbaojiao) in the early stages (i.e. 5 h, 10 h and 25 h post Foc TR4 inoculation, respectively). The expression of some differentially expressed (DE) miRNAs and their predicted target genes was studied by using quantitative real time PCR (qRT-PCR). Totally, 254 known miRNAs from 31 miRNA families and 28 novel miRNAs were identified. Differential expression analysis identified 84, 77 and 74 DE miRNAs at the three respective Foc TR4 infection time points compared with control healthy banana (CK). GO and KEGG analysis revealed that most of the predicted target genes of DE miRNAs (DET) were implicated in peroxisome, fatty acid metabolism, auxin-activated signaling pathway, sulfur metabolism, lignin metabolism and so on, and many known stress responsive genes were identified to be DETs. Moreover, expected inverse correlations were confirmed between some miRNA and their corresponding target genes by using qRT-PCR analysis. Our study revealed that miRNA play important regulatory roles during the banana-Foc TR4 interaction by regulating peroxidase, fatty acid metabolism, auxin signaling, sulfur metabolism, lignin metabolism related genes and many known stress responsive genes.
Similar content being viewed by others
Introduction
Fusarium wilt or Panama disease, the most destructive, notorious and still uncontrollable fungal disease of banana caused by soil-borne Fusarium oxysporum f. sp. cubense (Foc), has devastated and continues to threaten banana production worldwide1,2. Currently, Foc has been classified into 3 races according to their banana hosts, i.e., Foc race 1 (Foc 1), Foc race 2 (Foc 2) and Foc race 4 (Foc 4). Foc 4 can be further divided into Foc tropical race 4 (Foc TR4) and Foc subtropical 4 (Foc STR4). Among all the Foc biotypes, Foc TR4 was considered as the most predominant and destructive one for its much wider range of hosts (can invade Cavendish banana and all the cultivars that are sensitive to the other 3 Focs) and distributions (stronger high temperature tolerance) than the others3.
The only practical strategy for controlling Panama disease is by using resistant varieties4. Although sources of resistance to Foc 4 have been found in wild banana species5, it has not been possible to introgress this resistance into current commercial cultivars by conventional breeding mainly due to the sterile nature of most polyploid banana cultivars6. Genetic transformation in banana offers the opportunity to overcome this obstacle7,8. To date, however, limited banana Foc resistance related genes have been isolated9,10.
RNA Seq and digital gene expression (DGE) analysis, powerful methods for analyzing the plant transcriptome, have been used for investigating the transcriptional changes that occur during the response of banana to Foc TR4. For example, Li et al.11 compared the transcriptome changes of a resistant Cavendish banana mutant ‘Brazilian’ and its susceptible wild-type ‘Nongke No 1’ by using RNA-Seq and digital gene expression analysis, and found that the banana basal defense mechanism participate in the recognition of pathogen-associated molecular pattern (PAMP) and the defense-related transcripts contribute to the Foc TR4 resistance of the resistant Cavendish banana mutant. Wang et al.12 compared the transcriptome differences of ‘Brazilian’ banana at 4 time points post Foc TR4 infection, i.e. 0, 2, 4, and 6 days post inoculation (dpi) by using DGE analysis, and found that genes involved in the phenylalanine metabolism, phenylpropanoid biosynthesis and alpha-linolenic acid metabolism pathways showed significant expression changes, which indicated that these genes might be essential for the banana response to Foc TR4 infection. Li et al.13 compared the early stage transcriptome changes of ‘Brazilian’ banana after Foc 1 and Foc TR4 infection using DGE, and found that the global gene expression changes caused by the two Focs’ infection were very similar and also discovered the significant activation of ethylene biosynthesis and signaling related gene. Bai et al.14 investigated the gene expression profile changes of the high resistant banana cultivar ‘Yueyoukang 1’ and Foc TR4 susceptible ‘Brazilian’ at 0.5, 1, 3, 5 and 10 dpi with Foc 4, and faster and stronger defense response and as well as milder HR and senescence reactions were recorded in the high resistant cultivar. Deng15 compared the transcriptome changes of the high resistant ‘Kangku No. 5’ and susceptible ‘Tianbaogaojiao’ and found that plant defense related genes such as genes encoding some transcription factors, SA, JA and ethylene signaling related proteins, phenylalanine metabolism and cell wall lignification related proteins were more significantly induced in ‘Kangku No. 5’, indicating that these genes might contribute to its high resistance.
Small RNAs (sRNA) are a distinct class of regulatory non-coding RNAs in plants and animals16. The majority of them are short-interfering RNAs (siRNA) and microRNAs (miRNA) involving in gene silencing. Many plant miRNAs exert their critical regulatory roles through interacting specifically with their target mRNAs to suppress their activities17,18. Recently, accumulated evidences have suggested that sRNAs are involved in plant-pathogen interactions19,20,21. For banana, however, miRNAs have only been studied in relation to fruit ripening22, root salt stress response23, variety miRNAome difference24 and genome wide miRNA identification25,26,27,28, no research has been reported on the miRNAs involving in banana-Foc interaction. Therefore, in the present study, to reveal the role of miRNAs in banana in response to Foc TR4 infection, Illumina sequencing was applied to reveal the changes in sRNAome profiles in roots of Foc TR4-inoculated ‘Tianbaojiao’ banana (Musa acuminata cv. Tianbaojiao) in the early stage (i.e. 5 h, 10 h and 25 h post Foc TR4 inoculation, respectively). The results generated in this study would be very helpful for the understanding of the complex roles of banana miRNAs in response to Foc TR4 infection in the early stage and could provide basis for the exploration of the miRNA regulated Foc TR4 responsive genes.
Materials and Methods
Plant materials and pathogen preparation
The tissue cultured ‘Tianbaojiao’ banana plants used in this study were provided by Institute of Horticultural Biotechnology, Fujian Agriculture and Forestry University, Fuzhou, China. Tissue cultured seedlings were moved to nutrient soil (FAFARD® #1VC, Canada) after four weeks’ rooting and two-day’s hardening. Then, the plants were kept at 28 °C, 60~80% relative humidity, a photoperiod of 12 h (1500 ± 200 lx). After about 6 weeks, plants with five leaves and healthy root system were moved to the modified Hoagland solution for liquid culture for about 1 week.
The GFP-labeled Foc TR4 was provided by College of Plant Protection, Fujian Agriculture and Forestry University, Fuzhou, China. After seven-day’ incubation at 28 °C in dark on PDA medium, fresh Foc TR4 was washed out by using sterilized water. After filtering by using sterilized six-layer cheese cloth to remove the mycelium, the spore solution concentration was adjusted to 5 × 106 chlamydospores/ml for pathogen inoculation.
Determining the time-points for harvesting the samples and Foc TR4 inoculation
To determine the sample harvesting time-point, the GFP-labeled Foc TR4 progression in banana roots was monitored under a confocal microscope (Olympus, FV1200) according to Li et al.11. At about 5 hours post inoculation (hpi), many chlamydospores were attached to the banana root. At 10 hpi, after water rinsing, chlamydospores could also be observed in root samples and chlamydospores began to germinate and develop into hyphae. At 25 hpi, many fungus began to be detected in the vascular tissues. To analyze the sRNA expression changes caused by Foc TR4 infection, the four time points, i.e. 0 hpi, 5 hpi, 10 hpi and 25 hpi, were selected for RNA-Seq analysis of the root sRNAomes.
After the harvesting time determination, the liquid cultured banana seedlings were divided into four groups: group I (R25), group II (R10) and group III (R5H) was respectively inoculated with GFP-labeled Foc TR4 25 h, 10 h and 5 h before root harvesting, group IV banana plants that were not inoculated with the Foc TR4 and their roots were served as control (RCK). Three seedlings were used for each treatment and all the root samples were harvested at the same time and washed with sterilized water to remove the attached chlamydospores and hyphae. Then, root samples were precooled in liquid nitrogen and stored at −80 °C freezer for further use.
SRNA sequencing and sequence processing
Total RNA was extracted by using the QIAGEN RNeasy plant mini kit (QIAGEN, Valencia, CA), and then was treated with RNase free DNase I (Promega, Madison, Wisconsin, USA). RNA degradation and contamination, purity, concentration and integrity was monitored or measured by using 1% agarose gel electrophoresis, NanoPhotometer® spectrophotometer (IMPLEN, CA, USA), Qubit® RNA Assay Kit in Qubit® 2.0 Flurometer (Life Technologies, CA, USA) and the Agilent Bioanalyzer 2100 system (Agilent Technologies, CA, USA) respectively. To reduce the influence of individual background differences, high-quality root RNA of the 3 seedlings from each group were equiweightly mixed and were sent to Novogene Bioinformatics Technology Co. Ltd. for sRNA sequencing. For small RNA library construction, a total amount of 3 μg total RNA was used. By using NEBNext® Multiplex Small RNA Library Prep Set for Illumina® (NEB, USA.), sRNA sequencing libraries were constructed according to the manufacturer’s manual. After library quality assessment, the libraries were sequenced on the Illumina Hiseq2500 platform and 50 bp single-end reads were generated.
Small RNA annotation and miRNA identification
After removing the low quality reads containing ploy-N, with 5′ adapter contaminants, without 3′ adapter or the insert tag, containing ploy A or T or G or C and low quality reads from raw data, the clean data were obtained and were used for further analysis. The clean small RNA tags were firstly mapped to reference banana genome (http://banana-genome.cirad.fr/content/download-dh-pahang) by using Bowtie29 with no mismatch allowed to analyze their expression and distribution. The sRNA tags matched to protein-coding genes, repeat sequences, rRNA, tRNA, snRNA, and snoRNA, small RNA tags were then removed. The left genome mapped unique sRNA were used to search for known miRNAs by blasting against miRBase 21.0 database using modified software mirdeep2. To explore the occurrence of miRNA families, miFam.dat (http://www.mirbase.org/ftp.shtml) was used to look for miRNA families. The remaining unannotated sRNAs were then subjected to novel miRNA prediction using miREvo30 and mirdeep231 through exploring the secondary structure, the Dicer cleavage site and the minimum free energy22.
Identification of the Foc TR4 responsive miRNAs and target gene prediction
For the identification of the Foc responsive miRNA or differentially expressed miRNA (DE miRNA), the read number of each miRNA was firstly normalized to TPM (transcript per million). MiRNA with |log2 (fold change)| >1 and qvalue <0.01were identified as differentially expressed miRNA (DE miRNA). The target genes of each miRNA were predicted according to the method described by Bi et al.22 using psRobot32. GO and KEGG enrichment analysis of the predicted target genes were further performed for the elucidation of the way miRNA participate in the banana response to Foc TR4 infection according to Bi et al.22.
Quantitative real-time PCR analysis for miRNAs and their target genes
Total RNA of the three seedlings of each treatment was used for the reverse transcription reactions of the miRNAs and their corresponding target genes by respectively using TransScript miRNA First-Strand cDNA Synthesis SuperMix (Trans) and PrimeScript® RT reagent Kit (TaKaRa). Then, qRT-PCR was adopted to confirm the sequencing result of thirteen selected miRNAs. These thirteen selected miRNAs include 2 miRNAs (mac-miR166a and mac-miR395b-3p) identified as DE miRNA in none of the three comparison and eleven miRNAs (mac-nmiR11, mac-nmiR21, mac-miR528-5p, mac-miR171a-3p.2, mac-miR408-3p.3, mac-miR156a.1, mac-miR393a.1, mac-miR397b.2, mac-miR167.1, mac-miR398 and mac-miR5658) identified as DE miRNA in at least one comparison. Poly(A) Tailing Kit (Ambion) was used to extend the 3’-end of all sRNAs, and the following PAP real-time quantitative PCR (PAP qRT-PCR) was performed on Lightcycler® 480II (Roche, Switzerland) using the SYBR® Premix Ex TaqTM II (Perfect Real Time) kit33, and U6 was used as the internal control. PCR conditions were as follows: initial denaturation at 95 °C for 1 min; followed by 45 cycles of 95 °C for 20 s, 54~60 °C for 20 s, and 72 °C for 20 s. For melting curve analysis, the fluorescence intensity data were collected in the range of 65~95 °C with a rate of 0.5 °C per 10 s at the end of the run. Three biological and three technical replicates were used for analyzing each miRNA to minimize quantification errors. Relative expression analysis of these miRNA were calculated by using the 2−ΔΔCT method according to Bi et al.22. To check the expression pattern of target genes, the expression analysis of the target genes of selected miRNAs were also determined by qRT-PCR using GAPDH (Glyceraldehyde-3-phosphate dehydrogenase) and EIF5A-2 (Eukaryotic initiation factor 5A-2) as the endogenous controls according to Chen et al.34. Fifteen target genes, including the zeta-carotene desaturase (ZDS) gene (target of mac-miR166a), the putative disease resistance protein RPM1 (RPM1) gene (target of mac-miR395b-3p), the jasmonate ZIM-domain protein (JAZ) gene (target of mac-nmiR11), the protein cup-shaped cotyledon 2 (CUC2) gene (target of mac-nmiR21), the polyphenol oxidase (PPO) gene (target of mac-miR528-5p), the scarecrow like protein 27 (SCL27) gene (target of mac-miR171a-3p.2), the transcription factor TGA21 (TGA21) gene (target of mac-miR408-3p.3), the putative selenium-binding protein (SBP) and the superoxide dismutase [Cu-Zn] (SOD) gene (targets of mac-miR398), the putative squamosa promoter-binding-like protein 16 (SPB16) gene (target of mac-miR156a.1), the laccase-25 (LAC) gene (target of mac-miR397b.2), the putative regulatory protein NPR1 (NPR1) and the glutathione S-transferase zeta class (GST) (targets of mac-miR5658), the transport inhibitor response 1-like protein (TIR) gene (target of mac-miR393a.1), and the auxin response factor 12 (ARF12) gene (target of mac-miR167.1) were simultaneously analyzed for their expression levels using qRT-PCR. Each reaction (20 µl) included 10 µl of SYBR® Premix Ex Taq™ II (Tli RNaseH Plus) (Takara, Japan), respectively 0.8 µl of forward and reverse primer, 1 µl cDNA template and 7.4 µl nuclease-free water.
QRT-PCR conditions of the target genes were as follows: initial denaturation at 95 °C for 1 min; followed by 40 cycles of 95 °C for 20 s, 58 °C for 20 s, and 72 °C for 20 s. For melting curve analysis, the fluorescence intensity data were collected in the range of 65~95 °C with a rate of 0.5 °C per 10 s at the end of the run. All the primers used in this study were listed in Supplemental Data S1. Gene expression levels were calculated based on the normalization factors (NFs) produced by GeNorm according to Zhou et al.35.
Results
Small RNA profile changes revealed by high-throughput sequencing in Foc TR4 inoculated banana roots
To study the expression of banana sRNAs in response to Foc TR4 infection and to identify miRNAs involved in banana-Foc TR4 interaction, we inoculated the liquid cultivated banana seedlings with GFP-labeled Foc TR4 spore solution. According to the GFP detection result (Fig. 1), root samples treated by Foc TR4 for different time length (0 h, 5 h, 10 h and 25 h) were collected for RNA isolation and then for small RNA profiling. SRNA sequencing on Illumina Hiseq. 2500 produced 13,745,947, 13,694,126, 12,568,504 and 12,027,866 raw reads for RCK, R5H, R10 and R25 groups respectively (Table 1). About 90% of the total reads remained after removing the low quality reads and the unwanted reads below standards. There were 12,558,692, 11,934,390, 11,462,346 and 11,066,024 clean reads for RCK, R5H, R10 and R25 group, respectively. Then, the 18–30 nt small RNA sequences were selected out, and 6,063,504 (RCK), 5,335,626 (R5H), 5,377,388 (R10), 5,864,215 (R25) sRNA sequences were used for further analysis (Table 1). By blasting against the banana genome, 3,913,326 (64.54% of the sRNA in RCK), 3,422,908 (64.15% of the sRNA in R5H), 3,626,001 (67.43% of the sRNA in R10) and 3,208,726 (54.72% of the sRNA in R25) sequences of the 18–30 nt sRNAs were perfectly mapped to the genome (Fig. 2 and Table 2). The number of sRNA sequences identified as putative known miRNAs ranged from 38,680 to 67,596 and the number of sRNA sequences identified as novel miRNAs ranged from 1,551 to 2,740 (Fig. 2 and Table 2).
The observation results of GFP-labeled Foc TR4 on the lateral roots of ‘Tianbaojiao’ banana for RCK (A) R5H (B) R10 (C) and R25 (D). For the control, no GFP was observed. (A) Many chlamydospores were attached on the roots ‘Tianbaojiao’ banana at 5 hpi. (B) After water rinsing, few germinated chlamydospores were observed on the roots of ‘Tianbaojiao’ banana at 10 hpi (C) and more were observed at 25 hpi (D). (A,B,D) were taken under 20× and (C) was taken under 40× magnification on the Olympus confocal microscope (FV1200).
Category results of the genome mapped total sRNA (A–D) and unique sRNA (E–H) of the four banana root libraries. (A,E) The RCK library; (B,F) the R5H library; (C,G) the R10 library; (D,H) the R25 library. RCK, R5H, R10 and R25 represent the root samples treated with Foc TR4 for 0 h, 5 h, 10 h and 25 h, respectively.
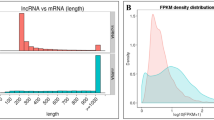
The sRNA length distribution results showed that more than 75% of the sRNAs were in the range of 18–25 nt and the 21 nt miRNAs take the largest part, followed by the 20 nt sRNAs (Fig. 3). This result was similar to the sRNAs of the 1-MCP treated banana fruit but differed a lot from the control and ethylene treated banana fruit22 (Bi et al. 2015), and also differed from the miRNA results from banana leaf 27, suggesting that the sRNA populations are different in different organs of the same plants and in the same organ under different treatments22. Our result was also different from many other plants such as Arabidopsis, rice, maize, tomato and citrus, for which the 24 nt sRNA were the most abundant. It can be concluded that the miRNA length distribution differed in different plants36,37,38,39.
Known and novel miRNAs identified in banana roots
For the identification of the known miRNAs, the genome mapped reads were searched against the mature miRNAs from miRBase database 21. Totally, 254 known miRNAs were identified (185, 212, 190 and 213 in RCK, R5H, R10 and R25 group, respectively), which belonged to 31 known miRNA families (Fig. 4 and Supplemental Data S2). MiR171 was the largest represented family containing 25 miRNA members, followed by miR159 family containing 24 miRNA members, and miR156, miR396 family also contained 20 miRNA members. The member numbers of miR160, miR166, miR167, miR169 and miR172 families were all more than 10. There were 9 members in miR395 family and 8 members in miR164. The member number of miR408, miR398, miR162, miR393, miR399, miR529, miR530, miR535, miR168, miR390, miR3630, miR444 and miR858 ranged from 2 to 6. While, other miRNA families had only 1 member.
The sRNA sequences that could be perfectly mapped to the banana genome but did not have a match in the miRBase were further subjected to novel miRNA prediction based on the plant miRNA annotation criteria17. In total, 28 novel miRNAs (22, 26, 27 and 23 for RCK, R5H, R10 and R25, respectively) were identified (Supplemental Data S3).
Identification of the differentially expressed miRNA (DE miRNAs)
After miRNA read number normalization, Pearson correlation analysis was performed to compare the miRNA expression correlation among the four sRNA libraries. Results showed that the R2 coefficient among all the four samples was all more than 0.8, indicating that no significant miRNA expression change was caused in the early Foc TR4 infection period. Notably, R2 coefficient between the Foc TR4 treated samples (R5H, R10 and R25) and the control (RCK) decreased with the increase of the Foc TR4 treatment time, implying that the influence of the pathogen on banana root miRNA expression became more and more obvious as treatment time went on (Supplemental Data S4).
Totally, 151 DE miRNAs, including 134 known miRNAs and 17 novel miRNAs, were identified by comparing the Foc TR4 treated groups (R5H, R10 and R25) with the control group (RCK) (Fig. 5). The DE miRNA number identified for R5H vs RCK, R10 vs RCK and R25 vs RCK was 84 (60 up-regulated, 24 down-regulated), 77 (42 up-regulated, 35 down-regulated) and 74 (69 up-regulated, 5 down-regulated) respectively. Seventeen DE miRNAs, including mac-miR156a.1, mac-miR167.1, mac-miR397a.1, mac-miR397a.2, mac-miR845, mac-miR164a.1, mac-miR397a.3, mac-miR397b-3p, mac-miR397.1, mac-nmiR13, mac-miR397.2, mac-miR160a-3p, mac-miR160d-3p, mac-miR164e, mac-miR397b.1, mac-miR397b.2 and mac-miR397.4, were identified in the three comparisons (Supplemental Data S5).
Cluster analysis and Venn diagram of the identified DE miRNAs in ‘Tianbaojiao’ banana roots. RCK, R5H, R10 and R25 represent the root samples treated with Foc TR4 for 0 h, 5 h, 10 h and 25 h, respectively. (A) Heatmap for the expression of the identified DE miRNAs in the four banana root libraries clustered by log10 (TPM + 1) value. The column represents different samples, and rows represent different DE miRNAs. Red color and blue color respectively means high and low expression. (B) Venn diagram of the DE miRNAs identified in the comparisons of R5H vs RCK, R10 vs RCK and R25 vs RCK.
Prediction and enrichment analysis of the DE miRNAs target genes
MiRNAs are functionally divergent, and the way they exert their critical regulatory roles is mainly through interacting specifically with their target mRNAs to suppress their activities17,18. So, for the function analysis of miRNA, the clarification of their target gene’s or genes’ function was very necessary. To further clarify the functions of the miRNAs, the putative target genes of these identified 254 known and 28 novel miRNAs were predicted, among which 255 miRNAs (250 known miRNAs and 5 novel miRNAs) were found to have predicted target genes. No target genes was identified for mac-miR167d.1, mac-miR8577, mac-miR1511-3p and mac-miR6478. For novel miRNAs, putative target genes only identified for mac-nmR3, mac-nmR5, mac-nmR8, mac-nmR21 and mac-nmR22.
Among the 151 DE miRNA, 1,012 putative target genes were identified for 137 DE miRNAs. The target gene number varied a lot for different DE miRNAs. The target of mac-miR5658 was found to be the most (130 predicted target genes including genes encoding MYB, RAP2, glucan endo-1,3-beta-glucosidase 3-like, aquaporins, syntaxins, WAT1-related proteins, regulatory protein NPR1-like, respiratory burst oxidase homolog protein H and so on, most of which were reported to be stress responsive), followed by mac-miR156h, mac-miR156a.1, mac-miR172c-3p, mac-miR156k, mac-miR172e-3p.1, whose target genes number was respectively 80, 79, 75, 64 and 62. The high abundance of predicted target genes suggests that these miRNAs may exert their roles through wide ways. While, only one target gene was identified for mac-miR3630-3p.1, mac-nmR3, mac-nmR8, mac-miR5538, mac-miR5141 and mac-miR3630-3p.2. Additionally, no predicted target genes was found for mac-miR8577 and some novel miRNAs (mac-nmR10, mac-nmR11, mac-nmR12, mac-nmR13, mac-nmR16, mac-nmR17, mac-nmR2, mac-nmR24, mac-nmR26, mac-nmR30, mac-nmR31, mac-nmR33 and mac-nmR9), which would make it difficult to understand their roles.
According to the target gene annotation result, the target genes of some miRNA families were found to be similar in different plant species, such as squamosa promoter-binding-like proteins (SPB) for miR156 and miR52940, ARF for miR160 and miR16741, nuclear TFY for miR16942, laccase for miR39743, MYB for miR858 and miR15922,44, scarerrow for miR17145, NAC for miR16446, AP2 for miR17247, EIN3 for miR39522. Moreover, IAA genes were found to be targeted by miR156, GDSL genes were predicted to be targets of miR164. PPO genes were predicted to be targeted by miR528. Laccase genes were also identified as targets of miR408 and miR397.
We also found that the number of the miRNA targeting the same gene also differed a lot (Supplemental Data S5), and the targeting miRNAs of the same gene are usually from the same miRNA family. For example, a transcription factor GAMYB-like gene (GSMUA_Achr10G29280_001) was predicted to be targets of 16 miR159s and 6 miR319s, which all belong to the miR159 family. A scarecrow-like protein 27 (GSMUA_Achr11G04360_001) was predicted to be targets of 22 miRNA members of miR171 family. The miRNAs of the same miRNA family targeting the same gene verified the function conservation of a miRNA family. The miRNAs targeting the same gene were even found to belong to different miRNA family. For example, the gene encoding scarecrow-like protein 15 (GSMUA_Achr4G07190_001) was predicted to be targeted by 21 miR171 family members and 5 miR408 members, the squamosa promoter-binding-like protein 12 gene (GSMUA_Achr10G02970_001) was predicted to be target of 15 miR156 family members, three miR529 family members and novel_22. Another squamosa promoter-binding-like protein 12 gene (GSMUA_Achr10G23280_001) was predicted target of 15 miR156 family members and novel_22. And a homeobox-leucine zipper protein HOX9 gene (GSMUA_Achr5G18530_001) was predicted to be target of 4 miRNAs of miR172 family and 13 members of miR166 miRNA family. The high diversity of the gene’s corresponding targeting miRNA suggested that the regulation of gene’s expression can be achieved by many miRNAs.
The predicted target genes of the DE miRNAs (DET) were then subjected to GO and KEGG analysis for their function. After GO enrichment analysis, most of the DETs were assigned to three GO ontologies, i.e. the biological process (BP), the cellular component (CC) and molecular function (MF) (Fig. 6).
GO enrichment analysis results of the predicted target genes of DE miRNAs identified in the comparisons of R5H vs RCK (A) R10 vs RCK (B) and R25 vs RCK. (C) RCK, R5H, R10 and R25 represent the root samples treated with Foc TR4 for 0 h, 5 h, 10 h and 25 h, respectively. BP: biological process; CC: cellular component; MF: molecular function.
From the aspect of BP, 53, 53 and 56 GO terms were significantly enriched in comparison of R5H vs RCK, R10 vs RCK and R25 vs RCK, respectively. Almost half of the enriched GO BP terms were biological process regulation related GOs such as ‘regulation of transcription’, ‘regulation of RNA biosynthetic process’, ‘regulation of RNA metabolic process’ and so on. Many other GOs such as ‘auxin-activated signaling pathway’, ‘cellular response to auxin stimulus’, ‘response to auxin’, ‘response to hormone’, ‘response to endogenous stimulus’, ‘phenylpropanoid catabolic process’, ‘lignin catabolic process’, ‘lignin metabolic process’ were also included. Notably, ‘auxin-activated signaling pathway’, ‘cellular response to auxin stimulus’ and ‘response to auxin’ GO terms were significantly enriched in all the three comparisons, and the ‘auxin-activated signaling pathway’ and the ‘cellular response to auxin stimulus’ term were even identified as the top two enriched GO terms in comparisons of R10 vs RCK and R25 vs RCK (Supplemental Data S6). The auxin signaling related genes included 17 auxin response factor genes (ARFs) targeted by members of miR156, miR160 and miR167. Moreover, ‘lignin catabolic process’ and ‘lignin metabolic process’ terms were also enriched in all the three comparisons. And the target genes enriched in the two GO terms were 4 laccase genes targeted by several miR408 and miR397 miRNA family members (Table 3). In addition, 4 polyphenol oxidase (PPO) genes targeted by mac-miR528-5p were found to be enriched in the ‘phenylpropanoid catabolic process’ (Table 3).
From the aspect of CC, 10 terms, including ‘nucleus’, ‘membrane-bounded organelle’, ‘intracellular membrane-bounded organelle’, ‘intracellular organelle’, ‘organelle’, ‘intracellular part’, ‘intracellular’, ‘cell’, ‘cell part’, ‘cellular component’ were enriched for all the three comparisons.
From the aspect of MF, 10, 12 and 9 terms were significantly enriched respectively for the three comparisons. Terms such as ‘DNA binding’, ‘nucleic acid binding’, ‘copper ion binding’, ‘hydroquinone: oxygen oxidoreductase activity’, ‘binding’, ‘GDP-L-fucose synthase activity’, ‘oxidoreductase activity, acting on diphenols and related substances as donors, oxygen as acceptor’, ‘DNA binding’, ‘nucleic acid binding’, ‘copper ion binding’ were enriched at least 2 comparisons. Terms of ‘nucleic acid binding transcription factor activity’, ‘transcription factor activity, sequence-specific DNA binding’ and ‘protein dimerization activity’ were only enriched in comparison of R5H vs RCK. Terms of ‘heterocyclic compound binding’, ‘organic cyclic compound binding’, ‘macromolecular complex binding’, ‘chromatin binding’, ‘sulfate adenylyltransferase (ATP) activity’ and ‘sulfate adenylyltransferase activity’ were only enriched in comparison of R10 vs RCK, and ‘oxidoreductase activity, acting on diphenols and related substances as donors’ and ‘catechol oxidase activity’ terms were found to be specifically enriched in comparison of R25 vs RCK.
KEGG pathway enrichment analysis showed that no significant pathway enrichment was identified, which may be due to the sample used being in the early Foc TR4 infection stage thereby causing no significant change. The top 20 pathways influenced by the Foc TR4 in the three early stages were shown in Fig. 7 and Supplemental Data S7. DETs involved in alanine, aspartate and glutamate metabolism, selenocompound metabolism, SNARE interactions in vesicular transport and some other 19 pathways were only enriched in one comparison. Cysteine and methionine metabolism, fatty acid metabolism, mRNA surveillance pathway, phenylalanine, tyrosine and tryptophan biosynthesis, protein export, purine metabolism, RNA transport pathways were found among the top 20 enriched pathways of two comparisons. Notably, pathways of peroxisome, fatty acid biosynthesis, fatty acid degradation, sulfur metabolism, nicotinate and nicotinamide metabolism, pyrimidine metabolism, pyruvate metabolism and metabolic pathways were found among the top 20 enriched pathways of all the three comparisons (Supplemental Datas S8 and S9), suggesting that the Foc TR4 greatly influenced these pathways in the early infection stages.
KEGG enrichment analysis results of the predicted target genes of DE miRNAs identified in the comparisons of R5H vs RCK (A) R10 vs RCK (B) and R25 vs RCK. (C) RCK, R5H, R10 and R25 represent the root samples treated with Foc TR4 for 0 h, 5 h, 10 h and 25 h, respectively. Only the top 20 enriched pathways of each comparison were shown.
Quantitative analysis for the expression of miRNAs and their target genes
The expression of 11 known miRNA and 2 novel miRNA identified in this study was analyzed using poly(A) miRNA-based qRT-PCR33 (Fig. 8). After Foc TR4 infection, although the expression of mac-miR166a was very significantly up-regulated at R5H and R10 (p-value < 0.01), their fold change were both less than 2 (Supplemental Data S2). The sequencing result also revealed it to be a not differentially expressed miRNA. The expression of mac-miR395b-3p was also slightly up-regulated at the early infection stages and a very significant change was found, although it was not identified as DE miRNA in the RNA-seq data (Supplemental Data S2). Mac-nmiR11 was identified as DE miRNA at R5H compared with RCK, and no expression was detected at R25 (Supplemental Data S2). Our qRT-PCR result showed that the miRNA’s relative expression was significant higher than RCK at R5H and R10H, and no significant difference were found between RCK and R25. Mac-nmiR21 was identified as a R25 specific novel miRNA in our sequencing result (Supplemental Data S2), and the highest relative expression of it was also found in R25 by using qRT-PCR. Mac-miR528-5p was identified as an up-regulated DE miRNA at R25 (Supplemental Data S2), and our qRT-PCR result also revealed that it was significantly up-regulated at R25. Moreover, its expression at R5H and R10 were also found to be very significantly higher than RCK (p-value < 0.01). Mac-miR171a-3p.2 was identified as a down-regulated DE miRNA at R5H and its highest expression was found at R10 in the RNA-Seq data (Supplemental Data S2). Our qRT-PCR result revealed that mac-miR171a-3p.2 expressed the highest at R10 but no significant expression change was found between R5H and RCK (p-value < 0.05). Mac-miR408-3p.3 was identified as an up-regulated DE miRNA at R5H and R25 in the RNA-Seq result (Supplemental Data S2). However, by using qRT-PCR, very significant up-regulation was only identified at R5H (p-value < 0.01) and its expression was found to be down-regulated at R25 compared with RCK. Mac-miR398 was identified as an up-regulated DE miRNA at R5H and R10 in the RNA-Seq result (Supplemental Data S2). Consistently, by using qRT-PCR, very significant expression change was also found at the same two time-point (p-value < 0.01). Mac-miR167.1 was an up-regulated at R5H and R25 but down-regulated at R10 compared with RCK according to our RNA-Seq results (Supplemental Data S2). The qRT-PCR result, however, revealed that this miRNA was significantly up-regulated at R5H and R10, but down-regulated at R25. Mac-miR156a.1 was identified as up-regulated DE miRNA at all the three Foc TR4 infected groups by RNA-seq (Supplemental Data S2). Its up-regulation at R5H was verified by qRT-PCR analysis, this miRNA was found to be significantly up-regulated at R5H but significantly down-regulated at R25 compared with RCK, and no significant expression difference was found between R10 and RCK. Mac-miR393a.1 was identified as up-regulated DE miRNA at R10 and R25 (Supplemental Data S2). Consistently, its up-regulation at the two time points was also verified by using qRT-PCR. Besides, its up-regulation was also found at R5H. Mac-miR397b.2 was identified as up-regulated DE miRNA at all the three Foc TR4 infected groups by RNA-seq (Supplemental Data S2). Although our qRT-PCR result showed it was down-regulated at R5H and its expression change at R10 compared with RCK was not significant, it was found to be up-regulated more than 4 fold at R25 compared with RCK. Mac-miR5658 was identified as a R5H specific novel miRNA in our sequencing result (Supplemental Data S2), and its expression at R5H was also found to be significant higher than RCK at R25 by using qRT-PCR. Overall, there were some consistencies between the sRNA high-throughput sequencing data and our RNA-Seq result. The differences might be caused by the calculation and principle differences of the two methods.
Quantitative real time PCR (qRT-PCR) result of the selected miRNAs and their corresponding target genes. Error bars represent the standard deviation. ZDS: Zeta-carotene desaturase; RPM1: disease resistance protein RPM1; JAZ: Jasmonate ZIM-domain protein; CUC2: protein CUP-SHAPED COTYLEDON 2; PPO: polyphenol oxidase; SCL27: scarecrow like protein 27; TGA21: transcription factor TGA21; SBP: selenium-binding protein; SOD: superoxide dismutase [Cu-Zn]; SPB16: putative squamosa promoter-binding-like protein 16; LAC: laccase-25; NPR1: putative regulatory protein NPR1; GST: glutathione S-transferase zeta class; TIR: transport inhibitor response 1-like protein; ARF12: auxin response factor 12. Different letters on the columns of the same color indicate significant difference at p-value < 0.05.
To investigate the expression patterns of the miRNAs’ target genes, the expression levels of one predicted target of these miRNAs were checked. Expected inverse relationship during the early Foc TR4 infection stage was identified between mac-nmiR21 and CUP, and mac-miR397b.2 and LAC. However, no contrasting expression pattern was found between the expression of the other miRNAs and their corresponding target genes, indicating that they may not target genes or these genes’ expression regulation were more complex22.
Discussion
Peroxisome and fatty acid metabolism were greatly influenced by Foc TR4 infection
Pathogen infection will induce the generation of reactive oxygen species (ROS)48. Peroxisomes are important producers of ROS and many antioxidants are located in them49. In this study, the target genes of several DE miRNA, such as mac-miR398, mac-miR398a-3p.1, mac-miR156a.1, mac-miR397.1 and so on, were involved in peroxisomes pathway. The pathway was found in the top 20 enriched pathways in all the three Foc TR4 infected groups compared with RCK, indicating that the Foc TR4 infection greatly influenced the peroxisome pathway.
Plant will induce rapid production of ROS in response to pathogen infection to inhibit their growth and invasion11,50. However, ROS accumulation will also cause damages to plant cells. Notably, all the DE miRNAs that own peroxisome related target genes were found to be up-regulated compared with RCK. Our qRT-PCR result also confirmed the up-regulation of miR398. Thus, it was deduced that the up-regulation of these miRNAs might be very necessary for decreasing the ROS damage to banana itself 50.
Peroxisomes accomplish a variety of biochemical functions, in which β-oxidation of fatty acids were included51,52. Fatty acids (FAs) are an important source of reserve energy and essential components of membrane lipids in all living organisms and are also biosynthetic precursors of cuticular components and phytohormone jasmonic acid (JA). Therefore, FA metabolic pathways play significant roles in pathogen defense53,54. Recently, FA has been identified as the major carbon source that transfer from host plant to symbiotic fungus55,56. And plant FAs were proved to be required for pathogen colonization56. In our present study, several DE miRNAs were predicted to target FA metabolism related genes such as biotin carboxylase 2, acetyl-CoA carboxylase 1 and two long chain acyl-CoA synthetase 9 genes. It was deduced that these miRNAs contribute a lot to the banana resistance by influencing the nutrient exchange between banana and Foc TR4.
Auxin pathway is an important target of Foc TR4 to facilitate its infection
The plant growth-defense tradeoffs regulation are dependent on the phytohormonal networks57. However, pathogens could disturb plant hormonal homeostasis to defeat plant immunity, modify babitate structure, enhance nutrient acquisition and facilitate infection and dissemination57.
Auxins are an important phytohormone for plant growth and disease resistance58,59,60,61. The reduction of auxin content was found to be significant for the enhancement of plant fungal pathogen resistance62,63. In Arabidopsis, F. oxysporum infection was reported to alter the auxin homeostasis in the root and promoted the pathogen colonization, and the auxin-signaling mutants were found to be more resistant to F. oxysporum64. Notably, the Foc 4 infection in banana could also increase the auxin content in banana65. All these suggested that pathogens could enhance the auxin biosynthesis of the plant to alter plant growth and development in their favor66. Conversely, in our study, many auxin signaling related auxin response factor genes (ARFs) were predicted to be targets of up-regulated DE miRNAs such as members of miR156, miR160 and miR167 families. This indicated that their expression was down-regulated. MiRNA-directed regulation of ARFs was proven to be essential for the root development and pivotal for modulating the expression of early auxin response genes67,68. Our qRT-PCR result also revealed that mac-miR167.1 was significantly up-regulated at R5H and R10. The up-regulation of their corresponding miRNAs suggested that Foc TR4 infection repressed the auxin response pathway. The suppression of the auxin response pathway could decrease the resistance of Arabidopsis to fungal pathogens60,69. The up-regulation of members of miR156, miR160 and miR167 families might function in banana-Foc TR4 interaction by repressing banana auxin response pathway.
Transport inhibitor response protein 1 (TIR1), the auxin receptor, is important for auxin signaling. In our present study, two TIR1 genes were predicted to be target genes of up-regulated miR393 members. And mac-miR393a.1, targeting a TIR1 gene (GSMUA_Achr5G26930_001), was identified to be significantly up-regulated in R5H, R10 and R25. MiR393 could enhance the plant disease resistance by negatively regulating genes for the F-box auxin receptors including TIR170. In cassava, higher miR393 expression was found in more anthracnose disease resistant cultivar, suggesting that this miRNA plays a role in fungal-infected cassava by influencing the auxin signaling71. The up-regulation of these miR393s in Foc TR4 infected banana root indicated that they might act in banana disease response by repressing auxin signaling.
Sulfate metabolism related miRNAs were Foc TR4 responsive
According to the KEGG pathway enrichment analysis, the sulfur metabolism were found to be enriched in all the three Foc TR4 infection stages, suggesting that sulfur metabolism were greatly influenced by the pathogen infection. Most of the DE miRNAs targeting sulfur metabolism related genes were up-regulated at least one stage, such as mac-miR171a.2, mac-miR171b.2 and mac-171c.1 targeting serine acetyltransferase 1 and mac-miR395t targeting ATP sulfurylase 1 genes. And our qRT-PCR results also verified the up-regulation of miR171 and miR395 member. The differential expression of these miRNAs might influence the sulfur metabolism in banana root. In plant, sulfur is an important component, and the sulfur nutritional status were directly related to plants’ health72. Moreover, sulfate metabolism was close related to the ethylene biosynthesis73, and Foc infection could significantly activate the ethylene biosynthesis and signaling related genes13. Additionally, accumulating evidences proved that its assimilation and metabolism are regulated by a variety of stresses74 and sufficient sulfate supply was found to be necessary for plant development due to sulfur induced resistance or sulfur enhanced defense75,76. Thus, it was concluded that miRNA-mediated sulfur metabolism changes might function in the Foc TR4 infection response of banana.
MiRNA participate in the banana wilt disease response by regulating lignin metabolism
Plant secondary metabolites, such as lignin, contributed greatly to an enhanced resistance to fungal diseases77. In Arabidopsis, miR397 and miR408 potentially regulate several laccase genes78. MiR397 could target the two laccase (lac) genes, LAC4 and LAC17, involved in lignification79,80. While the over-expression of miR408 in Arabidopsis changed the root gravitropic response81. In Populus trichocarpa, miR408 showed regulatory role in suppressing lignin deposition, and it was near basent in the major lignification tissue-the developing xylem82. MiR397 was also found to be the negative regulator of LAC genes affecting lignin content. In our present study, four lac genes were found to be targeted by the banana miR397 and miR408 family members. And our RNA-Seq result identified that the DE miRNAs targeting LAC genes were all significantly up-regulated by Foc TR4. Moreover, mac-miR397b.2 targeting LAC gene was found to be up-regulated more than 4 times at R25 compared with RCK according to our qRT-PCR result. It suggested that the lignin deposition in banana root was suppressed in response to Foc TR4 infection.
MiR528 is a monocot-specific miRNA. In this study, the Foc TR4 responsive mac-miR528-5p was predicted to target 4 PPO genes. PPO function a lot in cell wall lignification. In the study of Deng15, the cell wall lignification related genes were significantly induced by Foc TR4 in Fusarium wilt resistant ‘Kangku No. 5′, indicating that these genes might contribute to its high resistance. In maize, miR528 was predicted to target lac genes and its negative regulation role on lac expression and lignin biosynthesis were also identified83. The up-regulation of mac-miR528-5p in Foc TR4 infected banana root indicated that Foc TR4 infection might influence the banana cell wall lignification. The up-regulation of miR397, miR408 and miR528 in banana root suggested that banana altered its lignin metabolism in response to Foc TR4 infection.
Mac-miR5658 was a stress related miRNA
NPR1, one predicted target of miR5658, is one of the best-studied resistance-related genes84. Aquaporins, predicted target genes of mac-miR5658, are important for root development and their expression was regulated by auxin6,85, which again could provide evidence for the role of auxin in banana-Foc TR4 interaction. Coincidentally, the gene encoding WAT1, a vacuolar auxin transport facilitator required for auxin homeostasis86, was also predicted to be target of miR5658. Moreover, SNARE interactions in vesicular transport was identified to be enriched at R5H. The genes involved in this pathways were syntaxins that targeted by mac-miR5658. Syntaxins have been function in mediating the root growth49, and they also contribute to the plant disease resistance especially to the fungal diseases87,88,89,90,91,92. Mac-miR5658 owns so many stress responsive or pathogenesis related target genes, and its significant up-regulation revealed by both RNA-Seq and qRT-PCR indicated that it may contributed a lot during the banana-Foc TR4 interaction. The function of this miRNA need to be further studied for the clarification of its role in banana disease resistance.
MiRNA plays important roles in plant-fungal pathogen interaction by regulating the plant genes’ expression or by silencing the virulence genes of fungal pathogens93. In this study, we identified 254 known miRNAs from 48 miRNA families and 28 novel miRNAs from roots of Foc TR4-inoculated ‘Tianbaojiao’ banana in the early stage (i.e. 5 h, 10 h and 25 h post Foc TR4 inoculation, respectively), which could provide basis for the exploration of the banana disease response and the pathogenesis of Foc TR4. Differential expression analysis identified 84, 77 and 74 differentially expressed (DE) miRNAs at the three Foc TR4 infection times respectively compared with control healthy banana (CK). Function analysis of the DETs revealed the role of banana miRNAs in the complex defense response to Foc TR4 infection mainly by regulating the expression of peroxisome, fatty acid metabolism, auxin-activated signaling pathway, sulfur metabolism and so on, which will lead to the cell wall lignification repression, fungi growth inhibition, phytohormone signaling and plant defense activation (Fig. 9).
The role of miRNAs in the complex defense response to the Foc TR4 infection. Cell wall is the first barrier to the pathogen48, at the early stages of the Foc TR4 infection, several miRNAs including miR397, miR408 and miR528 were induced, which will lead to the down-regulation of LAC and PPO genes and ultimately the suppression of the cell wall lignification78,79,80,81,82. The Foc TR4 infection will induce the accumulation of ROS48, which will also cause damage to the plant cell11,50. The expression of banana miR156, miR397 and miR398 was induced, which will repress the expression of peroxisome related and FA metabolism related genes. The repression of peroxisome related genes will alleviate the harmful effect of ROS, while the FA metabolism change will influence the nutrient exchange between banana roots and Foc TR4 and the metabolism of JA51,52,53,54. The auxin content in banana could be enhanced by the Foc TR4 infection65, which might be helpful for the infection of pathogen60,69. To suppress the auxin signaling, miR156, miR160, miR167 and miR393 were up-regulated. The up-regulation of miR171 and miR395 will influence the sulfur metabolism and the ethylene biosynthesis72,73, which might contribute to the banana defense to Foc TR4. Mac-miR5658 was predicted to target many stress or pathogen responsive genes, indicating that it plays a role in regulation of banana-Foc TR4 interactions.
Data Availability
All the data generated or analyzed during this study are included in this published article and its Supplementary Information Files.
References
Ploetz, R. C. Fusarium wilt of banana. Phytopathology 105, 1512–1521 (2015).
Ordonez, N. et al. Worse comes to worst: bananas and panama disease-when plant and pathogen clones meet. PLos Pathog 11 (2015).
Garcia-Bastidas, F. et al. First report of Fusarium oxysporum f. sp cubense tropical race 4 associated with panama disease of banana outside Southeast Asia. Plant Dis 98, 694 (2014).
Ploetz, R. C. Panama disease: an old nemesis rears its ugly head: part 1. the beginnings of the banana export trades. Plant Health Progress 1–10 (2005).
Ploetz, R. C. & Pegg, K. G. Fusarium wilt diseases of banana, abaca and enset. (CABI Publishing, Wallingford, UK, 2000).
Reinhardt, H. et al. Tonoplast aquaporins facilitate lateral root emergence. Plant Physiol 170, 1640–1654 (2016).
Becker, D. K., Dugdale, B., Smith, M. K., Harding, R. M. & Dale, J. L. Genetic transformation of Cavendish banana (Musa Spp. Aaa Group) Cv ‘Grand Nain’ via microprojectile bombardment. Plant Cell Rep 19, 229–234 (2000).
Khanna, H., Becker, D., Kleidon, J. & Dale, J. Centrifugation assisted agrobacterium tumefaciens-mediated transformation (CAAT) of embryogenic cell suspensions of banana (Musa Spp. Cavendish Aaa and Lady Finger Aab). Mol. Breeding 14, 239–252 (2004).
Ghag, S. B., Shekhawat, U. K. S. & Ganapathi, T. R. Transgenic banana plants expressing a stellaria media defensin gene (Sm-Amp-D1) demonstrate improved resistance to Fusarium oxysporum. Plant Cell Tissue Org 119, 247–255 (2014).
Dale, J. et al. Transgenic Cavendish bananas with resistance to Fusarium wilt tropical race 4. Nat Commun 8, 1496 (2017).
Li, C. Y. et al. Transcriptome profiling of resistant and susceptible Cavendish banana roots following inoculation with Fusarium oxysporum f. sp. cubense tropical race 4. BMC Genomics 13, 374 (2012).
Wang, Z. et al. De Novo characterization of the banana root transcriptome and analysis of gene expression under Fusarium oxysporum f. sp. cubense tropical race 4 infection. BMC Genomics 13, 650 (2012).
Li, C. Q. et al. Analysis of banana transcriptome and global gene expression profiles in banana roots in response to infection by race 1 and tropical race 4 of Fusarium oxysporum f. sp. cubense. BMC Genomics 14, 851 (2013).
Bai, T. T. et al. Transcriptome and expression profile analysis of highly resistant and susceptible banana roots challenged with Fusarium oxysporum f. sp cubense tropical tace 4. PLos One 8, e73945 (2013).
Deng, G. M. Molecular mechanism research of Fusarium oxysporum f.sp cubense tropical race4 (FocTR4) early development and its interaction with disease sensitive and resistant bananas. Ph. D. Dissertation, South China Agricultural University, Guangzhou, 2014. (in Chinese) (2014).
Lakhotia, N. et al. Identification and characterization of miRNAome in root, stem, leaf and tuber developmental stages of potato (Solanum tuberosum L.) by high-throughput sequencing. BMC Plant Biol 14, 6 (2014).
Meyers, B. C. et al. Criteria for annotation of plant microRNAs. Plant Cell 20, 3186–3190 (2008).
Gu, M. et al. Identification of microRNAs in six solanaceous plants and their potential link with phosphate and mycorrhizal signaling. J. Integr Plant Biol 56, 1164–1178 (2014).
Zhang, W. X. et al. Bacteria-responsive microRNAs regulate plant innate immunity by modulating plant hormone networks. Plant Mol Biol 76, 205–206 (2011).
Zhao, H. W. et al. Small RNA profiling reveals phosphorus deficiency as a contributing factor in symptom expression for Citrus Huanglongbing disease. Mol Plant 6, 301–310 (2013).
Yang, L. & Huang, H. Roles of small RNAs in plant disease resistance. J Integr Plant Biol 56, 962–970 (2014).
Bi, F. C., Meng, X. C., Ma, C. & Yi, G. J. Identification of miRNAs involved in fruit ripening in Cavendish bananas by deep sequencing. BMC Genomics 16, 776 (2015).
Lee, W. S. et al. Transcripts and microRNAs responding to salt stress in Musa acuminata colla (AAA Group) cv. berangan roots. PLos One 10, e127526 (2015).
Ghag, S. B., Shekhawat, U. K. S. & Ganapathi, T. R. Small RNA profiling of two important cultivars of banana and overexpression of miRNA156 in transgenic banana plants. PLos One 10, e127179 (2015).
D’Hont, A. et al. The banana (Musa acuminata) genome and the evolution of monocotyledonous plants. Nature 488, 213–217 (2012).
Davey, M. W. et al. A draft Musa balbisiana genome sequence for molecular genetics in polyploid, inter- and intra-specific musa hybrids. BMC Genomics 14, 683 (2013).
Wen, J. Z. et al. A contig-based strategy for the genome-wide discovery of microRNAs without complete genome resources. PLos One. 9, e88179 (2014).
Chai, J. et al. Bioinformatic identification and expression analysis of banana microRNAs and their targets. PLos One. 10, e123083 (2015).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Bio 10, R25 (2009).
Wen, M., Shen, Y., Shi, S. H. & Tang, T. miREvo: an integrative microRNA evolutionary analysis platform for next-generation sequencing experiments. BMC Bio 13, 140 (2012).
Friedlaender, M. R., Mackowiak, S. D., Li, N., Chen, W. & Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res 40, 37–52 (2012).
Wu, H. J., Ma, Y. K., Chen, T., Wang, M. & Wang, X. J. PsRobot: a web-based plant small RNA meta-analysis toolbox. Nucleic Acids Res 40, 22–28 (2012).
Shi, R. & Chiang, V. L. Facile means for quantifying microRNA expression by real-time PCR. Biotechniques 39, 519–525 (2005).
Chen, L. et al. Validation of reference genes for RT-qPCR studies of gene expression in banana fruit under different experimental conditions. Planta 234, 377–390 (2011).
Zhou, Z. W. et al. Validation of reference genes for gene expression studies in post-harvest leaves of tea plant (Camellia sinensis). PeerJ 7, e6385 (2019).
Rajagopalan, R., Vaucheret, H., Trejo, J. & Bartel, D. P. A diverse and evolutionarily fluid set of microRNAs in Arabidopsis thaliana. Gene. Dev 20, 3407–3425 (2006).
Morin, R. D. et al. Comparative analysis of the small RNA transcriptomes of Pinus contorta and Oryza sativa. Genome Res 18, 571–584 (2008).
Yates, L. A., Norbury, C. J. & Gilbert, R. J. C. The long and short of microrna. Cell 153, 516–519 (2013).
Guo, Y. Q. et al. Identification of drought-responsive miRNAs and physiological characterization of tea plant (Camellia sinensis L.) under drought stress. BMC Plant Biol 17, 211 (2017).
Morea, E. G. O. et al. Functional and evolutionary analyses of the miR156 and miR529 families in land plants. BMC Plant Biol. 16, 40 (2016).
Xing, H. Y. et al. Genome-wide identification and expression profiling of auxin response factor (ARF) gene family in maize. BMC Genomics 12, 178 (2011).
Sorin, C. et al. A miR169 isoform regulates specific NF-YA targets and root architecture in Arabidopsis. New Phytol 202, 1197–1211 (2014).
Lu, S. F. et al. Ptr-miR397a is a negative regulator of laccase genes affecting lignin content in Populus trichocarpa. Proc Natl Acad Sci USA 110, 10848–10853 (2013).
Reyes, J. L. & Chua, N. H. ABA induction of miR159 controls transcript levels of two MYB factors during Arabidopsis seed germination. Plant J 49, 592–606 (2007).
Ma, Z. X. et al. Arabidopsis miR171-targeted scarecrow-like proteins bind to GT cis-elements and mediate gibberellin-regulated chlorophyll biosynthesis under light conditions. PLos Genet 10, e1004519 (2014).
Lee, M. H., Jeon, H. S., Kim, H. G. & Park, O. K. An Arabidopsis NAC transcription factor NAC4 promotes pathogen- induced cell death under negative regulation by microRNA164. New Phytol 214, 343–360 (2017).
Aukerman, M. J. & Sakai, H. Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 15, 2730–2741 (2003).
Cheng, C. Z., Zhang, Y. Y., Zhong, Y., Yang, J. W. & Yan, S. T. Gene expression changes in leaves of Citrus sinensis (L.) Osbeck infected by Citrus tristeza virus. J Hortic Sci Biotech 91, 466–475 (2016).
Ichikawa, M. et al. Syntaxin of plant proteins SYP123 and SYP132 mediate root hair tip growth in Arabidopsis thaliana. Plant Cell Physiol 55, 790–800 (2014).
Torres, M. A., Jones, J. & Dangl, J. L. Reactive oxygen species signaling in response to pathogens. Plant Physiol 141, 373–378 (2006).
Kubo, Y. Function of peroxisomes in plant-pathogen interactions. Subcell Biochem 69, 329–345 (2013).
Kessel-Vigelius, S. K. et al. An engineered plant peroxisome and its application in biotechnology. Plant Sci 210, 232–240 (2013).
Kachroo, A. & Kachroo, P. Fatty acid-derived signals in plant defense. Annu Rev Phytopathol 47, 153–176 (2009).
Cao, S. F., Yang, Z. F., Cai, Y. T. & Zheng, Y. H. Antioxidant enzymes and fatty acid composition as related to disease resistance in postharvest loquat fruit. Food Chem. 163, 92–96 (2014).
Luginbuehl, L. H. et al. Fatty acids in arbuscular mycorrhizal fungi are synthesized by the host plant. Science 356, 1175–1178 (2017).
Jiang, Y. N. et al. Plants transfer lipids to sustain colonization by mutualistic mycorrhizal and parasitic fungi. Science 356, 1172–1175 (2017).
Ma, K. W. & Ma, W. Phytohormone pathways as targets of pathogens to facilitate infection. Plant Mol Biol 91, 713–725 (2016).
Zhang, J., Peng, Y. L. & Guo, Z. J. Constitutive expression of pathogen-inducible OsWRKY31 enhances disease resistance and affects root growth and auxin response in transgenic rice plants. Cell Res 18, 508–521 (2008).
Mathesius, U. The role of auxin in root-symbiont and root-pathogen interactions: from development to defense. Progress in Botany 71, 185–210 (2010).
Fu, J. et al. Manipulating broad-spectrum disease resistance by suppressing pathogen-induced auxin accumulation in rice. Plant Physiol 155, 589–602 (2011).
Petti, C., Reiber, K., Ali, S. S., Berney, M. & Doohan, F. M. Auxin as a player in the biocontrol of Fusarium head blight disease of barley and its potential as a disease control agent. BMC Plant Biol 12, 224 (2012).
Domingo, C. F., Andres, F., Tharreau, D., Iglesias, D. J. & Talon, M. Constitutive expression of OsGH3.1 reduces auxin content and enhances defense response and resistance to a fungal pathogen in rice. Mol Plant Microbe Interact 22, 201–210 (2009).
Qi, L. L. et al. Arabidopsis thaliana plants differentially modulate auxin biosynthesis and transport during defense responses to the necrotrophic pathogen Alternaria brassicicola. New Phytol 195, 872–882 (2012).
Kidd, B. N. et al. Auxin signaling and transport promote susceptibility to the root-infecting fungal pathogen Fusarium oxysporum in Arabidopsis. Mol plant Microbe Interact 24, 733 (2011).
Tang, Q. F. et al. Endogenous hormone change in banana infected by Fusarium oxysporum f. sp. cubense. Journal of South China Agricultural University 27, 55–57 (in Chinese) (2006).
Ludwig-Müller, J. Auxin and the interaction between plants and microorganisms, Auxin and Its Role in Plant Development, pp. 413–434 (2014).
Mallory, A. C., Bartel, D. P. & Bartel, B. MicroRNA-directed regulation of Arabidopsis Auxin Response Factor17 is essential for proper development and modulates expression of early auxin response genes. Plant Cell 17, 1360–1375 (2005).
Wang, J. W. et al. Control of root cap formation by microRNA-targeted auxin response factors in Arabidopsis. Plant Cell 17, 2204–2216 (2005).
Llorente, F. et al. Repression of the auxin response pathway increases Arabidopsis susceptibility to necrotrophic fungi. Mol. Plant 1, 496–509 (2008).
Navarro, L. et al. A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science 312, 436–439 (2006).
Pinweha, N., Asvarak, T., Viboonjun, U. & Narangajavana, J. Involvement of miR160/miR393 and their targets in cassava responses to anthracnose disease. J Plant Physiol 174, 26–35 (2015).
Droux, M. Sulfur assimilation and the role of sulfur in plant metabolism: a survey. Photosynth Res 79, 331–348 (2004).
Iqbal, N. et al. Cross-talk between sulfur assimilation and ethylene signaling in plants. Plant Signaling & Behavior 8, e22478 (2013).
Lee, H. G. et al. Altered sulfate metabolism of Arabidopsis caused by beet severe curly top virus infection. The Plant Pathology Journal 21, 355–360 (2005).
Király, L. et al. Sulfate supply influences compartment specific glutathione metabolism and confers enhanced resistance to Tobacco mosaic virus during a hypersensitive response. Plant Physiol. Bioch 59, 44–54 (2012).
Bloem, E., Haneklaus, S. & Schnug, E. Milestones in plant sulfur research on sulfur-induced-resistance (SIR) in Europe. Front Plant Sci 5, 779 (2015).
Boubakri, H. et al. Thiamine modulates metabolism of the phenylpropanoid pathway leading to enhanced resistance to Plasmopara viticola in grapevine. BMC Plant Biol 13, 31 (2013).
Abdel-Ghany, S. E. & Pilon, M. MicroRNA-mediated systemic down-regulation of copper protein expression in response to low copper availability in Arabidopsis. J Biol Chem 283, 15932–15945 (2008).
Berthet, S. et al. Disruption of LACCASE4 and 17 results in tissue-specific alterations to lignification of Arabidopsis thaliana stems. Plant Cell 23, 1124–1137 (2011).
Wang, C. Y. et al. MiR397b regulates both lignin content and seed number in Arabidopsis via modulating a laccase involved in lignin biosynthesis. Plant Biotechnol J 12, 1132–1142 (2014).
Li, H. S. et al. Small RNA deep sequencing and the effects of microRNA408 on root gravitropic bending in Arabidopsis. Microgravity Sci Tec 27, 495–503 (2015).
Lu, S. F. et al. Novel and mechanical stress-responsive microRNAs in Populus trichocarpa that are absent from Arabidopsis. Plant Cell 17, 2186–2203 (2005).
Sun, Q. et al. MicroRNA528 affects lodging resistance of maize by regulating lignin biosynthesis under nitrogen-luxury conditions. Mol Plant 11, 806–814 (2018).
Sugano, S. et al. Role of OsNPR1 in rice defense program as revealed by genome-wide expression analysis. Plant Mol Biol 74, 549–562 (2010).
Péret, B. et al. Auxin regulates aquaporin function to facilitate lateral root emergence. Nat Cell Biol 14, 991 (2012).
Ranocha, P. et al. Arabidopsis WAT1 is a vacuolar auxin transport facilitator required for auxin homoeostasis. Nat Commun 4, 2625 (2013).
Collins, N. C. et al. SNARE-protein-mediated disease resistance at the plant cell wall. Nature 425, 973–977 (2003).
Assaad, F. F. et al. The PEN1 syntaxin defines a novel cellular compartment upon fungal attack and is required for the timely assembly of papillae. Mol Biol Cell 15, 5118–5129 (2004).
Monika, K., Nühse, T. S., Kim, F. & Peck, S. C. The syntaxin SYP132 contributes to plant resistance against bacteria and secretion of pathogenesis-related protein 1. Proc Natl Acad Sci USA 104, 11850–11855 (2007).
Lai, Z. B., Wang, F., Zheng, Z. Y., Fan, B. F. & Chen, Z. X. A critical role of autophagy in plant resistance to necrotrophic fungal pathogens. Plant J. 66, 953–968 (2011).
Pant, S. R. et al. Syntaxin 31 functions in Glycine max resistance to the plant parasitic nematode Heterodera glycines. Plant Mol Biol 85, 107–121 (2014).
Yun, H. S. & Kwon, C. Vesicle trafficking in plant immunity. Curr Opin Plant Biol 40, 34–42 (2017).
Cai, Q. et al. Plants send small RNAs in extracellular vesicles to fungal pathogen to silence virulence genes. Science 360, 1126–1129 (2018).
Acknowledgements
This work was supported by the Natural Science Funds for Distinguished Young Scholar of the Fujian Agriculture and Forestry University (xjq201721), the Postdoctoral Funds of the Fujian Agriculture and Forestry University (132300197), the Earmarked Fund for China Agriculture Research System (CARS-31-15), the Construction of Plateau Discipline of Fujian Province (102/71201801101), the Natural Science Foundation of China (31601713), and the Educational and Scientific Research Program for Young and Middle-aged Instructor of Fujian Province (JAT160166).
Author information
Authors and Affiliations
Contributions
C.C. and Z.L. designed the work; C.C., X.S., F.L., N.T. and R.M. performed the experiments and wrote the paper; C.C. analyzed the data; D.L., F.L. and N.T. helped to prepare the plant materials and pathogen. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cheng, C., Liu, F., Sun, X. et al. Identification of Fusarium oxysporum f. sp. cubense tropical race 4 (Foc TR4) responsive miRNAs in banana root. Sci Rep 9, 13682 (2019). https://doi.org/10.1038/s41598-019-50130-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-50130-2
- Springer Nature Limited
This article is cited by
-
Identification of self- and pathogen-targeted miRNAs from resistant and susceptible Theobroma cacao variety to black pod disease
Scientific Reports (2024)
-
Secreted in Xylem (SIX) genes in Fusarium oxysporum f.sp. cubense (Foc) unravels the potential biomarkers for early detection of Fusarium wilt disease
Archives of Microbiology (2024)
-
Contemporary narratives about asymmetries in responsibility in global agri-food value chains: the case of the Ecuadorian stakeholders in the banana value chain
Agriculture and Human Values (2023)
-
Fusarium wilt in banana: unraveling molecular aspects of host–pathogen interaction and resistance mechanism
Vegetos (2023)
-
MiRNA fine tuning for crop improvement: using advance computational models and biotechnological tools
Molecular Biology Reports (2022)