Abstract
To date, the investigation of genes involved in Al resistance has focused mainly on microarrays and short periods of Al exposure. We investigated genes involved in the global response under Al stress by tracking the expression profile of two inbred popcorn lines with different Al sensitivity during 72 h of Al stress. A total of 1003 differentially expressed genes were identified in the Al-sensitive line, and 1751 were identified in the Al-resistant line, of which 273 were shared in both lines. Genes in the category of “response to abiotic stress” were present in both lines, but there was a higher number in the Al-resistant line. Transcription factors, genes involved in fatty acid biosynthesis, and genes involved in cell wall modifications were also detected. In the Al-resistant line, GST6 was identified as one of the key hub genes by co-expression network analysis, and ABC6 may play a role in the downstream regulation of CASP-like 5. In addition, we suggest a class of SWEET transporters that might be involved in the regulation of vacuolar sugar storage and may serve as mechanisms for Al resistance. The results and conclusions expand our understanding of the complex mechanisms involved in Al toxicity and provide a platform for future functional analyses and genomic studies of Al stress in popcorn.
Similar content being viewed by others
Introduction
Aluminum (Al) is the third most abundant element in the earth’s crust. In acid soils with pH values at or below 5, the phytotoxic species Al3+ is solubilized in soil solution and becomes one of the most important abiotic stresses that limit crop production. Al stress occurs in approximately 30% of the world’s arable soils and in more than 50% of potentially arable land. Of this total, approximately 60% is located in tropical and subtropical regions and negatively impacts the food supply chain. The phytotoxic form Al3+ inhibits root growth, thereby altering water and nutrient absorption and consequently reducing plant development1,2,3.
Plants use multiple strategies against Al stress, and two types of mechanisms have been described: (1) Al exclusion, which prevents the entrance of Al into the root apex, and (2) tolerance mechanisms, where Al enters the plant and is detoxified and sequestered3. The well-characterized exclusion mechanism is dependent on organic acid (OA) exudation from the root apex4,5. Citrate release from the root is an important mechanism against Al stress in maize, which has been identified from the citrate transporter Multidrug and Toxic Compound Extrusion 1 (ZmMATE1)6. However, this mechanism is not well correlated with Al resistance, suggesting that other Al resistance mechanisms are operating in the roots of maize7. Organic acids such as malate, citrate, and oxalate can chelate Al and attenuate Al toxicity3. Al exposure induces malate secretion in wheat8, Arabidopsis9, and rapeseed10; citrate secretion in sorghum11, barley12, rice bean13, rice14, wheat15,16, and common maize17; and oxalate secretion in buckwheat18, spinach19, and tomato20.
Maize is widely grown on acid soils throughout the tropics and subtropics, causing yield losses of up to 69%17,21. In Brazil, maize cultivation has reached around 4156.6 thousand hectares of planted area as of the 2019/2020 harvest22. The cultivation of popcorn (Zea mays var. everta) is expanding with high demand in Brazil and USA, thus attracting the attention of breeders to obtain populations and hybrids that are adapted to Brazilian conditions23. In popcorn, Al toxicity affects root development and causes several types of damage and cell disorganization in the apical region, which compromise plant growth and nutrient uptake24. The land requirements and cultivation aspects of popcorn cultivation are similar to those of common maize, so it is essential to obtain genotypes that are tolerant of acid soils and to identify candidate genes for use in popcorn breeding programs.
To our knowledge, no studies have investigated the transcriptome profile of popcorn under Al stress, and it remains unclear what genes and mechanisms are involved in the transcriptional regulation of popcorn under Al stress. The transcriptional response of common maize roots has been tracked using a microarray approach after 1, 2, 6, and 24 h of Al exposure in a hydroponic system25,26, and in the early developmental stages, the expression patterns of miRNAs were investigated in maize roots under Al stress27. Unlike hydroponic experiments, Mattiello et al.28 characterized the transcriptional profile of maize roots during one and three days of growth in soil containing toxic levels of Al using a microarray.
To study the genetic control of Al tolerance in popcorn, Rahim et al.24 screened 18 inbred popcorn lines and performed relative root growth (RRG), hematoxylin staining, Al content, scanning electron microscopy, and stereoscopic analyses after seven days of stress treatment (160 μM Al3+) to identify inbred lines with Al sensitivity. They classified the 11–60 line as the most Al-sensitive line with the lowest RRG values, the greatest Al accumulation, and intense epidermal degradation in the root tips. They also classified the 11–133 line as an Al-resistant line with the highest RRG value, lowest Al accumulation, and lowest damage to the root apices.
To date, transcriptional expression with RNA sequencing techniques has been used to study Al toxicity tolerance mechanisms in other crops such as sugarcane29, buckwheat30, and tea plants31, allowing the identification of new genes involved in the response to this abiotic stress. The investigation of Al-responsive genes in maize has focused mostly on the early developmental stages and using a microarray approach. We believe that in long-term Al exposure, a robust maintenance mechanism is activated and that several components work at the same time. In this study, we present a high-throughput RNA sequencing method to track the transcriptional response of two contrasting popcorn inbred lines, the Al-resistant 11–133 and Al-tolerant 11–60 lines, to uncover candidate genes related to mechanisms of Al toxicity and tolerance in popcorn.
Results
We generated a range of ~ 38.5–46.5 million clean reads after a quality process for each sample and obtained around 52.87% GC content. An average of 80.10% of the reads were mapped, and from this total, only 6.33% presented multiple alignments with the B73 reference genome using the default parameters (Supplementary Table S1). We performed a Principal Component Analysis (PCA) to compare the differentially expressed genes (DEGs), which revealed that the DEGs from the control and + Al treatments were clustered together for both lines, but the lines were grouped into separated clusters, showing differences in the genetic background between both inbred lines (Supplementary Fig. S1). After 72 h of stress treatment, the inbred lines with different Al sensitivity presented visual phenotypic differences. The Al-sensitive line demonstrated changes in root development and a decreased number of roots (Fig. 1).
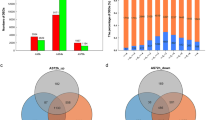
We detected a total of 1003 DEGs in the Al-sensitive line (467 down-regulated and 536 up-regulated) along with 1751 DEGs in the Al resistant line (942 down-regulated and 809 up-regulated) at a False Discovery Rate (FDR) of q < 0.01 (Fig. 2; Supplementary Table S2). The different numbers of DEGs in both inbred lines show that the Al-resistant line has a broader response to Al stress than the Al-sensitive line, regulating multiple pathways against Al damage. Furthermore, we found 273 common DEGs in both inbred lines under Al stress (Fig. 3a; Supplementary Table S3). The functional analysis indicated that the broadest range of these genes is involved in “response to chemicals,” followed by “response to stress” and “biological process” (Fig. 3a). There were 114 DEGs that were up-regulated in the Al-sensitive line and presented down-regulation in the Al resistant line. There were also 96 genes that had the opposite behavior, showing down-regulation in the Al-sensitive line and up-regulation in the Al resistant line (Fig. 3a; Supplementary Table S2).
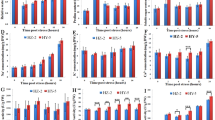
Gene Ontology (GO) enrichment of DEGs. (a) Heatmap of the differentially expressed genes shared in both inbred lines—ST/SC: contrast between Al-sensitive line under treatment/control conditions; TT/TC: contrast between Al-resistant line under treatment/control conditions. (b) Biological process. (c) Molecular function. (d) Cellular component. *The numbers of genes were divided by 10. Heatmap was generated in R version 3.6.2 (https://www.r-project.org/).
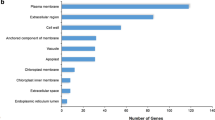
A Gene Ontology (GO) analysis was performed to categorize DEGs into different groups. For the Al-resistant line, the majority of the DEGs were classified as the “response to stress” category, but in the Al-sensitive line was highly enriched in the “biosynthetic process” category. In the biological process category, several terms involved in stress response were much more enriched in the Al-resistant line than in the Al-sensitive line (Fig. 3b). A total of 427 DEGs were clustered in the “response to abiotic stimulus” in the Al-resistant line, which is almost 4 times more than in the Al-sensitive line. This was also observed in the terms “signal transduction” and “cell growth” (Fig. 3b). The terms “protein binding” and “nucleotide binding” in the molecular function category (Fig. 3c) and “plasma membrane”, “nucleus”, and “membrane” in the cellular component category (Fig. 3d) were highly enriched in both inbred lines.
Differentially expressed transporters were also found in both inbred lines (Tables 1 and 2), including Natural resistance-associated macrophage protein (Nramp), aquaporins, Sugars Will Eventually be Exported Transporter (SWEET), Al-activated malate transporter (ALMT), MATE, and ABC transporters. The majority of transporters regulated in both inbred lines were ABC transporters from the G and B families. We detected an increase of regulation of the gene ABC G member 29 (Zm00001d043598) in the Al-resistant line (Table 2), while the same gene presented an opposite regulation profile in the Al-sensitive line (Table 1).
Nramp aluminum transporter 1 (Nrat1, Zm00001d014391) was up-regulated, while Nramp6 (Zm00001d019327) was down-regulated in the Al-sensitive line (Table 1). We also found four genes encoding aquaporin proteins in only the Al-resistant line (Table 2). Two Aquaporin PIP2-2 genes (Zm00001d005410 and Zm00001d014285) were up-regulated, while Aquaporin TIP3-1 (Zm00001d048520) and an Aquaporin PIP-2-2 ortholog (Zm00001d022608) were down-regulated. We also identified seven SWEET transporters, and all of them were up-regulated in both inbred lines except SWEET 6b, which was down-regulated in the Al-resistant line (Table 2).
Genes related to reactive oxygen species (ROS) protection were present in both inbred lines, but there were almost 2 times more in the Al-resistant line than in the Al-sensitive line. We detected up-regulated genes with glutathione S-transferase (GST) activity (Table 3), and the majority were in the Al-resistant line. Additionally, genes playing a role in ROS scavenging, such as peroxidase (POD), catalase (CAT), reductase (RE), and cytochrome P450, were detected in both inbred lines (Supplementary Table S3).
Interactive Pathways Explorer analysis was performed using the KEGG Orthology (KO) database, which revealed that various pathway maps in the Al-resistant line were highly modified under aluminum stress in comparison with the Al-sensitive line (Supplementary Figs. S2 and S3). Curiously, in the Al-resistant line, we detected several genes in the “lipid metabolism” pathway (Supplementary Fig. S2), in contrast with just a few genes in the Al-sensitive line (Supplementary Fig. S3). Several transcription factors (TFs) were detected among our differentially expressed genes. There were 32 down-regulated and 23 up-regulated TFs in the Al-sensitive line, and the Al-resistant line presented 89 down-regulated and 38 up-regulated TFs (Supplementary Table S4). Among these, we found TFs that belonged to the families AP2/EREBP, MYB, bHLH, and WRKY. We also found TFs that are exclusively overexpressed in the Al-resistant line, including ZF-HD, ARF, and E2F/DP families.
From the list of DEGs, nine genes were selected for experimental validation by RT-qPCR. The selected genes are related to abiotic stress: transcription factor HY5-like (HY5), pectin methyltransferase (PME), SWEET 12a, ALMT 10, brassinosteroid catabolism 2 (BRAS), glutathione S-transferase U16 (GST), MYB DNA-binding (MYB), SNF1-related protein kinase regulatory subunit beta-1 (SNF 1), and xyloglucan endotransglucosylase/hydrolase protein 21 (XET). The biological validation was confirmed by RT-qPCR and comparison to RNA-seq data (Supplementary Fig. S4).
Discussion
Aluminum toxicity is one of the main factors limiting crop cultivation in acidic soils. For more than 50 years, breeders have explored genetic diversity to improve Al resistance in several crops, especially in tropical breeding programs. Some critical Al toxicity events are initiated at the transcriptional, biochemical, and physiological levels. To date, several Al-tolerance mechanisms have been described, but much more is needed to uncover the complex response of Al stress. To date, no study has investigated the global differential responses of resistance to aluminum in popcorn plants. In the current study, a high-throughput RNA sequencing approach was used to measure the transcriptome changes in popcorn (Zea mays var. everta) roots under a long period (72 h) of Al exposure.
Al toxicity occurs when Al comes into contact with the cell walls, plasma membranes, and cytoplasm of apical root cells32. In the cell walls, expansins modify the cellulose and non-cellulosic components, thereby loosening and modifying the plant cell walls during growth and adaptation to biotic and abiotic stress33. Recently, the expansin HvEXPA1 was found to be inducible in barley roots under Al stress and to participate in root-cell elongation and regulation of the loosening of the root cell walls34. However, the relationship between expansins (EXP) and Al stress is still poorly understood. In our study, 10 EXPs were detected in the Al-resistant line (Supplementary Table S2). Two EXPA11s (Zm00001d048418 and Zm00001d032883) were up-regulated in the Al-resistant line but down-regulated in the Al-sensitive line, suggesting that these specific EXPs could play an important role in the cell-wall modification under Al stress.
Besides genes involved in cell wall modification, genes encoding membrane transporters are necessary for Al tolerance35. Transporters are some of the most important components in plant Al resistance and play a role in the plasma membrane and tonoplast by participating in exclusion and tolerance mechanisms3. The ABC transporter families found in both inbred lines (Tables 1, 2) suggest that these transporters work together with other detoxifying systems to increase the tolerance response in the Al-resistant line. In the co-expression network, the ABC6 transporter (Zm00001d042953) interacts with the protein Casparian strip membrane domain (CASP)-like 5 (Zm00001d010038) (Fig. 4a).
Unlike the Al-resistant line, the same CASP protein was down-regulated, while the ABC6 was not detected in the Al-sensitive line’s co-expression network (Fig. 4b). CASPs are involved in the formation and location of Casparian strips, nutrient uptake, and stress resistance36. Its expression is increased in cotton during cadmium stress37 and in Tamba black soybean38 during Al stress, thus affecting plant root growth. This interaction suggests the importance of ABC6 in the downstream regulation of CASP-like 5 under Al stress in popcorn roots.
Nrat1 is a specific Al transporter identified in rice that uptakes Al into cells for sequestration to vacuoles and is required for the initial steps of internal Al detoxification39. The best candidate gene as a homolog of OsNrat1, Zm00001d014391, was up-regulated in the Al-sensitive line. Although our Al-sensitive inbred line can respond with multiple mechanisms against the absorption of Al ion in popcorn roots, we propose that this is not sufficient to support this stress due to the metabolic unbalance caused by the Al toxicity conditions. In addition, aquaporins are a group of highly conserved membrane proteins that facilitate water transport across biological membranes40. Our observations suggest that tonoplast aquaporins are likely involved in the Al-tolerance response, but it remains unclear whether Aquaporin PIP proteins play an important role in response to Al stress in popcorn.
Cellular efflux of sugar plays an important role in the maintenance of sugar efflux in phloem loading, nectar secretion, and maternal efflux for filial tissue development41. Plants need to maintain rigid regulation in the storage and transport of vacuolar sugar to deal with adverse environmental conditions42. The role of SWEET in Al-tolerance responses remains unclear, but it plays an important role in adverse conditions, such as salt, osmotic, oxidative, and cold stress, as well as water deficit conditions in Arabidopsis43,44,45. The present work demonstrates a broad up-regulation of SWEET transporters under Al stress (Table 2). Thus, we suggest that these SWEET transporters might play a role in maintaining the tight regulation of vacuolar sugar storage and conferring greater flexibility to adapt to the Al-stress environment.
Membrane transport proteins generally mediate the exudation of organic acids in response to Al stress35. To model the temporal process featuring the response of the intracellular gene expression profile upon Al stress, the Interactive Pathways Explorer was applied to visualize and customize the various pathway maps using the KO database. In the tricarboxylic acid cycle, down-regulated genes were detected in the via from isocitrate, and up-regulated genes were detected in the via from oxaloacetate in the Al-resistant line (Supplementary Fig. S2). The organization of carboxylic acid metabolism in plants is highly dependent on the metabolic and physiological demands of the cell46, and the Al-stress conditions might induce this non-cyclic flux in Al-resistant line. We also observed up-regulated genes involved in the glyoxylate cycle in the Al-resistant line (Supplementary Fig. S2). To our knowledge, no evidence has been presented demonstrating glyoxylate-cycle changes in plants under Al stress, and this hypothesis needs further investigation.
Lipids are the major component of cell membranes, and their composition changes are widely found under various abiotic stresses47,48. Up-regulated genes were involved in lipid metabolism in the Al-resistant line (Supplementary Fig. S2), while these genes were mostly down-regulated or not detected in the Al-sensitive line (Supplementary Fig. S3). Abiotic stress induces changes in the fatty acid composition of plant membrane lipids due to the ability to adjust membrane lipid fluidity by changing the level of unsaturated fatty acids49. In wheat roots, increased expression of lipid transfer proteins enhanced cutin layer thickness in an Al-tolerant near isogenic line under 3 and 7 days of Al exposure, thus protecting root cells from Al damage50. In the Al-resistant line, up-regulated genes are involved in cutin, suberin, and wax biosynthesis, and down-regulated genes are involved in the fatty acid degradation pathway in the Al-resistant line (Supplementary Fig. S2). These results indicate that fatty acids might contribute to the membrane integrity of the Al-resistant inbred line under Al stress.
In several organelles, such as mitochondria, chloroplasts, and peroxisomes, Al stimulates the emergence of ROS, leading to oxidative damage and consequent cell toxicity35,51. Several DEGs involved in the antioxidant system and ROS scavenging were identified in our work (Supplementary Table S3). These data are in agreement with those from Mattiello et al.28, who detected an up-regulation response of several ROS-related genes in maize growing in acid soils. They suggested that this mechanism acts before the oxidative stress occurs. However, they did not detect the induction of superoxide dismutase (SOD) after 72 h of Al exposure, and the same was observed in our study. Testing two contrasting maize lines under different concentrations of aluminum ions, Giannakoula et al.52 showed that the anionic POD isoforms and SOD isoforms increased with increasing Al stress in the tolerant line. In the same way, the CAT enzyme acts as an auxiliary antioxidant that works selectively with either SOD or POD during the peroxidation caused by Al stress as a major enzyme responsible for root growth53.
The GSTs are also involved in Al-toxicity response in maize. Xu et al.25 identified an increase of GST expression in roots, resulting in a strong Al tolerance in maize. In the same way, Cançado et al.54 indicate that a GST may play a role in Al stress alleviation in maize roots. Consistently with these results, GSTs were mostly up-regulated in the Al-resistant line (Table 3) in our RNA-seq data. In the co-expression network analysis, significant patterns of protein–protein interaction were identified between several GSTs, among which GST6 (Zm00001d027541) interacts with a high number of proteins in the Al-resistant line (Fig. 4c). The GSTs also interact with other important components involved in the abiotic stress response, such as PER72 (Zm00001d009373), heat shock protein 20 (HSP20, Zm00001d030346), and plasma membrane-associated protein (PMAP, Zm00001d009932), thus demonstrating a central role of GSTs in the Al-stress response.
Several classes of TFs were identified in our work. Members of MYB, bHLH, AP2/EREBP, WRKY, and NAC families have been identified to be differentially expressed under Al toxicity in different species in previous work30,55. In common maize, a number of TFs were detected in hydroponic25,26 and acid soil experiments28. In addition, miRNA module-involving TFs such as NAC and MYB appear to play a role in the regulation of crown and seminal root development27. TFs belonging to these families were found with different expression regulation in both inbred lines (Supplementary Table S4) and may be involved in the downstream regulation of the expression of genes responsive to Al toxicity.
Finally, we selected a list of candidate genes involved in the global response to Al toxicity in popcorn (Fig. 5), which opens up an opportunity for the development of molecular markers in popcorn breeding programs. The classes of transporters, EXPs, genes related to ROS protection, hormones signaling, pathogenesis-related (PR) proteins, and TFs were significantly involved in the Al response in popcorn roots. These genes increase our understanding of responses to Al toxicity. Beyond that, SWEET transporters involved in Al stress may act in the regulation of vacuolar sugar storage under Al toxicity. The results open new avenues and could further help us to understand the mechanisms of Al toxicity and tolerance that are regulated at the transcriptional level in long-term exposure of popcorn roots.
Heatmap of UPGMA clustering of selected candidate genes differentially expressed in Al-resistant inbred line. SC Al-sensitive control, ST Al-sensitive treatment, TC Al-resistant control, TT Al-resistant treatment. Heatmap was generated in R version 3.6.2 (https://www.r-project.org/).
Methods
Plant materials
Seeds from two contrasting inbred popcorn lines developed by the Popcorn Breeding Program of the Universidade Federal de Viçosa were used in this study: 11–133 (Al-resistant) and 11–60 (Al-sensitive). These genotypes were selected based on a previous study to screen inbred popcorn lines with different Al sensitivity24. The inbred line 11–133 was highly resistant to Al toxicity, presenting statistical significance with the greatest RRG (0.15–0.37), no damage on root apices, lower hematoxylin staining score, and low Al accumulation (926.4 μg/g). The inbred line 11–60 was the most sensitive to Al toxicity, presenting the lowest RRG (0.02–0.06), strong hematoxylin staining, epidermal degradation, and high Al accumulation (1660.3 μg/g).
Growth conditions
First, we treated seeds with fungicide (Captan 400) and germinated them at 25 °C ± 1 °C in a growth chamber for 7 days. Seedlings with uniform growth were picked randomly and transferred to a nutritive solution with constant aeration to acclimate for 24 h. The nutrient solution composition was 1 mM KCl, 1.5 mM NH4NO3, 1 mM CaCl2, 45 µM KH2PO4, 200 µM MgSO4, 500 µM Mg(NO3)2, 155 µM MgCl2, 11.8 µM MnCl2·4H2O, 33 µM H3BO3, 3.06 µM ZnSO4·7H2O, 0.8 µM CuSO4.5H2O, 1.07 µM Na2MoO4·H2O, and 77 µM Fe-EDTA56,57. Then, the treatment group was subjected to aluminum stress with 540 µM of AlCl3 (160 μM Al3+) at pH 4.5 for 72 h. The seedlings were maintained in a growth chamber at 25 °C with a 12/12-h light/dark cycle. Roots from three biological replicates were collected and immediately frozen in liquid nitrogen.
RNA isolation and transcriptome sequencing
RNA from roots was isolated with Trizol LS reagent (Invitrogen, USA) according to the manufacturer’s protocol. The RNA was treated with DNase I Amp Grade (Thermo Scientific) to remove contaminated DNA and then quantified by spectrophotometry (NanoDrop 2000c, Thermo Scientific). The RNA integrity was verified by electrophoresis on 1.6% agarose gel in the presence of ethidium bromide. After quantification, the RNA samples were sent to Macrogen Inc. (Seoul, South Korea), where the libraries were generated using the TruSeq Stranded mRNA kit and sequenced using the Illumina HiSeq 2500 platform.
Read preprocessing and differential expression analysis
Quality control was first performed using the FastQC program (version 0.11.8)58 to check the sequencing quality and identify reads with adapter contamination. Then, the raw reads were trimmed and filtered, and adapters were removed using Trimmomatic (version 0.38)59. The clean reads of all 12 samples were aligned to the maize reference genome (B73 RefGenv4) using Bowtie2 (version 2.3.3.1)60 and TopHat (version 2.1.1)61 with default settings for all parameters. The Cuffdiff (v2.2.1)62 program was used with default parameters to calculate gene expression levels and to identify DEGs in terms of fragments per kilobase per million mapped reads (FPKM). We considered DEGs showing FDR < 0.01 and a log2 fold change value (treated/control) > 1 as up-regulated genes and those with values < − 1 as down-regulated genes. To assess the line groups, PCA was conducted using the stats (version 3.4.4) R-package and plotted in Prism 5 (GraphPad).
Gene function annotation and pathway analysis
Functional enrichment of all DEGs in both lines was conducted using OmicsBox (version 1.2.4) (http://biobam.com/omicsbox). Protein sequences from each DEG were subjected to a similarity search against the UniRef Enriched KEGG Orthology (UEKO) database (http://maxixe.icb.ufmg.br/ueko/) using BLAST (version 2.7.1)63. A script was developed to parse the output and return the KO identifier from each corresponding gene in both lines. The pathway analysis was carried out via Interactive Pathways Explorer (iPath) (version 3) (https://pathways.embl.de/) using the KO identifier. Heatmaps were produced using heatmaply (version 1.0.0)64 R-package.
Co-expression network
The DEGs from each line were used to construct an interaction network. The first-degree of interaction was retrieved from STRING (version 10.5) (https://string-db.org). The minimum required interaction score set was 0.7, and the selected active interaction source was “co‐expression.” The resulting protein–protein interaction network was used as an input for downstream analysis in Cytoscape (version 3.7.1).
Real-time qPCR
RNA from three independent replicates from each treatment was treated with DNase I Amplification Grade (Invitrogen, USA), and the cDNA was synthesized from 2 µg of RNA using the SuperScript Reverse Transcriptase kit (Invitrogen, USA). Real-time qPCR for nine genes identified as differentially expressed in at least one of the inbred lines was performed with an ABI 7500 (Applied Biosystems, USA). The primers were designed using Primer Express software (Applied Biosystems, USA), and the specificity was confirmed by BLAST in the Phytozome database (Supplementary Table S5).
The real-time qPCR reactions were performed using 1 μL of cDNA diluted to 1:10, 5 μL of forward and reverse primers mixed at 1.5 μM (each primer), and 6 μL of SYBR Green PCR Master Mix. The experiment was conducted using three biological replicates for each genotype (different samples from the ones used for the RNA-Seq experiment) and two technical replicates. The maize 18S rRNA was used as an endogenous control: 18S-Fw: GACTACGTCCCTGCCCTTTG and Rev-18S: TCACCGGACCATTCAATCG. The relative expression was estimated using the 2−ΔCt method. The results and the statistical analysis were plotted using GraphPad Prism.
Consent to participate
All authors consented to participate of this research.
Declaration of use of plant material
The popcorn seeds used in this article followed the national standards required by Ministry of Agriculture, Livestock and Supply (MAPA), agency that regulates production, processing, repackaging, storage, analysis or seed trading activities in Brazil, according to Decree Nº. 10.586, of December 18, 2020, which regulates Law Nº. 10.711, of August 5, 2003. We emphasize that none of the seeds were collected for this work, once they belong to the Germplasm Bank of UFV and come from several cycles of interpopulation recurrent selection and more recently have been evaluated for some abiotic stresses by the Popcorn Breeding Program of UFV and that all works have the institution's full consent for its realization.
Data availability
The data sets supporting the results of this article are available in the NCBI SRA repository, http://www.ncbi.nlm.nih.gov/bioproject/508768.
References
von Uexküll, H. R. & Mutert, E. Global extent, development and economic impact of acid soils. Plant Soil 171, 1–15 (1995).
Kochian, L. V. et al. Mechanisms of metal resistance in plants: aluminum and heavy metals. In Progress in Plant Nutrition: Plenary Lectures of the XIV International Plant Nutrition Colloquium (eds. Horst, W. J. et al.) 109–119 (Springer Netherlands, 2002). https://doi.org/10.1007/978-94-017-2789-1_8.
Kochian, L. V., Piñeros, M. A., Liu, J. & Magalhaes, J. V. Plant adaptation to acid soils: The molecular basis for crop aluminum resistance. Annu. Rev. Plant Biol. 66, 571–598 (2015).
Ma, J. F., Ryan, P. R. & Delhaize, E. Aluminium tolerance in plants and the complexing role of organic acids. Trends Plant Sci. 6, 273–278 (2001).
Kochian, L. V., Hoekenga, O. A. & Piñeros, M. A. How do crop plants tolerate acid soils? Mechanisms of aluminum tolerance and phosphorous efficiency. Annu. Rev. Plant Biol. 55, 459–493 (2004).
Maron, L. G. et al. Two functionally distinct members of the MATE (multi-drug and toxic compound extrusion) family of transporters potentially underlie two major aluminum tolerance QTLs in maize. Plant J. 61, 728–740 (2010).
Piñeros, M. A., Shaff, J. E., Manslank, H. S., Carvalho Alves, V. M. & Kochian, L. V. Aluminum resistance in maize cannot be solely explained by root organic acid exudation. A comparative physiological study. Plant Physiol. 137, 231–241 (2005).
Delhaize, E., Ryan, P. R. & Randall, R. J. Aluminum tolerance in wheat (Triticum aestivum L.) II. Aluminum-stimulated excretion of malic acid from root apices. Plant Physiol. 103, 695–702 (1993).
Hoekenga, O. A. et al. AtALMT1, which encodes a malate transporter, is identified as one of several genes critical for aluminum tolerance in Arabidopsis. Proc. Natl. Acad. Sci. U. S. A. 103, 9738–9743 (2006).
Ligaba, A., Katsuhara, M., Ryan, P. R., Shibasaka, M. & Matsumoto, H. The BnALMT1 and BnALMT2 genes from rape encode aluminum-activated malate transporters that enhance the aluminum resistance of plant cells. Plant Physiol. 142, 1294–1303 (2006).
Magalhaes, J. V. et al. A gene in the multidrug and toxic compound extrusion (MATE) family confers aluminum tolerance in sorghum. Nat. Genet. 39, 1156–1161 (2007).
Furukawa, J. et al. An aluminum-activated citrate transporter in barley. Plant Cell Physiol. 48, 1081–1091 (2007).
Yang, X. Y. et al. A de novo synthesis citrate transporter, Vigna umbellata multidrug and toxic compound extrusion, implicates in Al-activated citrate efflux in rice bean (Vigna umbellata) root apex. Plant Cell Environ. 34, 2138–2148 (2011).
Yokosho, K., Yamaji, N. & Ma, J. F. An Al-inducible MATE gene is involved in external detoxification of Al in rice. Plant J. 68, 1061–1069 (2011).
Ryan, P. R., Raman, H., Gupta, S., Horst, W. J. & Delhaize, E. A second mechanism for aluminum resistance in wheat relies on the constitutive efflux of citrate from roots. Plant Physiol. 149, 340–351 (2009).
Tovkach, A. et al. Transposon-mediated alteration of TaMATE1B expression in wheat confers constitutive citrate efflux from root apices. Plant Physiol. 161, 880–892 (2013).
Maron, L. G. et al. Aluminum tolerance in maize is associated with higher MATE1 gene copy number. Proc. Natl. Acad. Sci. U. S. A. 110, 5241–5246 (2013).
Ma, J. F., Zheng, S. J., Matsumoto, H. & Hiradate, S. Detoxifying aluminium with buckwheat [12]. Nature 390, 569–570 (1997).
Jian, L. Y., Shao, J. Z., Yun, F. H. & Matsumoto, H. Aluminium resistance requires resistance to acid stress: A case study with spinach that exudes oxalate rapidly when exposed to Al stress. J. Exp. Bot. 56, 1197–1203 (2005).
Yang, J. L. et al. Aluminum regulates oxalate secretion and plasma membrane H+-ATPase activity independently in tomato roots. Planta 234, 281–291 (2011).
Ngoune Tandzi, L., Mutengwa, C., Ngonkeu, E. & Gracen, V. Breeding maize for tolerance to acidic soils: A review. Agronomy 8, 84 (2018).
CONAB. Acompanhamento da safra brasileira de grãos—safra 2019/20. 4° levantamento. 1–104 (Accessed 25 January 2020); https://www.conab.gov.br/info-agro/safras/graos/boletim-da-safra-de-graos.
Moterle, L. M. et al. Combining ability of popcorn lines for seed quality and agronomic traits. Euphytica 185, 337–347 (2012).
Rahim, F. et al. Identification of contrasting tropical popcorn inbreds for studying aluminum toxicity tolerance inheritance. Euphytica 215, 47 (2019).
Xu, L. et al. Transcriptomic responses to aluminum (Al) stress in maize. J. Integr. Agric. 17, 1946–1958 (2018).
Maron, L. G. et al. Transcriptional profiling of aluminum toxicity and tolerance responses in maize roots. New Phytol. 179, 116–128 (2008).
Kong, X. et al. System analysis of microRNAs in the development and aluminium stress responses of the maize root system. Plant Biotechnol. J. 12, 1108–1121 (2014).
Mattiello, L., Kirst, M., da Silva, F. R., Jorge, R. A. & Menossi, M. Transcriptional profile of maize roots under acid soil growth. BMC Plant Biol. 10, 196 (2010).
Rosa-Santos, T. M. et al. Molecular mechanisms underlying sugarcane response to aluminum stress by RNA-Seq. Int. J. Mol. Sci. 21, 7934 (2020).
Xu, J. M. et al. Transcriptome analysis of al-induced genes in buckwheat (Fagopyrum esculentum Moench) root apex: New insight into al toxicity and resistance mechanisms in an al accumulating species. Front. Plant Sci. 8, 1141 (2017).
Li, Y. et al. An RNA-Seq transcriptome analysis revealing novel insights into aluminum tolerance and accumulation in tea plant. Planta 246, 91–103 (2017).
Horst, W. J., Wang, Y. & Eticha, D. The role of the root apoplast in aluminium-induced inhibition of root elongation and in aluminium resistance of plants: A review. Ann. Bot. 106, 185–197 (2010).
Cosgrove, D. J. Growth of the plant cell wall. Nat. Rev. Mol. Cell Biol. 6, 850–861 (2005).
Liu, W., Feng, X., Chen, Z.-H., Zhang, G. & Wu, F. Transient silencing of an expansin HvEXPA1 inhibits root cell elongation and reduces Al accumulation in root cell wall of Tibetan wild barley. Environ. Exp. Bot. 165, 120–128 (2019).
Chauhan, D. K. et al. Aluminum toxicity and aluminum stress-induced physiological tolerance responses in higher plants. Crit. Rev. Biotechnol. 41, 715–730 (2021).
Roppolo, D. et al. A novel protein family mediates Casparian strip formation in the endodermis. Nature 473, 380–383 (2011).
Chen, H. et al. Analysis of potential strategies for cadmium stress tolerance revealed by transcriptome analysis of upland cotton. Sci. Rep. 9, 86 (2019).
Wei, Y. et al. Plasma membrane proteomic analysis by TMT-PRM provides insight into mechanisms of aluminum resistance in tamba black soybean roots tips. PeerJ 8, e9312 (2020).
Xia, J., Yamaji, N., Kasai, T. & Ma, J. F. Plasma membrane-localized transporter for aluminum in rice. Proc. Natl. Acad. Sci. 107, 18381–18385 (2010).
Jang, J. Y., Kim, D. G., Kim, Y. O., Kim, J. S. & Kang, H. An expression analysis of a gene family encoding plasma membrane aquaporins in response to abiotic stresses in Arabidopsis thaliana. Plant Mol. Biol. 54, 713–725 (2004).
Chen, L. SWEET sugar transporters for phloem transport and pathogen nutrition. New Phytol. 201, 1150–1155 (2014).
Chandran, D. Co-option of developmentally regulated plant SWEET transporters for pathogen nutrition and abiotic stress tolerance. IUBMB Life 67, 461–471 (2015).
Zhou, A., Ma, H., Feng, S., Gong, S. & Wang, J. DsSWEET17, a tonoplast-localized sugar transporter from Dianthus spiculifolius, affects sugar metabolism and confers multiple stress tolerance in Arabidopsis. Int. J. Mol. Sci. 19, 1564 (2018).
Le Hir, R. et al. Disruption of the sugar transporters AtSWEET11 and AtSWEET12 affects vascular development and freezing tolerance in Arabidopsis. Mol. Plant 8, 1687–1690 (2015).
Chen, L.-Q. et al. Sucrose efflux mediated by SWEET proteins as a key step for phloem transport. Science (80-). 335, 207–211 (2012).
Sweetlove, L. J., Beard, K. F. M., Nunes-Nesi, A., Fernie, A. R. & Ratcliffe, R. G. Not just a circle: Flux modes in the plant TCA cycle. Trends Plant Sci. 15, 462–470 (2010).
Maejima, E., Osaki, M., Wagatsuma, T. & Watanabe, T. Contribution of constitutive characteristics of lipids and phenolics in roots of tree species in Myrtales to aluminum tolerance. Physiol. Plant. 160, 11–20 (2017).
Huynh, V.-B., Repellin, A., Zuily-Fodil, Y. & Pham-Thi, A.-T. Aluminum stress response in rice: effects on membrane lipid composition and expression of lipid biosynthesis genes. Physiol. Plant 146, 272–284 (2012).
Upchurch, R. G. Fatty acid unsaturation, mobilization, and regulation in the response of plants to stress. Biotechnol. Lett. 30, 967–977 (2008).
Guo, P. et al. Transcriptional analysis between two wheat near-isogenic lines contrasting in aluminum tolerance under aluminum stress. Mol. Genet. Genom. 277, 1–12 (2007).
Kochian, L. V., Piñeros, M. A. & Hoekenga, O. A. The physiology, genetics and molecular biology of plant aluminum resistance and toxicity. Plant Soil 274, 175–195 (2005).
Giannakoula, A., Moustakas, M., Syros, T. & Yupsanis, T. Aluminum stress induces up-regulation of an efficient antioxidant system in the Al-tolerant maize line but not in the Al-sensitive line. Environ. Exp. Bot. 67, 487–494 (2010).
Wang, L., Fan, X.-W., Pan, J.-L., Huang, Z.-B. & Li, Y.-Z. Physiological characterization of maize tolerance to low dose of aluminum, highlighted by promoted leaf growth. Planta 242, 1391–1403 (2015).
Cançado, G. M. A. et al. Glutathione S-transferase and aluminum toxicity in maize. Funct. Plant Biol. 32, 1045 (2005).
Arenhart, R. A. et al. New insights into aluminum tolerance in rice: The ASR5 protein binds the STAR1 promoter and other aluminum-responsive genes. Mol. Plant 7, 709–721 (2014).
Famoso, A. N. et al. Development of a novel aluminum tolerance phenotyping platform used for comparisons of cereal aluminum tolerance and investigations into rice aluminum tolerance mechanisms. Plant Physiol. 153, 1678–1691 (2010).
Magnavaca, R., Gardner, C. O. & Clark, R. B. Inheritance of aluminum tolerance in maize. In Genetic Aspects of Plant Mineral Nutrition (eds. Gabelman W. H. & Loughman, B. C.) 201–212 (Springer Netherlands, 1987). https://doi.org/10.1007/978-94-009-3581-5_18.
Andrews, S. FastQC: A quality control tool for high throughput sequence data. Available at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (Accessed 13 November 2017) (2010).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Kim, D. et al. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Galili, T., O’Callaghan, A., Sidi, J. & Sievert, C. heatmaply: An R package for creating interactive cluster heatmaps for online publishing. Bioinformatics 34, 1600–1602 (2018).
Acknowledgements
We thank the National Council for Scientific and Technological Development (CNPq), the Brazilian Federal Agency for Support and Evaluation of Graduate Education (Capes; Finance Code 001), and the Foundation for Research Support of Minas Gerais State (Fapemig) for financial support. We are grateful to the Núcleo de Análises de Biomoléculas of the Universidade Federal de Viçosa for providing the facilities for the data analysis.
Author information
Authors and Affiliations
Contributions
J.M.S.V. and M.D.B. conceived the study and supervised the experiments. VBP conducted the experiments. V.B.P., P.G.F., P.M.P.V. and T.A.O.M. performed data analysis. V.B.P. and M.D.B. designed the primers and performed real-time qPCR. V.B.P. wrote the manuscript; J.M.S.V., M.D.B., J.V.M., and P.M.P.V. critically reviewed the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pinto, V.B., Ferreira, P.G., Vidigal, P.M.P. et al. Uncovering the transcriptional response of popcorn (Zea mays L. var. everta) under long-term aluminum toxicity. Sci Rep 11, 19644 (2021). https://doi.org/10.1038/s41598-021-99097-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-99097-z
- Springer Nature Limited
This article is cited by
-
Transcriptome-based strategies for identifying aluminum tolerance genes in popcorn (Zea mays L. var. everta)
Scientific Reports (2023)
-
Tolerance mechanisms to aluminum in popcorn inbred lines involving aluminum compartmentalization and ascorbate–glutathione redox pathway
Planta (2023)
-
Deciphering the major metabolic pathways associated with aluminum tolerance in popcorn roots using label-free quantitative proteomics
Planta (2021)