Abstract
Burkholderia pseudomallei (Bp) is the causative agent of melioidosis, a disease endemic to the tropics. Melioidosis manifests in various ways ranging from acute skin lesions to pneumonia and, in rare cases, infection of the central nervous system. Bp is a facultative intracellular pathogen and it can infect various cell types. The Bp intracellular lifecycle has been partially elucidated and is highly complex. Herein, we have identified a transcriptional regulator, BP1026B_II1198, that is differentially expressed as Bp transits through host cells. A deletion mutant of BP1026B_II1198 was attenuated in RAW264.7 cell and BALB/c mouse infection. To further characterize the function of this transcriptional regulator, we endeavored to determine the regulon of BP1026B_II1198. RNA-seq analysis showed the global picture of genes regulated while ChIP-seq analysis identified two specific BP1026B_II1198 binding regions on chromosome II. We investigated the transposon mutants of these genes controlled by BP1026B_II1198 and confirmed that these genes contribute to pathogenesis in RAW264.7 murine macrophage cells. Taken together, the data presented here shed light on the regulon of BP1026B_II1198 and its role during intracellular infection and highlights an integral portion of the highly complex regulation network of Bp during host infection.
Similar content being viewed by others
Introduction
Bacteria move through various niches responding and reacting to each specific environment they encounter. For this to occur, bacteria must transduce environmental cues through complex signaling pathways that lead to transcription of specific factors required for survival in each situation. These networks are highly complex and require the coordination of many components. Coordinated response to environmental signals often require transcription factors that can modulate RNA polymerase binding to a promoter region, leading to the activation or repression of an operon or gene1. Binding of transcription factors to specific DNA sequences is often controlled through interactions with small ligands, other proteins, and/or by covalent modification1,2,3. Control of transcription factors can be mediated through several mechanisms including turnover, sequestration, and synthesis that can also be coordinated by other transcription factors1. Additionally, a transcription factor can bind to several promoter regions, which in turn can be controlled by several transcription factors on a local and/or global scale1. Taken together, this highlights the highly complex system that bacteria use to coordinate a transcriptional response to each environment they confront. Pathogens, in particular, encounter many environments during the course of an infection, requiring them to have a well-coordinated regulation system in place.
Burkholderia pseudomallei (Bp) is a facultative intracellular pathogen that causes the disease melioidosis. Bp is endemic in tropical regions with increasing prevalence due to better diagnostics and rising awareness, having a predicted global mortality rate of 54 percent4. Bp encounters many different environments as it infects most tissues of the human body, leading to the formation of localized abscesses, bacteremia, septic shock, and sometimes death5,6. While infecting a cell, Bp has a dynamic lifecycle that can be broken down into multiple stages: attachment to the host cell, host cell entry, vacuole escape, cytoplasmic replication, and protrusion towards neighboring cells culminating with the spread of infection7. In the vacuole, the Burkholderia secretion apparatus (T3SSBsa), a type III secretion system, must be expressed to allow entry into the host cell cytoplasm8. Within the cytoplasm, Bp can use its secondary flagella or polymerize host cell actin using bacterial expressed BimA to move freely8,9. Eventually, Bp protrudes towards neighboring cells and uses a virulence-associated type VI secretion system (T6SS-5) to fuse host cell membranes, forming multinucleated giant cells (MNGCs)10,11,12,13. Although we understand the general flow of the Bp intracellular lifecycle, there is much more we do not know about how this pathogen transits through a host and causes the many forms of the disease we cumulatively call melioidosis.
To cause such a wide array of clinical manifestations and be able to transit through the intracellular environment with such precision, Bp requires many virulence factors, some of which are described above. As recently demonstrated, Bp goes through drastic transcriptional changes, with 1,953 genes showing unique patterns of expression during the intracellular infection process7. This finding suggests that Bp has a highly complex regulatory network that is required to coordinate the differential expression of these genes during the progression of infection, from the initial stage of attachment to eventual protrusion towards neighboring cells. The regulation of Bp virulence mechanisms during intracellular infection has been studied to a small extent and mainly focuses on the control of T3SSBsa, T6SS-5, and BimA. The transcriptional regulator BprP regulates expression of the structural components of the T3SSBsa and another transcriptional regulator BsaN14,15. BsaN with its chaperone, BicA, regulate expression of type III effectors and accessory proteins, the two-component sensor-regulator VirAG that is responsible for downstream expression of T6SS-5, BimA, and BprC, a transcriptional regulator essential for T6SS-5 expression14,15,16. Beyond the specific control of characterized virulence pathways, a LysR-type transcriptional regulator GvmR (globally acting virulence and metabolism regulator) was identified to control a broad network of genes involved in virulence and metabolism, highlighting the tie between expression of virulence factors and nutrient limitation within the host17. This study advances the understanding of Bp virulence regulation by describing a new transcriptional regulator that is differentially expressed throughout the intracellular lifecycle and required for complete pathogenesis.
Results
BP1026B_II1198 is differentially regulated as Bp transits through the host cell
To understand how intracellular pathogenesis of Bp is controlled, we first have to identify transcriptional regulators that control virulence mechanisms during intracellular infection. We identified BP1026B_II1198, a PadR-like transcriptional regulator that is differentially regulated during the Bp intracellular ‘transitome’7. BP1026B_II1198 shows similarities to PadR transcriptional regulators with an N-terminal winged helix-like DNA binding domain (Pfam: PF03551) and a C-terminal virulence activator alpha domain (Pfam: PF10400)18,19. BP1026B_II1198 is expressed while Bp is initially internalized within endocytic vesicles and later during protrusion as Bp spreads toward neighboring cells (Fig. 1a). In contrast, BP1026B_II1198 is significantly down-regulated while Bp replicates within the cytoplasm (Fig. 1a). This suggested that BP1026B_II1198 could play a regulatory role during the Bp intracellular lifecycle.
BP1026B_II1198 is differentially regulated during cell infection and critical for intracellular survival. (a) Single cell ‘transitome’ analysis7 shows that BP1026B_II1198 is up-regulated in the vacuole and protrusion while down-regulated in the cytoplasm during infection of RAW264.7 cells. Boxes represent three biological replicates and an average (Ave)7. (b) A mutant in BP1026B_II1198 shows a reduced ability to form wildtype level MNGCs. When the mutant is complemented it recovers beyond wildtype level of MNGC size. Individual MNGC diameters are shown as dots. (c) The BP1026B_II1198 mutant shows a significant reduction in intracellular replication at 8 and 24 h after infection compared to wildtype Bp 1026b. Complementation of the BP1026B_II1198 mutant recovers the intracellular replication to wildtype levels. (d) The BP1026B_II1198 mutant and complement grow to similar levels of wildtype Bp 1026b in vitro. Data in bar graphs and growth curve represent means ± s.e.m and analyzed via unpaired t-test. P values presented are as follows: ** p < 0.01, ****p < 0.0001.
A BP1026B_II1198 mutant is defective in MNGC formation and intracellular replication
To further investigate the role of BP1026B_II1198, a deletion mutant was created using modified knockout-recombineering20. This mutant was used to infect RAW264.7 cells to monitor MNGC formation and intracellular replication. The BP1026B_II1198 mutant strain showed a significant decrease in its ability to form MNGCs, marked by a 41% decrease in average MNGC diameter at 24 h post infection (hpi) (Fig. 1b). Complementation of the BP1026B_II1198 mutant restored this mutation beyond wildtype levels (Fig. 1b). RAW264.7 cells infected with the BP1026B_II1198 mutant also showed a significant decrease in intracellular replication at 8 and 24 hpi further suggesting that this transcriptional regulator is important for intracellular pathogenesis (Fig. 1c). Complementation of the BP1026B_II1198 mutant nullified this defect allowing wildtype levels of intracellular replication (Fig. 1c). The BP1026B_II1198 mutant showed a seven percent reduction in growth compared to wildtype Bp in vitro (Fig. 1d). This slight decrease in growth indicates that the reductions in MNGC formation and intracellular replication are likely due to defects in pathogenesis rather than in vitro fitness.
BP1026B_II1198 is required for complete pathogenesis in vivo
Beyond cell culture infection, we wanted to assess the importance of BP1026B_II1198 during an in vivo infection. BALB/c mice were infected via the intranasal route with a lethal dose (4,500 CFU/mouse) of wildtype Bp 1026b and an equivalent dose of the BP1026B_II1198 mutant strain. Within four days of the initiation of infection, all mice infected with wildtype Bp 1026b showed significant signs of acute murine melioidosis resulting in 100% lethality (Fig. 2a). In contrast, mice infected with the BP1026B_II1198 mutant survived for the duration of the study (62 days) indicating that this gene is important for in vivo pathogenesis of Bp (Fig. 2a). Even though mice challenged with the BP1026B_II1198 mutant showed no clinical symptoms at day 62 post infection, there was persistent infection in the lungs and spleens of the majority of the mice and one mouse showing high levels of infection within the liver (Fig. 2b). This level of persistence demonstrates a shift in the behavior of Bp in the absence of the BP1026B_II1198 gene. This result indicates that BP1026B_II1198 is important for in vivo Bp pathogenesis.
BP1026B_II1198 is attenuated in the BALB/c mouse model. (a) Mice infected with wildtype Bp 1026b (n = 5) did not survive beyond day four post challenge while mice infected with the BP1026B_II1198 mutant (n = 5) were able to survive for the duration of the study (62 days). Comparison of both experimental groups by Log-rank (Mantel-Cox) test shows a significant difference (P = 0.0031) in survival. Mice were challenged with a lethal dose (4500 CFU/mouse) via the intranasal route. (b) Lungs, livers, and spleens of the surviving mice were harvested, homogenized, and plated revealing persistent infection within the lungs of four mice, liver of one mouse, and spleen of three mice. No detectable bacteria were found in the lungs of one mouse, livers of four mice, and spleens of two mice (circles along x-axis).
BP1026B_II1198 regulates hypothetical proteins, secondary metabolite biosynthesis, extracellular matrix, efflux, and type VI secretion
To better understand how BP1026B_II1198 controls Bp pathogenesis, we expressed BP1026B_II1198 in trans and compared the mRNA profile to a control strain with an empty vector control via RNA-seq analysis. Samples were analyzed using Rockhopper21 identifying differential regulation of a subset of Bp genes confirming that BP1026B_II1198 is a transcriptional regulator (Fig. 3b). Genes with a q-value < 0.01 and a log2 fold-change (FC) ≥ 1 or ≤ -1 were selected for further analysis (Fig. 3b, Table S1). There were 88 genes that were up-regulated (log2FC ≥ 1) in the presence of BP1026B_II1198 while 94 genes were down-regulated (log2FC ≤ -1) (Fig. 3). Genes with a log2FC ≥ 1 or ≤ -1 were analyzed by COG functional predictions revealing an overabundance (57.1%) of genes annotated as encoding hypothetical proteins, hypothetical RNA transcripts, or uncharacterized proteins that are controlled by BP1026B_II1198 (Fig. 3a). Of these proteins of unknown function, 54 genes are up-regulated and 50 genes down-regulated by BP1026B_II1198 (Fig. 3a). While this analysis shows the individual genes that are controlled, it is valuable to identify larger regions of the genome that are controlled by BP1026B_II1198 to gain another perspective on its transcriptional landscape. When we analyze the entire RNA-seq data set (Fig. 3b) using WoPPER22, some additional pathways that are responsible for secondary metabolite biosynthesis, components of the extracellular polymeric matrix, and multidrug efflux appear to be controlled by BP1026B_II1198 (Figs. S1 and S2).
Hypothetical proteins are disproportionately controlled by BP1026B_II1198. (a) Genes with a q-value < 0.01 and a log2FC ≤ -1 or ≥ 1 were analyzed by COG functional predictions. There are 88 genes that are up-regulated and 94 genes that are down-regulated by BP1026B_II1198. Hypothetical proteins, hypothetical RNA transcripts and uncharacterized proteins represent 79.8% of the genes controlled by BP1026B_II1198. Of these unknown proteins 68% are up-regulated and 32% are down-regulated. (b) Scatter plot showing distribution of all genes present in RNA-seq analysis. Genes with a q-value < 0.01 and a log2FC ≤ -1 or ≥ 1 are highlighted in red.
Two nonribosomal peptide synthetase regions that produce the siderophores malleobactin23 and pyochelin24 are up-regulated by BP1026B_II1198 in cluster 12 (Fig. S1) and cluster 7 (Fig. S2), respectively, suggesting the involvement of this regulator in iron acquisition during infection. BP1026B_II1198 also up-regulates the polyketide synthase cluster producing another siderophore, malleilactone (Cluster 2, Fig. S2). Malleilactone is involved in the pathogenesis of Caenorhabditis elegans and Dictyostelium discoideum25. Beyond controlling secondary metabolite pathways, BP1026B_II1198 also down-regulates components of the extracellular polymeric matrix tied to contact dependent growth inhibition, BcpA (Cluster 18, Fig. S2)26,27,28. The genes encoding the RND efflux pump, BpeEF-OprC, responsible for multidrug efflux and clinical resistance to antimicrobials29,30, are up-regulated in the presence of BP1026B_II1198 (Cluster 3, Fig. S2). Mutation of the bpeEF-oprC efflux pump in clinical and environmental isolates of Bp showed increased susceptibility to trimethoprim, a component of trimethoprim-sulfamethoxazole that is used in oral eradication phase antimicrobial therapy of melioidosis30. Although we see low levels of up-regulation of bpeEF-oprC, we suspect that the up-regulation is indirect and could involve other regulatory factors. BP1026B_II1198 also up-regulates a region of the genome that includes the type VI secretion system (T6SS-5) that is required for complete pathogenesis and allows intercellular spread and MNGC formation (Cluster 15, Fig. S2)8,11,31. This supports the observation that the BP1026B_II1198 mutant showed reduced MNGC formation in RAW264.7 cells (Fig. 1b). The regulation of known pathways and the numerous hypothetical proteins indicates that the BP1026B_II1198 regulation network is tied to predicted and confirmed pathogenic processes, emphasizing the role of BP1026B_II1198 during pathogenesis.
Specific genes controlled by BP1026B_II1198 have potential role during pathogenesis
To further our understanding of this regulation network, we investigated individual genes that are highly regulated by BP1026B_II1198 with a log2FC ≥ 2 or ≤ -2 (Fig. 4a). BP1026B_I0955 and BP1026B_I0956 are significantly up-regulated by BP1026B_II1198 with log2FCs of 2.16 and 2.03, respectively (Fig. 4a). These genes encode putative SufC and SufB subunits of the SufBCD complex responsible for the assembly of iron-sulfur (Fe-S) clusters in proteins32. Genes encoding NifU and SufD are additional components of this Fe-S cluster assembly pathway that are also up-regulated to log2FCs of 1.06 and 1.58 (Table S1). Another redox related protein, BP1026B_II1579, is up-regulated to a log2FC of 2 and encodes a SCO1/SenC family protein predicted to be involved in the assembly of cytochrome c oxidase (Fig. 4a). Interestingly, BP1026B_II1198 also up-regulates a putative nitrate transporter, BP1026B_I1020 (narK-1), with a log2FC of 2.09, and a potential Burkholderia specific nitric oxide forming nitrite reductase, BP1026B_II1580 (nirK), with a log2FC of 2.99 (Fig. 4a)33. Other components involved in nitrogen metabolism, including narG and narK-2, are also up-regulated (log2FC 1.93 and 1.42) by BP1026B_II1198 (Table S1). The regulation of these pathways by BP1026B_II1198 suggests that they play a role in Bp intracellular survival and virulence similar to how nitrate transport/reduction and nitrite reduction have been tied to virulence in Pseudomonas aeruginosa34,35. In addition to the control of metabolic pathways, BP1026B_II1198 also up-regulates BP1026B_II1736, a type III secretion protein HrpB4 (Fig. 4a).
Genes highly regulated by BP1026B_II1198 show involvement during Bp intracellular survival. (a) Normalized transcripts of genes showing a log2 FC ≤ -2 or ≥ 2 in the presence of BP1026B_II1198 compared to an empty vector control. (b) Predicted functions of genes controlled by BP1026B_II1198. Percent of wildtype replication (%WT) in RAW264.7 cells is shown 24 hpi. Several transposon mutants were not available in the sequence-defined transposon mutant library of Bp 1026b (NA). (c) Transposon mutants present in the sequence defined transposon mutant library of Bp 1026b were analyzed for the ability to replicate in RAW264.7 murine macrophages for 24 h. Intracellular replication represented as a percent of wildtype 24 hpi and the average of triplicates represented in (b). (d) Schematic diagram showing genes involved in Bp intracellular survival (replicating < 80% wildtype) by indirect control of BP1026B_II1198. Data in bar graphs represent means ± s.e.m and analyzed via unpaired t-test. P values presented are as follows: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Many genes that showed large alterations in expression by BP1026B_II1198 are annotated as hypothetical proteins or have no known function (Fig. 3a). BP1026B_I1775 is predicted to be a lipoprotein that shows high amino acid identity to CpaD, a putative pilus assembly protein, in environmental bacteria and is up-regulated by BP1026B_II1198 (Fig. 4a). A GntR family transcriptional regulator, BP1026B_II1506 is also up-regulated and could cause downstream changes in gene expression leading to variations in pathogenesis and/or intracellular survival (Fig. 4a). BP1026B_I0528, BP1026B_I0950, BP1026B_II1464, BP1026B_II1581, and BP1026B_II1769 are hypothetical proteins that are up-regulated by BP1026B_II1198 while BP1026B_II0722 is a hypothetical protein that is down-regulated (Fig. 4a).
Mutant analysis identifies potential virulence-associated genes controlled by BP1026B_II1198
To test the role these genes play in pathogenesis, we used mutants present in the sequence-defined transposon mutant library of Bp 1026b for intracellular replication analysis in RAW264.7 cells36. Genes with normalized transcripts showing a log2 FC ≤ -2 or ≥ 2 in the presence of BP1026B_II1198 compared to the empty vector control were selected for analysis (Fig. 4a). Four of these genes, BP1026B_I0528, BP1026B_I0950, BP1026B_II1464, and BP1026B_II1769, had no mutants in the library and were not analyzed. Monolayers of RAW264.7 cells were infected with each transposon mutant present in the library and the level of intracellular replication determined at 24 h post infection (hpi). Transposon mutants of BP1026B_II0722 and BP1026B_II1580 replicate to wildtype levels (99–100%) after 24 h of infection indicating that these genes have no singular pathogenic function (Fig. 4b-c). In contrast, transposon mutants in BP1026B_II0956, BP1026B_I1020, BP1026B_II1581, and BP1026B_II1736 replicate between 74.9% and 82.6% of wildtype implying these genes have a small role in pathogenesis (Fig. 4b,c). BP1026B_II1736 is part of T3SS-2 that has been previously tied to virulence in S. lycopersicum but the T3SS-2 mutant showed wildtype levels of cytotoxicity in THP-1 cells37. This indicates that BP1026B_II1736 and the T3SS-2 cluster could be involved in intracellular survival but does not contribute to cytotoxicity. Mutants of BP1026B_I0956 and BP1026B_I1020, components of the SUF pathway and nitrate transport/reduction, exhibited decreased pathogenesis indicating that these genes and pathways are important for Bp intracellular survival (Fig. 4b,c). Transposon mutants of BP1026B_I0955, BP1026B_I1775, and BP1026B_II1506 showed the greatest decrease in intracellular survival, replicating at 61.7%, 59.3%, and 63.2% of wildtype, respectively (Fig. 4b,c). Finally, the SCO1/SenC family protein, BP1026B_II1579, replicates at 120.65% of wildtype indicating that protein is involved in reducing the intracellular survival of wildtype Bp within host cells (Fig. 4b,c). Transposon mutants showing a major decrease in intracellular replication were able to replicate at wildtype levels when grown in vitro indicating that the defects identified during RAW264.7 cell infection are not due to decreased fitness (Fig. S3a). This data suggests that BP1026B_II1198 controls several factors that contribute to intracellular pathogenesis to varying degrees (Fig. 4d).
BP1026B_II1198 directly binds to two intergenic regions on chromosome II
BP1026B_II1198 regulates numerous pathways/genes previously identified to be involved in virulence and many potential pathways/genes that have unknown functions during pathogenesis (Fig. 4, Fig. S1, Fig. S2). To determine the direct regulation network of BP1026B_II1198, we used a ChIP-seq strategy as previously described2,3 but modified for Burkholderia species. All peaks identified were plotted by genome position on each Bp chromosome revealing two major peaks located on chromosome II (Fig. 5a,b). When the –log10 of the q-value was compared to the fold-enrichment of each peak, we see the same two peaks further isolated from the background (Fig. 5c). These two peaks suggest that BP1026B_II1198 binds to two intergenic regions between divergently transcribed genes (Fig. 6a). Peak 1 is a 227 bp sequence in the intergenic region between levanase, levansucrase precursor, BP1026B_II0599-0600, and a LacI family regulator protein, BP1026B_II0601 (Fig. 6a). Peak 2 is a 508 bp region between BP1026B_II1006, a DNA-binding protein, and BP1026B_II1007, an oxidoreductase (Fig. 6a). By comparing the sequences of each of these peaks using MEME38, a 12 bp conserved region was identified (Fig. 5d). The sequence, AAAAGTAXXTTA, has seven and three conserved base pairs separated by a two base pair spacer and is oriented in the same direction on both peaks suggesting that transcriptional control from each region will be in the same direction. Emphasizing this point, the RNA-seq data shows that BP1026B_II1198 regulates genes in the same orientation by positively regulating BP1026B_II0600 and negatively regulating BP1026B_II1006 while showing no transcriptional control of genes in the opposite orientation, BP1026B_II0601 and BP1026B_II1007 (Fig. 6b,c).
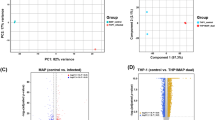
ChIP-seq identifies two regions of BP1026B_II1198 binding on chromosome II. Peaks identified using MACS276 by comparison to an empty vector negative control are plotted by peak height and genome position on chromosome I (a) and chromosome II (b). (b) Two peaks were identified on chromosome II and are labeled as peak 1 in blue and peak 2 in red. (c) The two identified peaks are further isolated from background when compared by statistical (-log10 q-value) and fold enrichment. Blue box shows peak 1 and red box shows peak 2. (d) Potential binding motif identified by analysis of peak 1 and peak 2 nucleotide sequences by MEME38. Each peak contains the same orientation of the potential binding motif.
BP1026B_II1198 has direct transcriptional control of genes involved in pathogenesis of RAW264.7 murine macrophages. (a) Schematic representation of the direct regulation network of BP1026B_II1198. BP1026B_II1198 binds to an intergenic region (peak 1) between BP1026B_II0600 and BP1026B_II0601 and an intergenic region (peak 2) between BP1026B_II1006 and BP1026B_II1007. (b) Normalized transcripts of genes surrounding peak 1 identifies that BP1026B_II1198 positively regulates BP1026B_II0600. (c) Normalized transcripts of genes around peak 2 identify that BP1026B_II1198 negatively regulates BP1026B_II1006. (d) Infection of RAW264.7 cells by transposon mutants of BP1026B_II0600 and BP1026B_II1006 compared to wildtype Bp 1026b showing various levels of intracellular replication defect. (e) Annotated function of BP1026B_II0600 and BP1026B_II1006 and corresponding intracellular replication when compared to wildtype Bp 1026b (%WT). (f) Model for the direct regulation network of BP1026B_II1198. Data in bar graphs represent means ± s.e.m and analyzed via unpaired t-test. P values presented are as follows: *p < 0.05.
Genes directly controlled by BP1026B_II1198 are important during intracellular infection
To investigate the roles BP1026B_II0600 and BP1026B_II1006 play during intracellular pathogenesis, a RAW264.7 macrophage intracellular replication assay was carried out. At 24 hpi a transposon mutant of BP1026B_II0600, a levansucrase precursor that is up-regulated by BP1026B_II1198, replicated intracellularly to 51.62% of wildtype Bp 1026b (Fig. 6d,e). This is a drastic decrease indicating that BP1026B_II0600 has a role during intracellular pathogenesis. While levansucrase is used as a counter-selection marker in the manipulation of the Bp genome, induction of BP1026B_II1198 in the presence of sucrose did not show an additional decrease in growth (Fig. S3c). On the other hand, a transposon mutant of BP1026B_II1006, a DNA-binding protein that is down-regulated by BP1026B_II1198, showed 82.89% wildtype intracellular replication at 24 hpi (Fig. 6d,e). Although it has a small decrease in the intracellular replication compared to wildtype, this decrease could be significant considering that BP1026B_II1006 is a DNA-binding protein and could have downstream transcriptional effects on other pathways like the ones identified by RNA-seq analysis. These data show that BP1026B_II1198 has a direct affect on the positive transcriptional control of BP1026B_II0600 and the negative transcriptional control of BP1026B_II1006 leading to variations in the pathogenesis during RAW264.7 murine macrophage infection (Fig. 6f).
Discussion
Bacterial transcriptional networks are highly complex and dependent on many variables to properly function. When bacteria are exposed to a unique environment they sense and respond through signaling pathways that lead to the expression of proteins/factors aiding in survival. This is especially true during infection, as pathogenic bacteria encounter various immune defenses within the host. Pseudomonas aeruginosa (Pa) is a model organism for the study of biofilms and regulation of virulence factor production, which are controlled by a complex network of transcription factors and signaling systems39. Only recently are the subtleties of the Pa transcription factor regulation network coming to light, highlighted by significant functional crosstalk between transcription factors and regulation systems40,41. Mycobacterium tuberculosis also has a well-documented, highly complex, regulatory network that it uses to navigate hypoxic environments42. To fully understand the pathogenic processes that allow for Bp host colonization and infection we must understand how these pathways are regulated. Some studies have started to shed light on the intricacies of Bp virulence regulation14,17,43,44. In the present study, we add to the knowledge of the Bp regulatory network during intracellular infection through the identification and characterization of BP1026B_II1198.
BP1026B_II1198 is annotated as a PadR-like transcriptional regulator. PadR regulators were first identified in Pediococcus pentosaceus and later characterized to act as repressors of phenolic acid decarboxylase, encoded by the padA gene in Lactobacillus plantarum45,46. Other PadR-like regulators have been identified in other survival processes including drug resistance, stress responses, biosynthesis of antibiotics, and carbon catabolism47,48,49,50,51,52,53. The Pfam database indicates that there are over 19,000 sequences containing a PadR-like domain and over 2,900 sequences representing the domain architecture of BP1026B_II1198 with the N-terminal PadR domain and a C-terminal virulence activator alpha domain19. PadR domains have also been shown to be in combination with many other domains with a total of 35 domain architectures identified in the Pfam database19. Bp does not contain a padA homolog or a phenolic acid metabolic pathway, and the varied nature of PadR-like proteins suggests broad and diverse functionalities. We therefore took a broad approach to study BP1026B_II1198.
BP1026B_II1198 is differentially expressed during the Bp intracellular lifecycle and is required for in vitro and in vivo pathogenesis, highlighted by the 100% attenuation of the BP1026B_II1198 mutant in the BALB/c mouse model. These two observations show that Bp precisely controls pathogenic function within the host. To further understand this contribution to pathogenesis we identified the genes controlled by BP1026B_II1198 using RNA-seq. We observed that 79.8% of genes differentially regulated in the presence of BP1026B_II1198 have no known functions. This is a striking increase from the 26.7% of genes annotated as hypothetical unknowns within the Bp 1026b genome, suggesting that many pathogenic and survival pathways are yet to be discovered. The presence of BP1026B_II1198 during the later stages of infection coincides with the regional expression around the T6SS-5 locus, aligning with the current understanding of the Bp lifecycle. In addition, expression of BP1026B_II1198 leads to the up-regulation of several secondary metabolite gene clusters malleobactin23, pyochelin24, and malleilactone25, that have been previously identified to be involved in iron acquisition and pathogenesis. The regulation of these known virulence mechanisms likely contribute to the attenuation of the BP1026B_II1198 mutant, although we speculate that there are other factors that are also involved.
We probed many genes that were regulated by BP1026B_II1198 to identify contributors to pathogenesis, including genes that may have a less direct pathogenic function. Although Fe-S clusters were first identified in 1962 to be involved during electron transport54, their function is diverse and has been implicated in the modulation of gene expression55. Fe-S clusters can respond to increased oxidative stress, iron levels within the host, or other environmental signals during infection leading to an adaptive response56. The up-regulation of genes involved in Fe-S assembly (BP1026B_I0955, BP1026B_I0956, NifU, SufD) by BP1026B_II1198, could be induced in response to the intracellular environment, where iron-limitation, reactive-oxygen-species, and other environmental cues requires the sensitive nature of Fe-S regulatory complexes. This result correlates with previous reports showing up-regulation of Fe-S assembly genes in Burkholderia mallei, a clonal derivative of Bp, when exposed to sub-inhibitory levels of nitric oxide57. Genes involved in nitrogen metabolism were also up-regulated, showing that BP1026B_II1198 is partially responsible for a shift in general metabolism during infection. In addition to this, the BP1026B_I1020 mutant showed a reduction during intracellular replication in RAW264.7 cells.
Other mutants of genes regulated by BP1026B_II1198 that showed reductions in RAW264.7 pathogenesis have less clues as to their function. BP1026B_II1506 is a GntR family transcriptional regulator that could have many downstream gene expression changes that contribute to the reduction in virulence while BP1026B_II1581 is a hypothetical protein that is worth further investigation. BP1026B_I1775 is a Sec-secreted lipoprotein with cleavage site located between amino acids 28 and 29 as identified by SignalP5.0 analysis58. The function of this protein should be investigated further and could potentially be manipulated as a novel therapeutic target or subunit vaccine component. BP1026B_II1736 sits within the second type III secretion system (T3SS-2) in Bp and encodes SctK, a cytoplasmic structural component of the type III injectasome59. This protein along with SctQ and SctL are referred to as the ‘sorting platform’ and play a critical role on the order in which effector proteins are secreted60. Beyond the implications the specific potential function of BP1026B_II1736, it has been shown that T3SS-2 is involved in the pathogenesis of Solanum lycopersicum (tomato) but not mammalian cells37. While T3SS-2 is not singly required for complete pathogenesis in mammalian models of infection61, our data suggest that this system could contribute to pathogenesis.
Large numbers of genes/pathways are controlled indirectly through the expression of BP1026B_II1198 as described above. To find the direct DNA binding regions of BP1026B_II1198 we used a ChIP-seq strategy. Through this approach we identified two major regions of DNA binding in intergenic regions of divergently transcribed genes. BP1026B_II1198 showed up-regulation of BP1026B_II0600 and down regulation of BP1026B_II1006. This suggests that BP1026B_II1198 has a dual function as an activator and as a repressor, similar to other PadR-type regulators previously described50. Subsequent mutant analysis of these genes showed a 52% and 83% reduction in wildtype intracellular replication. BP1026B_II0600 encodes a levansucrase precursor, which to our knowledge, has yet to be described as a virulence factor during infection of mammalian cells. However, in plant pathogens like Pseudomonas syringae, levansucrase is critical for the formation of levans that contribute to exopolysaccaride production62,63. Because Bp is an incidental host of mammals, and freely lives within the environment, the expression of levansucrase could indicate that this gene is used for adaptation to multiple conditions. BP1026B_II1006 is annotated as a DNA binding protein but further analysis using Phyre264 predicts structural similarities to an Fe-S complex containing transcriptional regulator, RrsR65. We predict that the down-regulation of BP1026B_II1006 by BP1026B_II1198 plays a major role in the indirect gene expression changes that occur in our data set. Although outside of the scope of the present work, this data warrants further analysis of the regulation network of BP1026B_II1006.
Here we show a snapshot of what is going on during intracellular infection through the lens of the transcriptional regulator BP1026B_II1198 that we would like to tentatively name Intracellular Pathogenesis Regulator A (IprA). The loss of IprA results in significantly reduced pathogenesis in RAW264.7 murine macrophages and in BALB/c mice. This investigation identified that IprA functions as a transcriptional regulator during Bp intracellular transit through the host by activation/repression of multiple factors and pathways. Although, no single virulence factor controlled by IprA shows the same drastic decrease in pathogenesis we believe the cumulative effect of the changes in transcription by IprA highlights its importance during pathogenesis. We believe that in the intracellular environment, many transcription factors, including IprA, work in unison to control how Bp responds to specific environments, allowing for infection to occur. The future study of many of these transcriptional regulators (i.e. BP1026B_II1006 and BP1026B_II1506) will increase the resolution of what we know about the Bp intracellular lifecycle and Bp gene expression within the host. Although the data represent the expression profile of one transcription factor, it is one step towards a better understanding of the intricacies of Bp intracellular pathogenesis.
Methods and materials
Bacterial strains, media and culture conditions
Escherichia coli strain EPMax10B (BioRad), E1869, and E1354 were used for cloning or plasmid mobilization into Bp as described previously66,67. Bp 1026b and its select agent regulation excluded analogue, Bp82, were used as described in specific sections below. Luria–Bertani (LB) medium (Difco) or 1 × M9 minimal medium supplemented with 20 mM glucose (MG) were used to culture all strains. MG media was supplemented with 0.3% glyphosate (GS) or 0.1% chlorinated phenylalanine (cPhe) when appropriate. Growth media for Bp82 was supplemented with adenine and thiamine as previously described68. Experiments involving the manipulation of Bp were conducted in a CDC-approved and registered facility at the University of Hawaiʻi at Mānoa or Colorado State University. All work with the select agent Bp was approved by internal review and adhere to recommendations set forth in the BMBL, 5th edition69 for BSL3 organisms.
Transposon mutants tested as potential virulence factors
The sequence-defined transposon mutant library of Bp 1026b was used as a source of mutants to test for potential virulence factors36. Mutants of BP1026B_I0528, BP1026B_I0950, and BP1026B_II1464 were not tested because the library did not have insertions in any of these genes36. Transposon mutants available in the transposon library were struck for isolation on media containing kanamycin and each insertion site sequenced to confirm correct transposon insertion location (data not shown). The transposon insertion in BP1026B_II1769 was not validated and confirmed to be present in another gene, BP1026B_II2115, a putative diguanylate cyclase. BP1026B_II1769 lacked a second transposon insertion. All other transposon mutants tested were confirmed to be in the gene of interest by sequencing.
Molecular methods and reagents
Molecular methods and reagents were used as described previously20,67,70,71. An in-frame deletion mutant of BP1026B_II1198 was generated in Bp 1026b using lambda-red recombineering with minor modification72. Lambda red genes were PCR amplified from pKaKa1 and co-incubated with the knockout fragment of BP1026B_II1198 and selection carried out on glyphosate. Glyphosate resistant colonies were purified and patched on MG + 0.3% GS and screened for presence of the gat gene to confirm the deletion BP1026B_II1198. The gat-pheS cassette was removed via Flp recombination and growth on MG + 0.1% cPhe. Phenotypes were confirmed after purification and patching on corresponding media (Fig. S4a). For expression of BP1026B_II1198 (complementation, RNA-seq, ChIP-seq), BP1026B_II1198 was PCR amplified from Bp 1026b with oligos 2778 (5’-GGT TGC CTC GCA TAT GTC CCT GCC CCA C) and 2779 (5’-TCG AAG CTT CAA GCC GAC AAC GAT CTT) digested with NdeI and HindIII and ligated into NdeI/HindIII-cut pAM3GIQ-3xTY1 (Fig. S4b). Confirmation of the resulting vector, pAM3GIQ-3xTY1-BP1026B_II1198, was carried out by EcoRV/PhsAI and NdeI/HindIII digests and the expression of BP1026B_II1198 confirmed via western blot analysis with a mouse anti-TY1-tag monoclonal antibody (Diagenode C15200054) and goat anti-mouse conjugated to HRP (Invitrogen G-21040) (Fig. S4c).
Growth analysis
Growth analysis was carried out as previously described with minor modifications68. All strains were first grown overnight in LB broth at 37 °C, harvested the following day, diluted to an OD630 of 0.1 in 200 µl of fresh LB, and placed into 96 well plate. Growth curves were carried out at 37 °C shaking and data was recorded with the BioTek ELx808IU plate reader, measurements taken at OD630 every 30 min.
Cell infection assays
Intracellular replication and MNGC formation assays were carried out as previously described73 with minor modifications. Briefly, RAW264.7 murine macrophages were seeded for infection at 80–90% confluence on Corning CellBIND culture plates, allowed to attach overnight, washed twice with 1XPBS, and infected the following day. For intracellular replication assays, Bp 1026b, the BP1026B_II1198 mutant, or the BP1026B_II1198 complement (mutant with pAM3GIQ-3xTY1-BP1026B_II1198) were allowed to infect monolayers at an MOI of 1:1, washed with 1XPBS, and then DMEM supplemented with 10% FBS, 0.1 mM IPTG to drive expression of the complement, 700 µg/mL amikacin and 700 µg/mL kanamycin were added to kill any extracellular bacteria. At 2, 8, and 24 hpi, infected monolayers were lysed with 0.2% Triton X-100. Serial dilutions of lysates were plated on LB and colony-forming units (CFU) per well were determined. MNGC formation assays were carried out as intracellular replication assays, with the exception that 1.2% low-melt agarose was added to DMEM after infection allowing the formation of MNGCs. At 24 h post infection, monolayers were fixed with 4% paraformaldehyde, agarose plugs removed, stained with 0.05% crystal violet, and MNGC diameters measured with the Zeiss Axio Observer D1 and the AxioVision 64 bit 4.9.1 software. Cell infection assays with mutants from the Bp 1026b::T24 transposon mutant library were carried out as described above although kanamycin was removed and 1,500 µg/mL amikacin added to remove extracellular bacteria post infection. Monolayers were lysed at 24 h post infection as described above. Percent wildtype replication (%WT) was calculated by the following formula: %WT = (CFUmutant/mean CFUWT) × 100.
Animal studies
BALB/c mice (4 to 6 weeks old) were purchased from Charles River Laboratory, allowed to acclimatize for a week, and then infected with Bp 1026b and the BP1026B_II1198 mutant as previously described7. Each infection group had 5 mice and the inoculating dose was 4,500 CFU via the intranasal route after mice were anesthetized with ketamine (100 mg/kg) and xylazine (10 mg/kg). Mice were monitored for 62 days post infection and organs (lungs, liver, and spleen) from surviving mice were harvested, homogenized in PBS, serial diluted, and plated on LB. The only surviving mice that survived the length of the study were mice infected with the BP1026B_II1198 mutant. The organ loads show the persistence of this mutant within the BALB/c mouse. No mice were excluded from the study or analysis. Survival curves were plotted using Prism software and analyzed by Log-rank (Mantel-Cox) test (GraphPad).
RNA-seq, RT-qPCR, and ChIP-seq analysis
RNA-seq and ChIP-seq analysis were carried out under the same conditions. Briefly, Bp82 expressing BP1026B_II1198 from pAM3GIQ-3xTY1-BP1026B_II1198 was grown overnight and sub-cultured to mid-log phase in LB + adenine + 0.1 mM IPTG in triplicate. An empty vector (pAM3GIQ-3xTY1) was used as a control. Total RNA was harvested using RNeasy Mini Kit (Qiagen) with on-column (Qiagen) and off-column (Epicentere) DNase digestion steps. RNA samples were sent to Tufts University Genomics Core (TUCF Genomics) for library preparation and Illumina 50 bp single-end reads were sequenced on the Illumina HiSeq 2500. RNA-seq data was analyzed with Rockhopper21 to identify individual genes controlled by BP1026B_II1198 and WoPPER22 to determine larger regions of the genome that are controlled by BP1026B_II1198. RNA-seq data was validated by RT-qPCR using BP1026B_I0774, BP1026B_II1384, and BP1026B_II0521 as housekeeping genes due to consistent expression (log2FC = 0) between conditions (Fig. S4d). Primers (Table S2) were designed with Integrated DNA Technologies Primer Quest software. Reverse transcription was carried out with Superscript III (Invitrogen) and Power Up SYBR Green master mix (Applied Biosystems) was used for amplification and quantitation on an Eppendorf Realplex 4 thermocycler. RT-qPCR data was analyzed as previously described7,74.
ChIP-seq was carried out as previously described with minor modifications2,3. Briefly, ChIP-seq samples were grown identically as RNA-seq samples, harvested, and fixed with 4% paraformaldehyde, followed by shearing of DNA–protein complexes with the Covaris M220 ultrafocused sonicator. Cell debris was removed and DNA–protein complexes were immunoprecipitated with anti-TY1-tag monoclonal antibody (Diagenode C15200054) and secondary antibodies conjugated to magnetic beads (Diagenode C03010022). DNA–protein complexes were washed, decrosslinked, treated with RNaseA and proteinase K, and purified with QIAquick PCR purification kit. Immunoprecipitated DNA was sent to TUCF Genomics where DNA libraries were prepped and 50 bp single-end reads were sequenced with the Illumina HiSeq 2500. ChIP-seq data was aligned to the Bp 1026b genome with Bowtie275, peaks called with MACS276, and consensus binding regions determined with MEME38.
Ethical standards
All animal studies described in this manuscript were approved by the Institutional Animal Care and Use Committee at the University of Hawaiʻi at Mānoa (Protocol No. 10-1073), conducted in compliance with the NIH (National Institutes of Health) Guide for the Care and Use of Laboratory Animals, and in compliance with ARRIVE guidelines.
Data availability
The datasets and materials generated during the current study are available from the corresponding author upon reasonable request. Any transfer of select agent materials needs additional approval from Bradley R. Borlee and the Responsible Official at Colorado State University. Any select agent transfer must be to a select agent registered facility, approved by the CDC, and comply with all select agent regulations (selectagents.gov).
References
Browning, D. F. & Busby, S. J. Local and global regulation of transcription initiation in bacteria. Nat. Rev. Microbiol. 14, 638–650 (2016).
Heacock-Kang, Y. et al. Two Regulators, PA3898 and PA2100, modulate the Pseudomonas aeruginosa multidrug resistance MexAB-OprM and EmrAB Efflux pumps and biofilm formation. Antimicrob. Agents Chemother. 62 (2018).
Heacock-Kang, Y. et al. Novel dual regulators of Pseudomonas aeruginosa essential for productive biofilms and virulence. Mol Microbiol 109, 401–414 (2018).
Limmathurotsakul, D. et al. Predicted global distribution of Burkholderia pseudomallei and burden of melioidosis. Nat Microbiol 1 (2016).
Cheng, A. C. & Currie, B. J. Melioidosis: epidemiology, pathophysiology, and management. Clin. Microbiol. Rev. 18, 383–416 (2005).
Wiersinga, W. J. et al. Melioidosis. Nat. Rev. Dis. Primers 4, 17107 (2018).
Heacock-Kang, Y. et al. The Burkholderia pseudomallei intracellular ‘TRANSITome’. Nat. Commun. 12, 1907 (2021).
French, C. T. et al. Dissection of the Burkholderia intracellular life cycle using a photothermal nanoblade. Proc. Natl. Acad. Sci. USA 108, 12095–12100 (2011).
Benanti, E. L., Nguyen, C. M. & Welch, M. D. Virulent Burkholderia species mimic host actin polymerases to drive actin-based motility. Cell 161, 348–360 (2015).
Kespichayawattana, W., Rattanachetkul, S., Wanun, T., Utaisincharoen, P. & Sirisinha, S. Burkholderia pseudomallei induces cell fusion and actin-associated membrane protrusion: a possible mechanism for cell-to-cell spreading. Infect. Immun. 68, 5377–5384 (2000).
Shalom, G., Shaw, J. G. & Thomas, M. S. In vivo expression technology identifies a type VI secretion system locus in Burkholderia pseudomallei that is induced upon invasion of macrophage. Microbiology 153, 2689–2699 (2007).
Toesca, I. J., French, C. T. & Miller, J. F. The Type VI secretion system spike protein VgrG5 mediates membrane fusion during intercellular spread by pseudomallei group Burkholderia species. Infect Immun. 82, 1436–1444 (2014).
Schwarz, S. et al. VgrG-5 is a Burkholderia type VI secretion system-exported protein required for multinucleated giant cell formation and virulence. Infect Immun. 82, 1445–1452 (2014).
Chen, Y. et al. Characterization and analysis of the Burkholderia pseudomallei BsaN virulence regulon. BMC Microbiol. 14, 206 (2014).
Sun, G. W. et al. Identification of a regulatory cascade controlling Type III Secretion System 3 gene expression in Burkholderia pseudomallei. Mol. Microbiol. 76, 677–689 (2010).
Chen, Y. et al. Regulation of type VI secretion system during Burkholderia pseudomallei infection. Infect Immun. 79, 3064–3073 (2011).
Duong, L. T. et al. GvmR - A Novel LysR-Type transcriptional regulator involved in virulence and primary and secondary metabolism of Burkholderia pseudomallei. Front Microbiol. 9, 935 (2018).
Winsor, G. L. et al. The Burkholderia Genome Database: facilitating flexible queries and comparative analyses. Bioinformatics 24, 2803–2804 (2008).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016).
Kang, Y. et al. Knock-out and pull-out recombinant engineering protocols for naturally transformable Burkholderia thailandensis and Burkholderia pseudomallei. Nat. Protoc. 6, 1085–1104 (2011).
McClure, R. et al. Computational analysis of bacterial RNA-seq data. Nucleic Acids Res 41, e140 (2013).
Puccio, S. et al. WoPPER: Web server for Position Related data analysis of gene Expression in Prokaryotes. Nucleic Acids Res. 45, W109–W115 (2017).
Alice, A. F., Lopez, C. S., Lowe, C. A., Ledesma, M. A. & Crosa, J. H. Genetic and transcriptional analysis of the siderophore malleobactin biosynthesis and transport genes in the human pothogen Burkholderia pseudomallei K96243. J. Bacteriol. 188, 1551–1566 (2006).
Biggins, J. B., Kang, H. S., Ternei, M. A., DeShazer, D. & Brady, S. F. The chemical arsenal of Burkholderia pseudomallei is essential for pathogenicity. J. Am. Chem. Soc. 136, 9484–9490 (2014).
Biggins, J. B., Ternei, M. A. & Brady, S. F. Malleilactone, a polyketide synthase-derived virulence factor encoded by the cryptic secondary metabolome of Burkholderia pseudomallei group pathogens. J. Am. Chem. Soc. 134, 13192–13195 (2012).
Garcia, E. C., Anderson, M. S., Hagar, J. A. & Cotter, P. A. Burkholderia BcpA mediates biofilm formation independently of interbacterial contact-dependent growth inhibition. Mol. Microbiol. 89, 1213–1225 (2013).
Garcia, E. C., Perault, A. I., Marlatt, S. A. & Cotter, P. A. Interbacterial signaling via Burkholderia contact-dependent growth inhibition system proteins. Proc. Natl. Acad. Sci. USA 113, 8296–8301 (2016).
Anderson, M. S., Garcia, E. C. & Cotter, P. A. The Burkholderia bcpAIOB genes define unique classes of two-partner secretion and contact dependent growth inhibition systems. PLoS Genet 8, e1002877 (2012).
Podnecky, N. L., Rhodes, K. A. & Schweizer, H. P. E ffl ux pump-mediated drug resistance in Burkholderia. Front. Microbiol. 6, 305 (2015).
Podnecky, N. L., Wuthiekanun, V., Peacock, S. J. & Schweizer, H. P. The BpeEF-OprC efflux pump is responsible for widespread trimethoprim resistance in clinical and environmental Burkholderia pseudomallei isolates. Antimicrob. Agents Chemother. 57, 4381–4386 (2013).
Schwarz, S. et al. Burkholderia type VI secretion systems have distinct roles in eukaryotic and bacterial cell interactions. PLoS Pathog 6, e1001068 (2010).
Wollers, S. et al. Iron-sulfur (Fe-S) cluster assembly: the SufBCD complex is a new type of Fe-S scaffold with a flavin redox cofactor. J. Biol. Chem. 285, 23331–23341 (2010).
Mangalea, M. R., Plumley, B. A. & Borlee, B. R. Nitrate sensing and metabolism inhibit biofilm formation in the opportunistic pathogen Burkholderia pseudomallei by reducing the intracellular concentration of c-di-GMP. Front. Microbiol. 8, 1353 (2017).
Van Alst, N. E., Picardo, K. F., Iglewski, B. H. & Haidaris, C. G. Nitrate sensing and metabolism modulate motility, biofilm formation, and virulence in Pseudomonas aeruginosa. Infect. Immun. 75, 3780–3790 (2007).
Van Alst, N. E., Wellington, M., Clark, V. L., Haidaris, C. G. & Iglewski, B. H. Nitrite reductase NirS is required for type III secretion system expression and virulence in the human monocyte cell line THP-1 by Pseudomonas aeruginosa. Infect. Immun. 77, 4446–4454 (2009).
Borlee, G. I. et al. Genome-scale analysis of the genes that contribute to Burkholderia pseudomallei biofilm formation identifies a crucial exopolysaccharide biosynthesis gene cluster. PLoS Negl Trop Dis 11, e0005689 (2017).
Lee, Y. H., Chen, Y., Ouyang, X. & Gan, Y. H. Identification of tomato plant as a novel host model for Burkholderia pseudomallei. BMC Microbiol. 10, 28 (2010).
Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994).
Balasubramanian, D., Schneper, L., Kumari, H. & Mathee, K. A dynamic and intricate regulatory network determines Pseudomonas aeruginosa virulence. Nucleic Acids Res. 41, 1–20 (2013).
Huang, H. et al. An integrated genomic regulatory network of virulence-related transcriptional factors in Pseudomonas aeruginosa. Nat. Commun. 10, 2931 (2019).
Galan-Vasquez, E., Luna, B. & Martinez-Antonio, A. The regulatory network of Pseudomonas aeruginosa. Microb. Inform. Exp. 1, 3 (2011).
Galagan, J. E. et al. The Mycobacterium tuberculosis regulatory network and hypoxia. Nature 499, 178–183 (2013).
Klaus, J. R. et al. Malleilactone is a Burkholderia pseudomallei virulence factor regulated by antibiotics and quorum sensing. J Bacteriol 200 (2018).
Losada, L., Shea, A. A. & DeShazer, D. A MarR family transcriptional regulator and subinhibitory antibiotics regulate type VI secretion gene clusters in Burkholderia pseudomallei. Microbiology 164, 1196–1211 (2018).
Barthelmebs, L., Lecomte, B., Divies, C. & Cavin, J. F. Inducible metabolism of phenolic acids in Pediococcus pentosaceus is encoded by an autoregulated operon which involves a new class of negative transcriptional regulator. J. Bacteriol. 182, 6724–6731 (2000).
Gury, J., Barthelmebs, L., Tran, N. P., Diviès, C. & Cavin, J.-F. Cloning, deletion, and characterization of PadR, the transcriptional repressor of the phenolic acid decarboxylase-encoding padA gene of Lactobacillus plantarum. Appl. Environ. Microbiol. 70, 2146–2153 (2004).
Nguyen, T. K. C., Tran, N. P. & Cavin, J.-F. Genetic and biochemical analysis of PadR-padC promoter interactions during the phenolic acid stress response in Bacillus subtilis 168. J. Bacteriol. 193, 4180–4191 (2011).
Lubelski, J. et al. LmrCD is a major multidrug resistance transporter in Lactococcus lactis. Mol. Microbiol. 61, 771–781 (2006).
Huillet, E., Velge, P., Vallaeys, T. & Pardon, P. LadR, a new PadR-related transcriptional regulator from Listeria monocytogenes, negatively regulates the expression of the multidrug efflux pump MdrL. FEMS Microbiol. Lett. 254, 87–94 (2006).
Flórez, A. et al. Transcriptional regulation of mithramycin biosynthesis in Streptomyces argillaceus: dual role as activator and repressor of the PadR-like regulator MtrY. Microbiology (Reading, England) 161 (2014).
Heravi, K. M., Lange, J., Watzlawick, H., Kalinowski, J. & Altenbuchner, J. Transcriptional regulation of the vanillate utilization genes (vanABK Operon) of Corynebacterium glutamicum by VanR, a PadR-like repressor. J Bacteriol 197, 959–972 (2015).
Madoori, P. K., Agustiandari, H., Driessen, A. J. M. & Thunnissen, A.-M.W.H. Structure of the transcriptional regulator LmrR and its mechanism of multidrug recognition. EMBO J. 28, 156–166 (2009).
De Silva, R. S. et al. Crystal structure of the virulence gene activator AphA from Vibrio cholerae reveals it is a novel member of the winged helix transcription factor superfamily. J. Biol. Chem. 280, 13779–13783 (2005).
Mortenson, L. E., Valentine, R. C. & Carnahan, J. E. An electron transport factor from Clostridium pasteurianum. Biochem. Biophys. Res. Commun. 7, 448–452 (1962).
Mettert, E. L. & Kiley, P. J. Fe-S proteins that regulate gene expression. Biochim. Biophys. Acta 1853, 1284–1293 (2015).
Miller, H. K. & Auerbuch, V. Bacterial iron-sulfur cluster sensors in mammalian pathogens. Metallomics 7, 943–956 (2015).
Jones-Carson, J. et al. Inactivation of [Fe-S] metalloproteins mediates nitric oxide-dependent killing of Burkholderia mallei. PLoS One 3, e1976 (2008).
Armenteros, J.J.A. et al. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. 37, 420–423 (2019).
Wagner, S. et al. Bacterial type III secretion systems: a complex device for the delivery of bacterial effector proteins into eukaryotic host cells. FEMS Microbiol Lett 365 (2018).
Lara-Tejero, M., Kato, J., Wagner, S., Liu, X. & Galan, J. E. A sorting platform determines the order of protein secretion in bacterial type III systems. Science 331, 1188–1191 (2011).
Warawa, J. & Woods, D. E. Type III secretion system cluster 3 is required for maximal virulence of Burkholderia pseudomallei in hamster infection model. FEMS Microbiol. Lett. 242, 101–108 (2005).
Osman, S. F., Fett, W. F. & Fishman, M. L. Exopolysaccharides of the phytopathogen Pseudomonas syringae pv. glycinea. J Bacteriol 166, 66–71 (1986).
Hettwer, U., Gross, M. & Rudolph, K. Purification and characterization of an extracellular levansucrase from Pseudomonas syringae pv. phaseolicola. J Bacteriol 177, 2834–2839 (1995).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocol. 10, 845–858 (2015).
Volbeda, A. et al. Crystal structure of the transcription regulator RsrR reveals a [2Fe-2S] cluster coordinated by Cys, Glu, and his residues. J. Am. Chem. Soc. 141, 2367–2375 (2019).
Kang, Y., Norris, M. H., Barrett, A. R., Wilcox, B. A. & Hoang, T. T. Engineering of tellurite-resistant genetic tools for single-copy chromosomal analysis of Burkholderia spp. and characterization of the B. thailandensis betBA-operon. Appl. Environ. Microbiol. 75, 4015–4027 (2009).
Norris, M. H., Kang, Y., Lu, D., Wilcox, B. A. & Hoang, T. T. Glyphosate resistance as a novel select-agent-compliant, non-antibiotic selectable marker in chromosomal mutagenesis of the essential genes asd and dapB of Burkholderia pseudomallei. Appl. Environ. Microbiol. 75, 6062–6075 (2009).
Propst, K. L., Mima, T., Choi, K. H., Dow, S. W. & Schweizer, H. P. A Burkholderia pseudomallei deltapurM mutant is avirulent in immunocompetent and immunodeficient animals: candidate strain for exclusion from select-agent lists. Infect Immun. 78, 3136–3143 (2010).
Wilson, D.E. & Chosewood, L.C. Biosafety in microbiological and biomedical laboratories (BMBL), 5th ed. Centers for Disease Control and Prevention, Atlanta, GA. (2007).
Norris, M. H. et al. The Burkholderia pseudomallei Deltaasd mutant exhibits attenuated intracellular infectivity and imparts protection against acute inhalation melioidosis in mice. Infect Immun. 79, 4010–4018 (2011).
Norris, M. H., Kang, Y., Wilcox, B. & Hoang, T. T. Stable site-specific fluorescent tagging constructs optimized for Burkholderia species. Appl. Environ. Microbiol. 76, 7635–7640 (2010).
Kang, Y. et al. Knockout and pullout recombineering for naturally transformable Burkholderia thailandensis and Burkholderia pseudomallei. Nat. Protoc. 6, 1085–1104 (2011).
Heacock-Kang, Y. et al. The heritable natural competency trait of Burkholderia pseudomallei in other Burkholderia species through comE and crp. Sci. Rep. 8, 12422 (2018).
Peirson, S. N., Butler, J. N. & Foster, R. G. Experimental validation of novel and conventional approaches to quantitative real-time PCR data analysis. Nucleic Acids Res. 31, e73 (2003).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Acknowledgements
This project was supported by the US National Institutes of Health (NIH)/National Institute of Allergy and Infectious Diseases (NIAID) grant number R21AI123913 awarded to Tung T. Hoang. We would like to thank Mark S. Blumhardt for helping make lambda red fragments for generating the BP1026B_II1198 deletion mutant. This is publication number 115 from the School of Life Sciences, University of Hawai'i at Mānoa.
Author information
Authors and Affiliations
Contributions
I.A.M., M.H.N., Y.H.K., and T.T.H. designed the experiments. I.A.M., M.H.N., J.Z.S., Y.H.K., and Z.S. conducted the experiments. I.A.M. analyzed the data. B.R.B. provided select-agent strains and contributed reagents and supplies. I.A.M. wrote the manuscript, and all authors contributed to editing of this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
McMillan, I.A., Norris, M.H., Zarzycki-Siek, J. et al. Identification of a PadR-type regulator essential for intracellular pathogenesis of Burkholderia pseudomallei. Sci Rep 11, 10405 (2021). https://doi.org/10.1038/s41598-021-89852-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-89852-7
- Springer Nature Limited