Abstract
Tuberculosis (TB), caused by Mycobacterium tuberculosis, is endemic in Pakistan. Resistance to both firstline rifampicin and isoniazid drugs (multidrug-resistant TB; MDR-TB) is hampering disease control. Rifampicin resistance is attributed to rpoB gene mutations, but rpoA and rpoC loci may also be involved. To characterise underlying rifampicin resistance mutations in the TB endemic province of Khyber Pakhtunkhwa, we sequenced 51 M. tuberculosis isolates collected between 2016 and 2019; predominantly, MDR-TB (n = 44; 86.3%) and lineage 3 (n = 30, 58.8%) strains. We found that known mutations in rpoB (e.g. S405L), katG (e.g. S315T), or inhA promoter loci explain the MDR-TB. There were 24 unique mutations in rpoA, rpoB, and rpoC genes, including four previously unreported. Five instances of within-host resistance diversity were observed, where two were a mixture of MDR-TB strains containing mutations in rpoB, katG, and the inhA promoter region, as well as compensatory mutations in rpoC. Heteroresistance was observed in two isolates with a single lineage. Such complexity may reflect the high transmission nature of the Khyber Pakhtunkhwa setting. Our study reinforces the need to apply sequencing approaches to capture the full-extent of MDR-TB genetic diversity, to understand transmission, and to inform TB control activities in the highly endemic setting of Pakistan.
Similar content being viewed by others
Introduction
Tuberculosis (TB), caused by Mycobacterium tuberculosis (MTB), affects ~ 10 million people annually1. Despite a recent reduction in TB incidence, increasing drug resistance is making infection control difficult1. New diagnostic tools and treatments are needed in developing countries to inform clinical and disease control activities2. The recommended combination of anti-TB drugs involves a 6-month standard regimen including first-line rifampicin, isoniazid, ethambutol, and pyrazinamide1. The detection of M. tuberculosis resistant to first-line rifampicin (RR-TB) and isoniazid, together called multidrug resistance (MDR-TB), is especially crucial in Pakistan3. The country is in the top eight for RR-TB and MDR-TB burden, with an estimated 6100 of its > 330,000 TB cases in 2019 presenting with RR-TB or greater resistance4.
Khyber Pakhtunkhwa is the third largest province of Pakistan, with an area of 74,521 km2 and a population size of approximately 30.5 million individuals. It has a history of armed conflicts, and TB control has been a neglected area of public health. Recently, a TB control program was launched at Hayatabad Medical Complex Peshawar to monitor disease incidence and perform drug susceptibility testing (DST) to assess levels of RR-TB and MDR-TB. RR-TB is mainly caused by the presence of mutations in the RNA polymerase β subunit (rpoB) gene. Mutations in an 81-bp core region of the rpoB gene, known as the RR-determining region (RRDR), account for 95% of the RR-TB detected in clinical isolates5. However, secondary mutations in the rpoA or rpoC genes can mitigate the initial fitness cost caused by a rpoB mutation6,7. About one-third of RR-TB involves rpoB mutant strains with companion rpoC or rpoA mutations8. Here, by sequencing fifty-one M. tuberculosis, we aimed to summarise the diversity and potential role of rpoB, rpoA, and rpoC mutations in the high MDR incidence setting of Khyber Pakhtunkhwa, to inform diagnostics tools for TB control and implementation. The analysis revealed some mixed MDR infections and heteroresistance, reflecting the complex dynamics of the high transmission setting in Pakistan, and reinforcing the need for robust diagnostic tools.
Materials and methods
A total of 4822 pulmonary and extrapulmonary samples positive for M. tuberculosis were processed between 2016 to 2019 by the provincial Reference Laboratory for TB at Hayatabad Medical Complex Peshawar. Samples were received from all the Programmatic drug management units in the province. All samples were processed using the N-acetyl-l-cysteine–sodium hydroxide (NALC–NaOH) concentration method, using Lowenstein–Jensen medium (LJ) solid culture and liquid MGIT tubes containing 7H9 media9. All samples were subject to phenotypic drug susceptibility testing (DST), performed through the automated BACTEC MGIT 960 system (BD Diagnostic Systems, NJ, USA)10. The DST for resistant isolates was repeated for confirmation of drug resistance profile. Resistance to isoniazid, rifampicin, and other drugs was assessed through the BACTEC MGIT 960 system, with established WHO critical concentrations11. As per WHO guidelines for MGIT DST, there is only a single concentration (1.0 umg/mL) for rifampicin. Fifty-one samples (4 pan-susceptible, 47 drug-resistant, including 44 MDR-TB) were selected for whole genome sequencing (WGS). The source of genomic DNA was MGIT Culture (Liquid Culture) and the same culture generation was used for the phenotypic and genotypic (WGS) characterisation. The study was approved by the Institutional Ethics Committee of KUST Kohat and Provincial Tuberculosis Reference Laboratory, Khyber Pakhtunkhwa (Ref. no.: PTP/PTRL-408/19). All study participants gave informed consent for the collection and use of their biological materials, and all methods were performed in accordance with the relevant guidelines and regulations.
Genomic DNA for the fifty-one isolates was extracted using the acetyl trimethylammonium bromide (CTAB) method12. The DNA underwent WGS at the London School of Hygiene and Tropical Medicine (LSHTM) using the Illumina MiSeq platform (QIAseq FX DNA Library Kit) with a 151 bp paired-end protocol (v3). The quality of the resulting raw sequence data (FASTQ files) was assessed using the FastQC tool (v0.11.8). Reads were trimmed using trimmomatic software (v0.39, LEADING:3 TRAILING:3 SLIDINGWINDOW:4:20 MINLEN:36). The filtered reads were aligned to the H37Rv genome (Accession No. AL123456.3) using Burrow-Wheeler Aligner (BWA)-mem (v0.6) software13. The GATK tool (HaplotypeCaller v4.1.4.1, -ERC GVCF)14 was used to call single nucleotide polymorphisms (SNPs) and insertions and deletions (indels), and the variants were annotated for function (e.g. non-synonymous mutations, frameshifts). The M. tuberculosis isolates were profiled for lineage and genotypic drug resistance using TB-Profiler software (v3.0.0), which requires a minimum of 10 reads to call variants by default15. Mixed infections were defined as samples with more than one lineage, predicted by TB-Profiler, and supported by > 200 biallelic genomic positions. Mutations were assigned to the strains by comparing their frequency in read coverage to the predicted frequency of the strains in each mixture based on genome-wide variants. All results were compared to a mutational database consisting of > 35 k M. tuberculosis spanning all lineages and drug resistance types16. The effects of mutations on RpoA, RpoB, and RpoC protein structures were investigated using the DynaMut server17. In particular, the effects on protein stability (changes in vibrational entropy, ΔΔG: kcal/mol) and flexibility (ΔΔSVib: kcal/mol-1.K-1) were assessed using models downloaded from the PDB (ID: 6c04). Negative predicted stability changes are destabilising, whilst positive changes are stabilising. Consensus genomes were constructed for all isolates using bcftools (v1.9)18. A multiple sequence alignment was constructed using the consensus genomes, and phylogenetic reconstruction was performed using iqtree software (v1.6.12, -m GTR + G)19.
Results
High quality data was generated for fifty-one isolates with a median of 5,186,316 read pairs per isolate. Trimming removed low-quality reads leading to the retention of on average 70.9% of pairs with both reads and 97.2% with at least one read. Classification of the reads using Kraken software reported high percentages classified as Mycobacterium (median: 99.7%). Mapping of the reads led to an average depth of 216-fold coverage, and 99% of the genome with at least tenfold coverage. Of the fifty-one M. tuberculosis isolates, the majority were from lineage 3 (30, 58.8%), and were sourced from female patients (n = 29, 56.9%) and previously treated cases (n = 30, 58.8%) (Table 1). We identified four mixed infections, consisting of different sub-lineages (4.8 and 2.2.1, 4.9 and 3, 3 and 2.2.1, 3 and 4.6.2.1) (Table 1; Supplementary Table S1; Fig. 1), potentially reflecting the high transmission setting. By comparing the allelic frequencies at each mutation position with the predicted frequency of the strains within a mixture, we assigned mutations to the individual strains wherever possible. The phenotypic and genotypic drug resistance profiles were concordant across all samples at the dominant strain level, but the genomic data revealed the complexity of the four mixed lineage infections. In particular, one sample contained a combination of MDR-TB and isoniazid-resistant strains, whilst the other three isolate samples were combinations of MDR-TB with MDR-TB (n = 2) or pan susceptible (n = 1) strains (Supplementary Table S1).
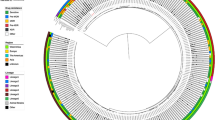
Phylogenetic tree of M. tuberculosis strains (n = 51), with their lineages, multi-drug resistance profiles and specific mutations in rpoB, rpoA, and rpoC. The phylogenetic tree was created using a maximum likelihood approach. The first vertical band to the right of the tree denotes the lineage. The second vertical band denotes the drug resistance phenotype. The squares show the presence of a specific mutation.
Isolates with isoniazid resistance (n = 45) presented with underlying haplotypes katG S315T (31/45), fabG1 − 15C > T + katG S315T (6/45), fabG1 − 15C > T (3/45), fabG1 − 15C > T + ahpC − 54C > T (3/45), fabG1 − 15C > T + katG D311G (1/45), and katG V1A (1/45) (Supplementary Table S1). The RR-TB and MDR-TB isolates (n = 46) all had established rifampicin related mutations: rpoB S450L (41/46), R552 (3/46), H445D/L/R (3/46), S874Y (3/46), and S450W/F (2/46) (Table 2). We found two potential new mutations in rpoB (K891Q, P471R) (Table 2), but they appeared on the same background as established resistance markers (rpoB H445L and S450L) (Table 3; Supplementary Table S1). Of the RR-TB and MDR-TB samples (n = 46), 34 (73.9%) had at least one potential compensatory mutation, the majority in rpoC (32/46) and a minority in rpoA (5/46), including three isolates with mutations in both genes. The most common rpoC mutations were V483G (10), E1092D (9), I491T (7), V483A (4), P1040S (3), and K891Q (2), and when observed previously, have been present in predominantly RR-TB strains (Table 2), thereby reflecting their potential compensatory role. The rpoA mutations included D253Y (2/46), V183G (2/46), and D190E (1/46), all appearing to be previously unreported (Table 2), but only V183G was present on the background without rpoC mutations (Table 3). Interestingly, the only potentially unreported rpoC mutation (V517E) was in a mixed infection (MDR-TB/isoniazid-resistant) with rpoB S450W and rpoA D190E present (Table 3, Supplementary Table S1).
We assessed in silico the impact of eight low frequency mutations (rpoB P471R, R552L, S874Y, K891Q; rpoA V183G, D190E; rpoC G332, V517E) on the stability and flexibility of appropriate protein structures (RpoA, RpoB, RpoC) (Table 4; Figs. 2, 3, 4). These mutations all have low frequency in our study (1–3 isolates) and in the 35 k dataset (< 30 isolates). Note, the impact of the potentially unreported rpoA D253Y mutation was not assessed, as residues of D253 for RpoA could not be modelled because of the eight mutations considered, all but two (rpoB R552L; rpoA D190E) had a potentially stabilizing effect. The rpoC G332R mutation had the highest stability and lowest flexibility effects (Table 4) and is found exclusively in RR-TB strains (Table 2). The rpoA D190E led to the least stable and highest flexibility effects. This mutation was present on the uncommon rpoB S450W and rpoC V517E mutation background, where each mutation is found almost exclusively in RR-TB strains (Table 2).
Effect of the rpoC G332R and V517E point mutations on RpoC protein dynamics. ΔΔG; Free energy difference. ΔΔSVib ENCoM; vibrational entropy energy. The effects have been predicted through the DynaMut online server. The increasing (red) and decreasing (blue) molecular flexibility effects of point mutations have been depicted. The total energy calculated for mutations shows a stabilizing effect on protein structure.
Discussion
Resistance to rifampicin (RR-TB) has been associated with mutations in an 81 bp hot-spot region of the rpoB gene. However, mutations in rpoA and rpoC, have also been linked to RR-TB, particularly in a compensatory role20. We identified 24 mutations across rpoA, rpoB and rpoC genes in 51 M. tuberculosis (46 RR-TB) isolates, including six mutations [rpoA (2), rpoB (3), and rpoC (1)], which were not found in a large collection (~ 35 k) of global clinical isolates. All RR-TB could be explained by known mutations, including the rpoB S450L (41/46 isolates), which is the most prevalent mutation globally (~ 64%)20 and in Pakistan (~ 74%21; ~ 71%22). Four rpoB mutations (R552L, S874Y, K891Q, P471R) appear not to have been reported by recent genetic studies from Khyber Pakhtunkhwa21,22,23. Some recently reported rpoB mutations in Pakistan (L430P, H445P/Y, L452P21; S450Q, P454H, G455D22; D435G21,22, D435Y21,23), which are of much lower frequency than the S450L mutation and sometimes reported in combination with it, were not present in our RR-TB isolates. Similarly, all isoniazid resistance could be explained by known mutations, including the katG S315T (37/45), which has a high frequency globally (~ 75%) and in Pakistan (> 89%21,23). The majority of RR-TB/MDR-TB isolates had mutations in rpoC (32/46), which is a locus known to restore the fitness of rifampicin-resistant bacteria carrying mutations in rpoB24. Other studies have found a lower prevalence of rpoC mutations in MDR isolates (< 30%)6,25,26, including a study in Pakistan26, and inconsistencies may be due to differences in sampling sites and strategies. Eight mutations in rpoC (G332R, N416S, V483A, V517E/L, G594E, P1040S, E1092D) appear not to have been reported in recent genetic studies from Khyber Pakhtunkhwa21,22,23. The evidence for a prominent compensatory role of rpoA in RR-TB is less strong. Our study identified only three rpoA mutations, of which two appear not to have been reported before, and were in the presence of rpoC mutations. In general, potential rpoA and rpoC compensatory mutations and mechanisms need to be validated experimentally in fitness studies, or through large-scale studies involving phylogenetic analysis of longitudinally collected strains.
Phenotypic DST is considered to be the gold standard method to infer resistance. However, some studies have reported that liquid culture systems (e.g. the MGIT platform) often fail to detect low-levels of RR-TB27,28, including those involving H445D/L/R mutations27,29, which have been linked to treatment failure or relapse30,31,32. The role of such mutations can be investigated using protein structural analysis33, and previous studies have shown that mutations may affect the structural dynamics of proteins, making them weak targets, resulting in drug resistance4,34,35. Our in silico analysis revealed that the rpoC G332R mutation, which is rare (< 0.05%) globally and exclusively in RR-TB isolates, led to high protein stability and less flexibility, which may be favorable features for compensatory effects. This mutation has been previously reported in a South African setting5, where it was observed to originate through homoplastic evolution and thus likely to confer a selective advantage. Several of the other low-frequency mutations in the three genes have been reported previously (rpoB R552L36, rpoB S874Y37 and rpoA V183G6) in different settings and have been proposed as potential compensatory mutations. For other low-frequency mutations including rpoB K891Q, rpoB P471R, rpoC V517E, rpoA D190E and rpoA D253Y, we did not find any previous reports in the literature. These mutations fall outside the rifampicin resistance determining regions and co-occur with known resistance mutations, potentially to compensate for the fitness impact of the resistance mutations.
A potential limitation of the phenotypic DST exposed by our genomic analysis is its robustness to mixed infections and heteroresistance. Only the WHO recommended concentration of the rifampicin drug was assessed, and dominant rifampicin resistance mutants within MDR-TB mixes can complicate the phenotypic interpretation of any low-level rifampicin resistance mutants. In our study, five instances of within-host resistance mechanism diversity were observed as the result of mixed infections. Three instances involved mixed-lineage infections, two of which were a mixture of MDR-TB strains and one was a mixture of a pan-susceptible and an MDR-TB strain. Interestingly, the MDR-TB mixtures contained different resistance-conferring mutations in rpoB, katG and the inhA promoter region, as well as likely compensatory mutations in rpoC and potentially rpoA. These mutations were found in the same proportions within the raw data as lineage variants, indicating the different strains carry resistance markers. Heteroresistance was also observed in isolates with a single lineage. This observation could be indicative of acquired resistance, where mutations have not become fixed in the within-host bacterial population. Almost all single-lineage RR-TB isolates contained the rpoB S450L at a completely fixed level in the sequenced population. Some of these strains possessed additional mutations in rpoB, rpoC and rpoA at unfixed proportions (e.g. rpoB R552L, rpoB S874Y, rpoC V483G and rpoA D190E). Based on this observation it is likely that these mutations were acquired after the rpoB S450L, which confers high-level resistance and serve as compensatory mutations to restore the fitness penalty of the resistance mutation.
The application of sequencing technologies in TB endemic regions will assist clinical management and personalised anti-TB drug approaches, as well as disease control through surveillance activities. The management of MDR-TB strains is essential for the control of TB. In our study, we found that the mutations underlying MDR-TB are established variants. Known and putative drug resistance markers in established loci are detectable using low-cost sequencing-based approaches (e.g. amplicon-based), suitable for a low resource setting. These methods are also robust to mixed infections and heteroresistance35,38, which were present in our study and likely driven by the high transmission setting. Large-scale approaches using whole genome sequencing and analysis will assist with understanding the epidemiology and risk groups or factors underpinning the transmission of TB, as well as provide insights into drug resistance and compensatory mechanisms. Such insights will inform the deployment of anti-TB drug regimens and disease control tools and strategies in high burden settings, such as Pakistan.
Conclusion
Whole genome sequencing for TB clinical management and disease control will have the greatest benefit in complex community outbreaks in endemic regions. Our study in the Khyber Pakhtunkhwa province in Pakistan has provided a characterization of circulating rifampicin resistance and MDR-TB mutations. Such insights will assist sequencing-informed diagnosis for proactive TB patient management, and the deployment of anti-TB drug regimens and surveillance activities.
Data availability
The raw sequence data is available from the EBI ENA (Accession number PRJEB43284).
Change history
19 July 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41598-021-94687-3
References
Global tuberculosis report 2020. Geneva: World Health Organization (2020).
Dutta, N. K. et al. Genetic requirements for the survival of tubercle bacilli in primates. J. Infect. Dis. 201, 1743–1752 (2010).
Prasad, R., Gupta, N. & Banka, A. Multidrug-resistant tuberculosis/rifampicin-resistant tuberculosis: Principles of management. Lung India 35, 78–81 (2018).
Ali, S. et al. Prevalence of multi-drug resistant Mycobacterium tuberculosis in Khyber Pakhtunkhwa—A high tuberculosis endemic area of Pakistan. Polish J. Microbiol. 69, 133–137 (2020).
De Vos, M. et al. Putative compensatory mutations in the rpoc gene of rifampin-resistant Mycobacterium tuberculosis are associated with ongoing transmission. Antimicrob. Agents Chemother. 57, 827–832 (2013).
Comas, I. et al. Whole-genome sequencing of rifampicin-resistant Mycobacterium tuberculosis strains identifies compensatory mutations in RNA polymerase genes. Nat. Genet. 44, 106–110 (2012).
Brandis, G., Wrande, M., Liljas, L. & Hughes, D. Fitness-compensatory mutations in rifampicin-resistant RNA polymerase. Mol. Microbiol. 85, 142–151 (2012).
Xu, Z. et al. Transcriptional approach for decoding the mechanism of rpoC compensatory mutations for the fitness cost in rifampicin-resistant Mycobacterium tuberculosis. Front. Microbiol. 9, 2895 (2018).
Kubica, G. P., Dye, W. E., Cohn, M. L. & Middlebrook, G. Sputum digestion and decontamination with N-acetyl-l-cysteine-sodium hydroxide for culture of mycobacteria. Am. Rev. Respir. Dis. 87, 775–779 (1963).
Siddiqi, S. & Gerdes, S. Foundation for Innovative New Diagnostics (FIND) MGITTM Procedure Manual for BACTECTM MGIT 960TM TB System. https://www.finddx.org/wp-content/uploads/2016/02/mgit_manual_nov2006.pdf (2006).
World Health Organization & World Health Organization. (2008). Policy guidance on drug-susceptibility testing (DST) of second-line antituberculosis drugs. World Health Organization. https://apps.who.int/iris/handle/10665/70500 (2008).
Minas, K., Mcewan, N. R., Newbold, C. J. & Scott, K. P. Optimization of a high-throughput CTAB-based protocol for the extraction of qPCR-grade DNA from rumen fluid, plant and bacterial pure cultures. FEMS Microbiol. Lett. 325, 162–169 (2011).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–95 (2010).
McKenna, A. et al. The genome analysis toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Phelan, J. E. et al. Integrating informatics tools and portable sequencing technology for rapid detection of resistance to anti-tuberculous drugs. Genome Med. 11, 41 (2019).
Napier, G. et al. Robust barcoding and identification of Mycobacterium tuberculosis lineages for epidemiological and clinical studies. Genome Med. 12, 114 (2020).
Rodrigues, C. H. M., Pires, D. E. V. & Ascher, D. B. DynaMut: Predicting the impact of mutations on protein conformation, flexibility and stability. Nucleic Acids Res. 46, W350–W355 (2018).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Shea, J. et al. Low-level rifampin resistance and rpoB mutations in Mycobacterium tuberculosis: An analysis of whole-genome sequencing and drug susceptibility test data in New York. J. Clin. Microbiol. https://doi.org/10.1128/jcm.01885-20 (2020).
Jabbar, A. et al. Whole genome sequencing of drug resistant Mycobacterium tuberculosis isolates from a high burden tuberculosis region of North West Pakistan. Sci. Rep. 9, 1–9 (2019).
Ahmad, B. et al. Mutation analysis for detection of drug resistance in Mycobacterium tuberculosis isolates from Khyber Pakhtunkhwa, Pakistan. J. Pak. Med. Assoc. 67, 1684–1688 (2017).
Khan, M. T. et al. Insight into the drug resistance whole genome of Mycobacterium tuberculosis isolates from Khyber Pakhtunkhwa, Pakistan. Infect. Genet. Evol. 92, 104861 (2021).
Andersson, D. I. The biological cost of mutational antibiotic resistance: Any practical conclusions?. Curr. Opin. Microbiol. 9, 461–465 (2006).
Coll, F. et al. Genome-wide analysis of multi- and extensively drug-resistant Mycobacterium tuberculosis. Nat. Genet. 50, 307–316 (2018).
Phelan, J. et al. Mycobacterium tuberculosis whole genome sequencing and protein structure modelling provides insights into anti-tuberculosis drug resistance. BMC Med. 14, 1–13 (2016).
Alexander, D. C. et al. Gene sequencing for routine verification of pyrazinamide resistance in Mycobacterium tuberculosis: A role for pncA but not rpsA. J. Clin. Microbiol. 50, 3726–3728 (2012).
Advani, J. et al. Whole genome sequencing of Mycobacterium tuberculosis clinical isolates from India reveals genetic heterogeneity and region-specific variations that might affect drug susceptibility. Front. Microbiol. 10, 309 (2019).
Alexander, K. A. et al. Novel Mycobacterium tuberculosis complex pathogen, M. mungi. Emerg. Infect. Dis. 16, 1296–1299 (2010).
Van Ingen, J., Aarnoutse, R., De Vries, G., Boeree, M. J. & Van Soolingen, D. Low-level rifampicin-resistant Mycobacterium tuberculosis strains raise a new therapeutic challenge. Int. J. Tuberc. Lung Dis. 15, 990–992 (2011).
Van Deun, A. et al. Disputed rpoB mutations can frequently cause important rifampicin resistance among new tuberculosis patients. Int. J. Tuberc. Lung Dis. 19, 185–190 (2015).
Shah, N. S. et al. Clinical impact on tuberculosis treatment outcomes of discordance between molecular and growth-based assays for Rifampin Resistance, California 2003–2013. Open Forum Infect. Dis. 3, ofw150 https://doi.org/10.1093/ofid/ofw150 (2016).
Khan, M. T., Rehaman, A. U., Junaid, M., Malik, S. I. & Wei, D. Q. Insight into novel clinical mutants of RpsA-S324F, E325K, and G341R of Mycobacterium tuberculosis associated with pyrazinamide resistance. Comput. Struct. Biotechnol. J. 16, 379–387 (2018).
Khan, M. T. et al. Structural and free energy landscape of novel mutations in ribosomal protein S1 (rpsA) associated with pyrazinamide resistance. Sci. Rep. 9, 1–12 (2019).
Khan, M. T. et al. Pyrazinamide resistance and mutations L19R, R140H, and E144K in Pyrazinamidase of Mycobacterium tuberculosis. J. Cell. Biochem. 120, 7154–7166 (2019).
Ko, D. H. et al. Application of next-generation sequencing to detect variants of drug-resistant Mycobacterium tuberculosis: Genotype-phenotype correlation. Ann. Clin. Microbiol. Antimicrob. 18, 1–8 (2019).
Ma, P. et al. Compensatory effects of M. tuberculosis rpoB mutations outside the rifampicin resistance-determining region. Emerg. Microbes Infect. 10, 743–752 (2021).
Shivakumar, K. V., Karunakar, P. & Chatterjee, J. Inhibition of NarL of Mycobacterium tuberculosis: an in silico approach. Interdiscip. Sci. Comput. Life Sci. 6, 292–299 (2014).
Acknowledgements
TGC is funded by Medical Research Council UK (Grant no. MR/M01360X/1, MR/N010469/1, MR/R025576/1, and MR/R020973/1) and BBSRC UK (Grant no. BB/R013063/1) grants. SC is funded by BloomsburySET and Medical Research Council UK grants (MR/M01360X/1, MR/R025576/1, and MR/R020973/1).
Author information
Authors and Affiliations
Contributions
A.S.K., J.E.P., T.G.C. and T.A.K. conceived and directed the project. A.S.K., M.T.K., S.A.L., M.Q., and T.A.K. coordinated and peformed sample collection and sample processing. S.C. and T.G.C. coordinated sequencing. J.E.P., M.T.K., G.N., S.A., O.M.C., S.Z., H.R., D.-Q.W. and T.G.C. contributed informatic tools and data. A.S.K., J.E.P. and M.T.K. performed bioinformatic and statistical analyses under the supervision of S.A., T.G.C. and T.A.K. All authors interpreted the results. A.S.K., J.E.P., T.G.C. and T.A.K. wrote the first draft of the manuscript. All authors commented on and edited the manuscript and approved the final version. A.J., J.E.P., S.J.W., and T.G.C. compiled the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained an error in the spelling of the author Otavio Cabral-Marques which was incorrectly given as Otavio Marques Cabral. The original Article has been corrected.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Khan, A.S., Phelan, J.E., Khan, M.T. et al. Characterization of rifampicin-resistant Mycobacterium tuberculosis in Khyber Pakhtunkhwa, Pakistan. Sci Rep 11, 14194 (2021). https://doi.org/10.1038/s41598-021-93501-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-93501-4
- Springer Nature Limited
This article is cited by
-
Large-scale statistical analysis of Mycobacterium tuberculosis genome sequences identifies compensatory mutations associated with multi-drug resistance
Scientific Reports (2024)
-
Whole genome sequencing of multidrug-resistant Mycobacterium tuberculosis isolates collected in the Czech Republic, 2005–2020
Scientific Reports (2022)