Abstract
Sjögren’s syndrome (SS) is a complex autoimmune disease associated with lymphocytic infiltration and secretory dysfunction of salivary and lacrimal glands. Although the etiology of SS remains unclear, evidence suggests that epithelial damage of the glands elicits immune and fibrotic responses in SS. To define molecular changes underlying epithelial tissue damage in SS, we laser capture microdissected (LCM) labial salivary gland epithelia from 8 SS and 8 non-SS controls for analysis by RNA sequencing (RNAseq). Computational interrogation of gene expression signatures revealed that, in addition to a division of SS and non-SS samples, there was a potential intermediate state overlapping clustering of SS and non-SS samples. Differential expression analysis uncovered signaling events likely associated with distinct SS pathogenesis. Notable signals included the enrichment of IFN-γ and JAK/STAT-regulated genes, and the induction of genes encoding secreted factors, such as LTF, BMP3, and MMP7, implicated in immune responses, matrix remodeling and tissue destruction. Identification of gene expression signatures of salivary epithelia associated with mixed clinical and histopathological characteristics suggests that SS pathology may be defined by distinct molecular subtypes. We conclude that gene expression changes arising in the damaged salivary epithelia may offer novel insights into the signals contributing to SS development and progression.
Similar content being viewed by others
Introduction
Sjögren's syndrome (SS) is a debilitating complex autoimmune disease presenting as exocrinopathy of salivary and lacrimal glands with frequent systemic extraglandular manifestations and an increased risk of non-Hodgkin’s lymphoma1,2,3,4,5,6. Damage to salivary and lacrimal glands from SS impairs their ability to produce saliva and tears, resulting in xerostomia and xerophthalmia. SS is a multifactorial disease that comprises various combinations of dry eyes, dry mouth, reduced tear and saliva secretion, presence of autoantibodies, focal lymphocytic infiltration of salivary glands, fatigue, vasculitis, joint and muscle pain, as well as peripheral nervous system dysfunction4,7. Although to date, the origin of SS remains unknown, the prevailing model for SS has been that loss of secretory function is secondary to lymphocytic infiltrates.
Since defects in the immune system have long been considered to be the cause of SS, the majority of studies and therapeutic approaches have focused on aberrant immune responses8,9,10. Analysis of genomic and epigenomic changes in SS has provided new insight into the understanding of disease pathogenesis9,11,12,13. Gene expression microarray studies on labial salivary glands and peripheral blood detected dysregulation of type 1 interferon (IFN)-inducible genes14. Additional upregulated genes involved in antigen presentation, lymphocyte development and activation, as well as interferon-induced chemokines were also identified. Interestingly, IFN-γ-inducible genes, such as Signal transducer and activator of transcription (STAT) family members STAT1 and STAT3, were also shown to be highly expressed in primary SS (pSS)15. Furthermore, IFN-mediated innate immune mechanisms have been implicated in the pathogenesis of pSS, where specifically 23 genes known to play a role in IFN signaling were identified16. Two GWAS studies have been performed in pSS and among the SS-associated non-HLA genes they identified STAT4 and IRF5 encoding transcription factors, BLK coding for B cell kinase as well as genes encoding the IL-12A cytokine. Genes involved in NF-κB signaling and CXCR5 chemokine production were also shown to be upregulated9,17. Collectively, these studies provided a strong support for the role of innate immunity, as well as adaptive immune mechanisms, in the pathogenesis of pSS. Despite these advances, treatment strategies for SS disease are almost inexistent, and molecular mechanisms that drive disease onset are poorly understood.
One consistent feature of human autoimmunity is the finding that there is an initial phase in which patients have many of the serological abnormalities of autoimmune disease before they develop clinical pathology18. In this respect, the immunobiology of the glandular epithelial cells within the target tissue likely plays a critical role in initiating events in SS. Indeed, accumulating evidence suggests that impaired glandular secretion develops independent of lymphocytic infiltration, and defective secretion precedes the immune response during SS onset, suggesting a role for the epithelium in the etiology of SS2. For example, aberrantly high expression of BMP growth factors, such as BMP6, has been implicated in the induction of salivary and lacrimal gland dysfunction independent of the autoantibodies and immune infiltration associated with SS19. Recent studies, including our own, have also shown that the structural integrity of the epithelium of SS patients is lost, and that these defects are associated with aberrant apical-basal polarity20,21,22,23,24,25. Aberrant polarity can arise in various ways, including environmental toxins, viral infection, physical damage, and unbalanced hormone levels, and therefore it may be an early event that triggers disease states, including SS. Since structural changes of the epithelium of salivary glands may drive inflammation, the aim of this study was to elucidate molecular details of how epithelial structural dysfunction contributes to SS. We used laser capture microdissection of glandular epithelial regions (acini and ducts) from resected labial salivary glands of patients fulfilling or not fulfilling SS criteria (8 SS and 8 non-SS controls) and processed them for global transcriptomes using bulk RNAseq analysis. Our studies provide evidence that damage to salivary gland epithelia is associated with changes in gene expression signatures that provide insights into deregulated pathways and cellular processes which may underlie predisposition to, and early onset of, SS. Collectively, these findings provide support for the theory that alterations in gene expression arising in the salivary epithelia contribute to the etiology of SS. We postulate that the molecular sub-groups reflect varying stages of disease predisposition and that they may offer novel insights into the signals contributing to the progression of SS.
Results
Clinical characteristics of patients
Sixteen samples of labial salivary glands were obtained from female patients who presented with sicca symptoms to the Department of Otolayngology/Head and Neck Surgery at Haukeland University Hospital, Bergen. Patients were assessed for primary Sjögren’s Syndrome (pSS) based on the American-European Classification Criteria26, where diagnosis of pSS required focal lymphocytic sialadenitis score (FS) of ≥ 1 or presence of anti-SSA/Ro or anti-SSB/La autoantibodies, as well as ocular symptoms of dry eyes, oral symptoms of dry mouth, positive Schirmer’s test, and reduced unstimulated salivary flow ≤ 1.5 mL in 15 min (at least four positive tests including symptoms, or three objective tests). We categorized samples into two clinically distinct groups, patients diagnosed with SS (named SS, n = 8) or without SS (named non-SS, n = 8). Clinical characteristics of patients are shown in Table 1. The patient mean age was 53 ± 12.4 years, with three patients in the SS group presenting with anti-SSA/Ro and ANA antibodies, of which also two were positive for anti-SSB/La antibodies. All patients reported dry eyes and the majority reported dry mouth. Three SS patients had FS of 1, three had a FS of 2, with one patient presenting with FS of 3 and one with FS of 4. Interestingly, FS of either 1 or 4 were associated with positivity for all anti-SSA/Ro, anti-SSB/La and ANA, suggesting that these may be independent variables in our sample group. Likewise, there was no association between germinal centers (GCs) and autoantibody positivity. Furthermore, two of the non-SS patients had a negligible salivary flow rate, and there was little correlation between salivary flow rate and auto-antibody positivity in SS patients.
Histopathological evaluation of H&E sections of biopsies from non-SS and SS patients revealed that SS samples were characterized by acinar atrophy, ductal dilation, detectable fat infiltration, and lymphocytic aggregates with FS ≥ 1, whereas non-SS tissues were generally normal in appearance with some acinar atrophy, ductal dilation, and scattered inflammation (Fig. 1). From analysis of clinical presentation, we observed that there was no correlation between SS and non-SS groups for the parameters of positive Schirmer’s test, saliva secretion, dry mouth or dry eyes. However, the presence of anti-SSA/Ro, anti-SSB/La, ANA autoantibodies, FS ≥ 1, and the presence of CD21+ GCs were only observed in the SS group. When assessed for age, the SS group was significantly older than the non-SS group (60.2 ± 10.7 years vs 45.6 ± 9.6 years respectively; p = 0.012).
Minor salivary glands biopsies. H&E staining of minor salivary gland biopsies from: (a) Non-SS control showing normal appearing salivary gland tissue with few lymphocytic cells, (b) Non-SS control with scattered lymphocytes and some interstitial fibrosis, (c) SS patient with FS = 1 showing a focal lymphocytic infiltrate and some acinar atrophy and (d) SS patient with FS = 2 with a germinal center within a large lymphocytic focal infiltrate.
Subclustering of epithelial-derived RNA-seq data reveals potential SS subgroups
To gain insight into the molecular characteristics of two groups with sicca symptoms, eight non-SS and eight SS, all samples were subjected to laser capture microdissection (LCM) of epithelial regions followed by RNAseq analysis to identify biomarkers restricted to the epithelial compartment of the labial salivary glands. We evaluated the entire set by principal component analysis (PCA) selected by the expression level of all differentially expressed (DE) genes. It was evident that the SS and non-SS samples did not entirely cluster together and rather three distinct transcriptomic subclusters were observed: a group containing non-SS samples (named Group A, n = 3), a group containing SS (named Group B, n = 3), and a group containing samples from both groups (named Group AB, n = 10) (Fig. 2a,b). The position of these groupings demonstrates different features of RNA expression in the epithelial tissues, suggesting that transcriptomic profiling of patient samples may characterize important molecular events in SS pathogenesis compared to grouping by clinical characteristics alone.
RNAseq of LCM isolated epithelia from 8 non-SS and 8 SS patients. (a) PCA of total gene expression data reveals segregation between non-SS (Group A) and SS (Group B) samples and similarities between non-SS and SS (Group AB), identifying a new sub-group with mixed clinical features but the same transcriptional profile. Groups were defined as: Group A (non-SS, n = 3), Group AB (mixed non-SS/SS, n = 10), and Group B (SS, n = 3). (b) Sample-sample distance heat map further clusters the sample set. (c) Comparison of outputs of the number differentially expressed (DE) genes at different filtered FDR cut-off levels. When the biopsies were analyzed for DE genes by RNAseq cluster, a greater number of DE genes were observed.
When assessed for age, Group B (69.3 ± 12.7 years) was significantly older than both the Group AB (52.2 ± 7.4 years; p = 0.020) and Group A (39.0 ± 6.6 years respectively; p = 0.002), whereas there was no significant age difference between Group A and Group AB. Clinical findings for the RNA-Seq groupings revealed that all samples in Group B had GCs with FS of ≥ 2. One of the patients in this group developed lymphoma during the course of this study. In the mixed Group AB, two of the five SS patients also had GCs and lower mean FS. There was no correlation observed for RNAseq grouping for the presence of anti-SSA/Ro, anti-SSB/La and ANA autoantibodies, salivary flow and Schirmer test. Likewise, no specific pattern regarding Group A patients was detected, except that all presented with positive Schirmer tests and a subjective feeling of dry eyes.
Differential expression testing of SS subgroups
Genome-wide differential expression testing was performed using DESeq2 for multiple group comparisons. First, we tested the clinical grouping of SS against non-SS. Second, we compared the RNAseq grouping of Group B against Group A samples to determine if there were differences in gene expression between clinical diagnosis of Sjögren’s and principal component analysis (PCA) for clustering. Third, the RNAseq Group A and Group B groups were compared to the Group AB.
RNAseq analysis of the epithelial regions for the groupings clinically diagnosed with SS showed no significantly altered differentially expressed (DE) genes compared to the non-SS group out of 25,444 total genes analyzed (Fig. 2c). However, when analyzed according to PCA clustering, there were 1247 DE genes between Group B and Group A at a FDR cut-off of 0.05 (Fig. 2c), further demonstrating a benefit of clustering the samples according to RNAseq data. Table 2 summarizes the top 10 DE upregulated and downregulated genes for both clinical Sjögren’s (SS vs non-SS) and RNAseq (Group B vs Group A vs Group AB) groupings.
Inter-group analysis with the Group AB subcluster revealed the following comparisons: 767 DE genes compared to the Group A (FDR cut-off = 0.05) and 1480 DE genes compared to Group B (FDR cut-off = 0.05). While strictly not statistically meaningful, the FDR-value ranking was used to define signatures to be annotated by pathway enrichment analysis.
Pathway analysis identifies epithelial signals associated with SS
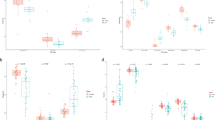
Gene set enrichment analysis (GSEA) was conducted for each of the groups, and hallmark gene sets were evaluated (Fig. 3a). Comparison of SS vs non-SS groupings revealed 8 signaling pathways that were statistically upregulated and 27 pathways downregulated in the SS group (Supplementary Table S1). Notably, immunological and autoimmune disease pathways were enriched in the SS group while cholesterol and protein-related metabolic pathways were downregulated (Fig. 3b,c). Interferon response genes were upregulated in both the clinical SS and RNAseq (Group B) groupings compared to non-SS and Group A samples, respectively, suggesting conservation of relevant processes reflected in RNA profiling. Further hallmark GSEA of comparisons between the subcluster groupings revealed 9 upregulated and 23 downregulated pathways for Group B vs Group A, 21 upregulated and 13 downregulated pathways for Group B vs Group AB, and 30 upregulated and 1 downregulated pathway for Group A vs Group AB (Fig. 3a).
Hallmark gene set analyses from the entire RNAseq dataset. (a) Heatmap of hallmark gene sets that are significantly up (red) or downregulated (blue) in the respective indicated pathology. (b) Gene set enrichment analysis (GSEA) of hallmark gene sets that are significantly enriched in SS (n = 8) vs non-SS (n = 8). (c) GSEA of hallmark gene sets that are significantly enriched in non-SS (n = 8) vs SS (n = 8). NES nominal enrichment score, FDR false discovery rate. A gene set shows significant enrichment at a FDR < 0.25.
Compared to Group AB, both Group A and Group B showed upregulated pathways in genes encoding components of the apical junction complex and apical surface of epithelial cells, epithelial–mesenchymal transition and coagulation, all indicative of changes in epithelial cell polarity. Significantly, the Group AB showed intermediary expression of genes involved in protein secretion and those activated through the mTORC1 complex as enrichment of genes in these pathways was low in Group B and high in Group A.
While the RNAseq data generated from the epithelia of non-SS and SS salivary glands showed complexity in gene signatures among patient samples, a closer comparison revealed genes that encode secreted proteins that may contribute to the disease. These included genes encoding lactotransferrin (LTF), matrix metalloproteinase 7 (MMP7), bone morphogenetic protein 3 (BMP3), interferon-inducible proteoglycan testican-2 (SPOCK2) as well as cytokines IL-15, IL-19 and BAFF (TNFSF13B). Both BAFF and IL-15 have been shown to be associated with B cell development, physiology and malignancy13,27. Furthermore, SPOCK2 was also identified as a downstream target of lncRNA-dependent miRNAs involved in the modulation of ECM genes in SS13.
RNAscope analysis reveals increased LTF, MMP7, and BMP3 expression in Sjögren’s labial salivary gland epithelia
Among the top genes whose transcriptional expression was increased in the RNAseq analysis of the SS grouping were LTF and in Group B were MMP7 and BMP3. LTF is a glycoprotein found in exocrine fluids, such as milk, saliva, tears, and nasal secretions, as well as in secondary granules of polymorphonuclear leukocytes (PMNs) and was previously shown to be upregulated on the protein level in saliva of SS patients28,29. MMP7 belongs to a family of ubiquitous proteolytic enzymes, the MMPs, and has been shown to be elevated in corneal specimens of pSS patients30. BMP3 is a member of the TGF-β superfamily of proteins, associated with inflammation in rheumatoid arthritis patients with concomitant inhibition of bone repair31.
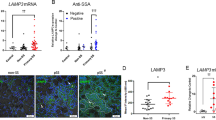
In order to validate the increased LTF, MMP7, and BMP3 expression identified in our RNAseq data, we performed independent RNAscope analyses for expression of these genes. This in situ hybridization approach is based on the amplification of target-specific signals but not non-specific background hybridization. Samples were counterstained with pan-cytokeratin in order to restrict analysis of RNA to epithelial regions and analyzed using the CellProfiler image analysis software32,33,34 (http://www.cellprofiler.org). To increase our sample size, we included an additional group of labial salivary gland samples obtained from the University of Oslo. Figure 4 illustrates combined RNAscope analysis of LTF, MMP7 and BMP3 expression from the Bergen samples described in Table 1 and Oslo samples described in Table 3. We note that the age of SS patients in the Oslo cohort was not significantly different than the non-SS group (51.0 ± 19.1 years vs 44.5 ± 13.1 years respectively) although the new Oslo cohort included SS samples from two male patients and the SS groups were mostly positive for autoantibodies (Table 3). Quantitation of the RNA showed significant increases in the epithelial percentage of LTF, MMP3, and BMP3 expression in the SS group compared to the non-SS group (Fig. 4a,b). The observed increases in LTF, MMP3 and BMP3 were not driven by age as there was no significant correlation between patient age and expression of either gene (Fig. 4, “Methods”, “Statistical analysis”). These results correlate with our RNAseq findings of increased epithelial expression of LTF, MMP7, and BMP3 in SS.
RNAscope validation of gene expression changes identified from RNAseq of SS epithelial tissues. (a) SS vs non-SS biopsies were probed for lactotransferrin (LTF), bone morphogenetic protein 3 (BMP3), and matrix metalloproteinase 7 (MMP7) RNA (green) by in situ RNAscope and counterstained with pan-cytokeratin to mark epithelium (pCK, red). Scale bars = 20 µm. (b) LTF, BMP3, and MMP7 RNAs were quantified in the respective pCK-positive tissues from patients in the Bergen (green) and Oslo (red) cohorts, revealing increased epithelial expression of LTF, BMP3, and MMP7 in SS tissues. The average % LTF, BMP3, and MMP7 area per patient sample in epithelial regions is shown. Data points represent each patient from Bergen and Oslo cohorts per condition of non-SS or SS. Each data point denotes the average of a range of 2000–4000 epithelial cells scored across multiple tissue fields from each patient sample (***p < 0.001 compared to non-SS group determined with unpaired t tests).
Discussion
Sjögren’s syndrome (SS) is a debilitating autoimmune disease with poorly understood etiology and limited biomarkers available for early detection and prognosis. In this study, we generated and annotated gene expression signatures of labial salivary gland epithelia from human SS and non-SS patients using RNAseq and computational analyses with a goal of obtaining new insights into molecular changes associated with this disease. Our studies provide evidence that damage to salivary gland epithelia is associated with changes in gene expression signatures that provide insights into deregulated pathways and cellular processes which may underlie predisposition to and early onset of SS. We postulate that the molecular sub-groups identified in our studies reflect varying stages of disease predisposition and that they may offer novel insights into the signals contributing to the development of SS.
PCA analysis showed a clear segregation of gene signatures between some non-SS and SS patients (Fig. 2). In addition, it revealed a novel subgroup of gene signatures derived from both non-SS and SS tissues that may reflect a mixed patient population. Interestingly, however, analyses of clinicopathological features of salivary epithelia from SS and non-SS patient cohorts did not reveal any consistent association with GCs and focus scores; indeed, there were no substantial differences among these patient groups in autoantibody positivity, saliva secretion, tear secretion, and subjective feeling of dry eyes or dry mouth. Our identification of a mixed gene expression signature subgroup (Group AB) sheds novel insights into molecular changes in labial salivary glands that may reflect early disease in individuals whose symptoms do not meet the criteria for SS diagnosis.
Better understanding of the molecular features that underlie different subsets of patients is essential for disease diagnostics, prognosis, and effective treatment interventions. The focal infiltration of the salivary gland provides the basis for the focus score used to describe the pathological changes of the salivary glands in the currently used classification criteria for pSS. While the European-American Consensus Group criteria26 include symptoms of dry mouth and dry eyes, the ACR-criteria include only objective criteria of reduced secretion of saliva and tears, ocular staining, positive focus score, and presence of anti-SSA/Ro in serum35. Both classification criteria for pSS require that the patients have anti-SSA/Ro antibodies and/or have a positive focus score, but do not include germinal centers. The epithelial gene expression profiles we describe show some association with clinical/pathological features of SS, but such molecular similarities are also evident in non-SS patients. This may be a consequence of the stochastic nature of clinical/pathological characteristics currently used to stratify patients into non-SS and SS groups.
Changes in SS epithelial and stromal tissue biology have been increasingly shown to play a role in SS pathobiology13,36. Previous studies revealed alterations in subpopulations of epithelial cells in NOD/ShiLtJ mouse model of SS, revealing a decrease in the tissue area of secretory cells in favor of ductal epithelial cells and expansion of keratin 5-positive basal cells37. Additional studies highlighted the key roles of the innate and adaptive immune systems, as well as stromal cells, in promoting the pro-inflammatory milieu38,39. A more recent report using an LCM approach coupled with RNAseq analyses described identification of cell-type and disease-specific markers in SS patients40. However, the latter study did not strictly focus on salivary gland epithelia, and thus precluded identification of gene signatures associated only with the epithelial compartment. Additionally, recent analyses of PBMCs from SS patients provided evidence for the involvement of the perturbed long non-conding RNAs (lncRNAs)—microRNAs axes in the expression of genes involved in the pathogenesis of SS, including those with roles in B cell function and extracellular matrix remodeling, among others13. Here, our results identifying IL-15 and BAFF cytokines with important functions in B cell physiology, as well as SPOCK2 proteoglycan contributing to the increasingly recognized roles of the stromal compartment in the pathogenesis of SS36, provide important new information into their epithelial origin in SS.
In addition to signals known to be enriched in SS, including IFN-γ and JAK/STAT-regulated genes, and genes encoding secreted factors implicated in immune responses, our studies identified LTF, also known as lactoferrin (LF), to be upregulated in expression in SS epithelia compared to non-SS tissues. Increased levels of salivary LTF with some relationship to the labial salivary gland histopathology grade were reported previously28, while no significant changes in LTF expression were detected in tear secretions in SS41. LTF is an iron-binding multifunctional glycoprotein known to function in the prevention of infections, late onset sepsis, and necrotizing enterocolitis29. LTF has been shown to stimulate a balanced T-helper-1/T-helper-2 cytokine immune response and to be a part of the innate defense, mainly at mucosa42. In our study, the observed increase in LTF gene expression in SS ductal epithelia suggests that it may be a relevant biomarker predictive of early disease with a potential causal effect on the induction of immune response.
Our data also show that MMP7 and BMP3 transcripts are upregulated in labial salivary glands from SS patients. MMP7 is a member of ubiquitous MMPs with important functions in tissue remodeling during development, but whose inappropriate expression has been linked to tissue destruction in autoimmune rheumatoid arthritis43 and with corneal melting in patients with primary SS30. This suggests that upregulation of MMP7 in labial tissues from SS patients is likely to contribute to salivary gland destruction. In contrast, relatively less is known about the role of BMP3 in pSS. Previous studies documented increased expression of BMP6 in salivary and lacrimal glands from human SS patients independent of autoantibodies19. To date, increased expression of BMP3 has been linked to immune infiltration in rheumatoid arthritis and inhibition of bone repair31. Thus, our RNAseq-based identification of BMP3 in salivary epithelia from SS patients may provide additional mechanistic insights into how inflammation induces BMP3 expression to drive epithelial dysfunction in pSS.
Accumulating evidence suggests a role for the epithelium in the etiology of SS. Impaired epithelial secretion has been reported to develop independently of lymphocytic infiltration and frequently, defective secretion precedes the immune response during SS onset2. Epithelial defects are often accompanied by pro-fibrotic changes in the stroma indicating remodeling of the extracellular matrix and fibroblast activation36. Here, our findings align alterations in gene expression signatures in the salivary epithelia with different degrees of compromised salivary and lacrimal glands’ function. Collectively, these studies offer a molecular explanation for exocrine tissue dysfunction as it progresses to SS and suggest that transcriptional signatures may serve as predictive biomarkers for early and advanced disease.
Methods
Patient selection and sample collection
The study protocol was approved by the Norwegian Regional Committee for Medical and Health Research Ethics of West (REK 2009/686) and South-East Norway (REK 2010/1292-1). The study was performed in compliance with the tenets of the Declaration of Helsinki. Prior to participation in the study, written informed consent was obtained from all participants. The data was de-identifed prior to analysis.
Labial salivary glands were obtained from a total of 27 patients with sicca symptoms. Based on the American-European Consensus Criteria26, these patients were sub-divided into 13 pSS (focus score (FS) 1–4) and 14 non-SS sicca controls (FS < 1). Inclusion criteria were: SS-patients fulfilling the AECG classification for pSS; non-SS controls were patients evaluated for SS, but not fulfilling the AECG classification. Importantly, none of the non-SS controls had a positive focus score or positive anti-SSA/Ro or anti-SSB/La autoantibodies. All participants provided informed consent prior to the initiation of this study. Clinical characteristics of patients whose minor salivary gland (SG) biopsies were analyzed for RNAseq are summarized in Table 1 (Bergen cohort) and for RNAscope analyses in Table 3 (Oslo cohort). These included assessment of objective tests of antinuclear antibodies (ANA, anti-SSA/Ro, anti-SSB/La), saliva secretion (unstimulated flow of whole saliva), tear secretion by Schirmer test, FS, and germinal center (GC)-like structures, as well as subjective symptoms noted by the patients (dry mouth and dry eyes).
Histopathology
Paraffin-embedded, formalin-fixed human salivary gland tissue sections, 5-µm-thick, were stained with hematoxylin and eosin (H&E) and focal lymphocytic infiltration areas in H&E slides were captured and assessed using Aperio digital technology (Leica Biosystem). Minor SG FS of SS patients was assessed based on the currently accepted criteria4. The sections were evaluated by one oral pathologist (KS) in order to determine their FS. The tissue sections were further assessed for the presence of GC-like structures both on H&E sections and with an additional staining with CD21 (a marker of follicular dendritic cells) to improve the reliability of GC identification44. Fresh frozen biopsies from the Bergen cohort characterized in Table 1, were used for RNAseq analyses. Additional minor SG biopsies from another patient cohort (Oslo cohort, Table 3) were used for immunofluorescence labeling and RNAscope analyses.
Laser capture microdissection (LCM) analysis
Fresh surgical specimens were preserved in the Tissue-TEK OCT compound and frozen in isopentane, followed by storage at − 80 °C. Tissue sections (5-μm-thick) were stained with an ethanol-based cresyl violet and eosin stain, followed by examination under the microscope and annotation of epithelium and inflammatory infiltrates on digital images. Acinar and ductal epithelium was subsequently captured by LCM using a Leica LMD6 microdissection system with regions containing inflammatory cells being excluded.
RNAseq analyses
RNA extraction and RNA sequencing
RNA was isolated from laser capture microdissected labial SG epithelia. Approximately 30–100 ng of RNA was isolated from each patient sample in 4 μl with concentration of 7.5–25 ng/μl. Libraries for RNAseq were prepared using the Clontech Mammalian SMARTer Stranded Total RNASeq Kit Pico Input and then sequenced on an Illumina HiSeq 2500 instrument.
Processing RNA-sequencing data
Paired-end RNA-sequencing reads for each sample were first trimmed for Illumina TruSeq adapters (AGATCGGAAGAGCACACGTCTGAACT CCAGTCAC and AGATCGGAAGAGCGTCGTGTAGGGAAAGAGTGTAGATCGGAAGA GCGTCGTGTAGGGAAAGAGTGTAGATCTCGGTGGTCGCCGTATCATT) using Cutadapt tool45, setting the minimum read quality threshold to 20 for the start and end of the reads, and 15 for the minimum read length. Reads were then aligned to a reference hg19 genome using Tophat246, with the number of allowed read mismatches, gap length and edit distance set to 1. Reads were then mapped to gene features using Ensembl hg19 (human genome release 19) genome annotations, and quantified as raw counts per feature using featureCount47 using default parameters.
RNA differential expression testing
Gene feature raw RNA expression counts were normalized and tested for differential expression between different sample groupings using DESeq248. In our analyses, we did not include age as a variable to be adjusted for when performing differential expression testing, although we are aware that age can be a confounding factor. Pairwise comparisons for the non-SS and SS RNAseq data provided sufficient coverage to estimate the differential expression of moderately to highly expressed genes.
Gene set enrichment analysis
For testing enrichment of pathways, genes were first ranked (genome-wide) based on their signed average log-fold change between group comparisons (e.g. Sjogren’s vs control), using the pre-ranked GSEA tool v2.1.1 given a specific differential expression ranked list (based on the ‘t-statistic’ of differential expression). Data were then tested for enrichment of annotated hallmark gene sets from the molecular signature database (MSigDB)49 using the pre-ranked Gene Set Enrichment Analysis (GSEA) tool50. Pathway-based enrichment testing allowed the identification of coordinated sets of genes, and GSEA computationally determined if a defined set of genes was of statistical significance. We note that we did not use any fold-change cut-off here in addition to an FDR threshold.
RNAscope analyses
Formalin-fixed, paraffin-embedded non-SS and SS patient minor SG biopsies from the Bergen and Oslo patient cohorts were cut into 4-μm-thick sections and mounted onto Superfrost Plus slides. Samples were deparafinized and in situ hybridization was conducted using the RNAscope 2.5 HD Assay Reagent Kit (ACDBio), probing for lactotransferrin Hs-LTF (LTF) RNA, matrix metalloproteinase 7 Hs-MMP7 (MMP7) RNA, and bone morphogenetic protein 3 (BMP3) RNA, Positive Control Probe Hs-PPIB, and Negative Control probe Dap-B according to the manufacturer’s instructions (ACDBio). After the final amplification, fast-red chromogenic detection was performed. The epithelial localizations of LTF, MMP7, and BMP3 were determined by counterstaining the tissue sections with pan-cytokeratin protein. Sections were blocked with 5% donkey serum-TBST (DS-TBST) and incubated in primary antibody 1:100 in 5% DS-TBST for 60 min, followed by incubation in 1:500 secondary goat anti-mouse conjugated with Alexa 647, counterstained with DAPI, and mounted in ProLong Gold Antifade and coverslipped. Image processing of the LTF/BMP3/MMP7 and pan-cytokeratin (pCK) signals was carried out on the free, open-source software CellProlifer v4.0.6 across all 2048X2048px images32,33,34 (http://www.cellprofiler.org). We utilized the ability to mask nuclei and epithelial cells, followed by merging the two masks to define a full epithelial cell. Nuclei outside the epithelial mask was discarded from further analysis. Following the estimation of epithelial cells, we measured the area of pixels of RNAscope probes within the of area covered by the epithelial mask to derive the percentage of area covered by a probe signal over all the epithelial areas (Supplementary Figure 1). This analysis was carried out on all patients on a minimum of 3 images per patient.
Statistical analysis
All analyses were performed with GraphPad Prism statistical software version 8.4.3 (GraphPad Software Inc., La Jolla, CA, USA). Unpaired t tests were performed to compare values among groups for RNAscope of LTF, BMP3, and MMP7 expression. Each data point represents a patient. In order to determine the impact of age on gene expression, simple linear regression was calculated for each condition (Non-SS and SS), where age was the independent variable and expression of genes (LTF, BMP3, MMP7) quantified in the RNAscope experiment was the dependent variable. p < 0.05 considered to be statistically significant.
References
Delaleu, N., Jonsson, M. V., Appel, S. & Jonsson, R. New concepts in the pathogenesis of Sjogren’s syndrome. Rheum. Dis. Clin. N. Am. 34, 833–845, vii (2008).
Jonsson, M. V., Delaleu, N., Brokstad, K. A., Berggreen, E. & Skarstein, K. Impaired salivary gland function in NOD mice: Association with changes in cytokine profile but not with histopathologic changes in the salivary gland. Arthritis Rheumatol. 54, 2300–2305 (2006).
Margaix-Munoz, M., Bagan, J. V., Poveda, R., Jimenez, Y. & Sarrion, G. Sjogren’s syndrome of the oral cavity. Review and update. Med. Oral Patol. Oral Cir. Bucal 14, E325-330 (2009).
Vivino, F. B. et al. Sjogren’s syndrome: An update on disease pathogenesis, clinical manifestations and treatment. Clin. Immunol. 203, 81–121 (2019).
Rischmueller, M., Tieu, J. & Lester, S. Primary Sjogren’s syndrome. Best Pract. Res. Clin. Rheumatol. 30, 189–220 (2016).
Nocturne, G. & Mariette, X. Advances in understanding the pathogenesis of primary Sjogren’s syndrome. Nat. Rev. Rheumatol. 9, 544–556 (2013).
Tzioufas, A. G., Kapsogeorgou, E. K. & Moutsopoulos, H. M. Pathogenesis of Sjogren’s syndrome: What we know and what we should learn. J. Autoimmun. 39, 4–8 (2012).
Kiripolsky, J., McCabe, L. G. & Kramer, J. M. Innate immunity in Sjogren’s syndrome. Clin. Immunol. 182, 4–13 (2017).
Lessard, C. J. et al. Variants at multiple loci implicated in both innate and adaptive immune responses are associated with Sjogren’s syndrome. Nat. Genet. 45, 1284–1292 (2013).
Haskett, S. et al. Identification of novel CD4+ T cell subsets in the target tissue of Sjogren’s syndrome and their differential regulation by the lymphotoxin/LIGHT signaling axis. J. Immunol. 197, 3806–3819 (2016).
Michael, D. G., Pranzatelli, T. J. F., Warner, B. M., Yin, H. & Chiorini, J. A. Integrated epigenetic mapping of human and mouse salivary gene regulation. J. Dent. Res. 98, 209–217 (2019).
Reale, M. et al. MicroRNA in Sjogren’s syndrome: Their potential roles in pathogenesis and diagnosis. J Immunol. Res. 2018, 7510174 (2018).
Dolcino, M. et al. Long non-coding RNAs modulate Sjogren’s syndrome associated gene expression and are involved in the pathogenesis of the disease. J. Clin. Med. 8, 1349 (2019).
Hjelmervik, T. O., Petersen, K., Jonassen, I., Jonsson, R. & Bolstad, A. I. Gene expression profiling of minor salivary glands clearly distinguishes primary Sjogren’s syndrome patients from healthy control subjects. Arthritis Rheumatol. 52, 1534–1544 (2005).
Ramos, H. L., Valencia-Pacheco, G. & Alcocer-Varela, J. Constitutive STAT3 activation in peripheral CD3(+) cells from patients with primary Sjogren’s syndrome. Scand. J. Rheumatol. 37, 35–39 (2008).
Gottenberg, J. E. et al. Activation of IFN pathways and plasmacytoid dendritic cell recruitment in target organs of primary Sjogren’s syndrome. Proc. Natl. Acad. Sci. U.S.A. 103, 2770–2775 (2006).
Li, H., Ice, J. A., Lessard, C. J. & Sivils, K. L. Interferons in Sjogren’s syndrome: Genes, mechanisms, and effects. Front. Immunol. 4, 290 (2013).
Jonsson, R., Theander, E., Sjostrom, B., Brokstad, K. & Henriksson, G. Autoantibodies present before symptom onset in primary Sjogren syndrome. JAMA 310, 1854–1855 (2013).
Yin, H. et al. Association of bone morphogenetic protein 6 with exocrine gland dysfunction in patients with Sjogren’s syndrome and in mice. Arthritis Rheumatol. 65, 3228–3238 (2013).
Enger, T. B. et al. The Hippo signaling pathway is required for salivary gland development and its dysregulation is associated with Sjogren’s syndrome. Lab. Investig. 93, 1203–1218 (2013).
Barrera, M. J. et al. Sjogren’s syndrome and the epithelial target: A comprehensive review. J. Autoimmun. 42, 7–18 (2013).
Barrera, M. J. et al. Aberrant localization of fusion receptors involved in regulated exocytosis in salivary glands of Sjogren’s syndrome patients is linked to ectopic mucin secretion. J. Autoimmun. 39, 83–92 (2012).
Perez, P. et al. Aberrant localization of ezrin correlates with salivary acini disorganization in Sjogren’s Syndrome. Rheumatology (Oxford) 49, 915–923 (2010).
Bahamondes, V. et al. Changes in Rab3D expression and distribution in the acini of Sjogren’s syndrome patients are associated with loss of cell polarity and secretory dysfunction. Arthritis Rheumatol. 63, 3126–3135 (2011).
Lai, Z. et al. Aquaporin gene therapy corrects Sjogren’s syndrome phenotype in mice. Proc. Natl. Acad. Sci. U.S.A. 113, 5694–5699 (2016).
Vitali, C. et al. Classification criteria for Sjogren’s syndrome: A revised version of the European criteria proposed by the American-European Consensus Group. Ann. Rheum. Dis. 61, 554–558 (2002).
Ding, J. et al. BAFF overexpression increases lymphocytic infiltration in Sjogren’s target tissue, but only inefficiently promotes ectopic B-cell differentiation. Clin. Immunol. 169, 69–79 (2016).
Jezequel, N. et al. Salivary lactoferrin in primary Sjogren’s syndrome. Clin. Exp. Rheumatol. 7, 123–125 (1989).
Gonzalez-Chavez, S. A., Arevalo-Gallegos, S. & Rascon-Cruz, Q. Lactoferrin: Structure, function and applications. Int. J. Antimicrob. Agents 33(301), e301–e308 (2009).
Brejchova, K., Liskova, P., Hrdlickova, E., Filipec, M. & Jirsova, K. Matrix metalloproteinases in recurrent corneal melting associated with primary Sjorgen’s syndrome. Mol. Vis. 15, 2364–2372 (2009).
Matzelle, M. M. et al. Inflammation in arthritis induces expression of BMP3, an inhibitor of bone formation. Scand. J. Rheumatol. 45, 379–383 (2016).
McQuin, C. et al. Cell Profiler 3.0: Next-generation image processing for biology. PLoS Biol. 16, e2005970 (2018).
Jones, T. R. et al. Scoring diverse cellular morphologies in image-based screens with iterative feedback and machine learning. Proc. Natl. Acad. Sci. U.S.A. 106, 1826–1831 (2009).
Jones, T. R. et al. Cell Profiler Analyst: Data exploration and analysis software for complex image-based screens. BMC Bioinform. 9, 482 (2008).
Shiboski, C. H. et al. 2016 American College of Rheumatology/European League Against Rheumatism classification criteria for primary Sjogren’s syndrome: A consensus and data-driven methodology involving three international patient cohorts. Ann. Rheum. Dis. 76, 9–16 (2017).
Asam, S., Neag, G., Berardicurti, O., Gardner, D. & Barone, F. The role of stroma and epithelial cells in primary Sjogren’s syndrome. Rheumatology (Oxford) https://doi.org/10.1093/rheumatology/kez050 (2019).
Gervais, E. M. et al. Changes in the submandibular salivary gland epithelial cell subpopulations during progression of Sjogren’s syndrome-like disease in the NOD/ShiLtJ mouse model. Anat. Rec. (Hoboken) 298, 1622–1634 (2015).
Colafrancesco, S. et al. CXCL13 as biomarker for histological involvement in Sjogren’s syndrome. Rheumatology (Oxford) 59, 165–170 (2020).
Barone, F. et al. IL-22 regulates lymphoid chemokine production and assembly of tertiary lymphoid organs. Proc. Natl. Acad. Sci. U.S.A. 112, 11024–11029 (2015).
Tandon, M., Perez, P., Burbelo, P. D., Calkins, C. & Alevizos, I. Laser microdissection coupled with RNA-seq reveal cell-type and disease-specific markers in the salivary gland of Sjogren’s syndrome patients. Clin. Exp. Rheumatol. 35, 777–785 (2017).
Dogru, M. et al. Lactoferrin in Sjogren’s syndrome. Ophthalmology 114, 2366–2367 (2007).
Donovan, S. M. The role of lactoferrin in gastrointestinal and immune development and function: A preclinical perspective. J. Pediatr. 173(Suppl), S16–S28 (2016).
Perez, P. et al. Increased acinar damage of salivary glands of patients with Sjogren’s syndrome is paralleled by simultaneous imbalance of matrix metalloproteinase 3/tissue inhibitor of metalloproteinases 1 and matrix metalloproteinase 9/tissue inhibitor of metalloproteinases 1 ratios. Arthritis Rheumatol. 52, 2751–2760 (2005).
Fisher, B. A. et al. Standardisation of labial salivary gland histopathology in clinical trials in primary Sjogren’s syndrome. Ann. Rheum. Dis. 76, 1161–1168 (2017).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal [S.l] 17, 10–12 (2011).
Kim, D. et al. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Liao, Y., Smith, G. K. & Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Subramanian, A. et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U.S.A. 102, 15545–15550 (2005).
Acknowledgements
This study was performed at Biogen Idec, and in the Departments of Biochemistry, Boston University School of Medicine and Department of Molecular and Cell Biology, Boston University School of Dental Medicine. Patient data and specimens were obtained from the University of Bergen and University of Oslo Biorepositories, Norway.
Funding
Studies described in this manuscript were supported in part by a Biogen Research Grant (XV and MAK), a National Institute of Dental and Craniofacial Research, National Institutes of Health R21 DE024954 (MAK) and a National Heart, Lung, and Blood Institute, National Institutes of Health R01 HL124392 (XV). ADC was the recipient of the American Association for Dental Research Student Research Fellowship.
Author information
Authors and Affiliations
Contributions
A.D.C., J.L.J., K.S., T.R., M.M., X.V., and M.A.K. participated in the conception and design of the experiments. A.D.C., A.T.-L., V.K., and R.J. performed the experiments and data analysis. A.D.C., J.L.J., K.S., X.V. and M.A.K. wrote the manuscript, which was revised by all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dela Cruz, A., Kartha, V., Tilston-Lunel, A. et al. Gene expression alterations in salivary gland epithelia of Sjögren’s syndrome patients are associated with clinical and histopathological manifestations. Sci Rep 11, 11154 (2021). https://doi.org/10.1038/s41598-021-90569-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-90569-w
- Springer Nature Limited
This article is cited by
-
Primary Sjögren’s syndrome: new perspectives on salivary gland epithelial cells
European Journal of Medical Research (2024)