Abstract
Yellow leaf disease caused by sugarcane yellow leaf virus (SCYLV) is one of the most prevalent diseases worldwide. In this study, six near-complete genome sequences of SCYLV were determined to be 5775–5881 bp in length. Phylogenetic analysis revealed that the two SCYLV isolates from Réunion Island, France, and four from China were clustered into REU and CUB genotypes, respectively, based on 50 genomic sequences (this study = 6, GenBank = 44). Meanwhile, all 50 isolates were clustered into three phylogroups (G1–G3). Twelve significant recombinant events occurred in intra- and inter-phylogroups between geographical origins and host crops. Most recombinant hotspots were distributed in coat protein read-through protein (RTD), followed by ORF0 (P0) and ORF1 (P1). High genetic divergences of 12.4% for genomic sequences and 6.0–24.9% for individual genes were determined at nucleotide levels. The highest nucleotide diversity (π) was found in P0, followed by P1 and RdRP. In addition, purifying selection was a main factor restricting variability in SCYLV populations. Infrequent gene flow between Africa and the two subpopulations (Asia and America) were found, whereas frequent gene flow between Asia and America subpopulations was observed. Taken together, our findings facilitate understanding of genetic diversity and evolutionary dynamics of SCYLV.
Similar content being viewed by others
Introduction
Sugarcane yellow leaf virus (SCYLV) is a causal pathogen of yellow leaf disease (YLD) in sugarcane worldwide1,2. The first occurrence of YLD, previously known as yellow leaf syndrome, was described on sugarcane cultivar H 65-7052 in Hawaii in 19883,4. Since then, the presences of SCYLV and sugarcane yellows phytoplasma (SCYP) were confirmed in plants during the 1990s and shared similar symptoms5,6,7,8. To discriminate the two diseases and their corresponding pathogens, Rott et al. proposed the disease caused by SCYLV was YLD, while leaf yellows was another disease resulting from SCYP2. Notably, a mix of SCYLV and SCYP infections has been found in sugarcane 9,10,11. Presently, SCYLV has been reported in more than 25 sugarcane growing countries, representing a major limitation to sugarcane production worldwide12.
SCYLV is phloem-limited and transmitted by various aphids in a semipersistent, circulative, and non-propagative manner, but not by mechanical transmission13. These aphids include Ceratovacuna lanigera, Melanaphis sacchari, Rhopalosiphum rufiabdominalis, and R. maidis13,14,15. Apart from aphid, long-distance transmission between sugarcane cultivation regions and countries is via SCYLV-infected cuttings15,16,17. SCYLV is limited to certain natural host plants, such as sugarcane (Sachharum spp. hybrids and other Sachharum species), species of Erianthus13,14,18, barley [Hordeum vulgare]19, grain sorghum [Sorghum bicolor]20, and Columbus grass [Sorghum almum]21. SCYLV infection has significantly affected the economic traits in sugarcane, such as cane growth, stalk diameter, number of millable canes, and sucrose content, which can lead to 10–50% loss in cane yield and 14% loss in sugar yield in plant or ratoon crops12. Moreover, about 30% yield reduction in asymptomatic sugarcane plants was reported in Thailand22.
In 2004, SCYLV was identified as a species of the Polerovirus genus of the Luteoviridae family23. Currently, 26 assigned species and some unassigned ones are included in the Polerovirus genus, which contains the representative species Potato leafroll virus (https://talk.ictvonline.org/taxonomy/). The genome of SCYLV is monopartite, 24–29 nm in diameter14, and consists of single-stranded, positive-sense, linear RNA (~ 6 kb) that include six open reading frames (ORFs 0–5) and three untranslated regions (UTRs)24,25. Additionally, two subgenomic RNAs (~ 2.4 and ~ 1.0 kb) in SCYLV found in either sugarcane25,26 or grain sorghum27 have been proposed during viral replication. Like other poleroviruses, translation of SCYLV proteins is carried out using a variety of strategies, such as leaky scanning, frameshifts, and read-through, to produce P1 (encoded by ORF1), RNA-dependent RNA polymerase (RdRP; P1–P2 fusion protein), and coat protein (CP) read-through protein (RTD), respectively17.
SCYLV has high genetic diversity among geographical origins and a dozen genotypes present worldwide which have been revealed by phylogenetic analysis based on complete genome sequences, including eight genotypes [BRA, CHN1, CHN3, CUB, HAW, IND, PER, and REU]28 and a few new genotypes, such as FLA1–FLA327,29. Two more SCYLV genotypes (COL in Colombia and CHN2 in China) were proposed based on partial genome fragments30,31. Ancestors of SCYLV evolved through RNA recombination between species of three genera, Luteovirus, Polerovirus, and Enamovirus24,25. Subsequently, numerous studies have shown that some novel SCYLV isolates were regenerated by RNA recombination among worldwide isolates, and potential recombination hotspots were distributed in ORF1/2 and ORF5 regions27,28,32. Overall, viral RNA recombination is known to be an important evolutionary force in the SCYLV genome and represents a formidable challenge in managing YLD.
Breeding new resistant cultivars is critical for successful control of sugarcane diseases33. However, plant resistance is currently limited to one or some specific viral strains and for a short time (a few years) because plant virus is able to rapidly circumvent host acquired genetic resistance17. Usage of healthy seed cane is another conventional and effective management strategy for control of viral disease, including YLD10,34. The present study analyzed the genetic variability, molecular evolution, and population constructs of SCYLV isolates collected from Réunion Island, France, and China along with all published sequences in GenBank. The results of this investigation will enrich current knowledge of SCYLV genomes and understanding of the genetic diversity and evolutionary dynamics of this virus, especially at the global level, providing a vital basis for designing strategies and management schemes for YLD.
Results
SCYLV genome sequencing and assembly
Two nearly complete genomes of SCYLV isolates (REU-YL11 and REU-YL15) from Réunion Island were assembled based on six overlapping genomic fragments. Meanwhile, four nearly complete genomes of SCYLV isolates (ZJWL002, ZJWL003, ZJWL007, and ZJWL012) from Zhejiang Province, China, were assembled according to three overlapping genomic fragments. The six nearly full genome sequences were determined to be 5775–5881 bp in size (Table S1) and have been deposited in the National Coalition Building Institute (NCBI) under accession numbers KY052165 and KY052166, MW439312, MW439313, MW446950, and MW446951, respectively.
Phylogenetic grouping of SCYLV isolates
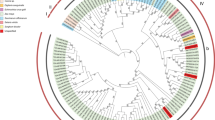
The 5′-UTR and 3′-UTR sequences of all tested SCYLV genomes were trimmed as most isolates lacked these sequences. After that, the nucleotide sequences (ORFs 0–5) of 50 SCYLV isolates, including six sequences obtained in the present study, were subjected to phylogenetic analyses. The Maximum Likelihood (ML) phylogenetic tree revealed a clear segregation of all 50 isolates into three major clades (Fig. 1). Clade I (phylogroup 1, G1) consisted of four genotypes (BRA, CHN3, HAW, and PER), including 22 SCYLV isolates from Brazil, China, India, Peru, and USA. Interestingly, one isolate (PI 157033) in this group originated from sorghum (Sorghum bicolor). Clade II (phylogroup 2, G2) was comprised of a unique REU genotype, including 10 SCYLV isolates from Réunion Island and Mauritius, among which two isolates (REU-YL11 and REU-YL15) determined herein, were clustered in this genotype. Clade III (phylogroup 3, G3) contained the remaining 18 SCYLV isolates from China, Colombia, Cuba, India, and USA, including four isolates (ZJWL002, ZJWL003, ZJWL007, and ZJWL012) obtained in the current study that clustered in the CUB genotype. Apart from CHN1, CUB, and IND genotypes involved in phylogroup G3, two SCYLV isolates (Sorg1_1 and Sorg2_2) from grain sorghum along with one isolate (FL84) from sugarcane were assigned to genotype FLA1/2, while one unique isolate (Sorg3_3) from grain sorghum was proposed to be in a distinct group [FLA3] (Fig. 1).
Maximum Likelihood (ML) phylogenetic tree constructed using IQ-TREE 1.6.12 based on nearly complete genome sequences [open reading frames (ORFs) 0–5] from 50 sugarcane yellow leaf virus (SCYLV) isolates. ML analysis was carried out using nucleotide identity distances and 1000 bootstrap replicates (Bootstrap values > 60% are shown at nodes). The scale bar indicates the number of substitutions per nucleotide. Different geographic regions are represented with red, green, and purple lines for Asia (filled square), Africa (filled circle), and America (filled triangle), respectively. The six SCYLV isolates from sugarcane obtained in the present study are indicated by the solid pentagram (filled star). A potato leafroll virus (PLRV; NCBI accession no. NC_001747) isolate served as an outgroup (filled diamond). The acronyms of each SCYLV genotype based on their originally geographical origins: BRA (Brazil), HAW (Hawaii), PER (Peru), CHN (China), REU (Reunion), FLA (Florida), IND (India), and CUB (Cuba).
Detection of recombination signals
Recombination analysis revealed that 12 clear recombinants were observed among the 50 SCYLV isolates that were under the threshold of at least four of the seven recombination detection algorithms and had a high acceptable P < 0.05 (Table 1). These recombination isolates included GZ-GZ18 and PI 157033 from group G1, MU-AB193 and MU-SC1233 from group G2, and IND2 along with Sorg1_1, Sorg2_2, and Sorg3_3 from phylogroup G3. Seven extremely significant recombination signals were found among isolates that were under the threshold of seven algorithms with a high acceptable P < 0.05. All 12 significant recombination signals were further confirmed by the SimPlot program (Fig. S1). Apart from intra-phylogroups, such as GZ-GZ18 originating from CHN-GD-WY19 (major parent) and FL86 (minor parent), recombination events commonly occurred in inter-phylogroups, such as four isolates (PI 157033, Sorg1_1, Sorg2_2, and Sorg3_3) from grain sorghum clustered in group G3 which were generated from isolates between phylogroups G1 and G3 (Table 1). Meanwhile, recombination events commonly occurred within sugarcane and between sugarcane and grain sorghum. Additionally, while recombination events distributed across the whole genome, most recombination breakpoints (67%) presented at the 3′-terminal of the genome, followed by 25% recombination breakpoints at the 5′-terminal of genome, and only two recombination breakpoints were positioned in the middle region of the genome. These results indicated that SCYLV genomic regions of 3′- and 5′-termini are recombination hotspots.
Sequence identity analysis within and between phylogroups
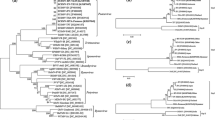
Pairwise sequence identity analysis revealed that nucleotide sequence identities ranged from 83.7 to 99.9% among genome sequences (ORFs 0–5) from the 50 isolates, indicating high sequence diversity between isolates (Table S2). Obvious divergences between phylogroups were found, especially for the comparisons of phylogroup G3 with phylogroups G1 and G2 having more than 12.4% nucleotide variation (Fig. 2a and Table S2). After three SCYLV isolates (Sorg1_1, Sorg2_2, and Sorg3_3) were excluded from the analysis of genetic diversity, ML phylogenetic trees and sequence identities demonstrated significant variation between SCYLV phylogroups based on each gene and corresponding protein from 47 SCYLV isolates (Fig. 2b–g and Table S2). High divergence in nucleotide (nt) and amino acid (aa) sequences of SCYLV genes/proteins between phylogroups were found: 3.0–23.9% (nt) and 2.0–31.7% (aa) for ORF0 (P0), 3.7–20.1% (nt) and 4.0–27.9% (aa) for ORF1 (P1), and 3.4–15.4% (nt) and 3.0–29.9% (aa) for ORF1-2 (RdRP). By contrast, high sequence identities of ORF3 (CP) and ORF4 (MP) between phylogroups were ≥ 94.9% (ORF3) and ≥ 96.2% (ORF4) at nt levels and ≥ 93.3% (CP) and ≥ 95.3% (MP) at aa levels, respectively.
Maximum Likelihood phylogenetic trees (leaf panel) constructed by IQ-TREE v1.6.12 and pairwise identity matrixes (right panel) constructed by SDT v1.2 among SCYLV isolates. (a) Fifty nearly-complete genome sequences of SCYLV. (b)–(g) Six nucleotide sequences of ORF 0, ORF 1, ORF 1–2, ORF 3, ORF 4, and ORF 3–5 from 47 SCYLV isolates (Sorg1_1, Sorg2_2, and Sorg3_3 were excluded), respectively. The acronyms of each SCYLV genotype: BRA (Brazil), HAW (Hawaii), PER (Peru), CHN (China), REU (Reunion), FLA (Florida), IND (India), and CUB (Cuba).
Genetic variation in individual genes among phylogroups
The 47 SCYLV isolates had complete ORFs and corresponding proteins, but slight differences in length in each ORF or protein (Table 2) were present due to 12 insertions/deletions (InDels) events occurring in 18 isolates (Fig. 3). The number of amino acids in these InDels ranged from one to 16 (Fig. 3a). These InDel events were distributed in different proteins: ORF0 encoding P0 (2 InDels), ORF1 encoding P1 (3 InDels), ORF1/2 encoding RdRP (4 InDels, pulsing 3 InDels in ORF1 region), and ORF3/5 encoding RTD (3 InDels). Notably, 15 of 18 isolates shared InDel 12, while the four SCYLV isolates hosted in a chewing cane clone Guangdong Huangpi (ZJWL002, ZJWL003, ZJWL007, and ZJWL012) from China specifically had InDel 10 (Fig. 3b). In addition, based on the sequence identities (nt and aa) and nucleotide diversity (π) in the overall SCYLV population (47 isolates was considered as a whole) or each phylogrouping subpopulation, higher genetic variation is present in the 5′-end of the viral genome, particularly in ORF0 (P0), and the lower genetic variation is present in the middle region of the viral genome, including ORF3 (CP) and ORF4 (MP) (Table 2). Among 47 SCYLV isolates, sequence identities of ORF0 were 91.7% and 88.6% at nt and aa levels, respectively, while nucleotide diversity (π) was 0.10735. By contract, ORF3 had 98.4% (nt) and 98.8% (aa) sequence identities, while nucleotide diversity (π) was 0.02242.
Schematic sketch of 12 Insertion/Deletion (InDel) events calculated by using amino acid sequences of 47 SCYLV isolates. (a) Position of InDel events in each ORF. (b) InDel events are indicated by the red pentagram (filled star), and the same InDel event occurring among SCYLV isolates is shown by the red dotted line. All isolate sequences (from ORF 0 to 5) are indicated by blue diagrams.
Neutrality tests and selection pressure analysis
In neutrality tests, Tajima’s D was used to assess the deviation from neutrality for all mutations among group-specific SCYLV gene sequences. The values of Tajima’s D were negative for CP and MP gene sequences in the overall population and in each phylogrouping subpopulation, indicating that purifying selection is likely acting on SCYLV populations as a result of these populations increasing (Table 2). However, Tajima’s D values were positive for P0, P1, RdRP, and RTD in the overall SCYLV population and for P1 and RdRP in the phylogroup-G2 subpopulation, indicating these populations were stable. However, all Tajima’s D statistics were not statistically significant (P > 0.05) in any populations, suggesting that all populations appear to be at demographic equilibrium.
To evaluate selection pressure on each SCYLV coding region, ratios of non-synonymous (dN) over synonymous (dS) mutation rates were calculated. Negative or purifying selection (dN/dS < 1) enhances the speed of elimination of deleterious mutations in genes and shapes a stable population genetic structure, whereas positive selection (dN/dS > 1) plays an important role in virus adaptability to environmental changes and new hosts. Except for MP, the dN/dS ratio was < 1 for all SCYLV coding regions in most of subpopulations, indicating that purifying selection was a main factor restricting variability in the SCYLV population (Table 2). However, the purifying selective pressure was not distributed uniformly across the SCYLV genome in a specific subpopulation. For example, for genes in the phylogroup-G2 subpopulation, the strongest purifying selection appeared in RdRP (dN/dS = 0.0884), but positive selection was found in MP (dN/dS = 1.3254). The dN/dS > 1 for MP in the phylogroup-G2 subpopulation suggests positive selection is acting on this gene in this subpopulation.
Population differentiation based on geographical origins
To shed light on the genetic relationship between different geographical populations, three statistics (Ks*, Z*, and Snn) were used to evaluate population genetic differentiation. Overall, significantly high values for the three statistics were obtained in all subpopulations based on each gene, indicating that the isolates between geographical populations had a very high genetic differentiation (Table 3). Furthermore, infrequent gene flow between Africa and the two subpopulations (Asia and America) were found, supported by an allele frequency across populations (Fst) > 0.33 (except for Fst > 0.20 in CP and MP) and a migration rate (Nm) < 1 for each gene (Table 3). However, frequent gene flow between Asian and American subpopulations was observed, as evidenced by an Fst < 0.1 (from 0.01581 in RdRP to 0.08389 in MP) and Nm > 1 (from 5.46 in MP and 31.13 in RdRP) for individual genes. Based on these statistics, geographical isolation likely played an important role in the SCYLV subpopulation structure of Africa. On the other hand, the five aforementioned parameters were also calculated for pairwise comparison between the SCYLV phylogrouping subpopulations. These results demonstrated significantly high values of Ks*, Z*, and Snn statistics for each gene between SCYLV phylogrouping subpopulations (except for MP gene in the comparison of G1 versus G2 subpopulations), indicating that the isolates between most phylogrouping subpopulations had a very high genetic differentiation and infrequent gene flow (Fst > 0.33 and Nm < 1; Table S3).
Discussion
YLD is a very common viral disease, and at least six SCYLV genotypes (BRA, CHN1, CHN2, CHN3, CUB, and PER or HAW) occurr in China28,30,31,35. Previous studies showed that BRA genotype is most prevalent in Chinese sugarcane planting areas, which is also highly distributed around the word, but other genotypes just occurred in limited regions28,30,31. The REU and CUB genotypes from Réunion Island and Cuba, respectively, were proposed by Abu et al.38 based on the geographical origins where they were first determined. The SCYLV isolate (CHN-GD-WY20) of the CUB genotype was also present in Guangdong Province, China, but only one isolate is available31. However, information is limited about SCYLV genome sequences of REU and CUB genotypes to date. In the present study, two genomic sequences of REU genotype and four genomic sequences of CUB genotype were obtained which enrich the genomic information available on the two SCYLV genotypes.
Apart from SCYLV genotype CUB occurring in China, this genotype is also distributed in India36 and Mauritius37. It is likely that this virus was transmitted by germplasm exchanges among the countries through vegetative propagation cuttings31,36,38. Furthermore, it is no surprise that the CUB genotype was present in both provinces of China because the sugarcane variety ‘Guangdong Huangpi’ (a chewing cane) was introduced from Guangdong Province where the CUB genotype exists to Zhejiang Province in 2010. Notably, FL180 (MH058009) along with chn1 (GU327735) were clustered in SCYLV genotype CHN1 in the present study, similar to Filloux et al.39, because the two isolates were clustered together in a branch based on each protein. The reason for the close relationship between the two isolates is likely that isolate chn1 hosted in sugarcane cultivar CP93-1309 was developed in Florida, USA, and then exported to China31, while FL180 (MH058009) hosted in cultivar CP00-1101 also originated from Florida39. Three isolates Sorg1_1 (KY960995), Sorg2_2 (KY960996), and Sorg3_3 (KY960997) hosted in grain sorghum in Florida were not identified as a specific genotype27,39, but they were recently proposed as FLA1, LFA2, and FLA3 genotypes, respectively, based on geographical origin29. Therefore, the current results suggest isolate FL84 (MH058007) from sugarcane together with two isolates (Sorg1_1 and Sorg2_2) clustered into a branch be assigned the FLA1/2 genotype, while Sorg3_3 is assigned a distinct FLA3 genotype29. More genomic sequences of SCYLV are needed to further discriminate between FLA1 and FLA2 genotypes.
Recombination is an important driving force in the evolution of poleroviruses, which contributes to genetic diversity in the virus population17,40. Numerous observations have indicated that RNA recombination occurs frequently in SCYLV based on nearly complete genome sequences27,28,32. Similar to observations by ElSayed et al. 27, the current data also demonstrated that recombination events among worldwide SCYLV isolates occurred in inter-phylogroups between geographical locations and crops (sugarcane and grain sorghum). These cases of genomic exchange are likely due to human exchange of sugarcane materials resulting in transmission through traditional geographical barriers between plant viruses28. Recombination events may create a new virus or strain, expand host range, or modify vector specificity40. The present findings further showed that most recombination hotspots were distributed in the SCYLV RTD protein. Notably, RTD protein encoded by poleroviruses is essential for aphid transmission and virus movement in plants17. Therefore, recombination hotspots of SCYLV isolates frequently present in RTD protein may help this virus accomplish jumping from sugarcane to grain sorghum as a host.
Other common sequence variation mechanisms, such as InDel and codon mutations, are also important driving forces in the evolution of plant viruses, including poleroviruses41. Sequence variation in the SCYLV genome may affect the degree of infection capacity, virulence, transmission, and symptoms28,42. The present data revealed the highest sequence divergence in P0 encoded by ORF0, followed by P1 encoded by ORF1. The P0 protein of poleroviruses, including SCYLV encoding a suppressor of RNA silencing, suppresses either local or both local and systemic gene silencing in host plants17,43. This SCYLV P0 protein affects disease symptoms43, and various P0 proteins from different viral genotypes diverge on suppression of host RNA silencing activity44, suggesting the performance of high sequence variability in SCYLV P0 probably contributes to different biological functions for this protein. On the other hand, P1 contains two putative domains, viral protein genome-linked (VPg) and a protease that releases VPg, when P1 is expressed by itself and not fused with P217. Observations from past studies suggest that deletions in ORF1 are associated with the high proliferation rate of the virus in susceptible plants26,30. Recombination together with mutation is an important factor in RNA viruses having a broad host range or using several vector species for transmission28,45.
The current data showed that geographic isolation contributed to shaping the genetic structure of SCYLV populations, particularly between Africa and two other regions (Asia and America), as evidenced by low-level gene flow and significant genetic differentiation between SCYLV geographic populations. This geographical distribution and phylogeny were consistent with our previous study based on P0 and P1 proteins46. Similarly, geographical isolation is responsible for the population structures of maize yellow mosaic virus, which infects sugarcane and maize (Zea mays) and is a novel species in the genus Polerovirus41. Additionally, the present findings showed that the six proteins encoded by SCYLV isolates from individual geographic populations were negative for purification selection, suggesting that this evolutionary driving force may enhance the stability of SCYLV population genetic structure. However, the MP gene of SCYLV from the African population is under positive selection, suggesting it may serve an important role in SCYLV adaptability to locational environment conditions. Other similar investigations have indicated that MP genes from viruses in the family Luteoviridae underwent positive selection pressure, but other genes were under purifying selection41,47.
Overall, the present study examined genetic variability and molecular evolution among SCYLV populations using comparative genomics, improving the current understanding of the genotypic variability of SCYLV populations throughout the world and enriching the taxonomic status of SCYLV isolates. High genetic diversity occurred among these SCYLV isolates. Recombination and InDels contributed to genetic variation during the evolutionary history of SCYLV. Geographical isolation (particularly in Africa) and purifying selection (except for MP gene) are important evolutionary driving forces shaping the SCYLV subpopulation. YLD is commonly managed by resistant cultivars and disease-free seedcane. Quarantine is also important to prevent exotic strains of SCYLV from entering counties where YLD is not present. The present results provide a research basis for transgenic breeding to generate new disease-resistant cultivars, as well as more valuable information for developing efficient molecular detection approaches of the virus during the quarantine stage. The present work also points out that to prevent the spread of YLD, healthy cutting is essential for transferring sugarcane material across borders.
Methods
Leaf sample collection and RNA extraction
Two symptomatic leaves were collected from a local sugarcane clone in Réunion Island in November 2015 while four asymptomatic leaves were collected from a chewing cane cultivar ‘Guangdong Huangpi’ in Wenling, Zhejiang Province, China, in June 2019. Different leaves were collected from different sugarcane plants from the same fields. The collection of leaf samples has been permitted by eRcane, Réunion Island, France. Total RNA was extracted from leaf samples using TRIzol Reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. The final concentration (100 ng/µL) of total RNA was used for SCYLV detection and viral genome cloning.
SCYLV detection by RT-PCR
The SCYLV-specific primers YLS111 and YLS462 designed by Dr. Mike Irey (US Sugar Corporation, Florida, USA) were used for SCYLV detection in the collected leaf samples with RT-PCR30. RT reactions were performed in a 10 µL total volume containing total RNA using a PrimeScript RT Reagent Kit (TaKaRa, Dalian, China) according to the manufacturer’s instructions. PCR reactions were conducted using Ex-Taq PCR Master Mix (TaKaRa Biotech, Dalian, China) and PCR cycling parameters were as follows: an initial denaturation at 94 °C for 2 min, 35 cycles of 94 °C for 30 s, 56 °C for 45 s, and 72 °C for 2 min, with a final 72 °C extension for 10 min.
Genome fragment cloning and assembly
In order to clone and assemble the complete genome sequences of SCYLV isolates, RT-PCR was carried out using five sets of primer pairs designed by Lin et al. 28 for Réunion Island samples and three sets of primer pairs designed herein for Chinese samples (Table S4). RT reactions were conducted with the same protocol used in SCYLV detection. PCR reactions were performed using Ex-Taq PCR Master Mix (TaKaRa Biotech) following the protocol described by Lin et al.28, with minor modification, such as an annealing temperature based on the Tm (°C) of each primer pair (Table S4). PCR-amplified genomic fragments were ligated into a pMD19-T vector (TaKaRa Biotech). Three independent clones for each amplicon were sequenced in both directions by Sangon Biotech Co, Ltd, (Shanghai, China). To remove errors caused by in vitro PCR, a consensus sequence was generated when the three independent clones per isolate showed ≥ 99% identity among three contigs. The full-length genome sequences of six SCYLV isolates (Réunion Island = 2, China = 4) were assembled from several overlapping fragments using DNAMAN 8 (Lynnon Biosoft, San Ramon, CA, USA).
Sequence alignment and phylogenetic analysis
In total, 50 complete genome sequences of SCYLV isolates (present study = 6, GenBank = 44) were used for sequence analysis. These isolates originated from different areas (Asia = 18, Africa = 10, and America = 22; Table S1). Six additional datasets of nucleotide and amino acid sequences from each gene and protein were also analyzed. All datasets were aligned using the ClustalW algorithm in MEGA748. Phylogenetic analysis was carried out using IQ-TREE version 1.6.12 software to construct a ML tree with TIM2e + R5 model and 1000 bootstrap replications49. The genome sequence of potato leafroll virus (GenBank accession no. NC_001747) was used as an outgroup. Meanwhile, pairwise sequence identity analysis was conducted by BioEdit version 7.1.950 and pairwise identity matrix inferred using SDT v1.251.
Test for recombination
Recombination analysis was tested among the 50 complete genome sequences (ORFs 0–5) using RDR4 v 4.6952. Seven different recombination detection algorithms (RDP, GENECONV, Chimaera, MaxChi, Bootscane, SISCAN, and 3Seq) implemented in RDP4 were used with default parameters. Recombination events that were detected by at least four of the seven detection algorithms with significant statistical support and Bonferroni-corrected P value cutoff of 0.05 were considered acceptable28,32. Furthermore, to verify the authenticity of the recombination events, Simplot 3.5.1 software was used to test putative recombination events according to the consistency between the potential recombination isolate and its major and minor parents53.
Calculation of population genetic parameters
Three SCYLV isolates (Sorg1_1, Sorg2_2, and Sorg3_3) from grain sorghum were excluded from the below-described analysis because their complete genes and proteins were not available in the GenBank database. InDel analysis was manually calculated based on the aligned sequences of SCYLV. Nucleotide diversity (π) between SCYLV groups was estimated according to Nei54. Neutrality testing of Tajima’s D55 was performed in DnaSP version 5.10.01 software56. To estimate selection pressure on each SCYLV coding region, the dN/dS ratio was calculated using the HYPhy package57. To assess genetic differentiation between SCYLV populations from different geographical origins (Asia, Africa, and America), three permutation statistical tests (Ks*, Z*, and Snn) were performed with 1000 replicates58,59. If the test statistics were strongly supported by P values < 0.05, the null hypothesis of no genetic differentiation was rejected. Meanwhile, the standardized variance of Fst (Subpopulation fixation index) and Nm (Number of migrants) were used for the degree of gene flow between SCYLV populations. If |Fst|> 0.33 or |Nm|< 1, infrequent gene flow is accepted to have occurred. If |Fst|< 0.33 or |Nm|> 1, frequent gene flow is considerable to have occurred41. All of the population genetic parameters were performed by DnaSP software.
Data availability
The data that support the findings of this manuscript are available from the corresponding author upon reasonable request.
References
Lockhart, B. E. L. & Cronje, C. P. R. Yellow leaf syndrome. In A Guide to Sugarcane Diseases (eds Rott, P. et al.) 291–295 (CIRAD-ISSCT, Montpellier, 2000).
Rott, P., Comstock, J. C., Croft, B. J., Kusalwong, A. & Saumtally, S. Advances and challenges in sugarcane pathology: a review of the 2003 pathology workshop. Proc. Int. Soc. Sugarcane Technol. Congr. 25, 607–614 (2005).
Schenck, S. Yellow Leaf Syndrome: A New Sugarcane Disease 98 (Hawaiian Sugar Planters Association, Honolulu, 1990).
Comstock, J. C., Irvine, J. E. & Miller, J. D. Yellow leaf syndrome appears on the United States mainland. Sugar J. 56, 33–35 (1994).
Lockhart, B. E. L., Irey, M. S. & Comstock, J. C. Sugarcane bacilliform virus, sugarcane mild mosaic virus, and sugarcane yellow leaf syndrome. In Sugarcane Germplasm Conservation and Exchange (eds Croft, B. J. et al.) 108–112 (Australian Centre for International Agricultural Research (ACIAR), Canberra, 1996).
Vega, J., Scagliusi, S. M. M. & Ulian, E. C. Sugarcane yellow leaf disease in Brazil: evidence of association with a luteovirus. Plant Dis. 81, 21–26 (1997).
Cronjé, C. P. R., Tymon, A. M. & Jones, P. Association of a phytoplasma with yellow leaf syndrome of sugarcane in Africa. Ann. Appl. Biol. 133, 177–186 (1998).
Arocha, Y., Gonzalez, L. & Peralta, E. L. First report of virus and phytoplasma pathogens associated with yellow leaf syndrome of sugarcane in Cuba. Plant Dis. 83, 1177 (1999).
Aljanabi, S. M., Parmessur, Y., Moutia, Y., Saumtally, S. & Dookun, A. Further evidence of the association of a phytoplasma and a virus with yellow leaf syndrome in sugarcane. Plant Pathol. 50, 628–636 (2001).
Parmessur, Y., Aljanabi, S., Saumtally, S. & Dookun-Saumtally, A. Sugarcane yellow leaf virus and sugarcane yellows phytoplasma: elimination by tissue culture. Plant Pathol. 51, 561–566 (2002).
Medeiros, L. A. et al. A survey of causal agents associated with sugarcane yellowing in Northeast Brazil. Crop Prot. 138, 105326 (2020).
Holkar, S. K. et al. Present status and future management strategies for Sugarcane yellow leaf virus: a major constraint to the global sugarcane production. Plant Pathol. J. 36, 536–557 (2020).
Schenck, S. & Lehrer, A. T. Factors affecting the transmission and spread of Sugarcane yellow leaf virus. Plant Dis. 84, 1085–1088 (2000).
Scagliusi, S. M. & Lockhart, B. E. L. Transmission, characterization and serology of a luteovirus associated with yellow leaf syndrome of sugarcane. Phytopathology 90, 120–124 (2000).
Li, J. G., Park, J. W., Mirkov, T. E. & Zhou, G. H. A new aphid vector, Ceratovacuna lanigera, of Sugarcane yellow leaf virus in south China. Indian J. Virol. 19, 95 (2008).
Gao, S. J., Lin, Y. H. & Chen, R. K. Advances in researches sugarcane yellow leaf disease and molecular biology of Sugarcane yellow leaf virus. Acta Phytophys. Sin. 39, 177–184 (2012).
Garcia-Ruiz, H., Holste, N. M. & LaTourrette, K. Poleroviruses (Luteoviridae). Reference Module in Life Sciences (Elsevier, Amsterdam, 2020).
Comstock, J. C., Miller, J. D. & Schnell, R. J. Incidence of Sugarcane yellow leaf virus in clones maintained in the world collection of sugarcane and related grasses at the United States National Repository in Miami, Florida. Sugar Tech. 3, 128–133 (2001).
Bouallegue, M., Mezghani-Khemakhem, M., Makni, H. & Makni, M. First report of Sugarcane yellow leaf virus infecting barley in Tunisia. Plant Dis. 98, 1016 (2014).
Wei, C., Hincapie, M., Larsen, N., Nuessly, G. & Rott, P. First report of Sugarcane yellow leaf virus infecting grain sorghum (Sorghum bicolor) in the United States. Plant Dis. 8, 1798 (2016).
Espinoza Delgado, H. V. et al. First report of Sugarcane yellow leaf virus infecting Columbus grass (Sorghum almum) in Florida. Plant Dis. 100, 1027 (2016).
Lehrer, A. T., Kusalwong, A. & Komor, E. High incidence of Sugarcane yellow leaf virus (SCYLV) in sugar plantations and germplasm collections in Thailand. Aust. Plant Dis. 3, 89–92 (2008).
Mayo, M. A. Changes to virus taxonomy 2004. Arch. Virol. 150, 189–198 (2005).
Smith, G. R., Borg, Z., Lockhart, B. E. L., Braithwaite, K. S. & Gibbs, M. J. Sugarcane yellow leaf virus: a novel member of the Luteoviridae that probably arose by inter-species recombination. J Gen. Virol. 81, 1865–1869 (2000).
Moonan, F., Molina, J. & Mirkov, T. E. Sugarcane yellow leaf virus: an emerging virus that has evolved by recombination between luteoviral and poleroviral ancestors. Virology 269, 156–171 (2000).
ElSayed, A. I., Weig, A. R. & Komor, E. Molecular characterization of Hawaiian Sugarcane yellow virus leaf genotypes and their phylogenetic relationship to strains from other sugarcane-growing countries. Eur. J. Plant Pathol. 129, 399–412 (2011).
Elsayed, A. I., Boulila, M., Odero, D. C. & Komor, E. Phylogenetic and recombination analysis of sorghum isolates of Sugarcane yellow leaf virus. Plant Pathol. 67, 221–232 (2018).
Lin, Y. H. et al. Genome characterization of Sugarcane yellow leaf virus from China reveals a novel recombinant genotype. Arch. Virol. 159, 1421–1429 (2014).
Boukari, W. et al. Screening for Sugarcane yellow leaf virus in sorghum in Florida revealed its occurrence in mixed infections with Sugarcane mosaic virus and a new marafivirus. Crop Prot. 139, 105373 (2021).
Gao, S. J. et al. Molecular characterization and phylogenetic analysis of Sugarcane yellow leaf virusisolates from China. Virus Genes 45, 340–349 (2012).
Wang, M. Q., Xu, D. L., Li, R. & Zhou, G. H. Genotype identification and genetic diversity of Sugarcane yellow leaf virus in China. Plant Pathol. 61, 986–993 (2012).
Chinnaraja, C. et al. Complete genome characterization of Sugarcane yellow leaf virus from India: evidence for RNA recombination. Eur. J. Plant Pathol. 135, 335–349 (2013).
Rott, P. C. et al. Controlling sugarcane diseases in Florida: a challenge in constant evolution. Int. Sugar J. 120, 138–142 (2018).
Boukari, W. et al. Field infection of virus-free sugarcane by Sugarcane yellow leaf virus and effect of yellow leaf on sugarcane grown on organic and on mineral soils in Florida. Plant Dis. 103, 2367–2373 (2019).
Lin, Y. H. et al. Molecular detection, genetic diversity and distribution of Sugarcane yellow leaf virus genotypes in China. Trop. Plant Pathol. 40, 200–207 (2015).
Viswanathan, R., Balamuralikrishnan, M. & Karuppaiah, R. Identification of three genotypes of Sugarcane yellow leaf virus causing yellow leaf disease from India and their molecular characterization. Virus Genes 37, 368–379 (2008).
Joomun, N. & Dookun-Saumtally, A. Occurrence of three genotypes of Sugarcane yellow leaf virus in a variety collection in Mauritius. Sugar Tech 12, 312–316 (2010).
Abu Ahmad, Y., Royer, M. & Daugrois, J. H. Geographical distribution of four Sugarcane yellow leaf virus genotypes. Plant Dis. 90, 1156–1560 (2006).
Filloux, D. et al. Viral metagenomic-based screening of sugarcane from Florida reveals occurrence of six sugarcane-infecting viruses and high prevalence of Sugarcane yellow leaf virus. Plant Dis. 102, 2317–2323 (2018).
Dombrovsky, A. et al. The complete genomic sequence of Pepper yellow leaf curl virus (PYLCV) and its implications for our understanding of evolution dynamics in the genus Polerovirus. PLoS ONE 8, e70722 (2013).
Sun, S. R. et al. Genetic variability and molecular evolution of maize yellow mosaic virus populations from different geographic origins. Plant Dis. https://doi.org/10.1094/PDIS-05-20-1013-RE (2020).
Abu Ahmad, Y. et al. Variation in infection capacity and in virulence exists between genotypes of Sugarcane yellow leaf virus. Plant Dis. 91, 253–259 (2007).
Mangwende, T. et al. The P0 gene of Sugarcane yellow leaf virus encodes an RNA silencing suppressor with unique activities. Virology 384, 38–50 (2009).
Lin, Y. H. et al. Molecular characterization and analysis of suppressing RNA silence of P0 protein encoded by Sugarcane yellow leaf virus. Sci. Agric. Sin. 47, 4627–4636 (2014).
Chare, E. R. & Holmes, E. C. A phylogenetic survey of recombination frequency in plant RNA viruses. Arch. Virol. 15, 933–946 (2006).
Zhao, C. H. et al. Population structure and selection pressure analysis among Sugarcane yellow leaf virus isolates based on P0 and P1 sequences. Trop. Plant Pathol. 41, 237–245 (2016).
Pagán, I. & Holmes, E. C. Long-term evolution of the Luteoviridae: time scale and mode of virus speciation. J. Virol. 84, 6177–6187 (2010).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. 33, 1870–1874 (2016).
Nguyen, L. D., Schmidt, H. A., Haeseler, A. V. & Minh, B. Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Hall, T. A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl. Acids Symp. Ser. 41, 95–98 (1999).
Muhire, B. M., Varsani, A. & Martin, D. P. SDT: a virus classification tool based on pairwise sequence alignment and identity calculation. PLoS ONE 9, e108277 (2014).
Martin, D. P., Murrell, B., Golden, M., Khoosal, A. & Muhire, B. RDP4: Detection and analysis of recombination patterns in virus genomes. Virus Evol. 1, vev003 (2015).
Lole, K. S. et al. Full-length human immunodeficiency virus type 1 genomes from subtype C-infected seroconverters in India, with evidence of intersubtype recombination. J. Virol. 73, 152–160 (1999).
Nei, M. Molecular Evolutionary Genetics (Columbia University Press, 1987).
Tajima, F. The effect of change in population size on DNA polymorphism. Genetics 123, 597–601 (1989).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Pond, S. L. K. & Muse, S. V. HyPhy: hypothesis testing using phylogenies. In Statistical Methods in Molecular Evolution (ed. Nielsen, R.) 125–181 (Springer, New York, 2005).
Hudson, R. R. A new statistic for detecting genetic differentiation. Genetics 155, 2011–2014 (2000).
Hudson, R. R., Boos, D. D. & Kaplan, N. L. A statistical test for detecting geographic subdivision. Mol. Biol. Evol. 9, 138–151 (1992).
Acknowledgements
This work was supported by the Earmarked Fund for the China Agricultural Research System (Grant No. CARS-170302), the Special Fund for Science and Technology Innovation in Fujian Agriculture and Forestry University, China (Grant No. CXZX2017348), and the project of Guizhou International Science and Technology Cooperation Base for Modern Agriculture, China (Grant No. 2019-5804).
Author information
Authors and Affiliations
Contributions
J.-J.L., E.-R.H., and W.-Q.B. performed the experiments, analyzed the data, and wrote a first draft of the manuscript. J.-S.C. and S.-R.S. participated in the experimental design and analyzed the data. S.-J.G. conceived and supervised the project, and critically reviewed the manuscript. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lu, JJ., He, EQ., Bao, WQ. et al. Comparative genomics reveals insights into genetic variability and molecular evolution among sugarcane yellow leaf virus populations. Sci Rep 11, 7149 (2021). https://doi.org/10.1038/s41598-021-86472-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-86472-z
- Springer Nature Limited