Abstract
Although numerous pathogenic mutations have been identified in various subunits of N-methyl-D-aspartate receptors (NMDARs), ionotropic glutamate receptors that are central to glutamatergic neurotransmission, the functional effects of these mutations are often unknown. Here, we combined in silico modelling with microscopy, biochemistry, and electrophysiology in cultured HEK293 cells and hippocampal neurons to examine how the pathogenic missense mutation S688Y in the GluN1 NMDAR subunit affects receptor function and trafficking. We found that the S688Y mutation significantly increases the EC50 of both glycine and d-serine in GluN1/GluN2A and GluN1/GluN2B receptors, and significantly slows desensitisation of GluN1/GluN3A receptors. Moreover, the S688Y mutation reduces the surface expression of GluN3A-containing NMDARs in cultured hippocampal neurons, but does not affect the trafficking of GluN2-containing receptors. Finally, we found that the S688Y mutation reduces Ca2+ influx through NMDARs and reduces NMDA-induced excitotoxicity in cultured hippocampal neurons. These findings provide key insights into the molecular mechanisms that underlie the regulation of NMDAR subtypes containing pathogenic mutations.
Similar content being viewed by others
Introduction
N-methyl-D-aspartate receptors (NMDARs) are a subclass of ionotropic glutamate receptors that play an essential role in mediating excitatory neurotransmission and synaptic plasticity in the mammalian central nervous system (CNS)1,2,3. NMDARs are tetramers comprised of two GluN1 subunits (with eight splice variants) together with two GluN2 (GluN2A through GluN2D) and/or two GluN3 (GluN3A and GluN3B) subunits3,4. GluN2A and GluN2B are the principal GluN2 subunits expressed in the forebrain of mature animals3,4,5. NMDARs can also occur as triheteromeric GluN1/GluN2A/GluN2B6,7,8,9 or GluN1/GluN2/GluN3A10,11,12 receptors as well as diheteromeric GluN1/GluN3A receptors13,14,15,16. All GluN subunits share a basic membrane topological structure consisting of four membrane domains (M1 through M4), an intracellular C-terminal domain (CTD), an extracellular N-terminal domain composed of the amino-terminal domain and the S1 segment of the ligand-binding domain (LBD), and an extracellular loop between M3 and M4 containing the S2 segment of the LBD. GluN1/GluN2 receptors are gated by the simultaneous binding of an agonist (glutamate or NMDA) to the LBD in the GluN2 subunit and a co-agonist (glycine or d-serine) to the LBD in the GluN1 subunit3,17,18,19. In contrast, GluN1/GluN3 receptors are gated by glycine binding to the LBD in the GluN3 subunit, with receptor desensitisation mediated by glycine binding to the LBD in the GluN1 subunit20,21,22,23. Thus, the LBD in the GluN1 subunit plays distinct functional roles in GluN1/GluN2 and GluN1/GluN3 receptors.
The number of NMDARs at the cell surface is regulated at multiple levels, including transcription/translation, processing in the endoplasmic reticulum (ER), trafficking to the cell surface, lateral diffusion through the membrane, and receptor internalisation, recycling, and degradation3,4,24,25. Most studies suggest that the intracellular CTD of GluN subunits plays a critical role in regulating the surface expression of NMDARs, including receptors that contain GluN2A and/or GluN2B subunits5. Interestingly, the extracellular part of mammalian GluN subunits is extremely large and contains the LBDs, which also play a critical role in regulating receptor trafficking26. For example, studies have shown that the structural integrity of the LBD in the GluN1 subunit of both GluN1/GluN2 and GluN1/GluN3 receptors27,28, as well as of the LBD in the GluN2B subunit of GluN1/GluN2B receptors29, are likely involved in the ER quality control of NMDARs. Furthermore, several studies found that specific agonists and co-agonists differentially regulate surface NMDARs; for example, d-serine regulates the mobility of GluN2A-containing NMDARs30, whereas glycine drives receptor internalisation31. However, precisely how structural changes in the LBD of GluN subunits regulate the surface delivery and function of specific NMDAR subtypes remains poorly understood.
Numerous pathogenic mutations have been identified in all GluN subunits‒encoding genes in patients with a variety of neuropsychiatric disorders and conditions32,33,34,35,36,37,38,39,40,41,42. The pathogenic mutations in GluN1 subunit have phenotypic similar with pathogenic mutations found in other GluN subunits, although they are usually associated with more sever course43. For example, the pathogenic mutations in GluN1 subunit were reported in patients with intellectual disability, cognitive dysfunction and development delay43,44,45,46,47,48, polymicrogyria47, oculomotor and movement disorders43,45,46, speech difficulties43,45, epilepsy46,47,48, postnatal microcephaly47, generalized cerebral atrophy44 or cortical blindness47. In addition, the pathogenic S688Y mutation in GluN1 subunit was associated with severe early infantile encephalopathy and some of the above symptoms43. Here, we used in silico modelling as well as electrophysiology, microscopy, and biochemistry in HEK293 cells and primary rat hippocampal neurons to examine the effect of the pathogenic S688Y mutation in the LBD of the GluN1 subunit on the trafficking and function of NMDARs. We found that the GluN1-S688Y subunit alters the functional properties of GluN1/GluN2A, GluN1/GluN2B, and GluN1/GluN3A receptors when expressed in both HEK293 cells and hippocampal neurons. Importantly, we also found that the S688Y mutation reduces surface delivery of GluN3A-containing NMDARs in both HEK293 cells and hippocampal neurons. Finally, we found that the S688Y mutation reduces both NMDA-induced Ca2+ influx and excitotoxicity in hippocampal neurons. Taken together, these findings reveal new insights in the role that this pathogenic mutation plays in regulating various NMDAR properties.
Results
The S688Y mutation in the GluN1 subunit causes a steric change in co-agonist binding
Here, we examined the effects of the previously reported S688Y mutation in the LBD of the GluN1 subunit43, focusing our study on the function and trafficking of NMDARs in cultured HEK293 cells and primary hippocampal neurons. As a first step, we performed molecular modelling using an in silico model of the human GluN1/GluN2A receptor49 to compare the structural properties of glycine and d-serine binding at the LBD between wild-type GluN1 and the GluN1-S688Y mutant subunit (Fig. 1). We first docked glycine at the LBD of wild-type GluN1, yielding an excellent root-mean-square deviation (RMSD) score of 0.332 Å, thus validating our approach. Consistently with crystallographic data49, glycine (shown in green in Fig. 1) binds via several hydrogen bonds and electrostatic interactions (Fig. 1a), including interactions between glycine’s carboxyl group and the guanidinium moiety in R52350, the backbone amide groups in T518 and S688, and the hydroxyl group in S688. In addition, the glycine molecule’s positively charged ammonium group forms bonds with the carboxylate in D732, the hydroxyl groups in T518 and S688, and one water molecule. In contrast with the reported crystal structure of glycine bound to the LBD in wild-type GluN1 subunit49, our model suggests that the carboxyl group in P516 is 4.7 Å from the glycine molecule; thus, a hydrogen bond between glycine and this residue is unlikely. Moreover, both W731 and Q405 are located relatively close to the glycine molecule and are presumably involved in other electrostatic interactions and/or water-mediated bridges, although this is difficult to estimate using docking studies. With respect to the interaction between d-serine (shown in yellow in Fig. 1b,d) and the LBD in wild-type GluN1, our model suggests that the d-serine molecule likely forms hydrogen bonds between its functional carboxyl group and the guanidinium moiety in the R523 residue and the backbone amides in S688 and T518 (Fig. 1b). In addition, the d-serine molecule’s hydroxyl group is located near the backbone amides in V689 and S688 and the hydroxyl group in S688, and the d-serine molecule’s positively charged ammonium moiety forms contacts with the carboxylate group in D732, the carbonyl oxygen in P516, and the hydroxyl group in T518. Finally, our docking model did not reveal any direct interaction between d-serine and a water molecule.
The S688Y mutation in the GluN1 subunit alters the binding of co-agonists in the receptor’s ligand-binding domain. (a–d) Glycine (in green; a,c) and d-serine (in yellow; b,d) binding to the LBD of wild-type GluN1 (a,b) and GluN1-S688Y (c,d). The critical amino acid residues involved in co-agonist binding are shown as dark blue carbon atoms; hydrogen bonds are depicted as dashed lines, and the rest of the receptor is shown in light grey. (e,f) Superimposed structures of the LBD in wild-type (S688) GluN1 (shown as dark blue carbon atoms) and mutant (Y688) GluN1 (shown as yellow carbon atoms), with the glycine (e) and d-serine (f) molecules shown in green (WT) and orange (Y688). The PyMOL Molecular Graphics System, Version 2.0.6, Schrödinger, LLC (https://pymol.org/2/) was used to make the figure.
With respect to the GluN1-S688Y subunit, we found that glycine binding is generally similar to the wild-type GluN1 subunit (Fig. 1c); the only apparent difference was within the vicinity of the Y688 residue, in which the S688Y mutation shifted the glycine molecule toward the vicinity of the carbonyl oxygen in P516 residue and away from the V689 residue. Interestingly, the hydrogen bond formed between the glycine molecule’s carboxyl group and the backbone amide in residue 688 residue was unaffected by the mutation. Importantly, hydroxyl group of Y688 is too far away from glycine to form the hydrogen bond; the interaction between positively charged ammonium moiety in glycine and the aromatic region of Y688 residue is the only that can be observed. In contrast with glycine, we found considerable differences in d-serine binding between the GluN1-S688Y and wild-type GluN1 subunits (Fig. 1d). Specifically, we found that the S688Y mutation appears to position the d-serine molecule far from the V689 residue; thus, no hydrogen bond is formed between the ligand’s hydroxyl group and the backbone amide in V689. Interestingly, we also found that a water molecule plays a role in anchoring the d-serine molecule’s hydroxyl group to the GluN1-S688Y subunit. The most striking finding is that the interaction between the d-serine molecule and the GluN1-S688Y subunit lacks the hydrogen bond between the backbone amide in V689 residue and the ligand’s hydroxyl moiety, as well as the hydroxyl-hydroxyl interaction between d-serine and residue 688. To help visualise the structural changes induced by the S688Y mutation, we superimposed the glycine-bound (Fig. 1e) and d-serine‒bound (Fig. 1f) structures of the wild-type and mutant GluN1 LBDs. Glycine-bound and d-serine-bound complexes generated a high degree of similarity with RMSD value of 0.140 Å.
In summary, our in silico modelling reveals that the presence of the more sterically demanding tyrosine at position 688 in the mutant GluN1 subunit decreases the apparent affinity of both glycine and d-serine for binding the LBD.
The GluN1-S688Y mutation alters the functional properties of human GluN1/GluN2A, GluN1/GluN2B, and GluN1/GluN3A receptors
To test our hypothesis that the S688Y mutation in GluN1 decreases the receptor’s affinity for glycine and d-serine, we performed whole-cell patch-clamp recordings in HEK293 cells expressing human (h) GluN1/GluN2A, GluN1/GluN2B, and GluN1/GluN3A receptors. Specifically, we co-expressed either wild-type or hGluN1-4a-S688Y together with hGluN2A (Fig. 2a,c), hGluN2B (Fig. 2e,g), or hGluN3A subunits (Fig. 2i).
The S688Y mutation in GluN1 alters the receptor’s ligand affinity and desensitisation properties. (a,c,e,g) Representative whole-cell patch-clamp recordings of HEK293 cells co-transfected with hGluN1-4a and hGluN2 (left) or hGluN1-4a-S688Y and hGluN2 (right). Currents were elicited by applying glutamate (100 µM or 10 mM) and either glycine or d-serine at the indicated concentrations; where indicated, the NMDAR antagonist 7-chlorokynurenic acid (KYNA, 10 µM) was applied. (b,d,f,h) Normalised steady-state concentration–response curves for cells expressing the indicated NMDAR subunits. Each data point represents the mean normalised steady-state current (± SEM). The EC50 value, Hill coefficient, and number of cells recorded for each group are listed in Table 1. (i) Representative whole-cell patch-clamp recordings of HEK293 cells co-expressing hGluN1-4a and hGluN3A (top) or hGluN1-4a-S688Y and hGluN3A (bottom). Currents were elicited by applying glycine at the indicated concentrations (µM). (j) Normalised peak concentration–response curves for cells expressing the indicated NMDAR subunits. Each data point represents the mean normalised peak current (± SEM). The EC50 value, Hill coefficient, and number of cells recorded for each group are listed in Table 2. (k) Summary of the τw of desensitisation measured in response to glycine in cells expressing the indicated NMDAR subunits.
First, we generated concentration–response curves for glycine (Fig. 2b,f) and d-serine (Fig. 2d,h) in the presence of 100 µM glutamate, as described previously51. The results are summarised in Table 1 and are consistent with previously reported EC50 values for glycine d-serine51. To generate concentration–response curves for receptors containing the GluN1-S688Y subunit, we measured the responses in the presence of 10 mM glutamate, because the GluN1-S688Y mutation might affect the cooperativity among the LBDs within the functional NMDAR heterotetramer (see Supplementary Fig. S1). As shown in Fig. 2 and summarised in Table 1, the S688Y mutation caused a significantly reduced potency for both glycine and d-serine, reflected by significantly increased EC50 values. Taken together, these findings support our modelling data, confirming that the S688Y mutation reduces the receptor’s affinity for both glycine and d-serine.
Unlike GluN2-containing receptors, GluN3-containing receptors are activated by glycine binding to the LBD in the GluN3A subunit and desensitised by glycine binding to the LBD in GluN120,21,22,23. We therefore measured currents induced by glycine at concentrations ranging from 30 µM to 10 mM in HEK293 cells expressing either hGluN1-4a/hGluN3A or hGluN1-4a-S688Y/hGluN3A receptors (Fig. 2i) and then analysed the peak concentration–response curve and the time constant for desensitisation, as described previously27. We found that cells expressing hGluN1-4a-S688Y/hGluN3A receptors were significantly less responsive to glycine compared to cells expressing wild-type receptors (Fig. 2j), with an ~ twofold decrease in glycine potency (Table 2). In addition, we found that hGluN1-4a-S688Y/hGluN3A receptors had an increased time constant (τw) of desensitisation compared to wild-type receptors (Fig. 2k). Thus, although GluN1-S688Y subunits are capable of forming functional GluN1/GluN2 and GluN1/GluN3A receptors, its presence significantly alters the receptor’s functional properties.
The S688Y mutation in GluN1 differentially regulates the surface delivery of GluN1/GluN2 and GluN1/GluN3 NMDARs in HEK293 cells
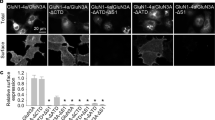
The D732A mutation in the glycine-binding site of GluN1 has been reported to reduce trafficking of GluN1/GluN2A receptors to the cell surface28; similarly, the integrity of the glutamate-binding site in GluN2B has been shown to regulate the trafficking of GluN1/GluN2B receptors29. We recently reported that the surface delivery of GluN1/GluN3A receptors is regulated by structural features in the glycine-binding sites of both GluN1 and GluN3A27. Here, we examined the effect of the S688Y mutation on the surface delivery of NMDARs expressed in HEK293 cells. To monitor expression, we co-transfected cells with either wild-type or hGluN1-4a-S688Y together with GFP-tagged rat GluN2A (GFP-rGluN2A), GFP-tagged rat GluN2B (GFP-rGluN2B), or GFP-tagged human GluN3A (GFP-hGluN3A; Fig. 3a–c). We then measured relative surface expression of the various NMDARs using fluorescence confocal microscopy. We found that the S688Y mutation did not affect the surface delivery of receptors containing either GFP-rGluN2A or GFP-rGluN2B subunits compared to the corresponding wild-type GluN1 subunits; in contrast, the mutation significantly reduced the surface delivery of receptors containing GFP-hGluN3A subunit (Fig. 3a–c). Similar results were obtained when we expressed the YFP-tagged hGluN1-1a-S688Y subunits together with hGluN2A, hGluN2B, or hGluN3A subunits compared to wild-type YFP-hGluN1-1a (Fig. 3d–f). Together, these data indicate that the S688Y mutation in GluN1 differentially regulates the surface expression of NMDARs in a subunit-dependent manner.
The S688Y mutation in GluN1 differentially regulates the surface expression of NMDAR subtypes in HEK293 cells. (a–c) Representative images of total and surface GFP-rGluN2A (a), GFP-rGluN2B (b), and GFP-hGluN3A (c) subunits measured in HEK293 cells lacking or expressing the indicated hGluN1-4a subunits. Shown below is the relative surface expression of GFP-rGluN2A, GFP-rGluN2B, and GFP-hGluN3A measured using fluorescence microscopy (n ≥ 192 cells per group); *p < 0.05 (one-way ANOVA followed by Dunnett’s post hoc test). (d–f) Summary of the relative surface expression of YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y subunits expressed alone or together with hGluN2A, hGluN2B, and hGluN3A measured using fluorescence microscopy (n ≥ 125 cells per group); *p < 0.05 (one-way ANOVA followed by Dunnett’s post hoc test).
The S688Y mutation in GluN1 reduces the surface delivery of GluN3A-containing subunits in hippocampal neurons
Next, we examined whether the S688Y mutation in GluN1 also affects the surface delivery of GluN3-containing NMDARs in hippocampal neurons. Because the endogenous GluN1 subunit is robustly expressed in neurons52, we used shRNA to knock down endogenous GluN1 while expressing shRNA-resistant YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y subunits, similarly as we employed previously53. We found that neurons expressing either YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y had similar levels of both total and surface hGluN1 subunits, as well as similar levels of endogenous GluN2A and GluN2B subunits (Fig. 4a,b). In contrast, and consistent with our findings with HEK393 cells, we found that neurons expressing the YFP-hGluN1-1a-S688Y subunit had reduced levels of surface GluN3A compared to neurons expressing YFP-hGluN1-1a (Fig. 4a,b). Similar results were obtained when we examined GluN subunits in the postsynaptic density (PSD) fraction isolated from cortical neurons expressing YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y (Fig. 4c,d). As additional confirmation, we used confocal microscopy to measure YFP-hGluN1-1a and YFP-hGluN1-1a-S688Y subunits in hippocampal neurons, finding similar surface expression (Fig. 4e,f). We also used electrophysiology to measure glycine-induced currents (in the presence of 1 mM NMDA) in hippocampal neurons expressing either YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y (Fig. 4g). Consistent with our previous findings, we found that neurons expressing the YFP-hGluN1-1a-S688Y subunit have a significantly shifted concentration–response curve for glycine, with an EC50 of 220 µM compared to 0.2 µM for neurons expressing the YFP-hGluN1-1a subunit (Fig. 4h). Taken together, these results indicate that in hippocampal neurons, the S688Y mutation in GluN1 reduces the surface delivery of GluN3A-containing NMDARs, but not GluN2A- or GluN2B-containing receptors, and alters the receptor’s glycine affinity.
The S688Y mutation in GluN1 alters the surface expression of GluN3A-containing NMDARs and alters the ligand affinity of NMDARs in hippocampal neurons. (a) Cell-surface biotinylation assay of dense cultures of hippocampal neurons co-expressing the GluN1 shRNA together with YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y. Surface NMDARs were biotinylated and then pulled down using streptavidin-agarose beads. Total input (2.5% of the lysate) and surface NMDAR subunits were then detected using the indicated antibodies by western blot analysis; the GFP antibody was used to detect the YFP-labelled GluN1 subunit. In the GluN1 blot, the arrow indicates the recombinant hGluN1-1a and hGluN1-1a-S688Y subunits, and the arrowhead indicates the endogenous GluN1 subunit. All biochemical results shown in (a) were cropped from the same blots and exposures for each antibody labelling (GFP, GluN1, GluN2A, GluN2B, GluN3A and α-tubulin; see Supplementary Fig. S2). (b) Summary of the relative surface expression of the indicated NMDAR subunits in neurons expressing YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y. For each subunit, the ratio of surface to total band intensity was measured and normalised to the corresponding hGluN1-1a‒expressing group (n = 6 each); *p < 0.05 (Student’s t-test). (c) Western blot analysis of NMDAR subunits in the postsynaptic density (PSD) isolated from cortical neurons expressing the GluN1 shRNA together with YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y. The positive and negative signals for PSD-95 and synaptobrevin, respectively, confirm that the samples contain the PSD fraction (see Supplementary Fig. S2). (d) Summary of the relative expression of the indicated NMDAR subunits in the PSD fraction, normalised to the corresponding hGluN1-1a‒expressing group (n = 5 each); *p < 0.05 (Student’s t-test). (e) Representative images of hippocampal neurons infected with YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y, showing total and surface GFP immunostaining. (f) Summary of the relative surface expression of YFP-hGluN1-1a and YFP-hGluN1-1a-S688Y in hippocampal neurons measured from a 10-µm2 region of interest in secondary and tertiary dendrites (n ≥ 55 segments from ≥ 11 cells); *p < 0.05 (Student’s t-test). (g) Representative whole-cell patch-clamp recordings of primary hippocampal neurons expressing YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y. Currents were elicited by applying glycine at the indicated concentrations in the presence of 1000 µM NMDA; where indicated, 10 µM KYNA was applied. (h) Steady-state concentration–response curves for hippocampal neurons expressing YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y; each data point represents the mean normalised steady-state current (± SEM). The EC50 value, Hill coefficient, and number of cells recorded for each group are listed in Table 3.
The S688Y mutation reduces NMDA-induced Ca2+ influx and excitotoxicity in hippocampal neurons
Excessive influx of Ca2+ through NMDARs can induce neuronal cell death54. To examine whether the S688Y mutation in GluN1 alters the cell’s susceptibility to excitotoxicity, we measured intracellular Ca2+ in neurons expressing either YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y subunits. Neurons were loaded with the Ca2+ indicator Fura-2 and then stimulated with either 10 or 1000 µM glycine together with 100 µM NMDA (Fig. 5a). We found that both 10 and 1000 µM glycine induced a robust increase in intracellular Ca2+ in cells expressing YFP-hGluN1-1a (Fig. 5a,b); in contrast, 1000 µM glycine induced a similar Ca2+ transient in YFP-hGluN1-1a-S688Y, whereas 10 µM glycine had no measurable effect on intracellular Ca2+ (Fig. 5a,c). These results support the notion that receptors containing GluN1-S688Y subunits have reduced NMDA-induced Ca2+ currents when expressed in hippocampal neurons.
The S688Y mutation in GluN1 reduces NMDA-induced Ca2+transients in hippocampal neurons. (a) The intracellular Ca2+ was measured using the Ca2+ indicator Fura-2 in hippocampal neurons expressing YFP-hGluN1-1a (upper row) or YFP-GluN1-1a-S688Y (bottom row). The representative F340/380 ratiometric images show ratio at resting time (t1) and after application of 100 μM NMDA together with 10 µM (t2) or 1000 μM (t3) glycine as indicated. The cells appear red in accordance with raised Ca2+ levels. Below are shown averaged traces (± SEM) of the F340/F380 ratio measured from n ≥ 45 neurons per group for YFP-hGluN1-1a (b) and YFP-GluN1-1a-S688Y (c). Horizontal bars indicate applications (30 s duration) and the times t1-t3 correspond with the ratiometric images in (a).
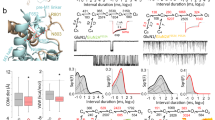
Finally, we examined whether the reduced Ca2+ influx in GluN1-S688Y‒containing receptors has physiologically relevant consequences by expressing either YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y in hippocampal neurons and treating the cells for 1 h with 100 µM NMDA together with glycine (10 or 100 µM) or d-serine (10 or 100 µM). Excitotoxicity was measured 23 h after treatment by staining cells with the nuclear marker Hoechst 33,342 and then performing complex image analysis (Fig. 6a,b; see "Methods"). We found that treating cells with control solution, 10 µM glycine alone, or 10 µM d-serine alone induced only a small degree of cell death in neurons expressing either YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y subunits (Fig. 6c,d). In contrast, treating YFP-hGluN1-1a‒expressing neurons with 10 µM glycine or 10 µM d-serine in the presence of 100 µM NMDA caused nearly 100% excitotoxicity, consistent with previous reports55; similar results were obtained with YFP-hGluN1-1a‒expressing neurons treated with 100 µM glycine or 100 µM d-serine in the presence of 100 µM NMDA (Fig. 6c,d). Importantly, however, YFP-hGluN1-1a-S688Y‒expressing neurons were less sensitive to NMDA-induced excitotoxicity, with significantly reduced cell death when treated with either 10 µM glycine or 10 µM d-serine in the presence of 100 µM NMDA (Fig. 6c,d).
The S688Y GluN1 reduces NMDA-induced excitotoxicity in hippocampal neurons. (a) Hippocampal neurons expressing YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y were treated with control solution or 100 µM NMDA together with 10 µM glycine 1 h; 23 h later, YFP-positive cells were analysed for excitotoxicity by staining with Hoechst 33,342. Shown are representative images. (b) The distribution of nuclear area reveals two distinct Gaussian peaks corresponding to pyknotic and non-pyknotic cells. The histogram was fitted with a 2-Gaussian distribution model (solid line), and the estimated individual Gaussian distributions are plotted as a dashed line (pyknotic cells) and a dotted line (non-pyknotic cells). (c) Distributions of nuclei areas measured for cells expressing YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y and treated as indicated. All nuclei were classified as either pyknotic (dark grey bars) or non-pyknotic (light grey bars) group. (d) Summary of neuronal cell death (calculated as described in the "Methods") measured in cells expressing YFP-hGluN1-1a or YFP-hGluN1-1a-S688Y and treated as indicated (n ≥ 3269 cells per condition from 4 independent experiments); *p < 0.05 and **p < 0.01 (one-way ANOVA supported by the Shapiro–Wilk test of normality at p < 0.05).
Discussion
Here, we focused our study on NMDARs in the mammalian CNS, as these receptors play an important role in a wide range of physiological processes such as learning and memory56,57, as well as neuropathological processes such as neurodegeneration54. Specifically, we characterised the functional effects of the pathogenic S688Y mutation in the ligand-binding domain of the GluN1 NMDAR subunit43 using in silico modelling and microscopic, biochemical, and electrophysiological analyses in HEK293 cells and rat hippocampal neurons.
Our experimentally measured EC50 values for both glycine and d-serine with respect to wild-type GluN1/GluN2 receptors expressed in HEK293 cells are similar to previously published values for receptors expressed in Xenopus oocytes3,58,59. Moreover, we found that expressing the GluN1-S688Y subunit significantly decreased the receptor’s affinity for both glycine and d-serine, with a more profound effect on glycine affinity. These findings are consistent with previous data showing that the S688A mutation produced a four-fold reduction in glycine potency60, the notion that the S688 residue in GluN1 plays a key role in ligand recognition61, as well as with our in silico modelling, which showed that the predicted interaction between the LBD in GluN1-S688Y differs slightly for glycine compared to d-serine. In addition, our finding that the S688Y mutation in GluN1 significantly increased the τw of desensitisation in GluN3A-containing receptors is consistent with previous reports showing that structural changes in the GluN1 LBD alter the desensitisation properties of GluN1/GluN3A receptors20,21,22,23,27. Importantly, our results obtained with hippocampal neurons ‒ which express endogenous GluN2 and GluN3 subunits ‒ support our findings in HEK293 cells; interestingly, however, we found that the change in glycine affinity induced by the S688Y mutation was less profound in neurons compared to HEK293 cells. This discrepancy have several possible explanations, including: (i) the presence of other NMDAR subtypes in hippocampal neurons compared to the subtypes we expressed in HEK293 cells, including triheteromeric9,62 and/or GluN3A-containing NMDARs10,63; (ii) possible differences in posttranslational modifications such as glycosylation64; (iii) possible differences in other proteins that interact with NMDARs65, and (iv) the use of different concentrations of glutamate (HEK293 cells) and NMDA (hippocampal neurons).
Our finding that the S688Y mutation in GluN1 subunit profoundly reduces the surface delivery of GluN3A-containing NMDARs is consistent with our recent report that the LBD’s sensitivity for glycine is the key factor that regulates the surface delivery of these types of NMDARs27. On the other hand, our finding that the S688Y mutation does not appear to affect the surface delivery of GluN1/GluN2A or GluN1/GluN2B receptors differs from a previous report that the D732A mutation in GluN1 affects the trafficking of GluN1/GluN2A receptors28; however, this difference may be explained by differences between the D732A and S688Y mutations with respect to changing the conformation of functional NMDAR heterotetramers, thereby affecting the surface delivery of the resulting GluN1/GluN2A receptors. It is interesting to note that the EC50 for l-glutamate is also correlated with reduced surface delivery of GluN1/GluN2B receptors29,66. Similarly, the NMDARs carrying several pathogenic mutations in the LBDs of GluN2A and GluN2B subunits exhibit clear correlation between the EC50 for l-glutamate and surface expression40. For example, the pathogenic E413G mutation in GluN2B subunit profoundly reduced the surface delivery of NMDARs40, likely by promoting the unbinding of l-glutamate and opening of the LBD67. On the other hand, other pathogenic mutations within the LBDs of GluN2A and GluN2B subunits revealed no clear correlation between the receptor’s EC50 for l-glutamate and surface expression, suggesting that additional mechanisms than the potency of l-glutamate regulate the surface delivery of the GluN1/GluN2 receptors40.
Our finding that the S688Y mutation in GluN1 reduces NMDA-induced excitotoxicity supports the notion that a pathogenic missense mutation in a GluN subunit can profoundly affect neuronal survival. Nevertheless, an open question is whether this in vitro effect is relevant in vivo, particularly given that specific de novo pathogenic mutations in patients are heterozygous. Moreover, whether the reduction in surface delivery of GluN3A-containing NMDARs is functionally relevant is currently unknown; however, it is interesting to speculate that a reduction in the surface expression of GluN3A-containing NMDARs may alter dendritic spine maturation68,69 and may change the neuron’s vulnerability to ischaemic events, given that the GluN3A subunit has been found to reduce neuronal apoptosis70,71.
In summary, our results indicate that the functional effects of putative pathogenic mutations in GluN subunits should be examined carefully and systematically using a variety of techniques, particularly given that endogenous NMDARs are comprised of a wide variety of GluN subunit combinations. Nevertheless, our findings provide new insights into the role that the ligand-binding domain in the GluN1 subunit plays in NMDAR trafficking and functioning.
Methods
In silico modelling
The structure of the GluN1/GluN2A LBD was obtained from the RCSB Protein Data Bank – PDB ID: 5KCJ49 (structure of the human GluN1/GluN2A LBD in complex with GNE6901, resolution 2.09 Å, no outliers according to Ramachandran et al.72). The numbering of the amino acid residues provided in the original GluN1/GluN2A receptor downloaded from the Protein Data Bank was revised based on the human full-length GluN1 subunit sequence (PubMed gene ID: 2902, NCBI Reference Sequence (RefSeq): NM_001185091.2) as follows: Q14 = Q405, P125 = P516, T127 = T518, R132 = R523, S181 = S688, V182 = V689, W224 = W731, and D225 = D732. The GluN1/GluN2A LBD structure was prepared using the DockPrep function of UCSF Chimera (v. 1.4). The position of the Y688 residue in the GluN1-S688Y/GluN2A LBD was obtained from the Dunbrack rotamer library as the most likely rotamer. Two structural water molecules were removed from the mutated receptor due to steric reasons. The energy of the ligand/GluN1/GluN2A LBD was minimised using UCSF Chimera (v. 1.4) with 1000 iterations. The wild-type and mutant GluN1/GluN2A LBDs were converted to pdbqt files using AutodockTools (v. 1.5.6)73, and the S688 and Y688 residues were set as flexible for docking, while the other amino acid residues were rigid. Three-dimensional structures of ligands/co-agonists were built using Open Babel (v. 2.3.1), minimised using Avogadro (v 1.1.0), and converted to pdbqt file format using AutodockTools73. The docking calculations were made using Autodock Vina (v. 1.1.2) with an exhaustiveness value of 874. The visualisation of the receptor-ligand interactions was prepared using the PyMOL Molecular Graphics System, v. 2.0 (Schrödinger LLC, Mannheim, Germany).
Mammalian expression vectors and lentiviruses
For this study, we used cDNA vectors expressing untagged human GluN1-4a (hGluN1-4a; NCBI RefSeq NM_001270610.1 was modified to the human version by changing the four amino acid residues (N159S, R212K, I267L, M415L) that differ between the rat and human GluN1-4a subunits)27, GluN2A (hGluN2A; NCBI Reference Sequence: NM_000833.5), GluN2B (hGluN2B; NCBI RefSeq: NM_000834.5), and GluN3A (hGluN3A; NCBI RefSeq: NM_133445.3) subunits27,75,76, GFP-tagged versions of rat GluN2A (GFP-rGluN2A; NCBI RefSeq: NM_012573.3) and GluN2B (GFP-rGluN2B; NCBI RefSeq: NM_012574.1) subunits, and GFP-tagged human GluN3A (GFP-hGluN3A; NCBI RefSeq: NM_133445.3)27,77. The YFP-tagged hGluN1-1a subunit (NCBI RefSeq NM_017010.2 was modified to the human version as described above)76 was cloned into the FHUGW lentivirus vector containing 20 sense nucleotides in the GluN1 target sequence (gac cgg aag ttt gcc aac ta; with a short hairpin (AAGCTT) and 20 antisense nucleotides cloned downstream of the H1 promoter) to knock down the endogenous GluN1 subunit, as described previously53. Silent mutations (gac cgC aaA ttC gcG aac ta; the mutated nucleotides are indicated in capital letters) were introduced in order to generate shRNA-resistant versions of YFP-hGluN1-1a and YFP-hGluN1-1aS688Y . All mutations were introduced using the Quick-Change site-directed mutagenesis kit (Agilent Technologies), and the full GluN-coding sequences were confirmed by sequencing. Lentiviruses expressing YFP-hGluN1-1a and YFP-hGluN1-1a-S688Y were prepared in HEK293T cells by co-transfecting the FHUGW lentiviral vector with Δ8.9 and VSVG as described previously53, and supernatants containing the viral particles were harvested 60 h after transfection.
Mammalian cell culture and primary hippocampal neurons
Human embryonic kidney 293 (HEK293) cells were maintained in Opti-MEM I media containing 5% (v/v) foetal bovine serum (FBS; Thermo Fisher Scientific). The cells were transfected using Lipofectamine 2000 (Thermo Fisher Scientific) as described previously64. After transfection, the HEK293 cells used for electrophysiology were dissociated with trypsin; the cells used for microscopy and biochemistry were cultured without the trypsinisation step in culture media containing 1% FBS, 20 mM MgCl2, 1 mM D,L-2-amino-5-phosphonopentanoic acid, and 3 mM kynurenic acid (to prevent cell death caused by excessive activation of NMDARs). The cells were plated on poly-L-lysine-coated glass coverslips and were used 24–48 h after transfection.
All procedures involving the use of laboratory animals were performed in accordance with the European Communities Council Directive November 24, 1986 (86/609/EEC) and animal care guidelines approved by the Institute of Experimental Medicine CAS Animal Care Committee. Primary cultures of hippocampal neurons were prepared from embryonic day 18 Wistar rats53. In brief, the hippocampi were removed, placed in cold Hank's Balanced Salt Solution containing 10 mM HEPES (pH 7.4), and incubated for 20 min at 37 °C in dissection media containing 0.1 mg/ml DNase I and 0.05% trypsin (Merck). The cells were then dissociated by trituration through a fire-polished glass pipette and resuspended in plating medium consisting of Neurobasal media with B-27 supplement and l-glutamine (Thermo Fisher Scientific). The cells were grown at a density of approximately 2 × 104 cells per cm2 on dishes coated with poly- l-lysine (Sigma). The neurons were fed every 7 days with fresh plating media, infected with lentiviruses after 5–7 in culture, and used for experiments 10 days after infection.
Electrophysiology
Whole-cell patch-clamp recordings were performed using an Axopatch 200B amplifier (Molecular Devices) at room temperature using intracellular recording solution containing (in mM): 125 gluconic acid, 15 CsCl, 5 BAPTA, 10 HEPES, 3 MgCl2, 0.5 CaCl2, and 2 ATP-Mg salt (pH adjusted to 7.2 with CsOH)75. Glass patch pipettes (3–6 MΩ tip resistance) were prepared using a model P-1000 micropipette puller (Sutter Instrument Co.). A microprocessor-controlled multi-barrel rapid perfusion system (with a time constant for solution exchange around the cell of approximately 20 ms) was used to apply the extracellular recording solution (ECS)75,78. The control ECS contained (in mM): 160 NaCl, 2.5 KCl, 10 HEPES, 10 glucose, 0.2 EDTA, and 0.7 CaCl2 (pH adjusted to 7.3 with NaOH)79. When recording hippocampal neurons, the ECS also contained 1 µM tetrodotoxin (TTX) and 10 µM bicuculine. pCLAMP 9 software (Molecular Devices) was used to record and analyse the NMDAR-induced currents recorded in voltage-clamp mode at a membrane potential of − 60 mV. The normalised steady-state and peak concentration–response data for each recording were best-fitted using the following equation: I = Imax/(1 + (EC50/[Agonist])h), where Imax is the maximum peak current in response to agonist, EC50 is the agonist concentration (in μM) that elicited the half-maximal response, [Agonist] is agonist concentration (in μM), and h is the apparent Hill coefficient. Data were fitted using SigmaPlot 14.0 (Systat Software, Inc.).
Surface expression analysis
HEK293 cells grown in 12-well plates were transfected with GluN subunit combinations using Lipofectamine 2000 as described previously27. Live-cell labelling of GFP/YFP-GluN subunits at the cell surface was performed using phosphate-buffered saline (PBS) containing 0.2% bovine albumin serum (BSA) and the rabbit anti-GFP primary antibody (1:1000; Merck) followed by an anti-rabbit antibody conjugated to either Alexa Fluor 555 (1:1000; Thermo Fisher Scientific; for HEK293 cells) or Alexa Fluor 647 (1:1000; Thermo Fisher Scientific; for hippocampal neurons)27,53,77. The stained cells were then fixed in 4% paraformaldehyde (PFA) in PBS for 20 min. To label the total pool of GluN subunits, fixed cells were permeabilised with 0.25% Triton X-100 (Serva) in PBS for 5 min. The cells were then labelled in PBS containing 0.2% BSA (HEK293 cells) or 3% normal goat serum (hippocampal neurons) containing the mouse anti-GFP antibody followed by an anti-mouse antibody conjugated to Alexa Fluor 488 (1:1000; Thermo Fisher Scientific)27,53,77. The labelled cells were mounted using ProLong Gold Antifade reagent (Thermo Fisher Scientific) and imaged using an Olympus FV10i confocal microscope with a 60x/1.35 oil immersion objective (for HEK293 cells) or a Leica SP8 confocal scanning microscope with a 63x/1.40 oil immersion apochromatic objective (for hippocampal neurons). The z-stack for all images was 0.3 µm, and the resolution was 512 × 512 pixels and 2048 × 2048 pixels for HEK293 cells and hippocampal neurons, respectively. The surface and total fluorescence intensities of HEK293 cells were analysed on whole-cell areas using ImageJ 1.52 N (National Institutes of Health, Bethesda, MD). For hippocampal neurons, the fluorescence intensity of the total and surface signals was analysed for 5 separate 10-µm segments of secondary or tertiary dendrites per neuron27,53,80. Prior to the intensity analysis, a z-stack projection was made with maximal intensity from the bottom of the cell to the top of the cell.
Biochemistry
For the surface biotinylation assay81, cultured hippocampal neurons were labelled with 0.5 mg/ml EZ-Link Sulfo-NHS-LC-biotin (Thermo Fisher Scientific) in PBS++ solution (0.01 M PBS, pH 7.4, supplemented with 1 mM MgCl2 and 0.1 mM CaCl2) for 20 min at 4 °C with gentle mixing. The remaining biotin was quenched with PBS++ containing 50 mM glycine. The biotinylated neurons were lysed, centrifuged, and the supernatant was incubated with 20 µl of streptavidin-agarose beads (Thermo Fisher Scientific) for 3 h at 4 °C. After washing the beads 3 times with lysis buffer, the bound proteins were eluted and analysed using western blot.
For PSD fractionation in neurons82, cultured cortical neurons were homogenised in hypotonic buffer (10 mM Tris–HCl, pH 8.0, 10 mM KCl, 0.1 mM EDTA) and passed through a 23-G needle 20 times to disrupt the plasma membrane. Sucrose was then added to a final concentration of 0.32 M. The nuclear pellet and any remaining intact neurons were removed by centrifugation at 800×g for 5 min. The supernatant was then centrifuged at 18,000×g for 20 min to obtain a crude synaptosome pellet. This pellet was resuspended in TNE buffer (50 mM Tris–HCl, pH 8.0, 150 mM NaCl, 2 mM EDTA) containing a protease inhibitor cocktail (Roche) and solubilised with 1% Triton X-100 for 5 min on ice. The samples were then centrifuged at 18,000×g for 15 min, and the resulting pellet was further solubilised in TNE buffer containing 1% SDS. After the insoluble material was removed, the supernatant was harvested as the PSD fraction.
Calcium imaging
Hippocampal neurons were pre-incubated for 30 min with Fura-2, AM (2.5 μM, Thermo Fisher Scientific) and the non-ionic detergent Pluronic F-127 (0.05% (w/v) in DMSO, Merck), and imaged in ECS containing 1 µM TTX and 10 µM bicuculine. The Fura-2 signal was captured using an inverted AxioObserver D1 microscope controlled with ZEN 2012 software (Zeiss) and equipped with a CCD camera and Lambda-DG4 fast illumination system (Sutter Instruments, Novato) for excitation at 340 and 380 nm. The fluorescence intensity of the Fura-2 emission was measured at 510 nm as a ratio of signals obtained after excitation at 340 and 380 nm. Data was sampled every 500 ms during the Ca2+ imaging. The cells were continuously perfused with ECS at 37 °C, and solutions were exchanged using a multiple capillary perfusion system consisting of a computer-controlled multichannel peristaltic pump (Reglo ICC, Ismatec). Traces of individual cells expressed as the F340/F380 ratio were horizontally aligned by their baseline and analysed using Matlab 2019b (MathWorks). An average trace was calculated as the mean of all recorded cells, with the shaded area corresponding to the standard error of the mean (SEM).
NMDA-induced excitotoxicity
Excitotoxicity was induced as described previously55. In brief, the cultured neurons were incubated overnight in 10% MEM (Thermo Fisher Scientific) and 90% salt-glucose (SG) medium containing 114 mM NaCl, 0.219% NaHCO3, 5.292 mM KCl, 1 mM MgCl2, 2 mM CaCl2, 10 mM HEPES, 30 mM glucose, 0.5 sodium pyruvate, and 0.1% phenol red. The following day, the media was replaced with 100% SG and the indicated concentrations of agonists/co-agonists were added to the neurons. After 1 h, the medium was replaced with 10% MEM and 90% SG medium; 23 h later, the neurons were stained with Hoechst 33,342 (5 µM, Molecular Probes) for 30 min, fixed in 4% PFA, and the YFP-hGluN1 subunits were labelled with rabbit anti-GFP primary and anti-rabbit Alexa Fluor 488 secondary antibodies as described above. The images (1024 × 1024 pixels with a pixel size of 1.243 × 1.243 µm, covering a field of 1272 × 1272 µm) were acquired using an Olympus FV10i confocal microscope with a 60x/1.35 oil immersion objective; the following three images were obtained for each field of view: the YFP signal (for infected cells), Hoechst 33,342 (to stain the nuclei), and a widefield image. Nuclear area was measured using ImageJ software (v. 1.52p), and custom-made macro scripts were used to automatically measure only the nuclei of infected cells (identified by YFP expression). We plotted all of the measured nuclei from a single experiment as a histogram containing control (with mostly non-pyknotic cells) and each tested conditions (mostly with a mixture of pyknotic and non-pyknotic cells). The histogram contained two clearly distinguishable groups corresponding to pyknotic cells and non-pyknotic cells. The MatLab function “fitgmdist()” was used to fit the histograms with two Gaussian functions. The data for each condition were then passed with the estimated Gaussian mixture model parameters in the MatLab function “cluster()”, which estimated the posterior probability of each nuclear area belonging to one of the distributions and classified the cells into two groups. The ratio of the number of cells in each group was then used to estimate the effects of mixing the two distributions and is expressed as the ratio of pyknotic and non-pyknotic cells. The percentage of pyknotic cells in each condition was then calculated as the number of cells classified as pyknotic divided by the total number of cells in that condition.
Statistical analysis
Except where indicated otherwise, all summary data are presented as the mean ± SEM. Group differences were analysed using the Student’s t-test or a one-way ANOVA followed by Dunnett’s post hoc text or Shapiro–Wilk normality test. Data were analysed using SigmaStat 3.5 (Systat Software, Inc.), and differences with a p value < 0.05 were considered significant.
Data availability
All materials will be provided promptly upon request without undue qualifications for material transfer agreement.
References
Paoletti, P., Bellone, C. & Zhou, Q. NMDA receptor subunit diversity: impact on receptor properties, synaptic plasticity and disease. Nat. Rev. Neurosci. 14, 383–400. https://doi.org/10.1038/nrn3504 (2013).
Sanz-Clemente, A., Gray, J. A., Ogilvie, K. A., Nicoll, R. A. & Roche, K. W. Activated CaMKII couples GluN2B and casein kinase 2 to control synaptic NMDA receptors. Cell Rep. 3, 607–614 (2013).
Traynelis, S. F. et al. Glutamate receptor ion channels: structure, regulation, and function. Pharmacol. Rev. 62, 405–496 (2010).
Sanz-Clemente, A., Nicoll, R. A. & Roche, K. W. Diversity in NMDA receptor composition: many regulators, many consequences. Neuroscientist https://doi.org/10.1177/1073858411435129 (2012).
Vieira, M., Yong, X. L. H., Roche, K. W. & Anggono, V. Regulation of NMDA glutamate receptor functions by the GluN2 subunits. J. Neurochem. https://doi.org/10.1111/jnc.14970 (2020).
Rauner, C. & Kohr, G. Triheteromeric NR1/NR2A/NR2B receptors constitute the major N-methyl-D-aspartate receptor population in adult hippocampal synapses. J. Biol. Chem. 286, 7558–7566. https://doi.org/10.1074/jbc.M110.182600 (2011).
Al-Hallaq, R. A., Conrads, T. P., Veenstra, T. D. & Wenthold, R. J. NMDA di-heteromeric receptor populations and associated proteins in rat hippocampus. J. Neurosci. 27, 8334–8343 (2007).
Stroebel, D., Carvalho, S., Grand, T., Zhu, S. & Paoletti, P. Controlling NMDA receptor subunit composition using ectopic retention signals. J. Neurosci. 34, 16630–16636. https://doi.org/10.1523/JNEUROSCI.2736-14.2014 (2014).
Hansen, K. B., Ogden, K. K., Yuan, H. & Traynelis, S. F. Distinct functional and pharmacological properties of Triheteromeric GluN1/GluN2A/GluN2B NMDA receptors. Neuron 81, 1084–1096 (2014).
Perez-Otano, I. et al. Assembly with the NR1 subunit is required for surface expression of NR3A-containing NMDA receptors. J Neurosci 21, 1228–1237 (2001).
Sasaki, Y. F. et al. Characterization and comparison of the NR3A subunit of the NMDA receptor in recombinant systems and primary cortical neurons. J. Neurophysiol. 87, 2052–2063. https://doi.org/10.1152/jn.00531.2001 (2002).
Perez-Otano, I., Larsen, R. S. & Wesseling, J. F. Emerging roles of GluN3-containing NMDA receptors in the CNS. Nat. Rev. Neurosci. 17, 623–635. https://doi.org/10.1038/nrn.2016.92 (2016).
Grand, T., Abi Gerges, S., David, M., Diana, M. A. & Paoletti, P. Unmasking GluN1/GluN3A excitatory glycine NMDA receptors. Nat. Commun. 9, 4769. https://doi.org/10.1038/s41467-018-07236-4 (2018).
Pina-Crespo, J. C. et al. Excitatory glycine responses of CNS myelin mediated by NR1/NR3 “NMDA” receptor subunits. J. Neurosci. 30, 11501–11505. https://doi.org/10.1523/JNEUROSCI.1593-10.2010 (2010).
Otsu, Y. et al. Control of aversion by glycine-gated GluN1/GluN3A NMDA receptors in the adult medial habenula. Science 366, 250–254. https://doi.org/10.1126/science.aax1522 (2019).
Chatterton, J. E. et al. Excitatory glycine receptors containing the NR3 family of NMDA receptor subunits. Nature 415, 793–798. https://doi.org/10.1038/nature715 (2002).
Kleckner, N. W. & Dingledine, R. Requirement for glycine in activation of NMDA-receptors expressed in Xenopus oocytes. Science 241, 835–837 (1988).
Clements, J. D. & Westbrook, G. L. Activation kinetics reveal the number of glutamate and glycine binding sites on the N-methyl-D-aspartate receptor. Neuron 7, 605–613 (1991).
Patneau, D. K. & Mayer, M. L. Structure-activity relationships for amino acid transmitter candidates acting at N-methyl-D-aspartate and quisqualate receptors. J. Neurosci. 10, 2385–2399 (1990).
Awobuluyi, M. et al. Subunit-specific roles of glycine-binding domains in activation of NR1/NR3 N-methyl-D-aspartate receptors. Mol. Pharmacol. 71, 112–122 (2007).
Kvist, T., Greenwood, J. R., Hansen, K. B., Traynelis, S. F. & Brauner-Osborne, H. Structure-based discovery of antagonists for GluN3-containing N-methyl-D-aspartate receptors. Neuropharmacology 75, 324–336 (2013).
Madry, C. et al. Principal role of NR3 subunits in NR1/NR3 excitatory glycine receptor function. Biochem. Biophys. Res. Commun. 354, 102–108 (2007).
Kehoe, L. A., Bernardinelli, Y. & Muller, D. GluN3A: an NMDA receptor subunit with exquisite properties and functions. Neural Plast. 2013, 145387. https://doi.org/10.1155/2013/145387 (2013).
Petralia, R. S., Al-Hallaq, R. A. & Wenthold, R. J. Trafficking and targeting of NMDA receptors. Biol. NMDA Recep. 1, 149–200 (2009).
Horak, M., Petralia, R. S., Kaniakova, M. & Sans, N. ER to synapse trafficking of NMDA receptors. Front. Cell Neurosci. 8, 394. https://doi.org/10.3389/fncel.2014.00394 (2014).
Hansen, K. B., Furukawa, H. & Traynelis, S. F. Control of assembly and function of glutamate receptors by the amino-terminal domain. Mol. Pharmacol. 78, 535–549 (2010).
Skrenkova, K. et al. Structural features in the glycine-binding sites of the GluN1 and GluN3A subunits regulate the surface delivery of NMDA receptors. Sci. Rep. 9, 12303. https://doi.org/10.1038/s41598-019-48845-3 (2019).
Hawkins, L. M. et al. Export from the endoplasmic reticulum of assembled N-methyl-d-aspartic acid receptors is controlled by a motif in the c terminus of the NR2 subunit. J. Biol. Chem. 279, 28903–28910. https://doi.org/10.1074/jbc.M402599200 (2004).
She, K., Ferreira, J. S., Carvalho, A. L. & Craig, A. M. Glutamate binding to the GluN2B subunit controls surface trafficking of N-methyl-D-aspartate (NMDA) receptors. J. Biol. Chem. 287, 27432–27445 (2012).
Ferreira, J. S. et al. Co-agonists differentially tune GluN2B-NMDA receptor trafficking at hippocampal synapses. Elife https://doi.org/10.7554/eLife.25492 (2017).
Nong, Y. et al. Glycine binding primes NMDA receptor internalization. Nature 422, 302–307. https://doi.org/10.1038/nature01497 (2003).
Hardingham, G. E. & Do, K. Q. Linking early-life NMDAR hypofunction and oxidative stress in schizophrenia pathogenesis. Nat. Rev. Neurosci. 17, 125–134 (2016).
Chen, W. et al. Functional evaluation of a de novo GRIN2A mutation identified in a patient with profound global developmental delay and refractory epilepsy. Mol. Pharmacol. 91, 317–330 (2017).
Hu, C., Chen, W., Myers, S. J., Yuan, H. & Traynelis, S. F. Human GRIN2B variants in neurodevelopmental disorders. J. Pharmacol. Sci. 132, 115–121. https://doi.org/10.1016/j.jphs.2016.10.002 (2016).
Li, D. et al. GRIN2D recurrent de novo dominant mutation causes a severe epileptic encephalopathy treatable with nmda receptor channel blockers. Am. J. Hum. Genet. 99, 802–816 (2016).
Tarabeux, J. et al. Rare mutations in N-methyl-D-aspartate glutamate receptors in autism spectrum disorders and schizophrenia. Transl. Psychiatry 1, e55 (2011).
Shen, Y. C. et al. Exomic sequencing of the ionotropic glutamate receptor N-methyl-D-aspartate 3A gene (GRIN3A) reveals no association with schizophrenia. Schizophr. Res. 114, 25–32. https://doi.org/10.1016/j.schres.2009.07.005 (2009).
Strehlow, V. et al. GRIN2A-related disorders: genotype and functional consequence predict phenotype. Brain 142, 80–92. https://doi.org/10.1093/brain/awy304 (2019).
Amin, J. B., Leng, X., Gochman, A., Zhou, H. X. & Wollmuth, L. P. A conserved glycine harboring disease-associated mutations permits NMDA receptor slow deactivation and high Ca(2+) permeability. Nat. Commun. 9, 3748. https://doi.org/10.1038/s41467-018-06145-w (2018).
Swanger, S. A. et al. Mechanistic insight into NMDA receptor dysregulation by rare variants in the GluN2A and GluN2B agonist binding domains. Am. J. Hum. Genet. 99, 1261–1280. https://doi.org/10.1016/j.ajhg.2016.10.002 (2016).
Platzer, K. et al. GRIN2B encephalopathy: novel findings on phenotype, variant clustering, functional consequences and treatment aspects. J. Med. Genet. 54, 460–470. https://doi.org/10.1136/jmedgenet-2016-104509 (2017).
Hornig, T. et al. GRIN3B missense mutation as an inherited risk factor for schizophrenia: whole-exome sequencing in a family with a familiar history of psychotic disorders. Genet Res (Camb) 99, e1. https://doi.org/10.1017/S0016672316000148 (2017).
Zehavi, Y. et al. De novo GRIN1 mutations: an emerging cause of severe early infantile encephalopathy. Eur. J. Med. Genet. 60, 317–320. https://doi.org/10.1016/j.ejmg.2017.04.001 (2017).
Lemke, J. R. et al. Delineating the GRIN1 phenotypic spectrum: a distinct genetic NMDA receptor encephalopathy. Neurology 86, 2171–2178. https://doi.org/10.1212/WNL.0000000000002740 (2016).
Redin, C. et al. Efficient strategy for the molecular diagnosis of intellectual disability using targeted high-throughput sequencing. J. Med. Genet. 51, 724–736. https://doi.org/10.1136/jmedgenet-2014-102554 (2014).
Ohba, C. et al. GRIN1 mutations cause encephalopathy with infantile-onset epilepsy, and hyperkinetic and stereotyped movement disorders. Epilepsia 56, 841–848. https://doi.org/10.1111/epi.12987 (2015).
Fry, A. E. et al. De novo mutations in GRIN1 cause extensive bilateral polymicrogyria. Brain 141, 698–712. https://doi.org/10.1093/brain/awx358 (2018).
Hamdan, F. F. et al. Excess of de novo deleterious mutations in genes associated with glutamatergic systems in nonsyndromic intellectual disability. Am. J. Hum. Genet. 88, 306–316. https://doi.org/10.1016/j.ajhg.2011.02.001 (2011).
Hackos, D. H. et al. Positive allosteric modulators of GluN2A-containing NMDARs with distinct modes of action and impacts on circuit function. Neuron 89, 983–999. https://doi.org/10.1016/j.neuron.2016.01.016 (2016).
Armstrong, N. & Gouaux, E. Mechanisms for activation and antagonism of an AMPA-sensitive glutamate receptor: crystal structures of the GluR2 ligand binding core. Neuron 28, 165–181. https://doi.org/10.1016/s0896-6273(00)00094-5 (2000).
Chen, W. et al. GRIN1 mutation associated with intellectual disability alters NMDA receptor trafficking and function. J. Hum. Genet. 62, 589–597. https://doi.org/10.1038/jhg.2017.19 (2017).
Prybylowski, K. et al. Relationship between availability of NMDA receptor subunits and their expression at the synapse. J. Neurosci. 22, 8902–8910 (2002).
Lichnerova, K. et al. Two N-glycosylation sites in the GluN1 subunit are essential for releasing N-methyl-d-aspartate (NMDA) receptors from the endoplasmic reticulum. J. Biol. Chem. 290, 18379–18390. https://doi.org/10.1074/jbc.M115.656546 (2015).
Hardingham, G. E. & Bading, H. Synaptic versus extrasynaptic NMDA receptor signalling: implications for neurodegenerative disorders. Nat. Rev. Neurosci. 11, 682–696 (2010).
McQueen, J. et al. Pro-death NMDA receptor signaling is promoted by the GluN2B C-terminus independently of Dapk1. Elife https://doi.org/10.7554/eLife.17161 (2017).
Nicoll, R. A. A brief history of long-term potentiation. Neuron 93, 281–290. https://doi.org/10.1016/j.neuron.2016.12.015 (2017).
Morris, R. G. NMDA receptors and memory encoding. Neuropharmacology 74, 32–40. https://doi.org/10.1016/j.neuropharm.2013.04.014 (2013).
Chen, P. E. et al. Modulation of glycine potency in rat recombinant NMDA receptors containing chimeric NR2A/2D subunits expressed in Xenopus laevis oocytes. J. Physiol. 586, 227–245. https://doi.org/10.1113/jphysiol.2007.143172 (2008).
Dravid, S. M. et al. Structural determinants of D-cycloserine efficacy at the NR1/NR2C NMDA receptors. J. Neurosci. 30, 2741–2754. https://doi.org/10.1523/JNEUROSCI.5390-09.2010 (2010).
Kuryatov, A., Laube, B., Betz, H. & Kuhse, J. Mutational analysis of the glycine-binding site of the NMDA receptor: structural similarity with bacterial amino acid-binding proteins. Neuron 12, 1291–1300. https://doi.org/10.1016/0896-6273(94)90445-6 (1994).
Furukawa, H. & Gouaux, E. Mechanisms of activation, inhibition and specificity: crystal structures of the NMDA receptor NR1 ligand-binding core. EMBO J. 22, 2873–2885. https://doi.org/10.1093/emboj/cdg303 (2003).
Tovar, K. R., McGinley, M. J. & Westbrook, G. L. Triheteromeric NMDA receptors at hippocampal synapses. J. Neurosci. 33, 9150–9160 (2013).
Rozeboom, A. M. et al. Evidence for glycinergic GluN1/GluN3 NMDA receptors in hippocampal metaplasticity. Neurobiol. Learn. Mem. 125, 265–273. https://doi.org/10.1016/j.nlm.2015.10.005 (2015).
Skrenkova, K. et al. N-glycosylation regulates the trafficking and surface mobility of GluN3A-containing NMDA receptors. Front. Mol. Neurosci. 11, 188. https://doi.org/10.3389/fnmol.2018.00188 (2018).
Lin, Y. C. et al. Genome dynamics of the human embryonic kidney 293 lineage in response to cell biology manipulations. Nat. Commun. 5, 4767. https://doi.org/10.1038/ncomms5767 (2014).
Laube, B., Hirai, H., Sturgess, M., Betz, H. & Kuhse, J. Molecular determinants of agonist discrimination by NMDA receptor subunits: analysis of the glutamate binding site on the NR2B subunit. Neuron 18, 493–503 (1997).
Wells, G. et al. The GluN2B-Glu413Gly NMDA receptor variant arising from a de novo GRIN2B mutation promotes ligand-unbinding and domain opening. Proteins 86, 1265–1276. https://doi.org/10.1002/prot.25595 (2018).
Kehoe, L. A. et al. GluN3A promotes dendritic spine pruning and destabilization during postnatal development. J. Neurosci. 34, 9213–9221. https://doi.org/10.1523/JNEUROSCI.5183-13.2014 (2014).
Fiuza, M., Gonzalez-Gonzalez, I. & Perez-Otano, I. GluN3A expression restricts spine maturation via inhibition of GIT1/Rac1 signaling. Proc. Natl. Acad. Sci. USA 110, 20807–20812. https://doi.org/10.1073/pnas.1312211110 (2013).
Wang, H. et al. The GluN3A subunit exerts a neuroprotective effect in brain ischemia and the hypoxia process. ASN Neuro 5, 231–242. https://doi.org/10.1042/AN20130009 (2013).
Martinez-Turrillas, R. et al. The NMDA receptor subunit GluN3A protects against 3-nitroproprionic-induced striatal lesions via inhibition of calpain activation. Neurobiol Dis 48, 290–298. https://doi.org/10.1016/j.nbd.2012.07.001 (2012).
Ramachandran, G. N., Ramakrishnan, C. & Sasisekharan, V. Stereochemistry of polypeptide chain configurations. J. Mol. Biol. 7, 95–000. https://doi.org/10.1016/S0022-2836(63)80023-6 (1963).
O’Boyle, N. M. et al. Open babel: an open chemical toolbox. J. Cheminform. 3, 33. https://doi.org/10.1186/1758-2946-3-33 (2011).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461. https://doi.org/10.1002/jcc.21334 (2010).
Hemelikova, K., Kolcheva, M., Skrenkova, K., Kaniakova, M. & Horak, M. Lectins modulate the functional properties of GluN1/GluN3-containing NMDA receptors. Neuropharmacology 157, 107671. https://doi.org/10.1016/j.neuropharm.2019.107671 (2019).
Vyklicky, V. et al. Surface expression, function, and pharmacology of disease-associated mutations in the membrane domain of the human GluN2B subunit. Front. Mol. Neurosci. 11, 110. https://doi.org/10.3389/fnmol.2018.00110 (2018).
Kaniakova, M. et al. Key amino acid residues within the third membrane domains of NR1 and NR2 subunits contribute to the regulation of the surface delivery of N-methyl-D-aspartate receptors. J. Biol. Chem. 287, 26423–26434 (2012).
Turecek, R. et al. Intracellular spermine decreases open probability of N-methyl-D-aspartate receptor channels. Neuroscience 125, 879–887. https://doi.org/10.1016/j.neuroscience.2004.03.003S0306452204001757 (2004).
Gazova, Z. et al. Multi-target-directed therapeutic potential of 7-methoxytacrine-adamantylamine heterodimers in the Alzheimer’s disease treatment. Biochim. Biophys. Acta Mol. Basis Dis. 607–619, 2017. https://doi.org/10.1016/j.bbadis.2016.11.020 (1863).
Kaniakova, M., Lichnerova, K., Vyklicky, L. & Horak, M. Single amino acid residue in the M4 domain of GluN1 subunit regulates the surface delivery of NMDA receptors. J. Neurochem. https://doi.org/10.1111/jnc.12002 (2012).
Lee, S. et al. Nedd4 E3 ligase and beta-arrestins regulate ubiquitination, trafficking, and stability of the mGlu7 receptor. Elife https://doi.org/10.7554/eLife.44502 (2019).
Terashima, A., Suh, Y. H. & Isaac, J. T. R. The AMPA receptor subunit GluA1 is required for CA1 hippocampal long-term potentiation but is not essential for synaptic transmission. Neurochem. Res. 44, 549–561. https://doi.org/10.1007/s11064-017-2425-3 (2019).
Acknowledgements
This work was supported by the Czech Science Foundation (20-12420S). The in silico study was also supported by MH CZ—DRO (University Hospital Hradec Kralove, Nr. 00179906), Long-term Development Plan (Faculty of Military Health Sciences), and the Ministry of Health of the Czech Republic (NU20-08-00296).
Author information
Authors and Affiliations
Contributions
M.H. and K.S. designed the study; K.S., J-m.S., K.H., M.K., J.N. and M.Ka. performed the experiments; K.S., K.H., M.K., S.K., and B.H.K. analysed the data; J.K., and T.K. performed the in silico analyses; and K.S., B.H.K., Y.H.S., and M.H. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Skrenkova, K., Song, Jm., Kortus, S. et al. The pathogenic S688Y mutation in the ligand-binding domain of the GluN1 subunit regulates the properties of NMDA receptors. Sci Rep 10, 18576 (2020). https://doi.org/10.1038/s41598-020-75646-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-75646-w
- Springer Nature Limited