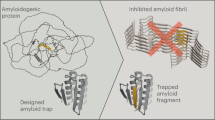
Abstract
Blocking the formation, growth, and breaking of amyloid fibrils by synthetic nanosystems could provide a treatment of neurodegenerative diseases. With this in mind, here atomistic molecular dynamics simulations are used to screen for nanoparticles (NPs), covered with different mixtures of ligands, including positively and negatively charged ligands, Aβ40-cut-peptide, and synthetic inhibitor ligands, in their selective coupling to Aβ40 peptides and their fibrils. The simulations reveal that only Aβ40-cut-peptide-covered NPs have strong and selective coupling to Aβ40 monomers. On the other hand, positive, positive-neutral, Janus, and peptide NPs couple to the beta sheet surfaces of Aβ40 fibrils and only the negative-neutral NPs couple to the fibril tips.
Similar content being viewed by others
Introduction
In recent years, NPs were used for bio-imaging, diagnostics, and delivery of drugs in biological systems1,2,3,4,5. NPs can be designed to selectively interact with and affect the functions of lipids6,7, proteins8,9, and various bio-molecular complexes. For example, NPs can have strong multivalent and selective interactions with viruses allowing them to act as broad spectrum virucidal drugs10.
In principle, NPs could be designed to specifically interact with amyloid fibrils, which are responsible for the development of Alzheimer, Parkinson and other neurological disorders. Some NPs were already shown to induce11,12,13 or prevent14,15 fibrillation of amyloid peptides, depending on their sizes, shapes, hydrophobicity and other parameters. It might be particularly interesting to prepare NPs that can affect the formation, growth, and breaking of amyloid fibrils structures16, in analogy to polymer-peptide conjugates that can break amyloid fibrils17.
Recently, molecular dynamics (MD) simulations were used to examine how selected NPs interact with amyloid peptides and affect their fibrillation by influencing the conformational change and the local concentration of free peptides12,18,19,20,21,22. Still, there is a lack of information how the NPs with different surface properties interact with free Aβ40 peptides and specifically with a fibril. In this work, we perform systematic atomistic MD simulations to screen NPs of different types selectively coupling with free Aβ40 peptides and their fibrils.
Results and Discussion
Figure 1(A) shows four ligands which are used to cover NPs in the present modeling: positive (Pos-lig; \({\text{NH}}_{3}^{+}\) terminal group), negative (Neg-lig; \({\text{SO}}_{3}^{-}\) terminal group), neutral (Neu-lig; NQTrp terminal group) and peptide ligands (Pep-lig). Except Pep-lig, all the ligands have two ethylene glycol groups (PEG) to reduce H-bonding between PEG oxygens and \({\text{NH}}_{3}^{+}\) terminal groups (Pos-lig)6. NQTrp is a quinone-tryptophan hybrid molecule, which is actually an inhibitor of an Aβ fibrillation process23. The amino acid sequence in Pep-lig, Cys-Glu-Leu-Val-Phe-Phe-Ala-Lys-Lys, follows the sequence, Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp, present in the exposed region of a β sheet of the Aβ40 fibril. The charged amino acids (Lys, Glu and Asp) of Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp are replaced with the oppositely charged amino acids (Glu, Lys, Lys) in Pep-lig, while keeping the remaining amino acids identical. The ligands are attached by thiol groups to the Au NP core (from Cysteine in Pep-lig).
Nanoparticle ligands screened in simulations. (A) (Pos-lig) PEG chain terminated with \({-\mathrm{NH}}_{3}^{+}\) group, (Neg-lig) PEG chain terminated with \({-\mathrm{SO}}_{3}^{-}\) group, (Pep-lig) peptide ligand, (Neu-lig) PEG chain terminated with NQTrp (quinone-tryptophan hybrid). Images of ligands created by VMD 1.9.3 software (https://www.ks.uiuc.edu/Research/vmd/). (B) The ligand composition in each nanoparticle.
Figure 1(B) describes five NPs prepared and simulated, each with a 2.2 nm diameter Au core and 90 attached and homogeneously (except Janus) distributed ligands. Since, Aβ40 is overall negative, three NPs are chosen to be positive (Pos, PosNQ and Pep), one is neutral (Janus) and one is negative (NegNQ). PosNQ and NegNQ NPs have 80 Pos-ligs and Neg-ligs, respectively, and 10 Neu-ligs, decreasing the NPs solubility in water. The number of Neu-lig ligands are optimized in such a way that they cannot be clustered due to van der Waals (vdW) coupling between their aromatic rings, since clustering of NQTrp prevents their coupling with the peptides5.
First, we simulated individual NPs together with (5) freely solvated Aβ40 peptides in a 150 mM NaCl solution box of (150 × 150 × 130 Å3) dimensions. Most NPs interacted rather weakly with the free peptides. However, after 100 ns of simulations, multiple peptides were found adsorbed on Pep NP (Fig. 2), matching well the peptides structure. To examine the NPs-peptides coupling, we calculated the number of heavy atoms of each peptide within 5 Å of different NPs and their interaction energies with these NPs. The highest number of contacts, 70–110, were observed among the five peptides coupled with Pep NP, individually presented in Fig. 2B (top). These peptides also had the largest coupling energies of −(40–80) kcal/mol (Fig. 2B (bottom)), mostly due to vdW coupling. These results are in line with the observation of peptide polymers capturing Aβ42 peptides24. The free Aβ40 peptides interacted with Pos and PosNQ NPs mainly by Coulombic coupling, while they interacted very little with Janus and NegNQ NPs (Figs. S1 and 2).
(A) A simulation snapshot of Pep NP with five Aβ40 peptides coupled to it, after 100 ns of equilibration. Positive, negative, polar and nonpolar amino acids of free peptides are shown by blue, red, green and white surface, respectively. In the case of Pep NP, terminal positively charged and hydrophobic amino acids are cyan and white, respectively. Images of molecular structures created by VMD 1.9.3 software (https://www.ks.uiuc.edu/Research/vmd/). (B) (top) Number of heavy atoms of the Aβ40 peptides within 5 Å of the Pep NP ligands. (bottom) Binding energy of each bound Aβ40 peptide with the Pep NP surface. The numbers in panel B are averaged over the last 5 ns of trajectories.
Next, we simulated the coupling of individual NPs to different regions of the Aβ40 fibril, consisting of two coupled protofilaments of a 13–14 nm length (29 peptides in each protofilament). Initially, each NP was placed close to one of the two exposed β sheet regions within the fibril. At the beginning of the simulation, the NPs were slowly diffusing on the surface, after 90–95 ns of simulations, the positively charged NPs (Pos, PosNQ, Pep) were strongly interacting with the β sheet surface, as shown in Fig. 3(A). For example, Pos NP was mostly nested on the negatively charged Glu22, Asp23 and neutral Val24 in the β sheet region, and often interacted with Asp1, Ala2 and Glu3 in the random chain regions of the fibril. All these interactions between Pos NP and the fibril gave a strong Coulombic coupling energy (on average of −290 kcal/mol), but a very small vdW decoupling energy (on average of 13 kcal/mol), as displayed in Fig. 3(B) for all NPs.
(A) NPs adsorbed on a Aβ40 fibril after 100 ns of simulations. Random chains regions of the fibril are not shown. In the case of fibril, positive, negative, polar and nonpolar amino acids are shown by blue, red green and white surface, respectively. For Pos, PosNQ, Janus and NegNQ, PEG chain, \({\text{NH}}_{3}^{+}\), \({\text{SO}}_{3}^{-}\) and NQTrp are shown by yellow, blue, red and green vdW representation, respectively. For Pep NP, terminal positively charged and hydrophobic amino acids are cyan and white, respectively. Pos, PosNQ, Pep and Janus NPs are coupled to the β sheet surface and NegNQ NP is coupled at the tip of the fibril. Images of molecular structures created by VMD 1.9.3 software (https://www.ks.uiuc.edu/Research/vmd/). (B) Coupling energies of NPs to the specified regions of the Aβ40 fibril, averaged over the last 5 ns of trajectories.
A rather different situation occured in PosNQ NP, having positively charged ammonium and neutral NQTrp terminal groups, differently interacting with the fibril (Fig. 3(A)). The Coulombic coupling energy (on average of −266 kcal/mol) was slightly smaller than in Pos NP, due to a smaller number of positively charged groups, but the vdW coupling energy, mostly due to NQTrp groups, was rather large (on average of −90 kcal/mol). Out of 10 NQTrp groups, 4 were close to the β sheet region and interacted with His14, Gln15, Lys16, Leu17, Val18, Phe20, Val24, Gly25, Ser26 and Asn27. Random chains of the Aβ40 fibril were also wrapping PosNQ NP by Asp1, Glu3, Arg5, Asp7, Ser8, Glu9, Tyr10, and Glu11.
Pep NP is more complex and directly designed to match the peptides, but this specific coupling is mostly active for free peptides (Fig. 2). It has an overall positive charge, which helped it to also nest on Glu22 of the fibril (Fig. 3(A)). The Coulombic (on average of −185 kcal/mol) and vdW (on average of −135 kcal/mol) coupling energies between Pep NP and the fibril were about the same (Fig. 3(B)), reflecting the matching structures of both coupled components. Random chains of the Aβ40 fibril were also relatively strongly interacting with the larger Pep NP by Asp1, Glu3, Phe4, Arg5, His6, Asp7, Ser8, Gly9, Tyr10, and Glu11.
Janus NP has asymmetrically distributed charges on its overall neutral surface. Therefore, its positive side mainly interacted with Phe20, Glu22, Asp23 and Val24 in the β sheet region, whereas its negative side interacted with His13, Lys16, Val18 and Phe20. Due to its overall neutrality, Janus NP has almost zero Coulombic coupling with the β sheet, since the attractive and repulsive coupling energies tend to cancel each other (special orientation). Nevertheless, the vdW coupling energy (on average of −46 kcal/mol) keeps Janus NP attached on the β sheet surface. Finally, when the negatively charged NegNQ NP was positioned close to the β sheet, it gained a strong repulsive Coulombic coupling energy (of ≈200 kcal/mol) and a weak attractive vdW coupling energy (of −60 kcal/mol) (Fig. S3), due to the presence of the NQTrp molecules, which prevented NegNQ NP from nesting on the fibril surface.
In short, Pos NP was initially nested on several Glu22 or Asp23 residues, but later it moved away to similar residues in different chains. Initially, the nature of binding of PosNQ, Janus and Pep NPs was quite similar like for Pos NP. For Janus, two types of ligands were interacting with oppositely charged residues, which increased their contact area. PosNQ and Pep NPs slowly extended their ligands to increase the number of contacts after their initial nesting.
In order to examine how the NP-coupling affected structural properties of the fibril, we calculated the average twist angle between two adjacent peptides in the absence/presence of NPs. In each peptide, we defined a vector going from the α-carbon atom of a 32nd amino acid residue to the 18th residue and measured the angle between two vectors of two neighboring peptides. 22 inner peptides out of 29 were used for calculation (see Methods). The simulation revealed that in the absence of NPs the average twist angle of the fibril is ≈6.48°. In the presence of NPs, the average twist angles of the interacting protofilament has increased to more than 7.5° (Pos: 7.7°, PosNQ: 7.93°, Pep: 8.08° and Janus: 7.62°) (Fig. 4). Figure 4 shows the distribution of twist angles in the last 5 ns, where the total number of counts in each plot is 10,500 (500 frames, each one contains 21 angles) and Fig. S5 shows average twist angles of each frame for all five systems. In all the cases, most of the twist angles are below 10° (fibril: 85.36%, Pos: 82.5%, PosNQ: 83.8%, Pep: 74.5%, Janus: 75.5%), but the presence of NPs extends the distribution tails. In Pos, PosNQ and Pep NPs, 9.3%, 8.7% and 8.6% of twist angles are ≥20°, respectively, whereas without NPs it is only 1.4%. The Janus NP has the largest percentage (23.33%) of twist angles of 10–20°. This enhancement of the twist angle was also observed when a D-enantiomeric amino acid peptide D3 (Arg-Pro-Arg-Thr-Arg-Leu-His-Thr-His-Arg-Asn-Arg) interacted with the Aβ fibril25.
We also evaluated the average fibril-NP contact area for NPs (Pos, PosNQ, Janus and Pep) with a favorable binding interaction to the fibril β-sheet to find out possible correlations with an induced twist of the protofilament. Figure S7 shows contact areas of NPs-fibril and NPs-fibril β-sheet (residue 12 to 40) over the last 5 ns. While the contact areas of Pos NP (full fibril: 6.83 nm2, fibril β-sheet: 4.43 nm2) are the smallest, the contact areas for NP-fibril β-sheet for PosNQ (fibril β-sheet: 13.59 nm2) and Janus (fibril β-sheet: 11.07 nm2) are comparable but 2.5–3 times larger than for Pos, due to the presence of two different types of ligands. Though the contact area of Pep NP with fibril β-sheet (fibril β-sheet: 10.01 nm2) is comparable to PosNQ and Janus, Pep NP contact area with full fibril (27.79 nm2) is the largest, as it has the longest ligands of all NPs studied, which make many contacts with the random coil regions of the fibril. The results show that NPs-fibril contact areas (Fig. S7) do not correlate with the induced twist angles in fibril β-sheet (Fig. 4). The twist angles induced by Pos, PosNQ, Janus and Pep are very similar, even though the Pos and Pep NP-fibril contact areas are significantly different than for the other two NPs.
We have also modeled coupling of NPs to the fibril tip, to find NPs that are potentially able to stop the fibril growth. From the five tested NPs, only NegNQ couple to the fibril tip, with an average Coulombic and vdW coupling energies of −108.6 and −35.1 kcal/mol, respectively (Fig. 3(A,B)). The positive (Pos, PosNQ, Pep) and Janus NPs experience a strong electrostatic repulsion at the tip (Fig. S4). Figure S6 shows selected equipotential surfaces of the Aβ40 fibril, where the light pink color (β sheet region) corresponds to −10.4 V and the light blue color (at the tip) corresponds to 2.6 V. This explains why positively charged NPs can be bound to the β sheet surface, whereas negatively charged NPs prefer to interact with the tip.
We used the same size of gold core (2.2 nm diameter) for all NPs, whereas the ligands have quite different lengths, especially in Pep NP. Free Aβ monomers can be easily adsorbed on the Pep NP ligands, due to their similar amino acid sequences, leading to significant vdW coupling. On the other hand, PosNQ and Pos NPs have a strong Coulombic coupling to free peptides (Fig. S2). Increasing their ligand length might increase the number of free peptides bound on PosNQ and Pos NPs at the cost of their reduced solubility, enhanced undesired protein interactions and smaller exposure of terminal charged groups. Addition of methylene groups (–CH2–) in the ligands can reduce their solubility and increase undesired protein interactions, whereas addition of ethylene glycol units can decrease the exposure of terminal \(N{H}_{3}^{+}\) groups. The exposed surface area of ligated PosNQ and Pos can be comparable to Pep NP by increasing NPs core size, which have more ligands and larger total surface charges, possibly causing larger toxicity26 in biological systems.
From the estimated size of the NP-fibril contact area, it is clear that the induced twist angle of the fibril β-sheet is not proportional to the contact area, which shows that functional group-specific Coulombic couplings are the primary controlling factor for changing the twist angle. In this context, NPs with smaller cores could be more effective for biomedical applications. By reducing their core size, the overall curvature of NPs would increase, which could cause more stretching of some of the ligands and increasing of the force with which they can act on the fibril surface through multivalent coupling10. This stronger coupling might induce larger twists of the fibril closer to the NPs.
Conclusion
In summary, the coupling of different NPs with the Aβ40 amyloid fibril was modeled by atomistic MD simulations. The results show that only Pep NP strongly couples with free peptides, and potentially affect their fibrillation. On the other hand, the positively charged NPs couple to the β sheet surface of the Aβ40 fibril, and locally deform it (helical twist), whereas the negatively charged NP couples to the tip of the fibril, and can potentially affect its growth. The inhibitor (NQTrp) molecules provide a number of heavy atoms coupling to amino acid residues, which increases the vdW coupling energy. The coupling of NPs to different polymorphs of the Aβ fibril might be different, due to different arrangements of exposed group sequences. These studies could help in designing nanomedicines for curing of neural diseases.
Methods
The initial structure of an Aβ40 fibril was prepared based on a protein structure (pdb ID 2LMO)27. In the coordinate file, first eight amino acid residues of each peptide were missing, which were added as disordered chains by the Modeller program28,29. Then, the fibril was extended up to 13–14 nm (29 peptides in each protofilament). The prepared fibril was simulated in a 150 mM NaCl solution for 5 ns. NPs were modeled with icosahedral gold shell core of a 2.2 nm diameter, where 90 ligands were homogeneously distributed. Masses of gold atoms were rescaled to mimic a solid gold core. Zero partial charges were assigned to the gold atoms bonded together with rigid bonds (b0 = 2.74 Å and large Kb). Ligands were attached to gold atoms via thiol groups. Angles, dihedrals in between ligands and gold cores were not considered. We placed the NPs at 5–10 Å above the fibril. The positively charged NPs were close to Glu22, Asp23 and the negatively charged NP was close to Lys16. In the case of Janus NP, the negative and positive surfaces were oriented towards Lys16/Glu22 and Asp23 respectively. Better selected initial positions of NPs reduce the exploration timescales for finding the NPs binding sites on the β-sheet surface of the fibril. All the simulated systems contained 400,000–600,000 atoms. After the initial minimization and warming to 300 K, ions and water molecules were equilibrated for 2 ns with restraining the movement of the NP core and the fibril backbone by a harmonic force constant of 1 kcal/mol Å2. Then, all the systems were simulated for 90–100 ns with releasing the NPs and the fibril. Visual Molecular Dynamics 1.9.3 (VMD) (https://www.ks.uiuc.edu/Research/vmd/)30 was used to visualize and create all the images of the ligands, NPs and the fibril.
The systems were simulated with a NAMD package31 using CHARMM general32,33,34 and protein forcefields35,36,37. In the simulations, we used a Langevin dynamics with a damping coefficient of γlang = 0.1 ps−1. Nonbonded vdW interactions were calculated with the Lennard-Jones (12,6) potential using a cut-off distance, d = 10 Å. Long range electrostatic interactions were calculated by the PME method38 in the presence of periodic boundary conditions and the MD integration time step was set to 2 fs. The simulations were performed using an NPT ensemble with P = 1 bar and T = 300 K. The simulations of five NPs at the tip of the fibril were also performed using similar conditions.
We performed the simulations of NPs binding on the fibril surface for 100 ns timescale, which is long enough for NPs to explore the fibril surface during the initial association process and bind to it if favourable. This timescale is too short to describe a full equilibration of the fibril without or with NPs bound to it. After simulating each system, we calculated the interaction energies (electrostatic and vdW interaction energy) of the NPs with fibril using the VMD energy analysis plugin considering the dielectric constant of water equals to 78.5. The average was performed over last 5 ns (500 frames). Despite this limited equilibration (100 ns of pre-equilibration), averaging of the data collected over the last 5 ns was specifically selected to describe the final state of the system reached in simulation to reveal the nature of binding of NPs to the fibril. We investigated the effect of NPs coupling to the β sheet surface of the fibril by calculating the average twist angle between two peptides. To calculate the twist angles, we first defined a vector from alpha carbon atoms of 32nd amino acid residue to 18th residue of each peptide and measured the angle between the two vectors of two consecutive peptides39. We calculated the average of the twist angles over the last 5 ns (500 frames) of the simulation trajectories. We calculated the twist angles from 4th to 25th monomer of each layer. In the calculations, the terminal peptides were not taken into account to remove the boundary effect of a fibril simulation in a water box. The calculation is performed using equation \({\theta }_{twist}=1/m{\sum }_{j=4}^{24}{\phi }_{j,j+1}\), where m is the number of peptide pairs, j and (j + 1) are the indices of two consecutive peptides and \({\phi }_{j,j+1}\) is a twist angle for a pair of peptides. Each distribution of twist angles was prepared using 10500 datapoints collected from 500 frames (each frame contains 21 twist angles).
We have separately calculated the contact area of NPs (Pos, PosNQ, Janus and Pep) with a full fibril and with a β-sheet of the fibril. To estimate the contact area, solvent accessible surface area (SASA) of NPs in the absence of fibril, in the presence of fibril and in the presence of fibril β-sheet were calculated using VMD SASA plugin. In this calculation, a radius of the solvent of 1.4 Å was considered. A sphere of the combined radius (solvent radius + vdWs radius of NP atom) was imaged surrounding each NP atom and 500 points were randomly distributed on the surface of the sphere. All the points were checked against the surface of all the neighboring atoms and the accessible number of points was multiplied with the assigned surface area that each point represents. In the case of NPs lacking the fibril, all the points at the sphere surface were checked against the surface of neighboring NP atoms, whereas in the case of NPs in the presence of fibril, the points were also checked against the fibril atom surface. Subtracting the SASA value of NPs in the presence of fibril and in the presence of fibril β-sheet from the NP SASA in the absence of fibril gave us the NP contact area with fibril and fibril β-sheet, respectively. In Fig. S7, we reported the average contact area of NPs with fibril and NPs with fibril β-sheet. The averaging was performed over last 5 ns (500 frames).
In the case of NP with free peptides, five Aβ40 peptides are randomly distributed surrounding each NP. The systems were kept in a 150 mM NaCl solution. At the beginning of the simulations, ions and water molecules were equilibrated for 2 ns. Then each system was simulated for 100 ns with the similar conditions like NP-fibril systems to identify the NPs that can capture freely solvated peptides due to their high mutual affinity. Then, we calculated the interaction energies of each peptide with NP surfaces using VMD energy analysis plugin considering water dielectric constant 78.5. We also calculated the number of heavy atoms of each peptide within 5 Å of the NP surfaces. All the analyses were performed on the last 5 ns (500 frames) of the simulations.
References
Bianco, A., Kostarelos, K. & Prato, M. Applications of carbon nanotubes in drug delivery. Curr. Opin. Chem. Biol. 9, 674–679, https://doi.org/10.1016/j.cbpa.2005.10.005 (2005).
Lademann, J. et al. Nanoparticles – an efficient carrier for drug delivery into the hair follicles. Eur. J. Pharm. Biopharm. 66, 159–164, https://doi.org/10.1016/j.ejpb.2006.10.019 (2007).
Medintz, I. L., Uyeda, H. T., Goldman, E. R. & Mattoussi, H. Quantum dot bioconjugates for imaging, labelling and sensing. Nat. Mater. 4, 435–446, https://doi.org/10.1038/nmat1390 (2005).
Alivisatos, P. The use of nanocrystals in biological detection. Nat. Biotechnol. 22, 47–52, https://doi.org/10.1038/nbt927 (2004).
Pearson, R. M. et al. Tuning the selectivity of dendron micelles through variations of the poly(ethylene glycol) corona. ACS Nano 10, 6905–6914, https://doi.org/10.1021/acsnano.6b02708, PMID: 27267700 (2016).
Hsu, H.-J. et al. Poly(ethylene glycol) corona chain length controls end-group-dependent cell interactions of Dendron micelles. Macromolecules 47, 6911–6918, https://doi.org/10.1021/ma501258c (2014).
Lu, B., Smith, T. & Schmidt, J. J. Nanoparticle-lipid bilayer interactions studied with lipid bilayer arrays. Nanoscale 7, 7858–7866, https://doi.org/10.1039/C4NR06892K (2015).
Shemetov, A. A., Nabiev, I. & Sukhanova, A. Molecular interaction of proteins and peptides with nanoparticles. ACS Nano 6, 4585–4602, https://doi.org/10.1021/nn300415x (2012).
Nel, A. E. et al. Understanding biophysicochemical interactions at the nano-bio interface. Nat. Mater. 8, 543–557, https://doi.org/10.1038/nmat2442 (2009).
Cagno, V. et al. Broad-spectrum non-toxic antiviral nanoparticles with a virucidal inhibition mechanism. Nat. Mater. 17, 195, https://doi.org/10.1038/nmat5053 (2017).
Linse, S. et al. Nucleation of protein fibrillation by nanoparticles. P. Natl. Acad. Sci. 104, 8691–8696, https://doi.org/10.1073/pnas.0701250104 (2007).
Gladytz, A., Abel, B. & Risselada, H. J. Gold-induced fibril growth: The mechanism of surface-facilitated amyloid aggregation. Angew. Chem. Int. Ed. 55, 11242–11246, https://doi.org/10.1002/anie.201605151 (2016).
Colvin, V. L. & Kulinowski, K. M. Nanoparticles as catalysts for protein fibrillation. P. Natl. Acad. Sci. 104, 8679–8680, https://doi.org/10.1073/pnas.0703194104, http://www.pnas.org/content/104/21/8679.full.pdf (2007).
Palmal, S., Jana, N. R. & Jana, N. R. Inhibition of amyloid fibril growth by nanoparticle coated with histidine-based polymer. J. Phys. Chem. C 118, 21630–21638, https://doi.org/10.1021/jp505613g (2014).
Yoo, S. I. et al. Mechanism of fibrillation inhibition of amyloid peptides by inorganic nanoparticles reveal functional similarities with proteins. Angew. Chem. Int. Ed. 50, 5110–5115, https://doi.org/10.1002/anie.201007824 (2011).
Barros, H. R., Kokkinopoulou, M., Riegel-Vidotti, I. C., Landfester, K. & Thérien-Aubin, H. Gold nanocolloid–protein interactions and their impact on β-sheet amyloid fibril formation. RSC Adv. 8, 980–986, https://doi.org/10.1039/C7RA11219J (2018).
Song, Y., Moore, E. G., Guo, Y. & Moore, J. S. Polymer-peptide conjugates disassemble amyloid β fibrils in a molecularweight dependent manner. J. Am. Chem. Soc. 139, 4298–4301, https://doi.org/10.1021/jacs.7b00289 (2017).
Brancolini, G. et al. Probing the influence of citrate-capped gold nanoparticles on an amyloidogenic protein. ACS Nano 9, 2600–2613, https://doi.org/10.1021/nn506161j (2015).
Radic, S., Davis, T. P., Ke, P. C. & Ding, F. Contrasting effects of nanoparticle-protein attraction on amyloid aggregation. RSC Adv. 5, 105498, https://doi.org/10.1039/c5ra20182a (2015).
Bellucci, L. et al. The interaction with gold suppresses fiber-like conformations of the amyloid β (16–22) peptide. Nanoscale 8, 8737–8748, https://doi.org/10.1039/C6NR01539E (2016).
Song, M. et al. Exploring the mechanism of inhibition of au nanoparticles on the aggregation of amyloid-β (16–22) peptides at the atom level by all-atom molecular dynamics. Int. J. Mol. Sci. 19, 1815, https://doi.org/10.3390/ijms19061815 (2018).
John, T. et al. Impact of nanoparticles on amyloid peptide and protein aggregation: a review with a focus on gold nanoparticles. Nanoscale 10, 20894–20913, https://doi.org/10.1039/C8NR04506B (2018).
Scherzer-Attali, R. et al. Complete phenotypic recovery of an alzheimer’s disease model by a quinone-tryptophan hybrid aggregation inhibitor. PLoS One 5, 1–15, https://doi.org/10.1371/journal.pone.0011101 (2010).
Luo, Q. et al. A self-destructive nanosweeper that captures and clears amyloid β-peptides. Nat. Commun. 9, 1802, https://doi.org/10.1038/s41467-018-04255-z (2018).
Funke, S. A. et al. Oral treatment with the d-enantiomeric peptide d3 improves the pathology and behavior of alzheimer’s disease transgenic mice. ACS Chem. Neurosci. 1, 639–648, https://doi.org/10.1021/cn100057j (2010).
Jia, Y.-P., Ma, B.-Y., Wei, X.-W. & Qian, Z.-Y. The in vitro and in vivo toxicity of gold nanoparticles. Chin. Chem. Lett. 28, 691–702, https://doi.org/10.1016/j.cclet.2017.01.021 (2017).
Petkova, A. T., Yau, W.-M. & Tycko, R. Experimental constraints on quaternary structure in alzheimer’s β-amyloid fibrils. Biochemistry 45, 498–512, https://doi.org/10.1021/bi051952q (2006).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815, https://doi.org/10.1006/jmbi.1993.1626 (1993).
Fiser, A., Do, R. & Sali, A. Modeling of loops in protein structures. Protein Sci. 9, 1753–1773, https://doi.org/10.1110/ps.9.9.1753 (2000).
Humphrey, W., Dalke, A. & Schulten, K. Vmd: Visual molecular dynamics. J. Mol. Graph. 14, 33–38, https://doi.org/10.1016/0263-7855(96)00018-5 (1996).
Phillips, J. C. et al. Scalable molecular dynamics with namd. J. Comput. Chem. 26, 1781–1802, https://doi.org/10.1002/jcc.20289 (2005).
Vanommeslaeghe, K. et al. Charmm general force field (cgenff): A force field for drug-like molecules compatible with the charmm all-atom additive biological force fields. J. Comput. Chem. 31, 671–690, https://doi.org/10.1002/jcc.21367 (2010).
Vanommeslaeghe, K. & MacKerell, A. D. Automation of the charmm general force field (cgenff) i: Bond perception and atom typing. J. Chem. Inf. Model. 52, 3144–3154, https://doi.org/10.1021/ci300363c (2012).
Yu, W., He, X., Vanommeslaeghe, K. & MacKerell, A. D. Extension of the charmm general force field to sulfonyl-containing compounds and its utility in biomolecular simulations. J. Comput. Chem. 33, 2451–2468, https://doi.org/10.1002/jcc.23067 (2012).
Best, R. B. et al. Optimization of the additive charmm all-atom protein force field targeting improved sampling of the backbone ϕ, ψ and side-chain χ 1 and χ 2 dihedral angles. J. Chem. Theory Comput. 8, 3257–3273, https://doi.org/10.1021/ct300400x (2012).
MacKerell, A. D. Jr. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616, https://doi.org/10.1021/jp973084f (1998).
Huang, J. & MacKerell, A. D. Charmm36 all-atom additive protein force field: Validation based on comparison to nmr data. J. Comput. Chem. 34, 2135–2145, https://doi.org/10.1002/jcc.23354 (2013).
Darden, T., York, D. & Pedersen, L. Particle mesh ewald: An nlog(n) method for ewald sums in large systems. J. Chem. Phys. 98, 10089–10092, https://doi.org/10.1063/1.464397 (1993).
Zheng, J., Jang, H., Ma, B., Tsai, C.-J. & Nussinov, R. Modeling the alzheimer aβ17–42 fibril architecture: tight intermolecular sheet-sheet association and intramolecular hydrated cavities. Biophys. J. 93, 3046–3057, https://dx.doi.org/10.1529%2Fbiophysj.107.110700 (2007).
Acknowledgements
The authors thank Prof. Francesco Stellacci for valuable discussions. Some of the calculations were performed at the UIC Extreme supercomputing cluster. Research reported in this publication (L.V.) was supported by the National Institute Of Allergy And Infectious Diseases of the National Institutes of Health under Award Number R03AI142553. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. The authors would also like to acknowledge the Research Open Access Publishing (ROAAP) Fund of the University of Illinois at Chicago for financial support towards the open access publishing fee for this article.
Author information
Authors and Affiliations
Contributions
Must include all authors, identified by initials, for example: P.K. conceived the experiment(s), S.S. and L.V. conducted the experiment(s), S.S. analyzed the results. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sen, S., Vuković, L. & Král, P. Computational screening of nanoparticles coupling to Aβ40 peptides and fibrils. Sci Rep 9, 17804 (2019). https://doi.org/10.1038/s41598-019-52594-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-52594-8
- Springer Nature Limited