Abstract
Crimean-Congo hemorrhagic fever (CCHF) virus, a highly pathogenic viral agent is responsible for severe fatal hemorrhagic infections in many parts of the world. The early diagnosis of CCHF infection is important for successful clinical management and epidemiological control. The nucleoprotein (NP) of CCHFV being highly conserved and immunogenic is used as early diagnostic marker. In this study, we report a rapid and sensitive double antibody based antigen capture ELISA to detect Crimean-Congo hemorrhagic fever virus (CCHFV). Highly specific polyclonal and monoclonal antibody against NP has been generated and used as capture and detector antibody respectively. The assay was able to detect viral nucleoprotein in different matrices including human serum, ticks and culture supernatant. The detection limit of the developed sandwich ELISA assay was 25 ng of purified antigen. Comparison with a real time RT-PCR revealed its detection limit to be 1000 genome equivalents of CCHFV. Further the assay was comparatively evaluated with a commercial kit employing gamma irradiated CCHFV, revealing a sensitivity and specificity of 100%. This newly developed sandwich ELISA (sELISA) with high sensitivity and specificity could be used as an efficient method for the detection of CCHF virus in humans, ticks and culture supernatant. The assay will be useful as alternate tool for diagnosis of acute infection and is amenable for screening of large scale samples in resource limited settings.
Similar content being viewed by others
Introduction
Crimean-Congo hemorrhagic fever (CCHF), a widespread tick borne disease is endemic in more than 30 countries in the world. The etiological agent CCHF virus belongs to the family Nairoviridae, genus Orthonairovirus and order Bunyavirales1. Since last two decades, CCHFV has spread rapidly throughout Africa, Europe and Asia and causing fatal hemorrhagic fever in naïve population. Every year, nearly 1000 cases are reported from Middle East and eastern European countries. World health organisation (WHO) has included CCHF virus in WHO R&D Blueprint as priority agent in order to opt immediate actions to prevent future epidemics. CCHFV is maintained in enzootic tick-vertebrate-tick cycle in nature, where ticks act as principal vector and reservoir2. Virus infected ticks, rearing in host (small and large animals) can transmit the virus to animals and humans via bite3,4.
CCHFV causes a fatal human infection with high mortality rate. Onset of disease coincides with sudden fever, myalgia, dizziness and bodyache leading to nausea, vomiting, diarrhoea, neural impairments, kidney and liver failure after 5th day of illness that is associated with higher mortality5,6. The disease also represents hemorrhagic symptoms characterized by petechial rash on mucosal surfaces, haematuria, epistaxis and gum bleeding7,8.
In spite of wide distribution and high mortality, vaccines for human use are not widely available. The antiviral ribavirin is non-specific and also suffers from limited efficacy. Supportive therapies including passive plasma transfusion and non-specific antivirals were attempted during initial stage of illness to reduce the mortality9,10. Therefore, a specific and sensitive diagnosis of CCHFV infection becomes crucial to initiate timely tracking and treatment of patients.
Currently, direct detection of CCHFV relies either on virus isolation in suitable animal model or molecular detection of virus. Both these methods are technically demanding and limited to advanced laboratory. The isolation of CCHFV involves high biohazard risk and can only be attempted in biosafety level-4 (BSL-4) laboratories11. Though isolation is a gold standard, however, it plays a limited role in diagnosis12. Although molecular assays such as real-time reverse transcriptase polymerase chain reaction (RT-PCR) can be useful for early and confirmatory diagnosis, yet most of these formats are expensive and requires high-end equipments13. Therefore, these formats may be good at laboratory level but difficult to implement in resource limited settings and field hospital at this point of time. Serological diagnostic kits for detection of IgM and IgG antibody are commercially available. However, by the time detectable antibody titer appears, the patient starts recovering and thus become less significant in clinical outcome of patient. Due to high mortality rate (40%), it is important to detect in early stage of infection i.e., during day 3–6 so that the supportive treatment of patient can be initiated. The antigen detection provides an easy alternative for virus detection at early stage of infection. In spite of CCHFV being a widely circulating virus, there is a lack of availability of antigen detection system around the world. Therefore, there is an urgent need to develop a safe ELISA assay for detection of CCHFV antigen in multiple matrices including serum, ticks and culture fluids. The present study is focused on development of safe, scalable and rapid sandwich ELISA for detection of CCHF virus in multiple matrices.
Results
Antibodies generation and titration
The polyclonal antibody (pAb) and monoclonal antibody (mAb) against recombinant nucleoprotein were raised in rabbit and mouse respectively following the standard protocol. The titer of the generated rabbit anti-rNP polyclonal antibody was found to be 1:64,000. The concentration of purified pAb was 2.0 mg/ml. Out of ten hybridoma clones, two highly reacting hybridoma clones were further shortlisted (Supplementary Figure 1).The monoclonal antibody 2 (CCHF_mAb2) showed higher titer of 1:3200 than CCHF_mAb1 of 1:1600 (Fig. 1). The concentration of purified mAb1 and mAb2 were 1.9 mg/ml and 2.2 mg/ml respectively.
Twofold titration of monoclonal antibody (CCHF_mAb1 and CCHF_mAb2) against recombinant NP in triplicates. Monoclonal antibody 1 (CCHF_mAb1) showed reactivity titre up to 1:1600 dilution against r-NP in indirect ELISA. Monoclonal antibody 2 (CCHF_mAb2) showed the reactivity up to 1:3200 dilution against r-NP in indirect ELISA with the cut-off value of 0.5 determined by twice the OD of negative control.
Characterization of generatdxed antibodies and conjugation of mAb with HRP
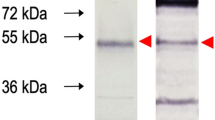
SDS-PAGE analysis of pAb and mAb showed the characteristic light (25 kDa) and heavy chains (60 kDa) (Fig. 2a).Western blot analysis revealed the immunoreactivity of pAb and mAb towards r-NP (56 kDa) (Fig. 2b). Following characterization, mAb_2 was successfully conjugated with HRP enzyme and the titer of the HRP conjugated mAb2 was found to be 1:1600 (Supplementary Figure 2).
(a) 10% SDS-PAGE analysis of antibodiesraised against recombinant nucleoprotein. Lane 1 was loaded with polyclonal antibody and Lane 2 was loaded with monoclonal antibody (CCHF_mAb2) revealing characteristic heavy chain (60 kDa) and light chain (25 kDa) of antibodies. (b) Western blot analysis using polyclonal (Lane 1) and monoclonal (CCHF_mAb2) (Lane 2) antibodies showing immune reactivity with 56 kDa recombinant nucleoprotein (NP) of CCHF virus.
The isotyping assay revealed that both the monoclonal antibodies (CCHF_mAb1 and CCHF_mAb2) belonged to IgG2b isotype (Fig. 3). Dominance of IgG2b represents a Th1 response towards CCHF infection.
Standardization of sandwich ELISA (sELISA)
The optimum working concentration of pAb and HRP_CCHF_mAb2 was determined with checkerboard titration. The optimum titer of pAb IgG and mAb IgG in assay was found to be 1:1000 and 1:100 respectively. The assay was optimized with one hundred and ten (n = 110) healthy human sera and twenty five (n = 25) healthy tick lysate to obtain minimal background with calculated threshold of twice the average of negative controls. The values of negative controls ranged from 0.2 to 0.37.
Evaluation of sandwich ELISA
Using the standardized protocol, the sELISA assay was evaluated with commercial NP antigen. The limit of detection was found to be 25 ng of viral antigen. The detection limit of the assay with in-house r-NP was also tested in parallel and similar detection limit of 25 ng was obtained (Table 1, Fig. 4). Further, on evaluation with CCHFV spiked in different matrices (culture supernatant, serum and tick lysate), sELISA was able to detect up to 10–3 dilution in culture supernatant and 10–2 dilution in both human sera and tick lysate due to matrix interference. The similar dilutions when tested in parallel with real time RT-PCR could detect up to 10–4 dilution. The comparison revealed a limit of detection of 1000 genome equivalents (GE) in terms of RNA copy number (Table 2).
Specificity of the CCHF sandwich ELISA assay
All the samples of DENV 1–4, KFDV, Yellow fever virus and apparently healthy individuals were found non-reactive with the developed sELISA assay (Supplementary Figure 3).
Comparative evaluation with commercial ELISA kit
The prototype was also subjected to comparative evaluation with commercial Vecto-Crimea CHF ELISA kit (Vector Best, Russia) using γ irradiated CCHFV spiked samples. The protocol was followed as per the user manual. Data was comparatively evaluated and found that the commercial kit was able to detect up to 10–2 dilution of virus; while in-house prototype sELISA was able to detect the virus up to 10–3 in culture supernatant and 10–2 in both healthy human sera and tick lysate (Table 3). Sensitivity and specificity of the assay was found to be 100% with commercial antigen ELISA. The percent agreement and correlation coefficient between the two assays were 100%. However, the Vector-best antigen ELISA detected the CCHF culture fluid upto10–2 whereas in-house sandwich ELISA detected the dilution up to 10–3.
Discussion
CCHFV infection causes a fatal hemorrhagic fever in humans with high mortality rate14. Early detection of CCHFV becomes more crucial due to limited availability of vaccines and effective antivirals. Early diagnosis of CCHFV via RT-PCR is an expensive protocol and is not amenable in remote settings. The current disease outbreaks like a box of Pandora demands simple detection systems to track the disease and control the outbreak situations. So far, commercial serological ELISA kit (Vecto-Crimea, Russia) is available for antigen detection but is not intended for human use (mentioned in kit’s manual). Other serological in-house assays are meant to detect antibodies in different ELISA formats either in indirect ELISA or in sandwich ELISA15,16,17. It has also been reported that CCHF in cell culture does not form a clear plaque, leading to difficulties in accurate virus quantification, crucial for the development of immunodiagnostics and vaccines18,19.
Keeping these limitations in mind, the current study was focussed to develop a double antibody based sandwich ELISA for early detection and quantification of CCHFV in different matrices including serum sample, tick and culture supernatant. The CCHFV nucleoprotein being the most conserved structural protein among all the circulating genotypes was targeted to develop the assay. Therefore, pAb and mAb were generated against native NP to serve as capture and detector molecules respectively in sELISA format.
Following optimization, the assay was evaluated with commercial antigen produced using heterologous mammalian expression host. This NP antigen was expressed using a gene derived from CCHF strain IbAr10200, belonging to genotype III20,21. Though the CCHF_mAb2 was generated using in-house r-NP of a CCHFV belonging to genotype IV as an antigen22, its cross reaction to a heterologous genotype confirms its applicability for multiple CCHFV genotypes. The sELISA assay was able to detect both the recombinant antigens with a limit of detection of 25 ng. Sandwich ELISA on evaluation with virus, revealed the detection limit up to 1000 RNA copies which is sufficient to detect the virus in early stage. It has been reported that CCHFV RNA titers ranges from 3.2 × 105 and 4.3 × 106 in mild and severe human cases respectively23. Another study reported a viral load ≥ 1 × 109 RNA copies/mL as marker for prediction of fatal outcome in CCHF infection24. Further, the viral load of CCHFV in tick to cause viremia in vertebrate is approximately 102.2 PFU, equivalent to 1.1 × 105 RNA copies25,26. This indicates that the assay can easily detect the virus in human and ticks in early stages of infection.
The assay also did not reveal cross reactivity with Dengue, KFD and Yellow fever virus. The comparative evaluation with commercial kit revealed a sensitivity and specificity of 100%. Furthermore, the sELISA picked an additional dilution up to 10–3 dilution of CCHFV in culture supernatant, indicating higher sensitivity compared to commercial kit. The sELISA detection assay reported in this study is a highly sensitive, specific and cost-effective tool for early diagnosis and surveillance of CCHFV in humans and ticks in endemic areas, particularly in resource limited settings.
Materials and methods
Chemicals and reagents
Recombinant nucleoprotein, Fruend’s complete and incomplete adjuvants (Sigma, USA), RPMI 1640 media and Bovine serum albumin (Thermo Fisher Scientific, USA), IgG purification kit, anti-mice HRP, anti-rabbit HRP conjugated secondary antibody (Sigma, USA), HRP-conjugation kit (Abcam, USA), Maxisorp plate (Nunc, Denmark), Mouse IgG isotyping kit and 3,3,5,5-tetramethylbenzidine (TMB) (Invitrogen, USA).
Preparation of antigen
Recombinant nucleoprotein (NP) was used as antigen to generate specific antibodies for capturing viral antigen. A recombinant E. coli clone expressing CCHF NP of strain NIV112143 (GenBank Accession no. JN572089) was revived and r-NP was purified as per Shrivastava et al.22. Briefly, cultured and harvested cells were suspended in lysis buffer (50 mM NaH2PO4, 300 mM NaCl, and 10 mM imidazole, pH 8.0) supplemented with phenylmethylsulfonylfluoride (PMSF), lysozyme, and protease inhibiting cocktail (Sigma-Aldrich, USA). The culture was incubated at 4 °C for 30 min and sonicated at 9 pulse on/off for 30 min. The sonicated culture was centrifuged at 10,000×g for 20 min and the supernatant was loaded on the pre-equilibrated polypropylene column containing Ni–NTA slurry for 2 h at 4 °C for efficient binding. The column was washed with 10 bed volumes of wash buffer (50 mM NaH2PO4, 300 mM NaCl, and 20 mM imidazole, pH 8.0) and eluted with elution buffer (50 mM NaH2PO4, 300 mM NaCl, and 250 mM imidazole, pH 8.0). The protein was concentrated and dialyzed with dialysis buffer (50 mM NaH2PO4 and 300 mM NaCl) and further utilized for immunization purpose.
Generation and purification of polyclonal antibody (pAb)
All animal experiments were approved by the Institutional Animal Ethics Committee (IAEC) constituted by the Committee for the Purpose of Control and Supervision of Experiments on Animals (CPCSEA), Ministry of Environment and Forestry, Government of India (Regd. no. 37/GO/Rbi/S/99/CPCSEA) vide protocol no. Viro-14/57/PKD. The affinity purified r-NP was used to raise polyclonal antibody in female New Zealand white rabbit. Purified NP (100 µg) along with Fruend’s complete adjuvant in 1:1 ratio (1 ml) was subcutaneously immunized in the 4 week old female rabbit, procured from DRDE Animal facility division. The booster doses were subsequently given twice within an interval of 15 days with Fruend’s incomplete adjuvant. On the day 7th of post second booster, 20 ml blood was collected from marginal ear vein of the rabbit and titrated using indirect ELISA. Immunoglobulin G was purified from antiserum with protein A sepharose column using Pure-1A antibody purification kit as per manufacturer’s instruction (Sigma, USA). Briefly, the serum was diluted with 3 volume of loading buffer and the culture supernatant was diluted with 1 volume of loading buffer. The diluted material was loaded to the column. The column was washed with 5 ml loading buffer per ml of resin. Following washing twice, the column was applied with elution buffer and the fractions were collected up to 10 ml. Assay elute fractions were analyzed for protein by absorbance at 280 nm. The positive protein fractions were pooled and neutralized with 1.5 M Tris base at pH 6.8. The protein was dialyzed in PBS (pH 7) at 4 °C for 4 h. The protein was further quantified with bicinchoninic acid (BCA) reagent and estimated at an absorbance of 562 nm. Purified IgG was titrated using indirect IgG ELISA22 and utilized for the development of sandwich ELISA for virus detection.
Generation and purification of monoclonal antibody (mAb)
Monoclonal antibodies were raised against purified recombinant NP of CCHFV in in-house bred 4 weeks old female Balb/C mice at M/s Imgenex, India. On the day 7 post immunization the mice were bled from retro-orbital plexus using heparinised capillary, and the serum was separated by centrifugation and further titrated with indirect ELISA. The mouse showing high antibody titre was sacrificed by cervical dislocation and the splenocytes were fused with myeloma cells as per Kohler-Milstein’s protocol. Briefly, 7 days post second booster dose, a single cell suspension of spleenocytes was prepared aseptically in RPMI-1640 media as primary culture. A fusion cell suspension was prepared with myeloma cells Sp2/0 as per the standard protocol using HAT media. Following culture for 7–12 days, hybridoma cells were expanded further for limiting dilution to obtain monoclonal antibody secreting hybridoma clones. Ten hybridoma clones were screened and titrated using indirect IgG ELISA against NP as an antigen. The high titered monoclonal antibodies (n = 2) were purified as described above and stored at − 20 °C.
Characterization of antibody
The purified polyclonal antibody and monoclonal antibody were electrophoresed on 10% reducing SDS-PAGE. For immunoblotting, recombinant NP was electrophoresed and subsequently transferred to nitrocellulose membrane. The membrane was blocked with 5% skimmed milk powder (SMP) in 0.1% PBST and incubated at 4 °C for overnight. The blot was washed and treated with 1:2000 and 1:500 dilution of purified polyclonal and monoclonal antibody respectively and incubated at 4 °C for 4 h. Following incubation, the membrane was washed with 0.1% PBST and treated with 1:5000 dilutions of anti-rabbit and anti-mouse HRP conjugated secondary antibody at 37 °C for 2 h. The blot was further developed with DAB/H2O2as substrate.
Titration of monoclonal antibody
Shortlisted monoclonal antibodies were titrated with indirect ELISA assay. Briefly, 300 ng of r-NP was coated in each well of microtiter plate (Nunc, Denmark) at 4 °C for overnight. The wells were subsequently blocked with 5% SMP for 4 h, followed by washing with 0.05% PBST to remove blocking buffer. Hybridoma cell supernatants in the twofold dilution (1:100 to 1:6400) were applied to wells at 37 °C for 60 min. Following incubation, wells were washed with wash buffer (0.1% PBST) and 1:5000 dilution of anti-mouse HRP secondary antibody (Sigma, USA) was added in each well and incubated at 37 °C for 60 min. Following incubation and washing, the reaction was developed with TMB/H2O2 solution (Invitrogen, USA). The reaction was stopped with 1 N HCl and absorbance was recorded at 450 nm.
Isotyping of monoclonal antibody
The sub-class of mAb (CCHF_mAb1 and CCHF_mAb2) were determined using mouse IgG isotyping kit (Cat. No. 88-50630-88, Invitrogen, USA). ELISA plate was coated with 100 µL/well of capture antibody in coating buffer. Each well was coated with isotype specific antibody and blocked with 250 µL of blocking buffer and incubated at room temperature for 2 h. Following incubation, 50 µL/well of assay buffer A (1×) was added to all wells and incubated at room temperature for 2 h. Wells were washed and incubated with detector antibody and incubated. Plate was washed and developed with TMB/H2O2 incubated at room temperature for 10 min and stopped with 1 N HCl. The absorbance was recorded at 450 nm wavelength.
HRP conjugation of monoclonal antibody
The purified CCHF_mAb2 was conjugated with horseradish peroxidase enzyme (HRP) using HRP conjugation kit (Abcam, USA) as per manufacturer’s instruction to accelerate the speed compared to conventional sandwich ELISA method.
Standardization of sandwich ELISA (sELISA)
The purified pAb and HRP_mAb2 was used as capture and detector antibodies respectively to determine the optimal concentration by checkerboard titration. In brief, 1:250, 1:500, 1:1000 and 1:1500, 1:2000 and 1:4000 dilutions of 1 mg/ml stock of polyclonal antibody were coated (100 μl/well) and incubated at 4 °C for overnight. Further the plates were blocked with 5% skimmed milk powder (SMP) in PBS and a fixed antigen concentration (100 μg/well) in healthy human serum and healthy tick lysates was added in each well for 1 h. Following incubation, the twofold serially diluted (initial concentration 1:50, 1:100, 1:500, 1:1000) HRP_mAb2 (2 mg/ml stock) was added in respective wells and incubated at 37 °C for 1 h. The plate was incubated at 37 °C for 1 h and washed, the reaction was then developed with TMB and stopped with 1 N HCl. The absorbance was recorded with ELISA plate reader (BioTek, USA).
Evaluation of sELISA with commercial antigen
The standardized sandwich ELISA was evaluated for its efficacy in detecting a mammalian cell (HEK) expressed nucleoprotein (The Native antigen, UK). The commercial antigen (NP) was spiked in 0.05% PBST, serially tenfold diluted and tested as per standardized protocol and the limit of detection was calculated. This was also compared with in house E. coli expressed r-NP following the same protocol.
Evaluation of sELISA with CCHF virus
Sandwich ELISA was evaluated with tenfold serial dilutions of gamma irradiated CCHF virus (10–1 to 10–5) in different matrices viz., culture supernatant, serum and tick lysate with standardized protocol. CCHF virus strain NIV1040505 (GenBank number MH396640-MH396642) was used in this study. For CCHFV inactivation, Co-60 source (Gamma Chamber; GC 5000) was used. For complete inactivation of the CCHFV, gamma irradiation dose of 24 kGy has been used for 2 h. Before further use, inactivation of the virus stock was confirmed by two blind passages in Vero cells. Serum and tick specimens referred to NIV during various CCHF outbreaks in India were used for the evaluation of the assay.
Quantitative real-time PCR (qRT-PCR) of CCHFV
The same dilutions were tested with real time RT-PCR and the data was comparatively analyzed27. Taqman real time RT-PCR was performed using TaqMan fast virus one-step master mix in 20 µl reaction mixture (Thermo Fisher Scientific, USA) using ABI7500 Dx real time PCR system (Applied Biosystems, USA). The primer/probe set used for the real time RT-PCR targeting S-gene of CCHFV were Forward Primer: 5′-CAAAGAAACACGTGCCGCTT-3′; Reverse Primer: 5′-ATTCACCTCGATTTTGTTTTCCAT-3′ and Probe: 5′-ACGCCCACAGTGTTCTCTTGAGTGTTAGCA-3′. PCR cycling conditions included 50 °C for 30 min and at 95 °C for 2 min followed by 45 two-step cycles at 95 °C for 15 s and at 55 °C for 60 s. Following cycling, the result was analyzed from the amplification plot.
Cross-reactivity analysis of sandwich ELISA
Cross reactivity of the sELISA assay was analysed with culture supernatants of Dengue virus serotypes [DENV-1, RR107 (KF289072); DENV-2, GWL18 (AY324614); DENV-3, ND143 (FJ644564); DENV-4, ND 73 (HM237348)], Kyasanur forest disease virus strain 12839 (MG720080) and Yellow fever virus 17D strain (KF769015).
Statistical analysis
All the generated data was analyzed with Medcalc 2.0 software.
Ethics statement
All methods are reported in accordance with ARRIVE guidelines. All methods were performed in accordance with the relevant guidelines and regulations. For animal experiments, the study protocol for all experiments was approved as protocol no. Viro-14/57/PKD by the Institutional Animal Ethics Committee (IAEC) constituted by the Committee for the Purpose of Control and Supervision of Experiments on Animals (CPCSEA), Ministry of Environment and Forestry, Government of India (Regd. no. 37/GO/Rbi/S/99/CPCSEA).
References
King, A. M. Q. et al. Changes to taxonomy and the International Code of Virus Classification and Nomenclature ratified by the International Committee on Taxonomy of Viruses (2018). Arch. Virol. 163, 2601–2631 (2018).
Spengler, J. R. et al. A chronological review of experimental infection studies of the role of wild animals and livestock in the maintenance and transmission of Crimean-Congo hemorrhagic fever virus. Antiviral Res. 135, 31–47 (2016).
Papa, A., Tsergouli, K., Tsioka, K. & Mirazimi, A. Crimean-Congo hemorrhagic fever: Tick-host-virus interactions. Front. Cell. Infect. Microbiol. 7, 213 (2017).
Spengler, J. R., Bergeron, É. & Spiropoulou, C. F. Crimean-Congo hemorrhagic fever and expansion from endemic regions. Curr. Opin. Virol. 34, 70–78 (2019).
Kilinc, C. et al. Examination of the specific clinical symptoms and laboratory findings of Crimean-Congo hemorrhagic fever. J. Vector Borne Dis. 53, 162–167 (2016).
Khazaei, Z., Darvishi, I., Amiri, M., Sohrabivafa, M. & Kamran, S. Crimean Congo hemorrhagic fever: A brief report regarding kidney involvement. J. Renal Inj. Prev. 7, 129–131 (2018).
Hoogstraal, H. The epidemiology of tick-borne Crimean-Congo hemorrhagicfever in Asia, Europe, and Africa. J. Med. Entomol. 15, 307–417 (1979).
Swanepoel, R. et al. Epidemiologic and clinical features of Crimean-Congo hemorrhagic fever in southern Africa. Am. J. Trop. Med. Hyg. 36, 120–132 (1987).
Ergonul, O. Treatment of Crimean-Congo hemorrhagic fever. Antiviral Res. 78, 125–131 (2008).
Ergonul, O. Crimean-Congo hemorrhagic fever virus: New outbreaks, new discoveries. Curr. Opin. Virol. 2, 215–220 (2012).
Shayan, S., Bokaean, M., Shahrivar, M. R. & Chinikar, S. Crimean-Congo hemorrhagic fever. Lab. Med. 46, 180–189 (2015).
Mostafavi, E., Pourhossein, B. & Chinikar, S. Clinical symptoms and laboratory findings supporting early diagnosis of Crimean-Congo hemorrhagic fever in Iran: Early Diagnosis of Crimean-Congo hemorrhagic fever. J. Med. Virol. 86, 1188–1192 (2014).
Mertens, M., Schmidt, K., Ozkul, A. & Groschup, M. H. The impact of Crimean-Congo hemorrhagic fever virus on public health. Antiviral Res. 98, 248–260 (2013).
Palomar, A. M. et al. Crimean-Congo hemorrhagicfever virus in ticks from migratory birds, Morocco. Emerg. Infect. Dis. 19, 260–263 (2013).
Samudzi, R. R., Leman, P. A., Paweska, J. T., Swanepoel, R. & Burt, F. J. Bacterial expression of Crimean-Congo hemorrhagic fever virus nucleoprotein and its evaluation as a diagnostic reagent in an indirect ELISA. J. Virol. Methods. 179, 70–76 (2012).
Sas, M. A. et al. A novel double-antigen sandwich ELISA for the species-independent detection of Crimean-Congo hemorrhagic fever virus-specific antibodies. Antiviral Res. 151, 24–26 (2018).
Schuster, I. et al. A competitive ELISA for species-independent detection of Crimean-Congo hemorrhagic fever virus specific antibodies. Antiviral Res. 134, 161–166 (2016).
Mourya, D. T. et al. Retrospective analysis of clinical information in Crimean-Congo haemorrhagic fever patients: 2014–2015, India. Indian J. Med. Res. 145, 673–678 (2017).
Mourya, D. T. et al. Detection, isolation and confirmation of Crimean-Congo hemorrhagic fever virus in human, ticks and animals in Ahmadabad, India, 2010–2011. PLoS Negl. Trop. Dis. 6, e1653 (2012).
Golden, J. W. et al. GP38-targeting monoclonal antibodies protect adult mice against lethal Crimean-Congo hemorrhagic fever virus infection. Sci. Adv. 5, eaaw9535 (2019).
Deyde, V. M., Khristova, M. L., Rollin, P. E., Ksiazek, T. G. & Nichol, S. T. Crimean-Congo hemorrhagicfever virus genomics and global diversity. J. Virol. 80, 8834–8842 (2006).
Shrivastava, N. et al. Development of multispecies recombinant nucleoprotein-based indirect ELISA for high-throughput screening of Crimean-Congo hemorrhagicfever virus-specific antibodies. Front. Microbiol. 10, 1822 (2019).
Duh, D. et al. Viral load as a predictor of Crimean-Congo hemorrhagic fever outcome. Emerg. Infect. Dis. 13(11), 1769–1772 https://doi.org/10.3201/eid1311.070222 (2007).
Cevik, M. A. et al. Viral load as a predictor of outcome in Crimean-Congo hemorrhagic fever. Clin. Infect. Dis. 45, e96–e100 (2007).
Durden, L. A., Logan, T. M., Wilson, M. L. & Linthicum, K. J. Experimental vector incompetence of a soft tick, Ornithodorossonrai (Acari: Argasidae), for Crimean-Congo hemorrhagic fever virus. J. Med. Entomol. 30, 493–496 (1993).
Bell-Sakyi, L., Kohl, A., Bente, D. A. & Fazakerley, J. K. Tick cell lines for study of Crimean-Congo hemorrhagic fever virus and other arboviruses. Vector-Borne Zoonotic Dis. 12, 769–781 (2012).
Liu, J. et al. Development of a TaqMan array card for acute- febrile-illness outbreak investigation and surveillance of emerging pathogens, including Ebola virus. J. Clin. Microbiol. 54, 49–58 (2016).
Acknowledgements
Authors are thankful to Dr D. K. Dubey, Director, DRDE, Gwalior, Dr.Priya Abraham, Director, NIV, Pune and Dr. D. T. Mourya, Former Director, NIV, Pune for their keen interest, constant support, and provision of necessary facilities for this study. This manuscript is assigned DRDE accession no. DRDE/VIRO/24/2020. The first author would also like to thank Department of Science and technology (DST), Govt. of India for providing INSPIRE fellowship.
Author information
Authors and Affiliations
Contributions
P.K.D. and N.S. conceived, designed the experiments. N.S. conducted the experiments, analysed data and wrote the MS. P.K.D. supervised the study, critically revised the MS for intellectual content. J.S.K. and A.S. performed the antibody generation experiments and reviewed the manuscript. P.D.Y., A.M.S. and R.J. performed the validation and comparative evaluation of the assay. All authors have approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shrivastava, N., Kumar, J.S., Yadav, P. et al. Development of double antibody sandwich ELISA as potential diagnostic tool for rapid detection of Crimean-Congo hemorrhagic fever virus. Sci Rep 11, 14699 (2021). https://doi.org/10.1038/s41598-021-93319-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-93319-0
- Springer Nature Limited
This article is cited by
-
Recent Advances in Crimean-Congo Hemorrhagic Fever Virus Detection, Treatment, and Vaccination: Overview of Current Status and Challenges
Biological Procedures Online (2024)
-
Identification and evaluation of multi-antigenic epitopes of immunodominant protein from the selected Crimean–Congo hemorrhagic fever virus genome towards the development of diagnostic and vaccine candidates by reverse vaccinology approach
Journal of Proteins and Proteomics (2024)
-
Crimean–Congo haemorrhagic fever virus
Nature Reviews Microbiology (2023)
-
Development of a novel double-antibody sandwich quantitative ELISA for detecting SADS-CoV infection
Applied Microbiology and Biotechnology (2023)