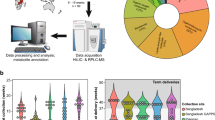
Abstract
The elucidation of dynamic metabolomic changes during gestation is particularly important for the development of methods to evaluate pregnancy status or achieve earlier detection of pregnancy-related complications. Some studies have constructed models to evaluate pregnancy status and predict gestational age using omics data from blood biospecimens; however, less invasive methods are desired. Here we propose a model to predict gestational age, using urinary metabolite information. In our prospective cohort study, we collected 2741 urine samples from 187 healthy pregnant women, 23 patients with hypertensive disorders of pregnancy, and 14 patients with spontaneous preterm birth. Using gas chromatography-tandem mass spectrometry, we identified 184 urinary metabolites that showed dynamic systematic changes in healthy pregnant women according to gestational age. A model to predict gestational age during normal pregnancy progression was constructed; the correlation coefficient between actual and predicted weeks of gestation was 0.86. The predicted gestational ages of cases with hypertensive disorders of pregnancy exhibited significant progression, compared with actual gestational ages. This is the first study to predict gestational age in normal and complicated pregnancies by using urinary metabolite information. Minimally invasive urinary metabolomics might facilitate changes in the prediction of gestational age in various clinical settings.
Similar content being viewed by others
Introduction
Pregnancy induces dynamic and temporal changes in maternal physiological profiles1. Recent reports suggested that various omics profiles (e.g., those of transcriptomics, epigenomics, metabolomics, and microbiomics) show substantial temporal changes during pregnancy2,3,4,5. Furthermore, changes in omics profiles are associated with pregnancy complications6,7.
To evaluate pregnancy status and predict pregnancy complications, several groups have attempted to characterize normal pregnancy via omics analyses of maternal biospecimens8,9,10. Specifically, Liang and colleagues have constructed models to predict gestational age based on plasma metabolites at the time of specimen collection11. The models could predict gestational age with high accuracy (i.e., correlation coefficient of 0.92 between predicted and actual gestational ages); those predictive models precisely reflected physiological adaptations in pregnant women. However, the models thus far have used omics information collected from maternal blood. Furthermore, there have not been investigations regarding whether predictive models show distinct results in cases with pregnancy complications, compared with women who have normal pregnancy. Therefore, less invasive methods are desired for the prediction of gestational age in women with normal and complicated pregnancies. Several studies have shown that urinary metabolome information is associated with blood metabolome information12,13. Moreover, urine can be collected in a minimally invasive manner at each routine antenatal visit in most clinical settings14.
The present study examined the dynamic urinary metabolomic profiles during pregnancy, then proposed a model to predict gestational age by means of urinary metabolomics analysis. Furthermore, by applying the predictive model to urinary metabolite data from women who subsequently developed pregnancy complications, this study revealed changes in predicted gestational ages, compared with actual ages, in women with complicated pregnancies.
Results
Maternal clinical backgrounds
Clinical backgrounds of study participants are shown in Table 1. Among the healthy pregnant women, the mean maternal age at the 40th gestational week and the mean pre-pregnancy body mass index were 32.7 ± 4.8 years, and 22.2 ± 4.7 kg/m2, respectively. Of the healthy cases, 51% were parous. The mean (± standard deviation) gestational age at delivery was 38.6 ± 1.2 weeks.
Compared with healthy pregnant women, the maternal age was higher among subjects with hypertensive disorders of pregnancy (HDP), while the gestational ages at delivery were lower in subjects with HDP or spontaneous preterm birth (SPTB). No other variables significantly differed between healthy pregnant women and subjects with complications.
Urinary metabolite profiles during normal pregnancy
We investigated urinary metabolomic profiles during normal pregnancy. Figure 1 shows hierarchical clustering of changes in the levels of urinary metabolites in healthy pregnant women (n = 187) during gestation. Relative levels of multiple metabolites changed dynamically according to gestational age. Approximately half of the metabolites were in cluster 1 (upper part of Fig. 1); the normalized levels of metabolites in this cluster increased with gestational age (correlation coefficient = 0.86). Cluster 1 included comparatively greater proportions of amino acids, peptides, and carbohydrates (Fig. 2, and Supplementary Table 1 online); for example, this cluster included sulfur-containing amino acids (e.g., cysteine and methionine) and monosaccharides (e.g., glucose and lactose). In contrast, the normalized levels of metabolites in cluster 3 were negatively correlated with gestational age (correlation coefficient = − 0.38). This cluster included greater proportions of purines and pyrimidines (e.g., cytosine, thymine, and uracil), compared with other clusters. Hydrophilic basic amino acids (e.g., histidine and lysine) were also present in this cluster. In contrast to metabolites in clusters 1 and 3, metabolites in cluster 2 did not show a clear linear relationship with gestational age (correlation coefficient = 0.24, see Supplementary Fig. 1 online). Comparatively greater proportions of amino acids and fatty acids (e.g., aromatic amino acids and stearic acid) were characteristic of cluster 2.
Changes in urinary metabolite levels during normal pregnancy. Levels of metabolites in urine samples from 187 healthy pregnant women were averaged for each gestational week, standardized by the z-score method, and presented as a heatmap. On the left, metabolite types are indicated using red, blue, yellow, green, and gray stripes; these indicate “amino acids, peptides, and analogs,” “carbohydrates and carbohydrate conjugates,” “fatty acids and conjugates,” “purines, pyrimidines, and their derivatives,” and “others,” respectively.
Predictive model to estimate gestational age
Using urinary metabolites from healthy pregnant women, we constructed a predictive model to estimate gestational age at the time of urine collection. Blue dots in Fig. 3 show actual and predicted days of gestation when leave-one-out cross validation was performed for each healthy pregnant woman. The root mean squared error between the actual and predicted days of gestation was 26.7, while the Pearson correlation coefficient was 0.86.
Prediction of gestational age in healthy pregnant women. Blue dots indicate actual days (horizontal axis) and predicted days (vertical axis) of gestation when leave-one-out cross validation was performed for each healthy pregnant woman using urine samples collected before delivery. Red dots indicate gestational ages estimated by the predictive model, which was trained with urine samples before delivery, using metabolite levels in samples collected after delivery.
Sixty-eight of the 184 metabolites consistently contributed to the predictive model; no metabolite coefficient reached zero during leave-one-out cross validation (see Supplementary Table 1 online). The top 10 metabolites with the greatest contributions to the predictive model are shown in Table 2. These contributions were evaluated by the absolute values of coefficients. Eight of these 10 metabolites were classified in either cluster 1 or 3 in Fig. 1. These results indicated that our predictive model successfully selected metabolites that were highly correlated with gestational age.
The results of enrichment analysis for the 68 metabolites are shown in Table 3. In cluster 1, metabolites related to cysteine metabolism were enriched. The metabolism of homocysteine, which is located upstream of cysteine metabolism, is considered crucial for the maintenance of a healthy pregnancy15. Metabolites related to vitamin and purine metabolic processes were significantly enriched in cluster 3.
Red dots in Fig. 3 indicate gestational age predicted by the model, using metabolite levels in urine samples collected after childbirth. Similar to the findings by Ghaemi et al., the predicted gestational age was significantly younger when our model was used with urine samples collected after childbirth than when our model was used with urine samples collected near delivery10.
Prediction of gestational age in complicated pregnancies
By entering urinary metabolite data from complicated pregnancies into the predictive model, we investigated potential differences in predicted gestational age between healthy and complicated pregnancies. After 20 weeks of gestation, the predicted gestational age was significantly older in HDP cases than in healthy pregnancies (Fig. 4). Comparison between healthy pregnancies and SPTB cases showed no significant differences in predicted gestational age or in metabolite levels (Fig. 4).
To analyze the effects of maternal age on the predictive model, we compared predicted ages between younger (< 35 years) and older (≥ 35 years) groups of women with normal pregnancies (see Supplementary Fig. 2 online). We found no significant differences between the two groups at any gestational age.
Subsequently, isocitric acid, 3-hydroxy-3-methylglutaric acid, and urocanic acid were identified as urinary metabolites that had significant associations with HDP onset (false discovery rate < 0.05) (see Supplementary Table 2 online). All three metabolites were classified in cluster 1 (Fig. 1); their urinary concentrations were higher in HDP cases than in healthy cases (see Supplementary Table 2 online).
As shown in Fig. 5, the levels of these three metabolites and the predicted gestational age were significantly associated with HDP onset at several weeks of gestation. Compared with each of the three metabolites, the predicted gestational age was more strongly associated with HDP onset.
Statistical analyses of associations of urinary metabolites or predicted gestational age with HDP onset. Urinary levels of isocitric acid, 3-hydroxy-3-methylglutaric acid, and urocanic acid, as well as the predicted gestational age, were significantly associated with HDP onset at several weeks of gestation. p-values calculated by the Wald test, adjusted for maternal age and urine type, are shown as a heatmap for each gestational week. Asterisks in panel A indicate statistical significance (p < 0.05).
Discussion
This study demonstrated dynamic changes in urinary metabolomic profiles during normal pregnancy. By means of machine learning algorithms, we constructed a predictive model that used urinary metabolite data to estimate gestational age during the progression of normal pregnancy. Moreover, we found that the predicted gestational age was significantly older than actual gestational age in cases with HDP.
This is a challenging attempt to predict gestational age in normal and complicated pregnancies by means of urinary metabolomics. Although a few studies have analyzed urinary metabolites in pregnant women, fewer than 50 metabolites were analyzed in each of those studies16,17,18.
We initially analyzed urine samples that were collected longitudinally from 187 healthy pregnant women, with the aim of elucidating urinary metabolite profiles during pregnancy. Hierarchical clustering showed that urinary metabolites comprised three distinct clusters (Fig. 1). The levels of metabolites contained in cluster 1, the largest cluster, increased with gestational age. These increased urinary metabolites during pregnancy might reflect the increased those in blood due to the reduction of maternal energy consumption associated with fetal and placental growth19,20. In a previous study, Diaz et al. used untargeted nuclear magnetic resonance-based metabolomics to examine temporal changes in the levels of 21 urinary metabolites during pregnancy; they identified 11 metabolites for which the levels significantly increased from the second trimester to the third trimester21. Three of those 11 metabolites (i.e., alanine, lactic acid, and threonine) were also identified in our study; all were included in cluster 1. Monosaccharides (e.g., glucose and lactose) were also included in cluster 1. The increased glucose level is consistent with the increased maternal insulin resistance that occurs during pregnancy22. A cohort study of 823 healthy pregnant women also showed similar results with respect to urinary lactose23. The increased lactose level was also present in blood metabolomics analyses; this was presumed to reflect maternal physiological changes to enable breastfeeding after delivery24.
Sixty-eight of the 184 metabolites with dynamic systematic changes consistently contributed to the predictive model; the coefficient of each metabolite never reached zero during leave-one-out cross validation. Therefore, these 68 metabolites are presumed to strongly reflect the progression of normal pregnancy. Enrichment analysis showed that 36 of the 68 metabolites (all in cluster 1) were related to cysteine metabolism. This suggests that homocysteine metabolism, which is located upstream of cysteine metabolism, is upregulated to maintain a healthy pregnancy15. Moreover, methionine, which is produced from homocysteine in the methionine cycle, was also classified in cluster 1, further supporting the potential upregulation of homocysteine metabolism.
The levels of metabolites classified in cluster 3 exhibited downward trends during pregnancy. This cluster included pyrimidines (e.g., cytosine, thymine, and uracil) and hydrophilic basic amino acids (e.g., histidine and lysine). Only a few previous studies have described the roles of, or changes in, these metabolites during pregnancy18,25. Furthermore, enrichment analysis of metabolites selected by the predictive model revealed significant enrichment of metabolites related to vitamins in cluster 3. Water-soluble vitamins (e.g., B6, B12, and C) play critical roles during pregnancy. For example, vitamin B12 is an important cofactor for DNA synthesis; it also participates in metabolic processes involving amino acids and fatty acids26. During pregnancy, vitamin B12 is supplied to the fetus by the mother through the placenta; the blood concentration of vitamin B12 is approximately twofold greater in the fetus than in the mother27. In addition, levels of vitamins B6, B12, and C in maternal blood have been reported to decrease during pregnancy27, which is consistent with our results. Because vitamins B6 and B12 are involved in homocysteine metabolism, the results of enrichment analysis for cluster 1 may reflect both maternal and fetal metabolism.
Using urinary metabolites, we successfully established a predictive model to estimate gestational age that exhibited performance equivalent to that of previously reported models involving plasma metabolites11. Therefore, gestational age could presumably be estimated from urinary metabolites. Because urine samples can be collected in a minimally invasive manner at each routine antenatal visit in most clinical settings, our results have potential clinical applications (e.g., in telemedicine using a medical examination kit).
Our predictive model mainly uses metabolites that demonstrate linear changes in levels during pregnancy. However, some metabolites demonstrate non-linear changes in levels (see Supplementary Fig. 1 online). For example, fatty acids play important roles in normal pregnancy; fatty acids reportedly accumulate in the mother during the first and second trimesters, then are released in the third trimester28. In the present study, many fatty acids were classified in cluster 2, such that they did not extensively contribute to the predictive model. However, the incorporation of metabolites with complex behaviors into our predictive model might enable more accurate prediction of gestational age.
Clinically, diagnosis of gestational weeks is conducted by using information regarding the last menstrual periods or fetal ultrasound findings at the early stage of pregnancy. This clinical information is sometimes inaccurate or inaccessible even in developed countries29,30,31. Therefore, estimation of gestational weeks by minimally invasive procedures is needed for various clinical settings in obstetrics, including determination of the due date, preterm birth and fetal growth. Taken together, our prediction model constructed by urinary metabolomics may provide useful clinical information for estimating gestational weeks in a variety of clinical settings.
The predicted gestational age was significantly older in cases who subsequently developed HDP. This result might be due to the presence of circulating metabolites caused by premature placental aging32. Premature placental aging associated with elevated oxidative stress and mitochondrial damages is known to cause placental insufficiency leading to HDP, including preeclampsia32,33,34. Urinary metabolite profiles also reportedly change with cellular senescence35. Therefore, the older predicted gestational age in the HDP cases, which implies aberrant acceleration of pregnancy, might be caused by changes in the urinary metabolite profile due to placental aging.
Importantly, we identified metabolites that were associated with the onset of HDP. For example, the increased level of 3-hydroxy-3-methylglutaric acid was significantly associated with the onset of HDP. In a rat model, 3-hydroxy-3-methylglutaric acid was able to induce physiological oxidative stress36. In addition, Nemeth et al. reported that the onset of HDP, including gestational hypertension, is related to increased maternal oxidative stress37. They indicated that insufficient capacity for glutathione recycling in patients with HDP might lead to reduced protection against oxidative stress. Furthermore, elevated oxidative stress is regarded as a manifestation of preeclampsia; for example, in the placentas of patients with preeclampsia, increased lipid peroxidation and decreased activities of some antioxidant enzymes have been identified38. Our results are suggestive of such physiological aberrations.
The increased levels of isocitric and urocanic acids also showed significant associations with HDP onset. Although isocitric acid is a component of the tricarboxylic acid cycle, the level of succinic acid (located downstream of isocitric acid) did not significantly differ between the healthy and HDP groups. This suggests reduced conversion of isocitric acid to oxalosuccinic acid, which requires nicotinamide adenine dinucleotide (NAD+). Therefore, the cellular NAD+ level may be reduced in cases who develop HDP39. Because NAD+ is required for DNA repair, DNA damage in the placenta caused by elevated oxidative stress associated with HDP might lead to a reduction of the NAD+/NADH ratio40. Urocanic acid is a histidine metabolite and a major component of ultraviolet light absorption in the skin. Its concentration in the plasma is reportedly elevated in cases with preeclampsia41, in agreement with our results.
In this study, we demonstrated that the predicted gestational age calculated by combining multiple metabolite levels could more accurately predict the onset of HDP, compared with each metabolite level alone (Fig. 5). This finding suggests that a combination of metabolites—each with weak explanatory power—could more accurately predict the risk, compared with individual metabolites; this was similar to the previously described polygenic risk score, which comprised the weighted sum of alleles associated with some traits42.
There were no significant differences in predicted gestational age or the levels of metabolites between the healthy and SPTB groups. Previous studies reported an association between urinary phthalate metabolites and SPTB43,44. Unfortunately, the metabolites could not be detected by gas chromatography-tandem mass spectrometry (GC–MS/MS) in our study. Although associations between maternal urinary or blood metabolites and SPTB development have rarely been described, various metabolites in cervicovaginal fluid, amniotic fluid, and neonatal urine are reportedly associated with SPTB45,46,47. SPTB often involves inflammatory changes caused by local bacterial infection in organs or tissues such as the vagina, decidua, placenta, and amniotic cavity48. Therefore, manifestations of SPTB may not be detected with urinary metabolites.
The present study had several limitations. First, all study participants were recruited at a single facility in Japan; therefore, our predictive model must be validated in independent facilities and in cohorts of patients with different ethnicities49. Second, we studied physiological changes in pregnant women solely on the basis of urinary metabolites; thus, comprehensive analyses that consider other omics data are warranted. In a previous prospective cohort study (i.e., the Maternity Log [MLOG] study), we obtained multi-omics information including the plasma metabolome, blood transcriptome, and urinary metabolome49. We hope that the validity of our predictive model will be verified by the analysis of relationships among urinary metabolites and other multi-omics data in the future. Third, the number of healthy pregnant women available for construction of the predictive model in this study was insufficient. In the TMM BirThree Cohort Study, which is a parent cohort of the MLOG study, 23,406 pregnant women were recruited50,51; moreover, urine samples were collected twice during pregnancy from each participant for use in the quantification of urinary metabolites. Urinary metabolite information from a very large number of pregnant women could further elucidate metabolomic changes during pregnancy; this might improve the predictive performance of our model. Finally, only relative quantification of metabolites was performed in this study. For metabolites that were found to have a significant association with the onset of HDP, we plan to increase the reliability of the evidence by absolute quantification in the near future.
In conclusion, we demonstrated dynamic changes in urinary metabolomic profiles of 184 metabolites during pregnancy, a predictive model was constructed by using urinary metabolite information to estimate gestational age at the time of urine specimen collection. The results suggested that urinary metabolite information is useful for understanding the normal progression of pregnancy, as well as for predicting the development of pregnancy complications. Minimally invasive urinary metabolomics might lead to breakthroughs in the analysis and management of healthy and complicated pregnancies in various clinical settings in the future.
Methods
Study setting
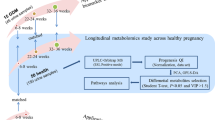
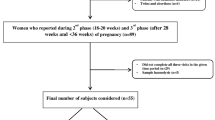
In total, 302 pregnant women were recruited at Tohoku University Hospital, Sendai, Japan, between September 2015 and November 2016; these women were previously included in the MLOG study49, which is an add-on study to the TMM BirThree Cohort Study50,51. Written informed consent was obtained from all participants by the genome medical research coordinators. The MLOG study was conducted under a collaborative research agreement among Tohoku Medical Megabank Organization, Tohoku University, and NTT DOCOMO, Inc. (Tokyo, Japan) and all procedures were in accordance with the Declaration of Helsinki. The study protocol was approved by the ethics committees of the Graduate School of Medicine (2014-1-704) and Tohoku Medical Megabank Organization (22017-1-085) at Tohoku University.
As shown in Supplementary Fig. 3 online, 187 healthy pregnant women were selected from the 302 recruited subjects; the data from these 187 women were used for the construction of a model to predict gestational age. Healthy pregnant women were defined as subjects who had a singleton pregnancy, gave birth at term, and did not develop any pregnancy complications. With respect to pregnancy complications observed in this study, 23 subjects developed HDP (excluding those with chronic hypertension) and 14 subjects gave SPTB. HDP was defined as gestational hypertension, preeclampsia, or superimposed preeclampsia, excluding chronic hypertension52,53. SPTB was defined as spontaneous labor at fewer than 37 weeks of gestation, excluding delivery by cesarean section.
Urine collection
Either early morning first urine or spot urine specimen (10 mL) was collected at each antenatal visit; 12 samples were obtained from each subject. In total, 2741 urine samples were collected from healthy, HDP, or SPTB cases (2140 early morning first urine and 601 spot urine samples). All samples were stored at − 80 °C immediately after collection, then subjected to GC–MS/MS analysis.
Chemical reagents
The following chemical reagents were used in this study: pyridine (Cat. No. Q003, Tokyo Chemical Industry Co. Ltd., Tokyo, Japan), methanol (Cat. No. 25185-76, Kanto Kagaku, Tokyo, Japan), chloroform (Cat. No. 07278-79, Kanto Kagaku), water (MilliQ, Millipore, Burlington, MA, USA), 2-isopropyl malic acid (Cat. No. 333115-100MG, Sigma-Aldrich, St. Louis, MO, USA), methoxyamine hydrochloride (Cat. No. 226904-25G, Sigma-Aldrich), and N-methyl-N-(trimethylsilyl) trifluoroacetamide (Cat. No. 1022-11061, GL Science, Saitama, Japan).
GC–MS/MS analysis
To determine metabolite levels in maternal urine samples, GC–MS/MS analysis was performed using a previously described method49,54. The urine metabolites were extracted by a robotic system (Microlab STARlet Robot System, Hamilton, Reno, NV, USA). Fifty microliter of each urine sample was added to 260 μL of extraction solvent, consisting of chloroform/methanol/water/internal standard solution at a ratio of 1/2.5/1/0.18 (vol/vol/vol/vol); the internal standard solution contained 0.5 mg/mL 2-isopropylmalic acid. Each sample was shaken at 37 °C for 30 min, then centrifuged at 6231×g for 5 min at 4 °C. The supernatant was transferred to a 1.5-mL tube and lyophilized with a freeze-drying system (FDS-1000, EYELA, Tokyo, Japan). For the first derivatization, the residue was resuspended using 20 mg/mL methoxyamine hydrochloride in pyridine (80 μL) at 37 Hz for 20 min in an ultrasonic bath (UT-10, Sharp, Osaka, Japan); it was incubated at 1200 rpm for 90 min at 30 °C. After centrifugation at 16,000×g for 3 min at 4 °C, the supernatant (40 μL) was transferred to a glass vial and placed on the multifunctional autosampler system (AOC6000, Shimadzu, Kyoto, Japan). The metabolites in the sample were automatically derivatized with 20 μL of N-methyl-N-(trimethylsilyl)trifluoroacetamide using the multifunctional autosampler. The sample was then analyzed with gas chromatography triple quadrupole mass spectrometry (TQ-8040, Shimadzu). The sample (1 μL) was exposed to an injector (split ratio 30:1); the targeted analytes, including 2-isopropylmalic acid (internal standard), were separated using a capillary column (BPX5: 30-m length, 0.25-μm film thickness, 0.25-mm inner diameter; Cat. No. SGE054101, Shimadzu GLC Ltd., Tokyo, Japan) under helium gas with constant linear velocity of 39.0 cm/sec. The inlet temperature was set to 250 °C; the oven temperature was initially set to 60 °C for 2 min, then changed to 330 °C at a rate of 15 °C/min, and finally maintained at this temperature for 3 min. The electron ionization voltage, ion source temperature, and MS interface temperature were 70 eV, 200 °C, and 280 °C, respectively. The metabolites were identified with the Smart Metabolites Database (Shimadzu) and the peak area was calculated by Traverse MS (Reifycs, Tokyo, Japan). To adjust for measurement errors during each GC–MS/MS analysis, normalization was first conducted using 2-isopropylmalic acid as an internal standard added to each sample (see Supplementary Fig. 4 online). Normalization was repeated using quality control samples (see Supplementary Fig. 4 online) mixed second-trimester human urine (IR100040-Donor-HG, Innovative Research, Inc., Novi, MI, USA), third-trimester human urine (IR100041-DONOR-JT, Innovative Research, Inc.), and our study urine samples, which were injected at intervals of 12 study samples, in accordance with the reference quality control method55. Normalized levels of metabolites in the quality control samples were assessed by calculating coefficients of variation; metabolites with coefficients of variation > 20% were eliminated. Finally, 184 metabolites fulfilled the criteria of the second normalization. Thereafter, to avoid the dilution effect of urine, the levels of those metabolites were compensated by creatinine concentration in the samples measured by a biochemical test (Detaminer-L CRE, Hitachi Chemical Diagnostics Systems Co., Ltd., Tokyo, Japan).
Data analysis
To investigate the profiles of urinary metabolites during normal pregnancy, we determined the average level of each metabolite according to gestational age in healthy pregnant women. The average levels were standardized for each metabolite by the z-score method, then subjected to hierarchical clustering analysis and trend analysis. The Ward variance minimization algorithm was selected for the clustering analysis56.
Subsequently, using the levels of urinary metabolites in healthy pregnant women, we constructed a predictive model to estimate gestational age at the time of urine collection. Elastic net regression was used for model construction, with the objective variable as the gestational age (days) and the explanatory variable as the logarithmic level of each of 184 metabolites. Elastic net regression has a substantial learning ability for features with multicollinearity, such as omics data10. The logarithms of metabolite levels were used to reduce the effects of outliers. Leave-one-out cross validation was performed; the data of one healthy pregnant woman were used as the test data, and after learning with the data of others, prediction was performed for the test data. This cycle was repeated according to the number of healthy pregnant women (n = 187). Then, the predictive performance of all combined test data was evaluated by the root mean squared error and Pearson correlation coefficient between actual and predicted gestational age.
Using the MBROLE 2.0 web-based analysis tool57, we performed enrichment analysis for 68 metabolites that consistently contributed to the prediction; no metabolite coefficient in the model reached zero during 187 cycles in the leave-one-out cross validation. Enrichment analysis was performed for each metabolite cluster, targeting the biofunctions defined by Human Metabolome Database58.
Finally, by entering urinary metabolite data from HDP or SPTB cases into the model constructed by data from all healthy pregnant women in this study, we examined whether outputs significantly differed between healthy and complicated pregnancies.
For 68 metabolites, we also identified those that were associated with the onset of HDP. The association between the onset and each metabolite according to gestational age was confirmed by using the Wald test for logistic regression, with the onset of HDP as the objective variable (i.e., healthy = 0 and HDP = 1) and the level of each metabolite as the explanatory variable.
Principal component analysis (PCA) of 2741 urine samples revealed significant differences between early morning first urine and spot urine samples with respect to both first and second principal components (see Supplementary Figs. 5 and 6 online). Therefore, when constructing our predictive model for gestational age, we used the levels of urinary metabolites in healthy pregnant women, as well as their urine sample types (i.e., early morning urine = 0 and spot urine = 1). In hierarchical clustering analysis, we used only early morning urine samples; these are less likely to be affected by each subject’s lifestyle or behavior. Furthermore, we found a significant difference in maternal age between healthy and HDP cases (Table 1). Therefore, to exclude the effects of maternal age and urine type, we evaluated the association between each metabolite and the onset of HDP by using the Wald test for logistic regression, with maternal age and urine type as covariances. Considering the number of metabolites (n = 68), we used the Benjamini–Hochberg method to compensate for multiple Wald tests. Metabolites with a false discovery rate of less than 0.05 at any one gestational week were defined as those significantly associated with HDP.
Python libraries as follows; (1) Seaborn59, (2) Scikit-learn60, and (3) Statsmodels61, were used for (1) the hierarchical clustering analysis and each figure visualization, (2) the trend analysis, predictive model construction and PCA, and (3) the Wald test for logistic regression, respectively.
Data are presented as the mean ± S.D. (standard deviation) unless otherwise indicated. Computational resources were provided by the Tohoku Medical Megabank Organization supercomputer system.
References
Granger, J. P. Maternal and fetal adaptations during pregnancy: Lessons in regulatory and integrative physiology. Am. J. Physiol. Regul. Integr. Comp. Physiol. 283, 1289–1292 (2002).
Hong, S. et al. Longitudinal profiling of human blood transcriptome in healthy and lupus pregnancy. J. Exp. Med. 216, 1154–1169 (2019).
Nelissen, E., van Montfoort, A., Dumoulin, J. & Evers, J. Epigenetics and the placenta. Hum. Reprod. Update 17, 397–417 (2011).
Lindsay, K. L. et al. Longitudinal metabolomic profiling of amino acids and lipids across healthy pregnancy. PLoS ONE 10, e0145794. https://doi.org/10.1371/journal.pone.0145794 (2015).
Koren, O. Host remodeling of the gut microbiome and metabolic changes during pregnancy. Cell 150, 470–480 (2012).
Benny, P. A., Alakwaa, F. M., Schlueter, R. J., Lassiter, C. B. & Garmire, L. X. A review of omics approaches to study preeclampsia. Placenta 92, 17–27 (2020).
Huynh, J., Xiong, G. & Bentley-Lewis, R. A systematic review of metabolite profiling in gestational diabetes mellitus. Diabetologia 57, 2453–2464 (2014).
Tarca, A. L. et al. Targeted expression profiling by RNA-Seq improves detection of cellular dynamics during pregnancy and identifies a role for T cells in term parturition. Sci. Rep. 9, 848 (2019).
Roberto, R. et al. The maternal plasma proteome changes as a function of gestational age in normal pregnancy: A longitudinal study. Am. J. Obstet. Gynecol. 217, 1–21 (2017).
Ghaemi, M. S. et al. Multiomics modeling of the immunome, transcriptome, microbiome, proteome and metabolome adaptations during human pregnancy. Bioinformatics 35, 95–103 (2019).
Liang, L. et al. Metabolic dynamics and prediction of gestational age and time to delivery in pregnant women. Cell 181, 1680–1692 (2020).
Feng, Q. et al. Integrated metabolomics and metagenomics analysis of plasma and urine identified microbial metabolites associated with coronary heart disease. Sci. Rep. 6, 22525. https://doi.org/10.1038/srep22525 (2016).
Do, K. T. et al. Network-based approach for analyzing intra- and interfluid metabolite associations in human blood, urine, and saliva. J. Proteome Res. 14, 1183–1194 (2015).
Miller, S. et al. Beyond too little, too late and too much, too soon: A pathway towards evidence-based, respectful maternity care worldwide. Lancet 388, 2176–2192 (2016).
Hague, W. M. Homocysteine and pregnancy. Best Pract. Res. Clin. Obstet. Gynaecol. 17, 459–469 (2003).
Maitre, L. et al. Urinary metabolic profiles in early pregnancy are associated with preterm birth and fetal growth restriction in the Rhea mother–child cohort study. BMC Med. 12, 110. https://doi.org/10.1186/1741-7015-12-110 (2014).
Austdal, M. et al. Metabolomic biomarkers in serum and urine in women with preeclampsia. PLoS ONE 9, e91923. https://doi.org/10.1371/journal.pone.0091923 (2014).
Liu, X. et al. Urinary metabolic variation analysis during pregnancy and application in gestational diabetes mellitus and spontaneous abortion biomarker discovery. Sci. Rep. 9, 2605 (2019).
Catalano, P. M., Drago, N. M. & Amini, S. B. Maternal carbohydrate metabolism and its relationship fetal growth and body composition. Am. J. Obstet. Gynecol. 172, 1464–1470 (1995).
Herrera, E. Metabolic adaptations in pregnancy and their implications for the availability of substrates to the fetus. Eur. J. Clin. Nutr. 54, 47–51 (2000).
Diaz, S. O. et al. Following healthy pregnancy by nuclear magnetic resonance (NMR) metabolic profiling of human urine. J. Proteome Res. 12, 969–979 (2012).
Barbour, A. L. et al. Cellular mechanisms for insulin resistance in normal pregnancy and gestational diabetes. Diabetes Care 30, 112–119 (2007).
Sachse, D. et al. Metabolic changes in urine during and after pregnancy in a large, multiethnic population-based cohort study of gestational diabetes. PLoS ONE 7, e52399. https://doi.org/10.1371/journal.pone.0052399 (2012).
Neville, M. C., Morton, J. & Umemura, S. Lactogenesis: The transition from pregnancy to lactation. Pediatr. Clin. N. Am. 48, 35–52 (2001).
Park, S., Park, J. Y., Lee, J. H. & Kim, S. Plasma levels of lysine, tyrosine, and valine during pregnancy are independent risk factors of insulin resistance and gestational diabetes. Metab. Syndr. Relat. Disord. 13, 64–70 (2015).
Scott, J. M. Folate and vitamin B12. Proc. Nutr. Soc. 58, 441–448 (1999).
Dror, D. K. & Allen, L. H. Interventions with vitamins B6, B12 and C in pregnancy. Paediatr. Perinat. Epidemiol. 26, 55–74 (2012).
Herrera, E. Implications of dietary fatty acids during pregnancy on placental, fetal and postnatal development: A review. Placenta 23, 9–10 (2002).
Swanson, D. L. et al. Including ultrasound scans in antenatal care in low-resource settings: Considering the complementarity of obstetric ultrasound screening and maternity waiting homes in strengthening referral systems in low-resource, rural settings. Semin. Perinatol. 43, 273–281 (2019).
Scott, K. et al. “I can guess the month … but beyond that, I can’t tell” an exploratory qualitative study of health care provider perspectives on gestational age estimation in Rajasthan, India. BMC Pregnancy Childbirth 20, 529. https://doi.org/10.1186/s12884-020-03201-6 (2020).
Martin, J. A. et al. Births: final data for 2017. Natl. Vital Stat. Rep. 67, 1–50 (2018).
Manna, S., McCarthy, C. & McCarthy, F. P. Placental ageing in adverse pregnancy outcomes: Telomere shortening, cell senescence, and mitochondrial dysfunction. Oxid. Med. Cell. Longev. 2019, 3095383 (2019).
Wang, Y., Lim, R. & Nie, G. Elevated circulating HtrA4 in preeclampsia may alter endothelial expression of senescence genes. Placenta 90, 71–81 (2020).
Nezu, M. et al. Nrf2 inactivation enhances placental angiogenesis in a preeclampsia mouse model and improves maternal and fetal outcomes. Sci. Signal. 10, eaam5711 (2017).
Hertel, J. et al. Measuring biological age via metabonomics: The metabolic age score. J. Proteome Res. 15, 400–410 (2016).
Wysocki, S. J. & Hahnel, R. 3-Hydroxy-3-methylglutaryl-coenzyme A lyase deficiency: A review. Inherit. Metab. Dis. 9, 225–233 (1986).
Nemeth, I., Orvos, H. & Boda, D. Blood glutathione redox status in gestational hypertension. Free Radic. Biol. Med. 30, 715–721 (2001).
Burton, G. J. & Jauniaux, E. Placental oxidative stress: From miscarriage to preeclampsia. J. Soc. Gynecol. Investig. 11, 342–352 (2004).
Fouquerel, E. & Sobol, R. W. ARTD1 (PARP1) activation and NAD+ in DNA repair and cell death. DNA Repair 23, 27–32 (2014).
Fujimaki, A. et al. Placental oxidative DNA damage and its repair in preeclamptic women with fetal growth restriction. Placenta 32, 367–372 (2011).
Sander, K. N. et al. Untargeted analysis of plasma samples from pre-eclamptic women reveals polar and apolar changes in the metabolome. Metabolomics 15, 157. https://doi.org/10.1007/s11306-019-1600-8 (2019).
Khera, A. V. et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 50, 1219–1224 (2018).
Ferguson, K. K., McElrath, T. F. & Meeker, J. D. Environmental phthalate exposure and preterm birth. JAMA Pediatr. 168, 61–67 (2014).
Meeker, J. D. et al. Urinary phthalate metabolites in relation to preterm birth in Mexico City. Environ. Health Perspect. 117, 1587–1592 (2009).
Amabebe, E. et al. Identifying metabolite markers for preterm birth in cervicovaginal fluid by magnetic resonance spectroscopy. Metabolomics 12, 67. https://doi.org/10.1007/s11306-016-0985-x (2016).
Menon, R. et al. Amniotic fluid metabolomic analysis in spontaneous preterm birth. Reprod. Sci. 21, 791–803 (2014).
Atzori, L. et al. 1 H NMR-based metabolomic analysis of urine from preterm and term neonates. Front. Biosci. 3, 1005–1012 (2011).
Di Renzo, G. C., Tosto, V. & Giardina, I. The biological basis and prevention of preterm birth. Best Pract. Res. Clin. Obstet. Gynaecol. 52, 13–22 (2018).
Sugawara, J. et al. Maternity Log Study: a longitudinal lifelog monitoring and multiomics analysis for the early prediction of complicated pregnancy. BMJ Open 9, e025939. https://doi.org/10.1136/bmjopen-2018-025939 (2018).
Kuriyama, S. et al. Cohort profile: Tohoku medical megabank project birth and three-generation cohort study (TMM BirThree cohort study): Rationale, progress and perspective. Nt. J. Epidemiol. https://doi.org/10.1093/ije/dyz169 (2020).
Sugawara, J. et al. Maternal baseline characteristics and perinatal outcomes: The Tohoku medical megabank project birth and three-generation cohort study. J. Epidemiol. https://doi.org/10.2188/jea.JE20200338 (2020).
Watanabe, K., Naruse, K., Tanaka, K., Metoki, H. & Suzuki, Y. Outline of definition and classification of “pregnancy induced hypertension (PIH)”. Hypertens. Res. Pregnancy 1, 3–4 (2013).
Metzger, B. E. International Association of Diabetes and Pregnancy Study Groups recommendations on the diagnosis and classification of hyperglycemia in pregnancy. Diabetes Care 33, 676–682 (2010).
Saigusa, D., Matsukawa, N., Hishinuma, E. & Koshiba, S. Identification of biomarkers to diagnose diseases and find adverse drug reactions by metabolomics. Drug Metab. Pharmacokinet. 20, 30434. https://doi.org/10.1016/j.dmpk.2020.11.008 (2021).
Saigusa, D. et al. Establishment of protocols for global metabolomics by LC-MS for biomarker discovery. PLoS ONE 11, e0160555. https://doi.org/10.1371/journal.pone.0160555 (2016).
Johnson, S. C. Hierarchical clustering schemes. Psychometrika 32, 241–254 (1967).
Lopez-Ibanez, J., Pazos, F. & Chagoyen, M. MBROLE 2.0: Functional enrichment of chemical compounds. Nucl. Acids Res. 44, 201–204 (2016).
Wishart, D. S. et al. HMDB: The human metabolome database. Nucl. Acids Res. 35, 521–526 (2007).
Waskom, L. M. Seaborn: Statistical data visualization. J. Open Source Softw. 6, 3021. https://doi.org/10.21105/joss.03021 (2021).
Pedregosa, F. et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Seabold, S. & Perktold, J. Statsmodels: Econometric and statistical modeling with Python. In Proceedings of the 9th Python in Science Conference 57–61 (2010).
Acknowledgements
We thank all past and present members of Tohoku Medical Megabank Organization at Tohoku University (present members are listed at https://www.megabank.tohoku.ac.jp/english/a200601/). We also thank Ryan Chastain-Gross, Ph.D., from Edanz Group (https://en-author-services.edanz.com/ac) for editing a draft of this manuscript. This study was supported by NTT DOCOMO, Inc., with a collaborative research agreement among NTT DOCOMO, Tohoku Medical Megabank Organization, and Tohoku University. The Tohoku Medical Megabank is supported by grants from the Reconstruction Agency, the Ministry of Education, Culture, Sports, Science and Technology (MEXT), and the Japan Agency for Medical Research and Development (AMED). This study was supported in part by AMED under grant numbers JP19km0105001 and JP19km0105002. This work was also partially supported by JSPS KAKENHI Grant Number JP18KT0015 and JP18H03326. The Radiation Effects Research Foundation (RERF) is a public interest foundation funded by the Japanese Ministry of Health, Labour and Welfare (MHLW) and the United States Department of Energy.
Author information
Authors and Affiliations
Contributions
Conceptualization, data analysis, and writing the manuscript, T.Y. and J.S.; advice or assistance for data analysis, D.O., Y.T., and S.K.; recruiting subjects, M.I., T.O., and S.K.; urine collection, N.M., R.Y., and N.M.; GC–MS/MS analysis, N.M., D.S., R.Y., and S.K.; editing the manuscript, O.T., H.M., M.I., S.H., and J.S.; reviewing the manuscript, all authors; supervision, M.N. and S.H.; project administration, J.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yamauchi, T., Ochi, D., Matsukawa, N. et al. Machine learning approaches to predict gestational age in normal and complicated pregnancies via urinary metabolomics analysis. Sci Rep 11, 17777 (2021). https://doi.org/10.1038/s41598-021-97342-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-97342-z
- Springer Nature Limited
This article is cited by
-
Hypertensive disorders of pregnancy: definition, management, and out-of-office blood pressure measurement
Hypertension Research (2022)