Abstract
The family Richtersiidae, although established recently with the use of phylogenetic methods, was considered potentially paraphyletic at the time of its erection. Until now, the family comprised four genera, Richtersius, Diaforobiotus, Adorybiotus and a newly erected genus Crenubiotus. However, the genetic characterisation for the latter two genera was very limited or absent. To address concerns about the phylogenetic affinity of these two genera, we present a multilocus phylogeny of the families Richtersiidae and Murrayidae based on four molecular markers (18S rRNA, 28S rRNA, ITS-2 and COI). Our results show a distinct evolutionary lineage composed of Adorybiotus and Crenubiotus, which is sister to Murrayidae. In order to accommodate the phylogenetic and morphological distinctiveness of this lineage, we erect a new family, Adorybiotidae fam. nov. The new taxon differs morphologically from other families in the superfamily Macrobiotoidea by a unique combination of traits: (1) the presence of tubercles/cushions with aggregations of microgranules on their surfaces present on all legs and on the dorso-caudal cuticle, (2) a system of internal septa in claws, and (3) buccal apparatus morphology. Moreover, in order to stabilise the taxonomy and nomenclature in the genus Crenubiotus, we redescribe its type species, Crenubiotus crenulatus, by means of integrative taxonomy and designate a new neotype based on a population from the original terra typica.
Similar content being viewed by others
Introduction
Tardigrades are a phylum of microinvertebrates which are found in freshwater, marine and limno-terrestrial environments throughout the world1. The first formally described tardigrade was Macrobiotus hufelandi C.A.S. Schultze, 18342 and currently over 1300 nominal taxa are recognised within the phylum3,4,5. Although research on tardigrade systematics started almost two centuries ago, only recently have studies aided by molecular phylogenetics begun to shed more light onto the relationships between taxa within the phylum, e.g.6,7,8,9,10,11. Thanks to genetic data for already known, as well as for newly detected species, the discovery and the delimitation of new high rank taxa, such as genera and families, have become more frequent. One example of this trend is the recent erection of the family Richtersiidae Guidetti et al., 201611, which currently comprises four genera: Adorybiotus Maucci & Ramazzotti, 198112, Crenubiotus Lisi et al., 202013, Diaforobiotus Guidetti et al., 201611 and Richtersius Pilato & Binda, 198914. Guidetti et al.11 explicitly demonstrated that the genera Adorybiotus, Diaforobiotus and Richtersius do not belong to Macrobiotidae Thulin, 192815 and a family of their own was established. However, at the same time, Guidetti et al.11 also stressed that the relationships within Richtersiidae need to be clarified with further molecular data as their results indicated a polyphyletic status of the family. More specifically, Adorybiotus was shown to be more closely related to Murrayidae than to the other members of Richtersiidae (Diaforobiotus + Richtersius). Nevertheless, based on morphological similarities, Adorybiotus was provisionally included within the Richtersiidae11. The same pattern of relatedness within the family was also recovered by the phylogenetic analysis in Guil et al.16, based—as in Guidetti et al.11—on two conservative ribosomal markers, 18S rRNA and 28S rRNA. Finally, very recently, Lisi et al.13 erected a new genus Crenubiotus within Richtersiidae based exclusively on morphological evidence. Crenubiotus comprises two species: C. crenulatus (Richters, 1904)17, originally described as a Macrobiotus from Svalbard, now the type species for the genus Crenubiotus, and C. revelator Lisi et al., 202013 discovered in Colombia. Lisi et al.13 refined the morphological characters defining the family Richtersiidae as follows: (1) the presence of an additional apophysis on the ventral lamina, (2) the presence of a dorsal apophysis on the buccal tube (reduced in Crenubiotus and Diaforobiotus), (3) a specific type of claws with a system of internal septa and clearly dentate lunulae on all legs, (4) the presence of cuticular pores (at least in some life stages), and (5) the presence of two macroplacoids in the bulbus.
In this study, we present an upgraded molecular phylogeny of the families Richtersiidae and Murrayidae based on four genetic markers (18S rRNA, 28S rRNA, ITS-2 and COI). New DNA sequences for six species/populations of Adorybiotus, Crenubiotus and Diaforobiotus, as well as two additional species of the family Murrayidae are added to the dataset available from earlier studies. Our results show that Adorybiotus and Crenubiotus form a clade that is more closely related to the family Murrayidae than to the other two genera of Richtersiidae (Richtersius and Diaforobiotus). These results, together with evident differences recovered by morphological analysis, led us to the erection of a new family within Macrobiotoidea. Finally, the analysis of a population of Crenubiotus crenulatus from the original terra typica (Svalbard, Norway) under the integrative taxonomy framework enabled us to redescribe the species and propose a new neotype that will stabilise the taxonomy and will help to uncover the diversity within this recently erected genus.
Results
Phylogenetic analysis
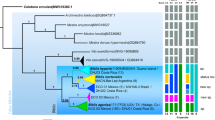
We have obtained good quality sequences of all four markers for all sequenced individuals. The phylogenetic reconstruction performed with BI and ML methods (Fig. 1) showed almost identical topologies, with lower support values for the ML tree. The superfamily Macrobiotoidea was recovered monophyletic and composed of four well supported clades (Fig. 1; the following sentences decribe the topology of the tree from the bottom to the top). The first clade represents the family Macrobiotidae, the second clade comprises a part of the family Richtersiidae (Richtersius and Diaforobiotus but not Adorybiotus and Crenubiotus), the third clade contains Murrayidae (i.e. Murrayon and Dactylobiotus), and the fourth clade is composed of Adorybiotus and Crenubiotus. The last three clades are more closely related to each other than to the family Macrobiotidae. Moreover, the family Richtersiidae is a sister group to Murrayidae + (Adorybiotus + Crenubiotus). Given the evident phylogenetic and morphological distinctiveness, the Adorybiotus + Crenubiotus clade is further elevated to the family level (see the next section below for more details). The majority of the genera in the families Richtersiidae, Murrayidae and in the new family Adorybiotidae were retrieved as monophyletic. The only two paraphyletic genera in our reconstruction were Murrayon and Adorybiotus. In the genus Murrayon, M. dianae was found to be sister to the Murrayon cf. pullari + the Dactylobiotus clade. A similar topology was found for Adorybiotus, with Adorybiotus sp. JP.008 being sister to Adorybiotus granulatus + Crenubiotus crenulatus.
Phylogenetic reconstruction of the superfamily Macrobiotoidea based on concatenated 18S rRNA + 28S rRNA + ITS-2 + COI nucleotide sequences. Topology and branch length of BI reconstruction. Nodes with BI support under 0.70 were collapsed. Values above branches indicate BI posterior probabilities (pp), values below branches indicate ML bootstrap (bs) support. Black circle and no value = full support in both analyses, i.e. 1.00 for BI or 100 for ML; grey circle = node supported in both analyses but not fully in at least one of them (* = full support, BI/ML values lower than 1.00/100 shown). Newly sequenced taxa/populations are bolded. Neotype or type populations are underlined.
Description of the new family
Systematic and taxonomic account
-
Phylum: Tardigrada Doyère, 184018.
-
Class: Eutardigrada Richters, 192619.
-
Order: Parachela Schuster et al., 198020 (restored in Morek et al.21).
-
Superfamily: Macrobiotoidea Thulin, 192815.
-
Family: Adorybiotidae fam. nov. Stec, Vecchi & Michalczyk.
-
ZooBank: urn:lsid:zoobank.org:act:CC69D220-D0D8-43CA-86AE-E998FB843D6D.
(Figs. 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12).
Figure 2 Adorybiotus cf. granulatus from Japan: buccal apparatus. (a) Dorsal projection of the entire buccal apparatus seen in NCM; left lower insert: ventral projection of the anterior portion of the buccal apparatus; right upper insert: ventral projection of the placoids. (b) Buccal crown and oral cavity followed by the buccal tube opening, frontal view. (c) Buccal crown, lateral view. (d) Placoids, dorsal view. The filled arrow indicates the cuticular hook on the T-shaped apophysis, empty arrows indicate the lateral triangular apophysis, filled indented arrowheads indicate the bulbous apophysis at the anterior end of the ventral lamina, filled flat arrowheads indicate constrictions in the macroplacoids, empty flat arrowheads indicate dorsal spikes, Roman numerals indicate the bands of teeth in the oral cavity. Scale bars in μm.
Figure 3 Figure 4 Adorybiotus cf. granulatus from Japan: cuticular granulation. (a) Granulation on the external surface of leg I (PCM). (b) Granulation on the internal surface of leg I (PCM). (c) Wide granulation band on the caudal end of the body and granulation on legs IV (PCM). (d) Granulation on the external surface of leg II (SEM). (e) Granulation on leg IV (SEM). (f,g) Details of the cuticular granulation (SEM). The flat arrow indicates the cuticular bulge/fold on the external leg surface, the empty flat arrowhead indicates the cuticular bulge/fold on the internal leg surface, the filled indented arrowhead indicates paired muscle attachments under claws, the arrow indicates the wide granulation band on the caudal cuticle. Scale bars in μm.
Figure 5 Adorybiotus cf. granulatus from Japan: eggs morphology. (a,b) Egg surface and processes seen in PCM. (c) Entire egg seen in SEM. (d,e) Details of egg processes morphology seen in SEM. (f) Details of the egg surface between processes seen in SEM. Arrowheads indicate short flexible filaments at the end of multifurcated process apices. Scale bars in μm.
Figure 6 Figure 7 Crenubiotus crenulatus s.s. (Richters, 1904) from Spitsbergen: cuticular granulation seen in PCM. (a) Granulation on the external surface of leg II. (b) Granulation on the internal surface of leg II. (c) The wide granulation band on the caudal cuticle and granulation on legs IV. The filled flat arrowhead indicates the cuticular bulge/fold on the external leg surface, the empty flat arrowhead indicates the cuticular bulge/fold on the internal leg surface, the arrow indicates the wide granulation band on the caudal cuticle. Scale bars in μm.
Figure 8 Crenubiotus crenulatus s.s. (Richters, 1904) from Spitsbergen: cuticular granulation seen in SEM. (a) Granulation on the external surface of leg III. (b) Granulation on the internal surface of leg III. (c) Granulation on leg IV. (d) The wide granulation band on the caudal cuticle and granulation on legs IV. (e) Sparse cuticular granulation and pores on the dorsal cuticle. (f) Sparse cuticular granulation and pores on the ventral cuticle. The filled flat arrow indicates the cuticular bulge/fold on the external leg surface, the empty flat arrowhead indicates the cuticular bulge/fold on the internal leg surface, the filled arrow indicates the wide granulation band on the caudal cuticle, empty arrows indicate the sparse cuticular granulation. Scale bars in μm.
Figure 9 Crenubiotus crenulatus s.s. (Richters, 1904) from Spitsbergen: claws. (a) Claws I (PCM). (b) Lunula II (PCM) (c) The cuticular swelling/thickening under lunula II (PCM). (d) Claws IV (PCM). (e) Lunula IV (PCM). (f) The cuticular swelling/thickening under lunula IV (PCM). (g) Claws I (SEM). (h) Claws IV (SEM). Photos (b,c) and e–f show the same lunula, respectively. Empty flat arrowheads indicate slight constrictions in the middle of the claws, the filled indented arrowhead indicates double muscle attachments below the claws, filled flat arrowheads indicate cuticular swellings/thickenings under lunulae. Scale bars in μm.
Figure 10 Crenubiotus crenulatus s.s. (Richters, 1904) from Spitsbergen: buccal apparatus seen in PCM. (a) Dorsal projection of the entire buccal apparatus. (b) Dorsal view of the oral cavity armature. (c) Ventral view of the oral cavity armature. (d) Lateral view of the anterior portion of the buccal apparatus. (e) Ventral view of placoids. Empty flat arrowheads indicate dorsal spikes, filled indented arrowheads indicate the ventral thickening/additional ridge on the ventral lamina, empty indented arrowheads indicate the third band of teeth in the oral cavity armature, the arrow indicates the putative dorsal residual apophysis, filled flat arrowheads indicate constrictions in the macroplacoids. Scale bars in μm.
Figure 11 Crenubiotus crenulatus s.s. (Richters, 1904) from Spitsbergen: eggs seen in PCM. (a–c) Details of egg processes and surface under a ×1000 magnification, upper inserts shows details of the ‘reticulation’ present in egg processes of a given egg. (d) Midsections of egg processes under a ×1000 magnification. Filled flat arrowheads indicate divided tips of the processes, filled indented arrowheads indicate faint septae, empty flat arrowheads indicate short thickenings/projections around processes bases, empty indented arrowheads indicate thickenings within the processes walls between main and distal parts of the processes, the arrow indicates elongated thickenings/projections around processes the bases. Scale bars in μm, scale bar for inserts in figures a–c are twice as long as those in the main photos.
Diagnosis:
Tubercles/cushions with aggregations of microgranules on their surfaces present on all legs and on the dorso-caudal cuticle. Cuticular pores present in all instars. Double Y-shaped claws, with the two branches forming an evident common tract of a variable length. Large, comb-like lunulae under claws on each leg, equipped with long and evenly distributed teeth. Buccal tube with the ventral lamina and a cuticular thickening (which can form a large apophysis) on the anterio-dorsal wall of the buccal tube. Two macroplacoids and a microplacoid positioned close to the second macroplacoid in the bulbus. Ornamented eggs without areolation on the surface between the egg processes laid freely to the environment.
Type genus:
Adorybiotus Maucci & Ramazzotti, 198112.
Family composition:
Adorybiotus (Figs. 2, 3, 4, 5), Crenubiotus Lisi et al., 202013. (Fig. 6, 7, 8,9, 10, 11, 12).
Adorybiotidae fam. nov., by the combination of morphological characters of animals and eggs is unique within the superfamily Macrobiotoidea, and it differs specifically from the family:
-
Macrobiotidae Thulin, 192815 by: large comb-like lunulae under claws on each leg equipped with long and evenly distributed teeth (lunulae smaller, often without teeth and when equipped with teeth, they are short and not as regularly arranged as in Adorybiotidae fam. nov.); and by the system of internal septa within claws on each leg as described by Lisi et al.13 (the system of internal septa absent in Macrobiotidae).
-
Richtersiidae Guidetti et al., 201611 by: the presence of tubercles/cushions with aggregations of microgranules on their surfaces on all legs and on the dorso-caudal cuticle (granulation on legs and on the dorso-caudal cuticle absent in Richtersiidae); the presence of the microplacoid in the bulbus (microplacoid absent in Richtersiidae); and by egg process morphology (processes in the shape of cones with wide bases and very narrow elongated apices or processes with concave bases in the shape of cooling towers and apices divided into 2–4 thick horizontal branches in Adorybiotidae fam. nov. vs egg process in the shape of elongated, thin, conical spikes in Richtersiidae).
-
Murrayidae Guidetti et al., 200522 by: cuticular pores (absent in Murrayidae); large comb-like lunulae under claws on each leg equipped with long and evenly distributed teeth (lunulae without teeth in Murrayidae); the system of internal septa within claws on each leg as described by Lisi et al.13 (the system of internal septa absent in Murrayidae); the presence of tubercles/cushions with aggregations of microgranules on their surfaces on all legs and on the dorso-caudal cuticle (only regular granulation present but tubercles/cushions absent in Murrayidae); and by claw morphology (the common tract of the claw longer than the half of the entire claw height in Adorybiotidae fam. nov. vs the common tract of the claw shorter than the half of the entire claw height in Murrayidae).
Genus:
Crenubiotus Lisi et al., 202013.
Crenubiotus crenulatus (Richters, 1904)17.
(Tables 1, 2, Figs. 6, 7, 89, 10, 11, 12).
Macrobiotus crenulatus Richters, 190417; Macrobiotus dentatus Binda, 197423.
Material examined:
39 animals, and 22 eggs. Specimens mounted on microscope slides in Hoyer’s medium (28 animals + 16 eggs), fixed on SEM stubs (10 + 6), processed for DNA sequencing (1 animal).
Neotype locality:
78°12′58.4"N, 15°20′45.1"E; 53 m asl: Norway, Svalbard, Spitsbergen, Lower part of the Bjørndalen (Bjørn valley; Nordenskiöld Land); moss from tundra on the valley slope; coll. 23.07.2016 by Wojciech Maciejowski.
Type depositories:
Neotype (slide NO.429.06 with 3 neoparatypes) and remaining 18 neoparatypes (slides: NO.429.*, where the asterisk can be substituted by any of the following numbers 03, 07–09) and 14 eggs (slides: NO.429.*: 01–05) and SEM stub 20.01 are deposited at the Institute of Zoology and Biomedical Research, Jagiellonian University, Gronostajowa 9, 30–387, Kraków, Poland. Six neoparatypes (slide NO.429.11) and two eggs (slide NO.429.10) are deposited in the Pilato and Binda collection at the University of Catania, Italy.
Redescription of Crenubiotus crenulatus (Richters, 1904)
Animals (measurements and statistics in Table 1): When alive, body almost transparent in juveniles and yellowish in adults; after fixation in Hoyer’s medium body transparent (Fig. 6). Eyes present, visible also in specimens mounted in Hoyer’s medium. Body cuticle with larger elliptical (0.8–2.0 µm in diameter) and smaller circular (0.3–0.8 µm) pores distributed randomly on the entire body cuticle with the largest elliptical pores being present in the dorso-cephalic and dorso-caudal cuticle (Figs. 7a–c, 8a–f). Patches of dense very visible granulation that comprises of small tubercles/cushions and aggregations of microgranules on their surfaces present on all legs and on the dorso-caudal cuticle (Figs. 7a–c, 8a–d). These tubercles are slightly less developed and evident compared to Adorybiotus (Fig. 4a–g) but still obvious (Figs. 7a–c, 8a–d). On legs I–III, the dense granulation comprises of two patches on the external and internal leg surfaces respectively, that are connected with each other by a narrower band of granulation extending from them on the proximal leg surface (Figs. 7a,b, 8a,b). Two pulvini are present on each leg I–III, one on the external and the other—on the internal leg surface (Figs. 7a,b, 8a,b). On legs IV, dense granulation covers evenly the dorsal and lateral leg surfaces (Figs. 7c, 8c,d). This dense granulation is also present as a wide granulation band on the caudal cuticle that extends across the terminal body segment from the left body side, through dorsal surface, to the right side (Figs. 7c, 8d). Beside these dense granulation patches, very fine granulation is present and evenly distributed on the entire body surface but visible only under SEM (Fig. 8a–f). Remarks: The large elliptical pores reported by Lisi et al.13 as located laterally to the mouth in C. revelator are also present in C. crenulatus, however their exact position and shape could not be determined as the mouth was retracted in almost all analysed specimens.
Claws slender, of the Richtersiidae type. Primary branches with distinct accessory points, a long, constricted in the middle, common tract with a system of internal septa, and with an evident stalk connecting the claw to the lunula (Fig. 9a–h). The common tract apparently longer than the half of the entire claw height (Fig. 9a,d,g–h). Large, comb-like, triangular lunulae with long and evenly distributed teeth present on all legs (Fig. 9a–h). Under PCM, the lower portion of the lunulae just above the dentation is evidently darker and visible as a dark arc (Fig. 9a–b, d–e). The lunulae are curved and clamped around the cuticular swelling/thickening present under them (Fig. 9g–h) what is well visible in SEM whereas in PCM on the lower focal plane it is visible as a darkening beneath the lunules (Fig. 9c,f). Paired muscle attachments present just below lunulae on legs I–III (Fig. 9a).
Mouth antero-ventral. Bucco-pharyngeal apparatus of modified “Macrobiotus type” (Fig. 10a), i.e. with ten peribuccal lamellae, a rigid buccal tube with the ventral lamina which is provided with an additional ventral thickening in its anterior portion, that appears as an elongated trapezoidal structure pointing towards the mouth opening in the ventral view (Fig. 10c) and is visible as a ridge in the lateral view (Fig. 10d). Based on LCM observations, the oral cavity armature is poorly developed and composed only of the third band of teeth (Fig. 10b,c). The first and the second band of teeth are absent or not visible under LCM (Fig. 10b,c). The teeth of the third band are located within the posterior portion of the oral cavity, anteriorly to the buccal tube opening (Fig. 10b,c). The third band of teeth is divided into the dorsal and the ventral portion. Under LCM, both the dorsal and the ventral portions are seen as two distinct transverse ridges and each of them forms a globular thickening at the medial extremity (Fig. 10b,c). Median teeth absent (Fig. 10b,c). Bulbus spherical (Fig. 10a), with triangular apophyses, three anterior cuticular spikes (typically only two are visible in any given plane) and two rod-shaped macroplacoids (2 < 1) and a microplacoid positioned close to the second macroplacoid (Fig. 10a,e). The first macroplacoid is anteriorly narrowed and constricted in the middle whereas the second has a sub-terminal constriction (Fig. 10e).
Eggs (measurements and statistics in Table 2): Laid freely, yellowish, spherical with conical processes with elongated apices and egg surface without areolation (Figs. 11a–d, 12a–f). The elongated process apices are often multifurcated into short flexible filaments (Figs. 11a–d, 12a–f). These elongated distal portions of the processes seem to be sometimes separated from the lower portion of the process by a faint septum (Fig. 11a) but this is caused a circular thickening on the inner process wall (Fig. 11d). The labyrinthine layer between the process walls is visible as a very faint reticular pattern with circular margins under LCM (Fig. 11a–d). The faint meshes are visible in the lower part of the processes but are not visible in the elongated upper part (Fig. 11a–d). The entire surface of processes is smooth under SEM (Fig. 12a–f). Processes attached to the egg by a ring of short thickenings, seen as dark projections visible only under LCM, which gives the process bases a jagged appearance (Fig. 11b–c). Only rarely some these projections might be elongated making the impression of connection between two neighbouring processes (Fig. 11b), however this character should be treated with a dose of caution as all the eggs were covered with debris, thus these thin connectors may be an artefact. Besides these structures, egg surface between processes appears as smooth under LCM (Fig. 11a–d), whereas it is slightly wrinkled in SEM (Fig. 12a–e).
Reproductive mode
The examination of 28 adults freshly mounted in Hoyer’s medium revealed no testes or spermathecae filled with spermatozoa, which suggests that the species is (at least facultatively) parthenogenetic.
DNA sequences
We obtained sequences for all four of the above-mentioned molecular markers from one of the two individuals destined for DNA extraction and sequencing, which are as follow: 18S rRNA (GenBank: MT812474), 994 bp long; 28S rRNA (MT812468), 735 bp long; ITS-2 (MT812606), 398 bp long; COI (MT808079), 629 bp long.
Phenotypic differential diagnosis
To date, the genus Crenubiotus comprises only two species: the nominal C. crenulatus and an extremely similar species, C. revelator, recently described from Colombia. Despite the overall similarity, C. crenulatus differs from C. revelator by: the absence of the circular median tooth in the ventral portion of the third band of teeth in the oral cavity (the median tooth present in C. revelator), the presence of only two lateral teeth in the ventral portion of the third band of teeth, which have thickenings in their medial extremity that resemble circular teeth (the teeth without thickenings in C. revelator), a shorter ventral lamina (19.8–25.3 µm [53.4–61.7] in C. crenulatus vs. 12.1–18.5 µm [44.5–50.7] in C. revelator), larger eggs (ranges of full and bare diameter for four C. crenulatus eggs: 122.4–126.5 µm and 88.5–93.9 µm vs. 97.8 and 78.1 µm of the sole known egg of C. revelator).
Discussion
By the analysis of both morphological and molecular data, we explicitly demonstrated the presence of a previously unrecognised phyletic lineage within the superfamily Macrobiotoidea which comprises two genera, Adorybiotus and Crenubiotus, for which genetic data were extremely limited or absent. To accommodate the phylogenetic and morphological distinctiveness of this group from the remaining three families within the Macrobiotoidea, we erected the new family Adorybiotidae fam. nov. Furthermore, in order to enhance taxonomic studies on the recently erected genus Crenubiotus and stabilise its nomenclature, we provided an integrative redescription of C. crenulatus based on the population from original terra typica and replaced the existing, inadequate neotype with the new one that is associated with DNA barcodes.
Two genera analysed in this study, Murrayon in the family Murrayidae and Adorybiotus in Adorybiotidae fam. nov., appear to be paraphyletic. As already shown by Bertolani et al.6, Murrayon cf. pullari IT.338 is more closely related to Dactylobiotus than to Murrayon dianae. However, the paraphyly should be treated with great caution because the M. dianae branch is exceptionally long, thus it could lead to topological artefacts. The unbalanced sequencing may also be the cause behind the paraphyly of Adorybiotus, as only the 18S rRNA was sequenced for Adorybiotus granulatus in Bertolani et al.6, in contrast to the other analysed populations of the Adorybiotidae fam. nov., for which a complete set of four markers was available. Thus, a better sampling within Murrayon and Adorybiotus, both in terms of the number of species and sequenced markers, is needed to verify the phyletic relationships within the two genera.
As was pointed out in the Introduction, thanks to the easier acquisition of genetic data and their use in phylogenetic studies, the relationships between major evolutionary lineages within the phylum Tardigrada are being gradually resolved. The increasing popularity of integrative taxonomy also contributed to the recognition of considerable and yet undescribed species diversity within this animal group, e.g.10,11,24,25,26. The recent years showed several times that old, outdated and inadequate tardigrade species descriptions or redescription are often the major obstacle in resolving the systematics within genera and species complexes24,25,27,28,29,30,31. In the families Richtersiidae and Adorybiotidae fam. nov., type species for all genera were described in the beginning of the twentieth century and are insufficient in details under modern standards of tardigrade taxonomy, while the original type series do not exist. However, Richtersius coronifer (Richters, 1903)32, the nominal species for the genus Richtersius, was recently redescribed by Stec et al.31 under the integrative taxonomy framework. Although a neotype for this species was previously established by Maucci and Ramazzotti12, its designation and redescription have been questioned and considered as an impediment for Richtersius taxonomy and nomenclature31,33. Thus, Stec and Michalczyk33 have formally designated a new neotype based on a population from the original locus typicus integratively examined in Stec et al.31. Importantly, Crenubiotus crenulatus (Richters, 1904)17, the type species of its genus, suffers from a similar problem. The species was originally described from the Svalbard Archipelago (specifically from Smeerenburg on Spitsbergen), where it seems to be a common element of the tardigrade fauna, e.g.34,35,36,37,38. Nonetheless, the diagnosis of this species has been questioned for many years, specifically concerning its possible synonymy with Macrobiotus echinogenitus Richters, 1904, also originally described from the Svalbard Archipelago13,17,39. The issue has been clarified to some extent by Binda40, who communicated that the types of C. crenulatus and M. echinogenitus are lost and therefore she redescribed and established neotypes for both those species. Recently, Lisi et al.13 provided further morphological details of the C. crenulatus neotype established by Binda40 in order to differentiate it from a new species from Colombia. Nevertheless, there are some more issues regarding the 1988 neotype designation that have to be stressed and appropriate actions should be undertaken. First of all, the neotype of C. crenulatus established by Binda comes from Italy (Valtellina) which is almost 3800 km away from the original locus typicus. This neotype locality designation does not comply with the International Code of Zoological Nomenclature (Article 75.3.6) that states that the neotype should came as nearly as practicable from the original type locality and, where relevant, from the same geological horizon or host species as the original name-bearing type41. This requirement was in force already when Binda40 made her designation42. Second, the neotype series is in a bad condition, which prevents a detailed morphological characterisation of the species13. Finally, taking into consideration the morphological similarity between C. crenulatus and C. revelator, which has been highlighted in this study and in Lisi et al.13, the future taxonomic studies on the genus Crenubiotus will be challenging without the use of DNA barcodes. Therefore, in order to stabilise the taxonomy and nomenclature within Crenubiotus, in agreement with to the International Code of Zoological Nomenclature, we established a new neotype from a population found at 180 km from the original locus typicus, on the same archipelago. With an integrative redescription that comprises detailed morphological data and associated DNA barcodes, future species identification will be much less problematic that it would have been with Binda types from northern Italy. Furthermore, as the same rule of ICZN that was mentioned above was violated when establishing the neotype of M. echinogenitus based on the population from Algeria, which is ca. 5500 km away from the Svalbard Archipelago, we propose to consider this designation as invalid too. As the original description of M. echinogenitus from Richters (1903)32 is vague and there has been confusion around its identity40, we suggest caution in assigning an individual to this species until it is redescribed with material from the original locus typicus.
As obstacles in the form of incomplete and outdated descriptions of the type species for Richtersius and Crenubiotus (Richtersiidae and Adorybiotidae fam. nov., respectively) have been now removed by Stec et al.31 and this study, respectively, attention should be paid to similar issues in the remaining two genera, Adorybiotus and Diaforobiotus. In this study, we presented some details of Adorybiotus morphology and genetics, but they were all based on an undetermined species from Japan that cannot be confidently identified until Adorybiotus granulatus (Richters, 1903)32, the type and the only species of the genus, is redescribed by means of integrative taxonomy. The species was described originally from Norway (Merok), and later redescribed by Maucci and Ramazzotti12 based also on a population from Norway (Steinkjer). Importantly, however, this designation can be questioned as Maucci and Ramazzotti12 did not fulfil properly the condition given in Article 75.3.4 of the Code41, which requires mentioning the “reasons for believing the name-bearing type specimen(s) (i.e. holotype, or lectotype, or all syntypes, or prior neotype) to be lost or destroyed, and the steps that had been taken to trace it or them” (this requirement was also in force already when Maucci and Ramazzotti12 made their designation42). This opens up the possibility of an extensive taxonomic study on A. granulatus that would result in an integrative redescription and a designation of a new neotype, when a suitable population from central Norway is found. The genus Diaforobiotus comprises currently only two subspecies D. islandicus islandicus (Richters, 1904)43 (the nominal subspecies) and D. islandicus nicaraguensis (Séméria, 1985)44. For years, numerous records of D. islandicus islandicus accumulated in the literature from localities all over the world45,46,47,48 but it has been demonstrated by Guidetti et al.11 as well as by this study that the genus definitely comprises more than one species. Nonetheless, since the types of D. islandicus islandicus do not exist and morphological details of the species are unknown, we consider descriptions of new taxa highly hazardous until the type species is redescribed by means of integrative taxonomy. The Diaforobiotus population from Iceland analysed in this study is a suitable candidate for the neotype population, which together with two other populations from Norway and Indonesia will be revised by us in the near future and published as an integrative revision of the genus. Such integrative redescriptions of the type taxa have opened and will continue to open the windows for describing species diversity within genera or species groups/complexes, further contributing to our understanding of evolution of microscopic invertebrates, including tardigrades.
Material and methods
Samples and specimens
To reconstruct the phylogeny of Richtersiidae and Murrayidae, along with already published data, we analysed eight new populations representing eight species isolated from moss or pond sediment samples collected from eight distinct localities (see Table 3 for details). In our study, by a population we mean a group of conspecific individuals found in a single sample. All samples were processed following a protocol described in detail in Stec et al.49.
DNA sequencing
Genomic DNA was extracted from individual animals following a Chelex 100 resin (BioRad) extraction method by Casquet et al.50 with modifications described in detail in Stec et al.51. Each specimen was mounted in water on a temporary microscope slide and examined under light microscope prior to DNA extraction. We sequenced four DNA fragments, three nuclear (18S rRNA, 28S rRNA, ITS-2) and one mitochondrial (COI) from 2–4 individuals per each of the six newly analysed populations. All fragments were amplified and sequenced according to the protocols described in Stec et al.51; primers with their original references are listed in Table 4. Sequencing products were read with the ABI 3130xl sequencer at the Molecular Ecology Lab, Institute of Environmental Sciences of the Jagiellonian University, Kraków, Poland. Sequences were processed in BioEdit ver. 7.2.552 and submitted to NCBI GenBank.
Phylogenetic analysis
The phylogenetic analyses were conducted using concatenated 18S rRNA + 28S rRNA + ITS-2 + COI sequences for Macrobiotoidea with Bertolanius volubilis (Durante Pasa & Maucci, 1975)53, Eohypsibius nadjae Kristensen, 198254, Hypsibius exemplaris Gąsiorek et al. 201828 and Ramazzottius subanomalus (Biserov, 1985)55 as outgroups. We choose isolates from the families Richtersiidae and Murrayidae with all four sequenced markers (18S rRNA, 28S rRNA, ITS-2 and COI) and isolates that overlap with the sequences produced in this study in the cases of 18S rRNA and 28S rRNA. However, there were four exceptions to this: two Adorybiotus granulatus (Richters, 1903)32 isolates, with only 18S rRNA sequences (HQ604961 and HQ604962), that were included as they are the only available sequences for the nominal species of the genus Adorybiotus (although the species identification is uncertain; see the Discussion for details). The other two exceptions were Murrayon dianae (Kristensen, 1982)54 and Dactylobiotus ovimutans Kihm et al., 202056 that were included to have a better representation of the Murrayidae. For the family Macrobiotidae, one to two species of each genus, for which sequences are available in GenBank, were included in the analysis (see Table 5 for GenBank accession numbers). In the analysed dataset, 82% (31/38) terminals had sequences for all four markers.
The 18S rRNA, 28S rRNA and ITS-2 sequences were aligned using MAFFT ver. 757,58 with the G-INS-i method (thread = 4, threadtb = 5, threadit = 0, reorder, adjustdirection, anysymbol, maxiterate = 1000, retree 1, globalpair input). The COI sequences were aligned according to their aminoacid sequences (translated using the invertebrate mitochondrial code) with the MUSCLE algorithm59 in MEGA760 with default settings (all gap penalties = 0, max iterations = 8, clustering method = UPGMB, lambda = 24). Alignments were visually inspected and trimmed in MEGA7. Aligned sequences were concatenated with an in-house R script provided by MV. Model selection and phylogenetic reconstructions were done on the CIPRES Science Gateway61. Model selection was performed for each alignment partition (6 in total: 18S rRNA, 28S rRNA, ITS-2 and three COI codons) with PartitionFinder262, partitions and models selection process and results are present in Supplementary Data S1. The BI phylogenetic reconstruction was done with MrBayes v3.2.663 without BEAGLE. Four runs with one cold chain and three heated chains were run for 20 million generations with a burning of 2 million generations, sampling a tree every 1000 generations. Posterior distribution sanity was checked with the Tracer v1.764 and with the R package RWTY65. MrBayes input file with the input alignment is available as Supplementary Data S2. ML phylogenetic reconstruction was performed with RAxML-HPC Black Box 8.2.1266 with 1000 bootstrap replicates and estimation of proportion of invariable sites (f = a, N = 1000, m = GTRCATI). The phylogenetic trees were visualised with FigTree v1.4.467 and the image was edited with Inkscape 0.92.368. The complete output trees from BI and ML analysis Supplementary Data S3.
Microscopy and imaging
Specimens for light microscopy were mounted on microscope slides in a small drop of Hoyer’s medium and secured with a cover slip, following the protocol by Morek et al.69. Slides were examined under an Olympus BX53 light microscope with phase and Nomarski differential interference contrasts (PCM and NCM, respectively; named collectively as light contrast microscopy, LCM), associated with an Olympus DP74 digital camera. In order to obtain clean and extended specimens for SEM, tardigrades were processed according to the protocol by Stec et al.49. Specimens were examined under high vacuum in a Versa 3D DualBeam Scanning Electron Microscope (SEM) at the ATOMIN facility of the Jagiellonian University, Kraków, Poland. All figures were assembled in Corel Photo-Paint X6, ver. 16.4.1.1281. For structures that could not be satisfactorily focused in a single LCM photograph, a vertical stack of 2–6 images were taken with an equidistance of ca. 0.2 μm and assembled manually into a single deep-focus image in Corel Photo-Paint.
Morphometrics and nomenclature
All measurements are given in micrometres (μm). Sample size was adjusted following recommendations by Stec et al.70. Structures were measured only if undamaged and their orientation was suitable. Body length was measured from the anterior extremity to the end of the body, excluding the hind legs. The terminology used to describe oral cavity armature and egg shell morphology follows Michalczyk and Kaczmarek71 and Kaczmarek and Michalczyk72 respectively. Macroplacoid length sequence is given according to Kaczmarek et al.73. Buccal tube length and the level of the stylet support insertion point were measured according to Pilato74. The pt index is the ratio of the length of a given structure to the length of the buccal tube expressed as a percentage74. Measurements of buccal tube widths, heights of claws and eggs follow Kaczmarek and Michalczyk72. Morphometric data were handled using the “Parachela” ver. 1.7 template available from the Tardigrada Register75 and are given in Supplementary Data S4. Tardigrade taxonomy follows Bertolani et al.6 with updates from Guidetti et al.11, Vecchi et al.76 and Morek et al.21.
Data availability
All data generated and analysed during this study are included in the article (and its Supplementary Information files). DNA sequences are deposited and available in GenBank.
References
Nelson, D. R., Guidetti, R. & Rebecchi, L. Phylum Tardigrada. In Ecology and General Biology: Vol. 1: Thorp and Covich’s Freshwater Invertebrates 4th edn (eds Thorp, J. H. & Rogers, D. C.) 347–380 (Elsevier, Amsterdam, 2015).
Schultze, C. A. S. Macrobiotus hufelandii animal e crustaceorum classe novum, reviviscendi post diuturnam asphixiam et aridiatem potens, etc. 8, 1 tab. C. Curths, Berlin, 6 pp, I Table (1834).
Guidetti, R. & Bertolani, R. Tardigrade taxonomy: an updated check list of the taxa and a list of characters for their identification. Zootaxa 845, 1–46 (2005).
Degma, P. & Guidetti, R. Notes to the current checklist of Tardigrada. Zootaxa 1579, 41–53 (2007).
Degma, P., Guidetti, R. & Bertolani, R. Actual Checklist of Tardigrada Species. https://iris.unimore.it/handle/11380/1178608#.XvOM-pMzblw (2020). Accessed 10 August 2020.
Bertolani, R. et al. Phylogeny of Eutardigrada: new molecular data and their morphological support lead to the identification of new evolutionary lineages. Mol. Phylogenet. Evol. 76, 110–126 (2014).
Fujimoto, S., Jørgensen, A. & Hansen, J. G. A molecular approach to arthrotardigrade phylogeny (Heterotardigrada, Tardigrada). Zool. Scr. 46, 496–505 (2017).
Gąsiorek, P., Stec, D., Morek, W. & Michalczyk, Ł. Deceptive conservatism of claws: distinct phyletic lineages concealed within Isohypsibioidea (Eutardigrada) revealed by molecular and morphological evidence. Contrib. Zool. 88(1), 78–132 (2019).
Gąsiorek, P., Morek, W., Stec, D. & Michalczyk, Ł. Untangling the Echiniscus Gordian knot: paraphyly of the “arctomys group” (Heterotardigrada: Echiniscidae). Cladistics 35(6), 633–653 (2019).
Gąsiorek, P. & Michalczyk, Ł. Phylogeny of Itaquasconinae in light of the evolution of the flexible pharyngeal tube in Tardigrada. Zool. Scr. 49(4), 499–515 (2020).
Guidetti, R. et al. Morphological and molecular analyses on Richtersius (Eutardigrada) diversity reveal its new systematic position and lead to the establishment of a new genus and a new family within Macrobiotoidea. Zool. J. Linn. Soc. 178(4), 834–845 (2016).
Maucci, W. & Ramazzotti, G. Adorybiotus gen. nov.: nouva posizione sistematica per Macrobiotus granulatus Richters, 1903 e per Macrobiotus coronifer Richters, 1903 (Tardigrada, Macrobiotidae). Mem. Ist. Ital. Idrobiol. 39, 153–159 (1981).
Lisi, O., Londoño, R. & Quiroga, S. Description of a new genus and species (Eutardigrada: Richtersiidae) from Colombia, with comments on the family Richtersiidae. Zootaxa 4822(4), 531–550 (2020).
Pilato, G. & Binda, M. G. Richtersius, nuove nome generico in sostituzione di Richtersia Pilato e Binda 1987 (Eutardigrada). Animalia 16, 147–148 (1989).
Thulin, G. Über die Phylogenie und das system der Tardigraden. Hereditas 11, 207–266 (1928).
Guil, N., Jørgensen, A. & Kristensen, R. M. An upgraded comprehensive multilocus phylogeny of the Tardigrada tree of life. Zool. Scr. 48(1), 120–137 (2019).
Richters, F. Arktische Tardigraden. Fauna Arct. 3, 494–508 (1904).
Doyère, P. L. N. Memoire sur les Tardigrades. Ann. Sci. Nat. Ser. 2 (Zool.) 14, 269–362 (1840).
Richters, F. Tardigrada. In Handbuch der Zoologie 3 (eds Kükenthal, W. & Krumbach, T.) 58–61 (Walter de Gruyter & Co., Berlin, 1926).
Schuster, R. O., Nelson, D. R., Grigarick, A. A. & Christenberry, D. Systematic criteria of the Eutardigrada. Trans. Am. Microsc. Soc. 99, 284–303 (1980).
Morek, W., Ciosek, J. & Michalczyk, Ł. Description of Milnesium pentapapillatum sp. nov., with an amendment of the diagnosis of the order Apochela and abolition of the class Apotardigrada (Tardigrada). Zool. Anz. 288, 107–117 (2020).
Guidetti, R., Gandolfi, A., Rossi, V. & Bertolani, R. Phylogenetic analysis of Macrobiotidae (Eutardigrada, Parachela): a combined morphological and molecular approach. Zool. Scr. 34, 235–244 (2005).
Binda, M. G. Tardigrada della Valtellina. Animalia 1, 201–216 (1974).
Guidetti, R., Cesari, M., Bertolani, R., Altiero, T. & Rebecchi, L. 2019) High diversity in species, reproductive modes and distribution within the Paramacrobiotus richtersi complex (Eutardigrada, Macrobiotidae. Zool. Lett. 5, 1 (2019).
Stec, D., Morek, W., Gąsiorek, P. & Michalczyk, Ł. Unmasking hidden species diversity within the Ramazzottius oberhaeuseri complex, with an integrative redescription of the nominal species for the family Ramazzottiidae (Tardigrada: Eutardigrada: Parachela). Syst. Biodivers. 16(4), 357–376 (2018).
Morek, W. & Michalczyk, Ł. First extensive multilocus phylogeny of the genus Milnesium Doyère, 1840 (Tardigrada) reveals no congruence between genetic markers and morphological traits. Zool. J. Linn. Soc. 188(3), 681–693 (2020).
Gąsiorek, P., Stec, D., Morek, W. & Michalczyk, Ł. An integrative redescription of Echiniscus testudo (Doyère, 1840), the nominal taxon for the class Heterotardigrada (Ecdysozoa: Panarthropoda: Tardigrada). Zool. Anz. 270, 107–122 (2017).
Gąsiorek, P., Stec, D., Morek, W. & Michalczyk, Ł. An integrative redescription of Hypsibius dujardini (Doyère, 1840), the nominal taxon for Hypsibioidea (Tardigrada: Eutardigrada). Zootaxa 4415(1), 45–75 (2018).
Grobys, D. et al. High diversity in the Pseudechiniscus suillus-facettalis complex (Heterotardigrada; Echiniscidae) with remarks on the morphology of the genus Pseudechiniscus. Zool. J. Linn. Soc. 188(3), 733–752 (2020).
Stec, D., Krzywański, Ł, Zawierucha, K. & Michalczyk, Ł. Untangling systematics of the Paramacrobiotus areolatus species complex by an integrative redescription of the nominal species for the group, with multilocus phylogeny and species delineation within the genus Paramacrobiotus. Zool. J. Linn. Soc. 188(3), 694–716 (2020).
Stec, D., Krzywański, Ł, Arakawa, K. & Michalczyk, Ł. A new redescription of Richtersius coronifer, supported by transcriptome, provides resources for describing concealed species diversity within the monotypic genus Richtersius (Eutardigrada). Zool. Lett. 6, 2 (2020).
Richters, F. Nordische Tardigraden. Zool. Anz. 27, 168–172 (1903).
Stec, D. & Michalczyk, Ł. Macrobiotus coronifer Richters, 1903 (type species for Richtersius Pilato & Binda, 1989): designating a new neotype from the original type locality described within the integrative taxonomy framework. Zootaxa 4858(2), 292–294 (2020).
Zawierucha, K. Tardigrada from Arctic tundra (Spitsbergen) with description of Isohypsibius karenae sp. n. (Isohypsibiidae). Pol. Polar Res. 34(4), 383–396 (2013).
Zawierucha, K., Coulson, S., Michalczyk, Ł & Kaczmarek, Ł. Current knowledge of the Tardigrada of Svalbard with the first records of water bears from Nordaustlandet (High Arctic). Polar Res. 32, 20886 (2013).
Zawierucha, K., Smykla, J., Michalczyk, Ł, Gołdyn, B. & Kaczmarek, Ł. Distribution and diversity of Tardigrada along altitudinal gradients in the Hornsund, West Spitsbergen (Arctic). Polar Res. 34, 24168 (2015).
Zawierucha, K., Zmudczyńska-Skarbek, K., Kaczmarek, Ł & Wojczulanis-Jakubas, K. The influence of a seabird colony on abundance and species composition of water bears (Tardigrada) in Hornsund (Spitsbergen, Arctic). Polar Biol. 39(4), 713–723 (2016).
Zawierucha, K., Węgrzyn, M., Ostrowska, M. & Wietrzyk, P. Tardigrada in Svalbard lichens: diversity, densities and habitat heterogeneity. Polar Biol. 40(7), 1385–1392 (2017).
Ramazzotti, G. & Maucci, W. Il Phylum Tardigrada. Mem. dell’Istituto Ital. Idrobiol. Pallanza 41, 1–1012 (1983).
Binda, M. G. Ridescrizione di Macrobiotus echinogenitus Richters, 1904 e sul valore di buona specie di Macrobiotus crenulatus Richters, 1904 (Eutardigrada). Animalia 15(1/3), 201–210 (1988).
ICZN. International Code of Zoological Nomenclature 4th edn. (The International Trust for Zoological Nomenclature, London, 1999).
ICZN. International Code of Zoological Nomenclature 2nd edn. (The International Trust for Zoological Nomenclature, London, 1964).
Richters, F. Isländische Tardigraden. Zool. Anz. 28, 373–377 (1904).
Séméria, Y. New tardigrades from Nicaragua. C. R. Acad. Sci. Paris 300 Sér. 3 1, 9–11 (1985).
McInnes, S. J. Zoogeographic distribution of terrestrial /freshwater tardigrades from current literature. J. Nat. Hist. 28, 257–352 (1994).
Kaczmarek, Ł, Michalczyk, Ł & McInnes, S. J. Annotated zoogeography of non-marine Tardigrada. Part II: South America. Zootaxa 3923(1), 1–107 (2015).
Kaczmarek, Ł, Michalczyk, Ł & McInnes, S. J. Annotated zoogeography of non-marine Tardigrada. Part III: North America and Greenland. Zootaxa 4203(1), 1–249 (2016).
McInnes, S. J., Michalczyk, Ł & Kaczmarek, Ł. Annotated zoogeography of non-marine Tardigrada Part IV: Africa. Zootaxa 4284(1), 1–74 (2017).
Stec, D., Smolak, R., Kaczmarek, Ł & Michalczyk, Ł. An integrative description of Macrobiotus paulinae sp. nov. (Tardigrada: Eutardigrada: Macrobiotidae: hufelandi group) from Kenya. Zootaxa 4052(5), 501–526 (2015).
Casquet, J., Thebaud, C. & Gillespie, R. G. Chelex without boiling, a rapid and easy technique to obtain stable amplifiable DNA from small amounts of ethanol-stored spiders. Mol. Ecol. Resour. 12, 136–141 (2012).
Stec, D., Kristensen, R. M. & Michalczyk, Ł. An integrative description of Minibiotus ioculator sp. nov. from the Republic of South Africa with notes on Minibiotus pentannulatus Londoño et al., 2017 (Tardigrada: Macrobiotidae). Zool. Anz. 286, 117–134 (2020).
Hall, T. A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl. Acids Symp. Ser. 41, 95–98 (1999).
Durante Pasa, M. V. & Maucci, W. Descrizione di tre nuove specie di Tardigradi della Scandinavia. Atti. Soc. Ital. Sci. Nat. Museo Civ. St. Nat. Milano 116(3–4), 244–250 (1975).
Kristensen, R. M. New abbarant Eutardigrades from homothermic springs on Disko Island, West Greenland. In Nelson D.R. ed, Proceedings of the Third International Symposium on Tardigrada 203–220 (East Tennessee State University Press, Johnson City, Tennessee, 1982).
Biserov, V. I. Hypsibius subanomalus sp. n. (Eutardigrada, Hypsibiidae) from the Astrakhan District. Zool. Zhurnal. 64, 131–135 (1985).
Kihm, J. H. et al. Integrative description of a new Dactylobiotus (Eutardigrada: Parachela) from Antarctica that reveals an intraspecific variation in tardigrade egg morphology. Sci. Rep. 10, 9122 (2020).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucl. Acids Res. 30, 3059–3066 (2002).
Katoh, K. & Toh, H. Recent developments in the MAFFT multiple sequence alignment program. Brief. Bioinform. 9, 286–298 (2008).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucl. Acids Res. 32(5), 1792–1797 (2004).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Miller, M. A., Pfeiffer, W. & Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In 2010 gateway computing environments workshop (GCE), 1–8 (2010).
Lanfear, R., Frandsen, P. B., Wright, A. M., Senfeld, T. & Calcott, B. PartitionFinder 2: new methods for selecting partitioned models of evolution formolecular and morphological phylogenetic analyses. Mol. Biol. Evol. 34(3), 772–773 (2016).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61(3), 539–542 (2012).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst. Biol. 67(5), 901–904 (2018).
Warren, D. L., Geneva, A. J. & Lanfear, R. RWTY (R We There Yet): an R package for examining convergence of Bayesian phylogenetic analyses. Mol. Biol. Evol. 34(4), 1016–1020 (2017).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9), 1312–1313 (2014).
Rambaut, A. FigTree, A Graphical Viewer of Phylogenetic Trees. (2007)
Bah, T. Inkscape: Guide to a Vector Drawing Program, 559 (Prentice Hall, Upper Saddle River, 2011).
Morek, W. et al. An experimental test of eutardigrade preparation methods for light microscopy. Zool. J. Linn. Soc. 178(4), 785–793 (2016).
Stec, D. et al. Estimating optimal sample size for tardigrade morphometry. Zool. J. Linn. Soc. 178(4), 776–784 (2016).
Michalczyk, Ł & Kaczmarek, Ł. A description of the new tardigrade Macrobiotus reinhardti (Eutardigrada, Macrobiotidae, harmsworthi group) with some remarks on the oral cavity armature within the genus Macrobiotus Schultze. Zootaxa 331, 1–24 (2003).
Kaczmarek, Ł & Michalczyk, Ł. The Macrobiotus hufelandi (Tardigrada) group revisited. Zootaxa 4363(1), 101–123 (2017).
Kaczmarek, Ł, Cytan, J., Zawierucha, K., Diduszko, D. & Michalczyk, Ł. Tardigrades from Peru (South America), with descriptions of three new species of Parachela. Zootaxa 3790(2), 357–379 (2014).
Pilato, G. Analisi di nuovi caratteri nello studio degli Eutardigradi. Animalia 8, 51–57 (1981).
Michalczyk, Ł & Kaczmarek, Ł. The Tardigrada Register: a comprehensive online data repository for tardigrade taxonomy. J. Limnol. 72, 175–181 (2013).
Vecchi, M. et al. Integrative systematic studies on tardigrades from Antarctica identify new genera and new species within Macrobiotoidea and Echiniscoidea. Invertebr. Syst. 30(4), 303–322 (2016).
Stec, D., Zawierucha, K. & Michalczyk, Ł. An integrative description of Ramazzottius subanomalus (Biserov, 1985) (Tardigrada) from Poland. Zootaxa 4300(3), 403–420 (2017).
Gąsiorek, P., Stec, D., Zawierucha, Z., Kristensen, R. M. & Michalczyk, Ł. Revision of Testechiniscus Kristensen, 1987 (Heterotardigrada: Echiniscidae) refutes the polar-temperate distribution of the genus. Zootaxa 4472(2), 261–297 (2018).
Mironov, S. V., Dabert, J. & Dabert, M. A new feather mite species of the genus Proctophyllodes Robin, 1877 (Astigmata: Proctophyllodidae) from the Long-tailed Tit Aegithalos caudatus (Passeriformes: Aegithalidae): morphological description with DNA barcode data. Zootaxa 3253, 54–61 (2012).
Astrin, J. J. & Stüben, P. E. Phylogeny in cryptic weevils: molecules, morphology and new genera of western Palaearctic Cryptorhynchinae (Coleoptera : Curculionidae). Invertebr. Syst. 22(5), 503–522 (2008).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 3, 294–299 (1994).
Prendini, L., Weygoldt, P. & Wheeler, W. C. Systematics of the Damon variegatus group of African whip spiders (Chelicerata: Amblypygi): evidence from behaviour, morphology and DNA. Org. Divers. Evol. 5, 203–236 (2005).
Zawierucha, K., Kolicka, M. & Kaczmarek, Ł. Re-description of the Arctic tardigrade Tenuibiotus voronkovi (Tumanov, 2007) (Eutardigrada; Macrobiotidae), with the first molecular data for the genus. Zootaxa 4196(4), 498–510 (2016).
Stec, D., Tumanov, D. T. & Kristensen, R. M. Integrative taxonomy identifies two new tardigrade species (Eutardigrada: Macrobiotidae) from Greenland. Eur. J. Taxon. 614, 1–40 (2020).
Stec, D., Arakawa, K. & Michalczyk, Ł. An integrative description of Macrobiotus shonaicus sp. nov. (Tardigrada: Macrobiotidae) from Japan with notes on its phylogenetic position within the hufelandi group. PLoS ONE 13(2), e0192210 (2018).
Coughlan, K., Michalczyk, Ł & Stec, D. Macrobiotus caelestis sp. nov., a new tardigrade species (Macrobiotidae: hufelandi group) from the Tien Shan mountains (Kyrgyzstan). Annal. Zool. 69(3), 499–513 (2019).
Kaczmarek, Ł et al. An integrative redescription of the nominal taxon for the Mesobiotus harmsworthi group (Tardigrada: Macrobiotidae) leads to descriptions of two new Mesobiotus species from Arctic. PLoS ONE 13(10), e0204756 (2018).
Itang, L. A. M., Stec, D., Mapalo, M. A., Mirano-Bascos, D. & Michalczyk, Ł. An integrative description of Mesobiotus dilimanensis, a new tardigrade species from the Philippines (Eutardigrada: Macrobiotidae: furciger group). Raffles Bull. Zool. 68, 19–31 (2020).
Guil, N. & Giribet, G. A comprehensive molecular phylogeny of tardigrades - adding genes and taxa to a poorly resolved phylum-level phylogeny. Cladistics 28(1), 21–49 (2012).
Pogwizd, J. & Stec, D. New records of Dactylobiotus parthenogeneticus Bertolani, 1982 provide insight into its genetic variability and geographic distribution. Folia Biol. (Kraków) 68(2), 57–72 (2020).
Acknowledgements
We are especially grateful to our colleagues Atsushi Suzuki, Brian Blagden, Artur Oczkowski, Piotr Gąsiorek, Wojciech Witaliński, Michala Bryndová and Claudio Ferrari for collecting and sending samples which enable us to conduct this study. The Jagiellonian University expedition in 2016 constituted the main party in the project “Southeastern Spitsbergen landscape-seascape and biodiversity dynamics under current climate warming”, which was funded by the Prince Albert II of Monaco Foundation (www.fpa2.org). However, the published results of this expedition are the sole responsibility of Jagiellonian University and under no circumstances can be regarded as reflecting the position of the Prince Albert II of the Monaco Foundation. Sample IT.338 was collected under sampling permit Protocol N.1128 by Parco Nazionale Appennino Tosco-Emiliano (Italy) issued to MV. The sampling in Celebes was supported by the Polish Ministry of Science and Higher Education via the Diamond Grant (DI2015 014945 to Piotr Gąsiorek, supervised by ŁM). We are also grateful to anonymous reviewers who improved the manuscript. The study was supported by the Preludium programme of the Polish National Science Centre (Grant No. 2018/31/N/NZ8/03096 to DS supervised by ŁM) and partially supported by supported by the Jagiellonian University (subsidy no. DS/D/WB/IZiBB/16/2019 awarded to DS). During this study, DS was a beneficiary of a National Science Centre scholarship to support doctoral research (No. 2019/32/T/NZ8/00348). Some of the analyses were carried out with the equipment purchased from the Sonata Bis programme of the Polish National Science Centre (Grant No. 2016/22/E/NZ8/00417 to ŁM). Open-access publication of this article was funded by the BioS Priority Research Area under the program “Excellence Initiative – Research University” at the Jagiellonian University in Kraków, Poland.
Author information
Authors and Affiliations
Contributions
D.S.: conceived the study, analysed the samples, performed the molecular and morphological analyses, assembled morphological figures, drafted the first version of the manuscript and partially funded the research; M.V.: conceived the study, performed phylogenetic analysis, prepared the phylogenetic figure, drafted the first version of the manuscript; W.M.: collected samples and drafted the first version of the manuscript; Ł.M.: conceived the study, supervised the research, prepared the phylogenetic figure, drafted the first version of the manuscript and partially funded the research. All authors contributed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Stec, D., Vecchi, M., Maciejowski, W. et al. Resolving the systematics of Richtersiidae by multilocus phylogeny and an integrative redescription of the nominal species for the genus Crenubiotus (Tardigrada). Sci Rep 10, 19418 (2020). https://doi.org/10.1038/s41598-020-75962-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-75962-1
- Springer Nature Limited
This article is cited by
-
The tardigrade Mesobiotus aradasi (Binda, Pilato & Lisi, 2005) is widely distributed along the Antarctic Peninsula
Polar Biology (2024)
-
Integrative taxonomy helps to revise systematics and questions the purported cosmopolitan nature of the type species within the genus Diaforobiotus (Eutardigrada: Richtersiusidae)
Organisms Diversity & Evolution (2023)
-
Description of a new species of Tardigrada Hypsibius nivalis sp. nov. and new phylogenetic line in Hypsibiidae from snow ecosystem in Japan
Scientific Reports (2022)