Abstract
Metabolites in plasma of healthy nursery pigs were quantified using nuclear magnetic resonance. Heritabilities of metabolite concentration were estimated along with their phenotypic and genetic correlations with performance, resilience, and carcass traits in growing pigs exposed to a natural polymicrobial disease challenge. Variance components were estimated by GBLUP. Heritability estimates were low to moderate (0.11 ± 0.08 to 0.19 ± 0.08) for 14 metabolites, moderate to high (0.22 ± 0.09 to 0.39 ± 0.08) for 17 metabolites, and highest for l-glutamic acid (0.41 ± 0.09) and hypoxanthine (0.42 ± 0.08). Phenotypic correlation estimates of plasma metabolites with performance and carcass traits were generally very low. Significant genetic correlation estimates with performance and carcass traits were found for several measures of growth and feed intake. Interestingly the plasma concentration of oxoglutarate was genetically negatively correlated with treatments received across the challenge nursery and finisher (− 0.49 ± 0.28; P < 0.05) and creatinine was positively correlated with mortality in the challenge nursery (0.85 ± 0.76; P < 0.05). These results suggest that some plasma metabolite phenotypes collected from healthy nursery pigs are moderately heritable and genetic correlations with measures of performance and resilience after disease challenge suggest they may be potential genetic indicators of disease resilience.
Similar content being viewed by others
Introduction
Metabolites are small molecules with molecular size < 1.5 kDa and metabolomics is the study of these small molecules to provide a more detailed and comprehensive understanding of how cells function. Metabolites are involved in cellular metabolism, such as energy conversion, growth, signaling, and stimulatory and inhibitory effects on enzymes1,2 and can, therefore, be considered as a bridge between genotype and phenotype. Metabolomics has been used in pharmacology studies, drug development, food technology, toxicology, plant biotechnology, and human health. In the latter case, metabolomics has been used successfully for disease diagnosis and determination of disease state, biomarker discovery, and risk determination3. Metabolite testing is increasingly gaining attention in animal science, including in animal genetics, animal health, and milk and meat quality4. Applications in livestock include discovery of disease biomarkers in dairy cows5,6, animal health7, growth performance8, feed efficiency in beef cattle and pigs9,10, and swine diseases11.
The large impact of infectious disease in swine on animal welfare and productivity has increased interest in selection for pigs that are less susceptible to pathogens and that are more disease resilient, which has been defined as the ability to maintain relatively undiminished performance levels when exposed to disease12,13. It has been proposed that resilience can be an effective approach to select for both disease resistance and tolerance in animals13. Results from studies with dairy cows, beef cattle, pigs including human medicine5,8,9,10, suggest that metabolomics can be useful in helping increase understanding of the processes involved in disease resilience. In addition, metabolomics offers the potential to identify new phenotypes or traits that can be used for selection of resilient animals. There is evidence in the literature that blood metabolites in humans and cattle are heritable14,15,16,17. Given the similarities that exist between porcine and human, blood metabolites in porcine might be expected to be heritable.
Heritability is a prerequisite for the use of a trait in breeding strategies and, ideally, indicators of resilience should be expressed in healthy animals, such that they can be measured in high-health nucleus farms to provide prediction of resilience in commercial environments18. It is therefore important to understand the genetic architecture of these potential new phenotypes.
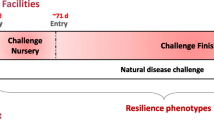
This study is part of a larger project entitled “Phenomics for genetic and genome-enabled improvement of resilience in pigs”, which had identification of predictors of resilience on young healthy pigs as one of its main objectives. The project uses a natural polymicrobial disease challenge model18 with collection of resilience related traits, including average daily gain (ADG), feed intake and feed intake duration (ADFI and ADFD), number of individual health treatments (nTRT) and mortality, residual feed intake (RFI), and feed conversion ratio (FCR). In addition, new resilience traits based on day-to-day variation in feeding patterns were proposed by Putz et al18. The overall purpose of the current study was to use data and plasma samples collected on healthy pigs prior to pathogen/disease challenge to estimate heritability of 44 metabolites and 2 amino acid indices, the phenotypic and genetic parameters of plasma metabolite concentration in relation to their subsequent performance, disease resilience, and carcass traits under the natural disease challenge.
Results
Heritability
Table 1 summarizes estimates of heritabilities and variance components of 44 metabolites measured in plasma of young healthy pigs and two amino acid indexes. Heritability estimates for 13 metabolites were negligible, zero or not estimable. Metabolites with zero or negligible heritability were: glycerol, creatine, l-arginine, 3-hydroxybutyric acid, ethanol, formate, acetoacetate, succinate, acetone, methanol and l-acetylcarnitine. Among the 33 heritable metabolites, 14 showed low estimates of heritability, ranging from 0.11 ± 0.08 to 0.19 ± 0.08, while 17 metabolites had moderately high estimates of heritability, ranging from 0.22 ± 0.09 to 0.42 ± 0.08. Finally, two metabolites, i.e. l-glutamic acid and hypoxanthine, had the highest estimates of heritability, 0.41 ± 0.09 and 0.42 ± 0.08, respectively. Low and moderate estimates of heritability were obtained for two amino acid indexes, i.e. ketogenic amino acids (ketoAA) (0.23 ± 0.08) and branched-chain amino acids (BCAA) (0.19 ± 0.08).
Phenotypic correlations
In general, phenotypic correlations of the metabolites with performance, resilience, and carcass traits were small (Supplementary Figure S1a). However, there were some significant phenotypic correlations. l-Glutamine (0.12 ± 0.04), l-ornithine (0.12 ± 0.04), betaine (0.18 ± 0.04), citric acid (0.21 ± 0.04), l-lysine (0.13 ± 0.04) and l-methionine (0.15 ± 0.04) showed positive phenotypic correlation estimates with quarantine nursery ADG (qNurADG) (P < 0.05), while creatinine (− 0.21 ± 0.04), l-histidine (− 0.20 ± 0.04) and isobutyric acid (− 0.22 ± 0.04) showed negative phenotypic correlation estimates with qNurADG (P < 0.05). In addition, l-glutamic acid showed positive phenotypic correlation estimates with ADFI (0.11 ± 0.04; P < 0.05) and l-aspartate showed positive phenotypic correlation estimates with ADFD (0.18 ± 0.06; P < 0.05). None of the phenotypic correlations of metabolites with challenging nursery ADG (cNurADG), finisher ADG (FinADG), treatment, and mortality rates, and carcass traits were significant (Supplementary Table S1; P > 0.05).
Moreover, we estimated the phenotypic correlations between the metabolites that are involved in the same pathway (glycine, serine, alanine and threonine metabolism) and the results showed that betaine was positively correlated with dimethylglycine and l-serine, and l-serine was positively correlated with l-methionine (Supplementary Table S2). In addition, betaine did have significant positive phenotypic correlations with l-glycine (Supplementary Table S3; P < 0.05).
Genetic correlations
Genotypic correlations of the metabolites with performance, resilience, and carcass traits were larger than phenotypic correlation (Supplementary Figure S1b) however with larger SE. Estimates of genetic correlation between metabolites and ADG in the three phases, qNurADG, cNurADG and FinADG are provided in Table 2. Overall, estimates of genetic correlations among plasma metabolites and ADG in the three phases were very low, with high SE. However, some metabolites showed significant correlation estimates, with the largest negative genetic correlation between plasma creatinine and qNurADG (− 0.60 ± 0.18). Dimethylglycine, betaine, L-methionine and L-serine showed positive genetic correlation estimates with qNurADG. Metabolites that were positively correlated with qNurADG are visualized in a compound network in Fig. 1. The metabolites that were genetically positively correlated with qNurADG are involved in metabolic pathways, including the glycine and serine, methionine and cysteine, glycerophospholipid, and glycosphingolipid pathways. We estimated the genetic correlations between the metabolites that are involved in the same pathway (glycine, serine, alanine and threonine metabolism) and the results showed no significant genetic correlations between these metabolites (Supplementary Table S2).
No significant genetic correlations were estimated between metabolites and cNurADG. Only two metabolites namely: l-glutamic acid and oxoglutarate were estimated to have significant (positive) genetic correlations with FinADG.
Table 3 shows estimates of genetic correlations of metabolites with the feed intake traits ADFI, ADFD, FCR, and RFI. l-Glutamic acid was positively correlated with ADFI (P < 0.001), while several other metabolites, including dimethylglycine and l-aspartate, tended to be positively genetically correlated with ADFI (0.05 < P < 0.1). Four metabolites that were positively correlated with ADFD (Table 3; P < 0.05): dimethylglycine, l-glycine, betaine and citric acid were visualized in a compound network (Fig. 2). These metabolites are involved in metabolic pathways such as the TCA cycle and the glycine and serine, methionine, and cysteine pathways but they were not significantly genetically correlated with each other (Supplementary Table S3).
In addition, the l-glutamine was positively correlated at the genotypic level with FCR (P < 0.05) and isobutyric acid was negatively correlated with RFI (P < 0.05).
Estimates of genetic correlations of metabolites with carcass traits are presented in Table 4. The only significant estimate was of citric acid with carcass back fat (CBF) (P < 0.05).
Finally, only oxoglutarate had a significant negative genetic correlation estimate with the number of treatments across the challenge nursery and finisher (Table 5; P < 0.05), while creatinine had a significant positive genetic correlation with mortality in the challenge nursery (Table 6; P < 0.05).
Discussion
The objectives of this study were to estimate (1) heritabilities of metabolites in plasma of young healthy pigs and (2) their genetic correlations with production, disease resilience, and carcass traits in growing pigs that were exposed to a natural polymicrobial disease challenge. The metabolomics samples were collected from healthy pigs at an average of 26 days of age in the quarantine nursery and the traits analyzed included those in the quarantine nursery, as well as in the challenge nursery and finisher, and carcass traits at slaughter. This study contributes to one of the main overarching objectives of the polymicrobial natural disease challenge model, which is to identify genetic predictors of resilience using samples from healthy pigs. Such predictors would be very useful for breeding for disease resilience, as they can be measured in high health genetic nucleus herds as indicator traits for disease resilience in commercial farms.
We measured the concentration of a panel of 44 metabolites in plasma, including amino acids, short chain fatty acids, sugars, alcohols, organic acids, amines, and TCA cycle intermediates, and urea cycle intermediates. Variation in metabolite concentration can be due to environmental effects, diet, gender, age, physiological conditions, and genetic effects. Literature indicates that in humans, approximately 50% of phenotypic differences in metabolite levels is due to genetics, but heritability estimates differ across metabolite classes16. Metabolites can be grouped into primary and secondary metabolites. Primary metabolites are directly involved in primary metabolic processes, such as normal growth, development, reproduction, and immune response, e.g. amino acids and products derived from glycolysis and the TCA cycle. Primary metabolites are highly conserved across species and serve as precursors for the synthesis of secondary metabolites19. For example, amino acids, in addition to being the building blocks of proteins, are also regulators of innate and adaptive immune responses in living cells. Many studies have demonstrated that glutamic acid, glutamine, histidine, methionine, leucine, isoleucine, and valine are functional regulators of macrophages, dendritic cells, and T-cells.20,21,22,23. Our results showed that 31 metabolites had low to moderate heritability and two metabolites had relatively high heritability (> 0.4). In a study of beef cattle, only 11 of 33 metabolites measured (29 in common with this study) were reported to be heritable17. Similar heritability estimates to Li et al.17 were reported here for betaine, creatinine, pyruvic acid and citric acid. In our study, other metabolites, such as ketone bodies (3-hydroxybutyric acid, acetoacetate and acetone), creatine, succinate, formate, and methylhistidine, showed negligible heritability estimates, suggesting that they are primarily influenced and manipulated by environmental effects such as diet, and/or age, health status etc.
Branched-chain amino acids, was calculated as the sum of l-leucine, l-isoleucine and l-valine and BCAA did not have higher heritability than the amino acids that contributed to the index.
Phenotypic correlations of metabolites with production, disease resilience, and carcass traits were generally very low. These results are in line with those reported previously in dairy cows by Buitenhuis et al.15. However, genetic correlation analyses found that dimethylglycine (0.28), betaine (0.39), l-methionine (0.45), and l-serine (0.54) were positively correlated with ADG in the quarantine nursery, while creatinine (− 0.60) was negatively correlated with this trait. These metabolites are involved in glycine, serine, alanine and threonine pathway, which suggests that this pathway might be a target to improve ADG in young healthy piglets. Indeed, supplementation with dimethylglycine has been shown to improve growth performance, significantly increasing total body weight gain and feed intake, and improving feed efficiency in low-birth-weight piglets24. We found that l-methionine (0.15), l-glutamine (0.12) and betaine (0.18), had positive phenotypic correlation estimates with qNurADG. Methionine supplementation has been reported to improve growth rate in nursery pigs25. Positive effects of methionine supplementation on intestine structure have also been observed, with greater average daily gain from days 7–14 of age and improved feed efficiency25.
Creatinine was genetically highly negatively correlated (− 0.60) with ADG in the quarantine nursery. Variation in creatinine concentration can arise as result of environmental and genetic factors. For example, phenotypic variation in creatinine concentration can be expected due to batch, transportation and fighting or re-grouping of animals. Creatinine is considered a waste product produced by muscles from the breakdown of creatine and is removed from the blood and released into urine by the kidneys. Typically, 95% of creatinine is found in muscle and an increase of creatinine concentration in blood is often considered as an indicator of kidney malfunction27. Conversion of muscle creatine into creatinine can reflect protein degradation28 and serum creatinine concentration has been proposed as an indicator of protein deposition29. Animals with low ADG are expected to have lower protein or muscle deposition and our results suggest that increased plasma creatinine concentration might be a genetic indicator of low ADG in healthy nursery piglets. In addition, creatinine was phenotypically negatively correlated with ADG of young healthy pigs (− 0.21). Moreover, creatinine was genetically positively correlated with mortality in the challenge nursery, suggesting that young healthy pigs that have higher plasma creatinine, genetically have lower ADG as healthy nursery pigs but are more likely to die when challenged by disease. Further research is necessary to validate creatinine as a potential genetic marker for ADG in healthy nursery pigs and as an early genetic indicator trait for mortality under disease.
Plasma oxoglutarate concentration in the quarantine nursery was genetically positively correlated with ADG in the finisher and negatively correlated with nTRT across the nursery and finisher. Moreover, l-glutamic acid showed positive genetic correlation estimates with ADG (0.72) and with ADFI (0.62) in the finisher. Oxoglutarate, also known as alpha-ketoglutarate, is a key organic acid of the TCA cycle and a source of glutamic acid and glutamine, and stimulates protein synthesis and inhibits protein degradation in muscles30,31. Positive effects of oxoglutarate on the protein synthesis and skeletal system have been reported in various farmed species, including turkeys32, pigs33,34, and sheep35,36. Our results suggest that young healthy pigs that had greater oxoglutarate and l-glutamic acid concentration, genetically have greater ADG in the finisher and received fewer health treatments across the challenge nursery and finisher. Interestingly, none of the metabolites were genetically correlated with ADG in the challenge nursery. For the data used here, Cheng et al.37 reported that the estimate of heritability of ADG was lower in the challenge nursery (0.19) than in the quarantine nursery (0.31) and in the finisher (0.30), which might have impacted the ability to accurately estimate the genetic correlations of metabolites with ADG in the challenge nursery.
Four metabolites (dimethylglycine, l-glycine, betaine and citric acid) had moderate to high positive genetic correlation estimates with ADFD (0.30–0.52). These metabolites are involved in the TCA cycle and in glycine, serine, alanine, and threonine metabolism. Interestingly, dimethylglycine and betaine were also positively correlated with ADG in the quarantine nursery, which suggests that pigs with greater dimethylglycine and betaine concentration genetically have greater ADG as young healthy pigs and might spend more time eating in the finisher stage.
A moderate positive genetic correlation (0.34) between citric acid and carcass backfat was estimated, which is in line with the fact that citric acid is involved in fatty acid metabolism. Citrate, the conjugated base of citric acid, is formed from oxaloacetate and acetyl-coenzyme A (acetyl-CoA) by citrate synthase. Once transported to the cytosol, citrate is converted to acetyl CoA and oxaloacetate. Acetyl-CoA is then converted to malonyl CoA and can be used as a substrate for fatty acid synthesis38. The genetic correlation observed suggests that young healthy pigs that have higher plasma content of citric acid genetically have higher carcass backfat when disease is present.
Interestingly, isobutyric acid was estimated to be negatively correlated, genetically, with RFI (− 0.38), which suggests that young healthy pigs that have higher plasma isobutyric acid content genetically have better feed efficiency under disease conditions. Isobutyric acid is a carboxylic or short chain fatty acid (SCFA) that is generated via microbial (gut) metabolism. Isobutyric acid has been described and quantified in the faeces of human, rats, horses, and pigs39. It has been reported that SCFAs such as acetic, propionic, and butyric acids, derive mostly from carbohydrates, while other SCFAs such as isobutyric acid are mostly derived from proteins40, specifically from degradation of branched-chain amino acids, and greater concentration of isobutyric acid could be indicative of better utilization of dietary protein by the microbiota41. Literature indicates that low RFI animals (efficient) have greater ileal isobutyric acid concentrations42. Thus, our results suggest that low RFI pigs genetically might have a better utilization of dietary protein by the microbiota. Further research is necessary to investigate the role of isobutyric acid as a potential genetic biomarker of RFI in pigs with or without disease.
In conclusion, the results suggest that some metabolic phenotypes measured in plasma of young healthy pigs are moderately heritable. The present work contributes to our understanding of the genetic parameters of plasma metabolite concentrations in young healthy pigs and their relationships with production, resilience, and carcass traits under disease. To the best of our knowledge, this is the first study to report estimates of heritabilities of plasma metabolite concentrations and of their genetic correlations with production, resilience, and carcass traits in pigs following polymicrobial disease challenge. Metabolites have the potential to be used in high health genetic nucleus herds as indicator traits for disease resilience in commercial farms. Further studies are warranted to validate the identified possible genetic indicators of resilience.
Material and methods
Animals, production, resilience and slaughter traits
This study was carried out in accordance with the Canadian Council on Animal Care guidelines (CCAC)43. All procedures were carried in accordance with the Animal Research: Reporting of In Vivo Experiments (ARRIVE) guidelines44. The Animal Care protocol was approved by the Animal Protection Committee of the Centre de Recherche en Sciences Animales de Deschambault (15PO283) and Animal Care and Use Committee at the University of Alberta (AUP00002227).
This study is part of a larger research project which investigates the underlying genetic mechanisms of disease resilience in grow-finisher pigs exposed to a natural polymicrobial disease challenge. The details of the polymicrobial challenge and phenotypes/traits collected were previously described in Putz et al.18, Cheng et al.37, and Bai et al.45. The challenge was established in late 2015 at the Centre de développement du porc du Québec inc (CDPQ) test station in Québec, Canada, with the aim to mimic a commercial farm with high disease pressure to maximize expression of genetic differences in resilience. For purpose of clarity, here we briefly describe the three phases of the experiment, which included a pre-challenge quarantine nursery (19 days on average beginning at 3 weeks of age), the challenge nursery (27 days on average), and lastly, the finishing phase (100 days on average). The natural disease challenge protocol was established by introducing naturally infected animals with known diseases into the challenge nursery and finisher barn at CDPQ. Some of the introduced pathogens included: viruses (PRRSV and swine influenza virus A), bacterial pathogens such as Brachyspira hampsonii, Haemophilus parasuis, Mycoplasma hyopneumoniae, Salmonella enterica serovar Typhimurium, and Streptococcus suis), and two parasites (Ascaris suum and Cystoisospora suis)45. Moreover, environmental enrichment (inedible point source objects) was applied in 50% of quarantine nursery pens in some of the batches, with the purpose to evaluate the impact of environmental enrichment on disease resilience. Pigs that received enrichment in the quarantine nursery, continued to receive enrichment in the challenge nursery and finisher37. Throughout the study feed was available ad libitum (the quarantine nursery, challenging nursery and finisher). In the quarantine nursery all pigs were fed the same commercial diet appropriate for pigs’ age and weight (Délice, Nourisson and Premier Age (Cie Alfred Couture ltée; Quebec, Canada)).
Production, resilience, and carcass traits were collected from a total of 3205 F1 crossbred (Landrace × Yorkshire) barrows. The phenotypes used in this study included: average daily gain recorded in the quarantine nursery, challenge nursery, and finisher stages. Finisher traits considered in the present study included feed intake and duration, feed conversion ratio, and residual feed intake. The number of parenteral treatments provided to individual pigs were also tabulated separately for the challenge nursery, finisher and combined challenge nursery and finisher as the number of treatments per 180 days. Mortality (0 = survived, 1 = died) in the challenge nursery, finisher, and across the nursery and finisher were also included37. Finally, carcass traits included: carcass weight, backfat, and loin depth, dressing percentage, and lean yield. Details of the recording and derivation of the resilience, production, and carcass traits can be found in Putz et al.18 and Cheng et al.37. Cheng et al.37 used two data sets for analysis of traits recorded in the finisher but in the current study we only use the data set that included phenotypic finisher data on pigs that survived to slaughter (survivor data), except for mortality. For the challenge nursery, phenotypes on all pigs were used. Details of the number of animals and traits considered here are described in Cheng et al.37. For purpose of brevity: data from 958 young healthy pigs were included in the present study for metabolomics analysis, nearly 3200 animals for nursery traits, around 2500 animals for finisher traits, and 2000 animals for carcass traits.
Blood samples
For metabolomics analysis we used blood samples collected from 958 young healthy animals which were introduced in 15 batches of 60 or 75 pigs. Blood was collected from the jugular vein into K2 ethylenediaminetetraacetic acid (EDTA) tubes (BD Vacutainer, Blood Collection Tubes, United States), on all pigs in the quarantine nursery at an average age of 26 days, 5 days post-arrival from their farm of origin45. After collection, the blood samples were centrifuged at 3000 rpm at 4 °C for 10 min, plasma collected and immediately frozen and stored at − 80 °C and only thawed for the metabolomics analysis. Two weeks after the first sampling, pigs were transferred to the test station and naturally exposed to multiple pathogens, as described in Putz et al.18 and Bai et al.45.
All animals (n = 3205) were genotyped using a 650k Affymetrix Axiom Porcine Genotyping Array by Delta Genomics (Edmonton AB, Canada). Raw Affymetrix SNP data were processed by Delta Genomics, separately for each cycle, with the Axiom Analysis Suite, using all defaults. Details of genotyping and quality control are described in Cheng et al.37 and Bai et al.45. After quality control, a total of 417,443 SNPs in 3205 pigs remained and were used for analysis.
Nuclear magnetic resonance spectroscopy and quality control
Forty-eight plasma metabolites were quantified using NMR, following established protocols at The Metabolomics Innovation Center at University of Alberta (TMIC), AB, Canada (https://www.metabolomicscentre.ca/).
Plasma samples were thawed on ice and a deproteinization step, involving ultra-filtration was performed as previously described46, in order to remove plasma macromolecules. Prior to filtration, 3 kDa cut-off centrifugal filter units (Amicon Microcon YM-3), were rinsed five times each with 0.5 mL of H2O and centrifuged (10,000 rpm for 10 min) to remove residual glycerol bound to the filter membranes. Aliquots of each plasma sample were then transferred into the centrifuge filter devices and spun (10,000 rpm for 20 min) to remove macromolecules (primarily protein and lipoproteins) from the sample. The filtrates were collected and the volumes for each sample were recorded. If the total volume of the sample was under 250 µL an appropriate amount of 150 mM KH2PO4 buffer (pH 7) was added and the dilution factor was annotated and taken into account in the analysis. Subsequently, 46.5 µL of a standard buffer solution (54% D2O:46% 1.75 mM KH2PO4 pH 7.0 v/v containing 5.84 mM DSS (2,2-dimethyl-2-silcepentane-5-sulphonate), 5.84 mM 2-chloropyrimidine-5 carboxylate, and 0.1% NaN3 in H2O) was added to the sample.
The plasma samples (250 µL) were transferred in 3 mm SampleJet NMR tubes for spectral analysis. All 1H-NMR spectra were collected on a 700 MHz Avance III (Bruker) spectrometer equipped with a 5 mm HCN Z-gradient pulsed-field gradient (PFG) cryoprobe. 1H-NMR spectra were acquired at 25 °C using the first transient of the NOESY pre-saturation pulse sequence (noesy1dpr), chosen for its high degree of quantitative accuracy47. All FID’s (free induction decays) were zero-filled to 250 K data points. The singlet produced by the DSS methyl groups was used as an internal standard for chemical shift referencing (set to 0 ppm) and for quantification all 1H-NMR spectra were processed and analyzed using an in-house version of the MAGMET automated analysis software package using a custom metabolite library. MAGMET allows for qualitative and quantitative analysis of an NMR spectrum by automatically fitting spectral signatures from an internal database to the spectrum48. Each spectrum was inspected by an NMR spectroscopist in order to minimize compound misidentification and misquantification.
Prior to statistical analysis, a quality control step was applied to the metabolite data. Four metabolites that were frequently (> 20%) below the limit of detection or with at least 20% missing values were removed from consideration. A total of 44 metabolites and two amino acid indexes remained in the dataset. Other missing values (15 data points) were replaced by the median value of each metabolite in the original data. First, we assessed the significance of each fixed (batch, enrichment), covariable (age), and random effects (pen and litter) using linear regression models implemented in R statistical software49. The residuals of the model were plotted and visually inspected for the presence of outliers, which were excluded from the dataset. Data normalization (log10) of metabolite concentrations that were not normally distributed (2-hydroxybutyrate, ethanol, 3-hydroxybutyric acid, l-alpha-aminobutyric acid, methanol and creatine) was done prior to statistical analysis. Two indexes were also computed for statistical analysis: (1) branched-chain amino acids (BCAA), which was calculated as the sum of l-leucine, l-isoleucine and l-valine and (2) ketogenic amino acids (ketoAA), calculated as the sum of l-lysine and l-leucine. A summary of descriptive statistics of all metabolites after quality control is in Supplementary Table S4.
Variance component analyses
Variance components were estimated by GBLUP using the BLUPF90 programs50. The general following mixed linear model was used to estimate the heritability of each metabolite:
where Yijk is the trait (metabolite); Batchi is the fixed batch effect (i = 1, …, 15); Ageijk is the covariate of age when the pig entered the quarantine nursery; Penj is the random effect of pen by batch corresponding the different phases (quarantine nursery, challenging nursery, or finisher), with Penj ~ N (0, σ2P) where σ2P is pen variance; Litterijk is the litter environmental effect, with Litterijk ~ N (0, σ2L) where σ2L is the litter environmental variance; uijk is the random additive genetic effect, with the vector u ~ N (0, Gσ2A), where G is the genomic relationship matrix and σ2A is the additive genetic variance; and eijk is the residual effect, with eijk ~ N (0, σ2e) where σ2e is the residual variance. The genomic relationship matrix, G, was created separately for each of the seven companies supplying pigs using the software preGSf9050 and the method described by VanRaden51, and then combined into one G matrix, with genetic relationships between companies set to zero in order to focus on within-company variance components, as described by Cheng et al.37. For six metabolites namely: l-ornithine, l-leucine, l-valine, l-asparagine, 3-methyl 2-oxovaleric acid and formate, environment enrichment was included as a fixed effect because the effect of enrichment was significant (P ≤ 0.05).
For metabolites with heritability estimates greater than 0.20, genetic correlations with production, disease resilience, and carcass traits were estimated using bivariate models. For these traits, phenotypes from batches that were not measured for metabolites were also included. Models used for these traits were as described by Cheng et al.37.
Heritability was estimated as σ2a/ (σ2a + σ2L + σ2e) and the standard error (SE) for the heritability estimate was calculated according to the Monte-Carlo method suggested by Meyer and Houle52. The proportion of variance explained by sow or maternal effects, referring to effects that are common to individuals with the same mother, was estimated as σ2L/ (σ2L + σ2e). Genetic correlations between two traits were estimated as the estimate of the genetic covariance from the bivariate analysis divided by the product of the genetic standard deviations for the two traits. Significance of estimates of heritabilities and genetic correlations were determined using likelihood ratio tests with 1 degree of freedom. For heritability estimates, the resulting P-values were divided by 2 because the estimates are restricted to be positive53.
Network visualization
For exploration and visualization of the biochemical pathway that metabolites are involved in, the Metscape plugin54 in Cytoscape 3.8.255 was used. The metabolites that had significant (P ≤ 0.05) genetic correlations with qNurADG and ADFD were used for pathway visualization. The file with the list of KEGG elements was loaded into Metscape to generate a compound network. Only metabolites with KEGG IDs were considered for compound network and pathway analysis. In a compound network, metabolites are represented as nodes and reactions are represented as edges. A compound node with an outgoing edge is a substrate, while a compound with an incoming edge is the product of a specific biochemical reaction. Finally, we estimated phenotypic and genetic correlations among metabolites that belong to the same pathway.
Data availability
The data analyzed in this study are not publicly available, because the data were generated on samples from commercially owned animals, but they can be made available by the corresponding author on reasonable request.
References
Lu, C. & Thompson, C. B. Metabolic regulation of epigenetics. Cell Met. 16, 9–17. https://doi.org/10.1016/j.cmet.2012.06.001 (2012).
Wellen, K. E, Hatzivassiliou, G, Sachdeva, U. M., Bui. T. V., Cross, J. R. & Thompson CB. ATP-citrate lyase links cellular metabolism to histone acetylation. Science. 324, 1076–1080. https://doi.org/10.1126/science.1164097 (2009).
Gomez-Casati, D. F., Zanor, M. I. & Busi, MV. Metabolomics in plants and humans: Applications in the prevention and diagnosis of diseases. BioMed. Res. Int. https://doi.org/10.1155/2013/792527 (2013).
Goldansaz, S. A., Guo, A. C., Sajed, T., Steele, M. A., Plastow G. S. & Wishart, D. S. Livestock metabolomics and the livestock metabolome: A systematic review. PLoS ONE. 12(5). e0177675. https://doi.org/10.1371/journal.pone.0177675 (2017).
Dervishi, E., Zhang, G., Mandal, R., Wishart. D. S. & Ametaj B. N. Targeted metabolomics: New insights into pathobiology of retained placenta in dairy cows and potential risk biomarkers. Animal. 12, 1050–1059. https://doi.org/10.1017/S1751731117002506 (2018).
Dervishi, E., Zhang, G., Hailemariam, D., Mandal, R., Wishart, D. S. & Ametaj, B. N. Urine metabolic fingerprinting can be used to predict the risk of metritis and highlight the pathobiology of the disease in dairy cows. Metabolomics. 14, 83. https://doi.org/10.1007/s11306-018-1379-z (2018).
Montgomery, S. P. et al. Plasma metabolites of receiving heifers and the relationship between apparent bovine respiratory disease, body weight gain, and carcass characteristics. J. Anim. Sci. 87, 328–333. https://doi.org/10.2527/jas.2008-0969 (2009).
Widmann, P. et al. A systems biology approach using metabolomic data reveals genes and pathways interacting to modulate divergent growth in cattle. BMC Genom. 14, 798. https://doi.org/10.1186/1471-2164-14-798 (2013).
Karisa, B. K. et al. Plasma metabolites associated with residual feed intake and other productivity performance traits in beef cattle. Livest. Sci. 165, 200–211. https://doi.org/10.1016/j.livsci.2014.03.002 (2014).
Carmelo, V. A. O., Banerjee, P., da Silva Diniz, W. J. & Kadarmideen, H. N. Metabolomic networks and pathways associated with feed efficiency and related-traits in Duroc and Landrace pigs. Sci. Rep. 10, 255. https://doi.org/10.1038/s41598-019-57182-4 (2020).
Malgarin, C. M., MacPhee, D. J. & Harding, J. C. S. Fetal metabolomic alterations following porcine reproductive and respiratory syndrome virus infection. Front. Mol. Biosci. 7, 430. https://doi.org/10.3389/fmolb.2020.559688 (2020).
Doeschl-Wilson, A. B., Villanueva, B. & Kyriazakis, I. The first step toward genetic selection for host tolerance to infectious pathogens: Obtaining the tolerance phenotype through group estimates. Front. Genet. 3, 265. https://doi.org/10.3389/fgene.2012.00265 (2012).
Mulder, H. A. & Rashidi, H. Selection on resilience improves disease resistance and tolerance to infections. J. Anim. Sci. 95, 3346–3358. https://doi.org/10.2527/jas.2017.1479 (2017).
Kettunen, J. et al. Genome-wide association study identifies multiple loci influencing human serum metabolite levels. Nat. Genet. 44, 269–276. https://doi.org/10.1038/ng.1073 (2012).
Buitenhuis, A. J, Sundekilde, U. K., Poulsen, N. A., Bertram, H. C., Larsen, L. B. & Sørensen, P. Estimation of genetic parameters and detection of quantitative trait loci for metabolites in Danish Holstein milk. J. Dairy Sci. 96, 3285–3295. https://doi.org/10.3168/jds.2012-5914 (2013).
Hagenbeek, F. A. et al. Heritability estimates for 361 blood metabolites across 40 genome-wide association studies. Nat. Commun. 11, 39. https://doi.org/10.1038/s41467-019-13770-6 (2020).
Li, J. et al. Genomic heritability and genome-wide association studies of plasma metabolites in crossbred beef cattle. Front. Gen. 11, 538600. https://doi.org/10.3389/fgene.2020.538600 (2020).
Putz, A. M. et al. Novel resilience phenotypes using feed intake data from a natural disease challenge model in wean-to-finish pigs. Front Genet. 9, 660. https://doi.org/10.3389/fgene.2018.00660 (2019).
Pott, D. M., Osorio, S. & Vallarino, J. G. From central to specialized metabolism: An overview of some secondary compounds derived from the primary metabolism for their role in conferring nutritional and organoleptic characteristics to fruit. Front. Plant Sci. 10, 835. https://doi.org/10.3389/fpls.2019.00835 (2019).
Yoneda, J, Ayatoshi Andou, A. & Takehana, K. Regulatory roles of amino acids in immune response. Curr. Rheumat. Rev. 5, 252–258. https://doi.org/10.2174/157339709790192567 (2009).
McGaha, T. L. et al. Amino acid catabolism: A pivotal regulator of innate and adaptive immunity. Immun. Rev. 249, 135–157. https://doi.org/10.1111/j.1600-065X.2012.01149.x (2012).
Wu, X., Zhang, Y., Liu, Z., Li, T. J. & Yin, Y. L. Effects of oral supplementation with glutamate or combination of glutamate and N-carbamylglutamate on intestinal mucosa morphology and epithelium cell proliferation in weanling piglets. J. Anim. Sci. 90, 337–339. https://doi.org/10.2527/jas.53752 (2012).
Liu, Y. et al. Roles of amino acids in preventing and treating intestinal diseases: Recent studies with pig models. Amino Acids 49, 1277–1291. https://doi.org/10.1007/s00726-017-2450-1 (2017).
Feng, Ch. et al. Effects of dimethylglycine sodium salt supplementation on growth performance, hepatic antioxidant capacity, and mitochondria-related gene expression in weanling piglets born with low birth weight. J. Anim. Sci. 96, 3791–3803. https://doi.org/10.1093/jas/sky233 (2018).
Shen, Y. B., Weaver, A. C. & Kim, S. W. Effect of feed grade l-methionine on growth performance and gut health in nursery pigs compared with conventional DL-methionine. J. Anim. Sci. 92, 5530–5539. https://doi.org/10.2527/jas.2014-7830 (2014).
Chen, Y. et al. l-Methionine supplementation maintains the integrity and barrier function of the small-intestinal mucosa in post-weaning piglets. Amino Acids 46(4), 1131–1142. https://doi.org/10.1007/s00726-014-1675-5 (2014).
Thongprayoon, C, Cheungpasitporn, W. & Kashani, K. Serum creatinine level, a surrogate of muscle mass, predicts mortality in critically ill patients. J. Thoracic Dis. 8, E305–E311. https://doi.org/10.21037/jtd.2016.03.62 (2016).
Hanset, R. & Michaux, C. Characterization of biological types of cattle by the blood levels of creatine and creatinine. J. Anim. Breeding Gen. 103, 227–240 (1986).
Cameron, N., McCullough, E., Troup, K. & Penman, J. Physiological responses to divergent selection for daily food intake or lean growth rate in pigs. J. Anim. Sci. 76, 27–34. https://doi.org/10.1017/S1357729800053285 (2003).
Harrison, A. P. & Pierzynowski, S. Biological effects of 2-oxoglutarate with particular emphasis on the regulation of protein, mineral and lipid absorption/metabolism, muscle performance, kidney function, bone formation and cancerogenesis, all viewed from a healthy ageing perspective state of the art-review article. J. Physiol. Pharmacol. 59, 91–106 (2008).
Wu, N. et al. Alpha-Ketoglutarate: Physiological functions and applications. Biomol. Ther. 24, 1–8. https://doi.org/10.4062/biomolther.2015.078 (2016).
Tatara, M. et al. Effects of alpha-ketoglutarate on bone homeostasis and plasma amino acids in turkeys. Poult Sci. 84, 1604–1609. https://doi.org/10.1093/ps/84.10.1604 (2005).
Kowalik, S., Śliwa, E., Tatara, M. R, Krupski, W, Majcher, P. & Studziński, T. Influence of alpha-ketoglutarate on mineral density and geometrical and mechanical parameters of femora during postnatal life in piglets. Bull. Vet. Inst. Pulawy. 49, 107–111 (2005).
Andersen, N. K., Tatara,M. R., Krupski, W., Majcher, P. & Harrison AP. The long-term effect of alpha-ketoglutarate, given early in postnatal life, on both growth and various bone parameters in pigs. J. Anim. Physiol. Anim. Nutr. 92, 519–528. https://doi.org/10.1111/j.1439-0396.2007.00742.x (2008).
Harrison, A. P, Tygesen, M. P, Sawa-Wojtanowicz, B., Husted, S. & Tatara, M. α-Ketoglutarate treatment early in postnatal life improves bone density in lambs at slaughter. Bone. 35, 204–209. https://doi.org/10.1016/j.bone.2004.03.016 (2004).
Tatara, M. et al. Bone development: The effect of short-term alpha-ketoglutarate administration on long-term mechanical properties of ribs in ram lambs. Small Rum. Res. 67, 179–183. https://doi.org/10.1016/j.smallrumres.2005.09.031 (2007).
Cheng, J. et al. Genetic analysis of disease resilience in wean-to-finish pigs from a natural disease challenge model. J. Anim. Sci. 98, 244. https://doi.org/10.1093/jas/skaa244 (2020).
Akram, M. Citric acid cycle and role of its intermediates in metabolism. Cell. Biochem. Biophys. 68, 475–478. https://doi.org/10.1007/s12013-013-9750-1 (2014).
Cardona, M. E. et al. Correlation between faecal iso-butyric and iso-valeric acids in different species. Microb. Ecol. Health Dis. 17, 177–182. https://doi.org/10.1080/08910600500356265 (2005).
Zarling, E. J. & Ruchim, M. A. Protein origin of the volatile fatty acids isobutyrate and isovalerate in human stool. J. Lab. Clin. Med. 109, 566–570 (1987).
Walsh, A. M., Sweeney, T., Bahar, B., Flynn, B. & O’Doherty, J. V. The effects of supplementing varying molecular weights of chitooligosaccharide on performance, selected microbial populations and nutrient digestibility in the weaned pig. Animal 7, 571–579. https://doi.org/10.1017/S1751731112001759 (2013).
McCormack, U. M. et al. Exploring a possible link between the intestinal microbiota and feed efficiency in pigs. Am. Soc. Microb. 83, e00380-e417. https://doi.org/10.1128/AEM.00380-17 (2017).
Canadian Council on Animal Care guidelines (CCAC). https://www.ccac.ca/en/certification/about-certification.
Animal Research: Reporting of In Vivo Experiments (ARRIVE). https://arriveguidelines.org.
Bai, X. et al. Exploring phenotypes for disease resilience in pigs using complete blood count data from a natural disease challenge model. Front. Genet. 11, 216. https://doi.org/10.3389/fgene.2020.00216 (2020).
Psychogios, N. et al. The human serum metabolome. PLoS ONE 6, e16957. https://doi.org/10.1371/journal.pone.0016957 (2011).
Saude, E. J, Slupksy, C. M. & Sykes, BD. Optimization of NMR analysis of biological fluids for quantitative accuracy. Metabolomics. 2, 113–23. https://doi.org/10.1007/s11306-006-0023-5 (2006).
Ravanbakhsh, S. et al. Accurate, fully-automated NMR spectral profiling for metabolomics. PLoS ONE 10, e0124219. https://doi.org/10.1371/journal.pone.0124219 (2015).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/. (2013).
Misztal, I., Tsuruta S., Strabel T., Auvray B., Druet T. & Lee D. H. BLUPF90 and related programs (BGF90). In Proceedings of the 7th World Congress on Genetics Applied to Livestock Production, London (2002).
VanRaden, P. M. Efficient methods to compute genomic predictions. J. Dairy Sci. 91, 4414–4423. https://doi.org/10.3168/jds.2007-0980 (2008).
Meyer, K. & Houle, D. Sampling based approximation of confidence intervals for functions of genetic covariance matrices. Proc. Assoc. Advmt. Anim. Breed. Genet. 20, 523–526 (2013).
Visscher, P. M. A note on the asymptotic distribution of likelihood ratio tests to test variance components. Twin Res. Hum. Genet 9, 490–495. https://doi.org/10.1375/183242706778024928 (2006).
Karnovsky, A. et al. Metscape 2 bioinformatics tool for the analysis and visualization of metabolomics and gene expression data. Bioinformatics 28(3), 373–380. https://doi.org/10.1093/bioinformatics/btr661 (2012).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Acknowledgements
This project was funded by Genome Canada, Genome Alberta (ALGP2), Genome Prairie, PigGen Canada, and USDA-NIFA Grant No. 2017-67007-26144. The authors acknowledge the members of PigGen Canada for providing the pigs and for helpful discussions, including: Canadian Centre for Swine Improvement, Fast Genetics, Genesus, Hypor, ALPHAGENE, Topigs Norsvin, DNA Genetics, the Canadian Swine Breeders Association, and Alliance Genetics Canada. We wish to thank Xuechun Bai, Yan Meng, Ziqi Yang and Jiehan Lim for their contribution in collecting and preparing the blood samples and The Metabolomics Innovation Center (TMIC) at University of Alberta, AB, Canada for their assistance and service during metabolomics work and Dr. Rupasri Mandal for providing input on the methodological description.
Author information
Authors and Affiliations
Consortia
Contributions
G.P., J.C.M.D., M.K.D, F.F., PGC, and J.C.S.H. conceived the experiment and obtained funding as the main applicants. E.D. T.Y. and J.C. analysed the results and E.D. wrote the first draft of the manuscript. All authors collaborated in the interpretation of results and discussion of the manuscript. All authors have read, reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dervishi, E., Yang, T., Dyck, M.K. et al. Heritability and genetic correlations of plasma metabolites of pigs with production, resilience and carcass traits under natural polymicrobial disease challenge. Sci Rep 11, 20628 (2021). https://doi.org/10.1038/s41598-021-99778-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-99778-9
- Springer Nature Limited
This article is cited by
-
Alginate oligosaccharide supplementation improves boar semen quality under heat stress
Stress Biology (2024)
-
Serum metabolic profile and metabolome genome-wide association study in chicken
Journal of Animal Science and Biotechnology (2023)
-
Breeding for disease resilience: opportunities to manage polymicrobial challenge and improve commercial performance in the pig industry
CABI Agriculture and Bioscience (2022)
-
Relationship between indirect genetic effects for growth, environmental enrichment, coping style and sex with the serum metabolome profile of pigs
Scientific Reports (2021)