Abstract
The molecular mechanism of muscle atrophy has been studied a lot, but there is no comprehensive analysis focusing on the denervated muscle atrophy. The gene network that controls the development of denervated muscle atrophy needs further elucidation. We examined differentially expressed genes (DEGs) from five denervated muscle atrophy microarray datasets and predicted microRNAs that target these DEGs. We also included the differentially expressed microRNAs datasets of denervated muscle atrophy in previous studies as background information to identify potential key microRNAs. Finally, we compared denervated muscle atrophy with disuse muscle atrophy caused by other reasons, and obtained the Den-genes which only differentially expressed in denervated muscle atrophy. In this meta-analysis, we obtained 429 up-regulated genes, 525 down-regulated genes and a batch of key microRNAs in denervated muscle atrophy. We found eight important microRNA-mRNA interactions (miR-1/Jun, miR-1/Vegfa, miR-497/Vegfa, miR-23a/Vegfa, miR-206/Vegfa, miR-497/Suclg1, miR-27a/Suclg1, miR-27a/Mapk14). The top five KEGG pathways enriched by Den-genes are Insulin signaling pathway, T cell receptor signaling pathway, MAPK signaling pathway, Toll-like receptor signaling pathway and B cell receptor signaling pathway. Our research has delineated the RNA regulatory network of denervated muscle atrophy, and uncovered the specific genes and terms in denervated muscle atrophy.
Similar content being viewed by others
Introduction
Denervated muscle atrophy refers to muscle atrophy caused by injury of nerves that innervate the muscle. Denervated muscle atrophy is the main cause of motor function loss after peripheral nerve injury. Due to the slow speed of nerve regeneration, it is difficult for regenerated axons to reach the target muscle in a short time. Skeletal muscle will atrophy and fibrosis, and eventually lead to poor recovery of muscle function or even failure to recover1,2,3. Over time, due to muscle fiber necrosis, connective tissue hyperplasia, muscle cell regeneration failure and a large loss of muscle cell, the muscles at nerve endings lose the ability to accept the regenerated motor axons4. In addition, depletion of skeletal muscle satellite cells, changes in related protein metabolism and enzyme activities, vascular bed remodeling and regulation of myogenic factors are all important reasons for denervated muscle atrophy5,6,7,8,9.
At present, the main methods of treating denervated muscle atrophy include electrical stimulation10,11, passive exercise12, and various drug treatments. Up-regulated breakdown of protein in skeletal muscle is a sign of atrophy, so all potential drugs target the proteolytic system to cure or prevent atrophy13. Nevertheless, the FDA has not recommended other drugs to treat muscle atrophy except for megestrol acetate (MA)14, because these drugs cannot effectively target whole proteolytic system, and the potential targets of denervated muscle atrophy have not been fully explored.
The balance between protein synthesis and degradation maintains muscle mass15,16. In atrophic muscles, muscle-specific E3 ubiquitin ligase and muscle-specific ring finger 1 (MuRF1) in the ubiquitin proteasome system (UPS) are activated, promoting protein degradation in an adenosine triphosphate-dependent manner17,18. The biological axis composed of histone deacetylases (HDACs) 4 and 5, transcription factors Dach2 and myogenin can control the expression of many denervation related genes, including atrogin and MuRF119,20,21,22. Myogenin directly mediates the transcriptional activation of the gene encoding E3 ubiquitin ligase, and the lockdown of HDAC4, HDAC5 or myogenin can alleviate denervated muscle atrophy23. In addition, the FoxO family is also an important regulator of atrogin and MuRF1. Overexpression of FoxOs can induce gene expression of atrogin and MuRF1, which will eventually lead to muscle atrophy24,25. Considering the complexity of these processes leading to denervated muscle atrophy, the identification of genes and pathways based on meta-analysis may help to understand the underlying molecular mechanisms.
Denervation, long-term bed rest, unloading of hind limbs, immobilization or microgravity can cause disuse muscle atrophy, of which the common features are decreased cross-sectional area of muscle fibers, decreased muscle strength, increased insulin resistance, and the transition of fiber types from slow to fast26. Decreased protein synthesis and increased protein degradation are the main reasons for the rapid loss of muscle protein due to denervation26. Current research focuses on the general signaling pathways and common key regulators that mediate disuse muscle atrophy26,27,28, but obviously, denervation is also accompanied by the loss of neurotrophic factors and a series of changes brought about by nerve injury. These denervation-specific pathological changes might be accompanied by specific transcriptome changes which remain to be discovered.
We systematically searched and integrated the gene expression data of denervated muscle atrophy to construct the microRNA-mRNA regulatory network. By comparing denervated muscle atrophy with disused muscle atrophy caused by other reasons, we have discovered the genes and terms that are specifically differentially expressed in denervated muscle atrophy.
Results
Data screening and inclusion criteria
This study obtained five gene expression data of denervated muscle atrophy after screening (Fig. 1)29,30,31. The details of these data were aggregated into a table (Table 1) and they were all from mouse models in different muscles, including tibialis anterior muscle, gastrocnemius muscle, and gastrocnemius triceps muscle. We only included the data sets that denervated for 7–14 days, because the muscle with denervation greater than 7 days has entered the stage of pathological atrophy31, and the gene expression profiles denervated for 7/14 days have high similarity, which can ensure the homogeneity of the included data sets32.
DEGs in denervated muscle atrophy
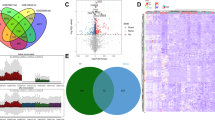
By crossing five data sets, we obtained 429 up-regulated mRNAs and 525 down-regulated mRNAs (Fig. 2a,b) (Table S1), GO analysis revealed the biological role of DEGs in denervated muscle atrophy (Table S2). The analysis showed the pathways enriched by up-regulated genes included cellular process, cellular developmental process, cell differentiation, anatomical structure development, system development, proteasome complex, cytoplasm, cytosol, intracellular part, intracellular, protein binding, binding, threonine-type peptidase activity, threonine-type endopeptidase activity and tubulin binding. The pathways enriched by down-regulated genes include generation of precursor metabolites and energy, energy derivation by oxidation of organic compounds, acetyl-CoA metabolic process, cellular respiration, aerobic respiration, mitochondrion cytoplasm, mitochondrial part, cytoplasmic part, mitochondrial membrane, catalytic activity, oxidoreductase activity, phosphoric ester hydrolase activity, cofactor binding and coenzyme binding (Fig. 2c,d). In order to avoid losing information, we performed GO analysis on the DEG of each data set, and then obtained GO terms that were commonly enriched in each data set (Table 2). We found that the up-regulated GO terms were mostly concentrated in cellular components, and the down-regulated GO terms were still dominated by energy metabolism.
DEGs and GO terms in denervated muscle atrophy were identified. Venn diagrams for (a) up-regulated mRNAs and (b) down-regulated mRNAs were drawn based on 5 denervated muscle atrophy datasets. GO analysis of DEGs in denervated muscle atrophy were conducted for (c) up- and (d) down-regulated mRNAs respectively (Each figure shows the top10 terms with the lowest p value). BP biological process, CC cellular component, MF molecular function.
A protein–protein Interaction (PPI) network was mapped based on DEGs to bring in more function-related proteins, and the nodes were sorted according to the degree of interaction between the nodes (Fig. 3). The top10 proteins are Gapdh, Mapk14, Jun, Cat, Casp3, Vegfa, Decr1, CS and Suclg1. The function and location of these genes were annotated respectively, and most of these genes are related to energy metabolism (Table 3).
PPI network in denervated muscle atrophy. (a) PPI network, the color was used to reflect the level of degree, the higher the degree, the darker the node color. (b) The top10 DEGs with the highest degree. (c) Partial enlarged view of PPI network. STRING v11.0 was used to generate protein interactions, and the resulting network was visualized using Cytoscape v3.7.2.
Identification of potential microRNAs in denervated muscle atrophy
MiRNA-mRNA target prediction identified miRNAs targeting down-regulated mRNAs and those targeting up-regulated mRNAs for den-7 days and den-14 days respectively. By retrieving, we obtained microRNA data sets denervated 7/14 days31,32, we intersected the up-regulated miRNAs in these two data sets with the miRNAs targeting down-regulated mRNAs in this Meta-analysis to obtain 9 miRNAs for den-7 days. Similarly, we obtained 2 down-regulated miRNAs for den-7 days, 5 up-regulated miRNAs and 3 down-regulated miRNAs for den-14 days (Fig. 4, Table 4). We predicted the target genes of these key miRNAs and found eight microRNA-mRNA interactions of which the mRNAs are the top 10 denervation-related mRNAs identified in this research (miR-1/Jun, miR-1/Vegfa, miR-497/Vegfa, miR-23a/Vegfa, miR-206/Vegfa, miR-497/Suclg1, miR-27a/Suclg1, miR-27a/Mapk14). Notably, seven interactions showed an opposite direction of expression between microRNAs and mRNAs (Table 5).
MicroRNAs identified in denervated muscle atrophy. MicroRNAs predicted as targeting mRNAs retrieved by our meta-analysis were further compared with microRNAs identified as differentially expressed in previous studies of denervated muscle atrophy for 7 days and 14 days respectively. (a) Venn for up-regulated miRNAs for Den-7 days. (b) Venn for down-regulated miRNAs for Den-7 days. (c) Venn for up-regulated miRNAs for Den-14 days. (d) Venn for down-regulated miRNAs for Den-14 days. Den-7 days denervation for 7 days, Den-14 days denervation for 14 days.
Specific genes in denervated muscle atrophy
By retrieving, we got three data sets of disuse muscle atrophy caused by unloading or casting (Table 6), genes only differentially expressed in denervated muscle atrophy were regarded as denervation specific genes, called den-genes. We got 187 up-regulated den-genes and 180 down-regulated den-genes (Fig. 5), then performed KEGG enrichment analysis on these genes and got top10 most enriched pathways (Fig. 6). We searched and found the expression of neurotrophic factors (NGF, BDNF, NT-3, NT-4, CNTF, Neuregulin-1 and Neuritin) changed in nerves or muscles after nerve injury and there was evidence that Insulin signaling pathway, MAPK signaling pathway, Neurotrophin signaling pathway, T cell receptor signaling pathway, Toll-like receptor signaling pathway, and B cell receptor signaling pathway were regulated by the above neurotrophic factors (Table 7). Furthermore, most of these terms in skeletal muscle showed different expression after electrical stimulation and Insulin signaling pathway, MAPK signaling pathway, Toll-like receptor signaling pathway, Neurotrophin signaling pathway, and VEGF signaling pathway show regulatory effects on ion channels and connexins (Table 8).
Specific mRNAs identified in denervated muscle atrophy. DEGs in denervated muscle atrophy retrieved by our meta-analysis (blue circle) were further compared with DEGs in disuse muscle atrophy caused by 7-day hindlimb casting (purple circle), 14-day hindlimb casting (red circle) and hindlimb unloading (green circle). (a) Venn for up-regulated genes. (b) Venn for down-regulated genes. Den denervation, cast 7 7-day hindlimb casting, cast 14 14-day hindlimb casting.
KEGG analysis of den-genes. Lines highlight KEGG network, with Jun, Mapk9, Calm2, Gys1, Ppp1cc, and Akt2 presenting the highest degree of interactions. Colors highlight KEGG pathways of the circle network components. The larger the circle, the higher the interaction degree identified. Cytoscape v. 3.7.2 was used to generate the resulting network. Permission has been obtained from Kanehisa laboratories for using KEGG pathway database33.
Discussion
In this study, we collected and aggregated gene expression data sets of denervated muscle atrophy, and obtained 429 up-regulated mRNAs and 525 down-regulated mRNAs. We drew a PPI network, and sorted the nodes according to the interaction degree. The top10 proteins were Gapdh, Mapk14, Jun, Cat, Casp3, Vegfa, Decr1, Cs and Suclg1. Mapk14, Jun, and Casp3 have been confirmed to be important in denervated muscle atrophy. (1) Mapk14: the three main categories of mitogen-activated protein kinase (MAPK) family proteins are as follows: extracellular signal-regulated kinase (ERK), c-Jun N-terminal kinase (JNK) and p38 MAPK (Mapk14). The members of the p38 MAPK family (p38α, p38β, p38γ and p38δ MAPK) act as transducers of cellular stress and various non-stress-related stimuli. Therefore, the p38 MAPK pathway has multiple functions and is involved in various cellular processes, including aging, apoptosis, cell cycle arrest, inflammation, and tumorigenesis63. The p38 MAPK pathway can mediate the expression of MuRF1 and Atrogin164,65, and knockdown of p38αMAPK can inhibit muscle atrophy caused by denervation66. (2) Jun: the protein deacetylase HDAC4 is strongly induced in muscles affected by motor neuron diseases such as ALS19. Denervation-induced HDAC4 activates AP1 (Jun) transcription factor by stimulating MAPK signaling to promote denervated muscle atrophy67. (3) Casp3: in the ubiquitin–proteasome pathway and apoptosis process, the activation of Caspase-3 (Casp3) is a common phenomenon. Caspase-3 regulates denervation-induced signal transduction through the mitochondrial-related cell death/apoptosis pathway, resulting in the loss of muscle mass. The lack of caspase-3 has a protective effect on denervated muscle atrophy68. The other proteins Gapdh, Cat, Vegfa, Decr1, Cs, and Suclg1 are mostly important links in redox and energy metabolism pathways69,70,71,72,73, and they may be potential targets in denervated muscle atrophy for treatment.
We included the differentially expressed miRNA expression data of denervated muscle atrophy in previous studies as background information31,32, identified the potential key microRNAs and eight microRNA-mRNA interactions of which the mRNAs are the top 10 denervation-related mRNAs identified in this research (miR-1/Jun, miR-1/Vegfa, miR-497/Vegfa, miR-23a/Vegfa, miR-206/Vegfa, miR-497/Suclg1, miR-27a/Suclg1, miR-27a/Mapk14). Most of the Key miRNAs we obtained showed important regulatory effects on skeletal muscle (Table 7). After 7-day hind limbs unloading in mice, the serum levels of muscle-specific miRNAs such as miR-1, miR-23a, miR-206 increased significantly, and could induce severe muscle atrophy74. MiR-1 has been confirmed to play an important role in skeletal muscle development75,76. MiR-23a can target Atrogin-1 and MuRF1 and inhibit their translation, and the ectopic expression of miR-23a can protect muscles from atrophy in vitro and in vivo, indicating that miR-23a is a key regulator of muscle atrophy77. Overexpression of miR-27a in mice with chronic kidney disease attenuated muscle loss, improved grip strength and reduced the expression of FoxO1, MuRF1and Atrogin140.
Denervation, long-term bed rest, unloading of hind limbs, immobilization or microgravity can cause disuse muscle atrophy. Current research focuses on the general signaling pathways and common key regulators that mediate disuse muscle atrophy26,27,28, but obviously, denervation is also accompanied by the loss of neurotrophic factors and a series of changes brought about by nerve injury. These denervation-specific pathological changes might be accompanied by specific transcriptome changes which remain to be discovered.
The nervous system controls skeletal muscle through two mechanisms: (1) neuromotor control, which causes muscle contraction through excitation and contraction coupling. (2) Neurotrophic control, which regulates muscles by releasing soluble factors from the nerve endings of motor neurons on the NMJ (Neuromuscular junction)78. NGF, BDNF, NT-3, NT-4, CNTF, Neuregulin-1 and Neuritin are neurotrophic factors differentially expressed in nerves or skeletal muscle after nerve injury79, and they were found to regulate Insulin signaling pathway, MAPK Signaling pathway, Neurotrophin signaling pathway, T cell receptor signaling pathway, Toll-like receptor signaling pathway and B cell receptor signaling pathway which enriched by den-genes (Table 8). The specific changes in skeletal muscle after nerve transection include increased membrane permeability, decreased membrane potential and increased membrane excitability. Most of these are caused by changes in the expression of ion channels and the insertion of connexins 39, 43, and 45 into the muscle membrane, which mediate skeletal muscle atrophy78,80. Insulin signaling pathway, MAPK signaling pathway, Toll-like receptor signaling pathway, Neurotrophin signaling pathway and VEGF signaling pathway showed the potential to regulate ion channels and connexins. Insulin signaling pathway and Toll-like receptor signaling pathway can affect the formation of neuromuscular junctions (Table 9). Therefore, Insulin signaling pathway and Toll-like receptor signaling pathway, as metabolic or inflammation-related terms, have been seldom studied in the neuromuscular systerm, but this study found that they may have important research value in the denervated muscle atrophy.
Current research focuses on the general signaling pathways and common key regulators that mediate disuse muscle atrophy, but obviously, denervation is also accompanied by the loss of neurotrophic factors and a series of changes brought about by nerve injury. These denervation-specific pathological changes must be accompanied by specific transcriptome changes which remain to be discovered. We systematically searched and integrated the gene expression data of denervated muscle atrophy to construct the microRNA-mRNA regulatory network. By comparing denervated muscle atrophy with disused muscle atrophy caused by other reasons, we have discovered the genes and terms that are specifically differentially expressed in denervated muscle atrophy.
Methods
Inclusion criteria for gene expression data
We performed a meta-analysis following the PRISMA Statement97, and found the gene expression data of denervated muscle atrophy by searching NCBI-GEO (http://www.ncbi.nlm.nih.gov/geo/) and PubMed (http://www.ncbi.nlm.nih.gov/pubmed). The keywords used were: “Denervated atrophy”, “Denervation AND muscle”, “nerve AND muscle”, and “Denervation AND muscle AND sequencing”. These meta-analysis searches comprised studies published between 2010 and 2020. Our inclusion criteria were (1) gene expression data in skeletal muscle with peripheral nerve injury, (2) the number of samples in each group should be greater than two, (3) the duration of denervation was 7 to 14 days, (4) all types of skeletal muscle were considered, (5) the inclusion of normal tissues for comparison, and (6) all gene expression analysis platforms were considered. Our exclusion criteria were (1) non-muscle samples, (2) non-mRNA datasets, and (3) review studies.
And we also searched NCBI-GEO (http://www.ncbi.nlm.nih.gov/geo/) and PubMed (http://www.ncbi.nlm.nih.gov/pubmed) to find the gene expression data of disuse muscle atrophy. The keywords used were: “Disuse atrophy”, “Disuse AND muscle”, “unloading AND muscle”, and “cast AND muscle”. These meta-analysis searches comprised studies published between 2010 and 2020. Our inclusion criteria were (1) gene expression data in skeletal muscle with disuse atrophy, (2) the number of samples in each group should be greater than two, (3) the duration of intervention was 7 to 14 days, (4) all types of skeletal muscle were considered, (5) the inclusion of normal tissues for comparison, and (6) all gene expression analysis platforms were considered. Our exclusion criteria were (1) non-muscle samples, (2) non-mRNA datasets, and (3) review studies.
In this study, each individual dataset was processed using edgeR packages to compare control samples and experiment samples to screen DEGs. The Benjamini & Hochberg false discovery rate method was used for p-value adjustment, and the default adj-p-value significance level cut-off is 0.05.
Meta-analysis of global gene expression data in denervated muscle atrophy
Meta-analysis was performed on the five datasets of denervated muscle atrophy using Vote counting generic ways of combining information98, and the DEGs that were common between all datasets were identified. The results were visualized using Venn diagrams by a graphing software (ORIGIN2019; OriginLab, Northampton, MA).
GO and KEGG enrichment analysis of DEGs
The DEGs were used to identify over-represented gene ontology categories and KEGG pathways by using DAVID 6.7 (https://david-d.ncifcrf.gov/). The p value has been corrected using FDR (false discovery rate). GO/KEGG enrichment with p < 0.05 was regarded as statistically significant and those with number of genes < 10 were removed. Bar chart of GO terms was drawn by Microsoft Excel 2016. KEGG network was visualized by Cytoscape software v. 3.7.2 54.
Protein–protein interaction (PPI) network construction for common DEGs
The DEGs were subjected to STRING v.11.0 database99 analysis to construct PPI networks. Minimum required interaction score was 0.4 (medium confidence). PPI networks were visualized by Cytoscape software v. 3.7.2100 and the degree which indicates the number of interactions of each node was calculated by cytoHubba101.
Identification of microRNAs as potential modulators of DEGs
The DEGs identified in our meta-analysis were used for microRNA prediction by DIANA-Tarbase102 to identify potential regulators in denervated muscle atrophy. Visualization of microRNA-gene interaction networks were generated using Cytoscape v3.7.2100 and the degree which indicates the number of interactions of each node was calculated by cytoHubba101.
References
Frostick, S. P. The physiological and metabolic consequences of muscle denervation. Int. Angiol. 14, 278–287 (1995).
Kraft, G. H. Fibrillation potential amplitude and muscle atrophy following peripheral nerve injury. Muscle Nerve 13, 814–821. https://doi.org/10.1002/mus.880130907 (1990).
Muheremu, A. & Ao, Q. Past, present, and future of nerve conduits in the treatment of peripheral nerve injury. Biomed. Res. Int. 2015, 237507. https://doi.org/10.1155/2015/237507 (2015).
Veltri, K., Kwiecien, J. M., Minet, W., Fahnestock, M. & Bain, J. R. Contribution of the distal nerve sheath to nerve and muscle preservation following denervation and sensory protection. J. Reconstr. Microsurg. 21, 57–70. https://doi.org/10.1055/s-2005-862783 (2005).
Viguie, C. A., Lu, D. X., Huang, S. K., Rengen, H. & Carlson, B. M. Quantitative study of the effects of long-term denervation on the extensor digitorum longus muscle of the rat. Anat. Rec. 248, 346–354. https://doi.org/10.1002/(SICI)1097-0185(199707)248:3%3c346::AID-AR7%3e3.0.CO;2-N (1997).
Liu, J. et al. Vascular bed-specific regulation of the von Willebrand factor promoter in the heart and skeletal muscle. Blood 117, 342–351. https://doi.org/10.1182/blood-2010-06-287987 (2011).
Nakao, R. et al. Atypical expression of circadian clock genes in denervated mouse skeletal muscle. Chronobiol. Int. 32, 486–496. https://doi.org/10.3109/07420528.2014.1003350 (2015).
Voytik, S. L., Przyborski, M., Badylak, S. F. & Konieczny, S. F. Differential expression of muscle regulatory factor genes in normal and denervated adult rat hindlimb muscles. Dev. Dyn. 198, 214–224. https://doi.org/10.1002/aja.1001980307 (1993).
Rodrigues Ade, C. & Schmalbruch, H. Satellite cells and myonuclei in long-term denervated rat muscles. Anat. Rec. 243, 430–437. https://doi.org/10.1002/ar.1092430405 (1995).
Modlin, M. et al. Electrical stimulation of denervated muscles: First results of a clinical study. Artif. Organs 29, 203–206. https://doi.org/10.1111/j.1525-1594.2005.29035.x (2005).
Helgason, T. et al. Monitoring muscle growth and tissue changes induced by electrical stimulation of denervated degenerated muscles with CT and stereolithographic 3D modeling. Artif. Organs 29, 440–443. https://doi.org/10.1111/j.1525-1594.2005.29073.x (2005).
Madsen, K., Franch, J. & Clausen, T. Effects of intensified endurance training on the concentration of Na, K-ATPase and Ca-ATPase in human skeletal muscle. Acta Physiol. Scand. 150, 251–258. https://doi.org/10.1111/j.1748-1716.1994.tb09684.x (1994).
Dutt, V., Gupta, S., Dabur, R., Injeti, E. & Mittal, A. Skeletal muscle atrophy: Potential therapeutic agents and their mechanisms of action. Pharmacol. Res. 99, 86–100. https://doi.org/10.1016/j.phrs.2015.05.010 (2015).
Busquets, S. et al. Megestrol acetate: Its impact on muscle protein metabolism supports its use in cancer cachexia. Clin. Nutr. 29, 733–737. https://doi.org/10.1016/j.clnu.2010.06.003 (2010).
Bonaldo, P. & Sandri, M. Cellular and molecular mechanisms of muscle atrophy. Dis. Model Mech. 6, 25–39. https://doi.org/10.1242/dmm.010389 (2013).
Goodman, C. A., Mayhew, D. L. & Hornberger, T. A. Recent progress toward understanding the molecular mechanisms that regulate skeletal muscle mass. Cell Signal 23, 1896–1906. https://doi.org/10.1016/j.cellsig.2011.07.013 (2011).
Bodine, S. C. et al. Identification of ubiquitin ligases required for skeletal muscle atrophy. Science 294, 1704–1708. https://doi.org/10.1126/science.1065874 (2001).
Gomes, M. D., Lecker, S. H., Jagoe, R. T., Navon, A. & Goldberg, A. L. Atrogin-1, a muscle-specific F-box protein highly expressed during muscle atrophy. Proc. Natl. Acad. Sci. U.S.A. 98, 14440–14445. https://doi.org/10.1073/pnas.251541198 (2001).
Cohen, T. J. et al. The histone deacetylase HDAC4 connects neural activity to muscle transcriptional reprogramming. J. Biol. Chem. 282, 33752–33759. https://doi.org/10.1074/jbc.M706268200 (2007).
Tang, H. & Goldman, D. Activity-dependent gene regulation in skeletal muscle is mediated by a histone deacetylase (HDAC)-Dach2-myogenin signal transduction cascade. Proc. Natl. Acad. Sci. U.S.A. 103, 16977–16982. https://doi.org/10.1073/pnas.0601565103 (2006).
Tang, H. et al. A histone deacetylase 4/myogenin positive feedback loop coordinates denervation-dependent gene induction and suppression. Mol. Biol. Cell 20, 1120–1131. https://doi.org/10.1091/mbc.E08-07-0759 (2009).
Moresi, V. et al. Myogenin and class II HDACs control neurogenic muscle atrophy by inducing E3 ubiquitin ligases. Cell 143, 35–45. https://doi.org/10.1016/j.cell.2010.09.004 (2010).
Macpherson, P. C., Wang, X. & Goldman, D. Myogenin regulates denervation-dependent muscle atrophy in mouse soleus muscle. J. Cell Biochem. 112, 2149–2159. https://doi.org/10.1002/jcb.23136 (2011).
Sandri, M. et al. Foxo transcription factors induce the atrophy-related ubiquitin ligase atrogin-1 and cause skeletal muscle atrophy. Cell 117, 399–412. https://doi.org/10.1016/s0092-8674(04)00400-3 (2004).
Kamei, Y. et al. Skeletal muscle FOXO1 (FKHR) transgenic mice have less skeletal muscle mass, down-regulated Type I (slow twitch/red muscle) fiber genes, and impaired glycemic control. J. Biol. Chem. 279, 41114–41123. https://doi.org/10.1074/jbc.M400674200 (2004).
Zhang, P., Chen, X. & Fan, M. Signaling mechanisms involved in disuse muscle atrophy. Med. Hypotheses 69, 310–321. https://doi.org/10.1016/j.mehy.2006.11.043 (2007).
Fisher, A. G. et al. Transcriptomic and epigenetic regulation of disuse atrophy and the return to activity in skeletal muscle. FASEB J. 31, 5268–5282. https://doi.org/10.1096/fj.201700089RR (2017).
Urso, M. L. Disuse atrophy of human skeletal muscle: Cell signaling and potential interventions. Med. Sci. Sports Exerc. 41, 1860–1868. https://doi.org/10.1249/MSS.0b013e3181a6458a (2009).
Furlow, J. D. et al. Altered gene expression patterns in muscle ring finger 1 null mice during denervation- and dexamethasone-induced muscle atrophy. Physiol. Genomics 45, 1168–1185. https://doi.org/10.1152/physiolgenomics.00022.2013 (2013).
Sartori, R. et al. BMP signaling controls muscle mass. Nat. Genet. 45, 1309–1318. https://doi.org/10.1038/ng.2772 (2013).
Weng, J., Zhang, P., Yin, X. & Jiang, B. The whole transcriptome involved in denervated muscle atrophy following peripheral nerve injury. Front. Mol. Neurosci. 11, 69. https://doi.org/10.3389/fnmol.2018.00069 (2018).
Soares, R. J. et al. Involvement of microRNAs in the regulation of muscle wasting during catabolic conditions. J. Biol. Chem. 289, 21909–21925. https://doi.org/10.1074/jbc.M114.561845 (2014).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Zhang, A. et al. miRNA-23a/27a attenuates muscle atrophy and renal fibrosis through muscle-kidney crosstalk. J Cachexia Sarcopenia Muscle 9, 755–770. https://doi.org/10.1002/jcsm.12296 (2018).
Qiu, H. et al. Regulatory axis of miR-195/497 and HMGA1-Id3 governs muscle cell proliferation and differentiation. Int. J. Biol. Sci. 13, 157–166. https://doi.org/10.7150/ijbs.17440 (2017).
Zhang, Y. et al. Beta-hydroxy-beta-methylbutyrate-induced upregulation of miR-199a-3p contributes to slow-to-fast muscle fiber type conversion in mice and C2C12 cells. J. Agric. Food Chem. 68, 530–540. https://doi.org/10.1021/acs.jafc.9b05104 (2020).
Hu, X. et al. Bta-miR-24-3p controls the myogenic differentiation and proliferation of fetal, bovine, skeletal muscle-derived progenitor cells by targeting ACVR1B. Animals (Basel). https://doi.org/10.3390/ani9110859 (2019).
Ling, Y. H. et al. miR-27b regulates myogenic proliferation and differentiation by targeting Pax3 in goat. Sci. Rep. 8, 3909. https://doi.org/10.1038/s41598-018-22262-4 (2018).
Chemello, F. et al. Transcriptomic analysis of single isolated myofibers identifies miR-27a-3p and miR-142–3p as regulators of metabolism in skeletal muscle. Cell Rep. 26, 3784–3797. https://doi.org/10.1016/j.celrep.2019.02.105 (2019).
Wang, B. et al. MicroRNA-23a and MicroRNA-27a mimic exercise by ameliorating CKD-induced muscle atrophy. J. Am. Soc. Nephrol. 28, 2631–2640. https://doi.org/10.1681/ASN.2016111213 (2017).
Przanowska, R. K. et al. miR-206 family is important for mitochondrial and muscle function, but not essential for myogenesis in vitro. FASEB J. 34, 7687–7702. https://doi.org/10.1096/fj.201902855RR (2020).
Williams, A. H. et al. MicroRNA-206 delays ALS progression and promotes regeneration of neuromuscular synapses in mice. Science 326, 1549–1554. https://doi.org/10.1126/science.1181046 (2009).
Morgoulis, D. et al. sPIF promotes myoblast differentiation and utrophin expression while inhibiting fibrosis in Duchenne muscular dystrophy via the H19/miR-675/let-7 and miR-21 pathways. Cell Death Dis. 10, 82. https://doi.org/10.1038/s41419-019-1307-9 (2019).
Sato, T., Yamamoto, T. & Sehara-Fujisawa, A. miR-195/497 induce postnatal quiescence of skeletal muscle stem cells. Nat. Commun. 5, 4597. https://doi.org/10.1038/ncomms5597 (2014).
Chen, J. F. et al. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat. Genet. 38, 228–233. https://doi.org/10.1038/ng1725 (2006).
Saika, T. et al. Effects of nerve crush and transection on mRNA levels for nerve growth factor receptor in the rat facial motoneurons. Brain Res. Mol. Brain Res. 9, 157–160. https://doi.org/10.1016/0169-328x(91)90142-k (1991).
Zhou, X. F. et al. Satellite-cell-derived nerve growth factor and neurotrophin-3 are involved in noradrenergic sprouting in the dorsal root ganglia following peripheral nerve injury in the rat. Eur. J. Neurosci. 11, 1711–1722. https://doi.org/10.1046/j.1460-9568.1999.00589.x (1999).
Abe, S. et al. Induction of nerve growth factor mRNA in a rat dorsal root ganglion after application of a tourniquet. Acta Neuropathol. 108, 183–188. https://doi.org/10.1007/s00401-004-0870-y (2004).
Amano, T., Yamakuni, T., Okabe, N., Sakimura, K. & Takahashi, Y. Production of nerve growth factor in rat skeletal muscle. Neurosci. Lett. 132, 5–7. https://doi.org/10.1016/0304-3940(91)90418-s (1991).
Wu, C. et al. A functional dynein-microtubule network is required for NGF signaling through the Rap1/MAPK pathway. Traffic 8, 1503–1520. https://doi.org/10.1111/j.1600-0854.2007.00636.x (2007).
Griesbeck, O., Parsadanian, A. S., Sendtner, M. & Thoenen, H. Expression of neurotrophins in skeletal muscle: Quantitative comparison and significance for motoneuron survival and maintenance of function. J. Neurosci. Res. 42, 21–33. https://doi.org/10.1002/jnr.490420104 (1995).
Meyer, M., Matsuoka, I., Wetmore, C., Olson, L. & Thoenen, H. Enhanced synthesis of brain-derived neurotrophic factor in the lesioned peripheral nerve: Different mechanisms are responsible for the regulation of BDNF and NGF mRNA. J. Cell Biol. 119, 45–54. https://doi.org/10.1083/jcb.119.1.45 (1992).
Chen, T. et al. Brain-derived neurotrophic factor increases synaptic protein levels via the MAPK/Erk signaling pathway and Nrf2/Trx axis following the transplantation of neural stem cells in a rat model of traumatic brain injury. Neurochem. Res. 42, 3073–3083. https://doi.org/10.1007/s11064-017-2340-7 (2017).
Omura, T. et al. Different expressions of BDNF, NT3, and NT4 in muscle and nerve after various types of peripheral nerve injuries. J. Peripher Nerv. Syst. 10, 293–300. https://doi.org/10.1111/j.1085-9489.2005.10307.x (2005).
Funakoshi, H. et al. Differential expression of mRNAs for neurotrophins and their receptors after axotomy of the sciatic nerve. J. Cell Biol. 123, 455–465. https://doi.org/10.1083/jcb.123.2.455 (1993).
Ye, J. et al. Effect of NT-3 on repair of spinal cord injury through the MAPK signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 24, 2165–2172. https://doi.org/10.26355/eurrev_202003_20481 (2020).
Smith, G. M., Rabinovsky, E. D., McManaman, J. L. & Shine, H. D. Temporal and spatial expression of ciliary neurotrophic factor after peripheral nerve injury. Exp. Neurol. 121, 239–247. https://doi.org/10.1006/exnr.1993.1091 (1993).
Martin, D. et al. Cachectic effect of ciliary neurotrophic factor on innervated skeletal muscle. Am. J. Physiol. 271, R1422–R1428. https://doi.org/10.1152/ajpregu.1996.271.5.R1422 (1996).
Kopp, D. M., Trachtenberg, J. T. & Thompson, W. J. Glial growth factor rescues Schwann cells of mechanoreceptors from denervation-induced apoptosis. J. Neurosci. 17, 6697–6706 (1997).
Kim, H. G. et al. Neuregulin 1 up-regulates the expression of nicotinic acetylcholine receptors through the ErbB2/ErbB3-PI3K-MAPK signaling cascade in adult autonomic ganglion neurons. J. Neurochem. 124, 502–513. https://doi.org/10.1111/jnc.12109 (2013).
Azuchi, Y. et al. Role of neuritin in retinal ganglion cell death in adult mice following optic nerve injury. Sci. Rep. 8, 10132. https://doi.org/10.1038/s41598-018-28425-7 (2018).
Yao, J. J., Zhao, Q. R., Lu, J. M. & Mei, Y. A. Functions and the related signaling pathways of the neurotrophic factor neuritin. Acta Pharmacol. Sin. 39, 1414–1420. https://doi.org/10.1038/aps.2017.197 (2018).
Cuenda, A. & Rousseau, S. p38 MAP-kinases pathway regulation, function and role in human diseases. Biochim. Biophys. Acta 1773, 1358–1375. https://doi.org/10.1016/j.bbamcr.2007.03.010 (2007).
Derbre, F. et al. Inhibition of xanthine oxidase by allopurinol prevents skeletal muscle atrophy: Role of p38 MAPKinase and E3 ubiquitin ligases. PLoS ONE 7, e46668. https://doi.org/10.1371/journal.pone.0046668 (2012).
Childs, T. E., Spangenburg, E. E., Vyas, D. R. & Booth, F. W. Temporal alterations in protein signaling cascades during recovery from muscle atrophy. Am. J. Physiol. Cell Physiol. 285, C391–C398. https://doi.org/10.1152/ajpcell.00478.2002 (2003).
Yuasa, K. et al. Targeted ablation of p38alpha MAPK suppresses denervation-induced muscle atrophy. Sci. Rep. 8, 9037. https://doi.org/10.1038/s41598-018-26632-w (2018).
Choi, M. C. et al. A direct HDAC4-MAP kinase crosstalk activates muscle atrophy program. Mol. Cell 47, 122–132. https://doi.org/10.1016/j.molcel.2012.04.025 (2012).
Plant, P. J., Bain, J. R., Correa, J. E., Woo, M. & Batt, J. Absence of caspase-3 protects against denervation-induced skeletal muscle atrophy. J. Appl. Physiol. 107, 224–234. https://doi.org/10.1152/japplphysiol.90932.2008 (2009).
Nakajima, H. et al. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) aggregation causes mitochondrial dysfunction during oxidative stress-induced cell death. J. Biol. Chem. 292, 4727–4742. https://doi.org/10.1074/jbc.M116.759084 (2017).
Nomura, K., Ono, K. & Ono, S. CAS-1, a C. elegans cyclase-associated protein, is required for sarcomeric actin assembly in striated muscle. J. Cell Sci. 125, 4077–4089. https://doi.org/10.1242/jcs.104950 (2012).
Blomme, A. et al. 2,4-dienoyl-CoA reductase regulates lipid homeostasis in treatment-resistant prostate cancer. Nat. Commun. 11, 2508. https://doi.org/10.1038/s41467-020-16126-7 (2020).
Wiegand, G. & Remington, S. J. Citrate synthase: Structure, control, and mechanism. Annu. Rev. Biophys. Biophys. Chem. 15, 97–117. https://doi.org/10.1146/annurev.bb.15.060186.000525 (1986).
El-Hattab, A. W. & Scaglia, F. Mitochondrial DNA depletion syndromes: Review and updates of genetic basis, manifestations, and therapeutic options. Neurotherapeutics 10, 186–198. https://doi.org/10.1007/s13311-013-0177-6 (2013).
Wang, F. et al. Serum miRNAs miR-23a, 206, and 499 as potential biomarkers for skeletal muscle atrophy. Biomed. Res. Int. 2017, 8361237. https://doi.org/10.1155/2017/8361237 (2017).
Wust, S. et al. Metabolic maturation during muscle stem cell differentiation is achieved by miR-1/133a-mediated inhibition of the Dlk1-Dio3 mega gene cluster. Cell Metab. 27, 1026–1039. https://doi.org/10.1016/j.cmet.2018.02.022 (2018).
Kukreti, H. et al. Muscle-specific microRNA1 (miR1) targets heat shock protein 70 (HSP70) during dexamethasone-mediated atrophy. J. Biol. Chem. 288, 6663–6678. https://doi.org/10.1074/jbc.M112.390369 (2013).
Sacheck, J. M. et al. Rapid disuse and denervation atrophy involve transcriptional changes similar to those of muscle wasting during systemic diseases. FASEB J. 21, 140–155. https://doi.org/10.1096/fj.06-6604com (2007).
Cisterna, B. A., Cardozo, C. & Saez, J. C. Neuronal involvement in muscular atrophy. Front. Cell Neurosci. 8, 405. https://doi.org/10.3389/fncel.2014.00405 (2014).
Kingham, P. J. & Terenghi, G. Bioengineered nerve regeneration and muscle reinnervation. J. Anat. 209, 511–526. https://doi.org/10.1111/j.1469-7580.2006.00623.x (2006).
Cea, L. A. et al. De novo expression of connexin hemichannels in denervated fast skeletal muscles leads to atrophy. Proc. Natl. Acad. Sci. U.S.A. 110, 16229–16234. https://doi.org/10.1073/pnas.1312331110 (2013).
La, G. et al. Proteomics and transcriptomics analysis reveals clues into the mechanism of the beneficial effect of electrical stimulation on rat denervated gastrocnemius muscle. Cell Physiol. Biochem. 52, 769–786. https://doi.org/10.33594/000000054 (2019).
Hung, W. L. et al. Attenuation of insulin signalling contributes to FSN-1-mediated regulation of synapse development. EMBO J. 32, 1745–1760. https://doi.org/10.1038/emboj.2013.91 (2013).
Yao, J. J. et al. Neuritin activates insulin receptor pathway to up-regulate Kv42-mediated transient outward K+ current in rat cerebellar granule neurons. J. Biol. Chem. 287, 41534–41545. https://doi.org/10.1074/jbc.M112.390260 (2012).
Massaro, C. M., Pielage, J. & Davis, G. W. Molecular mechanisms that enhance synapse stability despite persistent disruption of the spectrin/ankyrin/microtubule cytoskeleton. J. Cell Biol. 187, 101–117. https://doi.org/10.1083/jcb.200903166 (2009).
Jeong Nam, Y., Kim, A., Sung Lee, M., Suep Sohn, D. & Soo Lee, C. KATP channel block inhibits the Toll-like receptor 2-mediated stimulation of NF-kappaB by suppressing the activation of Akt, mTOR, JNK and p38-MAPK. Eur. J. Pharmacol. 815, 190–201. https://doi.org/10.1016/j.ejphar.2017.09.014 (2017).
Zhu, J. et al. Involvement of the delayed rectifier outward potassium channel Kv2.1 in methamphetamine-induced neuronal apoptosis via the p38 mitogen-activated protein kinase signaling pathway. J. Appl. Toxicol. 38, 696–704. https://doi.org/10.1002/jat.3576 (2018).
House, C. D. et al. Voltage-gated Na+ channel activity increases colon cancer transcriptional activity and invasion via persistent MAPK signaling. Sci. Rep. 5, 11541. https://doi.org/10.1038/srep11541 (2015).
Wang, Q. et al. The involvement of the ERK-MAPK pathway in TGF-beta1-mediated connexin43-gap junction formation in chondrocytes. Connect Tissue Res. 60, 477–486. https://doi.org/10.1080/03008207.2019.1593394 (2019).
Ballard, S. L., Miller, D. L. & Ganetzky, B. Retrograde neurotrophin signaling through Tollo regulates synaptic growth in Drosophila. J. Cell Biol. 204, 1157–1172. https://doi.org/10.1083/jcb.201308115 (2014).
Gao, X. et al. Toll-like receptor 3 controls QT interval on the electrocardiogram by targeting the degradation of Kv4.2/4.3 channels in the endoplasmic reticulum. FASEB J. 33, 6197–6208. https://doi.org/10.1096/fj.201801464R (2019).
Min, H. et al. Association of TRPV1 and TLR4 through the TIR domain potentiates TRPV1 activity by blocking activation-induced desensitization. Mol. Pain 14, 1744806918812636. https://doi.org/10.1177/1744806918812636 (2018).
Chen, Y. et al. Inhibition of connexin 43 hemichannels alleviates cerebral ischemia/reperfusion injury via the TLR4 signaling pathway. Front. Cell Neurosci. 12, 372. https://doi.org/10.3389/fncel.2018.00372 (2018).
Steele, A. D. & Yi, C. H. Neuromuscular denervation: Bax up against the wall in amyotrophic lateral sclerosis. J. Neurosci. 26, 12849–12851. https://doi.org/10.1523/jneurosci.4086-06.2006 (2006).
Hur, J., Jeong, H. J., Park, J. & Jeon, S. Chloride channel 4 is required for nerve growth factor-induced TrkA signaling and neurite outgrowth in PC12 cells and cortical neurons. Neuroscience 253, 389–397. https://doi.org/10.1016/j.neuroscience.2013.09.003 (2013).
Colley, B. S., Cavallin, M. A., Biju, K., Marks, D. R. & Fadool, D. A. Brain-derived neurotrophic factor modulation of Kv1.3 channel is disregulated by adaptor proteins Grb10 and nShc. BMC Neurosci. 10, 8. https://doi.org/10.1186/1471-2202-10-8 (2009).
Kanugula, A. K. et al. Novel noncanonical regulation of soluble VEGF/VEGFR2 signaling by mechanosensitive ion channel TRPV4. FASEB J. 33, 195–203. https://doi.org/10.1096/fj.201800509R (2019).
Moher, D., Liberati, A., Tetzlaff, J., Altman, D. G. & Group, P. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. BMJ 339, b2535. https://doi.org/10.1136/bmj.b2535 (2009).
Ramasamy, A., Mondry, A., Holmes, C. C. & Altman, D. G. Key issues in conducting a meta-analysis of gene expression microarray datasets. PLoS Med. 5, e184. https://doi.org/10.1371/journal.pmed.0050184 (2008).
von Mering, C. et al. STRING: A database of predicted functional associations between proteins. Nucleic Acids Res. 31, 258–261. https://doi.org/10.1093/nar/gkg034 (2003).
Kohl, M., Wiese, S. & Warscheid, B. Cytoscape: Software for visualization and analysis of biological networks. Methods Mol. Biol. 696, 291–303. https://doi.org/10.1007/978-1-60761-987-1_18 (2011).
Chin, C. H. et al. cytoHubba: Identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 8(Suppl 4), S11. https://doi.org/10.1186/1752-0509-8-S4-S11 (2014).
Paraskevopoulou, M. D., Vlachos, I. S. & Hatzigeorgiou, A. G. DIANA-TarBase and DIANA suite tools: Studying experimentally supported microRNA targets. Curr. Protoc. Bioinform. 55, 11–18. https://doi.org/10.1002/cpbi.12 (2016).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant 82072162), Beijing Municipal Natural Science Foundation (Grant 7192215), and National Natural Science Foundation of China (Grant 81971177).
Author information
Authors and Affiliations
Contributions
X.Y. designed research, X.G. and B.J. analyzed data and performed research, X.G. wrote the paper, B.J. and Z.Q. recorded the data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gu, X., Jin, B., Qi, Z. et al. Identification of potential microRNAs and KEGG pathways in denervation muscle atrophy based on meta-analysis. Sci Rep 11, 13560 (2021). https://doi.org/10.1038/s41598-021-92489-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-92489-1
- Springer Nature Limited
This article is cited by
-
Neutrophil–macrophage communication via extracellular vesicle transfer promotes itaconate accumulation and ameliorates cytokine storm syndrome
Cellular & Molecular Immunology (2024)