Abstract
Repulsive guidance molecules (RGMs) are evolutionarily conserved proteins implicated in repulsive axon guidance. Here we report the function of the Caenorhabditis elegans ortholog DRAG-1 in axon branching. The axons of hermaphrodite-specific neurons (HSNs) extend dorsal branches at the region abutting the vulval muscles. The drag-1 mutants exhibited defects in HSN axon branching in addition to a small body size phenotype. DRAG-1 expression in the hypodermal cells was required for the branching of the axons. Although DRAG-1 is normally expressed in the ventral hypodermis excepting the vulval region, its ectopic expression in vulval precursor cells was sufficient to induce the branching. The C-terminal glycosylphosphatidylinositol anchor of DRAG-1 was important for its function, suggesting that DRAG-1 should be anchored to the cell surface. Genetic analyses suggested that the membrane receptor UNC-40 acts in the same pathway with DRAG-1 in HSN branching. We propose that DRAG-1 expressed in the ventral hypodermis signals via the UNC-40 receptor expressed in HSNs to elicit branching activity of HSN axons.
Similar content being viewed by others
Introduction
Axon branching is a fundamental process for proper axon projection to target tissues and for the formation of correct synapses, both of which are important for development of the functional neural network. Axon branching begins with the formation of an actin-rich filopodium from the existing axon followed by extension of microtubules along the actin filaments1. Formation of filopodia and subsequent neurite extension involve various regulators for actin and microtubule polymerization and bundling. The location and the polarity of axon branching are dictated by extracellular cues, along with cytoskeletal activities. Axon guidance molecules are involved in this process.
Netrin-1 is a secreted guidance molecule that induces local filopodial protrusions in the axon shaft, which give rise to branches in the cortical neurons. In contrast, SEMA3A represses cortical axon branching2. Ephrins are membrane-bound molecules that abolish branching of thalamic axons3. In addition to these well-known guidance molecules, repulsive guidance molecules (RGMs) also repress axon branching in cortical neurons and mossy fibers of the hippocampus4,5. In vertebrates, RGMs are glycosylphosphatidylinositol (GPI)-linked membrane proteins that constitute a family with four members: RGMa, RGMb (DRAGON), RGMc (hemojuvelin), and RGMd6. RGMa was first discovered as an axon guidance cue that has a repulsive activity to retinal axons. RGMa is expressed in the embryonic tectum in an anterior-to-posterior concentration gradient and functions during the development of the retinotectal projection7.
By binding to the membrane receptor neogenin, RGMs function in axon guidance and neuronal survival as well as inhibition of axonal regeneration8,9,10. RGMs also bind bone morphogenetic proteins (BMPs) in the regulation of iron homeostasis and endochondral bone development11,12,13. Although the function of RGMs in axon guidance as a result of growth cone repulsion has been well studied, their role in axon branching is still elusive. Because of the lethality of knock-out mice and the functional redundancy of RGM proteins, the functions of RGMs have been mostly analyzed using in vitro culture systems14.
The nematode C. elegans has a single ortholog of RGM, DRAG-1. Loss-of-function mutations in drag-1 result in a small body size as well as genetic suppression of the mesodermal coelomocyte loss phenotype of sma-9 mutants15. drag-1 function in the hypodermal cells is required for the control of body size, while drag-1 function in the mesodermal M cell lineage controls coelomocyte differentiation15,16. In the present study, we showed that drag-1 expressed in the hypodermis functions in the formation of branches of hermaphrodite-specific neurons (HSNs), which innervate egg-laying muscles of the vulva17. Small body size mutants sma-1 and sma-5 also exhibited HSN axon branching defects although they act in genetic pathways distinct from that of drag-1. Genetic analyses suggested that unc-40 acts in the same pathway as drag-1. Because DRAG-1 binds the receptor UNC-4016 and UNC-40 is expressed in HSNs18, our findings suggest that DRAG-1 acts on the receptor UNC-40 to induce axon branching of HSNs.
Results
drag-1 mutants are defective in axon branching of HSNs
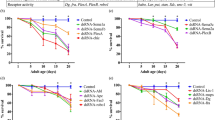
drag-1 encodes the sole C. elegans ortholog of the RGM family of proteins. We isolated a deletion allele of drag-1, tk81 (Fig. 1a). drag-1(tk81) animals had a smaller body size compared to wild type. The drag-1(tm3773) deletion mutant obtained from the National Bioresource Project showed a similar phenotype (Fig. 1a–c, Supplementary Fig. S1a,b). Because RGM family proteins act in axon guidance in mammals, we examined axonal morphology using a pan-neuronal GFP reporter ncIs219. We found no gross defects in axon trajectories in the mutants compared with wild type. We then examined the morphology of the axon of hermaphrodite specific neurons (HSNs) using kyIs262[unc-86p::myrGFP; odr-1::RFP]20 as a transgenic marker. In wild type animals, the cell bodies of the bilateral HSNs are positioned slightly posterior to the vulva. They extend a single axon toward the ventral nerve cord. After reaching the nerve cord, the axon is redirected dorsally and anteriorly and reaches the lateral position of the developing vulva, where it turns again, this time in a ventral and anterior direction toward the ventral nerve cord21. The projection of HSN axons was not affected in the drag-1 mutants (Supplementary Table S1). Although the axon usually sprouts one or two dorsal branches at the vulva in the wild type, the number of axons with branches was significantly reduced in the drag-1 mutants (Fig. 2a,b). To examine whether the branching defect is caused by loss of drag-1 function, we introduced a plasmid containing a fragment of the wild-type gene (drag-1p::drag-1) into the drag-1 mutants as a transgene. The transgene fully rescued the branching defect (Fig. 2b), confirming the function of DRAG-1 in HSN axon branching. Because drag-1 mutants did not accumulate fertilized eggs in the uterus, they were not egg-laying defective.
Gene structure and mutant phenotype of drag-1 animals. (a) Structure of drag-1 and mutation sites of the tk81 and tm3773 alleles. The exon and intron regions were determined by sequencing cDNA generated from isolated drag-1 mRNA. Black, yellow, blue, and magenta boxes indicate N-terminal signal peptide, partial von Willebrand factor type D domain, hydrophobic region, and C-terminal GPI-anchor signal sequence, respectively15. Bars depict the region of the cDNA used for expressing the antigenic peptide for producing antibodies and the respective mutation sites. tk81 is a 494-bp deletion within exon 3, which is expected to produce a truncated polypeptide that is missing the C-terminal 278 amino acids. tm3773 is an 892-bp deletion spanning from intron 2 to exon 3 (WormBase). (b) Body length phenotype of drag-1(tk81) mutants. Body length of young adult hermaphrodites. tk81 mutants had shorter bodies compared with wild type. Anterior is to the left. Scale bar: 50 μm. (c) Quantification of body length of young adult hermaphrodites for wild-type and drag-1 mutant animals. Significant difference was determined by Student’s t-test. ***P < 0.001. n = 60 for each strain.
HSN branching phenotypes. (a) HSN axon branching. Upper panels: Confocal micrographs of HSNs in wild-type and drag-1 mutant young-adult hermaphrodites with the unc-86p::myrGFP transgene. Lower panels: Schematic representations of HSN morphology. Yellow and red arrowheads depict the HSN cell body and axonal branches, respectively. Anterior is to the left, dorsal top. Scale bar: 10 μm. (b) Quantification of the HSN branching phenotypes in various strains. Percentages of HSNs having no branches are shown. Branching phenotypes were scored using fluorescence microscopy. drag-1(tk81) and drag-1(tm3773) mutants were compared with those transgenic for drag-1p::drag-1. Significant differences were determined by Fisher’s exact test. ***P < 0.001, **P < 0.01. n = 180 for each strain.
drag-1 acts in parallel pathways with sma-1 and sma-5
Because drag-1 mutants result in a small body size (Sma) phenotype, we examined whether other sma mutants affect HSN axon branching. Among the eight sma mutants examined, sma-2(e502), sma-3(wk28), sma-4(e729), sma-6(wk7), and sma-9(wk55) did not show HSN branching defects (Fig. 3a). We found HSN branching defects similar to that observed in the drag-1 mutants in sma-1(e30), sma-5(n678), and sma-8(e2111) (Fig. 3b). sma-1 and sma-5 encode βH-spectrin and MAP kinase, respectively22,23. sma-8(e2111) is a dominant mutation for which the causative gene has not yet been identified. We produced double mutants between drag-1 mutants and these sma mutants and found that all double mutants exhibited HSN branching defects stronger than those observed in the respective single mutants (Fig. 3b). Because drag-1(tk81) and drag-1(tm3773) mutants are putative null alleles, these results suggested that sma-1 and sma-5 act in pathways different from that of drag-1 to regulate HSN branching. Because of the dominancy of the sma-8(e2111) mutation, the relationship between drag-1 and sma-8 remains to be determined.
Genetic interactions between drag-1 mutants and sma mutants. Percentages of HSNs having no branches are shown. (a) sma-2(e502), sma-3(wk28), sma-4(e729), sma-6(wk7), and sma-9(wk55) mutants were compared with wild type. (b) drag-1(tk81) and drag-1(tm3773) mutants were compared with sma-1(e30), sma-5(n678), and sma-8(e2111) mutants and with double mutants consisting of drag-1(tk81) or drag-1(tm3773) in combination with individual sma mutations. Significant differences were determined by Fisher’s exact test. ***P < 0.001, **P < 0.01, *P < 0.05. NS not significant. n = 180 for each strain.
GPI anchoring of DRAG-1 is important for HSN branching
Because DRAG-1 is thought to be modified by a GPI-anchor, we placed the GFP or Venus coding region right upstream of the putative cleavage site of the C-terminal pro-peptide sequence (drag-1p::drag-1::GFP::GPI or drag-1p::drag-1::Venus::GPI) to keep the GPI-anchor signal intact. These constructs rescued the branching defect of the mutants (Fig. 4a,b). We examined whether the GPI anchoring is important for DRAG-1 to act in axon branching. A construct with the deleted C-terminal GPI-anchor signal sequence (drag-1p::drag-1::GFP::∆GPI) failed to rescue the branching defects of drag-1 mutants (Fig. 4a,b). We also examined a construct in which the C-terminal GPI-anchor signal was replaced by the transmembrane domain of the LIN-12/Notch receptor (drag-1p::drag-1::GFP::lin-12TM). This chimeric protein, which is potentially localized to the plasma membrane, failed to rescue the mutant defects (Fig. 4a,b). These results suggested that DRAG-1 needs to be anchored to the plasma membrane by the GPI anchor rather than being embedded as a transmembrane protein.
Rescue experiments of drag-1 mutants with modified DRAG-1 proteins. (a) Schematic representation of the GFP fusion constructs. The GFP or Venus coding sequence was inserted between amino acid (aa) 395 and 396 of the drag-1 coding region, just prior to the cleavage site of the C-terminal pro-peptide for drag-1p::drag-1::GFP::GPI. The C-terminal GPI-anchor signal (aa 387–408) was deleted from drag-1p::drag-1::GFP::GPI for drag-1p::drag-1::GFP::∆GPI. The lin-12 transmembrane domain (aa 907–934) (shown in purple) was connected with drag-1p::drag-1::GFP::∆GPI for drag-1p::drag-1::GFP::lin-12TM15. (b) Percentages of HSNs having no branches are shown for the drag-1(tk81) mutants expressing the different rescuing constructs shown in (a). Data for drag-1p::drag-1::Venus::GPI is also shown. Significant differences were determined by Fisher’s exact test. *P < 0.05. NS not significant. n = 180, 107, 105, 91, and 109 for drag-1p::drag-1::Venus::GPI, drag-1p::drag-1::GFP::GPI, drag-1p::drag-1::GFP::∆GPI #1, drag-1p::drag-1::GFP::∆GPI #2, and drag-1p::drag-1::GFP::lin-12TM, respectively. The #1 and #2 refer to two independently isolated transgenic lines.
Expression of DRAG-1
To understand how DRAG-1 functions, we examined its expression pattern using both transcriptional and translational reporters, as well as immunostaining using anti-DRAG-1 antibodies that we generated. Expression of a transcriptional drag-1p::Venus reporter was detected in the pharynx, intestine and the syncytial hypodermis from late embryos to adult stages. Notably, Venus expression was not detected in hypodermal seam cells and the vulval hypodermis (Fig. 5a). Unlike the transcriptional reporter described above, we could only detect Venus expression in the pharynx using the functional translational fusion construct drag-1p::drag-1::Venus::GPI (Fig. 5b). We therefore raised polyclonal antibodies against a DRAG-1 peptide corresponding to amino acids 130–368 (Fig. 1a). Immunostaining experiments were performed using the L4 to young adult stage animals in which HSN branches are formed24. The antibodies detected no signals in non-transgenic wild-type animals, but detected signals in the pharynx, intestinal cells, hypodermal seam cells, and ventral hypodermal cells (except in the vulval region) in animals transgenic for either drag-1p::drag-1 or drag-1p::drag-1::Venus::GPI. (Fig. 5c). The expression patterns were consistent with the observation using anti-GFP in animals expressing drag-1p::drag-1::GFP::GPI15. Thus, it is likely that the level of expression of endogenous DRAG-1 is low. Alternatively, DRAG-1 expression might be upregulated only transiently in HSN branch formation. The hypodermal signals detected by the anti-DRAG-1 antibodies appeared in a granular pattern in the cytoplasm. This could be due to the accumulation of DRAG-1 in the ER or Golgi apparatus as a result of over-expression of drag-1. We detected no DRAG-1 expression in the nervous system including HSNs even in the over-expressed condition.
drag-1 expression. (a) drag-1p::Venus expression. Expression was detected from late embryos to the adult stage in the pharynx, intestine, and hypodermis. White arrows, dotted arrows and white arrowheads correspond to the pharynx, intestine and hypodermis, respectively. Yellow arrowheads and asterisk point to seam cells and developing vulval epithelium which do not express Venus, respectively. Scale bar: 100 μm. (b) Expression of drag-1p::drag-1::Venus::GPI. The drag-1p::drag-1::Venus::GPI plasmid was injected into unc-119(e2498) animals at 150 ng/μl with 30 ng/μl of pBSII KS(–) and 20 ng/μl pDP#MM016B. DIC (upper) and fluorescence (lower) images of an L4 stage animal are shown. Venus expression was detected only in the pharynx. Anterior is to the left. Scale bar: 100 μm. (c) Immunostaining using anti-DRAG-1. L4 to young-adult animals expressing drag-1p::drag-1::Venus::GPI were fixed and stained with anti-DRAG-1. DRAG-1 expression was detected in the pharynx, intestine, hypodermal seam cells, and ventral hypodermal cells (arrows) except the vulval hypodermis (asterisk). Scale bar: 50 μm.
DRAG-1 functions in hypodermal cells for axon branching
We have shown that drag-1 is important for axon branching, but drag-1 is not expressed in the nervous system. To determine the tissues in which DRAG-1 expression is important for axon branching, we expressed drag-1 under tissue-specific promoters. We found that hypodermal expression of DRAG-1 using the rol-6 promoter (rol-6p::drag-1)25 rescued the branching defect, whereas expression in the pharyngeal muscle (myo-2p::drag-1)26 or in the intestine (elt-2p::drag-1)27 did not (Fig. 6). These results indicated that DRAG-1 functions in hypodermal cells to induce axon branching of the HSNs. Because we detected DRAG-1 expression in both ventral hypodermis and lateral hypodermal seam cells, we tested whether the expression in the seam cells could rescue the HSN branching defect of drag-1 mutants. drag-1 expression under the seam cell specific SCM promoter28 failed to rescue the HSN branching defect (Fig. 6). We also attempted to specifically express drag-1 in the ventral hypodermis, but we failed to find an appropriate promoter for this purpose. To circumvent this problem, we drove the expression of drag-1 under the dab-1 promoter, which drives gene expression in the developing vulval precursor cells29. Interestingly, the dab-1p::drag-1 construct successfully rescued the branching defect (Fig. 6). Thus even though drag-1 is not normally expressed in the vulval precursor cells, forced expression of drag-1 in these cells is sufficient for HSN branching, again supporting a role of DRAG-1 functioning in hypodermal cells for axon branching.
Tissue-specific rescue experiments of drag-1 mutants. Percentages of HSNs having no branches are shown. drag-1(tk81) and drag-1(tm3773) mutants were compared with those transgenic for rol-6p::drag-1, myo-2p::drag-1, elt-2p::drag-1, SCMp::drag-1 and dab-1p::drag-1. Two independently isolated transgenic lines (#1 and #2) were examined for each transgene. Significant differences were determined by Fisher’s exact test relative to the corresponding drag-1 mutants. **P < 0.01, *P < 0.05. NS not significant. n = 180 for each strain.
drag-1 acts in the same pathway as unc-40 in HSN axon branching
Neogenin is a receptor for RGMa for axonal growth cone guidance8,30 and UNC-40 is the sole ortholog of neogenin in C. elegans. We examined the HSN axons of unc-40(e271) null allele. The axons of the unc-40 mutants mostly failed to extend to the ventral side and the branches were rarely observed at the vulva or other locations (Supplementary Fig. S2, Table S1). However, we observed branching defects in unc-40(e271)/+heterozygotes with similar penetrance as in the drag-1 mutants. unc-40(e271)/+heterozygotes exhibited no abnormality in HSN axon trajectory (Supplementary Table S1). The branching defect in unc-40(e271)/+was not enhanced when combined with drag-1 null mutants (Fig. 7). Therefore, UNC-40 acts in the same genetic pathway as DRAG-1 in HSN axon branching.
drag-1 does not enhance unc-40/+with respect to HSN branching defects. Percentages of HSNs having no branches are shown. drag-1(tk81) and drag-1(tm3773) mutants were compared with unc-40(e271)/+heterozygotes and with drag-1(tk81) unc-40(e271)/+and drag-1(tm3773) unc-40(e271)/+double mutants. Significant differences were determined by Fisher’s exact test. **P < 0.01, *P < 0.05. NS not significant. n = 90 for each strain.
Discussion
In this study, we have shown that the sole C. elegans repulsive guidance molecule DRAG-1 functions in hypodermal cells to regulate axon branching of the HSNs that form synapses with the vulval egg-laying muscles in hermaphrodites. We further showed that DRAG-1 functions in the same genetic pathway as the neogenin homolog UNC-40 in regulating axon branching.
DRAG-1 has been previously shown to function in the BMP signaling pathway to regulate body size and mesoderm development15. sma-6 (BMP type I receptor); sma-2, -3, and -4 (Smads); and sma-9 (BMP antagonist schnurri) are components of the BMP signaling pathway in C. elegans. Mutations in these genes result in a small body size (Sma) phenotype. However, none of the sma mutants in the BMP pathway affected HSN axon branching. Instead, we observed HSN axon branching defects in sma-1(e30) and sma-5(n678) mutants, which are not involved in the BMP signaling. These results suggest that BMP signaling does not regulate HSN axon branching and that the function of DRAG-1 in regulating HSN axon branching is independent of BMP signaling.
Unlike BMP signaling, two genes known to function in regulating body size, sma-1 and sma-5, also play a role in regulating HSN axon branching. We have shown that SMA-1 and SMA-5 appear to act in pathways parallel to that of DRAG-1. sma-1 encodes βH-spectrin, which is a very large spectrin found in invertebrates such as C. elegans and Drosophila22. The submembrane skeletal network is primarily formed from α2β2 spectrin tetramers, each composed of two α-spectrin and two β-spectrin subunits31. The spectrin network interacts with peripheral actin filaments to act in synapse function, muscle sarcomere structure, and axonal outgrowth32,33. Although SMA-1 function in shaping cells in the hypodermis and pharyngeal muscles has been reported34,35, its function in neuronal cells is unknown. SMA-1 may function in HSNs for branch formation by regulating the actin filaments. SMA-5/MAP kinase is specifically expressed in the intestine to control intestinal tube stability and body size23. Because the intestine has no direct contact with HSNs, it is possible that SMA-5 indirectly affects branching of the neuron. Because 60% of HSNs produce at least one branch in the drag-1 null mutant background and 40–50% of HSNs still make branches even in drag-1; sma-1 or drag-1; sma-5 double mutants, additional mechanisms must exist to control branch formation in HSNs. These additional factors/pathways include KAL-1/anosmin-1 and EGL-17/fibroblast growth factor (FGF), which act through SAX-7/L1CAM and EGL-15/FGF receptor to regulate HSN branching36. Additionally, the immunoglobulin superfamily protein SYG-1, which is expressed in HSN, is required for HSN branching and appropriate synaptogenesis with egg-laying muscles37.
How does DRAG-1 functions to regulate HSN axon branching? We believe that hypodermally expressed DRAG-1 acts through the neogenin receptor UNC-40 that is expressed in the HSNs to regulate HSN axon branching. We found via immunostaining that DRAG-1 is expressed in hypodermal seam cells and in the ventral hypodermis with the exception of the vulval epithelium. Furthermore, the defective axon branching of drag-1 mutants can be rescued by expressing drag-1 under the hypodermis specific rol-6 promoter or under the vulval epithelial precursor cell-specific dab-1 promoter, but not the hypodermal seam cell-specific SCM promoter. The dab-1 promoter drives gene expression in the descendants of vulval precursor cells P5.p, P6.p and P7.p during the Pn.pxx stage and in the descendants of P5.p and P7.p during the Pn.pxxx stage29. It is during this period that the HSN axons contact with some of these vulval precursor cells, defasciculate from the ventral nerve cord and form branches24,37,38. Thus, it is likely that DRAG-1 ectopically expressed in the vulval precursor cells signals the HSN axon to induce branching. In wild type animals DRAG-1 is expressed in the ventral hypodermis, which is directly adjacent to the ventral nerve cord, and the HSN axons fasciculate with the ventral nerve cord twice at regions posterior and anterior to the vulva21. Thus, DRAG-1 from the hypodermal cells can bind to the neogenin receptor UNC-40 that is expressed in the HSNs to induce HSN branching in a non-cell-autonomous fashion. This is consistent with UNC-40 being an ortholog of vertebrate neogenin, which acts as a receptor for RGM proteins30,39. Moreover, DRAG-1 physically interacts with the extracellular domain of UNC-40 in C. elegans16, similar to the interaction observed between human RGMc and neogenin40. Neither DRAG-1 with its GPI-anchor sequence deleted (therefore as a potential secreted form) nor DRAG-1 fused with the LIN-12 transmembrane domain (therefore as a potential membrane-anchored form) rescued the axon branching defects of drag-1 mutants. The latter was unexpected because the same construct significantly rescues the drag-1 defect in the control of mesodermal cell differentiation15. With respect to mesodermal cell differentiation, DRAG-1 and UNC-40 are expressed and function in the same cells to promote BMP signaling. In HSN branching, however, they are expressed in different cells that contact each other. Thus, it is possible that the modes of signaling from DRAG-1 to UNC-40 may differ depending on the cellular context. Studies of vertebrate RGM and neogenin proteins have yielded similar findings showing that they either function in the same cells or in different cells to regulate different processes14.
In summary, we provide in vivo evidence that RGM proteins function to promote axon branching in C. elegans. Our finding contrasts with the observation that RGM proteins suppress the branching of axons in the mammalian brain4,5. RGMs may function in both ways depending on the tissues or the phases of organogenesis. Further research is needed to understand the precise function of RGMs in axon branching.
Methods
Strains and culture conditions
Culture and handling of C. elegans were as described41. The following strains were used: N2 (wild type, WT), drag-1(tk81) (this work), drag-1(tm3773) (National Bioresource Project), unc-119(e2498)42, sma-1(e30), sma-2(e502), sma-3(wk28), sma-4(e729), sma-5(n678), sma-6(wk7), sma-8(e2111), sma-9(wk55), unc-40(e271)41,43,44,45. HSNs were visualized using an integrated transgene kyIs262[unc-86p::myrGFP; odr-1::RFP]20. drag-1(tk81) was isolated by the trimethylpsoralen and UV irradiation method46.
Analysis of HSN branching
HSN branching phenotype was analyzed with confocal or fluorescence microscopy using young adult hermaphrodites. Young adult hermaphrodites were selected from the mixed population by having mature vulva and no fertilized eggs in the uterus.
Plasmid construction
drag-1p::drag-1::GFP::GPI, drag-1p::drag-1::GFP::∆GPI, and drag-1p::drag-1::GFP::lin-12TM correspond to pJKL849, pCX192, and pCX194, respectively15. To produce drag-1p::Venus, the drag-1 promoter region was PCR amplified from genomic DNA using primers 5′-TAGCCTGCAGGTTTCCGAAGACAGGGGAACATGGAA-3′ and 5′-GTTCGTCGACACTCTGTCAAGTCTTCTCATCTCACG-3′, digested with PstI and SalI, and cloned into the PstI and SalI sites of pPD95.75. To produce drag-1p::drag-1::Venus, the drag-1 coding region was PCR amplified from genomic DNA with primers 5′-GTCAGTCGACATGTCAATAGTCTATCTCG-3′ and 5′-CATGGGTACCAAGCATAACAATGATAAAAGAGC-3′, digested with SalI and KpnI, and cloned into the SalI and KpnI sites of drag-1p::Venus. To produce drag-1p::drag-1, drag-1p::drag-1::Venus was PCR amplified with primers 5′-GATCGCTAGCCTTGTCTGGTGTCAAAAATAATAGG-3′ and 5′-TCGCTAGCTCAGCATAACAATGATAAAAGAGCAAAA-3′, digested with NheI, and self-ligated. To produce drag-1p::drag-1::Venus::GPI, the Venus coding region was PCR amplified from pPD95.75 with primers 5′-GCATGGGCCCAGGGTACCGGTAGAAAAAATGAGT-3′ and 5′-GCATGGGCCCTTTGTATAGTTCATCCATGCCAAG-3′, digested with ApaI, and ligated into drag-1p::drag-1::GFP::GPI (pJKL849) in which the green fluorescent protein (GFP) coding region had been deleted by ApaI digestion. rol-6 and myo-2 promoter regions were PCR amplified with primers 5′-CAGTGCATGCCGAGAAGAGTCCGGTGTGAA-3′ and 5′-CAGTGTCGACCTGGAAATTTTCAGTTAGATCTAAAG-3′, and 5′-CAGTGCATGCGTGAGCAAGTGTGCGGCATC-3′ and 5′-CAGTGTCGACTTCTGTGTCTGACGATCGAGGG-3′, respectively. These PCR fragments were digested with SphI and SalI and ligated with drag-1p::drag-1 in which the drag-1 promoter region was deleted by SphI and SalI digestion to produce rol-6p::drag-1, myo-2p::drag-1. elt-2 promoter region was PCR amplified from genomic DNA with primers 5′-CAGTCCTGCAGGGTGACCGCTCAAAATAAAAGG-3′ and 5′-CAGTCTCGAGTCTATAATCTATTTTCTAGTTTCTA-3′. The PCR fragment was digested with SbfI and XhoI and ligated with drag-1p::drag-1 in which the drag-1 promoter region was deleted by SbfI and SalI digestion to produce elt-2p::drag-1. To produce SCMp::drag-1, the SCM region was PCR amplified from genomic DNA with primers 5′-ATGAAATAAGCTTGCATGCCTGCAGCCAAGCTTGCATGCCTGCAG-3′ and 5′-CGAGATAGACTATTGACATGTCGACTCCTTTGGCCAATCCCGGG-3′, and the resulted fragment was fused with drag-1p::drag-1 in which the drag-1 promoter region was deleted by PstI and SalI digestion using In-Fusion Cloning Kit (Takara Bio). To produce dab-1p::drag-1, the dab-1 promoter region was PCR amplified from genomic DNA with primers 5′-AAATAAGCTTGCATGCAAGATTATCCCAAATTGTGGACCGT-3′ and 5′-CTATTGACATGTCGACTGTTTCGAGAGAACCTTTAGAAATAAGATT-3′, and the resulted fragment was fused with drag-1p::drag-1 in which the drag-1 promoter region was deleted by SphI and SalI digestion using In-Fusion Cloning Kit (Takara Bio).
Production of transgenic animals
We injected DNA mixtures into the gonads of unc-119(e2498), drag-1(tk81); unc-119(e2498); kyIs262 or drag-1(tm3773); unc-119(e2498); kyIs262 adult hermaphrodites47. For transgenic rescue experiments, test plasmids were injected at 10–20 ng/μl with 160–170 ng/μl of pBSII KS(–) and 20 ng/μl of unc-119+ plasmid pDP#MM016B42. For immunohistochemistry, drag-1p::drag-1 and drag-1p::drag-1::Venus::GPI plasmids were injected at 150 ng/μl with 30 ng/μl of pBSII KS(–) and 20 ng/μl pDP#MM016B.
Production of antibodies
The RNA sample extracted from wild-type worms was treated with SuperScript III reverse transcriptase (Invitrogen) using a primer 5′-TCAGCATAACAATGATAAAAGAGC-3′ designed to anneal at the 3′-end of the coding region, and single-strand cDNA was produced. The double-strand cDNA was amplified by PCR using a primer designed to anneal at the SL1 splice leader sequence 5′-GGTTTAATTACCCAAGTTTGAG-3′ and the 3′-end primer48. The region coding for DRAG-1 peptide from I131 to E368 was amplified using primers 5′-GTCACATATGATAATGTTCAATGGCTCCGTGC-3′ and 5′-GTCACTCGAGTTCTTTCTGGAACCGAGCATG-3′, digested with NdeI and XhoI, and ligated into the pET-19b vector using the NdeI and XhoI sites. The resulting antigenic peptide of DRAG-1 was expressed as a histidine-tagged fusion protein in Escherichia coli and was used to immunize rabbits. The generated antibody was affinity purified.
Immunohistochemistry
Immunohistochemistry was performed as described49. The DRAG-1 antibody was used as the primary antibody at 4 μg/ml. Alexa 594–labeled donkey anti-rabbit IgG (Life Technologies) was used as the secondary antibody at a dilution of 1:500.
Microscopy
Nomarski and fluorescence microscopy was performed using an Axioplan 2 microscope equipped with Axiocam CCD camera (Zeiss). Confocal laser scanning microscopy was conducted with LSM5 (Zeiss) equipped with a C-apochromat 63× (water immersion; NA, 1.2) lens controlled by PASCAL version 3.2 SP2 or ZEN software (Zeiss).
Data availability
Strains and plasmids are available upon request. The authors state that all data necessary for confirming the conclusions presented in the article are represented fully within the article.
References
Sainath, R. & Gallo, G. Cytoskeletal and signaling mechanisms of neurite formation. Cell Tissue Res. 359, 267–278. https://doi.org/10.1007/s00441-014-1955-0 (2015).
Dent, E. W., Barnes, A. M., Tang, F. & Kalil, K. Netrin-1 and semaphorin 3A promote or inhibit cortical axon branching, respectively, by reorganization of the cytoskeleton. J. Neurosci. 24, 3002–3012. https://doi.org/10.1523/JNEUROSCI.4963-03.2004 (2004).
Mann, F., Zhukareva, V., Pimenta, A., Levitt, P. & Bolz, J. Membrane-associated molecules guide limbic and nonlimbic thalamocortical projections. J. Neurosci. 18, 9409–9419 (1998).
Yoshida, J., Kubo, T. & Yamashita, T. Inhibition of branching and spine maturation by repulsive guidance molecule in cultured cortical neurons. Biochem. Biophys. Res. Commun. 372, 725–729. https://doi.org/10.1016/j.bbrc.2008.05.124 (2008).
Shibata, K. et al. Repulsive guidance molecule a regulates hippocampal mossy fiber branching in vitro. NeuroReport 24, 609–615. https://doi.org/10.1097/WNR.0b013e3283632c08 (2013).
Camus, L. M. & Lambert, L. A. Molecular evolution of hemojuvelin and the repulsive guidance molecule family. J. Mol. Evol. 65, 68–81. https://doi.org/10.1007/s00239-006-0241-5 (2007).
Monnier, P. P. et al. RGM is a repulsive guidance molecule for retinal axons. Nature 419, 392–395. https://doi.org/10.1038/nature01041 (2002).
Matsunaga, E. et al. RGM and its receptor neogenin regulate neuronal survival. Nat. Cell Biol. 6, 749–755. https://doi.org/10.1038/ncb1157 (2004).
Kitayama, M., Ueno, M., Itakura, T. & Yamashita, T. Activated microglia inhibit axonal growth through RGMa. PLoS ONE 6, e25234. https://doi.org/10.1371/journal.pone.0025234 (2011).
Tassew, N. G. et al. Modifying lipid rafts promotes regeneration and functional recovery. Cell Rep. 8, 1146–1159. https://doi.org/10.1016/j.celrep.2014.06.014 (2014).
Babitt, J. L. et al. Bone morphogenetic protein signaling by hemojuvelin regulates hepcidin expression. Nat. Genet. 38, 531–539. https://doi.org/10.1038/ng1777 (2006).
Zhou, Z. et al. Neogenin regulation of BMP-induced canonical Smad signaling and endochondral bone formation. Dev. Cell 19, 90–102. https://doi.org/10.1016/j.devcel.2010.06.016 (2010).
Wang, R. H. et al. A role of SMAD4 in iron metabolism through the positive regulation of hepcidin expression. Cell Metab. 2, 399–409. https://doi.org/10.1016/j.cmet.2005.10.010 (2005).
Siebold, C., Yamashita, T., Monnier, P. P., Mueller, B. K. & Pasterkamp, R. J. RGMs: Structural insights, molecular regulation, and downstream signaling. Trends Cell Biol. 27, 365–378. https://doi.org/10.1016/j.tcb.2016.11.009 (2017).
Tian, C. et al. The RGM protein DRAG-1 positively regulates a BMP-like signaling pathway in Caenorhabditis elegans. Development 137, 2375–2384. https://doi.org/10.1242/dev.051615 (2010).
Tian, C. et al. The neogenin/DCC homolog UNC-40 promotes BMP signaling via the RGM protein DRAG-1 in C. elegans. Development 140, 4070–4080. https://doi.org/10.1242/dev.099838 (2013).
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans. Philos. Trans. R. Soc. Lond. B Biol. Sci. 314, 1–340. https://doi.org/10.1098/rstb.1986.0056 (1986).
Tang, X. & Wadsworth, W. G. SAX-3 (Robo) and UNC-40 (DCC) regulate a directional bias for axon guidance in response to multiple extracellular cues. PLoS ONE 9, e110031. https://doi.org/10.1371/journal.pone.0110031 (2014).
Shioi, G. et al. Mutations affecting nerve attachment of Caenorhabditis elegans. Genetics 157, 1611–1622 (2001).
Adler, C. E., Fetter, R. D. & Bargmann, C. I. UNC-6/Netrin induces neuronal asymmetry and defines the site of axon formation. Nat. Neurosci. 9, 511–518. https://doi.org/10.1038/nn1666 (2006).
Garriga, G., Desai, C. & Horvitz, H. R. Cell interactions control the direction of outgrowth, branching and fasciculation of the HSN axons of Caenorhabditis elegans. Development 117, 1071–1087 (1993).
McKeown, C., Praitis, V. & Austin, J. sma-1 encodes a βH-spectrin homolog required for Caenorhabditis elegans morphogenesis. Development 125, 2087–2098 (1998).
Geisler, F. et al. A novel function for the MAP kinase SMA-5 in intestinal tube stability. Mol. Biol. Cell 27, 3855–3868. https://doi.org/10.1091/mbc.E16-02-0099 (2016).
Chia, P. H., Chen, B., Li, P., Rosen, M. K. & Shen, K. Local F-actin network links synapse formation and axon branching. Cell 156, 208–220. https://doi.org/10.1016/j.cell.2013.12.009 (2014).
Kramer, J. M. & Johnson, J. J. Analysis of mutations in the sqt-1 and rol-6 collagen genes of Caenorhabditis elegans. Genetics 135, 1035–1045 (1993).
Okkema, P. G., Harrison, S. W., Plunger, V., Aryana, A. & Fire, A. Sequence requirements for myosin gene expression and regulation in Caenorhabditis elegans. Genetics 135, 385–404 (1993).
Fukushige, T., Hawkins, M. G. & McGhee, J. D. The GATA-factor elt-2 is essential for formation of the Caenorhabditis elegans intestine. Dev. Biol. 198, 286–302 (1998).
Koh, K. & Rothman, J. H. ELT-5 and ELT-6 are required continuously to regulate epidermal seam cell differentiation and cell fusion in C. elegans. Development 128, 2867–2880 (2001).
Kamikura, D. M. & Cooper, J. A. Lipoprotein receptors and a disabled family cytoplasmic adaptor protein regulate EGL-17/FGF export in C. elegans. Genes Dev. 17, 2798–2811. https://doi.org/10.1101/gad.1136103 (2003).
Rajagopalan, S. et al. Neogenin mediates the action of repulsive guidance molecule. Nat. Cell Biol. 6, 756–762. https://doi.org/10.1038/ncb1156 (2004).
Bennett, V. & Gilligan, D. M. The spectrin-based membrane skeleton and micron-scale organization of the plasma membrane. Annu. Rev. Cell Biol. 9, 27–66. https://doi.org/10.1146/annurev.cb.09.110193.000331 (1993).
Hammarlund, M., Davis, W. S. & Jorgensen, E. M. Mutations in β-spectrin disrupt axon outgrowth and sarcomere structure. J. Cell Biol. 149, 931–942 (2000).
Moorthy, S., Chen, L. & Bennett, V. Caenorhabditis elegans β-G spectrin is dispensable for establishment of epithelial polarity, but essential for muscular and neuronal function. J. Cell Biol. 149, 915–930 (2000).
Praitis, V., Ciccone, E. & Austin, J. SMA-1 spectrin has essential roles in epithelial cell sheet morphogenesis in C. elegans. Dev. Biol. 283, 157–170. https://doi.org/10.1016/j.ydbio.2005.04.002 (2005).
Raharjo, W. H., Ghai, V., Dineen, A., Bastiani, M. & Gaudet, J. Cell architecture: Surrounding muscle cells shape gland cell morphology in the Caenorhabditis elegans pharynx. Genetics 189, 885–897. https://doi.org/10.1534/genetics.111.132449 (2011).
Diaz-Balzac, C. A., Lazaro-Pena, M. I., Ramos-Ortiz, G. A. & Bulow, H. E. The adhesion molecule KAL-1/anosmin-1 regulates neurite branching through a SAX-7/L1CAM-EGL-15/FGFR receptor complex. Cell Rep. 11, 1377–1384. https://doi.org/10.1016/j.celrep.2015.04.057 (2015).
Shen, K., Fetter, R. D. & Bargmann, C. I. Synaptic specificity is generated by the synaptic guidepost protein SYG-2 and its receptor, SYG-1. Cell 116, 869–881. https://doi.org/10.1016/s0092-8674(04)00251-x (2004).
Asakura, T., Ogura, K. & Goshima, Y. UNC-6 expression by the vulval precursor cells of Caenorhabditis elegans is required for the complex axon guidance of the HSN neurons. Dev. Biol. 304, 800–810. https://doi.org/10.1016/j.ydbio.2007.01.028 (2007).
Cole, S. J., Bradford, D. & Cooper, H. M. Neogenin: A multi-functional receptor regulating diverse developmental processes. Int. J. Biochem. Cell Biol. 39, 1569–1575. https://doi.org/10.1016/j.biocel.2006.11.009 (2007).
Yang, F., West, A. P. Jr., Allendorph, G. P., Choe, S. & Bjorkman, P. J. Neogenin interacts with hemojuvelin through its two membrane-proximal fibronectin type III domains. Biochemistry 47, 4237–4245. https://doi.org/10.1021/bi800036h (2008).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Maduro, M. & Pilgrim, D. Identification and cloning of unc-119, a gene expressed in the Caenorhabditis elegans nervous system. Genetics 141, 977–988 (1995).
Savage-Dunn, C. et al. Genetic screen for small body size mutants in C. elegans reveals many TGFβ pathway components. Genesis 35, 239–247. https://doi.org/10.1002/gene.10184 (2003).
Watanabe, N., Nagamatsu, Y., Gengyo-Ando, K., Mitani, S. & Ohshima, Y. Control of body size by SMA-5, a homolog of MAP kinase BMK1/ERK5, in C. elegans. Development 132, 3175–3184. https://doi.org/10.1242/dev.01895 (2005).
Hedgecock, E. M., Culotti, J. G. & Hall, D. H. The unc-5, unc-6, and unc-40 genes guide circumferential migrations of pioneer axons and mesodermal cells on the epidermis in C. elegans. Neuron 4, 61–85. https://doi.org/10.1016/0896-6273(90)90444-k (1990).
Kubota, Y., Kuroki, R. & Nishiwaki, K. A fibulin-1 homolog interacts with an ADAM protease that controls cell migration in C. elegans. Curr. Biol. 14, 2011–2018. https://doi.org/10.1016/j.cub.2004.10.047 (2004).
Mello, C. C., Kramer, J. M., Stinchcomb, D. & Ambros, V. Efficient gene transfer in C.elegans: Extrachromosomal maintenance and integration of transforming sequences. EMBO J. 10, 3959–3970 (1991).
Krause, M. & Hirsh, D. A trans-spliced leader sequence on actin mRNA in C. elegans. Cell 49, 753–761. https://doi.org/10.1016/0092-8674(87)90613-1 (1987).
Kim, H. S. et al. VAB-10 spectraplakin acts in cell and nuclear migration in Caenorhabditis elegans. Development 138, 4013–4023. https://doi.org/10.1242/dev.059568 (2011).
Acknowledgements
We thank Noriko Nakagawa and Nami Okahashi for technical assistance. Some nematode strains used in this work were provided by the Caenorhabditis Genetics Center, which is funded by the National Institutes of Health National Center for Research Resources and by Shohei Mitani through the National Bioresource Project for the nematode. This work was supported by NIH R01 GM103869 and R35 GM130351 to J.L., and a Grant-in-Aid for Scientific Research by Ministry of Education, Culture, Sports, Science and Technology to K.N.
Author information
Authors and Affiliations
Contributions
K.N. designed research; K.T., H.-S.K., C.Y., K.K., Y.K., Y.S., C.T. and J.L. performed research and analyzed data; K.N. and J.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tsutsui, K., Kim, HS., Yoshikata, C. et al. Repulsive guidance molecule acts in axon branching in Caenorhabditis elegans. Sci Rep 11, 22370 (2021). https://doi.org/10.1038/s41598-021-01853-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-01853-8
- Springer Nature Limited
This article is cited by
-
A memristive circuit for self-organized network topology formation based on guided axon growth
Scientific Reports (2024)