Abstract
Rapid modulation of RNA function by endoribonucleases during physiological responses to environmental changes is known to be an effective bacterial biochemical adaptation. We report a molecular mechanism underlying the regulation of enolase (eno) expression by two endoribonucleases, RNase G and RNase III, the expression levels of which are modulated by oxygen availability in Escherichia coli. Analyses of transcriptional eno-cat fusion constructs strongly suggested the existence of cis-acting elements in the eno 5′ untranslated region that respond to RNase III and RNase G cellular concentrations. Primer extension and S1 nuclease mapping analyses of eno mRNA in vivo identified three eno mRNA transcripts that are generated in a manner dependent on RNase III expression, one of which was found to accumulate in rng-deleted cells. Moreover, our data suggested that RNase III-mediated cleavage of primary eno mRNA transcripts enhanced Eno protein production, a process that involved putative cis-antisense RNA. We found that decreased RNase G protein abundance coincided with enhanced RNase III expression in E. coli grown anaerobically, leading to enhanced eno expression. Thereby, this posttranscriptional up-regulation of eno expression helps E. coli cells adjust their physiological reactions to oxygen-deficient metabolic modes. Our results revealed a molecular network of coordinated endoribonuclease activity that post-transcriptionally modulates the expression of Eno, a key enzyme in glycolysis.
Similar content being viewed by others
Introduction
Genome-wide analyses of mRNA abundance at single-gene resolution have facilitated the identification of ribonuclease RNA targets in bacteria1,2,3,4,5,6,7. These studies showed that RNase III and RNase E endoribonucleases control the stability of mRNAs ranging from hundreds to thousands of genes, whereas RNase G, a paralog of the N-terminal catalytic domain of RNase E, affects the abundance of fewer mRNAs1,2,3,4. The C-terminal domain of RNase E, which is absent in RNase G, interacts with PNPase 3′ → 5′ exoribonuclease, RhlB RNA helicase, and the glycolytic enzyme enolase, forming multicomponent ribonucleolytic complexes termed RNA degradosomes8,9,10. In addition, it also acts as a negative modulator through binding to an inhibitor protein, RraA or RraB, at distinct sites2,11,12,13. In some cases, an RNA binding protein, Hfq, as well as small non-coding antisense RNAs, are associated with the RNA cleavage activity of RNase E (e.g. SgrS and pts mRNA, for a review see14).
A large body of evidence has been accumulated in recent years showing the importance of antisense RNA in the regulation of gene expression6,15,16. For instance, antisense RNAs regulate acid resistance17,18 and type I toxin-antitoxin production (for a review, see19) in Escherichia coli. The Salmonella AmgR/mgtC system has also been shown to be dependent on antisense RNA regulation20. In addition, RNase III, a highly conserved double-stranded RNA-specific endoribonuclease, has been shown to cleave target RNA transcripts that form intra-RNA molecular stem-loop structures by interacting with antisense RNAs6,21,22. This antisense RNA-mediated RNase III cleavage involves both cis- and trans-antisense RNAs transcribed from the same or different genomic loci of target genes, respectively6,21.
The single-strand-specific endoribonucleases RNase G and E are involved in the expression of genes encoding enzymes involved in the major carbon metabolism pathways. RNase E-deficient cells show reduced phosphoenolpyruvate carboxylase production, limiting the conversion of phosphoenolpyruvate to oxaloacetic acid23. RNase E also directly decays ptsG mRNA, which encodes the major glucose transporter, when the glycolytic pathway is blocked24. In addition, deletion of the rng gene, encoding RNase G, results in increased steady-state levels of adhE, eno, glk, pgi, and tpiA mRNAs, proteins that are involved in carbon metabolism1,5,25,26. Moreover, increased mRNA abundance of the eno and tpiA genes is directly associated with protein expression levels in rng-deleted cells26,27, while deletion of cra, encoding a catabolite repressor/activator, in rng-deleted E. coli cells also results in increased pyruvate production in the medium28. However, the mechanisms underlying these rng-mediated alterations in the levels of enzymes and products associated with carbon metabolism remain largely unknown.
While RNase E and G generally destabilise their target mRNAs, RNase III can both decay and process mRNAs. For instance, RNase III cleaves the 5′ UTR of target mRNAs to yield translationally active mRNAs18,29,30. It has also been suggested that many mRNAs can be stabilised by RNase III in E. coli, likely due to ribosomal protection of RNase III-processed mRNAs from ribonucleases4. The enzymatic activity of E. coli RNase III is known to be regulated through stress induced by entry into stationary phase, temperature and osmotic changes, and exposure to aminoglycosides4,31,32,33,34.
While investigating the molecular mechanisms underlying the negative regulation of eno gene expression by RNase G in E. coli, we observed that eno expression was also positively regulated by RNase III. Therefore, in this study, we examined the functional roles of RNase G and RNase III in eno expression, and characterised factors involved in this endoribonucleases-mediated regulation of eno expression.
Results
Effects of cellular concentrations of RNase III and/or RNase G on eno expression
RNase III has been shown to control rng mRNA stability by cleaving its coding region34. Therefore, we first tested whether RNase G-mediated down-regulation of eno expression is associated with RNase III by measuring eno expression levels in wild-type (WT), rnc- (Δrnc), and/or rng-deleted (Δrng) strains. Enolase expression increased approximately 1.6-fold in the Δrng strain compared to that in the WT strain, as has been previously reported27 (Fig. 1a). Deletion of the rnc gene (Δrnc) resulted in an approximately 30% decrease in eno expression. We reasoned that this decreased eno expression was due to increased expression levels of RNase G resulting from the stabilisation of rng mRNA in Δrnc cells. Indeed, the expression levels of RNase G increased 8.8-fold in the Δrnc strain compared to those in WT cells (Fig. 1a). However, inconsistent with the above results, we observed that eno expression levels decreased by approximately 15% in the rnc and rng double-mutant strain (Δrnc rng); eno expression levels in the Δrnc rng strain were expected to be similar to those in the Δrng strain if RNase G alone is solely responsible for the posttranscriptional regulation of eno expression (Fig. 1b). The steady-state levels of eno mRNA were highly correlated with expression levels of Eno in the strains used in Fig. 1a–c). These results suggested the existence of RNase III-mediated positive regulation of eno expression independent of RNase G, in addition to the positive regulation of eno expression via destabilisation of rng mRNA.
Regulation of Eno expression by RNase III and/or RNase G. (a) Effects of rnc and/or rng deletion on the expression level of eno. Escherichia coli MG1655 strains (WT, Δrng, Δrnc, and Δrnc rng) were grown in LB medium at 37 °C to mid-log phase and harvested for western blot analysis of Eno, Rng, and Rnc using protein-specific polyclonal antibodies. The expression levels of Eno, Rng, and Rnc were compared by setting those of WT to 1. (b) Independent modulation of Eno expression levels by RNase G and RNase III. Western blotting was performed as described for (a) using Δrnc rng strains harbouring pPM30, pRNG3, or pRNC3. The expression levels of Eno, Rng, and Rnc were compared by setting those of Δrnc rng harbouring pPM30 to 1. (c) Effects of rnc and/or rng deletion on the eno mRNA abundance. Total cellular RNA was extracted from cultures grown to an OD600 of 0.6 using an RNeasy mini prep kit. The number of amplicons of enolase and other rnpB mRNA amplified from the cDNAs of the (left) WT, Δrnc. Δrng and Δrnc rng strains (right) harbouring pPM30, pRNC3, or pRNG3. The eno mRNA expression levels were compared by setting those of WT or Δrnc rng harbouring pPM30 to 1. PCR products were resolved in an 1.5% agarose gel. (d) Identification of the regulatory DNA region that affected the eno expression levels. Top: Schematic diagram of the eno-cat reporter. Bottom: Effects of RNase G and RNase III expression levels on the degree of chloramphenicol resistance of MG1655 cells. MG1655 WT, Δrng, and Δrnc cells harbouring pERS1 were transformed with pPM30, pRNG3 (RNase G), or pRNC3 (RNase III). The transformants were grown in LB containing 1 mM IPTG to an OD600 of 0.6, diluted, and spotted on LB agar containing 0 (Cm 0) or 75 (Cm 75) μg ml−1 chloramphenicol. For (a,b), the S1 protein was used as an internal standard to evaluate the amount of cell extract in each lane. For (c), the rnpB mRNA was used as an internal standard to evaluate the amount of cell extract in each lane. For (a–c), the data are presented as means ± s. e. m. of at least three independent experiments.
The above results prompted us to test whether eno expression is directly regulated by RNase III cleavage in vivo. The Δrnc rng strain was separately transformed with plasmids that can overexpress either RNase G (pRNG3) or RNase III (pRNC3), following which eno expression levels were assessed. As shown in Fig. 1b, RNase III overexpression in the absence of RNase G (Δrnc rng + pRNC3) resulted in an approximately 67% increase in eno expression compared to the same strain harbouring an empty vector (pPM30), demonstrating the positive effect of cellular concentrations of RNase III on eno expression. Enolase expression was decreased by approximately 31% when RNase G was overexpressed in the Δrnc rng strain (Δrnc rng + pRNG3) (Fig. 1b). Enolase expression levels were not significantly affected by different cellular concentrations of RNase E, which plays a central role in mRNA decay in E. coli and presents substrate specificity and cleavage sites similar to those of RNase G (Supplementary Fig. 1).
Identification of regulatory DNA region for eno expression
The above results indicate that Eno protein abundance is associated with cellular concentrations of both RNase G and RNase III. To determine whether the upstream region of the eno CDS contains the regulatory DNA region that responds to cellular RNase III and RNase G activity, we generated an eno-cat transcriptional fusion reporter construct (pERS1) expressing mRNA containing the 3′ end (403 nucleotides) of the pyrG coding sequence (CDS), the eno 5′ UTR (87 nucleotides), and the first 20 amino acids of the eno CDS (60 nucleotides) fused to the chloramphenicol acetyltransferase (cat) coding region (Fig. 1d). To compare the degree of chloramphenicol resistance among cells expressing different levels of RNase G and RNase III using this reporter system, MG1655 WT, Δrng, and Δrnc cells harbouring pERS1 were transformed with pPM30, pRNG3 (RNase G), or pRNC3 (RNase III), the replication origin of which is derived from pSC10135, thus compatible with pACYC177-derived pERS1. The results showed that, compared to the WT strain harbouring an empty vector (pPM30), overexpression of RNase G rendered the WT strain less resistant to chloramphenicol while overexpression of RNase III resulted in increased resistance to chloramphenicol (Fig. 1d). Overexpression of RNase G rendered the Δrng strain more sensitive to chloramphenicol compared to the same strain harbouring an empty vector (Fig. 1d). In addition, when RNase III was overexpressed, the degree of chloramphenicol resistance of the Δrnc strain was restored to that of the WT strain harbouring an empty vector (Fig. 1d). Collectively, these results showed a good correlation between the activities of the eno-cat fusion and the cellular concentrations of RNase G and RNase III (Fig. 1d), suggesting that the RNase G and RNase III-responsive regulatory DNA region is present between −490 to +60 region of the eno CDS.
We further characterised the RNase G and RNase III-responsive regulatory DNA regions by generating a pERS1-derived plasmid that did not contain the pyrG CDS (pERS2) and, using this reporter system, compared the degree of resistance to chloramphenicol among cells expressing different levels of RNase G and RNase III. We observed a good correlation between the activities of the eno-cat fusion and the cellular concentrations of RNase G and RNase III (Supplementary Fig. 2). The degree of chloramphenicol resistance in all the strains decreased because the removal of the DNA segment from the coding region of pyrG pERS1 resulted in a decreased synthesis of the primary eno transcripts (see below). These results suggest that RNase G and RNase III-responsive cis-acting elements were present between the −87 and +60 region of the eno CDS.
Identification of RNase cleavage sites in eno mRNA
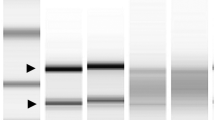
The experimental results shown in Fig. 1d and Supplementary Fig. 2 raised a possibility that RNase III and/or RNase G regulate eno expression by cleaving the 5′ UTR of eno mRNA. To examine whether different eno mRNA species were generated in a manner dependent on the expression of RNase III and/or RNase G, we performed a primer extension analysis using total RNA extracted from the WT, Δrnc, Δrng, and Δrnc rng strains and a 5′ end 32P-labelled primer (eno + 52R) designed to bind downstream of the N-terminal eno coding region. The results showed four cDNA bands (1, 2, 3, and 4) that were generated in a manner dependent on RNase III expression (Fig. 2a). When eno mRNA transcripts were further analysed by S1 nuclease mapping, three of the four putative cleavage products (1, 3, and 4) identified in the primer extension analysis were also detected (Fig. 2b). Canonical RNase III cleavage sites, characterised by a two nucleotide overhang at the 3′ end, were generated by cleavage sites 1 and 3 in the secondary structure of the 5′ UTR of eno mRNA predicted by the M-fold program (http://unafold.rna.albany.edu) (Fig. 2c). RNase III cleavage at sites 2 and 4 did not generate typical RNase III cleavage products. In addition, we observed a significant accumulation of band 3 in the rng-deleted strain in both the primer extension and S1 nuclease mapping analyses. These results suggest that RNase G degrades eno mRNA that is cleaved by RNase III at site 3. Putative cleavage sites 1 and 3 have been suggested as the 5′ ends of eno transcripts that were synthesised from two promoters (enoP4 and enoP6) in the intergenic region between eno and pyrG36,37. If products 1 and 3 were generated by RNase III cleavage, then the other closest transcription start site (enoP7) was located −211 base pairs (bp) from the start codon of the eno coding region, which explains why the 5′ end of the primary eno mRNAs was not detected in our primer extension and S1 nuclease mapping analyses.
Identification of RNase cleavage sites in eno mRNA in vivo. (a) Primer extension analysis of the 5′ UTR of eno mRNA in vivo. Total RNA was isolated from MG1655 strains (WT, Δrng, Δrnc, and Δrnc rng) and hybridised with the 5′ end 32P-labelled primer (eno + 52 R). Synthesised cDNA products were separated on a 6% polyacrylamide gel containing 8 M of urea. Sequencing ladders were synthesised with the same primers used for cDNA synthesis and PCR DNA encompassing the eno gene was used as a template. (b) S1 nuclease mapping. Total RNA was hybridised with the 5′ end 32P-labelled DNA probe. The DNA: RNA complex was treated with 1 U of S1 nuclease and separated in denaturing gel as described above. (c) Predicted eno 5′ UTR secondary structure and RNase cleavage sites. The secondary structure was inferred using the M-fold program. RNase III (1, 2, 3, and 4) cleavage sites identified in (a) and (b) are indicated. The putative Shine–Dalgarno sequence and start codon are indicated as blue and red colours, respectively.
To demonstrate biochemically the cleavage of eno mRNA by RNase III, an in vitro cleavage assay was performed using a model hairpin RNA of eno mRNA containing the RNase III cleavage sites (Supplementary Fig. 3). This synthetic RNA was labelled with 32P at the 3′ end to test for subsequent cleavage of RNase III-cleaved RNAs by RNase G. The results showed that RNase III cleaved the synthetic RNA at several sites that did not correspond to the sites identified by primer extension and S1 nuclease mapping analyses. We also observed that the intact synthetic RNA was not cleaved by RNase G, whereas RNA products generated by RNase III cleavage were cleaved by RNase G at the region encompassing the Shine–Dalgarno sequence of eno mRNA. These results indicated that other factors are involved in generating the RNase III cleavage products identified by the primer extension and S1 nuclease mapping analyses.
Identification of candidate cis-antisense RNA and involvement of the corresponding chromosomal region in eno expression
In vitro cleavage assays using synthetic eno RNA and purified RNase III and RNase G did not generate the cleavage products identified by primer extension and S1 nuclease mapping analyses. Therefore, we suspected that other unknown factors were involved in RNase III- and/or RNase G-mediated cleavage of eno mRNA. First, we analysed cDNA from RNA-seq data of E. coli MG165538 strain using an Artemis annotation tool39 to examine whether antisense RNA is involved in eno mRNA cleavage because RNase III can process mRNAs when duplexed with antisense RNAs17,18. Interestingly, a large number of cDNAs were found that appear to be synthesised from RNA species complementary to that of the eno mRNA 5′ UTR (Supplementary Fig. 4). These RNA species appeared to be synthesised from DNA segments encompassing the 5′ end of the eno and the 3′ end of the pyrG coding regions. The 5′ and 3′ ends of these antisense RNAs that were deduced from cDNAs were located at positions +451, +272, −36, and −83 (designated as TIS1, TIS2, TIS3, and TIS4, respectively) from the transcriptional start site of the eno coding region, and at positions +1,419, +1,265, +736, and +696 from the transcriptional start site of the pyrG coding region (designated as TTS1, TTS2, TTS3, and TTS4, respectively) (Fig. 3a and Supplementary Fig. 4).
Restoration of RNase III-dependent regulation of eno-cat expression by putative cis-antisense RNA expression. (a) Schematic diagram showing the transcriptional initiation and termination sites (TISs and TTSs, respectively) of putative cis-antisense RNA. The 5′ and 3′ termini of putative cis-antisense RNAs that were inferred from cDNAs were located at positions +451, +272, −36, and −83 (designated as TIS1, TIS2, TIS3, and TIS4, respectively) from the start codon of the eno coding region, and at positions +1,419, +1,265, +736, and +696 from the start codon of the pyrG coding region (designated as TTS1, TTS2, TTS3, and TTS4, respectively). The secondary structure of the TTSs was inferred using the M-fold program. (b) Effects of alterations in the eno 5′ UTR on the regulation of eno-cat expression. Top: schematic representation of the region encompassing the eno and pyrG genes in W3110 WT and W3110PBADeno strains. Bottom: degree of chloramphenicol resistance of W3110 and W3110PBADeno strains harbouring pERS1. The W3110 and W3110PBADeno strains harbouring pERS1 were transformed with pPM30, pRNG3 (RNase G), or pRNC3 (RNase III). The transformants were grown to an OD600 of 0.6 in LB containing 1 mM IPTG and 0.01% arabinose, diluted, and spotted on LB agar containing 0.01% arabinose and 0 (Cm 0) or 100 (Cm 100) μg ml−1 chloramphenicol. (c) Schematic representation of pERS1-derived plasmids that additionally contain a DNA segment encompassing the eno and pyrG genes (from + 875 of the pyrG coding region to + 748, + 320, or + 200 of the eno coding region). (d) Minimal inhibitory concentrations (MICs) of MG1655 harbouring pERS1, pERS-AS748, pERS-AS320, or pERS-AS200 against chloramphenicol. Measurements of the MICs were performed independently, in triplicate, in LB containing various chloramphenicol concentrations; significant differences are indicated with different letters (one-way analysis of variance [ANOVA] followed by the Student–Newman–Keuls test, P < 0.0001). (e) Northern blot analysis of putative cis-antisense RNA. Total RNA was isolated from the MG1655 strains harbouring pERS1or pERS-AS748 and used for northern blot analysis. (f) In vitro cleavage analysis of the half-length synthetic eno mRNA with or without cis-antisense RNA. The 5′ end-labelled eno transcript (1 pmol) was incubated with purified RNase III (1 pmol) in cleavage buffer with or without MgCl2 at 37 °C. Cleavage products were identified using size markers generated by alkaline hydrolysis and RNase T1 digestion. RNase T1 cleaves 3′ of single-stranded G nucleotides and cleavage products are indicated in blue bold characters. (g) The predicted secondary structure of eno 5′ UTR (red) and cis-antisense RNA (blue) encompassing RNase III cleavage sites. This secondary structure was inferred using the M-fold program.
To determine whether antisense RNA is involved in RNase III- and/or RNase G-mediated regulation of eno expression, we generated an E. coli strain containing alterations in the 5′ UTR of eno to prevent the production of putative antisense RNA(s) by inserting an exogenous DNA fragment (PBAD promoter). This strain, designated W3110PBADeno, was derived from strain TM44724 by removing the cat gene and replacing a segment of the intergenic region between the eno and pyrG genes (−45 to −28 from the eno coding region) with an arabinose-inducible promoter (PBAD) and a transcriptional terminator (Fig. 3b). In W3110PBADeno cells, eno expression reached endogenous MG1655 cell levels when 0.001% arabinose was added to the culture (Supplementary Fig. 5).
The pERS1 plasmid was co-transformed with pPM30 (empty vector), pRNG3, or pRNC3 into W3110PBADeno cells and the degree of chloramphenicol resistance was compared among the strains. RNase III expression-dependent eno-cat activity was lost in W3110PBADeno cells harbouring pERS1 (Fig. 3b). Overexpression of RNase G still resulted in decreased eno-cat activity, implying that eno-cat mRNA synthesised in W3110PBADeno cells serves as an RNase G substrate. These results indicate that the endogenous eno 5′ UTR has a role in RNase III-mediated regulation of eno-cat expression, possibly through cis-antisense RNA synthesised from the opposite DNA strand of the endogenous eno and pyrG genes.
Restoration of RNase III-dependent regulation of eno-cat expression by putative cis-antisense RNA expression
The loss of RNase III-dependent regulation of eno-cat expression in W3110PBADeno cells harbouring pERS1 (Fig. 3b) suggested that production of putative cis-antisense RNA from the opposite strand of the eno and pyrG genes was perturbed in the W3110PBADeno strain. To test this possibility, we constructed a pERS1-derived plasmid that additionally contained a DNA segment from the eno and pyrG genes (from +875 of the pyrG coding region to +748 of the eno coding region) (Fig. 3c). The pERS-AS748 plasmid was transformed into W3110PBADeno cells and the degree of chloramphenicol resistance of the resulting strains was compared. The results showed that pERS-AS748 could restore RNase III-dependent eno-cat expression in W3110PBADeno cells, suggesting the ability of pERS-AS748 to express antisense RNA (Fig. 3d).
To characterise the main promoter and 5′ end of the putative antisense RNA, we constructed pERS-AS748-derived plasmids containing different DNA segments encompassing the eno and pyrG genes. These pERS-AS748 derivatives contained a DNA segment from +875 of the pyrG coding region to +320 (pERS-AS320) or +200 (pERS-AS200) of the eno coding region. These plasmids were co-transformed with pPM30, pRNG3, or pRNC3 into W3110PBADeno cells and the resulting strains were tested for the degree of resistance to chloramphenicol (Fig. 3d). The results showed that pERS-AS320, but not pERS-AS200, could restore RNase III-dependent regulation of eno-cat expression in W3110PBADeno cells, indicating that the promoter for antisense RNA expression was present in the region between +200 and +320 of the eno coding region, and that TIS2 was likely to be the transcriptional initiation site (Fig. 3d). Northern blot analysis of antisense RNA showed the existence of an RNA transcript approximately 500 nucleotides long in the WT strain harbouring pERS-AS748 (Fig. 3e). The transcript was not detected in the strains that harboured pERS1, probably because its endogenous level was low (Fig. 3e). Collectively, these results indicate that the putative cis-antisense RNA was synthesised from TIS2 and terminated at TTS1 (Supplementary Fig. 4).
Effects of putative antisense RNA on RNase III cleavage activity on eno mRNA
To investigate whether the putative cis-antisense RNA is required for RNase III cleavage of eno mRNA, we performed an in vitro cleavage assay using purified RNase III, synthetic eno mRNA (−96 to +448 of the eno CDS), and cis-antisense RNA transcripts. RNase III-mediated cleavage of the 5′ end-labelled synthetic eno transcripts generated one major and several minor cleavage products in a cis-antisense RNA-dependent manner (Fig. 3e). One major band corresponded to cleavage product 1 (Fig. 3f,g, and Supplementary Fig. 6) that was identified in the primer extension and S1 nuclease mapping analyses of eno mRNA (Fig. 2a,b). Other bands (2–4 in Fig. 2a,b) were not readily identified in these in vitro cleavage assays (Fig. 3f). These results indicate a possible function of the putative cis-antisense RNA for eno mRNA cleavage by RNase III.
Regulation of Eno expression by oxygen availability
Previous studies have shown that Eno expression is up-regulated when E. coli cells are grown anaerobically40,41,42. For this reason, we hypothesised that Eno expression can be up-regulated when E. coli cells are grown anaerobically since glycolysis plays a major role in energy production in the absence of oxygen. To investigate whether oxygen availability affects the RNase-mediated regulation of eno expression, we measured the steady-state levels of Eno, RNase G, and RNase III in WT cells under aerobic–anaerobic–aerobic alternating conditions43 (Fig. 4a). Western blot analysis showed that, compared to WT E. coli cells grown under the initial aerobic conditions (t0), Eno expression levels increased by 1.9-fold after shifting to anaerobic conditions (t3), coinciding with a 2.0-fold increase and an approximately 22% decrease in RNase III and RNase G expression levels, respectively (Fig. 4b). The alterations in the expressions levels of these proteins were restored when the cultures reverted to aerobic conditions (t5) (Fig. 4b). Steady-state levels of eno mRNA were well correlated with Eno expression levels under aerobic–anaerobic–aerobic alternating conditions, whereas those of cis-antisense RNA increased continuously during anaerobic–aerobic switching growth conditions (t2–t6) (Fig. 4c). Therefore, Eno expression levels are modulated by RNase G and RNase III in response to oxygen availability.
Expression levels of Eno, Rng, Rnc, and cis-antisense RNA depending on oxygen availability. (a) Schematic representation of the aerobic–anaerobic–aerobic alternating experiment. (b) Expression profiles of Eno, Rng, and Rnc depending on oxygen availability. WT MG1655 cells were cultured to each time point under aerobic or anaerobic conditions at 37 °C and then harvested for western blot analysis of Eno, Rng, and Rnc using protein-specific polyclonal antibodies. Blue, red, and yellow bars indicate the relative expression levels of Eno, Rng, and Rnc, respectively. The expression levels of Eno, Rng, and Rnc were compared by setting those of t0 to 1. Significant differences are indicated with different letters (one-way ANOVA followed by the Student–Newman–Keuls test; Small English letters indicate the difference from expression levels of Eno; Large English letters indicate the difference from expression levels of Rng; Greek symbols indicate the difference from expression levels of Rnc). (c) Analysis of cis-antisense RNA and eno mRNA expression in WT MG1655 cells using RT-PCR. The cDNA was synthesised from the total RNA extracted at each time point using the primers designed to bind cis-antisense RNA and eno mRNA. The PCR products were resolved in an 1.5% agarose gel. Significant differences are indicated with different letters (one-way ANOVA followed by the Student–Newman–Keuls test; Small English letters indicate the difference from expression levels of cis-antisense RNA; Large English letters indicate the difference of expression levels of eno mRNA). (d) Effect of Eno expression levels on the growth of W3110PBADeno cells. Cultures of W3110PBADeno cells were grown in LB medium containing 0.2% glucose under anaerobic conditions to the early log phase (OD600 = 0.05) and different concentrations of arabinose (0%, 0.01%, or 0.1%) were added to induce Eno synthesis. As a control, WT W3110 cells were grown in the same way described above and 0.1% arabinose was added. Cultures were further grown (5 h after induction) and were monitored by measuring the OD600. Cultures were harvested for western blot analysis of Eno using protein-specific polyclonal antibodies. The grey and blue bars indicate the OD600 values and the relative expression levels of Eno, respectively. The expression levels of Eno were compared by setting those of W3110 to 1. N.S., not significant. For (b,d), the S1 protein was used as an internal standard to evaluate the amount of cell extract in each lane. For (c), the rnpB mRNA was used as an internal standard to evaluate the amount of cell extract in each lane. For (b–d), the data are presented as means ± s. e. m. of at least three independent experiments.
We further investigated the effect of Eno expression levels on the growth of E. coli cells under low oxygen conditions by measuring growth yields of W3110PBADeno cells depending on the Eno expression levels. Cultures of W3110PBADeno cells were grown in Luria-Bertani (LB) medium containing 0.2% glucose under anaerobic conditions to the early log phase (OD600 = 0.05) and different concentrations of arabinose (0%, 0.01%, or 0.1%) were added to induce Eno synthesis. After 5 h, we measured the optical density (OD600) of the cultures and Eno expression levels in W3110PBADeno cells. The results showed that Eno expression levels were very well correlated with growth yields (Fig. 4d). These results suggested that Eno expression can affect the growth of E. coli cells when they are grown anaerobically because glycolysis plays a major role in energy production in the absence of oxygen.
Discussion
Enolase is highly conserved in organisms from bacteria to humans44. It is an enzyme that catalyzes a reaction of glycolysis and also, known to be associated with several biological and pathophysiological processes45,46,47. The presence of enolase in the RNase E-based RNA degradation machinery regulates cell morphology in E. coli under anaerobic conditions43. The negative regulation of eno expression by RNase G at the posttranscriptional level has been previously shown in E. coli1,27. In this study, we identified additional regulatory pathways for eno expression that involve RNase III and cis-antisense RNA. Our findings imply that RNase III-mediated cleavage of sense-antisense eno mRNA is required for efficient eno mRNA translation. This enhanced eno mRNA translation by RNase III cleavage of the primary eno transcript might result from the removal of a large hairpin in the upstream of a putative Shine-Dalgarno sequence and the start codon (Fig. 2c), allowing ribosomes more accessible to these sites. In addition, eno expression is further enhanced by accelerated RNase III cleavage of rng mRNA through increased RNase III activity in E. coli cells grown anaerobically, thereby contributing to the adjustment of physiological reactions in E. coli cells to oxygen-deficient metabolic modes (Fig. 5). In this regard, it is worthwhile to note that the abundance of mRNAs encoding other glycolytic enzymes was affected by the cellular concentrations of RNase G1 (e.g. glk, pgi, and tpiA) and RNase III4 (e.g. glk and ptsA), implying that these endoribonucleases, expression levels of which are regulated upon oxygen availability, play an important role in modulating the expression level of glycolytic enzymes.
A model for the molecular mechanism involved in RNase III- and RNase G-mediated regulation of eno expression in response to oxygen availability in E. coli. RNase III and RNase G coordinately regulate eno expression in response to oxygen availability in E. coli. Low oxygen activates RNase III activity and promotes the degradation of rng mRNA, leading to decreased expression of RNase G. Enhanced RNase III-mediated cleavage of primary eno mRNA transcripts promotes Eno protein production under anaerobic conditions. This RNase III-mediated processing involves cis-antisense RNA synthesised from the eno coding region to that of pyrG in the opposite direction of mRNA synthesis of these genes.
To understand the basis for the oxygen availability–dependent expression of the rnc gene, we analysed the promoter sequence. Bioinformatic analysis of these promoters (the online database Prodoric48) indicated the existence of sequences that were similar to the binding sequences for the oxygen-sensitive transcription factors fumarate nitrate reductase (FNR) and aerobic respiratory control (ArcA) in the promoter regions of rnc and rng (Supplementary Fig. 7). This complex regulatory system has been extensively studied in E. coli, where the DNA-binding proteins FNR and ArcA sense changes in oxygen availability and control the expression of many genes either alone or in cooperation with other regulators42,49,50,51,52,53. Further studies are required to characterise the molecular mechanisms for the regulation of RNase III expression in response to oxygen availability.
Although genome-wide analyses of RNase III cleavage sites implied the existence of cis-antisense RNAs that form duplexes with RNase III target mRNAs5,6. ArrS is the only cis-antisense RNA that has been experimentally suggested to have a role in sense mRNA processing17,18. Overexpression of ArrS resulted in the generation of gadE T3 mRNA, which contains a sequence complementary to ArrS in its 5′ UTR, leading to the production of more stable gadE T2 mRNA17,18. However, there is no direct experimental evidence showing ArrS-mediated RNase cleavage of gadE T3 mRNA. In the case of eno, genetic complementation and in vitro cleavage assays indicated the existence of cis-antisense RNA and its function in RNase III-mediated processing of the primary eno transcript (Fig. 3 and Supplementary Fig. 3). However, further studies are needed to unveil the detailed mechanisms underlying this endoribonucleases-mediated regulatory pathway for eno expression, including investigation of cis-antisense RNA biogenesis, sense-antisense RNA interactions, and enhanced translation of RNase III-processed eno mRNAs.
The involvement of both RNase III and RNase G in RNA function has been best characterised for 16S rRNA maturation in E. coli26,54. It was subsequently reported that incomplete processing of the 16S rRNA 5′ terminus by RNase G, which is down-regulated by increased RNase III activity on rng mRNA under aminoglycoside antibiotic stress, led to increased aminoglycoside resistance in E. coli cells34. A similar pathway involving RNase G and RNase III appears to be present in Salmonella Typhimurium34. These studies, together with our current study, suggest that bacteria commonly adopt endoribonuclease activity-mediated posttranscriptional regulatory pathways for rapid physiological adjustment to environmental changes.
Methods
Strains and plasmids
E. coli strains were grown at 37 °C in Luria-Bertani (LB) medium with or without 0.2% glucose supplemented with the appropriate antibiotics, under aerobic or anaerobic conditions. TM447 and W3110PBADeno strains were cultured in the same medium supplemented with 0.001% L-arabinose24. For anaerobic growth of E. coli cells, a 30 ml cylindrical bottle containing a sterilised stir bar was fully filled with LB medium with 0.2% glucose, sealed with sealing tape, and then cultured on a magnetic stirrer55. For aerobic–anaerobic–aerobic alternating experiments, the cells were grown aerobically to OD600 ~0.15 (t0) as described above. The culture was subsequently shifted to anaerobic conditions and then returned to aerobic conditions. Aliquots of the cells at 0 (t0), 60 (t1), 180 (t2), 360 (t3), 390 (t4), 510 (t5), and 570 (t6) min were drawn for western blots and reverse transcriptase-polymerase chain reaction (PCR).
Both Δrnc and Δrng have been previously described34,56. To generate Δrnc rng, the rng open reading frame was deleted in the Δrnc strain using the procedure described by Datsenko and Wanner57. The primers used were: 5′-rng-KO (5′-GTGAGAAAAGGGATAAACATGACGGCTAATTGTTAGTAAACGTAACGGTGTAGGCTGGAGCTGCTTC-3′) and 3′-rng-KO (5′-TTACATCATTACGACGTCAAACTGCTCCTGGTTATAGAGCGGTTCAATATTCCGGGGATCCGTCGACC-3′), and pKD1357 was used as a template. To construct the pERS1 plasmid expressing the eno-cat fusion, PstI and BamHI sites were created by overlap-extension PCR using the following primers: Eno1 (5′-ATCTGCAGGCGGCCGCTGTGGCGCTGATTACCGAGT-3′), Eno2 (5′-TATCCAGTGATTTTTTTCTCAGTCGGGTTACCACGGGAGT-3′), Eno3 (5′-GAGAAAAAAATCACTGGATA-3′), and RMC-MscI (5′-CCTTGTCGCCTTGCGTATAA-3′). Two PCR products were obtained using primers Eno1/Eno2 and Eno3/RMC-Msc I and the PCR products were combined and amplified using the Eno1 and RMC-Msc1 primers. The resulting fragment was cloned into the pACYC177 using PstI and BamHI. To construct the pERS2 plasmid expressing the eno-cat fusion that doesn’t contain the pyrG CDS, overlap-extension PCR was performed using primers Eno4 (5′-ATCTGCAGGTAAAAAAGTTAGAGCGGCA-3′)/Eno2 and Eno3/RMC-MscI in the same manner as pERS1. The pRNC3 plasmid was constructed by subcloning the NotI and XbaI fragment from pRNC1 containing the RNase III coding region into the same sites in pPM304. To express antisense RNA, a DNA fragment encoding a putative antisense eno RNA was amplified using the primers eno-nhe748R (5′-ATGCTAGCAAGCTGCGCAGTCCATCGCC-3′) and pyrG-nhe856F (5′-TAGCTAGCCCGGTAAGTGAAGTCACCAT-3′). The fragment was cloned into pERS1 using the NheI site, resulting in pERS-AS748. The pERS-AS320 and pERS-AS200 plasmids were constructed in a similar way, except that the 5′ end of the DNA segment was amplified using the eno-nhe 320 R primer (5′-TAGCTAGCCCGAATTTGGATTTGTTTTC-3′) for pERS-AS320 and eno-nhe200R (5′-GCGCTAGcGCTTTGGTTACGCCTTTACC-3′) for pERS-AS200. The mutant TM447 strain with a mutation in the eno 5′ UTR was obtained from Dr. H. Aiba24. The cat gene located upstream of eno in TM447 was removed using pCP20, resulting in W3110PBADeno. The pRNG3 and pPM30 plasmids have been previously described1.
Antibody purification and western blot analysis
Western blot analysis was carried out as previously described4,58. Polyclonal antibodies against Eno and RNase III were obtained from rabbits inoculated with purified His-tagged Eno and RNase III. Enolase and RNase III were purified from E. coli BL21(DE3) strains containing pET15b-enolase-His and pET15b-RNase III-His, respectively, using Ni-NTA agarose (Qiagen, Hilden, Germany). The proteins were eluted from columns using 125 mM imidazole and then concentrated and stored as previously described59,60. Polyclonal antibodies against RNase G, RNase E, and S1 were obtained from Dr. Stanley N. Cohen. Western blot images were obtained using an Amersham Imager 600 (GE Healthcare, Buckinghamshire, UK) and quantified using Quantity One (Bio-Rad Laboratories, Hercules, CA, USA). The ribosomal S1 protein was used as the control.
Isolation of total RNA and reverse tanscriptase (RT)-PCR
Total cellular RNA was extracted from the cultures grown to an OD600 of 0.6 using an RNeasy mini prep kit (Qiagen). Following confirmation of the quality and quantity of the extracted total RNA with a Nanodrop 2000 instrument (Thermo Fisher Scientific, Waltham, MA, USA) the RT-PCR analysis was performed as previously described61. Briefly, cDNA was synthesised from 1 μg of total RNA using the PrimeScript 1st strand cDNA Synthesis Kit system for RT-PCR (Takara, Otsu, Japan) according to the manufacturer’s instructions. PCR primer sequences were designed according to the eno, cis-antisense RNA, and rnpB (as a standard control) genes in GenBank and products were obtained using primers, Eno RT F (5′-ATGTCCAAAATCGTAAAAAT-3′) and Eno RT R (5′- CATCTTTGCCAATCAGCGCC-3′) for eno mRNA; antisense RNA F (5′-TCACGGGAACCAGTAGAAGC-3′) and antisense RNA R (5′-TACGCGTTGTTTGTCTGGAG-3′) for cis-antisense RNA; and rnpB RT F (5′- TTGCTCCGGGTGGAGTTTAC-3′) and rnpB RT R (5′-GTGCAACAGAGAGCAAACCG-3′) for rnpB mRNA.
Measurement of minimal inhibitory concentrations (MICs)
The MICs were measured as previously described62. Briefly, overnight cultures grown in LB medium supplemented with the appropriate antibiotics were diluted 1:100 in the same medium and incubated for 2 h. At an optical density 0.6 at 600 nm (OD600), 1 × 102–106 cells were spotted on LB agar plates containing different concentrations of antibiotics, or approximately 1 × 104 cells were added to the same medium containing increasing concentrations of antibiotics. The cultures were grown for an additional 12 h and the lowest antibiotic concentrations that completely inhibited growth were designated as the MICs.
Primer extension analysis
The procedure for primer extension analysis has been described previously63. Briefly, total RNAs were hybridised with a 5′ end 32P-labelled primer (eno + 60R: 5′-AGTCGGGTTACCACGGGAGT-3′) at 65 °C for 15 min, slowly cooled to 42 °C for 2 h, and then incubated at 42 °C for 1 h with AMV reverse transcriptase for cDNA synthesis (New England Biolabs, Ipswich, MA, USA). The amplicons were separated on 9% polyacrylamide gels containing 8 M urea and a sequencing reaction was performed and used as a molecular weight marker. Autoradiography was generated using a Packard Cyclone Phosphor Imager (PerkinElmer, Waltham, MA, USA) as previously described64.
In vitro cleavage assay
Synthetic RNAs containing full-length and antisense eno sequences were synthesised from PCR DNA products using a MEGAshortscript T7 Kit (Thermo Fisher Scientific) according to the manufacturer’s instructions. The DNA products were PCR-amplified from MG1655 genomic DNA using the following primers: T7-eno −96F (5′-TAATACGACTCACTATAGGGCGAAGTAAGTAAAAAAGTT-3′), eno 448R (5′-TCGGAACCGGCATAGAGTAT-3′) for the full-length transcript, and T7-eno 273R (5′-CT TAATACGACTCACTATAGGGTCAATGCCAGCCTGATCTT-3′) and pyrG1153F (5′-CCACCTGCATACCCAGGCAA-3′) for antisense RNA. The RNAs were purified using a MEGAclear Kit and labelled at the 5′ end using [γ-32P] ATP. An aliquot (4 pmol) of 32P-5′ end labelled RNA was incubated with 1 ng of purified RNase III with 0.25 mg ml−1 yeast tRNA (Thermo Fisher Scientific), 20 U of RNase inhibitor (Takara), and RNase III cleavage buffer, with or without MgCl2 65. Samples were removed at the indicated time points and separated on an 8% polyacrylamide gel containing 8 M urea.
S1 nuclease mapping
Probes for S1 mapping were prepared by PCR amplification using the primers eno1 (5′-ATCTGCAGGCGGCCGCTGTGGCGCTGATTACCGAGT-3′) and eno10R (5′-TTTTGGACATTAGGTTTTCC-3′), after labelling of the 5′ ends of the primers with [γ-32P]ATP using T4 polynucleotide kinase at 37 °C for 1 h. The labelled eno 10 R primer and the unlabelled Eno1 primer generated a 500 bp eno probe. Total RNA (50 μg) was coprecipitated with 0.2 μg of the 5′ end 32P-labelled DNA probe and washed with 80% EtOH. The dried pellets were resuspended in 20 μl of hybridisation buffer and annealed at 52 °C overnight after denaturing at 80 °C for 10 min. After overnight incubation, S1 digestions were performed by adding 300 µl of an S1 nuclease mix (Promega, Madison, WI, USA) and incubating at 37 °C for 1 h. The protected fragments were electrophoresed alongside the sequence ladders obtained with the labelled primer used for probe preparation.
Northern blot analysis
E. coli MG1655 (WT, Δrnc, and Δrng) cells harbouring pERS1 and MG1655 WT harbouring pERS-AS748 were grown at 37 °C to an OD600 of 0.6. Total RNA samples were prepared from the cultures using an RNeasy mini prep kit (Qiagen). Total RNA samples (40 µg) were denatured at 65 °C for 15 min in an equal volume of formamide loading buffer and separated by electrophoresis on a 1.2% GTG agarose gel containing 0.66 M formaldehyde. The gels were blotted onto a nylon membrane (Hybond-XL blotting membrane, Amersham) using Turboblotter (GE Healthcare). The pyrG1608F 5′ end 32P-labelled oligo probes (5′-CCCGCTGTTTGCAGGCTTTGTG-3′) were hybridised. The procedure for northern blot analysis was as previously described1. The size markers generated by internally labeled transcripts.
Quantification and statistical analyses
All the statistical details of the experiments are included in the figure legends. Multiple comparison analysis was performed by the Student–Newman–Keuls test using SAS v.9.2 (SAS Institute, Cary, NC, USA), and the Student’s t-test was used for comparisons with controls using SigmaPlot (Systat Software, San Jose, CA, USA). The data are presented as means ± s. e. m., and P-values < 0.05 were considered significant66,67.
Data availability
The datasets generated during and/ or analysed during the current study are available from the corresponding author on request.
References
Lee, K., Bernstein, J. A. & Cohen, S. N. RNase G complementation of rne null mutation identifies functional interrelationships with RNase E in Escherichia coli. Mol Microbiol 43, 1445–1456 (2002).
Lee, K. et al. RraA. a protein inhibitor of RNase E activity that globally modulates RNA abundance in E. coli. Cell 114, 623–634 (2003).
Bernstein, J. A., Khodursky, A. B., Lin, P. H., Lin-Chao, S. & Cohen, S. N. Global analysis of mRNA decay and abundance in Escherichia coli at single-gene resolution using two-color fluorescent DNA microarrays. Proc Natl Acad Sci USA 99, 9697–9702, https://doi.org/10.1073/pnas.112318199 (2002).
Sim, S. H. et al. Escherichia coli ribonuclease III activity is downregulated by osmotic stress: consequences for the degradation of bdm mRNA in biofilm formation. Molecular Microbiology 75, 413–425, https://doi.org/10.1111/j.1365-2958.2009.06986.x (2010).
Gordon, G. C., Cameron, J. C. & Pfleger, B. F. RNA Sequencing Identifies New RNase III Cleavage Sites in Escherichia coli and Reveals Increased Regulation of mRNA. Mbio 8, https://doi.org/10.1128/mBio.00128-17 (2017).
Altuvia, Y. et al. In vivo cleavage rules and target repertoire of RNase Ill in Escherichia coli (vol 46, pg 10380, 2018). Nucleic Acids Res 46, 10530–10531, https://doi.org/10.1093/nar/gky816 (2018).
Lim, B. et al. Regulation of Escherichia coli RNase III activity. J Microbiol 53, 487–494, https://doi.org/10.1007/s12275-015-5323-x (2015).
Kaberdin, V. R. et al. The endoribonucleolytic N-terminal half of Escherichia coli RNase E is evolutionarily conserved in Synechocystis sp. and other bacteria but not the C-terminal half, which is sufficient for degradosome assembly. Proc Natl Acad Sci USA 95, 11637–11642, https://doi.org/10.1073/pnas.95.20.11637 (1998).
Miczak, A., Kaberdin, V. R., Wei, C. L. & Lin-Chao, S. Proteins associated with RNase E in a multicomponent ribonucleolytic complex. Proc Natl Acad Sci USA 93, 3865–3869, https://doi.org/10.1073/pnas.93.9.3865 (1996).
Py, B., Higgins, C. F., Krisch, H. M. & Carpousis, A. J. A DEAD-box RNA helicase in the Escherichia coli RNA degradosome. Nature 381, 169–172, https://doi.org/10.1038/381169a0 (1996).
Gao, J. et al. Differential modulation of E. coli mRNA abundance by inhibitory proteins that alter the composition of the degradosome. Mol Microbiol 61, 394–406, https://doi.org/10.1111/j.1365-2958.2006.05246.x (2006).
Seo, S. et al. RraAS1 inhibits the ribonucleolytic activity of RNase ES by interacting with its catalytic domain in Streptomyces coelicolor. Journal of Microbiology 55, 37–43, https://doi.org/10.1007/s12275-017-6518-0 (2017).
Park, N. et al. Crystal structure of Streptomyces coelicolor RraAS2, an unusual member of the RNase E inhibitor RraA protein family. J Microbiol 55, 388–395, https://doi.org/10.1007/s12275-017-7053-8 (2017).
De Lay, N., Schu, D. J. & Gottesman, S. Bacterial small RNA-based negative regulation: Hfq and its accomplices. J Biol Chem 288, 7996–8003, https://doi.org/10.1074/jbc.R112.441386 (2013).
Storz, G., Vogel, J. & Wassarman, K. M. Regulation by Small RNAs in Bacteria: Expanding Frontiers. Mol Cell 43, 880–891, https://doi.org/10.1016/j.molcel.2011.08.022 (2011).
Thomason, M. K. et al. Global Transcriptional Start Site Mapping Using Differential RNA Sequencing Reveals Novel Antisense RNAs in Escherichia coli. Journal of Bacteriology 197, 18–28, https://doi.org/10.1128/Jb.02096-14 (2015).
Aiso, T., Murata, M. & Gamou, S. Transcription of an antisense RNA of a gadE mRNA is regulated by GadE, the central activator of the acid resistance system in Escherichia coli. Genes Cells 16, 670–680, https://doi.org/10.1111/j.1365-2443.2011.01516.x (2011).
Aiso, T., Kamiya, S., Yonezawa, H. & Gamou, S. Overexpression of an antisense RNA, ArrS, increases the acid resistance of Escherichia coli. Microbiology 160, 954–961, https://doi.org/10.1099/mic.0.075994-0 (2014).
Wen, J. & Fozo, E. M. sRNA Antitoxins: More than One Way to Repress a Toxin. Toxins 6, 2310–2335, https://doi.org/10.3390/toxins6082310 (2014).
Lee, E. J. & Groisman, E. A. An antisense RNA that governs the expression kinetics of a multifunctional virulence gene. Mol Microbiol 76, 1020–1033, https://doi.org/10.1111/j.1365-2958.2010.07161.x (2010).
Lybecker, M., Zimmermann, B., Bilusic, I., Tukhtubaeva, N. & Schroeder, R. The double-stranded transcriptome of Escherichia coli. Proc Natl Acad Sci USA 111, 3134–3139, https://doi.org/10.1073/pnas.1315974111 (2014).
Lioliou, E. et al. Global regulatory functions of the Staphylococcus aureus endoribonuclease III in gene expression. PLoS Genet 8, e1002782, https://doi.org/10.1371/journal.pgen.1002782 (2012).
Tamura, M., Moore, C. J. & Cohen, S. N. Nutrient Dependence of RNase E Essentiality in Escherichia coli. Journal of Bacteriology 195, 1133–1141, https://doi.org/10.1128/Jb.01558-12 (2013).
Morita, T., Kawamoto, H., Mizota, T., Inada, T. & Aiba, H. Enolase in the RNA degradosome plays a crucial role in the rapid decay of glucose transporter mRNA in the response to phosphosugar stress in Escherichia coli. Mol Microbiol 54, 1063–1075, https://doi.org/10.1111/j.1365-2958.2004.04329.x (2004).
Manow, R. et al. Partial deletion of rng (RNase G)-enhanced homoethanol fermentation of xylose by the non-transgenic Escherichia coli RM10. J Ind Microbiol Biotechnol 39, 977–985, https://doi.org/10.1007/s10295-012-1100-6 (2012).
Li, Z., Pandit, S. & Deutscher, M. P. RNase G (CafA protein) and RNase E are both required for the 5′ maturation of 16S ribosomal RNA. Embo J 18, 2878–2885, https://doi.org/10.1093/emboj/18.10.2878 (1999).
Kaga, N., Umitsuki, G., Nagai, K. & Wachi, M. RNase G-dependent degradation of the eno mRNA encoding a glycolysis enzyme enolase in Escherichia coli. Biosci Biotechnol Biochem 66, 2216–2220, https://doi.org/10.1271/bbb.66.2216 (2002).
Sakai, T., Nakamura, N., Umitsuki, G., Nagai, K. & Wachi, M. Increased production of pyruvic acid by Escherichia coli RNase G mutants in combination with cra mutations. Appl Microbiol Biot 76, 183–192, https://doi.org/10.1007/s00253-007-1006-9 (2007).
Aristarkhov, A., Mikulskis, A., Belasco, J. G. & Lin, E. C. Translation of the adhE transcript to produce ethanol dehydrogenase requires RNase III cleavage in Escherichia coli. J Bacteriol 178, 4327–4332, https://doi.org/10.1128/jb.178.14.4327-4332.1996 (1996).
Kristiansen, K. I., Weel-Sneve, R., Booth, J. A. & Bjoras, M. Mutually exclusive RNA secondary structures regulate translation initiation of DinQ in Escherichia coli. RNA 22, 1739–1749, https://doi.org/10.1261/rna.058461.116 (2016).
Kim, K. S., Manasherob, R. & Cohen, S. N. YmdB: a stress-responsive ribonuclease-binding regulator of E. coli RNase III activity. Genes Dev 22, 3497–3508, https://doi.org/10.1101/gad.1729508 (2008).
Kavalchuk, K., Madhusudan, S. & Schnetz, K. RNase III initiates rapid degradation of proU mRNA upon hypo-osmotic stress in Escherichia coli. RNA Biol 9, 98–109, https://doi.org/10.4161/rna.9.1.18228 (2012).
Lim, B. & Lee, K. Stability of the osmoregulated promoter-derived proP mRNA is posttranscriptionally regulated by RNase III in Escherichia coli. J Bacteriol 197, 1297–1305, https://doi.org/10.1128/JB.02460-14 (2015).
Song, W. et al. Antibiotic stress-induced modulation of the endoribonucleolytic activity of RNase III and RNase G confers resistance to aminoglycoside antibiotics in Escherichia coli. Nucleic Acids Res 42, 4669–4681, https://doi.org/10.1093/nar/gku093 (2014).
Cohen, S. N., Chang, A. C., Boyer, H. W. & Helling, R. B. Construction of biologically functional bacterial plasmids in vitro. Proc Natl Acad Sci USA 70, 3240–3244, https://doi.org/10.1073/pnas.70.11.3240 (1973).
Shimada, T., Fujita, N., Maeda, M. & Ishihama, A. Systematic search for the Cra-binding promoters using genomic SELEX system. Genes Cells 10, 907–918, https://doi.org/10.1111/j.1365-2443.2005.00888.x (2005).
Olvera, L. et al. Transcription analysis of central metabolism genes in Escherichia coli. Possible roles of sigma38 in their expression, as a response to carbon limitation. PLoS One 4, e7466, https://doi.org/10.1371/journal.pone.0007466 (2009).
Thomason, M. K. & Storz, G. Bacterial Antisense RNAs: How Many Are There, and What Are They Doing? Annual Review of Genetics, Vol 44 44, 167–188, https://doi.org/10.1146/annurev-genet-102209-163523 (2010).
Rutherford, K. et al. Artemis: sequence visualization and annotation. Bioinformatics 16, 944–945, https://doi.org/10.1093/bioinformatics/16.10.944 (2000).
Smith, M. W. & Neidhardt, F. C. Proteins induced by anaerobiosis in Escherichia coli. J Bacteriol 154, 336–343 (1983).
Gonzalez, J. E., Long, C. P. & Antoniewicz, M. R. Comprehensive analysis of glucose and xylose metabolism in Escherichia coli under aerobic and anaerobic conditions by (13)C metabolic flux analysis. Metab Eng 39, 9–18, https://doi.org/10.1016/j.ymben.2016.11.003 (2017).
Tamarit, J., Cabiscol, E. & Ros, J. Identification of the major oxidatively damaged proteins in Escherichia coli cells exposed to oxidative stress. J Biol Chem 273, 3027–3032, https://doi.org/10.1074/jbc.273.5.3027 (1998).
Murashko, O. N. & Lin-Chao, S. Escherichia coli responds to environmental changes using enolasic degradosomes and stabilized DicF sRNA to alter cellular morphology. Proc Natl Acad Sci USA 114, E8025–E8034, https://doi.org/10.1073/pnas.1703731114 (2017).
Lopez-Lopez, M. J. et al. Biochemical and Biophysical Characterization of the Enolase from Helicobacter pylori. Biomed Res Int 2018, 9538193, https://doi.org/10.1155/2018/9538193 (2018).
Pancholi, V. Multifunctional alpha-enolase: its role in diseases. Cell Mol Life Sci 58, 902–920 (2001).
Kim, J. W. & Dang, C. V. Multifaceted roles of glycolytic enzymes. Trends Biochem Sci 30, 142–150, https://doi.org/10.1016/j.tibs.2005.01.005 (2005).
Capello, M., Ferri-Borgogno, S., Cappello, P. & Novelli, F. Alpha-Enolase: a promising therapeutic and diagnostic tumor target. FEBS J 278, 1064–1074, https://doi.org/10.1111/j.1742-4658.2011.08025.x (2011).
Eckweiler, D., Dudek, C. A., Hartlich, J., Brotje, D. & Jahn, D. PRODORIC2: the bacterial gene regulation database in 2018. Nucleic Acids Res 46, D320–D326, https://doi.org/10.1093/nar/gkx1091 (2018).
Chattopadhyay, S., Wu, Y. & Datta, P. Involvement of Fnr and ArcA in anaerobic expression of the tdc operon of Escherichia coli. J Bacteriol 179, 4868–4873, https://doi.org/10.1128/jb.179.15.4868-4873.1997 (1997).
Crofts, A. A. et al. Enterotoxigenic E. coli virulence gene regulation in human infections. Proc Natl Acad Sci USA 115, E8968–E8976, https://doi.org/10.1073/pnas.1808982115 (2018).
Bearson, S. M., Albrecht, J. A. & Gunsalus, R. P. Oxygen and nitrate-dependent regulation of dmsABC operon expression in Escherichia coli: sites for Fnr and NarL protein interactions. BMC Microbiol 2, 13 (2002).
Lazazzera, B. A., Beinert, H., Khoroshilova, N., Kennedy, M. C. & Kiley, P. J. DNA binding and dimerization of the Fe-S-containing FNR protein from Escherichia coli are regulated by oxygen. J Biol Chem 271, 2762–2768, https://doi.org/10.1074/jbc.271.5.2762 (1996).
Park, D. M. & Kiley, P. J. The influence of repressor DNA binding site architecture on transcriptional control. Mbio 5, e01684–01614, https://doi.org/10.1128/mBio.01684-14 (2014).
Gegenheimer, P., Watson, N. & Apirion, D. Multiple pathways for primary processing of ribosomal RNA in Escherichia coli. J Biol Chem 252, 3064–3073 (1977).
Jaejin Lee, D.-H. L. Jeon, C. O. & Lee, K. RNase G controls tpiA mRNA abundance in response to oxygen availability in Escherichia coli. J Microbiol 57, https://doi.org/10.1007/s12275-019-9354-6 (2019).
Lim, B. et al. RNase III controls the degradation of corA mRNA in Escherichia coli. J Bacteriol 194, 2214–2220, https://doi.org/10.1128/JB.00099-12 (2012).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97, 6640–6645, https://doi.org/10.1073/pnas.120163297 (2000).
Kime, L., Jourdan, S. S. & McDowall, K. J. Identifying and characterizing substrates of the RNase E/G family of enzymes. Methods Enzymol 447, 215–241, https://doi.org/10.1016/S0076-6879(08)02212-X (2008).
Amarasinghe, A. K., Calin-Jageman, I., Harmouch, A., Sun, W. & Nicholson, A. W. Escherichia coli ribonuclease III: affinity purification of hexahistidine-tagged enzyme and assays for substrate binding and cleavage. Methods Enzymol 342, 143–158 (2001).
Sun, J. et al. Overexpression and characterization of a novel cold-adapted and salt-tolerant GH1 beta-glucosidase from the marine bacterium Alteromonas sp. L82. J Microbiol 56, 656–664, https://doi.org/10.1007/s12275-018-8018-2 (2018).
Baek, J., Choi, E. & Lee, E. J. A rule governing the FtsH-mediated proteolysis of the MgtC virulence protein from Salmonella enterica serovar Typhimurium. J Microbiol 56, 565–570, https://doi.org/10.1007/s12275-018-8245-6 (2018).
Kim, H. M. et al. Functional relationships between the AcrA hairpin tip region and the TolC aperture tip region for the formation of the bacterial tripartite efflux pump AcrAB-TolC. J Bacteriol 192, 4498–4503, https://doi.org/10.1128/JB.00334-10 (2010).
Song, W. et al. Divergent rRNAs as regulators of gene expression at the ribosome level. Nat Microbiol 4, 515–526, https://doi.org/10.1038/s41564-018-0341-1 (2019).
Cho, M. et al. Vacuolar zinc transporter Zrc1 is required for detoxification of excess intracellular zinc in the human fungal pathogen Cryptococcus neoformans. J Microbiol 56, 65–71, https://doi.org/10.1007/s12275-018-7475-y (2018).
Sim, M. et al. Two tandem RNase III cleavage sites determine betT mRNA stability in response to osmotic stress in Escherichia coli. PLoS One 9, e100520, https://doi.org/10.1371/journal.pone.0100520 (2014).
Ramasamy, S., Arumugam, A. & Chandran, P. Optimization of Enterobacter cloacae (KU923381) for diesel oil degradation using response surface methodology (RSM). J Microbiol 55, 104–111, https://doi.org/10.1007/s12275-017-6265-2 (2017).
Yi, Y. J. et al. Potential use of lactic acid bacteria Leuconostoc mesenteroides as a probiotic for the removal of Pb(II) toxicity. J Microbiol 55, 296–303, https://doi.org/10.1007/s12275-017-6642-x (2017).
Acknowledgements
We are grateful to Dr. Hiroji Aiba for providing the TM447 strain. This research was supported by grants from the National Research Foundation (NRF) (NRF-2018R1A5A1025077 to KL, NRF-2019R1I1A1A01063517 to ML, NRF-2017R1D1A1B03028392 to MS and NRF-2019R1I1A1A01063708 to MJ).
Author information
Authors and Affiliations
Contributions
N.-C.H., J.-C.C. and K.L. conceived the research plan; M.L., M.J., M.S., S.-H.S. and H.-L.K. performed most of the experiments and analysed the data; J.L. and M.R. performed statistical analysis. J.-H.Y. and Y.H. discussed the results and analysed the data. K.L. supervised the project. N.-C.H., J.-C.C. and K.L. wrote the article with contributions of all the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, M., Joo, M., Sim, M. et al. The coordinated action of RNase III and RNase G controls enolase expression in response to oxygen availability in Escherichia coli. Sci Rep 9, 17257 (2019). https://doi.org/10.1038/s41598-019-53883-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-53883-y
- Springer Nature Limited
This article is cited by
-
Comparative Transcriptomic Analysis of Flagellar-Associated Genes in Salmonella Typhimurium and Its rnc Mutant
Journal of Microbiology (2024)
-
The effect of two ribonucleases on the production of Shiga toxin and stx-bearing bacteriophages in Enterohaemorrhagic Escherichia coli
Scientific Reports (2021)
-
Trans-acting regulators of ribonuclease activity
Journal of Microbiology (2021)