Abstract
Glutamine:fructose-6-phosphate aminotransferase (GFAT) and phosphofructokinase (PFK) are enzymes related to chitin metabolism. RNA interference (RNAi) technology was used to explore the role of these two enzyme genes in chitin metabolism. In this study, we found that GFAT and PFK were highly expressed in the wing bud of Nilaparvata lugens and were increased significantly during molting. RNAi of GFAT and PFK both caused severe malformation rates and mortality rates in N. lugens. GFAT inhibition also downregulated GFAT, GNPNA, PGM1, PGM2, UAP, CHS1, CHS1a, CHS1b, Cht1-10, and ENGase. PFK inhibition significantly downregulated GFAT; upregulated GNPNA, PGM2, UAP, Cht2-4, Cht6-7 at 48 h and then downregulated them at 72 h; upregulated Cht5, Cht8, Cht10, and ENGase; downregulated Cht9 at 48 h and then upregulated it at 72 h; and upregulated CHS1, CHS1a, and CHS1b. In conclusion, GFAT and PFK regulated chitin degradation and remodeling by regulating the expression of genes related to the chitin metabolism and exert opposite effects on these genes. These results may be beneficial to develop new chitin synthesis inhibitors for pest control.
Similar content being viewed by others
Introduction
Chitin is a linear polymer composed of N-acetylglucosamine units connected by β-1, 4-glycoside bonds and is the second most abundant biopolymer in nature. It is widely distributed in fungi, nematodes, and arthropods1. In insects, chitin is a major component of the exoskeleton, trachea, and the peritrophic matrix that lines the midgut epithelium1,2,3,4. Chitin enhances the mechanical strength of the insect exoskeleton and protects intestinal epithelial cells from abrasion by rough foods, penetration by toxins, and infection by microorganisms3. However, as the size of insects increases, their growth and development depend on the periodic synthesis and degradation of chitin3,5.
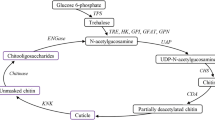
In nature, chitin exists in three crystal structures and is classified as α-, β-, and γ-chitin according to the arrangement of chitin chains2,3. In insects, chitin synthesis can be carried out in the microvilli of epidermal cells or intestinal epithelial cells, beginning with trehalose followed by a series of enzymatic reactions to eventually form chitin6. Previous studies have shown that the chitin synthetic pathway consists of at least eight enzymes2. The first enzyme involved in chitin synthesis is trehalase (TRE), which hydrolyzes trehalose to β-D-glucose7. When TRE gene expression was silenced in Leptinotarsa decemlineata, the chitin content decreased accordingly8. Chitin biosynthesis occurs via the hexosamine pathway (HP)9. β-D-glucose can be phosphorylated by hexokinase (HK); it is then converted to fructose-6-phosphate (F-6-P) through isomerization by glucose-6-phosphate isomerase (GPI) and amino and acetyl groups are added via glutamine:fructose-6-phosphate aminotransferase (GFAT) and glucosamine-6-phosphate N-acetyltransferase (GNPNA) to form N-acetylglucosamine-6-phosphate. Phosphoacetylglucosamine mutase (PGM) transforms its phosphate group to form N-acetylglucosamine-1-phosphate. Subsequently, N-acetylglucosamine-1-phosphate is combined with uridine nucleoside triphosphate (UTP) to form UDP-N-acetylglucosamine (UDP-GlcNAc), the final product of HP as well as the precursor substance for chitin, under the action of UDP-N-acetylglucosamine pyrophosphorylase (UAP). Finally, N-acetylglucosamine (GlcNAc) polymers, also known as chitin, is formed by the action of chitin synthase (CHS)4,5,10,11. Two kinds of CHS, CHS1 and CHS2, were identified in insects12. Insect CHS1 and CHS2 are differentially expressed and regulated during growth and development. CHS1 is mainly responsible for chitin synthesis in cuticle and trachea cells11, while CHS2 is only responsible for chitin synthesis on midgut peritrophic membrane3,11. However, hemipteran insects lack peritrophic membrane, and CHS2 has not been found in hemipteran insects at present.
The physical properties of chitin itself are soft and form the hard exoskeleton of insects by combining with other proteins, but it limits the growth of insects. Therefore, along with chitin synthesis, its timely degradation is also necessary13. Chitin polymers can be enzymatically digested by chitinases, which are divided into endochitinases, exochitinases, and β-1, 4-N-acetyl-glucosaminidases14. Endochitinases and exochitinases are responsible for degrading long-chain chitin into short-chain chitooligosaccharides; β-1, 4-N-acetyl-glucosaminidases then hydrolyze short-chain chitooligosaccharides into GlcNAcs15.
GFAT is the first and rate-limiting enzyme of HP, and converts fructose 6-phosphate and glutamine into glucosamine 6-phosphate and glutamate16. In human keratinocytes, GFAT1 silencing reduced the content of UDP-GlcNAc and hyaluronan17. The protein sequence of the GFAT gene is very similar in different types of organisms such as bacteria, insects, yeast, and mammals. GFAT is expressed in different mammalian tissues and may have different functions18. Phosphorylation plays an important role in regulating enzyme activity19; Li et al. have shown that the increase in GFAT activity can be achieved by phosphorylation of AMP-activated protein kinase at Ser243, a new regulation in addition to phosphorylation of protein kinase A20. In addition, a lot of experimental data demonstrate that GFAT activity can be inhibited by UDP-GlcNAc, the last product of HP21. GFAT plays an important regulatory role in shrimp's resistance to environmental stress, proliferation of mammalian cardiomyocytes and cancer cells through catalyzing hexosamine synthesis22,23,24. However, little is known about GFAT in insects, especially the relationship between GFAT and insect chitin metabolism.
Unlike GFAT, which is directly involved in chitin synthesis, phosphofructokinase (PFK) is a highly conserved and the main rate-limiting enzyme in the glycolytic pathway. Its activity significantly affects glucose consumption and energy production25. Both PFK in prokaryotic and eukaryotic cells catalyze phosphorylation of F-6-P to fructose-1,6-bisphosphate, but eukaryotic PFK is more than twice the size of bacterial PFK26. In vertebrates, PFK is activated by ADP, AMP, and fructose-2,6-diphosphate, whereas it is inhibited by physiological concentrations of ATP and citrate27. However, citrate does not inhibit PFK in insects28. PFK catalyzes the irreversible reaction in the glycolytic pathway and plays an essential role in glycometabolism, then is involved in the regulation of locomotion of different animals and other physiological activities29. However, there are few studies on the relationship between PFK and chitin metabolism in insects.
Rice serves as a staple food for more than half the world’s population, and is a source of calories for billions of people worldwide30. However, its production and storage has long been threatened by approximately 800 species of insect pests31. Nilaparvata lugens (Stål) (Hemiptera: Delphacidae), a hemimetabolous brown planthopper (BPH), is one of the most destructive and notorious insect pests32. It possesses high fecundity, sucks rice sap, oviposits in rice tissues, and transmits viruses such as grassy stunt virus and rugged stunt virus33. Use of chemical pesticides such as etofenprox has led to resistance in N. lugens; thus, integrated pest management is required to develop selective and environmentally safe biopesticides34. There is no chitin production in plants and vertebrates including humans and thus, the chitin metabolism pathway can be used as a safe target for pest control. During the past few years, some third-generation insecticides such as benzoylphenylurea related to the inhibition of arthropod pest chitin synthesis have been developed and commercialized, which can result in abortive molting and egg hatching as a consequence of chitin synthesis inhibition in the course of cuticle formation, thereby achieving the purpose of controlling pests35,36,37. A series of enzymes are involved directly or indirectly in the synthesis and hydrolysis of chitin. Therefore, we have selected the most critical enzymes in the HP and glycolytic pathway, GFAT and PFK, respectively, and have explored their exact effects on chitin metabolism in N. lugens using RNA interference (RNAi) technology in this study to evaluate their potential as targets for novel pesticide.
Results
Expression of GFAT and PFK in different developmental stages of N. lugens
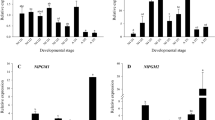
GFAT and PFK expression levels in different developmental stages of N. lugens were determined by qRT-PCR. These results showed that the expression trends of NlGFAT and NlPFK were relatively similar. Expression of GFAT increased on the 3rd day of the fourth instar nymph but decreased on the 2nd day of fifth instar nymph, subsequently remained at relatively low levels before molting, and significantly increased (p < 0.05) during the adult stage (Fig. 1A). During the nymph stage, PFK expression was the highest on the 3rd day of fourth instar nymph and remained at low and stable levels in other nymph stages (Fig. 1B). During the adult stage, the expression of PFK in male adults was significantly (p < 0.05) increased, whereas the expression of PFK in female adults showed no changes (Fig. 1B). Interestingly, there were significant gender differences in the expression of both GFAT and PFK (p < 0.05) (Fig. 1A,B).
Expression levels of GFAT (A) and PFK (B) genes in different developmental stages of Nilaparvata lugens. GFAT, glutamine:fructose-6-phosphate aminotransferase; PFK, phosphofructokinase. Expression levels were measured by quantitative real-time PCR, with 18S RNA as the internal control. Values are means ± SE from three independent measurements. The age of N. lugens was defined as follows: 4th—1d, 1st day of fourth instar nymph; 4th—2d, 2nd day of fourth instar nymph; 4th—3d, 3rd day of fourth instar nymph; 5th—1d, 1st day of fifth instar nymph; 5th—2d, 2nd day of fifth instar nymph; 5th—3d, 3rd day of fifth instar nymph; AM, male adult; AF, female adult. Different letters indicate significant differences according to Duncan’s test (p < 0.05).
GFAT and PFK expression in different tissues of N. lugens
According to the qRT-PCR results, the relative expression levels of GFAT and PFK were both the highest in the wing bud and were significantly (p < 0.05) higher than their expression levels in the head, leg, ovary, cuticle, midgut, and fat body (Fig. 2A,B). GFAT expression was also high in the cuticle of N. lugens, and its expression in the head, ovary, and fat body was relatively low (Fig. 2A). PFK is also highly expressed in the head of N. lugens, second only to the wing bud, and its expression levels in the legs, ovaries, cuticles, midgut, and fat body are similar and low (Fig. 2B). Overall, the results showed that the expression levels of GFAT and PFK were quite different among several N. lugens tissues, indicating that both their expression patterns are tissue-specific.
Expression levels of GFAT (A) and PFK (B) in seven different tissues of Nilaparvata lugens. GFAT, glutamine:fructose-6-phosphate aminotransferase; PFK, phosphofructokinase. The head, leg, ovary, cuticle, fat body, midgut, and wing bud of N. lugens used to detect the tissue expression level of genes, were evenly collected from individuals of nymphs in different instars and from adults. Gene expression levels were measured by quantitative real-time PCR with 18S RNA as the internal control. Values are means ± SE from three independent measurements. Different letters indicate significant differences according to Duncan’s test (p < 0.05).
The relative expression levels of target genes and phenotype observation after dsRNA injection
The results of qRT-PCR showed that the relative expression levels of NlGFAT (Fig. 3A) and NlPFK (Fig. 3B) were significantly decreased at 48 h and 72 h after dsGFAT and dsPFK injection, respectively, which suggests that dsRNA successfully inhibited the expression of target genes. In addition, after dsGFAT and dsPFK separate injection into the nymphs, three kinds of abnormal phenotypes including only molting difficulties, only wing deformities, and both, were observed (Fig. 3C). As seen above, silencing of both NlGFAT and NlPFK impacted on the development of N. lugens.
Relative expression levels of GFAT (A) and PFK (B) after dsGFAT and dsPFK injection, and phenotype changes (C) after dsRNA injection in Nilaparvata lugens. GFAT, glutamine:fructose-6-phosphate aminotransferase; PFK, phosphofructokinase. The N. lugens used for the microinjection of RNAi were at the 1st day of 5th instar nymph stage. dsGFAT or dsPFK injection were used as the test groups whereas dsGFP injection was used as the control group. Gene expression levels were measured by quantitative real-time PCR with 18S RNA as the internal control. Values are means ± SE from three independent measurements. Different letters indicate significant differences according to Duncan’s test (p < 0.05).
Malformation rates and mortality rates of N. lugens after dsRNA injection
As shown in the figures, after 48 h or 72 h of GFAT inhibition, the malformation rates were 19.19% and 23.77% (Fig. 4A), respectively, and the mortality rates were 25.63% and 32.51% (Fig. 4B), respectively. Similarly, after dsPFK injection, the malformation rates were 13.76% at 48 h and 16.14% at 72 h (Fig. 4A), and the mortality rates were 18.06% at 48 h and 25.59% at 72 h (Fig. 4B). Thus, the mortality and malformation rates of N. lugens showed significant increases (p < 0.01) after dsGFAT or dsPFK injection alone; both GFAT and PFK thus have a great influence on the growth and development of N. lugens individuals.
Malformation rates (A) and mortality (B) rates of Nilaparvata lugens at 48 h and 72 h after dsRNA injection. GFAT, glutamine:fructose-6-phosphate aminotransferase; PFK, phosphofructokinase. The N. lugens used for the microinjection of RNAi were at the 1st day of the 5th instar nymph stage and dsGFAT or dsPFK injection were used as the test groups whereas dsGFP injection was used as the control group. This experiment had three biological repeats. *, significant differences (P < 0.05); **, extremely significant differences (p < 0.01). The deformity rate of the dsGFP treatment group was zero, with “●” indicating the data.
Relative expression levels of chitin biosynthesis-related genes after dsRNA injection
One GFAT, GNPNA, UAP and two PGM genes were identified in N. lugens6. When the PFK of N. lugens was inhibited, the relative expression levels of GFAT were decreased extremely significantly (p < 0.01) at 48 h and 72 h (Fig. 5A), but the expression of GNPNA, PGM2, and UAP showed an extremely significant increase (p < 0.01) at 48 h and an extremely significant decrease (p < 0.01) at 72 h (Fig. 5B,D,E). In contrast, the expression of PGM1 showed little change at 48 h whereas it increased extremely significantly (p < 0.01) at 72 h (Fig. 5C).
Relative expression levels of chitin biosynthesis-related genes at 48 h and 72 h after dsRNA injection. GFAT, glutamine:fructose-6-phosphate aminotransferase; PFK, phosphofructokinase; GAPNA, glucosamine-6-phosphate N-acetyltransferase; PGM, phosphoacetylglucosamine mutase; UAP, UDP-N-acetylglucosamine pyrophosphorylase. The expression of GFAT (A), GNPNA (B), PGM1 (C), PGM2 (D), and UAP (E) was determined by quantitative real-time PCR with 18S RNA as the internal control. The N. lugens used for RNAi microinjection were at the 1st day of the 5th instar nymph stage and dsGFAT or dsPFK injection were used as test groups whereas the dsGFP injection was used as the control group. Values are the average of three sets of data and standard errors were calculated. *, significant differences (P < 0.05); **, extremely significant differences (p < 0.01).
The qRT-PCR results showed that the relative expression levels of GFAT, GNPNA, PGM1, and UAP were significantly decreased (p < 0.01) at 48 h and 72 h after dsGFAT injection (Fig. 5B,C,E) whereas that of PGM2 decreased extremely significantly (p < 0.01) at 48 h but showed little change at 72 h (Fig. 5D).
Relative expression levels of chitin degradation-related genes after dsRNA injection
There were 12 chitinase-like genes in N. lugens, including ten chitinases (Cht), one imaginal disc growth factor (IDGF) and one endo-β-N-acetylglucosaminidase (ENGase)1. At 48 h and 72 h after inhibiting the expression of NlGFAT using RNAi, the relative expression levels of all chitin degradation-related genes showed extremely significant declines (p < 0.01) (Fig. 6A–C,E–,J,L), whereas that there was little change in Cht4 expression at 48 h and in IDGF expression at 48 h and 72 h (Fig. 6D,K).
Relative expression levels of chitin degradation-related genes at 48 h and 72 h after dsRNA injection. GFAT, glutamine:fructose-6-phosphate aminotransferase; PFK, phosphofructokinase; Cht, chitinase; IDEF, imaginal disc growth factor; ENGase, endo-β-N-acetylglucosaminidase. Nilaparvata lugens larvae at the 1st day of the 5th instar stage were divided into three groups and injected with dsGFP, dsGFAT, and dsPFK, respectively. Insects were collected and used to determine the relative expression levels of Cht1 to Cht10 (A to J), IDGF (K), and ENGase (L) at 48 h and 72 h after dsRNA injection. Three replicates were performed per group. *, significant differences (P < 0.05); **, extremely significant differences (p < 0.01).
After injecting dsPFK into N. lugens on the 1st day of the 5th instar nymph, the relative expression levels of Cht2 to Cht8 (Fig. 6B–H), Cht10 (Fig. 6J) and ENGase (Fig. 6L) showed extremely significant increases (p < 0.01) at 48 h. But their subsequent trends were not the same. The expression levels of Cht2, Cht3, Cht4, Cht6, and Cht7 decreased significantly (p < 0.05) (Fig. 6B,C) and extremely significantly (p < 0.01) (Fig. 6D,F,G), whereas the expression levels of Cht8, Cht10, ENGase showed extremely significant declines at 72 h (p < 0.01) (Fig. 6H,J,L); the expression of Cht5 was restored to the same levels as the control group (Fig. 6E). Further, silencing PFK gene did not affect the expression of Cht1 and IDGF (Fig. 6A,K), but affected Cht9 as its expression decreased significantly (p < 0.05) at 48 h and showed an extremely significant increase (p < 0.01) at 72 h (Fig. 6I).
Relative expression levels of chitin synthases after dsRNA injection
N. lugens possesses one copy of CHS1 and there are two transcript variants (CHS1a and CHS1b) in N. lugens38. At both 48 h and 72 h after GFAT gene silencing by RNAi, the relative expression of CHS1, CHS1a, and CHS1b showed an extremely significant (p < 0.01) decline (Fig. 7A–C). However, an almost converse expression pattern was observed after dsPFK injection, the relative expression of CHS1, CHS1a, and CHS1b showed an extremely significant (p < 0.01) increase at 48 h and 72 h (Fig. 7A–C), except for CHS1a at 72 h (Fig. 7B).
Relative expression levels of CSH1 (A), CHS1a (B), and CHS1b (C) at 48 h and 72 h after dsRNA injection. CHS, chitin synthase. Nilaparvata lugens larvae on the 1st day of 5th instar stage were divided into three groups and injected with dsGFP, dsGFAT, and dsPFK, respectively. The dsGFP-treatment group was used as the control group. Three replicates were performed per group. *, significant differences (p < 0.05); **, extremely significant differences (p < 0.01).
Discussion
GFAT catalyzes the rate-limiting step of the UDP-GlcNAc synthesis pathway. Because of its role in the development of insulin resistance in type 2 diabetes39, studies on GFAT are mostly focused on mammals, whereas there have been rather few studies on the GFAT gene in insects for a long time and it has only been reported in Drosophila melanogaster, Aedes aegypti, Haemaphysalis longicornis at present21,40,41. Northern blot analysis of Drosophila melanogaster and Aedes aegypti showed two bands for GFAT1, the ratios of which varied in different developmental stages; GFAT1 was localized by whole mount in situ hybridization to chitin synthesis-related tissues, suggesting that DmGFAT and AeGFAT are involved in chitin synthesis21,40,42. Our studies have shown that NlGFAT was expressed in all stages after the 4th instar, and that there was a significant difference in its expression during the molt period from the 5th instar to the adult stage (Fig. 1A). In addition, NlGFAT was highly expressed at the wing bud and cuticle (Fig. 2A), which contains significant amounts chitin43,44. Therefore, GFAT and chitin metabolism are closely related. As rate limiting enzyme in the glycolytic pathway, PFK is closely related to diabetic cardiomyopathy45, and there have been few studies on PFK gene in insects. In Spodoptera litura, transcriptional expression of PFK occurs at a stable and low level during the period from larval stage to pupa, but its enzyme activity decreased dramatically in the pre-pupae and was recovered in pupae during metamorphosis46. In N. lugens, the expression levels of PFK were decreased dramatically during the period of the 4th instar to 5th instar, but were increased extremely significantly during the 5th instar nymph to adult stage (Fig. 1B). Glycolysis in the cytosol could produce ATP, the chemical energy in cells, which is used to run the reactions that maintain viability, growth, and proper function of individuals47. Flight muscles are tissues that require a large supply of energy48. In our experiments, NlPFK showed the highest expression in wing buds (Fig. 2B), which is consistent with the requirement of energy. Overall, the expression levels of GFAT and PFK change significantly during molting, consistent with the pace of chitin metabolism, and high expression in chitinous tissues, suggesting a link between them and chitin metabolism.
RNAi is a biological process that may be mediated by exogenous dsRNA, which is sliced into small RNAs, causing endogenous complementary mRNA silencing49. RNAi is considered as an important tool for gene function research50. The results of qRT-PCR showed that the relative expression levels of NlGFAT (Fig. 3A) and NlPFK (Fig. 3B) were significantly decreased after dsRNA injection, respectively, which suggesting that dsRNA successfully inhibited the expression of target genes. In the present study, we obtained many interesting experimental results after GFAT-knockdown or PFK-knockdown using RNAi. Most insects possess two CHS genes (CHS1 and CHS2), but N. lugens possesses only CHS1 with two transcript variants (CHS1a and CHS1b)38. RNAi against NlCHS1 and NlCHS1a causes high mortality rates and severe morphological malformations38,51. Knockdown of NlTRE1 could downregulate CHS1 and CHS1a and cause abnormal phenotypes44, and knockdown of TPS1 could downregulate the expression of CHS1, CHS1a, and CHS1b, resulting in extremely high malformation and mortality rates in N. lugens52, as well as HK-knockdown also could result in the downregulation of CHS1, CHS1a, CHS1b53. In our study, the mRNA levels of CHS1, CHS1a, and CHS1b were acutely decreased at 48 h and 72 h after dsGFAT injection (Fig. 7). In addition, molting difficulties and wing deformities were observed with dsGFAT injection (Fig. 3); the malformation rates and mortality rates of N. lugens were also increased extremely significantly after dsGFAT injection, compared to the dsGFP injection group (Fig. 4A,B). These results are consistent with previous studies. In locusts, reduced expression of miR-71 and miR-263 increased CHS1 and CHS10 mRNA expression, thus resulting in molting defects54. Similarly, in these experiments, reduced expression of NlPFK increased CHS1, CHS1a, and CHS1b (Fig. 7), along with high malformation rates and mortality rates in N. lugens (Fig. 4).
To further investigate the effects of GFAT and PFK genes on chitin metabolism in N. lugens, we detected the expression of chitin synthesis pathway genes after silencing GFAT and PFK. When TRE1-1, TRE1-2, and TRE2 in N. lugens were co-inhibited using RNAi, the relative expression levels of GFAT, GNPNA, PGM1, PGM2 and UAP were decreased significantly, but the relative expression of PGM2 was increased significantly at 72 h55. In addition, the same effects were achieved by injecting validamycin, a kind of trehalase inhibitor6. This suggested that TRE could regulate chitin synthesis by regulating the transcriptional levels of other enzymes involved in chitin synthesis, and that PGM1 and PGM2 might be functionally complementary6,55. HK-knockdown could also result in downregulation of GFAT, GNPNA, and UAP53. In our experiment, when dsGFAT was injected into the 5th instar nymph of N. lugens, other than PGM2 being expressed normally at 72 h compared with the control group (Fig. 5D), the relative expression levels of GFAT, GNPNA, PGM1, PGM2, and UAP were dramatically decreased (Fig. 5), similar to the result of N. lugens TRE and HK gene inhibition6,53,55. Therefore, silencing GFAT expression directly leads to impaired chitin synthesis by inhibiting the chitin pathway genes.
The role of PFK in regulating energy metabolism during insect development has been studied in Spodoptera litura, but it is unclear whether it affects the chitin synthesis pathway46. Radiometric glycolysis assays have demonstrated that low rates of glycolysis did not affect the overall level of incorporation of glucose-derived carbon into HP, but low PFK activity promotes channeling of F-6-P into HP56. In our study, when PFK was inhibited, the mRNA levels of GFAT were sharply declined at 48 h and 72 h (Fig. 5A). However, contrary to the interference results of GFAT, the expression levels of GNPNA, PGM2 and UAP were increased sharply at 48 h after PFK inhibition, but decreased significantly after 72 h (Fig. 5B,D,E), and the expression of PGM1 was still in contrast to PGM2 (Fig. 5C). We speculated that when PFK is inhibited, more fructose-6-phosphate flows into HP, as shown by radioactive glycolysis experiments56. Therefore, inhibition of PFK expression might have promoted CHS transcription by upregulating chitin synthesis pathway genes, which is contrary to GFAT inhibition. However, inhibition of NlPFK resulted in reduced transcription levels of NlGFAT, which may indicate the existence of other regulatory pathways.
Chitinases belong to family 18 glycosylhydrolases and are essential enzymes for chitin degradation and remodeling in insects57,58. 12 chitinase-like genes were identified in N. lugens, including 10 chitinases (Cht), one imaginal disc growth factor (IDGF) and one endo-β-N-acetylglucosaminidase (ENGase)1. Among these 12 genes, RNAi targeting Cht1, Cht 5, Cht 7, Cht 9, and Cht 10 caused a lethal phenotype in N. lugens, whereas RNAi against Cht2, Cht3, Cht4, Cht6, Cht8, IDGF, ENGase had little effect on the morphology and survival of N. lugens1. Our results show that the relative expression levels of Cht1, Cht5, Cht7, Cht9, and Cht10 were decreased extremely significantly at 48 h and 72 h after dsGFAT injection (Fig. 6A,E,G,I,J), and that Cht5, Cht7, Cht9, and Cht10 were upregulated extremely significantly with PFK inhibition (Fig. 6E,G,I,J). These all lead to the occurrence of a lethal phenotype (Fig. 3), consistent with previous studies1. Thus, in addition to chitin synthesis, GFAT and PFK also affect the degradation of chitin, and play contrasting roles in the degradation process.
In summary, silencing of GFAT or PFK affects the synthesis and degradation of chitin by interfering with the transcription levels of crucial chitin metabolizing enzymes, thus resulting in extremely high malformation rates and mortality rates. Moreover, GFAT and PFK have opposite effects on chitin synthesis in N. lugens. All the above results provide theoretical support for the discovery of new targets for pest control. However, all measurements were based on transcriptional levels, but data on protein levels are lacking, so measurements of enzyme activity and related metabolites at the tissue level, rather than at the individual level, will be considered.
Methods
Insect sourcing and culture conditions
The N. lugens used in this study were provided by the China National Rice Research Institute (Hangzhou, China), and the variety of all rice (Oryza sativa L.) cultivars was Taichung Native 1 (TN1) planted in cement tanks from April to October and in a greenhouse or growth chamber during winter. Insects were reared on fresh TN1 rice seedlings in an artificial climate chamber at 26 ± 1 ºC, 70% relative humidity, and 16 L:8 D (light:dark) photoperiod55. All experiments were performed under the same conditions. Developmental stages were synchronized by collecting new eggs laid by N. lugens, and the instar was judged based on the hind foot and antennae of the nymph.
Collection and dissection of N. lugens in different developmental stage
N. lugens individuals used in gene expression stage analyses were obtained from the 4th instar nymphs on their first day, and after every 24 h until they reached the adult stage; 10 individuals were taken from each stage. Besides, female adults and male adults were also collected separately. The N. lugens used to detect the tissue expression level of genes were collected from 50 individuals of adults, and with a 1:1 ratio of male to female. The head, leg, ovary, cuticle, fat body, midgut, and wing bud of N. lugens were dissected in a saline solution (0.75% NaCl) under an EZ4 microscope (Leica, Germany). Three biological replicates were used for each developmental stage and tissue sample. All samples were kept at − 80 °C until RNA extraction.
Total RNA extraction and cDNA synthesis
Tissues and whole bodies of N. lugens were used to extract the total RNA with TRIzol reagent (Invitrogen, Carlsbad, California, USA), following the manufacturer’s instructions. Total RNA integrity was determined by 1% agarose gel electrophoresis, and the RNA concentration and purity were determined by measuring the sample absorbance at 260 nm on a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA)44; the purified RNA was stored at -80℃ for future experiments. First-strand complementary DNA (cDNA) synthesis was performed using the PrimeScript RT reagent kit with gDNA Eraser (Takara, Kyoto, Japan) following the manufacturer’s instructions and was stored at − 20 ℃.
NlGFAT and NlPFK expression in several tissues and developmental stages using quantitative real-time polymerase chain reaction (qRT-PCR)
cDNA synthesis and qRT-PCR were performed to analyze the distribution of NlGFAT and NlPFK using gene-specific primers (Table 1). Using 1 µg of total RNA as template, and a specifically designed Nl18S primer pair (Table 1) the stability of 18S RNA was demonstrated in a PCR performed under the following conditions: 95 °C for 5 min; 28 cycles at 95 °C for 30 s, 60 °C for 30 s, and 72 °C for 30 s; and a final extension at 72 °C for 10 min44,55.
The expression of NlGFAT and NlPFK in several tissues and developmental stages was estimated by qRT-PCR with a SYBR Green master mix (SYBR Green Premix Ex Taq, Takara, Japan) in a Bio-Rad CFX96 Real-Time PCR Detection System (Bio-Rad Laboratories Inc., Hercules, CA, USA). Each PCR was performed in a 20 µL volume, containing 1 µL cDNA, 1 µL (10 µM) of each primer, 7 µL ultrapure water, and 10 µL SYBR buffer44,55. The reactions were performed under the following conditions: preincubation at 95 ºC for 2 min; 39 cycles of 95 ºC for 5 s and annealing at 59 °C for 30 s; and a melting curve at 65–95 °C. Amplification of 18S RNA was used as an internal control44,55.
Double-stranded RNA (dsRNA) synthesis and injections
The N. lugens cDNA template and specific primers (Table 2) were used to amplify the NlGFAT and NlPFK genes with reverse transcription polymerase chain reaction (RT-PCR). The reaction procedure is set as follows: preincubation at 95 ºC for 3 min, 35 cycles at 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 1 min; and a final extension at 72 °C for 10 min. Purified GFAT and PFK amplicons were transcribed to synthesize dsRNA using the T7 RiboMax Express RNAi System (Promega Corporation, Madison, WI)44. A green fluorescence protein (GFP) gene amplicon was also used to synthesize dsRNA for being control group. Sense and anti-sense strands were separately produced using PCR and were then mixed for annealing. Reactions were incubated for 10 min at 70 °C and then placed on ice for 20 min. Finally, dsRNAs were purified with 95% ethanol and 4.4 M sodium acetate (pH 5.2), then washed with 70% ethanol, air dried, and redissolved with DEPC. The integrity and quantity of dsRNAs were determined by spectrophotometer with Nanodrop 2000 (Thermo Fisher Scientific) and agarose gel electrophoresis44.
Using an IM-31 microinjector (NARISHIGE, Tokyo, Japan), dsGFAT and dsPFK (3000 ng of each) were injected into the abdomen of N. lugens on the 1st day of the 5th instar nymphs. Control groups were injected with dsGFP.
Sample statistics, collection and phenotype observations after injection
After dsRNA was injected into fifth-instar larvae of N. lugens, the malformation rates and mortality rates of N. lugens were counted at 48 h and 72 h, respectively. In addition, insects were randomly collected (excluding abnormal individuals) at 48 h and 72 h after injection to detect the relative expression of chitin metabolism-related genes. Collected samples were stored at -80℃. Photographs of abnormal insects were taken in different dsRNA injection treatments.
Quantification of chitin metabolism-related gene expression levels
N. lugens treated with dsRNA were used to extract the total RNA using the TRIzol reagent (Invitrogen, Carlsbad, California, USA), then first-strand cDNA synthesis was performed using the PrimeScript RT reagent kit with gDNA Eraser (Takara, Kyoto, Japan). Relative expression levels of chitin metabolism-related genes were estimated by qRT-PCR using gene-specific primers (Table 1) with a SYBR Green master mix (SYBR Green Premix Ex Taq, Takara, Japan) in a Bio-Rad CFX96 Real-Time PCR Detection System (Bio-Rad Laboratories Inc., Hercules, CA, USA). The specific steps have been mentioned previously. The 2−△△CT method was used for analyzing relative gene expression59.
Statistical analyses
In this study, all data were analyzed using one-way analysis of variance (ANOVA) and are shown as the mean ± standard error (SE) of three biological replicates. Data on developmental and tissues expression patterns were analyzed using Duncan’s test. In Duncan’s test, different letter indicates a significant difference (p < 0.05). Other data was analyzed using the Tukey’s test. In Tukey’s test, a double asterisk indicates an extremely significant difference in mRNA levels (p < 0.01), and an asterisk indicates a significant difference (p < 0.05).
Data availability
The datasets generated or analysed during the current study are not publicly available but are available from the corresponding author on reasonable request.
References
Xi, Y. et al. Chitinase-like gene family in the brown planthopper Nilaparvata lugens. Insect Mol. Biol. 24, 29–40 (2015).
Merzendorfer, H. & Zimoch, L. Chitin metabolism in insects: structure, function and regulation of chitin synthases and chitinases. J. Exp. Biol. 206, 4393–4412 (2003).
Zhu, K. Y., Merzendorfer, H., Zhang, W., Zhang, J. & Muthukrishnan, S. Biosynthesis, turnover, and functions of chitin in insects. Annu. Rev. Entomol. 61, 177–196 (2016).
Liu, X., Cooper, A. M. W., Zhang, J. & Zhu, K. Y. Biosynthesis, modifications and degradation of chitin in the formation and turnover of peritrophic matrix in insects. J. Insect Physiol. 114, 109–115 (2019).
Merzendorfer, H. Insect chitin synthases: a review. J. Comp. Physiol. B-biochem. Syst. Environ. Physiol. 176, 1–15 (2006).
Tang, B. et al. Suppressing the activity of trehalase with validamycin disrupts the trehalose and chitin biosynthesis pathways in the rice brown planthopper Nilaparvata lugens. Pestic. Biochem. Physiol. 137, 81–90 (2017).
Shukla, E., Thorat, L. J., Nath, B. B. & Gaikwad, S. M. Insect trehalase: physiological significance and potential applications. Glycobiology 25, 357–367 (2015).
Shi, J. F. et al. Physiological roles of trehalose in Leptinotarsa larvae revealed by RNA interference of trehalose-6-phosphate synthase and trehalase genes. Insect Biochem. Mol. Biol. 77, 52–68 (2016).
Kato, N., Mueller, C. R., Wessely, V., Lan, Q. & Christensen, B. M. Mosquito glucosamine-6-phosphate N-acetyltransferase: cDNA, gene structure and enzyme kinetics. Insect Biochem. Mol. Biol. 35, 637–646 (2005).
Cohen, E. Chitin synthesis and inhibition: a revisit. Pest Manag. Sci. 57, 946–950 (2001).
Chen, J. et al. Different functions of the insect soluble and membrane-bound trehalase genes in chitin biosynthesis revealed by RNA interference. PLoS ONE 5, e10133. https://doi.org/10.1371/journal.pone.0010133 (2010).
Chen, X. et al. Disruption of Spodoptera exigua larval development by silencing chitin synthase gene A with RNA interference. Bull. Entomol. Res. 98, 613–619 (2008).
Kramer, K. J. & Koga, D. Insect chitin: physical state, synthesis, degradation and metabolic regulation. Insect Biochem. 16, 851–877 (1986).
Niu, X. et al. Heterologous expression and characterization of a novel chitinase (ChiEn1) from coprinopsis cinerea and its synergism in the degradation of chitin. J. Agric. Food Chem. 65, 6943–6956 (2017).
Qu, M., Ma, L., Chen, P. & Yang, Q. Proteomic analysis of insect molting fluid with a focus on enzymes involved in chitin degradation. J. Proteome Res. 13, 2931–2940 (2014).
Denzel, M. S. & Antebi, A. Hexosamine pathway and (ER) protein quality control. Curr. Opin. Cell Biol. 33, 14–18 (2015).
Oikari, S. et al. Hexosamine biosynthesis in keratinocytes: roles of GFAT and GNPDA enzymes in the maintenance of UDP-GlcNAc content and hyaluronan synthesis. Glycobiology 26, 710–722 (2016).
Oki, T., Yamazaki, K., Kuromitsu, J., Okada, M. & Tanaka, I. cDNA cloning and mapping of a novel subtype of glutamine:fructose-6-phosphate amidotransferase (GFAT2) in human and mouse. Genomics 57, 227–234 (1999).
Ardito, F., Giuliani, M., Perrone, D., Troiano, G. & Lo Muzio, L. The crucial role of protein phosphorylation in cell signaling and its use as targeted therapy (review). Int. J. Mol. Med. 40, 271–280 (2017).
Li, Y. et al. Identification of a novel serine phosphorylation site in human glutamine: fructose-6-phosphate amidotransferase isoform 1. Biochemistry 46, 13163–13169 (2007).
Graack, H. R., Cinque, U. & Kress, H. Functional regulation of glutamine:fructose-6-phosphate aminotransferase 1 (GFAT1) of Drosophila melanogaster in a UDP-N-acetylglucosamine and cAMP-dependent manner. Biochem. J. 360, 401–412 (2001).
Liu, Y., Cai, D. X., Wang, L., Li, J. Z. & Wang, W. N. Glucosamine: fructose-6-phosphate amidotransferase in the white shrimp Litopenaeus vannamei: characterization and regulation under alkaline and cadmium stress. Ecotoxicology 24, 1754–1764 (2015).
Zhou, L. et al. Glutamine-fructose-6-phosphate transaminase 2 (GFPT2) promotes the EMT of serous ovarian cancer by activating the hexosamine biosynthetic pathway to increase the nuclear location of β-catenin. Pathol. Res. Pract. 215, 152681 (2019).
Tran, D. H. et al. Chronic activation of hexosamine biosynthesis in the heart triggers pathological cardiac remodeling. Nat. Commun. 11, 1771 (2020).
Jojima, T. & Inui, M. Engineering the glycolytic pathway: a potential approach for improvement of biocatalyst performance. Bioengineered 6, 328–334 (2015).
Poorman, R. A., Randolph, A., Kemp, R. G. & Heinrikson, R. L. Evolution of phosphofructokinase–gene duplication and creation of new effector sites. Nature 309, 467–469 (1984).
Martínez-Costa, O. H., Hermida, C., Sánchez-Martínez, C., Santamaría, B. & Aragón, J. J. Identification of C-terminal motifs responsible for transmission of inhibition by ATP of mammalian phosphofructokinase, and their contribution to other allosteric effects. Biochem. J. 377, 77–84 (2004).
Nunes, R. D. et al. Unique PFK regulatory property from some mosquito vectors of disease, and from Drosophila melanogaster. Parasites Vectors. 9, 107. https://doi.org/10.1186/s13071-016-1391-y (2016).
Hassan, A., Huang, Q., Xu, H., Wu, J. & Mehmood, N. Silencing of the phosphofructokinase gene impairs glycolysis and causes abnormal locomotion in the subterranean termite Reticulitermes chinensis Snyder. Insect Mol. Biol. https://doi.org/10.1111/imb.12672 (2020).
Zheng, X. et al. Use of banker plant system for sustainable management of the most important insect pest in rice fields in China. Sci. Rep. 7, 45581. https://doi.org/10.1038/srep45581 (2017).
Liu, Q., Hallerman, E., Peng, Y. & Li, Y. Development of Bt rice and Bt maize in China and their efficacy in target pest control. Int. J. Mol. Sci. 17, 1561. https://doi.org/10.3390/ijms17101561 (2016).
Backus, E. A., Serrano, M. S. & Ranger, C. M. Mechanisms of hopperburn: an overview of insect taxonomy, behavior, and physiology. Annu. Rev. Entomol. 50, 125–151 (2005).
Sun, H., Yang, B., Zhang, Y. & Liu, Z. Metabolic resistance in Nilaparvata lugens to etofenprox, a non-ester pyrethroid insecticide. Pestic. Biochem. Physiol. 136, 23–28 (2017).
Ghanbari, F., Moattar, F., Monavari, S. M. & Arjmandi, R. Human health risk assessment of organophosphorus pesticide in rice crop from selected districts of Anzali International Wetland basin, Iran. Hum. Exp. Toxicol. 36, 438–444 (2017).
Gangishetti, U. et al. Effects of benzoylphenylurea on chitin synthesis and orientation in the cuticle of the Drosophila larva. Eur. J. Cell Biol. 88, 167–180 (2009).
Merzendorfer, H. Chitin synthesis inhibitors: old molecules and new developments. Insect Sci. 20, 121–138 (2013).
Tian, X., Zhang, C., Xu, Q., Li, Z. & Shao, X. Azobenzene-benzoylphenylureas as photoswitchable chitin synthesis inhibitors. Org. Biomol. Chem. 15, 3320–3323 (2017).
Wang, Y. et al. Chitin synthase 1 gene and its two alternative splicing variants from two sap-sucking insects, Nilaparvata lugens and Laodelphax striatellus (Hemiptera: Delphacidae). Insect Biochem. Mol. Biol. 42, 637–646 (2012).
Dai, W., Dierschke, S. K., Toro, A. L. & Dennis, M. D. Consumption of a high fat diet promotes protein O-GlcNAcylation in mouse retina via NR4A1-dependent GFAT2 expression. Biochimica et Biophysica Acta-mol. Basis Dis. 1864, 3568–3576 (2018).
Kato, N., Dasgupta, R., Smartt, C. T. & Christensen, B. M. Glucosamine:fructose-6-phosphate aminotransferase: gene characterization, chitin biosynthesis and peritrophic matrix formation in Aedes aegypti. Insect Mol. Biol. 11, 207–216 (2002).
Huang, X. et al. Characterization of glutamine: fructose-6-phosphate aminotransferase from the ixodid tick, Haemaphysalis longicornis, and its critical role in host blood feeding. Int. J. Parasitol. 37, 383–392 (2007).
Kato, N., Mueller, C. R., Fuchs, J. F., Wessely, V. & Lan, Q. Christensen BM, Regulatory mechanisms of chitin biosynthesis and roles of chitin in peritrophic matrix formation in the midgut of adult Aedes aegypti. Insect Biochem. Mol. Biol. 36, 1–9 (2006).
Pesch, Y. Y., Riedel, D. & Behr, M. Drosophila Chitinase 2 is expressed in chitin producing organs for cuticle formation. Arthropod Struct. Dev. 46, 4–12 (2017).
Zhang, L. et al. Study on the effect of wing bud chitin metabolism and its developmental network genes in the brown planthopper, Nilaparvata lugens, by knockdown of TRE gene. Front. Physiol. 8, 750. https://doi.org/10.3390/ijms17101561 (2017).
Bockus, L. B., et al. Cardiac insulin signaling regulates glycolysis through phosphofructokinase 2 content and activity. J. Am. Heart Assoc. 6, e007159, https://doi.org/10.1161/JAHA.117.007159 (2017).
Hu, D. et al. Dynamics and regulation of glycolysis-tricarboxylic acid metabolism in the midgut of Spodoptera litura during metamorphosis. Insect Mol. Biol. 25, 153–162 (2016).
Mookerjee, S. A., Gerencser, A. A., Nicholls, D. G. & Brand, M. D. Quantifying intracellular rates of glycolytic and oxidative ATP production and consumption using extracellular flux measurements. J. Biol. Chem. 292, 7189–7207 (2017).
Van der Horst, D. J. Insect adipokinetic hormones: release and integration of flight energy metabolism. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 136, 217–226 (2003).
Han, H. RNA Interference to knock down gene expression. Methods Mol. Biol. 1706, 293–302 (2018).
Agrawal, N. et al. RNA interference: biology, mechanism, and applications. Microbiol. Mol. Biol. Rev. 67, 657–685 (2003).
Li, T., Chen, J., Fan, X., Chen, W. & Zhang, W. MicroRNA and dsRNA targeting chitin synthase a reveal a great potential for pest management of the hemipteran insect Nilaparvata lugens. Pest Manag. Sci. 73, 1529–1537 (2017).
Yang, M. et al. Knockdown of two trehalose-6-phosphate synthases severely affects chitin metabolism gene expression in the brown planthopper Nilaparvata lugens. Pest Manag. Sci. 73, 206–216 (2017).
Pan, B. Y., et al. Glucose utilization in the regulation of chitin synthesis in brown planthopper. J. Insect Sci. 19, 3, https://doi.org/10.1093/jisesa/iez081 (2019).
Yang, M. et al. miR-71 and miR-263 jointly regulate target genes chitin synthase and chitinase to control locust molting. PLoS Genet. 12, e1006257. https://doi.org/10.1371/journal.pgen.1006257 (2016).
Zhao, L. et al. Functional characterization of three trehalase genes regulating the chitin metabolism pathway in rice brown planthopper using RNA interference. Sci. Rep. 6, 27841. https://doi.org/10.1038/srep27841 (2016).
Gibb, A. A. et al. Integration of flux measurements to resolve changes in anabolic and catabolic metabolism in cardiac myocytes. Biochem. J. 474, 2785–2801 (2017).
Arakane, Y. & Muthukrishnan, S. Insect chitinase and chitinase-like proteins. Cell. Mol. Life Sci. 67, 201–216 (2010).
Zhang, J. et al. Comparative genomic analysis of chitinase and chitinase-like genes in the African malaria mosquito (Anopheles gambiae). PLoS ONE 6, e19899. https://doi.org/10.1371/journal.pone.0019899 (2011).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Acknowledgements
We thank Dr. Qiang Fu (China National Rice Research Institute, 359 Tiyuchang Rd., Hangzhou, Zhejiang, China) and Hong-Xing Xu (Zhejiang Academy of Agricultural Sciences, 198 Shiqiao Rd., Hangzhou, Zhejiang, China) for their kind help. This work was supported by National Natural Science Foundation of China (Grant No. 31672081).
Author information
Authors and Affiliations
Contributions
B.T. and C.-D.X. conceived and designed the work. L.-Y.Q., Y.-K.L., S.-S.W., B.-Y.P. and Y.L. carried out the experiments. L.-Y.Q. and Y.-K.L. performed the analysis. C.-D.X. and Y.-K.L. wrote the manuscript. B.T., L.-Y.Q. and S.-G.W. involved in interpreting data and revising manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xu, CD., Liu, YK., Qiu, LY. et al. GFAT and PFK genes show contrasting regulation of chitin metabolism in Nilaparvata lugens. Sci Rep 11, 5246 (2021). https://doi.org/10.1038/s41598-021-84760-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-84760-2
- Springer Nature Limited