Abstract
In mixed infections, the bacterial susceptibility differs significantly compared to monocultures of bacteria, and generally the concentrations of antibiotics required for the treatment increases drastically. For S. aureus and P. aeruginosa dual species biofilms, it has been numerously reported that P. aeruginosa decreases S. aureus susceptibility to a broad range of antibiotics, including beta-lactams, glycopeptides, aminoglycosides, macrolides, while sensitizes to quinolones via secretion of various metabolites. Here we show that S. aureus also modulates the susceptibility of P. aeruginosa to antibiotics in mixed cultures. Thus, S. aureus—P. aeruginosa consortium was characterized by tenfold increase in susceptibility to ciprofloxacin and aminoglycosides compared to monocultures. The same effect could be also achieved by the addition of cell-free culture of S. aureus to P. aeruginosa biofilm. Moreover, similar increase in antibiotics efficacy could be observed following addition of S. aureus suspension to the P. aeruginosa mature biofilm, compared to P. aeruginosa monoculture, and vice versa. These findings open promising perspectives to increase the antimicrobial treatment efficacy of the wounds infected with nosocomial pathogens by the transplantation of the skin residential microflora.
Similar content being viewed by others
Introduction
Bacterial fouling is an important factor that strongly affects acute and chronic wounds healing and prevents wound scratch closure1. Besides the physical obstruction of the cells, pathogenic bacteria produce various virulence factors including toxins and proteases that also affect cytokine production by keratinocytes, induce apoptosis of the host cells and cause inflammation2,3,4,5,6.
S. aureus and P. aeruginosa are one of the most widespread pathogens causing various nosocomial infections, including pneumonia on the cystic fibrosis background, healthcare associated pneumonia and chronic wounds7,8,9,10. During infection, bacterial cells are embedded into a self-produced extracellular matrix of organic polymers this way forming either mono- or polymicrobial biofilms11,12 which drastically reduce their susceptibility to both antimicrobials and the immune system of the host13,14. Current data suggests that bacterial pathogenicity is promoted during polymicrobial infections and recovery is delayed in comparison with monoculture infections15,16,17. Accordingly, interspecies interactions between S. aureus and P. aeruginosa within mixed biofilms attracted major attention in recent years including both in vitro15,18,19,20 and in vivo studies16,21.
P. aeruginosa is known as a common dominator in polymicrobial biofilm-associated infections due to multiple mechanisms allowing its rapid adaptation to the specific conditions of the host. In particular, P. aeruginosa produces multiple molecules to compete with other microorganisms for space and nutrients. The main anti-staphylococcal tools of P. aeruginosa are siderophores and 2-n-heptyl-4-hydroxyquinoline N-oxide (HQNO), the inhibitor of the electron transport chain of S. aureus. Their presence shifts S. aureus to a fermentative mode of growth, eventually leading to reduced S. aureus viability22,23,24,25,26, forces increased S. aureus biofilm formation27,28 and transition of S. aureus into small-colony variants (SCVs)29. SCV is a well-characterized phenotype detected in various diseases, including cystic fibrosis and device-related infections27,28,29,30. SCVs appear as small, smooth colonies on a culture plate and grow significantly slower compared to wild type colonies. Remarkably, switch to the SCV phenotype improves the survival of S. aureus under unfavorable conditions, as it exhibits increased resistance to beta-lactams, glycopeptides, aminoglycosides, macrolides, as well as intracellular survival29,30,31,32,33. By contrast, the P. aeruginosa cell-free culture liquid increased the sensitivity of S. aureus biofilms to multiple antimicrobial compounds, including fluoroquinolones and membrane-targeting antibacterial agents and antiseptic chloroxylenol, while promotes the tolerance to beta-lactams, glycopeptides, aminoglycosides, macrolides19.
P. aeruginosa was shown to suppress S. aureus during co-culture in vitro in both planktonic and biofilm forms34,35,36,37 and has been observed as the dominant pathogen in S. aureus-P. aeruginosa mixed infections16. Despite of the antagonistic relationship of S. aureus and P. aeruginosa38,39, many studies reported their mutual association both in vitro18,20 and in acute and chronic wounds embedded in a mixed biofilm8,40,41,42,43,44, with S. aureus typically residing on the wound surface, whereas P. aeruginosa being rather observed in the deep layers15,44,45,46,47. Interestingly, in mixed P. aeruginosa—S. aureus biofilms from cystic fibrosis patients S. aureus was shown to be dominating during childhood, with P. aeruginosa prevalence increasing with aging and worsening patient prognosis48,49,50. During the biofilm formation P. aeruginosa produces three main exopolysaccharides, namely alginate, Pel, and Psl, which form an extracellular matrix in the biofilm exhibiting both structural and protective functions51,52,53,54. Under prevalent Pel secretion, loose biofilm structures are formed55 and thus S. aureus is able to penetrate into the biofilm55. It has been recently suggested that frequent false-negative detection of S. aureus and P. aeruginosa together in diagnostic cultures of sputum of cystic fibrosis patients could be attributed to the existence of S. aureus as SCVs that are more difficult to detect due to their small size and fastidious growth requirements30,32.
The impact of S. aureus on P. aeruginosa susceptibility to antimicrobials is poorly described. S. aureus secretes various exoproducts including adhesines, enzymes and polysaccharides and peptides56. Among them, the abundantly secreted staphylococcal protein A (SpA) was shown to inhibit the biofilm formation by P. aeruginosa clinical isolates via affecting type IV pili and the exopolysaccharide Psl57. Nevertheless, the effect of these metabolites on antibiotic susceptibility of other bacteria remains unexplored.
Investigations on alternative treatment options against biofilm-associated infections are largely based upon using specialized agents (such as quaternary ammonium compounds, curcumin or chlorquinaldol) or enzymatic treatment that in combinations with antibiotics provide high local drug concentrations avoiding systemic adverse effects58,59,60,61,62,63,64. While many approaches to targeting staphylococcal biofilms were reported62,65,66,67,68,69, only few successive ways of targeting P. aeruginosa are known64,70,71,72. Among various compounds exhibiting anti-biofilm activities, the derivatives of 2(5H)-furanone have been reported to inhibit biofilm formation by Staphylococci73,74,75,76,77. While many of these approaches exhibited promising results against staphylococcal monocultures, their efficiency against polymicrobial biofilms remains questionable.
Here we demonstrate that S. aureus also modulates the susceptibility of P. aeruginosa in mixed biofilms. Thus, the efficiency of antimicrobials active against both bacterial species like ciprofloxacin and aminoglycosides in mixed biofilms increased nearly tenfold in comparison with corresponding monocultures and the very same effect could be obtained in presence of cells-free supernatants of S. aureus when added to mature P. aeruginosa biofilms. These data suggest bi-directional influences of both bacteria on their antibiotic susceptibility in mixed infections, the fact that should be taken in account when considering an optimized strategy of polymicrobial infections treatment.
Results
Modeling the S. aureus–P. aeruginosa mixed biofilm
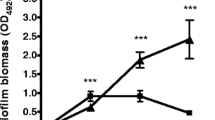
Despite of known antagonistic interactions between S. aureus and P. aeruginosa39, they are still the most common pathogens evoking wound infections and forming mixed biofilms on their surfaces41,42,44. We have simulated in vitro different situations where either S. aureus suspension was added to the preformed 24-h old biofilm of P. aeruginosa or, vice versa, P. aeruginosa was added to the preformed 24-h old biofilm of S. aureus. As a control, both strains were inoculated simultaneously and grown for 48 h with the broth exchange after 24 h of cultivation. Both S. aureus and P. aeruginosa were able to penetrate into the preformed biofilm of the other bacterium (Fig. 1). Irrespective of which bacterium initially preformed the biofilm and which one was added later, the ratio of their CFUs in the biofilm after 24 h cultivation remained around 1:10 with the prevalence of the first biofilm former (Fig. 1A,B), and was 1:1 when both bacteria were inoculated simultaneously (Fig. 1C).
In vitro simulation of the S. aureus-P. aeruginosa mixed biofilm. (A) P.aeruginosa suspension in a fresh broth was added to the preformed 24-h old biofilm of S. aureus or (B) S. aureus was added to the preformed 24-h old biofilm of P. aeruginosa and cultivation was continued for the next 24 h. (C) As a control, both strains were inoculated simultaneously and were grown for 48 h with the broth exchange after 24 h of cultivation. The biofilms were then assessed with either crystal-violet staining of wells bottom (upper lane) or differential CFUs counting. The median values with IQRs from six independent measurements are shown.
Next to analyze the biofilm structure and cells distribution in the matrix, the S. aureus—P. aeruginosa mixed biofilm was grown in imaging cover slips, live/dead stained with SYTO9/PI and analyzed with confocal laser scanning microscopy. Both S. aureus and P. aeruginosa formed 20–25 µm-thick biofilms when growing as monocultures (Fig. 2A,B). While the mixed biofilm was of similar thickness, it appeared more rigid in comparison with monoculture ones and the fraction of non-viable cells was similar to monocultures (compare Fig. 2A,B and C and Fig. S1A) suggesting stability of S. aureus—P. aeruginosa consortium under the conditions used. By using differential staining of S. aureus and P. aeruginosa (Fig. 2D and Fig. S1A) we have also analyzed the distributions of S. aureus (red-stained) and P. aeruginosa (blue-stained) over the biofilm layers and evaluated their relative fractions in each layer (Fig. 2E). Interestingly, in the mixed biofilm, S. aureus appeared as microcolonies within the biofilm (see white arrow in Fig. 2C,D) tended to distribute in the upper layers of the biofilm, while P. aeruginosa dominated in its lower layers (see Fig. 2E).
Mono- and polymicrobial biofilms formed by S. aureus and P. aeruginosa. Cells were grown without any antimicrobial (A–F) or in presence of 2(5H)-furanone derivative (F105, G–L) specifically inhibiting the biofilm formation by S. aureus and exhibiting no effects on P. aeruginosa. The 48-h old biofilms were stained by Syto9/PI (A–C, G–I) or ViaGram Red+ (D, J) to differentiate S. aureus (stained in red) and P. aeruginosa (stained in blue) and assessed by CLSM. (E, K) The distribution of S. aureus and P. aeruginosa in the biofilm layers expressed as their relative fractions. The repression of S. aureus ica-operon in mixed biofilm by F105 was monitored by detection of GFP in ica-GFP strain (F, L). The scale bars indicate 10 µm. S. aureus microcolonies in a mixed biofilm are shown by arrows.
In the last decades different approaches to inhibit the biofilm formation by various bacteria were developed59,60,62, appearing nowadays more successful in prevention of S. aureus biofilm formation65,66,68. Therefore we simulated the S. aureus—P. aeruginosa mixed biofilm formation under the conditions of biofilm-preventing treatment. For that, bacteria were cultivated in the presence of a derivative of 2(5H)-furanone denoted as F105, identified in recent study as an efficient inhibitor of biofilm formation by S. aureus77,78 , while exhibiting no significant effect against P. aeruginosa (Table S1). When S. aureus was grown for 48 h in the presence of 2.5 µg/ml of F105, no biofilm was formed and cells growth was delayed, while most of the cells remained viable in both surface-adherent (Fig. 2G) and planktonic state (Fig. S2A). As well, the repression of the biofilm formation was confirmed by evaluation of the ica-GFP expression in presence of F105. No GFP was observed in cells grown in presence of F105 in S.aureus pRB-ica-gfp (Fig. S3C, D), while the constitutive expression of GFP in S.aureus pC-tuf-gfp79 was not affected under these conditions (Fig. S4B, D) confirming the inhibition of the biofilm formation. As expected, no significant effect of F105 on P. aeruginosa viability could be observed (Fig. 2H, Table S1). Moreover, the biofilm formation was slightly increased as determined by crystal violet staining (Fig. S5) and CLSM (compare Fig. 2B,H).
When S. aureus and P. aeruginosa were grown together in the presence of F105, S. aureus microcolonies were also observed, similarly to the control (compare Fig. 2D,J). No GFP in S. aureus pRB-ica-gfp was observed (Fig. 2L), suggesting that in the presence of F105 S. aureus cells are able to form clusters inside the biofilm, presumably dominated by P. aeruginosa, despite of its antagonistic pressure (see white arrows on Fig. 2I,J). Next, in marked contrast to the control, S. aureus cells were observed predominantly in the middle layers of the biofilm (compare Fig. 2E,K).
The microscopic data were also validated by CFU counting in the biofilm; by using mannitol salt agar plates and cetrimide agar plates the bacterial species were differentiated and their CFUs were counted separately (Fig. S2). In the presence of F105 the amount of adherent viable S. aureus cells decreased by six orders of magnitude in monoculture, suggesting complete inhibition of the biofilm formation, while no significant differences in CFUs of P. aeruginosa could be observed. In a mixed biofilm, the S. aureus to P. aeruginosa ratio remained unchanged in the control, while the fraction of viable S. aureus cells decreased slightly in the presence of F105, this way confirming CLSM data and supporting the hypothesis that S. aureus is apparently able to embed and hide thereby in the biofilm formed by P. aeruginosa when its own biofilm formation is repressed.
Atomic force microscopy
The atomic force microscopy of both monocultures and mixed biofilms of S. aureus—P. aeruginosa confirmed the CLSM data. Thus, in control wells the biofilms of monocultures of both strains formed a typical confluent multilayer biofilm (Fig. 3A,B), in mixed biofilm S. aureus was prevalently distributed in the upper layers (Fig. 3C). Interestingly, the adhesion force of the mixed biofilm was threefold lower compared to S. aureus monoculture biofilm and twofold lower compared to P. aeruginosa monoculture biofilm (Fig. 4), suggesting more irregular structure of the mixed biofilm55. When growing with F105, only P. aeruginosa could be observed on the biofilm surface in the mixed culture, suggesting that S. aureus was hidden into the lower biofilm layers. Since the adhesion force of the mixed biofilm in the presence of F105 was similar to that one in the monoculture P. aeruginosa (Figs. 3F, 4), we assumed that the biofilm matrix under these conditions was presumably formed by P. aeruginosa.
Atomic force microscopy (Peak Force Tapping mode) of mono- and polymicrobial biofilms formed by S. aureus and P. aeruginosa. Cells were grown without any antimicrobials (A–C) or in presence of F105 specifically inhibiting the biofilm formation by S. aureus cells (D–F) for 48 h, then the plates were washed, fixed with glutardialdehyde and analyzed with AFM. (I) Sensor height (topography); (II) 3D reconstruction of height channel image; (III) adhesion.
The adhesion force of S. aureus and P. aeruginosa monoculture and mixed biofilms. To repress the biofilm formation F105 was added up to 2.5 µg/ml. Asterisks denote statistical significant difference was confirmed by the Kruskal–Wallis statistical test at p < 0.05. The average values with SDs from 4 independent measurements are shown.
S. aureus and P. aeruginosa susceptibility to antibiotics in mixed cultures
The effect of various conventional antibiotics on preformed mono- and polymicrobial biofilms was studied. The 48-h old monoculture and mixed biofilms were prepared in 24-well adhesive plates in either absence or presence of F105 to repress the biofilm formation by S. aureus itself. Then the biofilms were washed with sterile 0.9% NaCl and wells were loaded with fresh broth supplemented with antibiotics at wide range of final concentrations to fill the range of their 1–16 × MBCs (see table S1 for MBC values). After 24 h incubation the amount of CFUs of both S. aureus and P. aeruginosa in both the culture liquid and the biofilm was determined by the drop plate assay and the distribution of cells in the mixed biofilm was assessed by CLSM.
First, the biofilm-eradicating activity was investigated for vancomycin, tetracycline, ampicillin and ceftriaxone, antibiotics conventionally used for S. aureus treatment but typically inefficient against P. aeruginosa (Figs. 5, S6, S7). In monoculture, vancomycin reduced the amount of viable S. aureus cells in the biofilm by 3 orders of magnitude only at 16 × MBC (Fig. 5A). In the culture liquid, 0.125 × MBC (corresponds to 1 × MIC, see Table S1) of vancomycin was sufficient to decrease of CFUs count by three orders of magnitude (Fig S6). Nevertheless, no complete death of cells could be observed. Apparently, this could be attributed to the protection of detached cell clumps where cells could be partially protected by residual EPS (Fig. S8). As expected, in the presence of F105 (2.5 µg/ml) neither biofilms nor cell clumps could be observed (Fig. S8), and bacteria were found completely dead both in the biofilm and in the culture liquid after 24-h exposition to the antibiotic at 1 × MBC (Fig. 5C, S6). Irrespective of either presence or absence of F105, P. aeruginosa remained resistant to the antibiotic (Fig. 5B,D).
The effect of vancomycin on viability of S. aureus and P.aeruginosa embedded into their mono- and polymicrobial biofilms. Vancomycin was added to 48 h-old biofilms grown in absence (A–B, E–F) or presence (C–D, H–J) of F105 to inhibit the biofilm formation by S. aureus. After 24 h incubation, the biofilms were washed twice with sterile 0.9% NaCl. The adherent cells were scratched, resuspended and CFUs were counted. The median values with IQRs from six independent measurements are shown. Asterisks denote statistical significant difference was confirmed by the Kruskal–Wallis statistical test at p < 0.05.Alternatively, biofilms were stained by Syto9/PI and biofilms were assessed by CLSM. The scale bars indicate 10 µm.
In a mixed culture, irrespective of the S. aureus biofilm formation repression by F105, viable S. aureus cells were identified within the biofilm, indicating that the efficiency of antibiotic reduced drastically (Fig. 5A,C, compare reds and violets). Statistical significance of this discrepancy was confirmed by the Kruskal–Wallis statistical test at p < 0.05. Similar inefficiency of vancomycin against S. aureus in mixed cultures was observed in the culture liquid (Fig. S6).
For a deeper understanding of the localization distribution and viability of bacteria in mixed biofilms under vancomycin treatment, the CLSM analysis has been performed. In the presence of F105 no S. aureus biofilm could be observed resulting in significant decrease of viable cells fraction after vancomycin treatment, in contrast to the biofilm-embedded cells (compare Fig. 5E,H). In the mixed biofilm, irrespective of the F105 treatment, S. aureus microcolonies with viable cells could be observed in the biofilm, similarly to the control (compare Fig. 2C, I with Fig. 5G, J and Fig. 2D, J with Fig. 6). In marked contrast to the control, where S. aureus was mostly localized in the top layers of the biofilm, after vancomycin treatment most of the S. aureus cells appeared in the lower and middle layers of the biofilm (compare Fig. 2E and Fig. 6), and mostly remained viable (Fig. S9).
The effect of vancomycin on S. aureus and P. aeruginosa distribution in mixed biofilms. Cells were grown in absence (A, C) or in presence of F105 specifically inhibiting the biofilm formation by S.aureus cells (B, D). Vancomycin (256 µg/mL corresponding to 8 × MBC for S. aureus) was added to 48 h-old biofilms. After 24 h incubation, the biofilms were stained by ViaGram Red+ to differentiate S. aureus (stained in red) and P. aeruginosa (stained in blue) and assessed by CLSM. The scale bars indicate 10 µm. (C, D) The distribution of S. aureus and P. aeruginosa in the biofilm layers are expressed as their relative fractions.
Similarly to vancomycin, treatment by ampicillin, tetracycline and ceftriaxone was almost inefficient against biofilm-embedded S. aureus, while under conditions of biofilm formation repression by F105, the 1–2 × MBC of antimicrobials led to the complete death of cells in 24 h (Fig. S7). In culture liquids, 0.25–0.5 × MBC of ampicillin and ceftriaxone provided the full death of S. aureus, 0.125 × MBC of tetracycline decreased the amount of viable S. aureus cells by 3 orders of magnitude. F105 reduced the amount of antimicrobials sufficient to kill bacteria completely by a factor of 4 (Fig. S7 A, C). In the mixed culture, even despite of the S. aureus biofilm formation repression with F105, S. aureus cells remained insensitive to any of antimicrobials tested in both culture liquid and biofilm (Fig. S7 A, C). Under double treatment by F105 and antimicrobials, the prevalence of P. aeruginosa in the biofilm was observed, in agreement with the CFU count data (Figs S7 and S10). Earlier, 2-n-heptyl-4-hydroxyquinoline N-oxide (HQNO) and siderophores pyoverdine and pyochelin produced by P. aeruginosa have been shown to decrease the susceptibility of S. aureus to vancomycin, beta-lactams and cephalosporines33, apparently explaining low efficiency of antibiotics on detached cells (Figs S6, S7). Nevertheless, CLSM analysis indicated considerable redistribution of S. aureus from preferred topical localization to the bottom layers of the biofilm (Fig. S10). Therefore, one could assume that P. aeruginosa in the upper layers of the biofilm apparently prevented the penetration of the antibiotic into the matrix this way being the complementary factor of S. aureus resistance to the treatment.
Next, we investigated the effect of broad-spectrum antimicrobials such as ciprofloxacin, amikacin and gentamycin which are similarly active against both S. aureus and P. aeruginosa (see Table S1). In contrast to the previous group of antimicrobials, high concentrations of ciprofloxacin efficiently eradicated both P. aeruginosa and S. aureus monocultures even in the biofilm-embedded form (Fig. 7A, B). Interestingly, when the mixed biofilm was treated, nearly tenfold lower concentration of antimicrobial was sufficient to obtain similar reduction of P. aeruginosa CFUs in the biofilm, although not for detached cells (Fig. 7B, Fig S11). Moreover, significant increase of susceptibility of both biofilm-embedded and detached S.aureus to ciprofloxacin was observed in mixed culture in contrast to monoculture (see Figs. 7A, S11 A). Finally, in the mixed biofilm the complete death of both P. aeruginosa and S. aureus could be achieved at 8 × MBC of ciprofloxacin, in marked contrast to monocultures (Fig. 7A, B).
The effect of ciprofloxacin on viability of S. aureus and P.aeruginosa embedded into their mono- and polymicrobial biofilms. Ciprofloxacin was added to 48 h-old biofilms grown in absence (A–B, E–F) or presence (C–D, H–J) of F105 to inhibit the biofilm formation by S. aureus. After 24 h incubation, the biofilms were washed twice with sterile 0.9% NaCl. The adherent cells were scratched, resuspended and CFUs were counted. The median values with IQRs from six independent measurements are shown. Asterisks denote statistical significant difference was confirmed by the Kruskal–Wallis statistical test at p < 0.05. Alternatively, biofilms were stained by Syto9/PI and biofilms were assessed by CLSM. The scale bars indicate 10 µm.
Similarly, 1–2 × MBC of aminoglycosides (amikacin or gentamicin) led to the complete death of both P. aeruginosa and S. aureus in mixed biofilm while reducing their CFUs in monocultures only by 2–3 orders of magnitude at 8 × MBCs (Fig. 8). For detached S. aureus cells the effect was less pronounced (Fig. S12), while the susceptibility of P. aeruginosa to both aminoglycosides in the culture liquid was slightly increased in contrast to ciprofloxacin (Fig. S11).
The effect of amikacin and gentamycin on viability of S. aureus and P.aeruginosa in mono- and polymicrobial biofilms. Antimicrobials were added to 48 h-old biofilms grown in absence (A–B) or presence (C–D) of F105 to inhibit the biofilm formation by S. aureus. After 24 h incubation, the biofilms were washed twice with sterile 0.9% NaCl. The adherent cells were scratched, resuspended and CFUs were counted. The median values with IQRs from six independent measurements are shown. Asterisks denote statistical significant difference was confirmed by the Kruskal–Wallis statistical test at p < 0.05.
In the presence of F105, just 0.5 × MBC of any tested antimicrobial was already sufficient for the complete death of S. aureus both detached and biofilm-embedded cells (see Fig. 5C and Fig. 7C), similarly to vancomycin, tetracycline, ampicillin and ceftriaxone (see Fig. 5 and Fig. S7). The presence of F105 did not affect the susceptibility of monoculture P. aeruginosa biofilm to antibiotics. In contrast, in mixed cultures the inhibition of S. aureus by F105 restored the susceptibility of P. aeruginosa back to the monoculture level, suppressing the observed high efficiency of antimicrobials against this bacterium in the mixed biofilm (compare pallets B and D in Figs. 7, 8, S11, S12). Surprisingly, the efficiency of aminoglycosides against S. aureus in mixed culture did not depend on presence of F105 (pallets C in Figs. 7, 8, S11, S12).
The CLSM analysis of S. aureus and P. aeruginosa monoculture and mixed biofilms treated with ciprofloxacin confirmed the CFUs counting data. In particular, while 8 × MBC did not affect either S. aureus or P. aeruginosa cells in monoculture biofilms (Fig. 7E,F), in the mixed biofilm a huge fraction of non-viable cells was observed (Fig. 7G). In marked contrast, repression of the S. aureus biofilm production by F105 led to a reversal with most P. aeruginosa cells green-stained while S. aureus identified as non-viable in mixed culture (Fig. 7J).
The distribution of bacteria in the mixed biofilm layers under treatment with ciprofloxacin was also assessed (Fig. 9). In contrast to vancomycin treatment, here S. aureus dominated in the upper layers of the mixed biofilm (compare Fig. 6 and 9 A and C) and remained alive, while P. aeruginosa were presumably dead (see Figs S9, S13, S14) suggesting no reversal protection of P. aeruginosa by S. aureus biofilm. On the other hand, double treatment by ciprofloxacin combined with F105 resulted in hiding of S. aureus in the bottom layers of the biofilm and increased resistance of P. aeruginosa. Treatment by amikacin and gentamycin led to considerably different distributions of bacteria over the biofilm layers with the prevalence of S. aureus in the bottom layers irrespective of its biofilm repression by F105 (Fig. S14, cells distribution patterns) but qualitatively similar bacterial survival patterns (see Fig. S12). Moreover, under single antibiotic treatment P. aeruginosa were presumably dead, while S. aureus remained viable (Fig. S14). In the presence of F105 P. aeruginosa remained alive and much less S. aureus cells could be observed in the biofilm, as almost all of them were identified as non-viable.
The effect of ciprofloxacin on S. aureus and P. aeruginosa distribution in mixed biofilms. Cells were grown normally (A, C) or in presence of F105 specifically inhibiting the biofilm formation by S. aureus cells (B, D). Ciprofloxacin (512 µg/mL corresponding to 8 × MBC for both bacteria) was added to 48 h-old biofilms. After 24 h incubation, the biofilms were stained by ViaGram Red+ to differentiate S. aureus (stained in red) and P. aeruginosa (stained in blue) and assessed by CLSM. The scale bars indicate 10 µm. (C, D) The distribution of S. aureus and P. aeruginosa in the biofilm layers are expressed as their relative fractions.
Recent data indicate that S. aureus forms so-called Small Colony Variants (SCV) when growing with P. aeruginosa or when treated with aminoglycosides due to the defects in respiration27,28,29,30,33. Cells with SCV-phenotype are much more robust against external stresses compared to regular S. aureus cells, and thus the observed drastic decrease of S. aureus susceptibility to antimicrobials in mixed biofilms could be potentially attributed to the SCV formation. To verify this hypothesis, next the CFUs were counted by plating the cells on LB-agar broth with colistin, which specifically kills P. aeruginosa but does not affect S. aureus. While the SCVs constituted up to 90% of S. aureus population (Fig. S15), that fits with previous reports31, the effect of higher efficiency of broad-spectrum antibiotics against mixed biofilms could nevertheless be observed (Fig. S15).
This fact together with the observation that the repression of S. aureus by F105 abrogated the effect of higher efficiency of aminoglycosides against P. aeruginosa in mixed biofilms indicates that the increase of broad-spectrum antibiotics efficacy against P. aeruginosa and S. aureus in mixed biofilms is less related or unrelated to SCV formation and rather governed by interspecies interaction between these bacteria. One of the main antagonistic tools of P. aeruginosa is the production of cyanide80. To test whether it could be one of mechanisms increasing the efficiency of antimicrobials in mixed cultures, the experiments have been performed by using S.aureus 0,349 pCXcydABsa80 strain insensitive to cyanide. Despite of cyanide resistance, similar drastic reduction of S. aureus CFU in mixed biofilms could be observed when treated with ciprofloxacin and aminoglycosides also ruling out the cyanide synthesis by P. aeruginosa as the key contributing mechanism and indicating that other factors lead the observed effects (Fig. S16).
Extracellular metabolites of S. aureus sensitize P. aeruginosa to antimicrobials
Our data suggest that complex interspecies interactions between S. aureus and P. aeruginosa in mixed biofilm significantly affect bacterial susceptibility to antimicrobials with different specificity. While the antagonistic tools of P. aeruginosa are well described to the date, possible mechanisms of S. aureus–P. aeruginosa interactions remain unexplored. Since the F105 led to the restoration of P. aeruginosa susceptibility to aminoglycosides in mixed biofilm back to the monoculture level, we assumed that S. aureus produces certain metabolites that sensitize P. aeruginosa to antimicrobials. To test this assumption, 48 h-old P. aeruginosa biofilms were washed and wells were filled with cells-free culture liquids of S. aureus or P. aeruginosa–S. aureus mixed culture supplemented with antimicrobials in their respective 0.03–8 × MBCs. As a control, a fresh nutrient broth has been used. After 24 h the viability of detached and biofilm-embedded P. aeruginosa cells were quantified by CFUs count.
As one can see from Fig. 10, cells-free supernatant of S. aureus monoculture led to significant drop in P. aeruginosa viability, leading to a complete death of cells already at 2 μg/ml concentrations for all antimicrobials tested. At the same time, cells-free supernatant of P. aeruginosa-S. aureus mixed culture resulted in a considerably weaker effect, while being comparable with co-culture experiments (compare Figs. 7, 8, S11, S12 and 10), also suggesting that the observed effect is not due to nutrients depletion and consequent starvation. Since P. aeruginosa also forms SCV-phenotype in presence of aminoglycosides81,82, plates were incubated additional five day to let growth the cells with suppressed metabolism. No significant increase of CFUs has been observed (twofold to threefold increase of colonies count in last dilution with the visible growth, not shown), suggesting bactericidal effect of S. aureus culture liquid.
The effect of S. aureus and S. aureus-P.aeruginosa cell-free culture liquid on viability and susceptibility of P.aeruginosa to antimicrobials. Fresh nutrient broth (red), cells-free culture liquids of S. aureus (black) or P. aeruginosa-S. aureus mixed culture (blue) supplemented with antimicrobials in their respective 0.03–8 × MBCs were added to 48 h-old biofilms of P. aeruginosa. After 24 h incubation, the biofilms were washed twice with sterile 0.9% NaCl. The adherent cells were scratched, resuspended and CFUs were counted. The median values with IQRs from six independent measurements are shown. Asterisks and double asterisks denote statistical significant difference between fresh nutrient broth and cell-free supernatant of S. aureus culture or cell-free supernatant of S. aureus-P.aeruginosa co-culture respectively (confirmed by the Kruskal–Wallis statistical test at p < 0.05).
In earlier works the effects of P. aeruginosa metabolites on S. aureus susceptibility to antimicrobials have been shown to be strain-specific83. Therefore, the observed effects of S. aureus culture liquid have been tested on various clinical strains of P. aeruginosa and S. aureus. Firstly, two-fold serial dilutions of a cell-free culture liquid of S. aureus ATCC 29,213 were prepared and added to 48-h old cultures of P. aeruginosa and after 24 h the viability of detached and biofilm-embedded P. aeruginosa cells has been evaluated in resazurin test (Fig S17, A, B). Various P. aeruginosa strains exhibited different sensitivity, with ATCC 27853D-5 strain being the most sensitive to the culture of S. aureus. Therefore, P. aeruginosa ATCC 27853D-5 has been used as a test strain in reverse experiment, where the anti-pseudomonas potential of extracellular metabolites produced by various clinical isolates S. aureus of has been tested. Here, the highest activity has been observed for S. aureus ATCC 29,213 strain, while other clinical isolates were also able to produce the anti-pseudomonas metabolite(s).
Taken together, these data clearly demonstrate that S. aureus produces extracellular metabolites which suppress the growth of P. aeruginosa and potentiate ciprofloxacine and aminoglycosides against P. aeruginosa biofilms.
Intervention of P. aeruginosa into S. aureus biofilm and vice versa as a possible way to enhance antimicrobial susceptibility
Our results indicate that under appropriate conditions both S. aureus and P. aeruginosa due to their antagonistic interactions appear more susceptible to broad-spectrum antimicrobials in polymicrobial biofilms, compared to their monoculture counterparts. Based on these data, we have suggested that also the susceptibility of monoculture biofilms could be increased by deliberate intervention of P. aeruginosa into preformed S. aureus biofilm, and vice versa.
To verify the efficacy of this approach, P. aeruginosa suspension (106 CFU/mL) was added to the 24 h-old S. aureus biofilm and bacteria were incubated for the next 24 h. Then the biofilm was washed by sterile saline and fresh broth containing different antimicrobials was added into the wells. After 24 h the number of P. aeruginosa and S. aureus CFUs was counted by using differential media.
The introduction of P. aeruginosa into S. aureus biofilm did not change the efficacy of any antibiotic against P. aeruginosa itself (Fig. 11, lane II). In contrast, 1 × MBC of ciprofloxacin led to the reduction of viable S. aureus in biofilm by 3 orders of magnitude, while in the monoculture 4–8 × MBC was required to achieve the same effect (Fig. 11, lane I, compare reds and violets). Amikacin and gentamycin, being almost inefficient against S. aureus monoculture biofilm up to 8 × MBC, were able to decrease the S. aureus CFUs in biofilm by 3 orders of magnitude already at 1–2 × MBC after introduction of P. aeruginosa with the most pronounced effect observed for gentamycin.
The susceptibility of P. aeruginosa and S. aureus after introduction of the antagonist into monoculture biofilms. (I–II) P. aeruginosa suspension in a fresh broth was added to the preformed 24-h old biofilm of S. aureus or (III–IV) S. aureus was added to the preformed 24-h old biofilm of P. aeruginosa and cultivation was continued for the next 24 h. Then antimicrobials were added and after 24 h incubation CFUs in biofilms were counted. The median values with IQRs from six independent measurements are shown. Asterisks show significant difference between CFUs number between monoculture and mixed biofilms.
In the reverse experiment, when S. aureus was added to the P. aeruginosa biofilm, a remarkable increase of ciprofloxacin efficacy against P. aeruginosa could be observed (Fig. 11, lane IV, compare blues and violets), while the susceptibility of S. aureus itself did not change significantly. The efficacy of aminoglycosides has increased only against S. aureus, while not against P. aeruginosa.
Discussion
Biofilm formation represents an important virulence factor of many bacteria, as the extracellular matrix drastically reduces their susceptibility to antimicrobials resulting in up to 1,000-fold higher tolerance to antibiotics of biofilm-embedded cells compared to their planktonic forms13,14,84. In contrast, polymicrobial communities are often characterized by concurrent interspecies interactions that significantly alter bacterial susceptibility to antimicrobials. Thus, it has been shown in several works that in S. aureus–P. aeruginosa mixed biofilms, the most common pathogenic agents causing various nosocomial infections7,8,9, various metabolites produced by P. aeruginosa increase the sensitivity of S. aureus biofilms to fluoroquinolones and membrane-targeting antibacterial agents and antiseptic chloroxylenol, while promotes the tolerance to beta-lactams, glycopeptides, aminoglycosides, macrolides19,25,83,85. Here we show that this effect is bilateral, and S. aureus also affects the susceptibility of P. aeruginosa to antibiotics in biofilms.
Despite of the antagonistic relationship between S. aureus and P. aeruginosa described in multiple studies38,39,86, these bacteria can be found in close association in acute and chronic wounds being embedded into mixed biofilms8,40,41,42,43,44. Our in vitro data show that the inoculation of S. aureus to the mature P. aeruginosa biofilm or vice versa leads to the formation of mixed biofilm, although with the prevalence of the first biofilm former (Fig. 1). The co-cultivation of both bacteria results in the formation of a more rigid biofilm, where S. aureus is located mainly in the upper layers, while P. aeruginosa can be found mostly in the lower layers of the biofilm (Fig. 2), in agreement with earlier data15,44,45,46,47.
We investigated the effect of two groups of antimicrobials on bacterial viability in mixed biofilms. The first group contained vancomycin, tetracycline, ampicillin and ceftriaxone that were active against S. aureus while leaving P. aeruginosa nearly unaffected. The second group included broad-spectrum antibiotics such as ciprofloxacin, gentamicin and amikacin that exhibited comparable MBC values against both studied bacteria (see Table S1). Additionally, we also simulated the biofilm-preventing treatment with earlier described compound F105, specifically affecting only S. aureus biofilm formation77. In control experiments with S. aureus monoculture biofilms, none of the antimicrobials exhibited any bactericidal effect at their 8–16 × MBCs, while 1 × MBC was already sufficient for the complete eradication of both adherent and detached cells under biofilm repression conditions with F105 (compare Figs. 5, 7, 8, S6, S7, S11, S12 reds on panels A and C). In addition, ciprofloxacin, gentamicin and amikacin at 8 × MBCs significantly reduced the number of CFUs of biofilm-embedded P. aeruginosa (Figs. 5, 7, 8, S6, S7, S11, S12 blues on panels B and D).
In mixed biofilms, S. aureus became significantly less susceptible to antimicrobials active specifically against S. aureus such as vancomycin, tetracycline, ampicillin and ceftriaxone irrespectively to the biofilm repressing agent F105 presence (see Figs. 5, S7). It has been reported previously that HQNO and siderophores produced by P. aeruginosa decrease the S. aureus susceptibility to various antibiotics33 that could explain the observed effect. Nevertheless, CLSM analysis revealed that S. aureus was re-localized under treatment with vancomycin, tetracycline, ampicillin to the middle and lower layers of the biofilm and remained viable (see Figs. 6, S9, S10). Notably, in mixed biofilms S. aureus formed cell clumps in the biofilm matrix (compare Fig. 2 C, D, I, J and Fig. 6). As well, the adhesion force of S. aureus–P. aeruginosa dual-species biofilm in the presence of F105 was similar to that of the P. aeruginosa monoculture biofilm (See Fig. 4), indicating that upper layers of the biofilm are apparently formed by P. aeruginosa, making them hardly permeable by antibiotics87,88. Therefore, the protection of S. aureus by the matrix of P. aeruginosa biofilm also cannot be excluded.
By contrast, when the S. aureus–P. aeruginosa mixed biofilms were treated with any of the broad-spectrum antimicrobials such as ciprofloxacin, gentamicin or amikacin, nearly tenfold lower concentrations were sufficient to achieve the same reduction in the CFUs number of both bacteria in the biofilm, in comparison with monoculture treatment (Figs. 7 and 8, compare violets with reds or blues on panels A and B). This effect was more pronounced for aminoglycosides, despite of low permeability of biofilm for them89. Both gentamicin and amikacin at already 1–2 × MBC led to the complete death of both biofilm-embedded and detached cells in dual-species biofilm, while in monocultures 8 × MBC was required to reduce the number of CFUs by 3–5 orders of magnitude (Figs. 8 and S12) despite of reported synergy of F105 with aminoglycosides77. In the meanwhile, the observed reduction of the S. aureus CFUs number was not a consequence of their transition into small colony variants in response to aminoglycosides and cyanide production by P. aeruginosa (see Figs S15, S16). The same effects were observed also for combinations of S. aureus–P. aeruginosa clinical isolates. Apparently, this effect could be attributed to rhamnolipids synthesis with P. aeruginosa, which increase S. aureus susceptibility to aminoglycosides83. In contrast to other works on antimicrobial susceptibility of S. aureus–P. aeruginosa dual species cultures, here we have shown that S. aureus also affects the susceptibility of P. aeruginosa biofilm-embedded cells to antimicrobials with the most pronounced effect for aminoglycosides. Moreover, tetracycline and ceftriaxone, while being inefficient against P. aeruginosa, at high concentrations significantly reduced the CFUs of this bacterium in the mixed biofilms (Fig. S7).
The reason of enhanced P. aeruginosa susceptibility remains discussible. In experiments with cell-free culture liquid of S. aureus the same effect could be observed (Fig. 10, S17), suggesting that presumably certain metabolite of S. aureus is responsible for that, while the exact active molecule remains to be identified.
Interestingly, under repression of the S. aureus biofilm formation by F105, the efficiency of ciprofloxacin and aminoglycosides against S. aureus did not change significantly, while the sensitivity of P. aeruginosa was restored to the level characteristic for its monoculture biofilm (Figs. 7 and 8, compare violets with reds or blues on panels C and D). This effect could be attributed to either the significant reduction of S. aureus fraction in the biofilm (see Fig. 9, S13, S14) or the repression of the antagonistic factors production by S. aureus due to complex changes in its cell metabolism in the presence of F10577,78. Nevertheless, the molecular basis of these complex interbacterial interactions that under certain conditions lead to a clear reversal in the antimicrobials susceptibility requires further investigations.
Finally, we have shown that S. aureus and P. aeruginosa are able to penetrate into each other’s mature biofilms (see Fig. 1 A and B) and by this intervention significantly affect the susceptibility of the mixed biofilm to antimicrobials (Fig. 11). When P. aeruginosa was introduced into S. aureus biofilm, all antimicrobials reduced the amount of CFUs of both bacteria in the biofilm by 3 orders of magnitude at 1–2 × MBC with more pronounced effect observed for gentamicin. In the reverse experiment, the inoculation of S. aureus to the mature P. aeruginosa biofilm significantly increased the efficacy of ciprofloxacin against P. aeruginosa.
From a broader perspective, we believe that artificial intervention of antagonistic bacteria into already preformed monoculture biofilms could be used to enhance their antimicrobial treatment efficacy. We suggest that this approach has a strong potential of further development towards innovative treatment of biofilm-associated infections such as introduction of the skin residential saprophytic microflora to the biofilms formed by nosocomial pathogens to increase antimicrobial treatment efficacy, as it has been suggested for the treatment of atopic dermatitis90. While in this work we demonstrated the synergy of interbacterial antagonism with antimicrobials using the well-studied S. aureus—P. aeruginosa model system, we believe that many other bacteria of normal body microflora are available to antagonize with nosocomial pathogens and thus can be used for the enhancement of microbial infections treatment by using microbial transplantation.
Materials and methods
Derivate of 2(5H)-furanone designed as F105 (3-chloro-5(S)-[(1R,2S,5R)-2-isopropyl-5-methylcyclohexyloxy]-4-[4-methylphenylsulfonyl]-2(5H)-furanone) was described previously77,91.
Bacterial strains and growth conditions
Staphylococcus aureus subsp. aureus (ATCC 29213) and Pseudomonas aeruginosa (ATCC 27853D-5) were used in this assay. Clinical isolates of S. aureus and P. aeruginosa strains were obtained from Republic Clinical Hospital (Kazan, Russia). The bacterial strains were stored in 10% (V/V) glycerol stocks at − 80 °C and freshly streaked on blood agar plates (BD Diagnostics) followed by their overnight growth at 35 °C before use. Fresh colony material was used to adjust an optical density to 0.5 McFarland (equivalent to 108 cells/mL) in 0.9% NaCl solution that was used as a working suspension. For the biofilm assay the previously developed BM broth (glucose 5 g, peptone 7 g, MgSO4 × 7H2O 2.0 g and CaCl2 × 2H2O 0.05 g in 1.0 L tap water)61,75,92 where both S. aureus and P. aeruginosa formed rigid biofilms in 2 days was used. The mannitol salt agar (peptones 10 g, meat extract 1 g, NaCl 75 g, D-mannitol 10 g, agar–agar 12 g in 1.0 L tap water, Oxoid) and cetrimide agar (Sigma) were used to distinguish S. aureus and P. aeruginosa, respectively, in mixed cultures. Bacteria were grown under static conditions at 35 °C for 24–72 h as indicated.
Construction of ica-gfp reporter construction
The 500-bp promoter region of icaA gene was amplified from the chromosomal DNA of S. aureus USA30093 by using primers icaA for and icaA rev (table S2), the gfp gene was amplified from the pCtuf-gfp plasmid31 by using gfp for and gfp rev (table S2). The PCR products were cloned into the expression vector pRB47394 digested by HindIII using an isothermal, single-reaction method for assembling multiple overlapping DNA molecules as described previously95 by obtaining a plasmid pRB-ica-gfp. The resulting plasmid was transformed in S. aureus ATCC 29,213 by electroporation as described previously96.
Biofilm assays
Biofilm formation was assessed in 24-well polystirol plates (Eppendorf) by staining with crystal violet as described earlier in97 with modifications. Bacteria with an initial density of 3 × 107 CFU/ml were seeded in 2 ml BM at 37 °C and cultivated for 48 h under static conditions. Then the culture liquid was removed and the plates were washed once with phosphate-buffered saline (PBS) pH = 7.4 and dried for 20 min. Then, 1 ml of a 0.5% crystal violet solution (Sigma-Aldrich) in 96% ethanol was added per well followed by incubation for 20 min. The unbounded dye was washed off with PBS. The bound dye was eluted in 1 ml of 96% ethanol, and the absorbance at 570 nm was measured on a Tecan Infinite 200 Pro microplate reader (Switzerland). Cell-free wells subjected to all staining manipulations were used as control.
The biofilms were additionally analyzed by confocal laser scanning microscopy (CLSM) on Carl Zeiss LSM 780 confocal microscope. Both mono- and mixed cultures of S. aureus and P. aeruginosa were grown on cell imaging cover slips (Eppendorf) under static conditions for 48 h in BM broth. Next one-half of the medium was replaced by the fresh one containing antimicrobials at final concentrations as indicated and cultivation was continued for the next 24 h. The samples were then stained for 5 min with the SYTO 9 (ThermoFisher Scientific) at final concentration of 0.02 μg/ml (green fluorescence) and propidium iodide (Sigma) at final concentration of 3 μg/ml (red fluorescence) to differentiate between viable and non-viable bacteria. To differentiate between gram-positive and gram-negative bacterial species ViaGram Red+ (ThermoFisher Scientific) was used. The microscopic images were obtained with a 1-μm Ζ-stacks.
Evaluation of antibacterial activity
The minimum inhibitory concentration (MIC) of antimicrobials was determined by the broth microdilution method in 96-well microtiter plates (Eppendorf) according to the recommendation of the European Committee for Antimicrobial Susceptibility Testing (EUCAST) rules for antimicrobial susceptibility testing98. Briefly, the 108 cells/mL bacterial suspension was subsequently diluted 1:300 with BM broth supplemented with various concentrations of antimicrobials in microwell plates to obtain a 3 × 105 cells/mL suspension. The concentrations of antimicrobials ranged from 0.25 to 512 mg/L. Besides the usual double dilutions, additional concentrations were included in between. The cultures were next incubated at 35 °C for 24 h. The MIC was determined as the lowest concentration of antimicrobials for which no visible bacterial growth could be observed after 24 h incubation.
To determine the MBC of antimicrobials the CFU/mL were further evaluated in the culture liquid from those wells without visible growth. 10 μl of the culture liquid from the wells with no visible growth were inoculated into 3 ml of LB broth followed by cultivation for 24 h. The MBC was determined as the lowest concentration of compound for which no visible bacterial growth could be observed according to the EUCAST of the European Society of Clinical Microbiology and Infectious Diseases (ESCMID)99.
To evaluate the viability of cells with resazurine assay, the resazurine solution was added to cells suspension until the final concentration of 100 μM, or 600 μM resazurine solution was added to the biofilms and incubation was followed at 25 ˚C for 15 min. The pink-stained wells were considered as containing viable cells, blue-stained wells were considered as containing non-viable cells.
Drop plate assay
To evaluate the viability of both detached and planktonic cells, a series of tenfold dilutions of liquid culture from each well were prepared in 3 technical repeats and dropped by 5 μl onto LB agar plates. CFUs were counted from the two last drops typically containing 5–15 colonies and further averaged. To evaluate the viability of the biofilm-embedded cells, the wells were washed twice with 0.9% NaCl in order to remove the non-adherent cells. The biofilms were also suspended in 0.9% NaCl by scratching the well bottoms with subsequent treatment in an ultrasonic bath for 2 min to facilitate the disintegration of bacterial clumps75. Viable cells were counted by the drop plate method as described above.
Atomic force microscopy (AFM)
Atomic force microscopy images of the air-dried microbial biofilms (mono- and dual species) were collected using Dimension Icon scanning probe microscope (Bruker, USA) operating in PeakForce Tapping™ mode. For AFM imaging in air the biofilms were grown in BM-broth on 34-mm plates (TC-treated, Eppendorf, 2 ml per plate) and treated with F105 in concentration 2.5 µg/ml as described above. Then the treated biofilms were washed with water and fixed with glutaraldehyde (0.1% aqueous solution) for 4 h. After subsequent washing with water the plates were dried in air and imaged at ambient conditions. ScanAsyst-Air probes (Bruker) having nominal length 115 µm, tip radius 2 nm, spring constant 0.4 N\m were used throughout. The images were obtained at 512 lines\scan at 0.8–0.9 Hz scan rate. The images were acquired in height (topography), peak force error and adhesion channels. The raw AFM imaging data obtained were processed and analysed using Nanoscope Analysis v.1.7. software (Bruker).
Statistical analysis
Experiments were carried out in six biological repeats with newly prepated cultures and medium in each of them. The fraction of non-viable cells in microscopic images was estimated as the relative fraction of the red cells among all cells in the combined images obtained by overlaying of the green and the red fluorescence microphotographs (10 images per each sample) by using BioFilmAnalyzer software100 with modifications101.The statistical significance of the discrepancy between monoculture and mixed biofilms treatment efficacy was determined using the Kruskal–Wallis statistical test with significance threshold at p < 0.05.
References
Kirker, K. R. & James, G. A. In vitro studies evaluating the effects of biofilms on wound-healing cells: a review. Apmis 125, 344–352. https://doi.org/10.1111/apm.12678 (2017).
Kirker, K. R. et al. Loss of viability and induction of apoptosis in human keratinocytes exposed to Staphylococcus aureus biofilms in vitro. Wound Repair Regener. 17, 690–699. https://doi.org/10.1111/j.1524-475X.2009.00523.x (2009).
Marano, R. J. et al. Secreted biofilm factors adversely affect cellular wound healing responses in vitro. Sci. Rep. 5, 13296. https://doi.org/10.1038/srep13296 (2015).
Secor, P. R. et al. Staphylococcus aureus Biofilm and Planktonic cultures differentially impact gene expression, mapk phosphorylation, and cytokine production in human keratinocytes. BMC Microbiol. https://doi.org/10.1186/1471-2180-11-143 (2011).
den Reijer, P. M. et al. Detection of alpha-toxin and other virulence factors in biofilms of Staphylococcus aureus on polystyrene and a human epidermal model. PLoS ONE https://doi.org/10.1371/journal.pone.0145722 (2016).
Kirker, K. R., James, G. A., Fleckman, P., Olerud, J. E. & Stewart, P. S. Differential effects of planktonic and biofilm MRSA on human fibroblasts. Wound Repair Regener. 20, 253–261. https://doi.org/10.1111/j.1524-475X.2012.00769.x (2012).
Harrison, F. Microbial ecology of the cystic fibrosis lung. Microbiol. SGM 153, 917–923. https://doi.org/10.1099/mic.0.2006/004077-0 (2007).
Fazli, M. et al. Nonrandom distribution of Pseudomonas aeruginosa and Staphylococcus aureus in chronic wounds. J. Clin. Microbiol. 47, 4084–4089. https://doi.org/10.1128/jcm.01395-09 (2009).
Tipton, C. D. et al. Temporal dynamics of relative abundances and bacterial succession in chronic wound communities. Wound Repair Regener. 25, 673–679. https://doi.org/10.1111/wrr.12555 (2017).
10Foundation, C. F. Vol. 2011 annual data report (2012).
Lewis, K. Riddle of biofilm resistance. Antimicrob. Agents Chemother. 45, 999–1007. https://doi.org/10.1128/aac.45.4.999-1007.2001 (2001).
Atshan, S. S. et al. Comparative proteomic analysis of extracellular proteins expressed by various clonal types of Staphylococcus aureus and during planktonic growth and biofilm development. Front. Microbiol. https://doi.org/10.3389/fmicb.2015.00524 (2015).
Cosgrove, S. E., Kaye, K. S., Eliopoulous, G. M. & Carmeli, Y. Health and economic outcomes of the emergence of third-generation cephalosporin resistance in Enterobacter species. Arch. Intern. Med. 162, 185–190. https://doi.org/10.1001/archinte.162.2.185 (2002).
Sanchez-Vizuete, P., Orgaz, B., Aymerich, S., Le Coq, D. & Briandet, R. Pathogens protection against the action of disinfectants in multispecies biofilms. Front. Microbiol. https://doi.org/10.3389/fmicb.2015.00705 (2015).
Dalton, T. et al. An in vivo polymicrobial biofilm wound infection model to study interspecies interactions. PLoS ONE https://doi.org/10.1371/journal.pone.0027317 (2011).
Seth, A. K. et al. Quantitative comparison and analysis of species-specific wound biofilm virulence using an in vivo, rabbit-ear model. J. Am. Coll. Surg. 215, 388–399. https://doi.org/10.1016/j.jamcollsurg.2012.05.028 (2012).
Pastar, I. et al. Interactions of methicillin resistant Staphylococcus aureus USA300 and Pseudomonas aeruginosa in polymicrobial wound infection. PLoS ONE https://doi.org/10.1371/journal.pone.0056846 (2013).
Cendra, M. D., Blanco-Cabra, N., Pedraz, L. & Torrents, E. Optimal environmental and culture conditions allow the in vitro coexistence of Pseudomonas aeruginosa and Staphylococcus aureus in stable biofilms. Sci. Rep. https://doi.org/10.1038/s41598-019-52726-0 (2019).
Orazi, G., Ruoff, K. L. & O’Toole, G. A. Pseudomonas aeruginosa increases the sensitivity of biofilm- grown Staphylococcus aureus to membrane-targeting antiseptics and antibiotics. Mbio https://doi.org/10.1128/mBio.01501-19 (2019).
Wijesinghe, G. et al. Influence of laboratory culture media on in vitro growth, adhesion, and biofilm formation of Pseudomonas aeruginosa and Staphylococcus aureus. Med. Principles Pract. 28, 28–35. https://doi.org/10.1159/000494757 (2019).
Millette, G. et al. Despite antagonism in vitro, Pseudomonas aeruginosa enhances Staphylococcus aureus colonization in a murine lung infection model. Front. Microbiol. https://doi.org/10.3389/fmicb.2019.02880 (2019).
Mashburn, L. M., Jett, A. M., Akins, D. R. & Whiteley, M. Staphylococcus aureus serves as an iron source for Pseudomonas aeruginosa during in vivo coculture. J. Bacteriol. 187, 554–566. https://doi.org/10.1128/jb.187.2.554-566.2005 (2005).
Ruger, M., Ackermann, M. & Reichl, U. Species-specific viability analysis of Pseudomonas aeruginosa, Burkholderia cepacia and Staphylococcus aureus in mixed culture by flow cytometry. BMC Microbiol. https://doi.org/10.1186/1471-2180-14-56 (2014).
Nguyen, A. T., Jones, J. W., Ruge, M. A., Kane, M. A. & Oglesby-Sherrouse, A. G. Iron depletion enhances production of antimicrobials by Pseudomonas aeruginosa. J. Bacteriol. 197, 2265–2275. https://doi.org/10.1128/jb.00072-15 (2015).
Filkins, L. M. et al. Coculture of Staphylococcus aureus with Pseudomonas aeruginosa drives S-aureus towards fermentative metabolism and reduced viability in a cystic fibrosis model. J. Bacteriol. 197, 2252–2264. https://doi.org/10.1128/jb.00059-15 (2015).
Tognon, M., Kohler, T., Luscher, A. & van Delden, C. Transcriptional profiling of Pseudomonas aeruginosa and Staphylococcus aureus during in vitro co-culture. BMC Genom. https://doi.org/10.1186/s12864-018-5398-y (2019).
Mitchell, G. et al. Staphylococcus aureus sigma B-dependent emergence of small-colony variants and biofilm production following exposure to Pseudomonas aeruginosa 4-hydroxy-2-heptylquinoline-N-oxide. BMC Microbiol. https://doi.org/10.1186/1471-2180-10-33 (2010).
Fugere, A. et al. Interspecific small molecule interactions between clinical isolates of Pseudomonas aeruginosa and Staphylococcus aureus from adult cystic fibrosis patients. PLoS ONE https://doi.org/10.1371/journal.pone.0086705 (2014).
Hoffman, L. R. et al. Selection for Staphylococcus aureus small-colony variants due to growth in the presence of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 103, 19890–19895. https://doi.org/10.1073/pnas.0606756104 (2006).
Proctor, R. A. et al. Small colony variants: a pathogenic form of bacteria that facilitates persistent and recurrent infections. Nat. Rev. Microbiol. 4, 295–305. https://doi.org/10.1038/nrmicro1384 (2006).
Biswas, L., Biswas, R., Schlag, M., Bertram, R. & Gotz, F. Small-colony variant selection as a survival strategy for Staphylococcus aureus in the presence of Pseudomonas aeruginosa. Appl. Environ. Microbiol. 75, 6910–6912. https://doi.org/10.1128/aem.01211-09 (2009).
Atalla, H., Gyles, C. & Mallard, B. Staphylococcus aureus small colony variants (SCVs) and their role in disease. Anim. Health Res. Rev. 12, 33–45 (2011).
Orazi, G. & O’Toole, G. A. Pseudomonas aeruginosa alters Staphylococcus aureus sensitivity to vancomycin in a biofilm model of cystic fibrosis infection. Mbio https://doi.org/10.1128/mBio.00873-17 (2017).
Lightbown, J. W. & Jackson, F. L. Inhibition of cytochrome systems of heart muscle and certain bacteria by the antagonists of dihydrostreptomycin: 2-alkyl-4-hydroxyquinoline N-oxides. Biochem. J. 63, 130–137 (1956).
Palmer, K. L., Mashburn, L. M., Singh, P. K. & Whiteley, M. Cystic fibrosis sputum supports growth and cues key aspects of Pseudomonas aeruginosa physiology. J. Bacteriol. 187, 5267–5277. https://doi.org/10.1128/jb.187.15.5267-5277.2005 (2005).
Baldan, R. et al. Adaptation of Pseudomonas aeruginosa in cystic fibrosis airways influences virulence of Staphylococcus aureus in vitro and murine models of co-infection. PLoS ONE https://doi.org/10.1371/journal.pone.0089614 (2014).
DeLeon, S. et al. Synergistic interactions of Pseudomonas aeruginosa and Staphylococcus aureus in an in vitro wound model. Infect. Immun. 82, 4718–4728. https://doi.org/10.1128/iai.02198-14 (2014).
Rendueles, O. & Ghigo, J. M. Multi-species biofilms: how to avoid unfriendly neighbors. FEMS Microbiol. Rev. 36, 972–989. https://doi.org/10.1111/j.1574-6976.2012.00328.x (2012).
Hotterbeekx, A. et al. The endotracheal tube microbiome associated with Pseudomonas aeruginosa or Staphylococcus epidermidis. Sci. Rep. https://doi.org/10.1038/srep36507 (2016).
Co, E. M., Keen, E. F. & Aldous, W. K. Prevalence of methicillin-resistant Staphylococcus aureus in a combat support hospital in Iraq. Mil. Med. 176, 89–93 (2011).
Keen, E. F. et al. Incidence and bacteriology of burn infections at a military burn center. Burns 36, 461–468. https://doi.org/10.1016/j.burns.2009.10.012 (2010).
Gomez, R. et al. Causes of mortality by autopsy findings of combat casualties and civilian patients admitted to a burn unit. J. Am. Coll. Surg. 208, 348–354. https://doi.org/10.1016/j.jamcollsurg.2008.11.012 (2009).
Martin, J. M., Zenilman, J. M. & Lazarus, G. S. Molecular microbiology: new dimensions for cutaneous biology and wound healing. J. Investig. Dermatol. 130, 38–48. https://doi.org/10.1038/jid.2009.221 (2010).
Gjødsbøl, K. et al. Multiple bacterial species reside in chronic wounds: a longitudinal study. Int. Wound J. 3, 225–231 (2006).
Kirketerp-Moller, K. et al. Distribution, organization, and ecology of bacteria in chronic wounds. J. Clin. Microbiol. 46, 2717–2722. https://doi.org/10.1128/jcm.00501-08 (2008).
Duan, K. M., Dammel, C., Stein, J., Rabin, H. & Surette, M. G. Modulation of Pseudomonas aeruginosa gene expression by host microflora through interspecies communication. Mol. Microbiol. 50, 1477–1491. https://doi.org/10.1046/j.1365-2958.2003.03803.x (2003).
Korgaonkar, A., Trivedi, U., Rumbaugh, K. P. & Whiteley, M. Community surveillance enhances Pseudomonas aeruginosa virulence during polymicrobial infection. Proc. Natl. Acad. Sci. USA 110, 1059–1064. https://doi.org/10.1073/pnas.1214550110 (2013).
Sagel, S. D. et al. Impact of pseudomonas and staphylococcus infection on inflammation and clinical status in young children with cystic fibrosis. J. Pediatr. 154, 183–188. https://doi.org/10.1016/j.jpeds.2008.08.001 (2009).
Hauser, A. R., Jain, M., Bar-Meir, M. & McColley, S. A. Clinical significance of microbial infection and adaptation in cystic fibrosis. Clin. Microbiol. Rev. 24, 29–70. https://doi.org/10.1128/cmr.00036-10 (2011).
Foundation, C. F. Patient registry annual data report, vol. 2013 (2014).
Leid, J. G. et al. The Exopolysaccharide alginate protects Pseudomonas aeruginosa biofilm bacteria from IFN-γ-mediated macrophage killing. J. Immunol. 175, 7512–7518 (2005).
Ryder, C., Byrd, M. & Wozniak, D. J. Role of polysaccharides in Pseudomonas aeruginosa biofilm development. Curr. Opin. Microbiol. 10, 644–648. https://doi.org/10.1016/j.mib.2007.09.010 (2007).
Colvin, K. M. et al. The pel polysaccharide can serve a structural and protective role in the biofilm matrix of Pseudomonas aeruginosa. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1001264 (2011).
Colvin, K. M. et al. The Pel and Psl polysaccharides provide Pseudomonas aeruginosa structural redundancy within the biofilm matrix. Environ. Microbiol. 14, 1913–1928. https://doi.org/10.1111/j.1462-2920.2011.02657.x (2012).
Chew, S. C. et al. Dynamic remodeling of microbial biofilms by functionally distinct exopolysaccharides. Mbio. https://doi.org/10.1128/mBio.01536-14 (2014).
Foster, T. J. Immune evasion by Staphylococci. Nat. Rev. Microbiol. 3, 948–958. https://doi.org/10.1038/nrmicro1289 (2005).
Armbruster, C. R. et al. Staphylococcus aureus protein A mediates interspecies interactions at the cell surface of Pseudomonas aeruginosa. Mbio https://doi.org/10.1128/mBio.00538-16 (2016).
Kali, A., Devaraj, B. P., Charles, M. V. & Seetha, K. S. Antibacterial synergy of curcumin with antibiotics against biofilm producing clinical bacterial isolates. J. Basic Clin. Pharm. 7, 93–96 (2016).
Percival, S. L. et al. Antiseptics for treating infected wounds: Efficacy on biofilms and effect of pH. Crit. Rev. Microbiol. 42, 293–309. https://doi.org/10.3109/1040841x.2014.940495 (2016).
Bortolin, M., Bidossi, A., De Vecchi, E., Avveniente, M. & Drago, L. In vitro antimicrobial activity of chlorquinaldol against microorganisms responsible for skin and soft tissue infections: comparative evaluation with gentamicin and fusidic acid. Front. Microbiol. https://doi.org/10.3389/fmicb.2017.01039 (2017).
Baidamshina, D. R. et al. Targeting microbial biofilms using Ficin, a nonspecific plant protease. Sci. Rep. https://doi.org/10.1038/srep46068 (2017).
Blackledge, M. S., Worthington, R. J. & Melander, C. Biologically inspired strategies for combating bacterial biofilms. Curr. Opin. Pharmacol. 13, 699–706. https://doi.org/10.1016/j.coph.2013.07.004 (2013).
Kaplan, J. B. Biofilm dispersal: mechanisms, clinical implications, and potential therapeutic uses. J. Dent. Res. 89, 205–218. https://doi.org/10.1177/0022034509359403 (2010).
Bayer, A. S. et al. Effects of alginase on the natural-history and antibiotic-therapy of experimental endocarditis caused by mucoid Pseudomonas-aeruginosa. Infect. Immun. 60, 3979–3985 (1992).
Park, J. H., Lee, J. H., Cho, M. H., Herzberg, M. & Lee, J. Acceleration of protease effect on Staphylococcus aureus biofilm dispersal. FEMS Microbiol. Lett. 335, 31–38. https://doi.org/10.1111/j.1574-6968.2012.02635.x (2012).
Izano, E. A., Amarante, M. A., Kher, W. B. & Kaplan, J. B. Differential roles of poly-N-acetylglucosamine surface polysaccharide and extracellular DNA in Staphylococcus aureus and Staphylococcus epidermidis biofilms. Appl. Environ. Microbiol. 74, 470–476. https://doi.org/10.1128/aem.02073-07 (2008).
Donelli, G. et al. Synergistic activity of dispersin B and cefamandole nafate in inhibition of staphylococcal biofilm growth on polyurethanes. Antimicrob. Agents Chemother. 51, 2733–2740. https://doi.org/10.1128/aac.01249-06 (2007).
Darouiche, R. O., Mansouri, M. D., Gawande, P. V. & Madhyastha, S. Antimicrobial and antibiofilm efficacy of triclosan and DispersinB (R) combination. J. Antimicrob. Chemother. 64, 88–93. https://doi.org/10.1093/jac/dkp158 (2009).
Selan, L., Berlutti, F., Passariello, C., Comodiballanti, M. R. & Thaller, M. C. Proteolytic-enzymes: a new treatment strategy for prosthetic infections. Antimicrob. Agents Chemother. 37, 2618–2621 (1993).
Alkawash, M. A., Soothill, J. S. & Schiller, N. L. Alginate lyase enhances antibiotic killing of mucoid Pseudomonas aeruginosa in biofilms. Apmis 114, 131–138. https://doi.org/10.1111/j.1600-0463.2006.apm_356.x (2006).
Alipour, M., Suntres, Z. E. & Omri, A. Importance of DNase and alginate lyase for enhancing free and liposome encapsulated aminoglycoside activity against Pseudomonas aeruginosa. J. Antimicrob. Chemother. 64, 317–325. https://doi.org/10.1093/jac/dkp165 (2009).
Nijland, R., Hall, M. J. & Burgess, J. G. Dispersal of biofilms by secreted, matrix degrading, bacterial DNase. PLoS ONE https://doi.org/10.1371/journal.pone.0015668 (2010).
Heck, R., Stuetz, A. Antimycotic-6-phenyl-2-hexen-4-ynamines. (1988).
Lonn-Stensrud, J., Landin, M. A., Benneche, T., Petersen, F. C. & Scheie, A. A. Furanones, potential agents for preventing Staphylococcus epidermidis biofilm infections?. J. Antimicrob. Chemother. 63, 309–316. https://doi.org/10.1093/jac/dkn501 (2009).
Kayumov, A. R. et al. New derivatives of pyridoxine exhibit high antibacterial activity against biofilm-embedded staphylococcus cells. Biomed. Res. Int. https://doi.org/10.1155/2015/890968 (2015).
Trizna, E., Latypova, L., Kurbangalieva, A., Bogachev, M. I. & Kayumov, A. 2(5H)-Furanone derivatives as inhibitors of staphylococcal biofilms. Bionanoscience 6, 423–426. https://doi.org/10.1007/s12668-016-0258-1 (2016).
Sharafutdinov, I. S. et al. Antimicrobial effects of sulfonyl derivative of 2(5H)-furanone against planktonic and biofilm associated methicillin-resistant and -susceptible Staphylococcus aureus. Front. Microbiol. 8, 2246. https://doi.org/10.3389/fmicb.2017.02246 (2017).
Sharafutdinov, I. S. et al. Unraveling the molecular mechanism of selective antimicrobial activity of 2(5H)-furanone derivative against Staphylococcus aureus. Int. J. Mol. Sci. 20, 694. https://doi.org/10.3390/ijms20030694 (2019).
Cramton, S. E., Gerke, C. & Gotz, F. In vitro methods to study staphylococcal biofilm formation. Microbial Growth Biofilms Pt a Dev. Mol. Biol. Aspects 336, 239–255. https://doi.org/10.1016/s0076-6879(01)36593-x (2001).
Voggu, L. et al. Microevolution of cytochrome bd oxidase in staphylococci and its implication in resistance to respiratory toxins released by Pseudomonas. J. Bacteriol. 188, 8079–8086. https://doi.org/10.1128/jb.00858-06 (2006).
Soares, A., Caron, F. & Etienne, M. Commentary: tolerance and resistance of pseudomonas aeruginosa biofilms to antimicrobial agents-how P. aeruginosa can escape antibiotics. Front. Microbiol. 10, 2164. https://doi.org/10.3389/fmicb.2019.02164 (2019).
Ciofu, O. & Tolker-Nielsen, T. Tolerance and resistance of pseudomonas aeruginosa biofilms to antimicrobial agents-how P. aeruginosa can escape antibiotics. Front. Microbiol. 10, 913. https://doi.org/10.3389/fmicb.2019.00913 (2019).
Radlinski, L. et al. Pseudomonas aeruginosa exoproducts determine antibiotic efficacy against Staphylococcus aureus. Plos Biol. 15, e2003981. https://doi.org/10.1371/journal.pbio.2003981 (2017).
Naicker, P. R., Karayem, K., Hoek, K. G. P., Harvey, J. & Wasserman, E. Biofilm formation in invasive Staphylococcus aureus isolates is associated with the clonal lineage. Microb. Pathog. 90, 41–49. https://doi.org/10.1016/j.micpath.2015.10.023 (2016).
Willner, D. et al. Metagenomic analysis of respiratory tract DNA viral communities in cystic fibrosis and non-cystic fibrosis individuals. PLoS ONE 4, e7370. https://doi.org/10.1371/journal.pone.0007370 (2009).
Michelsen, C. F. et al. Evolution of metabolic divergence in Pseudomonas aeruginosa during long-term infection facilitates a proto-cooperative interspecies interaction. ISME J. 10, 1323–1336. https://doi.org/10.1038/ismej.2015.220 (2016).
Abdi-Ali, A., Mohammadi-Mehr, M. & Alaei, Y. A. Bactericidal activity of various antibiotics against biofilm-producing Pseudomonas aeruginosa. Int. J. Antimicrob. Agents 27, 196–200. https://doi.org/10.1016/j.ijantimicag.2005.10.007 (2006).
Mulcahy, H., Charron-Mazenod, L. & Lewenza, S. Extracellular DNA chelates cations and induces antibiotic resistance in pseudomonas aeruginosa biofilms. PLoS Pathog. 4, e1000213. https://doi.org/10.1371/journal.ppat.1000213 (2008).
Shigeta, M. et al. Permeation of antimicrobial agents through Pseudomonas aeruginosa biofilms: a simple method. Chemotherapy 43, 340–345. https://doi.org/10.1159/000239587 (1997).
Jabra-Rizk, A. M. Pathogenesis of polymicrobial biofilms. Open Mycol. J. 5, 39–43 (2011).
Sharafutdinov, I. et al. The antimicrobial activity of 2 (5H)-furanone derivative on Staphylococcus aureus. FEBS J. 283, 200–201 (2016).
Kayumov, A. R. et al. Inhibition of biofilm formation in Bacillus subtilis by new halogenated furanones. J. Antibiot. 68, 297–301. https://doi.org/10.1038/ja.2014.143 (2015).
Diep, B. A. et al. Complete genome sequence of USA300, an epidemic clone of community-acquired meticillin-resistant Staphylococcus aureus. Lancet 367, 731–739. https://doi.org/10.1016/s0140-6736(06)68231-7 (2006).
Bruckner, R. A series of shuttle vectors for Bacillus-subtilis and Escherichia-coli. Gene 122, 187–192. https://doi.org/10.1016/0378-1119(92)90048-t (1992).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343-U341. https://doi.org/10.1038/nmeth.1318 (2009).
Augustin, J. & Gotz, F. Transformation of Staphylococcus-epidermidis and other Staphylococcal species with plasmid DNA by electroporation. FEMS Microbiol. Lett. 66, 203–207 (1990).
Merritt, J. H., Kadouri, D. E. & O’Toole, G. A. Growing and analyzing static biofilms. Curr. Protocols Microbiol. 22, 1 (2005).
Leclercq, R. et al. EUCAST expert rules in antimicrobial susceptibility testing. Clin. Microbiol. Infection 19, 141–160. https://doi.org/10.1111/j.1469-0691.2011.03703.x (2013).
European committee for antimicrobial susceptibility testing (EUCAST) of the European society of clinical microbiology and infectious diseases (ESCMID). Terminology relating to methods for the determination of susceptibility of bacteria to antimicrobial agents. Clin. Microbiol. Inf. 6, 503–508, https://doi.org/10.1046/j.1469-0691.2000.00149.x (2000).
Bogachev, M. I. et al. Fast and simple tool for the quantification of biofilm-embedded cells sub-populations from fluorescent microscopic images. PLoS ONE 13, e0193267. https://doi.org/10.1371/journal.pone.0193267 (2018).
Acknowledgements
This work was supported by the Russian Science Foundation (Projects No. 15-14-00046 and 20-64-47014) and performed in the framework of the Russian Government Program of Competitive Development of Kazan Federal University. Atomic force microscopy experiments were funded by Russian Federation presidential grant MD-2153.2020.3. (recipient: Rawil Fakhrullin)
Author information
Authors and Affiliations
Contributions
E.T., M.Y., D.B., F.A., A.M. and E.R. performed the experiments. A.Ka. and R.F. conducted the experiments. A.Kh. and A.Ku. synthesized F105. E.T., M.Y., A.Ka. and M.B. analyzed the results, prepared figures and graphs and wrote the manuscript. All the authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Trizna, E.Y., Yarullina, M.N., Baidamshina, D.R. et al. Bidirectional alterations in antibiotics susceptibility in Staphylococcus aureus—Pseudomonas aeruginosa dual-species biofilm. Sci Rep 10, 14849 (2020). https://doi.org/10.1038/s41598-020-71834-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-71834-w
- Springer Nature Limited
This article is cited by
-
Staphopain mediated virulence and antibiotic resistance alteration in co-infection of Staphylococcus aureus and Pseudomonas aeruginosa: an animal model
BMC Biotechnology (2024)
-
Strain-specific interspecies interactions between co-isolated pairs of Staphylococcus aureus and Pseudomonas aeruginosa from patients with tracheobronchitis or bronchial colonization
Scientific Reports (2022)
-
Polymicrobial biofilms of ocular bacteria and fungi on ex vivo human corneas
Scientific Reports (2022)
-
In vitro virulence activity of Pseudomonas aeruginosa, enhanced by either Acinetobacter baumannii or Enterococcus faecium through the polymicrobial interactions
Archives of Microbiology (2022)
-
Novel hole-pillar spacer design for improved hydrodynamics and biofouling mitigation in membrane filtration
Scientific Reports (2021)