Abstract
Small ruminant lentiviruses (SRLVs) are found in sheep in Germany and Iran. SRLVs have been classified into four genotypes: A–C and E. Genotype A has been subdivided into 20 subtypes. Previous studies suggested that, first, the ancestors of genotype A are those SRLVs found in Turkey, second, the evolution of SRLVs is related to the domestication process, and, third, SRLV infection was first observed in sheep in Iceland and the source of that infection was a flock imported from Germany. This study generated, for the first time, partial SRLV sequence data from German and Iranian sheep, enhancing our knowledge of the genetic and evolutionary relationships of SRLVs, and their associations with the domestication process. Based on 54 SRLV sequences from German and Iranian sheep, our results reveal: (1) SRLV subtypes A4, A5, A11, A16 and A21 (new) are found in German sheep and A22 (new) in Iranian sheep. (2) Genotype A has potentially an additional ancestor (A22), found in Iran, Lebanon and Jordan. (3) Subtype A22 is likely an old version of SRLVs. (4) The transmission routes of some SRLVs are compatible with domestication pathways. (5) This study found no evidence of Icelandic subtype A1 in German sheep.
Similar content being viewed by others
Introduction
Small ruminant lentiviruses (SRLVs), which comprise maedi-visna virus (MVV) and caprine arthritis encephalitis virus (CAEV), belong to the genus Lentivirus and the family Retroviridae. SRLVs can cause progressive multisystem disease in sheep involving lungs, joints, mammary gland and the central nervous system1. There is no cure or vaccine available against SRLV infection. SRLV-related diseases are distributed worldwide among sheep and goats, resulting in considerable economic losses2.
Like in other lentiviruses, the SRLV genome includes three structural genes, coding for the group-specific antigens (gag), the polymerase (pol) and the envelope (env). The gag gene encodes the matrix (MA) protein (p17), capsid (CA) protein (p25) and nucleocapsid (NC) protein (p14)3. Both gag and pol genes are relatively conserved, and phylogenetic analyses of SRLVs have been established based on these two genes4.
SRLV isolates can be classified into four genotypes, A–C and E4,5,6. Genotypes A and B are widespread and refer to MVV-like and CAEV-like viruses, respectively. MVV-like and CAEV-like strains have been first described in sheep and goats, respectively, and considered strictly host-specific for a long time. However, there are nowadays several studies indicating that most strains can cross the species barrier (reviewed by Minardi da Cruz et al.7). Genotype A is the most heterogeneous group and has so far been subdivided into 20 subtypes, A1 to A204,8,9,10,11,12,13,14,15. Two recently published studies have to be noted, one by Olech et al.13 that defines SRLV subtype A18, and the other by Colitti et al.15 that defines SRLV subtypes A18 and A19. In the present study, the SRLVs found by Colitti et al.15 are renamed from ‘A18’ to ‘A19’ and from ‘A19’ to ‘A20’. Genotype B contains three subtypes, B1 to B34,16. SRLVs restricted to certain geographical areas have been assigned to other genotypes: genotype C is divided into two subtypes and refers to Norwegian isolates15,17,18, genotype D was found in few isolates originating from Switzerland and Spain, but they are now re-classified as genotype A4,6,19; genotype E comprises subtypes E1 and E2 and was isolated in Italy9,20.
Lentiviruses have a deep evolutionary history and have evolved alongside their mammalian hosts21,22,23,24,25. During the process of evolution, exogenous retroviruses (e.g. Jaagsiekte sheep retrovirus; a retrovirus that has many similarities with SRLVs and imposes pulmonary adenocarcinoma disease in sheep) have inserted into the germline of the infected host, leading to endogenous retroviruses (ERVs)26. In a study by Chessa et al.27, the presence of six variants of endogenous Jaagsiekte retrovirus (enJSRVs) was examined in 65 global domestic sheep, to investigate the history of sheep domestication. During the first wave of domestication, early domesticated sheep, which were morphologically wild but managed, appeared in the ancient Fertile Crescent region, including parts of Iran, Iraq, Turkey, Syria and Jordan, approximately 10,000 to 8,000 years before present (YBP)28,29. The domestication of goats started in the same region between 500 and 1,000 years earlier at about 11,000 YBP28,30. During the second wave of domestication, sheep with ‘modern’ features, typical of present-day breeds (e.g. with woolly fleeces and polled), appeared in West Asia and other areas in the world at approximately 6,000 YBP27. Host molecular genetic data combined with archaeological evidence indicates that sheep were distributed from the Fertile Crescent to the West and East likely during both waves of domestication27,31,32. For instance, during the first wave of domestication, the European mouflon migrated through the Mediterranean Basin to the islands of Corsica and Sardinia at around 7,000 YBP21,27,32,33. During the second wave, sheep with improved production traits were introduced from the East into Europe at the beginning of the 5th millennium YBP34.
In parallel with researches on host evolution, recent studies have demonstrated the usefulness of pathogens to elucidate the evolution of their hosts across time and location21,35,36,37. In this respect, besides enJSRVs27, investigating the phylogeny of SRLVs has the potential to enhance our knowledge of sheep and goat domestication16,20,21. The identification of SRLV subtype B3 in sheep/goats from Italy and Turkey as well as the finding that some bulk milk samples from Turkish sheep and goats were reactive against antigen derived from genotype E (a genotype found in Sardinia and other parts of Italy), supports the hypothesis of migration of domesticated sheep from the Fertile Crescent into the Mediterranean Basin during the Neolithic age16. However, phylogenetic studies involving SRLV sequences from other regions of the Fertile Crescent (except Turkey), which would potentially enhance our knowledge of the domestication process within the domestication origin itself, have been absent until now. While few SRLV sequences from Jordan and Lebanon are available in the database, no SRLV sequence information has been published from Iran, Iraq and Syria.
Historically, it has been suggested that maedi-visna was introduced to Iceland through German sheep2,6,38,39,40. In the early 1930s, a set of Karakul sheep (n = 20) was imported from Halle in Germany to Iceland. After several months, signs of maedi-visna were observed in some Icelandic sheep flocks, which hosted German Karakul rams38. The German Karakul flock originated from Astrakhan (Russia)41. The Icelandic SRLV strain (subtype A1) was first characterised by Sigurdsson et al.38 about 15 years after the observation of the first signs of maedi-visna in the Icelandic sheep flocks. Subtype A1 had already been detected in many countries6. However, German SRLVs have not yet been characterised.
In Iran, SRLV infection was first diagnosed in sheep using histopathological methods in 200142. Following this first survey, SRLV infection was reported in different parts of Iran, using serological methods or PCR techniques43,44,45,46.
Currently, researchers have suggested three domestication pathways, from West Asia (Iran and Turkey) to Europe and Africa32,47. Germany is located on the end terminal of the Danubian pathway, Italy is located on the northern Mediterranean pathway and Morocco is located on the southern Mediterranean pathway47. In this respect, Iran belongs to the ancient Fertile Crescent region, where the initial domestication of sheep and goats occurred28,29,30. Also, the geographical position of Germany for investigating domestication events is important, as it has long been connected with the ancient Fertile Crescent via the Danube River32,47. This study aimed at generating, for the first time, SRLV sequence data from German and Iranian sheep, for which no such data has been available until now. The results were expected to enhance our knowledge of the genetic and evolutionary relationships of German and Iranian SRLVs, as compared to those from other countries, as well as their associations with the domestication process.
Results
SRLV sequences from German and Iranian sheep flocks
All DNA samples (n = 54), were successfully amplified using gag-pol primers. None of the sequences showed evidence of recombination. Information on gag-pol SRLV sequences collected from different sheep flocks in Germany and Iran, and the accession numbers are given in Table 1. Notably, due to unmatched sequence data for the alignment (Supplementary Fig. S1 online), genetic and phylogenetic analyses on gag-pol sequences correspond to a part of the gag gene. Based on initial analyses, 23 out of the 54 gag sequences were selected for phylogenetic and genetic analyses. Of the 23 selected sequences, 17 gag sequences belonged to 13 German sheep flocks and 6 corresponded to 6 Iranian sheep flocks.
Phylogenetic analysis of SRLVs based on gag fragment
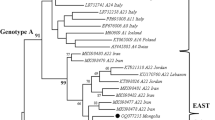
The constructed phylogenetic tree with gag sequences is shown in Fig. 1. According to the phylogenetic analysis, the gag sequences from Germany (n = 17), which were distributed in five distinct clusters, were strongly related to genotype A. The tree showed that these sequences were affiliated to subtypes A4 (flock 12), A5 (flocks 2–4), A11 (flocks 8, 10 and 13), A16 (flock 1) and potentially to a new subtype, which could be tentatively named A21 (flocks 3 and 5–11). Not all of these clusters were supported with high bootstrap values. The six Iranian SRLV sequences belonged to genotype A with a bootstrap value of 96%. Furthermore, the Iranian SRLV sequences clustered together with the Jordanian (KT898826 and KT921318) and Lebanese (KU170760) SRLV prototypes. This cluster was tentatively named A22.
Phylogenetic tree indicates SRLV subtypes and their differing geographical distribution (country). The analysis was performed using the Maximum Likelihood method and was based on the Tamura-Nei model61. The analysis involved a total of 399 bp from 65 nucleotide sequences: 17 German gag sequences (labelled by solid red circles), six Iranian sequences (labelled by solid blue circles), two Jordanian SRLV sequences (KT898826 and KT921318), a Lebanese SRLV sequence (KU170760) and 39 reference SRLV sequences originating from different geographical areas (retrieved from the GenBank database). The position of the analysed fragments was related to coding regions of the capsid gene (p25) and the nucleocapsid gene (p14) located on the gag fragment (nucleotide: 1114–1506; numbering according to prototype strain K151462. The difference in the evolutionary rate among sites was considered using the discrete gamma distribution ( + G parameter = 0.5202) and the invariable sites ( + I = 21.68% sites). The numbers on the nodes indicate the percentage of bootstrap values obtained from 1,000 replicates. Bootstrap values of less than 60% were excluded from nodes. The branch lengths show the number of substitutions per site. Evolutionary analyses were performed in MEGA760. The two suggestive clades of genotype A, “European/Turkish clade” and “Middle Eastern/Iranian clade” are shown with arrows.
Genetic distance analyses of German and Iranian SRLV sequences based on gag gene
As all German and Iranian gag sequences tend to be genotype A, a cut-off value was determined for assigning a new subtype within genotype A. The mean genetic distances between available gag reference data (A1–A20 except for A6, A10, A14 and A15) was 14.67% (SE = 0.19%, 95%CI = 14.29–15.05%). The value of 14.67% was the used cut-off value for defining a new subtype within genotype A. German gag sequences of the proposed subtype A21 (n = 9), had mean sequence similarity of 11.62% (range = 0.06–15.01%). The following mean genetic distances between the new subtype (A21) and different genotypes were observed: A: 15.15%, range = 13.94–16.81%; B: 23.11%, range = 22.01–24.17%; C: 23.61%, range = 21.88–25.95%; E: 28.50%, range = 27.42–29.46% (Table 2 and Supplementary Table S1 online). Based on phylogeny, the most similar subtype with cluster A21 was A18. In two out of nine gag sequences of cluster A21 (MN233132 and MN132145), genetic distances compared to A18 were higher than the cut-off value. Observation of these two sequences in cluster A21 that genetically differed from A18 (>14.67), fulfils the criterion of proposing a new subtype for a cluster in SRLVs4. Thus, all nine sequences of cluster A21 constitute subtype A21 (new). In other German gag sequences, the genetic distances with the most similar subtypes (corresponding to subtypes A4, A5, A11 and A16 based on phylogeny) were lower than the cut-off value (<14.67). Therefore, other German gag sequences did not constitute new subtypes and joined the reference subtypes A4 (MN233148), A5 (MN233105, MN233107 and MN233108), A11 (MN233124, MN233143 and MN233151) and A16 (MN233104). Notably, according to phylogeny (Fig. 1), the most similar subtypes with the three sequences of MN233143, MN233124 and MN233151 were A11 and A20 (both SRLV subtypes from Italy). Based on genetic distance analysis these three sequences were more similar with A11 than with A20. More details are shown in Table 2 and Supplementary Table S1 online.
The gag sequences of six Iranian sheep constituted a cluster (subtype A22) within genotype A. The mean genetic similarity between Iranian SRLVs was 10.62% and varied from 5.34% to 12.72%. The following mean genetic distances between Iranian SRLV sequences (A22) and different genotypes were observed: A: 18.29%, range = 17.85–18.89%; B: 23.45%, range = 22.31–24.77%; C: 24.13%, range = 22.65–25.45%; E: 28.73%, range = 27.87–29.84% (Table 2 and Supplementary Table S1 online). The genetic divergences between six Iranian, two Jordanian (KT898826 and KT921318) and one Lebanese (KU170760) gag sequences were lower than the cut-off value (<14.67). Therefore, all these nine gag sequences together constitute subtype A22. More details are shown in Table 2 and Supplementary Table S1 online.
Amino acid (aa) sequence analysis based on the gag-pol fragment
The alignment of 65 gag-pol aa sequences (up to 214 aa) is shown in Fig. 2 and Supplementary Fig. S2 online. The reference gag-pol sequence in both Fig. 2 and Supplementary Fig. S2 online is the Iranian SRLV strain of BKH1 (accession number: MK098477). Except for the reference gag-pol sequence (MK098477), the gag-pol aa sequences in Fig. 2 (n = 32) and Supplementary Fig. S2 online (n = 32) are not identical. Therefore, the sum of gag-pol aa sequences that have been aligned with MK098477 is 64, and altogether a total of 65 gag-pol aa sequences is shown in Fig. 2 and Supplementary Fig. S2 online.
Amino acid sequence alignments of the SRLV gag-pol fragment. The reference sequence is the SRLV sequence of strain BKH1 (accession number: MK098477) from the Iranian province of Chaharmahal-Va-Bakhtiari. For this amino acid comparison, 6 Iranian (labelled by solid blue circles) and 17 German sequences (labelled by solid red circles) were compared with different SRLV subtypes/genotypes. Immunodominant epitopes 2 and 3, the major homology region (MHR), the double glycine motif (GG) and insertions (at position 172 or 173) are delineated with boxes. The major core protein (p25) and nucleic acid-binding protein (p14) are separated with left and right arrows (p25 ≤ position 160; p14 ≥ position 161). Dashes and dots indicate deletions and identical residues, respectively.
Both German and Iranian SRLV sequences share similar epitope patterns with other sequences of genotype A. Also, they did not share the double glycine “GG” motif48, as other sequences of genotype A (Fig. 2). When comparing amino acid sequence substitutions at epitopes 2, major homology region (MHR) and epitopes 3, few alterations were observed mostly in the middle part of the conserved SRLV domains found in Germany and Iran.
All the Iranian SRLVs, as well as the Jordanian (KT898826 and KT921318) and Lebanese (KU170760) strains, contained an insertion at position 173, which was not seen in any other small ruminant lentiviruses (Fig. 2 and Supplementary Fig. S2 online). A second insertion at position 172 was found exclusively in two SRLV gag-pol sequences in sheep flocks from the Iranian province of Chaharmahal-Va-Bakhtiari. Interestingly, these insertions (positions 172 or 173) are found exclusively in SRLV subtype A22 and as well in other lentiviruses, including bovine immunodeficiency virus (BIV), human immunodeficiency virus type 1 (HIV1), simian immunodeficiency virus (SIV), feline immunodeficiency virus (FIV), equine infectious anemia virus (EIAV) and human immunodeficiency virus type 2 (HIV2) (Supplementary Fig. S3 online).
Geographical distribution of SRLV subtypes found in this study
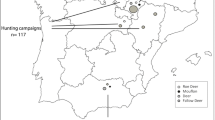
SRLV subtypes of A4, A5, A11, A16, A21 and A22 were observed in this study. Except for A21, the other SRLVs have also been found in other countries. The qualitative aspects of the phylogeography were presented in Fig. 3 by showing the geographic dispersal of pairwise SRLV subtypes of A5, A11 and A22. Transmission routes of SRLV subtypes A5 (solid red line), A11 (solid black line) and A22 (solid green line) follow the Danubian pathway, northern Mediterranean pathway and the ancient Fertile Crescent, respectively. As subtype A4 was only determined in Germany and Switzerland4, and subtype A16 was only observed in Germany and Poland12, for simplicity, their transmission routes were not shown in Fig. 3.
Putative transmission routes of SRLV subtypes A5, A11 and A22 based on their geographical distribution. The Danubian domestication pathway, corresponding to the Danube River, rises in Germany, flows through different European countries, and finally reaches the Black Sea. The northern Mediterranean pathway comprises Turkey and southern parts of Europe, including Italy. The ancient Fertile Crescent region incorporates Iran, Iraq, Turkey, Syria, Lebanon and Jordan (based on Harlan and Zohary57). The map was created using online Scribble Maps (https://www.scribblemaps.com/). For more information concerning printing permissions and inclusion of logos please visit https://help.scribblemaps.com/knowledgebase/articles/878916-printing-permissions-book-offline-etc. The online Scribble Maps used the databases of “© OpenStreetMap”, “© Mapbox” and “Improve this map”. The OpenStreetMap provides free geographic data and is available under the Open Database Licence (https://www.openstreetmap.org/copyright). To learn more about the Mapbox, visit https://www.mapbox.com/about/maps/. Further information regarding the use of “Improve this map” is available at https://www.mapbox.com/map-feedback/. The URL for linking to the map in Fig. 3 is https://www.scribblemaps.com/maps/view/Figure_3_/XUxAbALsVL.
Discussion
Maedi-visna disease, already distributed among the sheep flocks of Germany41,49, was first detected in the German East (1967) and then in the West (1972)41. It was reported in southwestern Iran in 200142 and later in other parts of the country43,44,45. In this study, for the first time, gag sequences of German and Iranian SRLVs were generated and analysed, using samples from five German states and three Iranian provinces. Our analyses identified a noticeable variation in German SRLV sequences that was absent in the Iranian ones. In the 13 studied German flocks, SRLV subtypes A4, A5, A11, A16 and A21 were found, with co-infection being identified in three flocks: flock 3 (A5 and A21), flocks 8 and 10 (A11 and A21) (Table 1). In the six Iranian flocks, only subtype A22 was found.
In earlier studies, migration of at least some SRLVs was related to the old times16,50. For example, subtype B3 and genotype E, which are rare, were found in Turkey and Italy (Sardinia and other regions)16. Turkey is one of the centres of sheep and goat domestication28,32, and the early agriculture of Sardinia is related to the Neolithic period21,27,32,33. Later, observations of various SRLV subtypes of genotype A (A2, A3, A5, A9 and A11) in sheep flocks of Turkey let Muz et al.50 propose that the ancestors of all SRLVs, especially genotype A, are Turkish SRLVs, and the evolution of some SRLVs is related to the domestication process or a more recent transmission pathway during the Ottoman Empire (about 14th–19th centuries). Our results are consistent with this scenario, which relates the evolution of SRLVs to the domestication process16,50.
So far, SRLV subtype A22 has been observed in sheep in Iran, Lebanon and Jordan but not in any European countries, including Turkey. The absence of A22 among small ruminants of Turkey and Europe is likely related to their hosts’ lineages. Several reports have shown that domestic sheep have different wild ancestors. Already, five lineages (A–E) have been identified in modern domestic sheep through global studies of mitochondrial DNA51,52,53,54,55. There is a relatively low percentage of lineages C–E, whilst the two most frequent lineages in domestic sheep are A and B, as identified by Hiendleder and colleagues51,52,55. These two lineages could be linked, but not completely, with modern fat- and thin-tailed where the first one is mostly found in Iran and Eastern Asia and the other one in Europe (including Turkey)56. Possibly, this could be a result of geographical separation of sheep lineages in the past. Accordingly, a long-term genetic differentiation took place between SRLV subtype A22, found in Iran, Lebanon and Jordan and other SRLVs of genotype A, found in Europe. Therefore, A22 could be suggested as an additional ancestor for genotype A of SRLVs (see Fig. 1; European/Turkish clade vs. Middle Eastern/Iranian clade).
In this study, we realised that the relationship between transmission routes of some SRLVs (A5, A11 and A22, this study; A3, A9, B3 and E reviewed by Ramírez et al.6) are compatible with domestication pathways of sheep. Three domestication pathways have been proposed from the Near East (Iran and Turkey) to the West and Africa32,47. The first pathway follows the Danube River from the Near East (Turkey) to Germany32,47. An excellent example of this distribution is the observation of SRLV subtype A5 in Turkey50 (start terminal), Slovenia10, Switzerland4 and Germany (end terminal). This subtype’s existence in countries lying on the route between Turkey and Germany (Fig. 3, red line) shows that the distribution is targeted. A further hint for the existence of this pathway is SRLV subtype A3, which has been identified in Turkey, Switzerland and Spain (reviewed by Ramírez et al.6). The second domestication pathway runs from the Near East via the Mediterranean route to southern Europe32,47, including Italy. SRLV subtype A11 hints at this distribution. It has been detected in Turkey50 and Italy8 and in the current study also in Germany (Fig. 3, black line). Other clues for the second domestication pathway could be related to the previously observed subtypes A9, B3 and genotype E in Turkey and Italy8,16,50. The third proposed domestication pathway runs from the Near East via the southern Mediterranean route to the northern parts of Africa32,47. A good support of this distribution is shown by the observations of A22 in Iran, Lebanon and Jordan; these observations are also compatible with the location of the ancient Fertile Crescent57 (Fig. 3, green line), where domestication originated. Therefore, our data reflect an association between the distribution of domestication and the transmission of some SRLVs. However, the distribution of some SRLV subtypes may not be related to antiquity. For example, subtype A16 was found only in Germany and Poland12. Subtype A21, the most common subtype in the North of Germany, is genetically close to subtype A1813 in Poland. A study by Olech et al.13 mentioned an epidemiological linkage between the small ruminants of Germany and Poland after the Second World War, which is supported by our results.
Amino acid insertions 172 or 173 were observed in SRLV subtype A22 but were not found in any other subtypes. Notably, we observed these insertions to be typical for subtype A22 and other lentiviruses but not for all other SRLVs. As these insertions are in a variable area of the gag-pol region (Supplementary Fig. S3 online), bioinformatics analyses may not show how these insertions have evolved in different lentiviruses. Observation of these insertions, however, confirms evolutionary processes within and between different lentiviruses. Furthermore, the hosts of the Jordanian isolates were sheep of the Awassi breed (M. Mazzei, University of Pisa, Italy, personal communication), which is a fat-tailed sheep with a history related to the second wave of domestication (about 5,000 years ago)58. Likewise, the Iranian samples came from three provinces close to the Zagros Mountains that are also known as primary centres of goat and sheep domestication29,30. The common presence of a unique variant of SRLVs (insertion at positions 172 or 173) among sheep from the countries of the ancient Fertile Crescent suggests that subtype A22 may represent an ancient precursor of modern SRLVs.
The detection of a single SRLV A subtype in Iranian, Jordanian and Lebanese sheep also suggests that these sheep have lived in isolation for centuries. The restricted variability points to this SRLV subtype as a potential marker to trace sheep domestication pathways. In contrast, the situation in Europe may be much more complex and the influence of trading overwhelming, confounding the potential link of SRLV genotypes to the domestication pathways, such as the Danubian or northern Mediterranean pathways. In this respect, Chessa and colleagues27 showed that tracking infection and endogenization of the Jaagsiekte sheep retrovirus in different sheep breeds permits the reconstruction of the domestication history of sheep. Unlike SRLVs, Jaagsiekte sheep retrovirus introduces a stable marker in the genome of the infected animals and may be more suitable to follow the domestication pathways of the species under study. Therefore, further studies on sheep genome variations using endogenous retroviruses may provide a more precise picture of the domestication pathways in Europe.
Earlier reports mentioned that maedi-visna (subtype A1) arrived in Iceland with Karakul sheep imported from Germany. Straub41 showed that this Karakul flock existed in Germany until 1970 but that the sheep of this flock had never shown any signs of maedi-visna. Currently, there is a meager number of Karakul sheep in Germany (maybe a single breeding flock), and this flock is not known to be SRLV positive. However, SRLVs are not strictly breed-specific. They can circulate from one region to another and as well from one breed to another. In this study, we found no evidence for the presence of the Icelandic subtype (A1) among 48 gag sequences in German sheep. However, it is possible that there are additional subtypes or genotypes of SRLVs present in German sheep flocks that have not yet been identified, including A1. More work is therefore needed to firmly rule out the hypothesis of distribution of subtype A1 via Karakul sheep from Germany to Iceland.
Shah et al.4 developed a systematic taxonomic classification for SRLVs, based on 1.8-kb gag-pol sequences. At that time and later, many strains have been included in the list of new SRLVs when the characteristic of a new strain was distinct enough from other strains. However, in different research projects, different fragments of gag-pol were sequenced. The available sequence data for the SRLV subtypes A12, A13, A14, A16, A17 and A18 were not fully matched to the sequence data of other SRLV subtypes of genotype A. As a result, the alignment and classification of SRLVs were limited to only 0.4-kb of gag gene (Supplementary Fig. S1 online). In this study, we used the same analysis for defining the cut-off value as it was used by Shah et al.4. As the 0.4-kb gag fragment we used in this study is more conserved than the 1.8-kb gag-pol fragment used by Shah et al.4, we adapted the cut-off value (14.67% instead 15%) in order to get an accurate classification.
The principal limitation of this study is the restricted number of sequences obtained. 48 samples were collected in Germany, 38 of which came from Schleswig-Holstein, 6 from Nordrhein-Westfalen, and the remaining 4 from Baden-Württemberg, Hessen and Bayern. It is highly unlikely that these sequences are representatives of the SRLV strains circulating in this country. The same applies to Iran, with only six sequences. The reason for unbalanced sequencing results from German flocks is the identification of more positive flocks in the North of Germany49. Although Germany is not free of SRLVs, there are not many infected flocks, especially of low susceptible breeds46. German Texel is a sheep breed susceptible to SRLV infection and it is mostly found in the state of Schleswig-Holstein (North of Germany)49. We found less/no positive flocks in other German states. The six Iranian SRLV sequences were collected in a previous study by collecting samples from 30 sheep flocks in six Iranian provinces. We found SRLV positive flocks only in three provinces of Western Azarbaijan, Kerman and Chaharmahal-Va-Bakhtiari46. Similar to the situation in Germany, the distribution of SRLV positive samples in these three Iranian provinces was not balanced. Finally, we decided to select two samples per each province. Therefore, performing follow up studies on characterisation of SRLVs in sheep flocks from additional regions in Germany and Iran is necessary to provide a more in-depth insight into the variability of SRLVs circulating in both countries.
Materials and Methods
Samples
For characterisation of German and Iranian SRLVs, a total of 54 DNA samples was selected from SRLV positive sheep flocks in Germany (13 flocks, n = 48 samples) and Iran (6 flocks, n = 6 samples)49. The SRLV infection status of German and Iranian sheep flocks were determined in earlier studies based on serological (IDEXX CAEV-MVV Total Ab ELISA, Ludwigsburg, Germany)49 and PCR (a semi-nested PCR on env gene)46 tests, respectively. Origin of flocks and the number of sequenced samples from each flock in this study are shown in Table 1.
Ethical approval
No live animals were used for this study. All blood samples were collected by trained veterinarians exclusively as part of a routine clinical examination and the residuals of the samples would have otherwise been discarded. According to the German regulations for animal protection, the German Animal Welfare Act (released on 8/5/2006, modified on 17/12/ 2018), this origin of samples obviates the need for an explicit ethics committee approval. Iran Veterinary Organization (IVO), a national authority, participated in collecting Iranian sheep samples (permission issued on 20/12/2014, No: 93/22/70521). The ethical responsibility for Iranian animal welfare was carried out in accordance with the legal regulations of the IVO.
PCR amplification and sequencing of SRLV proviral DNA
To characterise the SRLVs found in Germany and Iran, genomic DNA of sampled sheep was analysed using a nested PCR targeting the gag-pol region of the provirus as described and utilised elsewhere8,9,50.
For sequencing purposes, PCR products of the second PCR were purified and directly sequenced with either ABI PRISM 3130 Genetic Analyzer or by LGC Genomics GmbH, Germany (https://shop.lgcgenomics.com). To control PCR errors, both the sense and antisense strands were sequenced performing three independent reactions for each sample.
Analyses of SRLV sequence data
The obtained SRLV provirus nucleotide sequences were trimmed and analysed using the ChromasPro software version 2.1.6 (Technelysium Pty Ltd, Tewantin, Australia). Recombination analyses were carried out using RDP4 package59. Multiple alignments of the nucleotide sequences and the deduced amino acid sequences were generated with Muscle-built in Mega version 7.0.2660.
For simplicity, only one or two sequences (Table 1) of each flock were selected and used for all analyses. The available sequences of the reference SRLV strains of genotypes A–C and E, and the SRLV strains most homologues to German and Iranian SRLVs (using the Basic Local Alignment Search Tool, BLAST) were included in the analysis. The sequence data of SRLV subtypes A6, A10, A15 were not available (a pol fragment defined these subtypes) for the analyses. Additionally, A14 had to be excluded from analyses because of the shortness of the relevant sequence part. All analyses were carried out based on 65 aligned sequences from various sources (Germany 17, Iran 6, Jordan 2, Lebanon 1 and 39 from reference SRLV subtypes).
The evolutionary relationships of German and Iranian SRLVs with other published sequences were investigated by constructing the phylogenetic tree from alignments of the gag region. The best-fitting nucleotide substitution model for phylogenetic analysis was the Tamura-Nei (TN93)61 model, with the gamma distribution (G) and invariant sites (I), and it was estimated using MEGA version 7.0.2660. The phylogeny was estimated using the maximum likelihood (ML) method. The percentage of bootstrap values were obtained based on 1,000 repetitions.
Genetic divergences between sequences were estimated with the p-distance model applying the gamma distribution parameter using Mega version 7.0.2660. Descriptive statistics of genetic distances were done with the SPSS program version 25.0 for Windows (IBM SPSS Statistics, Armonk, NY: IBM Corp). A cut-off value for assigning new subtypes within genotype A was determined according to the gag fragment (399 bp). This was done by calculating the mean genetic distances among SRLV subtypes of A1–A20 (except for A6, A10, A14 and A15).
The deducted gag-pol amino acid sequences (214 aa) of 6 Iranian and 17 German gag-pol sequences were aligned with all available sequence data. The purposes of this alignment were: (1) Checking amino acid variations of epitopes 2 and 3, especially the N-terminus and C-terminus, as such substitutions may affect the sensitivity of ELISA. (2) Checking amino acid variations, with regard to epitopes 2 and 3, and other conserved motifs (GG and MHR), to be sure whether our sequence data have been genotyped correctly. Previous studies have shown some specific differences between different genotypes of A-C and E9,15. (3) Finding potential amino acid variation/s in our sequence data that presumably is linked to other lentiviruses of the family Retroviridae.
Data availability
All data generated and analysed in this study are included in this published article and its supplementary information files.
References
Narayan, O. et al. The lentiviruses of sheep and goats in The Retroviridae 2 (ed. Levy, J.A.) 229–255 (Springer, Boston, MA, 1993).
Peterhans, E. et al. Routes of transmission and consequences of small ruminant lentiviruses (SRLVs) infection and eradication schemes. Vet. Res. 35, 257–274, https://doi.org/10.1051/vetres:2004014 (2004).
Pépin, M., Vitu, C., Russo, P., Mornex, J.-F. & Peterhans, E. Maedi-visna virus infection in sheep: a review. Vet. Res. 29, 341–367, https://hal.archives-ouvertes.fr/hal-00902532 (1998).
Shah, C. et al. Phylogenetic analysis and reclassification of caprine and ovine lentiviruses based on 104 new isolates: evidence for regular sheep-to-goat transmission and worldwide propagation through livestock trade. Virology 319, 12–26, https://doi.org/10.1016/j.virol.2003.09.047 (2004).
Reina, R. et al. Prevention strategies against small ruminant lentiviruses: An update. Vet. J. 182, 31–37, https://doi.org/10.1016/j.tvjl.2008.05.008 (2009).
Ramírez, H., Reina, R., Amorena, B., de Andrés, D. & Martínez, H. A. Small ruminant Lentiviruses: Genetic variability, tropism and diagnosis. Viruses 5, 1175–1207, https://doi.org/10.3390/v5041175 (2013).
Minardi da Cruz, J. S., Singh, D., Lamara, A. & Chebloune, Y. Small ruminant lentiviruses (SRLVs) break the species barrier to acquire new host range. Viruses 5, 1867–1884, https://doi.org/10.3390/v5071867 (2013).
Giammarioli, M. et al. Phylogenetic analysis of small ruminant lentivirus (SRLV) in Italian flocks reveals the existence of novel genetic subtypes. Virus Genes 43, 380–384, https://doi.org/10.1007/s11262-011-0653-1 (2011).
Grego, E. et al. Genetic characterization of small ruminant lentivirus in Italian mixed flocks: evidence for a novel genotype circulating in a local goat population. J. Gen. Virol. 88, 3423–3427, https://doi.org/10.1099/vir.0.83292-0 (2007).
Kuhar, U., Barlič-Maganja, D. & Grom, J. Phylogenetic analysis of small ruminant lentiviruses detected in Slovenia. Vet. Microbiol. 162, 201–206, https://doi.org/10.1016/j.vetmic.2012.08.024 (2013).
Olech, M., Rachid, A., Croisé, B., Kuźmak, J. & Valas, S. Genetic and antigenic characterization of small ruminant lentiviruses circulating in Poland. Virus Res. 163, 528–536, https://doi.org/10.1016/j.virusres.2011.11.019 (2012).
Olech, M., Valas, S. & Kuźmak, J. Epidemiological survey in single-species flocks from Poland reveals expanded genetic and antigenic diversity of small ruminant lentiviruses. PLoS One 13, e0193892, https://doi.org/10.1371/journal.pone.0193892 (2018).
Olech, M., Murawski, M. & Kuźmak, J. Molecular analysis of small-ruminant lentiviruses in Polish flocks reveals the existence of a novel subtype in sheep. Arch. Virol. 164, 1193–1198, https://doi.org/10.1007/s00705-019-04161-9 (2019).
Pisoni, G. et al. Genetic analysis of small ruminant lentiviruses following lactogenic transmission. Virology 407, 91–99, https://doi.org/10.1016/j.virol.2010.08.004 (2010).
Colitti, B. et al. A new approach for Small Ruminant Lentivirus full genome characterization revealed the circulation of divergent strains. PLoS One 14, e0212585, https://doi.org/10.1371/journal.pone.0212585 (2019).
Bertolotti, L. et al. Characterization of new small ruminant lentivirus subtype B3 suggests animal trade within the Mediterranean Basin. J. Gen. Virol. 92, 1923–1929, https://doi.org/10.1099/vir.0.032334-0 (2011).
Gjerset, B., Storset, A. K. & Rimstad, E. Genetic diversity of small-ruminant lentiviruses: Characterization of Norwegian isolates of Caprine arthritis encephalitis virus. J. Gen. Virol. 87, 573–580, https://doi.org/10.1099/vir.0.81201-0 (2006).
Gjerset, B., Rimstad, E., Teige, J., Soetaert, K. & Jonassen, C. M. Impact of natural sheep-goat transmission on detection and control of small ruminant lentivirus group C infections. Vet. Microbiol. 135, 231–238, https://doi.org/10.1016/j.vetmic.2008.09.069 (2009).
Reina, R. et al. Molecular characterization and phylogenetic study of Maedi Visna and Caprine Arthritis Encephalitis viral sequences in sheep and goats from Spain. Virus Res. 121, 189–198, https://doi.org/10.1016/j.virusres.2006.05.011 (2006).
Reina, R. et al. Small ruminant lentivirus genotype E is widespread in Sarda goat. Vet. Microbiol. 144, 24–31, https://doi.org/10.1016/j.vetmic.2009.12.020 (2010).
Gifford, R. J. Viral evolution in deep time: lentiviruses and mammals. Trends Genet. 28, 89–100, https://doi.org/10.1016/j.tig.2011.11.003 (2012).
Zhang, C., De Silva, S., Wang, J.-H. & Wu, L. Co-evolution of primate SAMHD1 and lentivirus Vpx leads to the loss of the vpx gene in HIV-1 ancestor. PLoS One 7, e37477, https://doi.org/10.1371/journal.pone.0037477 (2012).
Yamada, E. et al. A naturally occurring bovine APOBEC3 confers resistance to bovine lentiviruses: implication for the co-evolution of bovids and their lentiviruses. Sci. Rep. 6, 33988, https://doi.org/10.1038/srep33988 (2016).
Takeuchi, J. S. et al. Coevolutionary dynamics between tribe Cercopithecini tetherins and their lentiviruses. Sci. Rep. 5, 16021, https://doi.org/10.1038/srep16021 (2015).
Worobey, M. et al. Island biogeography reveals the deep history of SIV. Science 329, 1487, https://doi.org/10.1126/science.1193550 (2010).
Garcia-Etxebarria, K., Sistiaga-Poveda, M. & Marina Jugo, B. Endogenous retroviruses in domestic animals. Curr Genomics 15, 256–265 (2014).
Chessa, B. et al. Revealing the history of sheep domestication using retrovirus integrations. Science 324, 532–536, https://doi.org/10.1126/science.1170587 (2009).
Zeder, M. A. Domestication and early agriculture in the Mediterranean Basin: Origins, diffusion, and impact. Proc. Natl. Acad. Sci. 105, 11597–11604, https://doi.org/10.1073/pnas.0801317105 (2008).
Zeder, M. A. Animal domestication in the Zagros: a review of past and current research. Paléorient 25, 11–25, http://www.jstor.org/stable/41496540 (1999).
Zeder, M. A. & Hesse, B. The initial domestication of goats (Capra hircus) in the Zagros Mountains 10,000 years ago. Science 287, 2254–2257, https://doi.org/10.1126/science.287.5461.2254 (2000).
Pereira, F. & Amorim, A. Origin and spread of goat pastoralism. e LS; https://doi.org/10.1002/9780470015902.a0022864 (2001).
Zeder, M. A. Out of the Fertile Crescent: The dispersal of domestic livestock through Europe and Africa in Hum. Dispersal Species Mov. From Prehistory to Present (ed. Boivin, N., Crassard, R. & Petraglia, M.) 261–296 (Cambridge University Press, 2017)
Vigne, J. D. Zooarchaeology and the biogeographical history of the mammals of Corsica and Sardinia since the last ice age. Mamm. Rev. 22, 87–96, https://doi.org/10.1111/j.1365-2907.1992.tb00124.x (1992).
Greenfield, H. J. et al. The origins of milk and wool production in the Old World: a zooarchaeological perspective from the Central Balkans [and comments]. Curr. Anthropol. 29, 573–593 (1988).
Kaewthamasorn, M. et al. Genetic homogeneity of goat malaria parasites in Asia and Africa suggests their expansion with domestic goat host. Sci. Rep. 8, 5827, https://doi.org/10.1038/s41598-018-24048-0 (2018).
Mühlemann, B. et al. Ancient human parvovirus B19 in Eurasia reveals its long-term association with humans. Proc. Natl. Acad. Sci. 115, 7557–7562, https://doi.org/10.1073/pnas.1804921115 (2018).
Otchere, I. D. et al. Comparative genomics of Mycobacterium africanum Lineage 5 and Lineage 6 from Ghana suggests distinct ecological niches. Sci. Rep. 8, 11269, https://doi.org/10.1038/s41598-018-29620-2 (2018).
Sigurdsson, B., Grímsson, H. & Pálsson, P. A. Maedi, a chronic, progressive infection of sheep’s lungs. J. Infect. Dis. 90, 233–241 (1952).
Blacklaws, B. A. et al. Transmission of small ruminant lentiviruses. Vet. Microbiol. 101, 199–208, https://doi.org/10.1016/j.vetmic.2004.04.006 (2004).
Thormar, H. The origin of lentivirus research: Maedi-visna virus. Curr. HIV Res. 11, 2–9, https://doi.org/10.2174/157016213804999212 (2013).
Straub, O. C. Maedi–Visna virus infection in sheep. History and present knowledge. Comp. Immunol. Microbiol. Infect. Dis. 27, 1–5, https://doi.org/10.1016/S0147-9571(02)00078-4 (2004).
Sayari, M. & Soltanian, S. Serological and pathological study of Maedi-like diseases in mammary glands of sheep of Ahwaz region. Pajouhesh Sazandegi (in Persian), 2–7 (2001).
Azizi, S. et al. Maedi in slaughtered sheep: A pathology and polymerase chain reaction study in southwestern Iran. Trop. Anim. Health Prod. 44, 113–118, https://doi.org/10.1007/s11250-011-9896-z (2012).
Norouzi, B., Razavizadeh, A. T., Azizzadeh, M., Mayameei, A. & Mashhadi, V. N. N. Serological study of small ruminant lentiviruses in sheep population of Khorasan-e-Razavi province in Iran. Veterinary Research Forum, 6, 245–249, PMC4611980 (2015).
Sasani, F., Javanbakht, J., Hemmatzadeh, F., Moghadam, M. R. & Hassan, M. A. M. Evaluation of histopathological on maedi disease with serological confirmation in North-East of Iran. Res. J. Infect. Dis. 1, 5, https://doi.org/10.7243/2052-5958-1-5 (2013).
Molaee, V., Otarod, V., Abdollahi, D. & Lühken, G. Lentivirus susceptibility in Iranian and German sheep assessed by determination of TMEM154 E35K. Animals. 9, 685, https://doi.org/10.3390/ani9090685 (2019).
Alberto, F. J. et al. Convergent genomic signatures of domestication in sheep and goats. Nat. Commun. 9, 813, https://doi.org/10.1038/s41467-018-03206-y (2018).
L’Homme, Y. et al. Molecular characterization and phylogenetic analysis of small ruminant lentiviruses isolated from Canadian sheep and goats. Virol. J. 8, 271, https://doi.org/10.1186/1743-422X-8-271 (2011).
Molaee, V., Eltanany, M. & Lühken, G. First survey on association of TMEM154 and CCR5 variants with serological maedi-visna status of sheep in German flocks. Vet. Res. 49, 36, https://doi.org/10.1186/s13567-018-0533-y (2018).
Muz, D. et al. First molecular characterization of visna/maedi viruses from naturally infected sheep in Turkey. Arch. Virol. 158, 559–570, https://doi.org/10.1007/s00705-012-1518-1 (2013).
Hiendleder, S., Mainz, K., Plante, Y. & Lewalski, H. Analysis of mitochondrial DNA indicates that domestic sheep are derived from two different ancestral maternal sources: no evidence for contributions from urial and argali sheep. J. Hered. 89, 113–120, https://doi.org/10.1093/jhered/89.2.113 (1998).
Hiendleder, S., Kaupe, B., Wassmuth, R. & Janke, A. Molecular analysis of wild and domestic sheep questions current nomenclature and provides evidence for domestication from two different subspecies. Proc. R. Soc. London. Ser. B Biol. Sci. 269, 893–904, https://doi.org/10.1098/rspb.2002.1975 (2002).
Pedrosa, S. et al. Evidence of three maternal lineages in Near Eastern sheep supporting multiple domestication events. Proc. R. Soc. B Biol. Sci. 272, 2211–2217, https://doi.org/10.1098/rspb.2005.3204 (2005).
Meadows, J. R. S., Cemal, I., Karaca, O., Gootwine, E. & Kijas, J. W. Five ovine mitochondrial lineages identified from sheep breeds of the near East. Genetics 175, 1371–1379, https://doi.org/10.1534/genetics.106.068353 (2007).
Meadows, J. R. S., Hiendleder, S. & Kijas, J. W. Haplogroup relationships between domestic and wild sheep resolved using a mitogenome panel. Heredity (Edinb). 106, 700, https://doi.org/10.1038/hdy.2010.122 (2011).
Bruford, M. W., Bradley, D. G. & Luikart, G. DNA markers reveal the complexity of livestock domestication. Nat. Rev. Genet. 4, 900, https://doi.org/10.1038/nrg1203 (2003).
Harlan, J. R. & Zohary, D. Distribution of wild wheats and barley. Science. 153, 1074–1080, https://doi.org/10.1126/science.153.3740.1074 (1966).
Sunderman, F. & Johns, M. Awassi fat tails: a chance for premium exports. J. Dep. Agric. West. Aust. Ser. 4 35, 99–105 (1994).
Martin, D. P., Murrell, B., Golden, M., Khoosal, A. & Muhire, B. RDP4: Detection and analysis of recombination patterns in virus genomes. Virus Evol., 1; https://doi.org/10.1093/ve/vev003 (2015).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874, https://doi.org/10.1093/molbev/msw054 (2016).
Tamura, K. & Nei, M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol. Biol. Evol. 10, 512–526, https://doi.org/10.1093/oxfordjournals.molbev.a040023 (1993).
Staskus, K. A. et al. Isolation of replication-competent molecular clones of visna virus. Virology 181, 228–240, https://doi.org/10.1016/0042-6822(91)90488-W (1991).
Acknowledgements
We want to thank Renate Latzke-Reinhardt and Lena Vogel from the Institute of Animal Breeding and Genetics, Justus Liebig University Giessen, Germany, for their excellent technical assistance. We also want to thank technical assistants of the Istituto Zooprofilattico Sperimentale dell’Umbria e delle Marche Togo Rosati, Perugia, Italy, who participated in our study. We are grateful to thank those sheep farmers who shared samples for this study.
Author information
Authors and Affiliations
Contributions
G.L. designed the research; G.L., V.O. and D.A. contributed to the collection of samples; V.M., M.B., G.M.D.M., G.L. and D.A. performed experiments; V.M. analysed data; V.M. generated figures; V.M. wrote the manuscript; G.L., S.R., M.B., G.M.D.M. and V.O. reviewed the manuscript; G.L. Supervision; All authors contributed to the final approval of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Molaee, V., Bazzucchi, M., De Mia, G.M. et al. Phylogenetic analysis of small ruminant lentiviruses in Germany and Iran suggests their expansion with domestic sheep. Sci Rep 10, 2243 (2020). https://doi.org/10.1038/s41598-020-58990-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-58990-9
- Springer Nature Limited
This article is cited by
-
Phylogenetic analysis of small ruminant lentiviruses in Mongolian sheep supports an ancient east-west split for the genotype A
Veterinary Research Communications (2024)
-
In vitro antiviral effect of ethanolic extracts from Azadirachta indica and Melia azedarach against goat lentivirus in colostrum and milk
Scientific Reports (2023)