Abstract
In recent years, accumulating evidences have shown that microRNA (miRNA) plays an important role in the exploration and treatment of diseases, so detection of the associations between miRNA and disease has been drawn more and more attentions. However, traditional experimental methods have the limitations of high cost and time- consuming, a computational method can help us more systematically and effectively predict the potential miRNA-disease associations. In this work, we proposed a novel network embedding-based heterogeneous information integration method to predict miRNA-disease associations. More specifically, a heterogeneous information network is constructed by combining the known associations among lncRNA, drug, protein, disease, and miRNA. After that, the network embedding method Learning Graph Representations with Global Structural Information (GraRep) is employed to learn embeddings of nodes in heterogeneous information network. In this way, the embedding representations of miRNA and disease are integrated with the attribute information of miRNA and disease (e.g. miRNA sequence information and disease semantic similarity) to represent miRNA-disease association pairs. Finally, the Random Forest (RF) classifier is used for predicting potential miRNA-disease associations. Under the 5-fold cross validation, our method obtained 85.11% prediction accuracy with 80.41% sensitivity at the AUC of 91.25%. In addition, in case studies of three major Human diseases, 45 (Colon Neoplasms), 42 (Breast Neoplasms) and 44 (Esophageal Neoplasms) of top-50 predicted miRNAs are respectively verified by other miRNA-disease association databases. In conclusion, the experimental results suggest that our method can be a powerful and useful tool for predicting potential miRNA-disease associations.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
As a small non-coding RNA (~22nt), MicroRNA (miRNA) plays a lot of critical managerial roles in cells. It is estimated that 1–4% of the genes in the human genome are miRNAs, with individual miRNAs regulating as many as 200 mRNAs1. miRNA usually binds to the 3′untranslation regions (UTRs) of the target mRNA through sequence-specific base pairs to inhibit the expression of target mRNA2,3,4,5. Because of this property, miRNAs can affect various biological processes and participate in a series of important processes in the life process6,7,8,9,10. In conclusion, it has been proved that miRNA plays a crucial role in biological processes. Understanding the molecular mechanism of disease is an important goal of biomedical researches. In this post-genome era, more and more contributions made by advanced high-throughput genome technologies are marching toward this goal. A lot of evidence indicates that miRNA plays a vital role in the development and progression of Human diseases1,11,12,13,14,15,16. For example, miR-195 expression levels are reduced in patients with Alzheimer’s disease (AD). Besides, over-expression of this miRNA can down-regulate the production of the AD amyloid-β17. Moreover, the expression of breast cancer patients’ serum miR-103 levels is significantly higher than that of healthy controls18. Therefore, we can believe that miRNA mutations, miRNA biogenic dysfunction, and miRNA’s target disorders may be associated with a variety of diseases, such as lung cancer19, lymphoma20, breast cancer21. However, to our knowledge, compared with a large number of cataloged miRNAs, systematic miRNA-disease association prediction methods are still insufficient. At the same time, the process of traditional laboratory experiments is very expensive and high time-consuming, so it is obvious that the computational method provides a new direction for large-scale miRNA-disease association prediction.
In recent years, a number of computational methods have been proposed to predict the associations between miRNA and disease. These methods can be classified according to their different strategies. For example, You et al.22 proposed a novel miRNA-disease association prediction model called PBMDA. This model constructs a heterogeneous graph composed of three interrelated subgraphs and then Depth-First-Search (DFS) algorithm is used to predict miRNA-disease associations. Chen et al.23 proposed a new bipartite network projection model for predicting potential associations between miRNA and disease (BNPMDA) based on miRNA functional similarity, disease semantic similarity, and the known human miRNA-disease associations. Zheng et al.24 developed a machine learning-based model for miRNA-disease association prediction (MLMDA). This method uses a deep auto-encoder neural network (AE), disease semantic similarity, miRNA sequence information, miRNA functional similarity and Gaussian association spectrum kernel similarity information to predict potential associations between miRNA and disease. Chen et al.25 established a model called WBSMDA. One of the advantages of this model is that it can be applied to diseases that are not associated with any miRNA, thus breaking through the limitations of most previous methods. You et al.26 put forward a new calculation method for the prediction of potential associations between miRNA and disease based on a personalized recommendation (PRMDA). In their study, a similarity network was widely used, taking into account the relevant miRNA and disease information for each miRNA-disease pair, thus recommending a high-priority potential miRNA-disease association. Jiang et al.27 proposed a calculation method to predict potential miRNA-disease associations by prioritizing the human microRNAome for diseases. It is a logical extension of earlier network-based approaches for predicting or prioritizing disease-associated protein-coding genes. They built a functionally-associated miRNA network and a human phenome-microRNAome network to examine whether functionally related miRNAs tended to be associated with diseases with similar phenotypes and prioritize miRNAs for human diseases. Shi et al.28 proposed a calculation method for miRNA and disease relationship prediction based on random walk analysis. They made a hierarchical clustering analysis on binary miRNA-disease networks to determine the miRNA-disease synergistic control module. Finally, the method yielded a good result, and provided a new perspective for predicting the relationship between miRNA and disease.
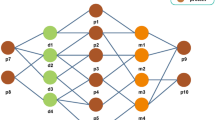
In this study, a network embedding-based heterogeneous information integration method is proposed to predict the potential associations between miRNA and disease. Firstly, a heterogeneous information network is established by combining the known associations between protein, miRNA, lncRNA, disease, and drug as shown in Fig. 1. After that, the network embedding method GraRep is adopted to learn the behavior information of miRNA and disease node in the network. As one of the network representation learning (NRL) models, the GraRep method can learn graph representations of the miRNA and disease nodes with global structural information. Secondly, the miRNA and disease nodes were converted to a vector by integrating the attribute information of the node itself (miRNA sequence information and disease semantic similarity) and the behavior information of them in the network to represent miRNA-disease pairs. Thirdly, 16427 known miRNA and disease pairs, which obtained from HMDD29 database, are used as positive samples and the same number of unrelated miRNA and disease pairs are randomly selected as negative samples, the two kinds of samples are combined to form the training samples. Finally, the prediction models are constructed based on the training samples by using the random forest, Fig. 2 shows the flowchart of our method. The model was evaluated through the 5-fold cross validation, and it performs well with high accuracy. To further test the effect of our method, we also conducted case studies of three major Human diseases. Our experiments prove that the network embedding method has great potential and provides a new direction for the prediction of miRNA and disease associations.
Materials and Methods
Heterogeneous information network construction
To systematically and comprehensively build the network of heterogeneous information, some known associations between miRNAs, lncRNAs, proteins, diseases, and drugs from multiple databases were downloaded. The source and version of the raw data are shown below: The miRNA-lncRNA association pairs are downloaded from the lncRNASNP2 database of Miao et al.30. The miRNA-protein association pairs are downloaded from the miRTarBase update 2018 database of Chou et al.31. The lncRNA-disease association pairs are downloaded from the lncRNASNP2 and LncRNADisease database of Miao et al.30 and Chen et al.32. The drug-disease association pairs are downloaded from the comparative toxicogenomics database: update 2019 of Davis et al.33. The lncRNA-protein association pairs are downloaded from the LncRNA2Target v2.0 database of Cheng et al.34. The drug-protein association pairs are downloaded from the DrugBank 5.0 database of Wishart et al.35. The protein-protein association pairs are downloaded from the STRING database in 2017 of Szklarczyk et al.36. The protein-disease association pairs are downloaded from the DisGeNET database of Piñero et al.37. The miRNA-disease association pairs are downloaded from the HMDD v3.0 database of Huang et al.29. After that, a series of operations such as unifying identifiers, de-redundancy, simplifying and deleting irrelevant items are conducted. The detailed data of the final experiment is shown in Table 1. In addition, we also classify and sort the above associations. Finally, we get different nodes as shown in Table 2.
Numerical miRNA sequence information
The sequences of miRNA are downloaded from miRbase38, to represent the attribute information of the miRNA node. To make the experiment less complicated, we select the 3-mer method and encode the miRNA sequence into a 64-dimensional feature vector, where each component represents the frequency of the occurrence of a 3-mer in the sequence (e.g. UGC, AUC, GUA).
Disease semantic similarity
The Medical Subject Heading (MeSH) database is a strict disease classification system, which can be used to effectively study the relationship between different diseases. Through this system, we can represent each disease with the Directed Acyclic Graph (DAG) achieved by MeSH of it. For example, for disease A, we can represent it as DAG(A) = (A, T(A), E(A)), where T(A) denotes all nodes in the DAG(A) that contain the disease A, E(A) indicating all disease link relationships in DAG(A)39. An example of gastrointestinal neoplasms’ DAG is shown in Fig. 3 below:
Therefore, we can select the disease semantic similarity calculated by DAG as the attribute information of disease according to the earlier method39. The semantic value of a disease D can be calculated as follows:
where ∆ is the semantic contribution factor and T(D) represents D and its all ancestor nodes. Observed results show that the two similar DAG ratios have higher disease similarity and the semantic similarity for disease \({d}_{i}\) and \({d}_{j}\) are defined as follows:
Stacked autoencoder
For the purpose of reducing the noise in the attribute information and normalizing it in a uniform dimension, we use a stacked autoencoder (SAE) to transform the original feature space into an appropriate subspace. SAE mainly consists of the following two steps: 1, the encoder projects x from the input layer to the hidden layer h through a mapping function f. 2, The decoder maps h in the hidden layer to y in the output layer through a mapping function g.
In this study, the ReLU function was selected as the activation function:
GraRep algorithms
Recently, many Network Representation Learning (NRL) methods have been proposed to learn vector representations of vertices in a network. GraRep40 is one of these methods. It factorizes different k-order proximity matrices and concatenates the embeddings learned from each proximity matrix. Specifically, GraRep takes into consideration the special relation matrix and extends the skip-gram model to capture the high order proximity of a network. It defines the k-step neighbors (k ≥ 1), and nodes that share a common k-step neighbor in the network should have similar and potential representations. Formally, the k-step representation of the learning node is composed of three steps. The first step is to obtain the k-step transition probability matrix \({A}^{k}\) for each k = 1, 2, … K. The second step is to use SVD method to factor the logarithmic probability matrix \({X}^{k}\) to obtain each k step representation:
where both U and V are orthonormal matrices and ∑ is a diagonal matrix that consists of an ordered list of singular values. The third step is to connect all k step representations, which can be represented as the following matrix:
More detailed algorithmic process participation can be seen in Table 3.
Node representation
The miRNA and disease nodes are represented by their intrinsic attribute information and behavior information with other nodes in the heterogeneous information network. The attribute information is respectively numerical miRNA sequence information and disease semantic similarity. In addition, in this paper, a network embedding method GraRep is used to obtain the behavior information of nodes in the entire network, before combining with their own attribute information. Their relationship with other nodes can be regarded as a functional representation based on the idea of collaborative filtering. Finally, they are converted into 128-dimensional vectors to represent known miRNA-disease associations.
Result and Discussion
Evaluate the performance of our method under the 5-fold cross validation
5-fold cross validation was used to evaluate the performance of our study, which randomly divided all data sets into five equal parts. In each validation, one part is used as the test set and the other four parts as the training set, so that test and training data do not overlap each other to ensure unbiased comparisons. The detailed result information of the proposed method is shown in Table 4. It can be seen from Table 4 that our proposed method exhibited the outcomes of average accuracy (Acc.), precision (Prec.), sensitivity (Sen.), matthews correlation coefficient (MCC), specificity (Spec.) and the areas under the ROC curve (AUC) of 85.11%, 88.75%, 80.41%, 70.53%, 89.81% and 91.25%, respectively.
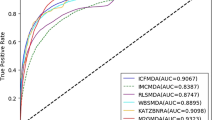
The receiver operating characteristic (ROC) curve is a functional image describing sensitivity. Here, its horizontal axis represents the False Positive Rate (FPR), which represents the ratio of all negative examples in the partitioning example to all negative cases (1-Specificity), where the larger the FPR and the more positive negative classes in the positive class are predicted. Besides, its vertical axis represents the True Positive Rate (TPR), which is used to represent the positive class coverage (Sensitivity). The larger the TPR and the more positive classes in the positive class are predicted. The AUC value indicates the areas under the ROC curve and it ranges from 0.1 to 1. AUC can be used as a numerical value to directly evaluate the quality of the classifier. We can see from Fig. 4 that the average AUC value obtained by our method is 0.9125. The Precision-Recall (PR) curve is another way to evaluate the performance of our method. It shows a trade-off between precision and sensitivity for all possible thresholds. From Fig. 5, we can see the PR curve corresponding to our method and the mean of the area under the precision-recall curve (AUPR) value is 0.9215. This once again proves that the good performance of our method.
Comparison of our method with different feature combinations
As we stated above, we use two different pieces of information to represent miRNA and disease in the entire network. Therefore, for the purpose of further testing the influence of various types of feature combinations on the classification results, we use attribute information, behavior information and attribute information plus behavior information to represent nodes respectively before conducting 5-fold cross-validation experiments. As the results of the final experiment shown in Table 5 and Fig. 6, there is a better performance in classification when we consider the attribute and behavior information simultaneously.
Comparison of our method with different classifiers
To further test the influence of the classifier in our model, we compared the performance of the four classifiers of Random Forest41, Decision Tree42, KNN43, and Naive Bayes44 under 5-fold cross validation. During the comparison experiment, we kept the same experimental environment, same training set and test set, and only changed the type of classifier. Similarly, we still use the six parameters: accuracy (Acc.), precision (Prec.), sensitivity (Sen.), matthews correlation coefficient (MCC), specificity (Spec.), and the areas under the ROC curve (AUC) as evaluation indicators. In the result, the Random Forest model yields average Acc., Prec., Sen., MCC, Spec. and AUC of 85.11 ± 0.37%, 88.75 ± 0.32%, 80.41 ± 0.73%, 70.53 ± 0.71%, 89.81 ± 0.33% and 91.25 ± 0.35%. Table 6 and Fig. 7 show the final comparison results. It can be seen that the Random Forest classifier has better performance and robustness than other classifiers, especially in the accuracy and AUC that can more represent the performance of the model, although our model is not as good as KNN and Naïve Bayes model are in sensitivy. In short, Random Forest is a better classifier for our model.
Case studies
In order to further test the prediction accuracy of our method, three Human diseases are selected for case studies. They are Colon neoplasms, Breast neoplasms, and Esophageal neoplasms, which are closely related to human health. We utilized the known miRNA-disease associations in HMDD V3.029 as the training set. The embedding representations of miRNA and disease are integrated with the attribute information of them (e.g. miRNA sequence information and disease semantic similarity) to represent these known miRNA-disease association pairs so that the input miRNAs and diseases can be identified by the classifier. Finally, the prediction model is constructed based on the training set by using random forest. After that, we constructed the test set for each investigated disease. The test set contains miRNAs in the heterogeneous information network and corresponding disease association pairs. In particular, the miRNA-disease association pairs already existing in the training set were deleted in the test set, including the disease-related miRNAs listed in Tables 7–9. Similarly, after converting the test set into the combination of embedding representations and attribute information, we use the prediction model to make predictions. After the completion of the prediction, the top 50 predicted miRNAs are selected and validated using two other miRNA-disease association databases, dbDEMC45 and miR2Disease46.
Colon neoplasms is a common malignant tumor in the gastrointestinal tract. As the most common part of colorectal cancer, it has an incidence rate which is second only to gastric and esophageal cancer. At the same time, as one of the most famous tumors, it plays a vital role in gene and cell growth. Moreover, since the early performance of colon neoplasms is not obvious, many patients have reached the late stage of its discovery so that they missed the best treatment opportunity47. More seriously, more and more studies have shown that patients with colon neoplasms disease are on the increase year by year48. In addition, the associations between miRNA and colon neoplasms has been discovered and confirmed by more and more experimental researchers, which proves once again that miRNA plays an important role in colon neoplasms. Therefore, there is an urgent need to predict the potential miRNA associated with colon neoplasms. For example, miR-143 and miR-145 are both confirmed to continue to be downregulated during colon neoplasms production12. In addition, miR-17 and miR-106a, which have been deleted in colon neoplasms and shown to use E2F1 as a target mRNA and inhibit the growth of colon neoplasms49. Therefore, we selected colon neoplasms as a case study to further test the accuracy of our method for the purpose of predicting potential miRNA-disease associations. According to dbDEMC and miR2Disease’s evidence, 45 of the top 50 predicted miRNAs are successfully confirmed (see Table 7). For example, the association between hsa-miR-206 and colon neoplasms has been confirmed by previous literature50. This method found that hsa-miR-206 can participate in the targeting and regulation of SLC44A1 and KLF13, thus participate in the occurrence and metastasis of colon cancer.
Esophageal neoplasms is another epidemic cancer, which is a deadly disease and one of the most common digestive tract tumors51. Its prevalence is due to the current poor eating habits. At present, research on it is still rare in the world. The most common symptom of patients with esophageal neoplasms is dysphagia, which can lead to pain, vomiting, weight loss, etc52. The most common method currently used for this disease is chemotherapy. Where appropriate, chemotherapy allows patients to achieve the longest remission period and prolong the survival of some patients. Some studies have shown that miRNAs can be considered as effective prognostic biomarkers for esophageal neoplasms53. Therefore, case studies of Esophageal Neoplasms were conducted on our method to select the most likely-associated miRNAs. According to dbDEMC and miR2Disease’s evidence, 44 of the top 50 predicted miRNAs were verified (see Table 8). For example, the association between hsa-miR-182-5p and esophageal neoplasms has been confirmed by previous method54. This method identified two new tumor suppressor miRNA, including miR-182-5p and miR-455-5p, of which has-miR-182-5p was confirmed to be associated with esophageal cancer.
Breast neoplasms is a kind of malignant tumor formed by the uncontrolled growth of abnormal breast cells55. Each year, more than 211,000 cases of invasive breast cancer are diagnosed in the United States56. In most cases, breast cancer occurs in women, but it can also occur in men. More than 1,600 cases of male breast cancer are diagnosed each year. Breast cancer in women remains a major medical problem with major public health and social implications. At present, breast cancer has posed a threat to women’s physical and mental health57. In addition, numerous experiments have proved that many miRNAs are related to breast neoplasms. Case studies of Breast Neoplasms were conducted on our method to select the most likely-associated miRNAs. According to dbDEMC and miR2Disease’s evidence, 42 of the top 50 predicted miRNAs were verified (see Table 9).
Conclusions
Prediction of the associations between miRNA and disease can not only help us better understand the important role of miRNA in the generation and development of diseases, but also greatly promote the diagnosis and treatment of diseases. In this article, we proposed a new method to predict the potential associations between miRNA and disease by extracting the embedding representation of miRNAs and diseases from the heterogeneous information network. After that, we used the GraRep method to get the behavior information of miRNAs and disease in the network before combining their attribute information to represent miRNA and disease nodes, respectively. Then, we put the final data set into the Random Forest classifier for training and prediction. The final experimental results show that our method performs well and it is better than the methods of using only attribute information and methods using only behavior information. In addition, the results of the case study also prove that our method can predict the potential miRNA-disease associations well and the associated miRNA of a given disease. Therefore, we believe that the proposed method will be a useful and efficient tool for predicting miRNA-disease associations in the future. Besides, the working code explored in this article is available at https://github.com/jiboya123/working-code.git.
References
Esquela-Kerscher, A. & Slack, F. J. Oncomirs—microRNAs with a role in cancer. Nature reviews cancer 6, 259 (2006).
Ambros, V. microRNAs: Tiny Regulators with Great Potential. Cell 107, 823–826 (2001).
Ambros, V. The functions of animal microRNAs. Nature 431, 350 (2004).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. cell 116, 281–297 (2004).
Meister, G. & Tuschl, T. Mechanisms of gene silencing by double-stranded RNA. Nature 431, 343 (2004).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. cell 136, 215–233 (2009).
Cheng, A. M., Byrom, M. W., Shelton, J. & Ford, L. P. Antisense inhibition of human miRNAs and indications for an involvement of miRNA in cell growth and apoptosis. Nucleic acids research 33, 1290–1297 (2005).
Karp, X. & Ambros, V. Encountering microRNAs in cell fate signaling. Science 310, 1288–1289 (2005).
Miska, E. A. How microRNAs control cell division, differentiation and death. Current opinion in genetics & development 15, 563–568 (2005).
Alshalalfa, M. & Alhajj, R. Using context-specific effect of miRNAs to identify functional associations between miRNAs and gene signatures. BMC bioinformatics 14, S1 (2013).
Care, A. et al. MicroRNA-133 controls cardiac hypertrophy. Nature medicine 13, 613 (2007).
Wiemer, E. A. The role of microRNAs in cancer: no small matter. European journal of cancer 43, 1529–1544 (2007).
Latronico, M. V., Catalucci, D. & Condorelli, G. Emerging role of microRNAs in cardiovascular biology. Circulation research 101, 1225–1236 (2007).
Liu, Z., Sall, A. & Yang, D. MicroRNA: an emerging therapeutic target and intervention tool. International journal of molecular sciences 9, 978–999 (2008).
Lu, M. et al. An analysis of human microRNA and disease associations. PloS one 3, e3420 (2008).
Nelson, P. T. & Keller, J. N. RNA in brain disease: no longer just” the messenger in the middle. Journal of Neuropathology & Experimental Neurology 66, 461–468 (2007).
Zhu, H.-C. et al. MicroRNA-195 downregulates Alzheimer’s disease amyloid-β production by targeting BACE1. Brain research bulletin 88, 596–601 (2012).
Wang, X., Wu, X., Yan, L. & Shao, J. Serum miR-103 as a potential diagnostic biomarker for breast cancer. Nan fang yi ke da xue xue bao= Journal of Southern Medical University 32, 631–634 (2012).
Esquela-Kerscher, A. et al. The let-7 microRNA reduces tumor growth in mouse models of lung cancer. Cell cycle 7, 759–764 (2008).
Chen, R. W. et al. Truncation in CCND1 mRNA alters miR-16-1 regulation in mantle cell lymphoma. Blood 112, 822–829 (2008).
Miller, T. E. et al. MicroRNA-221/222 confers tamoxifen resistance in breast cancer by targeting p27Kip1. Journal of biological chemistry 283, 29897–29903 (2008).
You, Z.-H. et al. PBMDA: A novel and effective path-based computational model for miRNA-disease association prediction. PLoS computational biology 13, e1005455 (2017).
Chen, X. et al. BNPMDA: bipartite network projection for MiRNA–disease association prediction. Bioinformatics 34, 3178–3186 (2018).
Zheng, K. et al. MLMDA: a machine learning approach to predict and validate MicroRNA–disease associations by integrating of heterogenous information sources. Journal of translational medicine 17, 260 (2019).
Chen, X. et al. WBSMDA: within and between score for MiRNA-disease association prediction. Scientific reports 6, 21106 (2016).
You, Z.-H. et al. PRMDA: personalized recommendation-based MiRNA-disease association prediction. Oncotarget 8, 85568 (2017).
Jiang, Q. et al. Prioritization of disease microRNAs through a human phenome-microRNAome network. BMC systems biology 4, S2 (2010).
Shi, H. et al. Walking the interactome to identify human miRNA-disease associations through the functional link between miRNA targets and disease genes. BMC systems biology 7, 101 (2013).
Huang, Z. et al. HMDD v3. 0: a database for experimentally supported human microRNA–disease associations. Nucleic acids research 47, D1013–D1017 (2018).
Miao, Y.-R., Liu, W., Zhang, Q. & Guo, A.-Y. lncRNASNP2: an updated database of functional SNPs and mutations in human and mouse lncRNAs. Nucleic acids research 46, D276–D280 (2017).
Chou, C.-H. et al. miRTarBase update 2018: a resource for experimentally validated microRNA-target interactions. Nucleic acids research 46, D296–D302 (2017).
Chen, G. et al. LncRNADisease: a database for long-non-coding RNA-associated diseases. Nucleic acids research 41, D983–D986 (2012).
Davis, A. P. et al. The comparative toxicogenomics database: update 2019. Nucleic acids research 47, D948–D954 (2018).
Cheng, L. et al. LncRNA2Target v2. 0: a comprehensive database for target genes of lncRNAs in human and mouse. Nucleic acids research 47, D140–D144 (2018).
Wishart, D. S. et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic acids research 46, D1074–D1082 (2017).
Szklarczyk, D. et al. The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible. Nucleic acids research, gkw937 (2016).
Piñero, J. et al. DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic acids research, gkw943 (2016).
Kozomara, A., Birgaoanu, M. & Griffiths-Jones, S. miRBase: from microRNA sequences to function. Nucleic acids research 47, D155–D162 (2018).
Wang, D., Wang, J., Lu, M., Song, F. & Cui, Q. Inferring the human microRNA functional similarity and functional network based on microRNA-associated diseases. Bioinformatics 26, 1644–1650 (2010).
Cao, S., Lu, W. & Xu, Q. GraRep: Learning Graph Representations with Global Structural Information. In proceedings of CIKM, 891–900 (2015).
Liaw, A. & Wiener, M. Classification and regression by randomForest. R news 2, 18–22 (2002).
Friedl, M. A. & Brodley, C. E. Decision tree classification of land cover from remotely sensed data. Remote sensing of environment 61, 399–409 (1997).
Peterson, L. E. K-nearest neighbor. Scholarpedia 4, 1883 (2009).
Murphy, K. P. Naive bayes classifiers. University of British Columbia 18, 60 (2006).
Yang, Z. et al. dbDEMC 2.0: updated database of differentially expressed miRNAs in human cancers. Nucleic acids research 45, D812–D818 (2017).
Jiang, Q. et al. miR2Disease: a manually curated database for microRNA deregulation in human disease. Nucleic acids research 37, D98–D104 (2008).
Pita-Fernández, S. et al. Diagnostic and treatment delay, quality of life and satisfaction with care in colorectal cancer patients: a study protocol. Health and quality of life outcomes 11, 117 (2013).
Chong, V., Abdullah, M., Telisinghe, P. & Jalihal, A. Colorectal cancer: incidence and trend in Brunei Darussalam. Singapore medical journal 50, 1085 (2009).
Díaz, R. et al. Deregulated expression of miR‐106a predicts survival in human colon cancer patients. Genes, Chromosomes and Cancer 47, 794–802 (2008).
Gao, P., He, M., Zhang, C. & Geng, C. Integrated analysis of gene expression signatures associated with colon cancer from three datasets. Gene 654, 95–102 (2018).
Zhang, Y. Epidemiology of esophageal cancer. World journal of gastroenterology: WJG 19, 5598 (2013).
Javle, M. et al. Palliation of malignant dysphagia in esophageal cancer: a literature-based review. The journal of supportive oncology 4(365-373), 379 (2006).
Xu, X.-L. et al. MicroRNA-17, microRNA-18a, and microRNA-19a are prognostic indicators in esophageal squamous cell carcinoma. The Annals of thoracic surgery 97, 1037–1045 (2014).
Wang, Y. et al. MicroRNA expression in esophageal squamous cell carcinoma: Novel diagnostic and prognostic biomarkers. Molecular medicine reports 15, 3833–3839 (2017).
Dunning, A. M. et al. A systematic review of genetic polymorphisms and breast cancer risk. Cancer Epidemiology and Prevention. Biomarkers 8, 843–854 (1999).
Lal, G. et al. Extracellular matrix 1 (ECM1) expression is a novel prognostic marker for poor long-term survival in breast cancer: a Hospital-based Cohort Study in Iowa. Annals of surgical oncology 16, 2280–2287 (2009).
Saslow, D. et al. Clinical breast examination: practical recommendations for optimizing performance and reporting. CA: a cancer journal for clinicians 54, 327–344 (2004).
Acknowledgements
This work is supported by the NSFC Excellent Young Scholars Program, under Grants 61722212, in part by the National Science Foundation of China under Grants 61873212, 61861146002, 61732012, in part by the West Light Foundation of the Chinese Academy of Sciences, Grants 2017-XBZG-BR-001. The authors would like to thank the editors and anonymous reviewers for their reviews.
Author information
Authors and Affiliations
Contributions
B.Y.J. conceived and designed the experiment, processed the data set and discussed the results with the help of Z.H.Y. and L.C. J.R.Z., D.A. and L.P.L. prepared data and performed experiments. All the authors contributed to the text of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ji, BY., You, ZH., Cheng, L. et al. Predicting miRNA-disease association from heterogeneous information network with GraRep embedding model. Sci Rep 10, 6658 (2020). https://doi.org/10.1038/s41598-020-63735-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-63735-9
- Springer Nature Limited
This article is cited by
-
A miRNA-disease association prediction model based on tree-path global feature extraction and fully connected artificial neural network with multi-head self-attention mechanism
BMC Cancer (2024)
-
A multi-source molecular network representation model for protein–protein interactions prediction
Scientific Reports (2024)
-
Synchronous Mutual Learning Network and Asynchronous Multi-Scale Embedding Network for miRNA-Disease Association Prediction
Interdisciplinary Sciences: Computational Life Sciences (2024)
-
Unsupervised Graph Representation Learning with Inductive Shallow Node Embedding
Complex & Intelligent Systems (2024)
-
Comprehensive applications of the artificial intelligence technology in new drug research and development
Health Information Science and Systems (2024)