Abstract
Plasmids may maintain antibiotic resistance genes in bacterial populations through conjugation, in the absence of direct selection pressure. However, the costs and benefits of conjugation for plasmid and bacterial fitness are not well understood. Using invasion and competition experiments with plasmid mutants we explicitly tested how conjugation contributes to the maintenance of a plasmid bearing a single extended-spectrum ß-lactamase (ESBL) gene (blaCTX-M-14). Surprisingly, conjugation had little impact on overall frequencies, although it imposed a substantial fitness cost. Instead, stability resulted from the plasmid conferring fitness benefits when rare. Frequency dependent fitness did not require a functional blaCTX-M-14 gene, and was independent of culture media. Fitness benefits when rare are associated with the core plasmid backbone but are able to drive up frequencies of antibiotic resistance because fitness burden of the blaCTX-M-14 gene is very low. Negative frequency dependent fitness can contribute to maintaining a stable frequency of resistance genes in the absence of selection pressure from antimicrobials. In addition, persistent, low cost resistance has broad implications for antimicrobial stewardship.
Similar content being viewed by others
Introduction
Understanding the selection pressures that maintain antimicrobial resistance (AMR) genes is key to understanding how resistance will respond to antibiotic usage and how we might develop interventions to extend the use of antibiotics in the face of resistance. Importantly, AMR gene frequencies show a varied and unpredictable response to reduced rates of usage: in some cases, resistance declines readily after antibiotics are withdrawn, in other cases resistance genes persist effectively in the absence of recent selection1,2,3. Moreover, even when selection pressure is intense AMR genes may not rise to fixation4. Understanding the fate of antibiotic resistance genes when antibiotic use is reduced or withdrawn is key to developing effective resistance management or ‘antibiotic stewardship’ strategies, especially those that rely on cycling of antimicrobials5.
The situation is further complicated by the role of bacterial plasmids, which are responsible for a substantial proportion of clinically important resistance and for the accumulation of multi-drug resistance by many of the most difficult to treat nosocomial bacteria6. Plasmids can maintain themselves, and the AMR genes they carry, as genetic parasites in the absence of direct selection pressure from antibiotics7. If rates of horizontal transfer (conjugation) are sufficient to offset fitness cost of carriage or loss from segregation, then plasmids may be stable or capable of increasing in frequency in bacterial populations without selection from antibiotic usage2,8.
In this study, we sought to explicitly test the importance of conjugation for stable maintenance of the IncK plasmid pCT in Escherichia coli in the absence of antibiotic selection. This plasmid carries a single resistance gene - the extended-spectrum ß-lactamase (ESBL) blaCTX-M-14 - which confers resistance to penicillins and third generation cephalosporins9. Invasion and competition experiments used wild type (highly conjugative) and conjugation-null plasmid mutants and a blue/white marker system based on inactivation of ß-galactosidase to track plasmid transfer (see Methods) and cost of engaging in conjugation. In addition we tested the contribution of a functioning blaCTX-M-14 for the fitness, persistence and stability of pCT and non-conjugative vectors (see Table 1 for a full list of mutants).
Results and Discussion
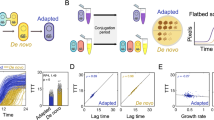
We expected that conjugation would enable plasmid persistence in the absence of antibiotics, or even facilitate the invasion of plasmids from low to high frequencies. However, experimental data showed that the pCT plasmid consistently increased in frequency regardless of ability to conjugate (Fig. 1A,B). Conjugation did increase plasmid frequencies overall (mixed model -likelihood ratio = 8.16, df = 1, P = 0.0043), but this effect was small when compared to the increase in frequency over several orders of magnitude through time (likelihood ratio = 322, df = 1, P < 0.0001). Plasmid frequencies declined after reaching a maximum of 10−2 −10−1 (Fig. 1A,B) (significant quadratic term- time2 likelihood ratios = 37.9 df = 1, P < 0.0001). In the wild type pCT transconjugants made up a substantial proportion of plasmid carriers after only five transfers (Fig. 1C), indicating that the similarities between conjugative and non-conjugative plasmids were not the result of low conjugation. The genetic background of the initial plasmid donor (wildtype or ΔlacZYA) had no impact on plasmid dynamics (Likelihood ratio = 1.97, df = 1, P = 0.16).
The invasion dynamics of wild type pCT plasmids (+symbols and solid lines) and conjugation null mutants (ΔtrbA mutant – open circles and dashed lines) in transfer experiments in complex nutrient rich LB broth initiated with a low frequency of plasmid carriers. Experiments were repeated using initial plasmid donors in the chromosomal MG1655 wild background (A – ‘blue donor’ with blue data and fitted models) and in the MG1655 Δlac mutant background (B – ‘white donor with black data and fitted models). Fitted models are quadratic glms with transfer and transfer2 as covariates. C The relative contribution of transconjugants (darker shading) and plasmids carriers having the original donor background (lighter shading) to the total bacterial population. Data are mean proportions from the 12 wild type plasmid replicates only, the blue panel shows data from blue donor and the grey panel data from white donors. D Relative fitness of MG1655 bacteria carrying the Δlac mutant over different initial frequencies, fitness >1 indicates beneficial plasmid carriage. Experiments were repeated with blue and white donors, as above.
The pCT plasmid is highly persistent, and previously, the mechanism accounting for persistence was inferred to be a near-zero fitness cost of carriage10. However, the dynamics of invasion from rare seen here, and the subsequent decline from high frequencies, are better explained by pCT conferring a negative frequency dependent (NFD) fitness advantage to its host, where fitness is higher when a genotype is present at lower frequencies in the population. Competition experiments using our conjugation null pCT∆trbA mutant confirmed that carriage of this plasmid was beneficial at low frequencies and costly at high frequencies (Fig. 1D; F1,70 = 35.0, P < 0.001). As above, there was no effect of bacterial background on relative fitness (F1,69 = 1.6, P = 0.21). Moreover, these fitness experiments indicate that equilibrium carriage (when plasmid free and plasmid carrying bacteria have equal fitness) is in the region of 10−1 – 10−2 cells, which corresponds to the maximum frequencies in the transfer experiment (Fig. 1A,B). In the early transfers we are able to show the relative contribution of conjugation and fitness effects to total carriage of wild type pCT (Fig. 1C). Importantly, even without conjugation, the original plasmid carriers were able to increase in frequency from 1 in 105 to 1 in 50 cells.
The focal pCT plasmid has few non-essential genes that are not required for transfer and maintenance: the typical IncI backbone accounts for most gene content9. Nevertheless, poorly characterized genes could encode metabolic functions responsible for frequency dependency fitness via resource competition11. However, plasmid carrying cells could also invade from rare in minimal media (M9 broth), where complex resources are absent (Fig. 2A,B). In addition, very high numbers of transconjugants appeared after only three transfers, suggesting a higher level of conjugation rate in minimal media (Fig. 2C). We measured pCT conjugation in both media in a separate experiment using a marked recipient strain, and confirmed that conjugation frequencies are approximately 30-fold higher in M9 than in LB (Fig. S1). Consistent with this observation, in minimal media the wild type plasmid increased in frequency more rapidly than the non-conjugative mutant (Fig. 2; time*plasmid interaction; Likelihood ratio = 32.2, df = 1, P < 0.0001), while there was no such effect in complex media (Fig. 1; time*plasmid interaction; Likelihood ratio = 2.54, df = 1, P = 0.11).
The invasion dynamics of wild type pCT plasmids (+symbols and solid lines) and conjugation null mutants (ΔtrbA mutant – open circles and dashed lines) in minimal media initiated with a low frequency of plasmid carriers in the (A) blue donor background and (B) white donor background (C) The relative contribution of transconjugants (darker shading) to the total bacterial population. Plasmids carriers having the original donor background (lighter shading) have also been plotted but these values are too low to be visible (max proportion = 0.002). Data are mean proportions from the 12 wild type plasmid replicates only. As in Fig. 1 the blue panel shows data from blue donor and the grey panel data from white donors.
Next, we investigated the importance of the resistance gene blaCTX-M−14 on bacterial fitness and patterns of NFD. Replacing the blaCTX-M-14 with a kanamycin resistance cassette substantially increased the fitness burden associated with pCT∆trbA (Fig. 3A, reduction in fitness relative to pCT∆trbA -0.18, t = −10.7, P < 0.0001), indicating that the fitness cost of blaCTX-M-14 might be particularly low. Knock-in genetic manipulation with a cloning vector showed that the fitness cost of blaCTX-M-14 was indistinguishable from zero in conventional competition experiments (Fig. 3B). Detoxifying ß-lactamase resistance, including that conferred by blaCTX-M-14 can confer NFD fitness to resistant bacteria in the presence of ß-lactam antibiotics12,13,14. Conceivably, this gene might provide an NFD advantage in the absence of antibiotic if it were detoxifying another substrate. However, deleting blaCTX-M-14 did not abolish the NFD fitness of conjugational null pCT∆trbA (Fig. 3A). Knock-in experiments also confirmed that NFD fitness was not associated with blaCTX-M-14 (Fig. 3B).
(A) Frequency-dependent fitness of WT plasmids in comparison to conjugation null pCTΔtrbA and double mutants also lacking the blaCTX-M-14 beta-lactamase gene pCTΔtrbAΔbla::aph. All three genotypes show strong negative frequency dependent fitness (F1,208 = 19.9, P < 0.0001) and there is a strong effect of genotype on average relative fitness (F2,206 = 86.5, P < 0.0001). Fitness data were generated independently to experiments in Fig. 1. (B) Impact of blaCTX-M-14 and initial frequency on fitness costs of carriage of non-conjugative vector pZS*2 R in competition experiments. Data points are independent repeats with a fitted linear model ± SE. Frequency did not affect the relative fitness of plasmids with and without this ESBL gene (F1,65 = 1.48, P = 0.23). The relative fitness of resistant plasmids carrying blaCTX-M-14 was estimated at 0.985 (confidence interval 0.967–1.002).
Evidence that conjugation determines a substantial proportion of the fitness costs imposed by plasmids has been indirect15,16,17. Here we used the wild type pCT and pCT∆trbA to assess the fitness cost associated with conjugation directly. This cost was substantial (Fig. 3, reduction in fitness relative to pCT∆trbA -0.15, t = −6.57, P < 0.0001). These competition experiments can explain why the loss of ability to conjugate had a minimal impact on plasmid dynamics: the gain in competitive fitness accompanying loss of conjugation compensates very closely for loss of ability to move between bacterial lineages. Only in minimal media was there a gain in plasmid level fitness for maintaining conjugation (Fig. 2). Experiments with diverse plasmids indicate that conjugation can readily offset the costs of plasmid carriage7, in this study the benefits of conjugation could only just offset the costs of conjugation in nutrient rich media. A high cost of conjugation can explain the existence of natural conjugation-null mutations in plasmids encoding virulence factors in Bacillus anthracis or multi-drug resistance in E. coli18,19.
Importantly, NFD was not sufficient to allow plasmid invasion under all circumstances. This mechanism can only increase frequency for plasmids encoding low cost resistance genes. For example, the mutant carrying kanamycin resistance could not have invaded populations as carriers always had a relative fitness less than that of plasmid-free cells (Fig. 3). NFD fitness has widespread implications for the maintenance of low cost AMR genes outside the laboratory, and could stabilize gene frequencies of resistance when drugs are withdrawn. The latest surveillance data in England shows that resistance to third generation cephalosporins in E. coli bacteraemia remained stable at 10%, despite considerable efforts to reduce prescription rates for these drugs20, while frequencies of particular resistant lineages such as ST131 also show strong stability21. Another potential outcome is that selection against plasmids at high frequencies might stabilize AMR gene frequencies, at least in the face of modest selection pressure. Important caveats here are that continuous selection from antibiotic usage can favour the transfer of genes to the chromosome, and this has been documented for E. coli22,23,24. In addition, data on prevalence of ESBL genes in Asia suggests that the frequency of these genes can reach very high levels (50–70%) under strong selection25.
Many plasmid encoded traits such as ß-lactamases, Cry toxins and detoxifying mercury resistance confer NFD fitness via social interactions in structured environments14,26,27, while colicin plasmids can confer positive frequency dependent fitness28, as well as negative density dependent fitness29. A wide range of secreted bacterial virulence factors are encoded on mobile elements, and these are commonly inferred to be cooperative public goods30,31,32. Current theory suggests that social traits accumulate on plasmids because horizontal transfer provides a means of reducing competition from cheaters32,33 or a means of increasing genetic relatedness30. An alternative, non-mutually exclusive explanation is that plasmids accumulate genes that drive frequency dependence, and that social interactions are just one route to this phenomenon. The existence of both positive and negative frequency dependent selection on plasmids suggests that fluctuating selection pressure is the important common factor here. Recent genomic analyses suggest that NFD may be widespread for accessory genes in Streptococcus pneumonia and E. coli and that mobile elements contain the majority of genes at intermediate frequency likely to drive NFD21,34. Mechanisms could include bacteria-host, bacteria-bacteria or bacteria-MGE interactions in addition to niche specialization. Here we show that bacteria-bacteria interactions are sufficient to produce strong NFD.
Methods
The strains used in this study are E. coli K-12 MG1655 and a MG1655 ΔlacZYA mutant. The MG1655 lac operon deletion was described previously12 and produced a Δlac mutant with a white phenotype in the presence of X-Gal & IPTG as opposed to the blue wild type phenotype. The focal plasmid in this study is pCT, an IncK plasmid, which carries the ESBL resistance gene blaCTX-M-14, which confers effective resistance to the third generation cephalosporin antibiotic cefotaxime9. We produced a conjugation null mutant of pCT via disruption of trbA, which encodes a putative regulation protein, at position 38848–40110 on pCT, as it is indispensable for plasmid transfer35. This mutant was produced by the Xercise protocol36. Diagnostic PCR was used to assess the insertion and subsequent deletion of the dif-CAT-dif fragment into MG1655. Diagnostic PCR was conducted using Taq (Qiagen, Manchester, UK) with an annealing temperature of 55 °C and using the primer F TrbA diag (5′-CGGCATCCAGGCAGGCATCA-3′) and R TrbA diag (5′-TTCAGCCCTGCCCGGTCATT-3′). Deletion used the primers F-TrbA del (5′–TTCTGCATCAACGGTATCAACAAGCACCGTTTCAGTTATTTCAGTGT GCTGGAATTCGCCCT-3′) and R-TrbA del (5′-ATTGTTCGCATTAATTCCACTC AGCCTCATCCCGAAATTTATCTATTTATCTGCAGAATTCGCCCTTCCT-3′). Electrocompetent cells were prepared according to the Bio-Rad protocol, and transformed with both pCT and pCTΔtrbA plasmids using MicroPulser electroporation apparatus (Bio-Rad). Phenotypic confirmation of loss of conjugative ability was confirmed using mating experiments37 and no trans-conjugants were identified from transfer experiments using pCTΔtrbA in this study. Additional plasmid mutants include a CTX-M-14 deletion mutant, disrupted by insertion of a kanamycin resistant cassette10, a gift of Laura Piddock (Univ. of Birmingham). To assess the impact of the blaCTX-M-14 gene on fitness, this gene was cloned into the pZS*2 R vector38, a low copy plasmid with a pSC101* replication origin and a kanamycin resistance marker. Amplification of blaCTX-M-14 from pCT used primers containing a XhoI restriction site and this product was ligated with pZS*2 R plasmid after digestion with XhoI. pZS-bla plasmids were transformed into MG1655; pZS*2 R control plasmid was transformed into MG1655 ΔlacZYA.
Plasmids were transformed into both host backgrounds. There is no detectable fitness difference between the wild type and the lacZYA knockout in broth (mean relative fitness of wild type = 1.006, SE = 0.02, n = 32); this fitness is also unaffected by relative frequency (F1,31 = 0.0001, P = 0.99). The plasmid transfer experiment replication used six replicates per treatment using wild type pCT and the conjugation null mutant (ΔtrbA). Transfers were initiated with two initial proportions of carriers, approximately 0.0001–0.00001, although these treatments had no impact on plasmid dynamics and were pooled in the analyses here. Transfer experiments were duplicated by switching the plasmid carrier strain between the wildtype and the ΔlacZYA ‘white’ mutant and competing plasmids with the opposing blue or white background genotype. The transfer regime used 1000x dilution into 1 ml of Lysogeny Broth (LB) in 24-well plates- transfers took place every 2–3 days. In transfer and competition experiments the proportion of blue and white mutants, as well as the frequency of pCT in both backgrounds was monitored by dilution plating cultures onto LB agar containing X-gal and IPTG (with and without ampicillin 100 μg/ml) each week. A replicate experiment used transfer in M9 broth and ran for 3 transfers only. To evaluate conjugation frequencies during the transfer experiment, we mixed the ΔlacZYA pCT donor strain and a MG1655 spontaneous mutant resistant to rifampicin39 in 50/50 ratios and measured donor, recipient and transconjugant densities after 24 h on ampicillin (100 μg/ml), rifampicin (100 μg/ml) and ampicillin + rifampicin respectively.
Measurement of bacterial fitness
Fitness measurements were calculated in competition experiments. Initially, separate cultures were prepared in overnight cultures of 1 ml LB broth (37 °C, 180 rpm). Competing cultures were then mixed at desired ratios and diluted 1:100 into fresh broth and co-cultured for 24 h to homogenize the physiology of competitors. Densities and relative frequencies (day1) were calculated by dilution plating and mixtures were again diluted at 1:100 and cultured for 24 hours. Measurement of frequencies on day 2 and day 1 were used to calculate relative growth rate (loge(final proportion/(initial proportion*100), relative fitness is then calculated as the ratio of the relative growth rates of the two competing strains40. By convention fitness was expressed as the relative growth of plasmid carriers/plasmid null strains, or of antibiotic resistant/antibiotic susceptible competitors.
Data analysis
The serial transfer experiments were designed such that a linear mixed-effects (lme) model could be used to analyze the resulting data, with the proportion of plasmid carriers as the response variable; time, plasmid type, donor type and transfer regime as fixed effects; and replicate as a random effect. Statistical modelling was conducted in R v3.2.2 using the nlme package41. Restricted maximum likelihood (REML) lme models were converted to maximum likelihood (ML) models to allow model comparison using mixed model ANOVA via Likelihood ratio tests or the Aikaike Information Criterion (AIC). Fitness experiments were analysed using generalized linear modeling, planned post hoc comparisons used treatment contrasts.
Data availability
The research data supporting this publication are openly available from the University of Exeter's institutional repository at: https://doi.org/10.24378/exe.2043. Mutants are available on request from BR.
References
Lipsitch, M. The rise and fall of antimicrobial resistance. Trends Microbiol 9, 438–444 (2001).
Enne, V. I. et al. Assessment of the fitness impacts on Escherichia coli of acquisition of antibiotic resistance genes encoded by different types of genetic element. J Antimicrob Chemo 56, 544–551 (2005).
Whittles, L. K., White, P. J. & Didelot, X. Estimating the fitness cost and benefit of cefixime resistance in Neisseria gonorrhoeae to inform prescription policy: a modelling study. PLoS Med 14 (2017).
Colijn, C. et al. What is the mechanism for persistent coexistence of drug-susceptible and drug-resistant strains of Streptococcus pneumoniae? J R Soc Interface 7, 905–919 (2010).
Raymond, B. Five rules for resistance management in the antibiotic apocalypse, a roadmap for integrated microbial management. Evol Appl, 1–13, (2019).
Hawkey, P. M. & Jones, A. M. The changing epidemiology of resistance. J Antimicrob Chem 64, i3–i10 (2009).
Lopatkin, A. J. et al. Persistence and reversal of plasmid-mediated antibiotic resistance. Nat Comm 8, 1689 (2017).
Vogwill, T. & Maclean, R. C. The genetic basis of the fitness costs of antimicrobial resistance: a meta‐analysis approach. Evol Appl 8, 284–295 (2015).
Cottell, J. L. et al. Complete sequence and molecular epidemiology of IncK epidemic plasmid encoding blaCTX-M-14. Emerg Inf Dis 17, 645–652 (2011).
Cottell, J. L., Webber, M. A. & Piddock, L. J. V. Persistence of transferable extended-spectrum-beta-Lactamase resistance in the absence of antibiotic pressure. Antimicrob Agents Chemother 56, 4703–4706 (2012).
MacLean, R. C. & Gudelj, I. Resource competition and social conflict in experimental populations of yeast. Nature 441, 498–501 (2006).
Medaney, F., Dimitriu, T., Ellis, R. J. & Raymond, B. Live to cheat another day: bacterial dormancy facilitates the social exploitation of β-lactamases. ISME J 10, 778–787 (2015).
Dugatkin, L. A., Perlin, M., Lucas, J. S. & Atlas, R. Group-beneficial traits, frequency-dependent selection and genotypic diversity: an antibiotic resistance paradigm. Proc. R. Soc. Lond. Ser. B-Biol. Sci. 272, 79–83 (2005).
Yurtsev, E. A., Chao, H. X., Datta, M. S., Artemova, T. & Gore, J. Bacterial cheating drives the population dynamics of cooperative antibiotic resistance plasmids. Mol. Syst. Biol. 9, 1–7 (2013).
Koraimann, G. & Wagner, M. A. Social behavior and decision making in bacterial conjugation. Front Cell Infect Microbiol 4 (2014).
Dahlberg, C. & Chao, L. Amelioration of the cost of conjugative plasmid carriage in Eschericha coli K12. Genetics 165, 1641–1649 (2003).
Haft, R. J. F., Mittler, J. E. & Traxler, B. Competition favours reduced cost of plasmids to host bacteria. ISME J 3, 761–769 (2009).
Hu, X., Van der Auwera, G., Timmery, S., Zhu, L. & Mahillon, J. Distribution, diversity, and potential mobility of extrachromosomal elements related to the Bacillus anthracis pXO1 and pXO2 virulence plasmids. Appl Env Microbiol 75, 3016–3028 (2009).
Woodford, N. et al. Complete Nucleotide Sequences of Plasmids pEK204, pEK499, and pEK516, Encoding CTX-M Enzymes in Three Major Escherichia coli Lineages from the United Kingdom, All Belonging to the International O25:H4-ST131 Clone. 53, 4472-4482, (2009).
Public Health England, (2016). English Surveillance Program for antimicrobial utilisation and resistance (ESPAUR). Public Health England, London, UK.
McNally, A. et al. Diversification of colonization factors in a multidrug-resistant Escherichia coli lineage evolving under negative frequency-dependent selection. mBio 10, 860–819 (2019).
Hall, J. P. J., Wood, A. J., Harrison, E. & Brockhurst, M. A. Source-sink plasmid transfer dynamics maintain gene mobility in soil bacterial communities. Proc Natl Acad Sci USA 113, 8260–8265 (2016).
Bergstrom, C. T., Lipsitch, M. & Levin, B. R. Natural selection, infectious transfer and the existence conditions for bacterial plasmids. Genetics 155, 1505–1519 (2000).
McNally, A. et al. Combined analysis of variation in core, accessory and regulatory genome regions provides a super-resolution view into the evolution of bacterial populations. PLoS Genet 12, e1006280–1006216 (2016).
Kaul, S. et al. One year trends in the gram-negative bacterial antibiotic susceptibility patterns in a medical intensive care unit in South India. Ind J Med Microbiol 25, 230 (2007).
Raymond, B., West, S. A., Griffin, A. S. & Bonsall, M. B. The dynamics of cooperative bacterial virulence in the field. Science 337, 85–88 (2012).
Ellis, R. J., Lilley, A. K., Lacey, S. J., Murrell, D. & Godfray, H. C. J. Frequency-dependent advantages of plasmid carriage by Pseudomonas in homogeneous and spatially structured environments. ISME J 1, 92–95 (2007).
Adams, J., Kinney, T., Thompson, S., Rubin, L. & Helling, R. B. Frequency-dependent selection for plasmid-containing cells of Escherichia coli. Genetics 91, 627–637 (1979).
Gerardin, Y., Springer, M. & Kishony, R. A competitive trade-off limits the selective advantage of increased antibiotic production. Nature Micro 1, 1–7 (2016).
Rankin, D. J., Rocha, E. P. C. & Brown, S. P. What traits are carried on mobile genetic elements, and why? Heredity 106, 1–10 (2010).
Nogueira, T. et al. Horizontal gene transfer of the secretome drives the evolution of bacterial cooperation and virulence. Curr Biol 19, 1683–1691 (2009).
Smith, J. The social evolution of bacterial pathogenesis. Proc R Soc Lond B i 268, 61–69 (2001).
Dimitriu, T. et al. Genetic information transfer promotes cooperation in bacteria. Proc Natl Acad Sci USA 111, 11103–11108 (2014).
Corander, J. et al. Frequency-dependent selection in vaccine-associated pneumococcal population dynamics. Nat Ecol Evol 1, 1950–1960 (2017).
Komano, T., Yoshida, T., Narahara, K. & Furuya, N. The transfer region of IncI1 plasmid R64: similarities between R64 tra and Legionella icm/dot genes. Mol Microbiol 35, 1348–1359 (2000).
Bloor, A. & Cranenburgh, R. An efficient method of selectable marker gene excision by Xer recombination for gene replacement in bacterial chromosomes. Appl Environ Micrbiol 72, 2520–2525 (2006).
Liebana, E. et al. Characterization of β-lactamases responsible for resistance to extended-spectrum cephalosporins in Escherichia coli and Salmonella enterica strains from food-producing animals in the United Kingdom. Microb Drug Resist 44, 1630–1634 (2004).
Chuang, J. S., Rivoire, O. & Leibler, S. Simpson’s paradox in a synthetic microbial system. Science 323, 272–275 (2009).
Dimitriu, T., Marchant, L., Buckling, A. & Raymond, B. Bacteria from natural populations transfer plasmids mostly towards their kin. Proc R Soc B 286, 20191110 (2019).
Lenski, R. E. Experimental Studies of Pleiotropy and Epistasis in Escherichia coli. I. Variation in Competitive Fitness Among Mutants Resistant to Virus T4. Evolution 42, 425 (1988).
Pinheiro, J. C., Bates, D. M., DebRoy, S. & Sarkar, D. Linear and non-linear mixed effect models. R package version (2007).
Acknowledgements
F.M. was supported by a joint AHVLA and Royal Holloway studentship. E.A. was supported by a grant from the Charles and Elsie Sykes Trust; B.R. received additional support from MRC grant MR/N013824/1. Thanks to Laura Piddock and John Chuang for the gift of mutants and/or vectors.
Author information
Authors and Affiliations
Contributions
B.R., F.M. and R.E. devised the study; F.M. and T.D. produced the mutants. T.D., F.M., J.F. and E.A. carried out experiments. F.M., T.D. and B.R. analyzed the data; B.R. wrote the first draft; all authors contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dimitriu, T., Medaney, F., Amanatidou, E. et al. Negative frequency dependent selection on plasmid carriage and low fitness costs maintain extended spectrum β-lactamases in Escherichia coli. Sci Rep 9, 17211 (2019). https://doi.org/10.1038/s41598-019-53575-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-53575-7
- Springer Nature Limited