Abstract
The ESR1 rs9340799 polymorphism has been frequently investigated with regard to its association with breast cancer (BC) susceptibility, but the findings have been inconclusive. In this work, we aimed to address the inconsistencies in study findings by performing a systematic review and meta-analysis. Eligible studies were identified from the Web of Science, PubMed, Scopus, China National Knowledge Infrastructure, VIP and Wanfang databases based on the predefined inclusion and exclusion criteria. The pooled odds ratio (OR) was then calculated under five genetic models: homozygous (GG vs. AA), heterozygous (AG vs. AA), dominant (AG + GG vs. AA), recessive (GG vs. AA + AG) and allele (G vs. A). Combined results from 23 studies involving 34,721 subjects indicated a lack of significant association between the polymorphism and BC susceptibility (homozygous model, OR = 1.045, 95% CI 0.887–1.231, P = 0.601; heterozygous model, OR = 0.941, 95% CI 0.861–1.030, P = 0.186; dominant model, OR = 0.957, 95% CI 0.875–1.045, P = 0.327; recessive model, OR = 1.053, 95% CI 0.908–1.222, P = 0.495; allele model, OR = 0.987, 95% CI 0.919–1.059, P = 0.709). Subgroup analyses by ethnicity, menopausal status and study quality also revealed no statistically significant association (P > 0.05). In conclusion, our results showed that the ESR1 rs9340799 polymorphism was not associated with BC susceptibility, suggesting its limited potential as a genetic marker for BC.
Similar content being viewed by others
Introduction
According to the World Health Organization statistics, breast cancer (BC) is the most common malignant tumor type, as well as a leading cause of mortality in the female population1,2. Like other malignancies, risk factors of BC are primarily genetic predisposition and environmental influences3. It has been reported that genetic background or familial history accounts for ~ 20–25% of overall BC incidence4. Among the ~ 80 genetic loci known to be associated with susceptibility to BC, the BRCA1 and BRCA2 loci carry the highest risk5,6. Together with other high- and medium-penetrance germline mutations located at the loci of TP53, PTEN, ATM, BRIP1, CHEK2 and PALB2, they made up ~ 15–20% of the genetic risk of BC7,8,9. Common low‑penetrance genetic polymorphisms account for the remaining risk for BC susceptibility10,11. On the other hand, environmental and lifestyle risk factors for BC include the consumption of oral contraceptive, cigarette smoking, alcohol consumption, breastfeeding and delayed age at first childbirth12,13,14.
Among these risk factors, it has been specifically pointed out that estrogen can act as a carcinogen, not only by causing chromosome segregation errors as well as structural chromosomal alterations, but also by stimulating the uncontrolled proliferation of mutated breast cells4,15,16. Mounting evidence from population-based studies has corroborated the association of endogenous and exogenous circulating estrogen in BC etiology and the increased risk of BC in premenopausal women17. The physiological receptors for estrogen are estrogen receptors (ER), which function to mediate the effect of estrogen on breast cells. Binding of estrogen to ER promotes the growth and differentiation of the normal breast cells and can lead to breast carcinogenesis18. There are two ER isoforms, i.e., ERα and ERβ. These two ER isoforms are respectively encoded by two distinct genes, ESR1 and ESR219. ERα has a higher level of expression in the breast tissue between these two isoforms, hence it is frequently implicated in BC development20.
The focus of this meta-analysis is the ERα-encoding gene, ESR1, which is highly polymorphic. Among the many polymorphisms in ESR1, the two best-studied ones are rs2234693 (also known as PvuII or 397T>C) and rs9340799 (also known as XbaI or 351A>G) polymorphisms. Both polymorphisms are located in intron 1, respectively at 1,397 bp and 351 bp upstream of exon 2 of the gene, and have been associated with several female cancers, including BC and endometrial cancer21,22,23. However, the association of the two polymorphisms with BC susceptibility has been described with conflicting results in many studies24,25,26,27. To sort out this inconsistency, a meta-analysis on rs2234693 was performed in 2018, and showed that the polymorphism was significantly associated with a decreased BC susceptibility28. As for rs9340799, a meta-analysis based on seven previous studies was reported by Zhang et al. in 2015, and found no significant association between the polymorphism and BC susceptibility under all three genetic models examined, even when the data were stratified into subgroups according to the ethnicity and source of controls29. In this current work, we attempted to perform an updated meta-analysis on the relationship between ESR1 rs9340799 polymorphism and BC susceptibility, by including a large number of additional studies that have been left out by Zhang et al. or have only been published after 2015.
Results
Study selection and characteristics
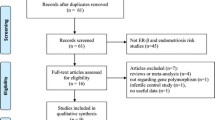
The study selection process is depicted in Fig. 1. The initial database and bibliographic searches identified 236 records (PubMed, N = 51; Scopus, N = 44; WoS, N = 114; Wanfang, N = 20; CNKI, N = 7; VIP, N = 0). After duplicated records were removed, 153 articles were screened by title and abstract. Thirty seven (37) articles were subsequently identified as potentially relevant and checked for eligibility by full-text review. Of these, 13 articles that did not meet the eligibility criteria and one article that had an inappropriate study design (as male controls were included in the analysis)30 were excluded. In addition, two articles were found to contain overlapping data18,31, and the one with the smaller sample size was excluded31. Ultimately, 22 articles comprising 23 studies were included for the quantitative synthesis of data3,18,21,24,25,27,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47.
The 23 included studies involved a total of 34,721 subjects (12,766 cases and 21,955 controls). Among the included studies, eight (from seven articles) reported data for pre- and postmenopausal women separately3,18,21,25,34,38,41, and four other studies included only postmenopausal women24,27,39,40. The remaining studies either did not mention the menopausal status or did not perform separate analyses for pre- and postmenopausal women. In terms of ethnicity, nine studies were conducted on Asians3,18,25,32,34,37,38,41,44, nine on Caucasian21,24,27,33,35,36,39,40,47, three on other ethnicities43,45,46, and two on mixed ethnicities21,42. All studies were case–control in design. Fifteen (15) of the studies were considered as having high quality, whereas eight had low quality (Supplementary Table S1 online). The characteristics of the included studies are summarized in Table 1.
Meta-analysis results
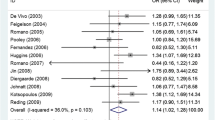
The meta-analysis results are shown in Table 2. Overall, no statistically significant association was observed between ESR1 rs9340799 polymorphism and BC susceptibility (homozygous model, OR = 1.045, 95% CI 0.887–1.231, P = 0.601; heterozygous model, OR = 0.941, 95% CI 0.861–1.030, P = 0.186; dominant model, OR = 0.957, 95% CI 0.875–1.045, P = 0.327; recessive model, OR = 1.053, 95% CI 0.908–1.222, P = 0.495; allele model, OR = 0.987, 95% CI 0.919–1.059, P = 0.709). The random-effects model was used in the above analyses as significant heterogeneity was present in all genetic models. The forest plots of the associations are presented in Fig. 2. Sensitivity analysis revealed that none of the individual studies had significant impact on the pooled OR (Supplementary Fig. S1 online).
Subgroup analyses
Subgroup analyses were performed based on the ethnicity (Asian vs. Caucasian) and menopausal status (premenopause vs. postmenopause) of the study subjects, as well as the quality of the studies (high quality vs. low quality). No statistical significant association was observed for all subgroups under all genetic models (P > 0.05; Table 2).
Although significant heterogeneity was observed in the overall analysis, several subgroups were found to have low heterogeneity based on the I2 value. In the homozygous model, low heterogeneity was found for Asians (I2 = 0.0%), premenopause (I2 = 0.0%), postmenopause (I2 = 0.0%) and low quality (I2 = 19.8%) subgroups. A similar observation was observed for the recessive model (Asians, I2 = 16.9%; premenopause, I2 = 0.0%; postmenopause, I2 = 0.0%; low quality, I2 = 45.4%). In heterozygous model, the Caucasian (I2 = 18.8%) and high quality (I2 = 34.9%) subgroups showed low heterogeneity, whereas in allele model, low heterogeneity was noted in premenopause (I2 = 24.5%), postmenopause (I2 = 47.5%) and low quality subgroups (I2 = 46.7%). All subgroups in the dominant model showed high heterogeneity (I2 > 50%).
Publication bias
No evidence of asymmetry was detected in the funnel plots of all genetic models (Fig. 3), indicating the absence of publication bias. This observation was corroborated by the results of Begg’s and Egger’s tests (homozygous model, Begg's test P = 0.529, Egger's test P = 0.625; heterozygous model, Begg's test P = 0.978, Egger's test P = 0.152; dominant model, Begg's test P = 0.800, Egger's test P = 0.366; recessive model, Begg's test P = 0.488, Egger's test P = 0.303; allele model, Begg's test P = 0.636, Egger's test P = 0.937).
Discussion
ERα, a member of the nuclear receptor superfamily, is encoded by a ~ 300 kb gene, ESR1, which is mapped to chromosomal locus 6q25.1 and contains eight exons. It has been documented that the human ESR1 gene contains at least nine promoters, whereby each promoter harbors multiple transcription factors-binding sites48. The ERα protein possesses DNA- and ligand-binding domains which are highly conserved49. It is depicted that ERα can mediate the effect of estrogen via several molecular pathways. Among these, the classical pathway is the best-known. In this direct pathway, unliganded ERα forms a cytosolic complex with Hsp90. Upon estrogen binding to the ligand-binding domains of ERα, the ERα-Hsp90 complex dissociates. Subsequently, ERα dimerizes and translocates to the nucleus. Following that, the DNA-binding domains of ERα, consisting of two functionally distinct zinc finger motifs, bind to a characteristic stretch of DNA sequence named the estrogen response elements in the promoters of the target genes to influence the process of transcription50.
Meanwhile, the tethered pathway entails protein–protein interaction or heterodimerization of ERα with other transcription factors such as AP1or NF-kB after ligand activation. This results in the indirect binding of DNA by ERα, contributing to the regulation of target genes including insulin-like growth factor 1, cathepsin D, progesterone receptor, transforming growth factor α, pS2, retinoic acid receptor α1, c-myc, etc., which are essential for cell proliferation and survival51. The nongenomic pathway typically involves a small plasma membrane population- and cytoplasm-based ERα52, which interacts with signaling proteins such as Src, mitogen-activated protein kinase (MAPK) and phosphoinositide 3-kinase. These signaling molecules can activate the phosphorylation of ERα and its coregulators53,54. This subsequently triggers signaling cascades via second messengers (SM), and eventually, it enhances nuclear ERα signaling without involving gene regulation. The last ER pathway is the ligand-independent pathway. In this case, ERs can become activated via crosstalk with other signaling pathways, e.g. the insulin-like growth factor-1 receptor and the epidermal growth factor receptor pathways55. In these instances, ERs are activated by phosphorylation to form dimers, to bind DNA, and regulate the expression of genes.
Notwithstanding, all models of ERα signaling pathways point to the vital role of ERα in the proliferation and survival of breast epithelial cells, as well as mammary tumorigenesis54. ER has been used as a molecular classifier for breast tumors, whereby BC can be graded as ER-positive and ER-negative. A large proportion (~ 75%) of BC are known to be ER-positive56. ERα-positive cases are often associated with more optimistic prognosis as they generally respond more positively to endocrine therapies, and are also sensitive to CDK4/6 inhibitors56,57. In contrast, ERα-negative BC is generally regarded as aggressive and metastatic malignancies58.
Given the important role of ERα in BC, its level and structure need to be tightly regulated to ensure an optimal functionality. The level and structure of a protein are known to be influenced by, among others, genetic polymorphisms59. For this reason, many genetic association studies have investigated the relationship between ESR1 polymorphisms and BC susceptibility. These polymorphisms include, but not limited to, rs9340799, rs3020364, rs9322335, rs2234693, rs1801132, rs2046210, rs3020314, rs1514348, rs3020314, rs1514348, rs1514348 and rs302031435,60,61,62,63.
Among these many polymorphisms, we have chosen to focus on rs9340799, an intronic polymorphism located just upstream of exon 2 of ESR1. This is because the rs9340799 polymorphism has been widely studied and conflicting results have been frequently obtained, and no recent meta-analysis has been carried out to address the inconsistencies in the study findings. For instance, while Wang et al. reported that the GG genotype of the polymorphism was associated with a reduced susceptibility to BC, Sierra‑Martínez et al. reported that the same genotype was associated with an increased susceptibility to BC36,45. Besides, Sakoda et al. did not find any significant association between the polymorphism and BC susceptibility34. The difference in the study findings could be attributed to the variations in allele frequency across different studies. These variations are particularly relevant in populations consisting of different ethnicities, as interethnic differences in allele frequencies have long been known64,65. Taking the examples above, while Wang et al.36 noted in a Caucasian population that the minor allele frequency (MAF) of the polymorphism was 0.369, Sakoda et al.34 found that the MAF was merely 0.192 in an Asian population. These variations can account for differences in gene expression and therefore, disease susceptibility66,67. It is thus important to take into account the population variations in the allele frequency when attempting to identify a genetic biomarker for early identification of a disease68. For this reason, heterogeneity tests and subgroup analysis by ethnicity need to be performed when pooling the results from different studies together, as were done in our meta-analysis.
It is noteworthy that most of these studies have centered on genetic association rather than deciphering the exact biological mechanisms. Nonetheless, it has been postulated that intronic polymorphisms such as the rs9340799 polymorphism of ESR1 may influence the cancer susceptibility by (i) being in linkage disequilibrium with another functional polymorphism in the same locus; (ii) influencing the expression of other genes through alterations to their transcriptional activity or mRNA stability; (iii) containing regulatory sequences which can impact gene expression via transcriptional regulation47,69. For these reasons, in this meta-analysis, we attempted to precisely re-examine the relationship between the ESR1 rs9340799 polymorphism and the susceptibility to BC. In doing so, we included 23 case–control studies from 22 systematically selected published articles. We performed the meta-analysis under five different genetic models, namely the homozygous, heterozygous, dominant, recessive, and allele models. Importantly, our analyses with all five genetic models failed to detect any significant association between the rs9340799 polymorphism and BC susceptibility. Under each genetic model, we further stratified our analysis based on the following subgroups: (i) ethnicity (Asian vs. Caucasian), (ii) menopausal status (premenopause vs. postmenopause), and (iii) study quality (high quality vs. low quality). Again, none of these subgroups showed any significant association. Notably, our finding was in agreement with that of the Zhang et al. even though we have included more studies (N = 23 vs. N = 7)29.
The major strength of our study is that we have analyzed data from a large population of meticulously selected studies; therefore, this study has strong statistical power. Besides, the chosen exposure, i.e., the rs9340799 polymorphism, is a discrete and well-defined parameter that can be genotyped with high precision using the available technologies. This allows a fair comparison to be made among independent studies, contributing to more consistent inter-laboratory or inter-study comparison. On the other hand, the major limitation of this study is that gene–gene or gene-environment interactions were not measured as most of the included studies did not report this information. Furthermore, our meta-analysis has so far focused on one polymorphism from ESR1. The analyses of more polymorphisms of ESR1 in future, either individually or in tandem, may further reveal the synergistic effects of such polymorphisms in influencing BC susceptibility70.
In conclusion, our overall results revealed no significant association between the rs9340799 polymorphism of ESR1 and the susceptibility to BC, despite the different genetic models considered. Each genetic model was further divided into subgroups based on ethnicity, study quality and menopausal status, but similarly, no statistically significant association was observed. Nevertheless, our conclusion warrants further studies, given that the ESR1 harbors many polymorphisms that await detailed investigation.
Methods
Literature search
A comprehensive literature search was performed in the Web of Science (WoS), PubMed, Scopus, China National Knowledge Infrastructure (CNKI), VIP and Wanfang databases up to January 21st, 2021, without language restriction. The following search terms were used: (ESR1 OR estrogen receptor) AND (XbaI OR rs9340799) AND (polymorphism or variant) AND (breast cancer OR breast neoplasm). Studies were selected if they fulfilled the following inclusion criteria: (i) were case–control and/or cohort studies which have investigated the association between ESR1 rs9340799 polymorphism and BC susceptibility, and (ii) reported the genotype and allele frequencies or contained necessary data to obtain the information. Studies were excluded if (i) they were not original research papers (e.g. review articles or commentaries), and (ii) the investigations were not performed on human subjects. The reference lists of the eligible studies were also manually screened to identify additional relevant studies. When overlapping data were found, we included only the study with the largest sample size. The study protocol was pre-registered with PROSPERO (registration number: CRD42021231912).
Data extraction and quality assessment
Three investigators independently extracted the following data from the included studies: name of the first author, publication year, location, ethnic group, sample size, genotype and allele frequencies, menopausal status, genotyping method, blinding status, genotyping success rate, and sources of controls. Discrepancies were resolved through discussion until a mutual agreement was reached. The P-values of the Hardy–Weinberg equilibrium (HWE) among the control group was calculated using a goodness-of-fit test. The Modified Newcastle–Ottawa Scale for Case–Control Studies of Genetic Association was used to assess the quality of the included studies71. Studies rated ≥ 6 stars were considered high quality.
Statistical analysis
STATA version 16.0 (StataCorp, College Station, Texas, USA) was used for the quantitative synthesis of the data. The association between ESR1 rs9340799 polymorphism and BC susceptibility was evaluated using the odds ratio (OR) for various genetic models, i.e. homozygous (GG vs. AA), heterozygous (AG vs. AA), dominant (AG + GG vs. AA), recessive (GG vs. AA + AG) and allele (G vs. A). A forest plot was also generated to graphically represent the findings. A fixed-effect model was used if the heterogeneity among the studies was low (Cochran’s Q P-value of > 0.1 and I2 value of < 50%). On the other hand, when heterogeneity was significant, a random-effects model was used. Sensitivity analysis was performed using the leave-one-out method for evaluating the robustness of the findings. Subgroup analyses were performed according to ethnicity (Asian vs. Caucasian), study quality (high quality vs. low quality), and menopausal status (premenopause vs. postmenopause). In most included studies, the ethnicity was explicitly stated, although the standards of classification (i.e. self-reported or via genetic analyses) was not known. However, when such information was not available, the populations were classified into different ethnicities based on the major ethnic group of the countries in which the subjects were recruited. Publication bias was evaluated using the Begg’s and the Egger’s tests, and through visual inspection of the funnel plot for asymmetry. For all analyses, the result was considered to be statistically significant when P < 0.05, unless otherwise stated.
References
Sapkota, Y. Germline DNA variations in breast cancer predisposition and prognosis: A systematic review of the literature. Cytogenet. Genome Res. 144, 77–91 (2014).
Smith, R. A. et al. Cancer screening in the United States, 2019: A review of current American Cancer Society guidelines and current issues in cancer screening. CA. Cancer J. Clin. 69, 184–210 (2019).
Hu, Z. et al. A multigenic study on breast cancer risk associated with genetic polymorphisms of ER Alpha, COMT and CYP19 gene in BRCA1/BRCA2 negative Shanghai women with early onset breast cancer or affected relatives. J. Cancer Res. Clin. Oncol. 133, 969–978 (2007).
Mitrunen, K. & Hirvonen, A. Molecular epidemiology of sporadic breast cancer: The role of polymorphic genes involved in oestrogen biosynthesis and metabolism. Mutat. Res. Rev. Mutat. Res. 544, 9–41 (2003).
Thompson, D. & Easton, D. F. Cancer incidence in BRCA1 mutation carriers. J. Natl. Cancer Inst. 94, 1358–1365 (2002).
Molina-Montes, E. et al. Cumulative risk of second primary contralateral breast cancer in BRCA1/BRCA2 mutation carriers with a first breast cancer: A systematic review and meta-analysis. Breast 23, 721–742 (2014).
Stratton, M. R. & Rahman, N. The emerging landscape of breast cancer susceptibility. Nat. Genet. 40, 17–22 (2008).
Peto, J. et al. Prevalence of BRCA1 and BRCA2 gene mutations in patients with early-onset breast cancer. J. Natl. Cancer Inst. 91, 943–949 (1999).
Antoniou, A. C. et al. A comprehensive model for familial breast cancer incorporating BRCA1, BRCA2 and other genes. Br. J. Cancer 86, 76–83 (2002).
Michailidou, K. et al. Large-scale genotyping identifies 41 new loci associated with breast cancer risk. Nat. Genet. 45, 353–361 (2013).
Tan, S. C. et al. Association between MIR499A rs3746444 polymorphism and breast cancer susceptibility: A meta-analysis. Sci. Rep. 10, 1–10 (2020).
Abdulrahman, G. O. & Rahman, G. A. Epidemiology of breast cancer in Europe and Africa. J. Cancer Epidemiol. 2012, 2 (2012).
Shen, Y., Li, D. K., Wu, J., Zhang, Z. & Gao, E. Joint effects of the CYP1A1 MspI, ERα PvuII, and ERα XbaI polymorphisms on the risk of breast cancer: Results from a population-based case-control study in Shanghai, China. Cancer Epidemiol. Biomarkers Prev. 15, 342–347 (2006).
Kord-Varkaneh, H. et al. Association between healthy eating index-2015 and breast cancer risk: A case-control study. Asian Pacific J. Cancer Prev. 21, 1363–1367 (2020).
Jefcoate, C. R. et al. Tissue-specific synthesis and oxidative metabolism of estrogens. J. Natl. Cancer Inst. Monogr. https://doi.org/10.1093/oxfordjournals.jncimonographs.a024248 (2000).
Zhu, B. T. & Conney, A. H. Functional role of estrogen metabolism in target cells: Review and perspectives. Carcinogenesis 19, 1–27 (1998).
Key, T. J. et al. Endogenous sex hormones and breast cancer in postmenopausal women: Reanalysis of nine prospective studies. J. Natl. Cancer Inst. 94, 606–616 (2002).
Shin, A. et al. Estrogen receptor alpha gene polymorphisms and breast cancer risk. Breast Cancer Res. Treat. 80, 127–131 (2003).
Katzenellenbogen, B. S. et al. Molecular mechanisms of estrogen action: Selective ligands and receptor pharmacology. J. Steroid Biochem. Mol. Biol. 74, 279–285 (2000).
Balfe, P. et al. Estrogen receptor α and β profiling in human breast cancer. Eur. J. Surg. Oncol. 30, 469–474 (2004).
Slattery, M. L. et al. ESR1, AR, body size, and breast cancer risk in Hispanic and non-Hispanic white women living in the Southwestern United States. Breast Cancer Res. Treat. 105, 327–335 (2007).
Dunning, A. M., Healey, C. S., Pharoah, P. D., Teare, M. D. & Ponder, B. A. A systematic review of genetic polymorphisms and breast cancer risk—PubMed. Cancer Epidemiol. Biomark. Prev. 8, 843–854 (1999).
Weiderpass, E. et al. Estrogen receptor α gene polymorphisms and endometrial cancer risk. Carcinogenesis 21, 623–627 (2000).
González-Zuloeta Ladd, A. M. et al. Estrogen receptor alpha polymorphisms and postmenopausal breast cancer risk. Breast Cancer Res. Treat. 107, 415–419 (2008).
Javed, S. et al. Combined effect of menopause age and genotype on occurrence of breast cancer risk in Pakistani population. Maturitas 69, 377–382 (2011).
Onland-Moret, N. C., Van Gils, C. H., Roest, M., Grobbee, D. E. & Peeters, P. H. M. The estrogen receptor a gene and breast cancer risk (The Netherlands). Cancer Causes Control 16, 1195–1202 (2005).
Wedrén, S. et al. Oestrogen receptor alpha gene haplotype and postmenopausal breast cancer risk: a case control study. Breast Cancer Res. 6, R437–R449 (2004).
Zhang, Z. L., Zhang, C. Z., Li, Y., Zhao, Z. H. & Yang, S. E. Association between ERα gene Pvu II polymorphism and breast cancer susceptibility. Medicine 97, 1 (2018).
Zhang, Y. et al. Association between ESR1 PvuII, XbaI, and P325P polymorphisms and breast cancer susceptibility: A meta-analysis. Med. Sci. Monit. 21, 2986–2996 (2015).
Andersen, T. I. et al. Oestrogen receptor (ESR) polymorphisms and breast cancer susceptibility. Hum. Genet. 94, 665–670 (1994).
Kang, H. J. et al. Polymorphisms in the estrogen receptor-alpha gene and breast cancer risk. Cancer Lett. 178, 175–180 (2002).
Lu, H. et al. Estrogen receptor alpha gene polymorphisms and breast cancer risk: A case–control study with meta-analysis combined. Asian Pac. J. Cancer Prev. 14, 6743–6749 (2014).
Ramalhinho, A. C., Marques, J., Fonseca-Moutinho, J. A. & Breitenfeld, L. Genetic polymorphims of estrogen receptor alpha -397 PvuII (T>C) and -351 XbaI (A>G) in a portuguese population: Prevalence and relation with breast cancer susceptibility. Mol. Biol. Rep. 40, 5093–5103 (2013).
Sakoda, L. C. et al. Selected estrogen receptor 1 and androgen receptor gene polymorphisms in relation to risk of breast cancer and fibrocystic breast conditions among Chinese women. Cancer Epidemiol. 35, 48–55 (2011).
Dunning, A. M. et al. Association of ESR1 gene tagging SNPs with breast cancer risk. Hum. Mol. Genet. 18, 1131–1139 (2009).
Wang, J. et al. Estrogen receptor alpha haplotypes and breast cancer risk in older Caucasian women. Breast Cancer Res. Treat. 106, 273–280 (2007).
Shen, Y., Li, D.-K., Wu, J., Zhang, Z. & Gao, E. Joint effects of the CYP1A1 MspI, ERalpha PvuII, and ERalpha XbaI polymorphisms on the risk of breast cancer: Results from a population-based case-control study in Shanghai, China. Cancer Epidemiol. Biomark. Prev. 15, 342–347 (2006).
Lu, X., Li, B., Wei, J. & Hua, B. The XbaI and PvuII gene polymorphisms of the estrogen receptor alpha gene in Chinese women with breast cancer. Zhonghua Wai Ke Za Zhi 43, 290–293 (2005).
Onland-Moret, N. C., van Gils, C. H., Roest, M., Grobbee, D. E. & Peeters, P. H. M. The estrogen receptor alpha gene and breast cancer risk (The Netherlands). Cancer Causes Control 16, 1195–1202 (2005).
Modugno, F. et al. Association of estrogen receptor alpha polymorphisms with breast cancer risk in older Caucasian women. Int. J. Cancer 116, 984–991 (2005).
Cai, Q. et al. Genetic polymorphisms in the estrogen receptor alpha gene and risk of breast cancer: Results from the Shanghai Breast Cancer Study. Cancer Epidemiol. Biomarkers Prev. 12, 853–859 (2003).
Comings, D. E., Gade-Andavolu, R., Cone, L. A., Muhleman, D. & MacMurray, J. P. A multigene test for the risk of sporadic breast carcinoma. Cancer 97, 2160–2170 (2003).
Carrillo-Moreno, D. I. et al. Association of rs2234693 and rs9340799 polymorphisms of ESR1 gene in breast cancer of Mexican population. J. BUON. 24, 1927–1933 (2019).
Dai, Z. et al. Genetic polymorphisms of estrogen receptor genes are associated with breast cancer susceptibility in Chinese women. Cancer Cell Int. 19, 11 (2019).
Sierra-Martinez, M. et al. Predictive polymorphisms for breast cancer in postmenopausal Mexican women. J. Cancer Res. Ther. 14, 640–646 (2018).
Atoum, M. F. & Alzoughool, F. Reduction in breast cancer susceptibility due to XbaI gene polymorphism of alpha estrogen receptor gene in Jordanians. Breast Cancer 9, 45–49 (2017).
Madeira, K. P. et al. Estrogen receptor alpha (ERS1) SNPs c454–397T>C (PvuII) and c454–351A>G (XbaI) are risk biomarkers for breast cancer development. Mol. Biol. Rep. 41, 5459–5466 (2014).
Koš, M., Reid, G., Denger, S. & Gannon, F. Minireview: Genomic organization of the human ERα gene promoter region. Mol. Endocrinol. 15, 2057–2063 (2001).
Hewitt, S. C. & Korach, K. S. Estrogen receptors: New directions in the new millennium. Endocr. Rev. 39, 664–675 (2018).
Whitesell, L. et al. HSP90 empowers evolution of resistance to hormonal therapy in human breast cancer models. Proc. Natl. Acad. Sci. U. S. A. 111, 18297–18302 (2014).
Ikeda, K., Horie-Inoue, K. & Inoue, S. Identification of estrogen-responsive genes based on the DNA binding properties of estrogen receptors using high-throughput sequencing technology. Acta Pharmacol. Sin. 36, 24–31 (2015).
Adlanmerini, M. et al. Mutation of the palmitoylation site of estrogen receptor a in vivo reveals tissue-specific roles for membrane versus nuclear actions. Proc. Natl. Acad. Sci. U. S. A. 111, 2 (2014).
Sun, Q., Liang, Y., Zhang, T., Wang, K. & Yang, X. ER-α36 mediates estrogen-stimulated MAPK/ERK activation and regulates migration, invasion, proliferation in cervical cancer cells. Biochem. Biophys. Res. Commun. 487, 625–632 (2017).
Arnal, J. F. et al. Membrane and nuclear estrogen receptor alpha actions: From tissue specificity to medical implications. Physiol. Rev. 97, 1045–1087 (2017).
Gee, J. M. et al. Epidermal growth factor receptor/HER2/insulin-like growth factor receptor signalling and oestrogen receptor activity in clinical breast cancer. Endocr. Relat. Cancer 12, 2 (2005).
Craig Allred, D., Brown, P. & Medina, D. The origins of estrogen receptor alpha-positive and estrogen receptor alpha-negative human breast cancer. Breast Cancer Res. 6, 240–245 (2004).
Louie, M. C. & Sevigny, M. B. Steroid hormone receptors as prognostic markers in breast cancer. Am. J. Cancer Res. 7, 1617–1636 (2017).
Chen, J. Q. & Russo, J. ERα-negative and triple negative breast cancer: Molecular features and potential therapeutic approaches. Biochim. Biophys. Acta Rev. Cancer 1796, 162–175 (2009).
Tan, S. C. Low penetrance genetic polymorphisms as potential biomarkers for colorectal cancer predisposition. J. Gene Med. 20, e3010 (2018).
Fjeldheim, F. N. et al. Polymorphisms in the estrogen receptor alpha gene (ESR1), daily cycling estrogen and mammographic density phenotypes. BMC Cancer 16, 1–12 (2016).
Hu, X. et al. Association of three single nucleotide polymorphisms of ESR1 with breast cancer susceptibility: A meta-analysis. J. Biomed. Res. 31, 213–225 (2017).
Lipphardt, M. F., Deryal, M., Ong, M. F., Schmidt, W. & Mahlknecht, U. ESR1 single nucleotide polymorphisms predict breast cancer susceptibility in the central European Caucasian population. Int. J. Clin. Exp. Med. 6, 282–288 (2013).
Chauhan, P., Yadav, R., Kaushal, V. & Kadian, L. Evaluation of genetic polymorphism in estrogen receptor? Gene as breast cancer risk. Biomed. Res. 30, 2 (2019).
Jp, I., Ee, N. & Ta, T. ‘Racial’ differences in genetic effects for complex diseases. Nat. Genet. 36, 1312–1318 (2004).
Tan, S. C., Ismail, M. P., Duski, D. R., Othman, N. H. & Ankathil, R. FAS c-671A>G polymorphism and cervical cancer risk: A case–control study and meta-analysis. Cancer Genet. 211, 2 (2017).
Mattei, J. et al. Disparities in allele frequencies and population differentiation for 101 disease-associated single nucleotide polymorphisms between Puerto Ricans and non-Hispanic whites. BMC Genet. 2009(10), 1–12 (2009).
Spielman, R. S. et al. Common genetic variants account for differences in gene expression among ethnic groups. Nat. Genet. 39, 226 (2007).
Goddard, K. A. B., Hopkins, P. J., Hall, J. M. & Witte, J. S. Linkage disequilibrium and allele-frequency distributions for 114 single-nucleotide polymorphisms in five populations. Am. J. Hum. Genet. 66, 216 (2000).
Tan, S. C. & Ankathil, R. Genetic susceptibility to cervical cancer: Role of common polymorphisms in apoptosis-related genes. Tumor Biol. 36, 2 (2015).
Timpson, N. J., Greenwood, C. M. T., Soranzo, N., Lawson, D. J. & Richards, J. B. Genetic architecture: the shape of the genetic contribution to human traits and disease. Nat. Rev. Genet. 19, 110–124 (2018).
Yang, M. et al. The polymorphisms of melatonin receptor 1B gene (MTNR1B) (rs4753426 and rs10830963) and susceptibility to adolescent idiopathic scoliosis: A meta-analysis. J. Orthop. Sci. 20, 593–600 (2015).
Acknowledgements
Research in SCT’s laboratory is supported by the Fundamental Research Grant Scheme of the Ministry of Higher Education, Malaysia (No. FRGS/1/2019/SKK08/UKM/02/9) and the Research University Grant of Universiti Kebangsaan Malaysia (No. GUP-2020-076).
Author information
Authors and Affiliations
Contributions
S.C.T. conceived and designed the study, screened and selected the eligible studies for meta-analysis, collected and extracted the data, appraised the study quality, performed statistical analysis. E.A.M.H. and M.A.K.S. independently screened and selected the studies, and extracted the data for analysis. H.K.-V. independently performed the quantitative data synthesis. T.Y.L. analyzed and interpreted the data and wrote the manuscript. M.A.I. critically revised the manuscript and provided significant input and feedback on the draft manuscript. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tan, S.C., Low, T.Y., Mohamad Hanif, E.A. et al. The rs9340799 polymorphism of the estrogen receptor alpha (ESR1) gene and its association with breast cancer susceptibility. Sci Rep 11, 18619 (2021). https://doi.org/10.1038/s41598-021-97935-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-97935-8
- Springer Nature Limited
This article is cited by
-
The Effect of Estrogen-Related Genetic Variants on the Development of Uterine Leiomyoma: Meta-analysis
Reproductive Sciences (2022)