Abstract
The patchy distribution of trees typical of savannas often results in a discontinuous distribution of water, nutrient resources, and microbial communities in soil, commonly referred to as “islands of fertility”. We assessed how this phenomenon may affect the establishment and impact of invasive plants, using the invasion of Opuntia stricta in South Africa’s Kruger National Park as case study. We established uninvaded and O. stricta-invaded plots under the most common woody tree species in the study area (Vachellia nilotica subsp. kraussiana and Spirostachys africana) and in open patches with no tree cover. We then compared soil characteristics, diversity and composition of the soil bacterial communities, and germination performance of O. stricta and native trees between soils collected in each of the established plots. We found that the presence of native trees and invasive O. stricta increases soil water content and nutrients, and the abundance and diversity of bacterial communities, and alters soil bacterial composition. Moreover, the percentage and speed of germination of O. stricta were higher in soils conditioned by native trees compared to soils collected from open patches. Finally, while S. africana and V. nilotica trees appear to germinate equally well in invaded and uninvaded soils, O. stricta had lower and slower germination in invaded soils, suggesting the potential release of phytochemicals by O. stricta to avoid intraspecific competition. These results suggest that the presence of any tree or shrub in savanna ecosystems, regardless of origin (i.e. native or alien), can create favourable conditions for the establishment and growth of other plants.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
The past few centuries have seen humans increasingly move plant species around the globe1,2. Some of these species have become invasive and pose significant threats to biodiversity by altering the composition and function of invaded ecosystems3,4,5. Despite achievements towards generalization and formulating regional and global strategies to deal with invasive species, it remains important to understand which factors are driving the success and impact of particular invasions in order to prioritize management actions and mitigate the impacts of invasive plants. This is especially urgent in protected areas6,7. However, this is no easy task, since invasion success and invader impact are often highly context-dependent, and can differ among taxa, the pathways through which species are introduced or disseminated, or the characteristics of recipient ecosystems (e.g., habitat type, altitude, nutrient availability, abundance of mutualists or natural enemies, disturbance frequency or socioeconomic characteristics and processes)8,9.
Savannas constitute the dominant vegetation type in Africa, covering about 60% of the continent10. Until recently, African savannas were thought to be less invaded by alien plants compared to other ecosystems11. Several reasons have been proposed for this, such as low rate of alien plant introductions in some regions, the adaptation of African savannas to fire (which might prevent the establishment of many alien plants), or the resistance of native plants to disturbance12. However, this trend is changing: in the last decades, alien plants (e.g., eucalypts, parthenium weed, or cacti) have been increasingly introduced, becoming invasive and threatening the conservation of savanna ecosystems in Africa, for example, by displacing native vegetation or modifying the characteristics of invaded areas12,13,14. Therefore, research is needed to understand the factors driving plant invasions in these ecosystems.
Savannas are seasonal ecosystems characterized by the presence of grasses and patchily distributed trees and/or shrubs10,15. In these ecosystems, woody species facilitate their own establishment and growth, as well as that of other plant species. Due to their extensive root systems, trees and shrubs extract moisture and nutrients from their surroundings, accumulate litter and exude nutrients and other phytochemicals into surrounding soils. This leads to an accumulation of soil nutrients, increased humidity, and modification of microbial communities under their canopies16. The patchy distribution of trees in savannas can therefore lead to the creation of so-called “fertility islands”17,18,19, with favorable conditions for the establishment and growth of other native plants20. Therefore, it is feasible that fertility islands may also aid the establishment, growth, and impact of alien plants in savanna ecosystems. Moreover, dense populations of invasive plants may themselves create fertility islands by dramatically modifying soil biotic and/or abiotic conditions21,22. These fertility islands may have positive or negative impacts on the establishment and growth of other plants23.
Kruger National Park (KNP) is South Africa’s flagship protected area, covering about 2 million ha of savanna. Situated in north-eastern South Africa along the border of Mozambique, KNP is one of the largest protected areas in the world24. Plant invasions in KNP are some of the most studied in African savannas, especially from a protected areas perspective12. Although 363 alien plant (including non-invasive) species have been recorded in KNP, Opuntia stricta is one of only a few species currently considered a high-impact transformer species in the park (others include, for example, Lantana camara and Parthenium hysterophorus)25,26. Therefore, the invasion of O. stricta in KNP provides a good opportunity to assess potential drivers and impacts of plant invasions in African savannas27.
Opuntia stricta was first introduced to Skukuza village in KNP as an ornamental plant in 1953 and soon became invasive, covering 30,000 ha by 199828. The invasion of O. stricta in KNP has been managed since 1985. An integrative management program, including chemical and biological control considerably reduced its density in the park (see Hoffmann et al.29). However, O. stricta is still expanding its range in both KNP30 and southern Africa31. In order to inform calls for management and understand its impacts, it is important to understand the biotic and/or abiotic factors driving, and responding to, O. stricta invasion.
Several hypotheses have been explored to explain the invasion success of O. stricta in KNP9. For example, Foxcroft et al.32 studied the influence of dispersal pathways, fire frequency, management history, soil type, substrate, vegetation type, and water regimes in the reproduction and survival of this species. However, besides the positive effect of the presence of the leguminous tree Vachellia nilotica subsp. kraussiana (hereafter Vachellia nilotica), none of the studied factors appeared to influence the invasion of O. stricta in the park32. Foxcroft and Rejmánek33 also tested the hypothesis that baboons (Papio ursinus) are major dispersal agents of O. stricta in KNP. Opuntia stricta, like many other Opuntia species, propagates both sexually, by producing large numbers of fleshy fruits and seeds34), and vegetatively, through plant fragments that root rapidly when in contact with soil35. Baboons eat the fruits and disperse both the seeds and fragments of O. stricta in KNP28. Foxcroft and Rejmánek33 therefore hypothesized that O. stricta plants would be more frequently found under trees structurally suitable for baboon roosting (e.g., Spirostachys africana) than under those that are not (e.g., V. nilotica), or on bare patches. However, they found O. stricta patches significantly more frequently under V. nilotica trees. Therefore, the landscape-scale factors driving O. stricta invasions in KNP remain unclear, and the potential influence of V. nilotica trees on the distribution of O. stricta requires further investigation.
The impact of O. stricta in KNP also deserves further study. Robertson et al.36 showed that, although the species does not affect native spider richness, diversity or community structure, it has an impact on native beetle assemblages in KNP. However, the impact of O. stricta on soil characteristics, soil bacterial communities, and the establishment of native plants, remains unknown.
Here, we compare soil characteristics (pH, humidity and nutrients), enzymatic activities, and the diversity and composition of bacterial communities between soils invaded by O. stricta and uninvaded soils, collected under the canopy of two native trees (V. nilotica and S. africana) and in open patches with no tree cover. With these data we aim to test whether (1) native trees and invasive O. stricta create fertility islands and, if so, whether (2) fertility islands influence the establishment of O. stricta. We also wanted to know (3) whether O. stricta modifies soils characteristics and soil bacterial communities in a different manner than native trees and (4) the effects of invasive O. stricta on soils affect the establishment of native trees. We hypothesise that native trees, but not O. stricta, will create fertility islands by increasing soil nutrient loads, humidity, soil enzyme activities and bacterial diversity and inducing changes in soil bacterial community composition. We also hypothesise that these fertility islands lead to enhanced performance of native trees and invasive O. stricta.
Results
Soil pH, humidity, nutrients and enzymatic activities
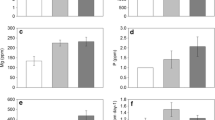
We found significant effects (p < 0.05) of invasion status (invaded, uninvaded) and/or tree cover (V. nilotica, S. africana, none) on soil humidity, organic matter, nitrogen, phosphorus, and phosphatase, β-glucosidase and urease activities (Table 1). Humidity, nutrients and phosphatase activity were higher under V. nilotica and S. africana canopies than under no canopy, and in invaded than in uninvaded areas. Moreover, in uninvaded areas, we generally found higher levels of nutrients and phosphatase activity under V. nilotica than under S. africana or in open patches with no trees. There were lower levels of ß-glucosidase and urease activities under the canopies of V. nilotica than under the canopies of S. africana or in open patches, regardless of invasion.
Soil bacterial community diversity and composition
We obtained 884,540 high-quality sequence reads after data cleaning, which resulted in 18,717 OTUs (97% similarity cut-off) representing 325 genera, 141 families, 75 orders, 40 classes, and 17 phyla. The following numbers of sequences remained unclassified: Genus level—43.9% of all sequences (388,210); Family level—28% (247,259); Order level—14.4% (127,477); Class level—3.7% (32,356); and Phylum level—3.2% (27,891) (Supplementary material). The most abundant phyla were Actinobacteria (50.8%), followed by Proteobacteria (32.9%) and Firmicutes (6.6%). At class level Actinobacteria (50.7%) were most abundant, followed by Alphaproteobacteria (27.7%), Bacilli (6.5%), Betaproteobacteria (2.9%), and Deltaproteobacteria (2.2%).
A total of 12 704 OTUs (861 232 reads, 97.4% of total read count) were shared between invaded and uninvaded soils (Fig. 1), whereas 3815 OTUs (16 172 reads, 1.8% of total read count) were unique to invaded soils and 2198 OTUs (7 136 reads, 0.8% of total sequence read count) to uninvaded soils. For tree cover, a core total of 6002 OTUs was shared by all soils (Fig. 1), and also made up the bulk of the sequence reads (834,286 reads, 94.3% of total read count). However, there were OTUs that were unique to each tree cover type: 720 OTUs (1839 reads, 0.21% of total read count) were unique to open areas, 1043 OTUs (2912 reads, 0.33% of total read count) were unique to V. nilotica soils, and 1054 OTUs (3439 reads, 0.39% of total read count) were unique to S. africana soils.
Distribution of OTUs (operational taxonomic units) between invasion (invaded and uninvaded; top left) and tree cover (V. nilotica, S. africana, none; top right) categories, together with relative abundances of soil bacterial taxa (Class | Phylum). Class-level relative abundances were calculated using the number of sequences for each taxon as a percentage of the total sequences for each invasion/cover combination. The “Other” category includes taxa that were unclassified at Class level and classes representing less than 0.5% of the total number of sequences.
Two-way ANOVAs indicated that the presence of native trees increased the diversity of bacterial communities, especially in uninvaded soils. Furthermore, all diversity metrics were significantly affected by invasion status, being higher in invaded than in uninvaded soils (Table 2, Fig. 2). The presence of V. nilotica, S. africana, and O. stricta also significantly altered soil bacterial community structure and composition based on Horn distances (Table 3, Fig. 3).
Non-metric Multidimensional Scaling (NMDS) plot for soil bacterial communities for various invasion (Invaded, Uninvaded) and tree cover (V. nilotica, S. africana, none) categories. PERMANOVA results confirmed the significant effects of both cover and invasion status on bacterial community composition. The relatively low stress coefficient indicates good ordination with no risk of drawing false inferences89.
Analysis of multivariate homogeneity of group dispersions indicated that uninvaded soils were significantly more over dispersed (BETADISPER F1,16 = 8.76, p = 0.009), suggesting that invasive O. stricta plants have a homogenizing effect on soil bacterial communities, i.e. areas invaded by O. stricta had more similar bacterial communities than uninvaded areas. The effect of tree cover did not, however, have a significant homogenizing effect (BETADISPER F2,15 = 0.15, p = 0.865).
Our linear discriminant analysis effect size (LEfSe) highlighted numerous OTUs (n = 94) that were significantly more abundant in invaded areas, irrespective of cover type (Fig. 4). This indicates that O. stricta invasion leads to a significant increase in the abundance of these OTUs. On the other hand, only three OTUs (two belonging to Rubrobacter and one to Blastococcus) were significantly more abundant in uninvaded soils.
Bacterial taxa that are significantly differentially abundant between status (invaded, uninvaded) and tree cover (V. nilotica, S. africana, none). Shown on the left is a heatmap of the taxa with Ln(Abundance) values, and on the right the LEfSe LDA scores for each taxon88. Only significant LDA scores are shown. The red and blue colours associated with the LDA scores indicate to which categories the taxa were associated with. Note that a small constant (0.01) was added to heatmap abundance values to avoid the undefined natural logarithm of 0.
Germination
The percentage and speed of germination of O. stricta were generally higher in soils collected under V. nilotica canopies and in soils from uninvaded areas (Table 4). The percentage and speed of germination of V. nilotica were significantly lower under uninvaded soils collected from underneath its own canopy. We found no significant differences in the percentage and speed of germination of S. africana.
Discussion
Woody species in savannas accumulate moisture and soil nutrients, and harbor unique bacterial communities under their canopies, creating “fertility islands”16. Accordingly, and in support of our hypothesis, our results show higher soil humidity, nutrient contents, and phosphatase activity under the canopies of V. nilotica and S. africana trees in KNP compared to open patches with no tree cover. Interestingly, the same soil physicochemical properties were also elevated as a result of O. stricta invasion. These results are surprising, since previous studies have shown that cacti in arid ecosystems generally provide little shade and limited amounts of aboveground litter, and therefore, that soils beneath them have similar humidity and nutrient levels than those in bare patches37,38.
Differences in morphology, physiology, or root symbiosis of woody species can cause differences in the characteristics of the fertility islands they create39. Accordingly, we found higher concentrations of soil nutrients and phosphatase activity under V. nilotica canopies than under S. africana canopies. This can be explained by the fact that acacias have rapid growth rates and the capacity to fix atmospheric nitrogen via rhizobium symbioses, and legumes in general have been repeatedly shown to induce changes in soil nutrient levels and cycles40,41,42,43,44. Moreover, V. nilotica has previously been found to increase soil nutrient concentrations due to high above- and belowground organic matter input45. In particular, the presence of V. nilotica trees in our uninvaded areas caused a significant increase in soil nitrogen content, which is a particularly limiting macronutrient in the study area10.
It has also been suggested that, due to leaf litter input, V. nilotica trees increase nutrient cycling45. However, we found significantly lower levels of β-glucosidase and urease activities under V. nilotica canopies. These results are unexpected, and more research is needed to unravel the mechanisms underlying these observations.
In agreement with our findings, previous studies have reported the phyla Actinobacteria and Firmicutes as abundant bacterial groups in semi-arid savannas46. Actinobacteria taxa are generally drought and heat resistant47, while many Firmicutes are spore-forming, an adaptation to harsh and unpredictable environmental conditions48.
Our results showed that the presence of V. nilotica, S. africana, and O. stricta altered soil bacterial composition and increased relative bacterial abundance and diversity compared to open patches with no tree cover. This likely reflects the increases in soil humidity and nutrient contents we observed under these plants49,50,51. However, increases in bacterial abundance and diversity were more pronounced in O. stricta-invaded soils and O. stricta invasion had a significant homogenizing effect on soil bacterial communities. These results suggest that fertility islands created by O. stricta might have stronger effects on the diversity and composition of bacterial communities of savanna soils than those created by the native woody species.
Seeds have various mechanisms to detect suitable conditions for establishment and to adjust their timing of germination accordingly52,53. Our results suggest that the germination of O. stricta in the study area might be facilitated by the presence of V. nilotica trees (also see32,33). Such enhanced germination kinetics may be the result of the higher soil nutrient levels present under the canopy of these trees compared with those under the canopy of S. africana or in open patches with no tree cover, which might trigger the germination of O. stricta seeds.
Timing of germination can also be adjusted as a response to chemicals released by conspecific or other plant species54. Such mechanisms can help plants to avoid intra- or inter-specific competition or to detect the presence of facilitating or nursing species, maximizing establishment potential55. Accordingly, while V. nilotica and S. africana trees appear to germinate equally well in invaded and uninvaded soils, we found O. stricta to have reduced germination performance (lower and slower) in invaded soils. These results suggest the potential release of phytochemicals by O. stricta, which may retard its own germination, i.e. the creation of negative soil-feedbacks, aiming to avoid potential intraspecific competition. Similarly, we also found V. nilotica to have lower germination performance in uninvaded soils collected under its own canopy. These results are not surprising, since previous studies have found different species in the genus Acacia (in which V. nilotica previously resided) to adjust their timing of germination as a response to chemically-induced signals released by adult plants56. Moreover, V. nilotica is known to produce several chemicals, including tannins, flavonoids, and phenolic acids, capable of stimulating or inhibiting seed germination57.
Overall, our results showed that native tree cover and invasive species can create fertility islands in savanna ecosystems, causing changes to the abiotic and biotic conditions of soils. This suggests that the presence of any tree or shrub in savanna ecosystems, regardless of origin (i.e. native or alien) or growth from (e.g. woody tree or succulent shrub), can result in the formation of fertility islands, which usually create favourable conditions for the establishment and growth of other plants20. We also found the presence of V. nilotica trees to be linked to higher increases in soil nutrient contents than the presence of S. africana trees and that these increases might favour the germination of invasive O. stricta seeds. These results provide a mechanistic basis to previous studies suggesting that establishment of O. stricta in KNP might be facilitated by the microenvironment created by V. nilotica trees32. Finally, invasive O. stricta did not affect the germination of native trees, but affected soil bacterial communities more strongly than native trees. Since the eradication of O. stricta in KNP is no longer feasible30, the consequences of these effects deserve further investigation.
Methods
All the local and national guidelines were followed in this study. A contract between the researchers and SANParks (permit NOVOA1292) provided all formal permission needed to conduct the research (which fulfils the NEMBA: Protected Areas Act).
Study site
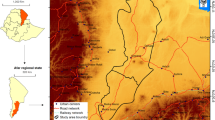
Our study site was located in the south of KNP (Fig. 5), approximately 1 km south-west of Skukuza Rest Camp (− 25.0049, 31.5852), in a typical savanna vegetation with sparse grass cover, open patches with no tree cover, and a patchy distribution of trees (Fig. 6). Two woody species, V. nilotica and S. africana, make up more than 90% of all trees in the study site, and their individuals are mixed in the landscape. The study site has low grass cover, minimising herbivory by native ungulates and due to the open patchy landscape fire in the immediate area is minimal. Moreover, the study site presents granitic geological substrate and thus has nutrient-poor soils58, suggesting a high importance of fertility islands for the establishment of plants in the area.
Study species
Vachellia nilotica (previously Acacia nilotica59), is a semi-deciduous tree in the Fabaceae (legume) family. It can grow up to 10 m in height and its branches are armed with straight, paired spines. Vachellia nilotica is widely distributed in southern Africa, from Tanzania to South Africa60. Spirostachys africana is a spineless deciduous tree from the Euphorbiaceae (spurge) family. It can grow up to 18 m in height, although it usually only reaches about 10 m. Spirostachys africana is also widely distributed in southern Africa61.
Opuntia stricta is a perennial, succulent, shrubby plant in the Cactaceae (cactus) family and is native to Cuba, Mexico, and the USA34. Commonly introduced around the world as an ornamental plant, O. stricta is currently considered invasive in 21 countries31. In its invasive ranges, O. stricta causes multiple ecological and socioeconomic impacts. It reduces agricultural production and biodiversity, causes loss of grazing potential, transforms habitats, and causes injuries to animals and people via its spines62.
Soil and seed collection
We collected samples of O. stricta-invaded and uninvaded soils under the canopies of V. nilotica and S. africana and in open patches with no tree cover. The identification of the plant species was undertaken by AN, LCF and JJLR. No voucher specimens were collected or deposited in a publicly available herbarium. In each of these six soil types, we randomly established five plots of 0.5 × 0.5 m. We did not establish more than one plot under the canopy of a single tree. In each plot, we took five subsamples from the top 10 cm of soil using a shovel. The subsamples within each plot were then sieved through a 2.0 mm mesh and homogenized into a single sample, resulting in 30 samples in total (2 invasion status [invaded, uninvaded] × 3 tree covers [V. nilotica, S. africana, none] × 5 plots × 1 sample per plot). All equipment was cleaned and sterilized with ethanol between collections to avoid cross contamination. For each soil sample we measured (1) pH, humidity, and nutrients (2) soil enzymatic activities, and (3) germination of O. stricta, V. nilotica and S. africana. Moreover, we randomly selected 3 samples per treatment, resulting in 18 samples in total (2 invasion status [invaded, uninvaded] × 3 tree covers [V. nilotica, S. africana, none] × 3 plots × 1 sample per plot), to measure the diversity and composition of soil bacterial communities. Soils used for pH, humidity, nutrients, and germination analyses were kept at room temperature. Soils used for enzymatic activity analysis were refrigerated at 4 °C and analysed within 3 days of collection. Soils used for DNA extraction (i.e. for next-generation sequencing analyses of bacterial communities) were kept on ice during transport and stored at −80 °C as soon as was possible.
Opuntia stricta, V. nilotica and S. africana seeds were also collected in the study area and stored in the dark at 4 °C. O. stricta is declared as invasive in the park, and therefore no permission was needed to collect its seeds. Seeds of the native trees were collected through the Skukuza plant nursery, which has permission to collect and store such seeds, as well as germinate and sell them.
Soil pH, humidity, nutrients and enzymatic activities analysis
Soil pH was determined by dilution with water (1:2.5; soil: distilled water)63, using a CRISON GLP 22 + pH & Ion-Meter. To determine soil humidity, we first weighed the fresh soil samples, and then dried (70 °C for 48 h) and reweighed them, calculating soil humidity as: (Fresh soil weight—Dry soil weight)/(Fresh soil weight) × 100. Soil nutrients (nitrogen, organic matter, and phosphorous) were measured by Labserve Laboratories (Nelspruit, South Africa).
We also analysed three enzymes that play key roles in soil nutrient cycling: β-glucosidase (E.C. 3.2.1.21), involved in carbon metabolism through the release of glucose from cellulose; urease (E.C 3.5.1.5), involved in the release of nitrogen by degrading urea to ammonium; and phosphatase (E.C. 3.1.3.1), involved in the release of phosphate from organic matter by hydrolyzing phosphate ester bonds52. We used the methods described by Tabatabai and Bremner64, Kandeler and Gerber65 and Allison and Vitousek66, for the β-glucosidase, urease, and phosphatase assays, respectively. Following the recommendation of German et al.67, we conducted the enzyme assays at environmental pH conditions and within 48 h of soil collection.
DNA extraction and next generation sequencing
We extracted whole genomic DNA from 0.25 g of soil using the PowerSoil® DNA extraction kit (MO BIO laboratories Inc., Carlsbad, CA, USA), following the manufacturer's protocol. We assessed DNA quality using the NanoDrop ND-1000 UV–Vis Spectrophotometer (Nanodrop Technologies, Wilmington, DE, USA). Part of the 16S rRNA gene (consisting of nine hypervariable regions: V1–V9) was targeted for amplification, since it is frequently used for the identification of bacterial taxa68,69. The more variable regions are useful for genus- or species-level identifications70. We targeted the V5-V7 hypervariable regions using primers 799F (5'-AAC MGG ATT AGA TAC CCK G-3') and 1391R (5'-GAC GGG CGG TGW GTR CA-3'). These primers are known for low non-specificity, and can accurately and reproducibly differentiate species71,72,73. Amplification was done with sample-specific barcodes in the forward primer, using a 30 cycle PCR and the HotStarTaq Plus Master Mix Kit (Qiagen, Valencia, CA, USA) under the following PCR conditions: 94 °C for 3 min, followed by 30 cycles of 94 °C for 30 s, 53 °C for 40 s and 72 °C for 1 min, followed by a final elongation at 72 °C for 5 min. We checked the PCR products on a 2% agarose gel to determine the success of amplification and the relative intensity of bands. Multiple PCR samples were then pooled together in equal proportions based on their molecular weight and DNA concentrations. Pooled samples were purified using calibrated Ampure XP beads (Agencourt Bioscience Corporation, Beverly, MA, USA) and were used to prepare DNA libraries following the Illumina TruSeq DNA library preparation protocol. We sequenced the samples using the Molecular Research LP next generation sequencing service (https://www.mrdnalab.com, Shallowater, TX, USA) on an Illumina MiSeq instrument (Illumina, San Diego, CA, USA) following the manufacturer’s guidelines.
Bioinformatics and taxonomic identification
We processed all raw MiSeq DNA sequence data following standard procedures as described in Schloss et al.74 using the mothur version 1.37.175. First, we removed low quality sequences and optimized the sequence lengths (to between 385 and 395 bp). We then aligned unique sequences to the SILVA-ARB reference database (release 123) to the same region of the 16S rRNA gene we sequenced and removed those columns that contained gaps only. Furthermore, independent of a reference database, we removed all the chimeric sequences using the uchime algorithm76 and the template as self, i.e. de novo removal. Subsequently, we clustered sequences into operational taxonomic units (OTUs) at the 97% DNA sequence similarity level. Representative sequences for OTUs were chosen as those that were most abundant in each cluster. We determined the taxonomic identity of each OTU with the ribosomal database project (RDP) Classifier77, and all sequences classified as chloroplast, mitochondria, and Archaea were removed. In order to standardize the number of reads across all samples, we subsampled (i.e. rarefied) equivalent reads from each sample. Rarefaction is believed to increase the false discovery rate78, but this is not true79 and instead, rarefying can lower sensitivity (false negatives) as a result of data discarding. Thus, recommendations are to rarefy to the highest depth possible80, which is what we did. Rarefaction is still considered a useful normalization technique, especially for uneven library sizes between groups, like here, and results in a higher PERMANOVA R2 for studied biological effects. However, it should still be noted that some OTUs can potentially be lost during rarefaction.
Germination experiments
Seeds of all three species were surface-sterilized for 5 min in 1% sodium hypochlorite, rinsed 3 times in distilled water, and dried at room temperature prior to the experiment to avoid fungal contamination. Thirty randomly selected seeds of O. stricta were sowed in petri dishes filled with 2 g of soil collected in invaded and uninvaded areas under the canopies of V. nilotica and S. africana and in open patches with no tree cover. Each treatment was replicated four times (n = 2 invasion scenarios [invaded, uninvaded] × 3 tree covers [V. nilotica, S. africana, none] × 4 replicates = 24 petri dishes). The same process was followed to test the germination of V. nilotica and S. africana in soils collected in invaded and uninvaded areas under their own canopy and in open patches with no tree cover (n = 2 species [V. nilotica, S. africana] × 2 invasion scenarios [invaded, uninvaded] × 2 tree covers [under its own canopy, none] × 4 replicates = 32 petri dishes). Petri dishes were moistened with 2 mL distilled water every 2 days and incubated in a germination chamber with periods of 12/12 h of light/dark and 25/15 °C day/night temperatures. Seeds were considered germinated when the seed coat was broken and the radicle visible (Posmyk et al. 2009). The number of germinated seeds was recorded every 2 days until no new seeds germinated. Total accumulated germination was calculated as Gt = [NT × 100]/N, where NT is the total germinated seeds in each treatment and N is the total number of seeds used in the treatment. Speed of accumulated germination was calculated as AS = [N1/1 + N2/2 + N3/3 … + Nn/n], where N1, N2, N3, Nn is the cumulative number of seeds which germinated at a 2-day interval 1, 2, 3, … N81.
Statistical analyses
All statistical analyses were conducted in the R statistical environment (version 3.5.1) (R Core Team 2017), unless otherwise specified. Where no package is indicated, functions used were from the vegan package version 2.3-382.
OTU accumulation curves were generated with the function specaccum to determine whether sampling was adequate to detect all OTUs present. Four diversity metrics were calculated from the sample x OTU matrix: OTU richness, the exponent of Shannon diversity, inverse Simpson diversity, and Pielou's evenness (OTU abundance equality)83,84. The exponent of Shannon and Inverse Simpson diversities were chosen since these metrics represent true diversities (i.e. "effective species"), unlike other diversity indices/entropies83,84. We calculated these various metrics with the function renyi. In order to investigate the influence of invasion (invaded vs. uninvaded) and tree cover (V. nilotica, S. africana, none) on the various diversity metrics, we performed two-way ANOVAs, including interaction effects. Significant differences between means were assessed using Tukey HSD post hoc tests.
For visualizing soil bacterial community composition, we performed Non-Metric Multidimensional Scaling (NMDS) using function metaMDS based on Horn similarity values85 for the 97% OTU table. Horn similarity values were calculated with the function sim.table in the vegetarian package86. To test for significant differences in soil bacterial community composition between different soils, we performed a Permutation Multivariate Analysis of Variance (PERMANOVA)87 with 9999 permutations using the function adonis.
We were interested whether the presence of native trees or invasive O. stricta homogenized soil bacterial communities. We tested this with the betadisper function (using 9999 permutations), which calculates differences in multivariate homogeneity of group variances between tree covers, and invaded and uninvaded soils.
To identify shifts in the abundance of OTUs due to the presence of native trees or the invasive O. stricta, we performed a linear discriminant analysis (LDA) effect size (i.e. LEfSe)88 using mothur75 at OTU level. LEfSe taxa are those that have shifted in abundance (identified using an alpha value of 0.05 and LDA score of > 2). Finally, we investigated which OTUs were broadly present in all soils as these are presumably not affected by invasion or tree cover.
Finally, in order to investigate the influence of invasion (invaded vs. uninvaded) and tree cover (V. nilotica, S. africana, none) on the soil pH, humidity, nutrients and enzymatic activities datasets and the germination indices, we performed two-way ANOVAs, including interaction effects. Significant differences between means were assessed using Tukey HSD post hoc tests.
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
Pyšek, P. et al. Naturalized alien flora of the world. Preslia 89, 203–274 (2017).
Pyšek, P. et al. Scientists’ warning on invasive alien species. Biol. Rev. (2020).
Vilà, M. et al. Ecological impacts of invasive alien plants: A meta-analysis of their effects on species, communities and ecosystems. Ecol. Lett. 14, 702–708 (2011).
Pyšek, P. et al. A global assessment of invasive plant impacts on resident species, communities and ecosystems: The interaction of impact measures, invading species’ traits and environment. Glob. Chang. Biol. 18, 1725–1737 (2012).
Le Roux, J. J. et al. Recent anthropogenic plant extinctions differ in biodiversity hotspots and coldspots. Curr. Biol. 29, 2912-2918.e2 (2019).
Hulme, P. E. et al. Greater focus needed on alien plant impacts in protected areas. Conserv. Lett. 7, 459–466 (2014).
Foxcroft, L. C., Pyšek, P., Richardson, D. M., Genovesi, P. & MacFadyen, S. Plant invasion science in protected areas: progress and priorities. Biol. Invasions 19, 1353–1378 (2017).
Novoa, A. et al. Invasion syndromes: A systematic approach for predicting biological invasions and facilitating effective management. Biol. Invasions 22, 1801–1820 (2020).
Foxcroft, L. C., Pickett, S. T. A. & Cadenasso, M. L. Expanding the conceptual frameworks of plant invasion ecology. Perspect. Plant Ecol. Evol. Syst. 13, 89–100 (2011).
Scholes, R. J. & Archer, S. R. Tree-grass interactions in savannas. Annu. Rev. Ecol. Syst. 28, 517–544 (1997).
Millennium Ecosystem Assessment. Ecosystems and Human Well-Being: Biodiversity Synthesis (Island Press, 2005).
Foxcroft, L. C., Richardson, D. M., Rejmánek, M. & Pyšek, P. Alien plant invasions in tropical and sub-tropical savannas: Patterns, processes and prospects. Biol. Invasions 12, 3913–3933 (2010).
Rejmánek, M., Huntley, B. J., Le Roux, J. J. & Richardson, D. M. A rapid survey of the invasive plant species in western Angola. Afr. J. Ecol. 55, 56–69 (2017).
Shackleton, R. T., Foxcroft, L. C., Pyšek, P., Wood, L. E. & Richardson, D. M. Assessing biological invasions in protected areas after 30 years: Revisiting nature reserves targeted by the 1980s SCOPE programme. Biol. Conserv. 243, 108424 (2020).
Skarpe, C. Dynamics of savanna ecosystems. J. Veg. Sci. 3, 293–300 (1992).
Okin, G. S. et al. Spatial patterns of soil nutrients in two southern African savannas. J. Geophys. Res. Biogeosci. 113, G2 (2008).
Ridolfi, L., Laio, F. & D’Odorico, P. Fertility island formation and evolution in dryland ecosystems. Ecol. Soc. 13, 5 (2008).
Perroni-Ventura, Y., Montaña, C. & Garcí-a-Oliva, F. Carbon-nitrogen interactions in fertility island soil from a tropical semi-arid ecosystem. Funct. Ecol. 24, 233–242 (2010).
Belnap, J. & Susan, L. P. Soil biota in an ungrazed grassland: response to annual grass (Bromus tectorum) invasion. Ecol. Appl. 51, 1261–1275. (2001).
Ludwig, F., Kroon, H., Prins, H. H. T. & Berendse, F. Effects of nutrients and shade on tree-grass interactions in an East African savanna. J. Veg. Sci. 12, 579–588 (2001).
Reinhart, K. O. & Callaway, R. M. Soil biota and invasive plants. New Phytol. 170, 445–457 (2006).
Weidenhamer, J. D. & Callaway, R. M. Direct and indirect effects of invasive plants on soil chemistry and ecosystem function. J. Chem. Ecol. 36, 59–69 (2010).
Levine, J. M., Pachepsky, E., Kendall, B. E., Yelenik, S. G. & Lambers, J. H. R. Plant-soil feedbacks and invasive spread. Ecol. Lett. 9, 1005–1014 (2006).
du Toit, J. T., Rogers, K. H. & Biggs, H. C. The Kruger Experience: Ecology and Management of Savanna Heterogeneity. (Island Press, 2003).
Foxcroft, L. C., Van Wilgen, N. J., Baard, J. A. & Cole, N. S. Biological invasions in South African National Parks. Bothalia 47, 11 (2017).
Pyšek, P. et al. Into the great wide open: do alien plants spread from rivers to dry savanna in the Kruger National Park?. NeoBiota 60, 61–77 (2020).
Kueffer, C., Pyšek, P. & Richardson, D. M. Integrative invasion science: Model systems, multi-site studies, focused meta-analysis and invasion syndromes. New Phytol. 200, 615–633 (2013).
Lotter, W. D. & Hoffmann, J. H. An integrated management plan for the control of Opuntia stricta (Cactaceae) in the Kruger National Park, South Africa. Koedoe 41, 63–68 (1998).
Hoffmann, J. H., Moran, V. C., Zimmermann, H. G. & Impson, F. A. C. Biocontrol of a prickly pear cactus in South Africa: Reinterpreting the analogous, renowned case in Australia. J. Appl. Ecol. 13737, 1365–2664. (2020).
Foxcroft, L. C., Rouget, M., Richardson, D. M. & MacFadyen, S. Reconstructing 50 years of Opuntia stricta invasion in the Kruger National Park, South Africa: Environmental determinants and propagule pressure. Divers. Distrib. 10, 427–437 (2004).
Novoa, A., Le Roux, J. J., Robertson, M. P., Wilson, J. R. U. & Richardson, D. M. Introduced and invasive cactus species: A global review. AoB Plants 7, 1 (2015).
Foxcroft, L. C., Hoffmann, J. H., Viljoen, J. J. & Kotze, J. J. Environmental factors influencing the distribution of Opuntia stricta, an invasive alien plant in the Kruger National Park, South Africa. S. Afr. J. Bot. 73, 109–112 (2007).
Foxcroft, L. C. & Rejmánek, M. What helps Opuntia stricta invade Kruger National Park, South Africa: Baboons or elephants?. Appl. Veg. Sci. 10, 265–270 (2007).
Anderson, E. F. The Cactus Family. (Timber Press, 2001).
Reyes-Agüero, J. A., Aguirre, R. J. R. & Valiente-Banuet, A. Reproductive biology of Opuntia: A review. J. Arid Environ. 64, 549–585 (2006).
Robertson, M. P. et al. Assessing local scale impacts of Opuntia stricta (Cactaceae) invasion on beetle and spider diversity in Kruger National Park, South Africa. Afr. Zool. 46, 205–223 (2011).
Butterfield, B. J. & Briggs, J. M. Patch dynamics of soil biotic feedbacks in the Sonoran Desert. J. Arid Environ. 73, 96–102 (2009).
Neffar, S., Chenchouni, H., Beddiar, A. & Redjel, N. Rehabilitation of degraded rangeland in drylands by Prickly Pear (Opuntia ficus-indica L.) plantations: Effect on soil and spontaneous vegetation. Ecol. Balk. 5, 63–76 (2013).
Garner, W. & Steinberger, Y. A proposed mechanism for the formation of ‘Fertile Islands’ in the desert ecosystem. J. Arid Environ. 16, 257–262 (1989).
Marchante, H., Elizabete M, & Helena, F. Invasion of the Portuguese dune ecosystems by the exotic species Acacia longifolia (Andrews) Willd.: effects at the community level. Plant invasions: ecological threats and management solutions. pp. 75–85 (2003).
Marchante, E. et al. Short-and long-term impacts of Acacia longifolia invasion on the belowground processes of a Mediterranean coastal dune ecosystem. Appl. Soil Ecol. 40(2), 210–217 (2008).
Yelenik, S. G., Stock, W. D. & Richardson, D. M. Ecosystem level impacts of invasive Acacia saligna in the South African fynbos. Restor. Ecol. 12(1), 44–51 (2004).
Werner, C. et al. High competitiveness of a resource demanding invasive acacia under low resource supply. Plant Ecol. 206(1), 83–96 (2010).
Le Maitre, D. C. et al. Impacts of invasive Australian acacias: implications for management and restoration. Divers. Distrib. 17(5), 1015–1029 (2011).
Bargali, K. & Bargali, S. S. Acacia nilotica: a multipurpose leguminous plant. Nat. Sci. 7, 11–19 (2009).
Rughöft, S. et al. Community composition and abundance of bacterial, archaeal and nitrifying populations in savanna soils on contrasting bedrock material in Kruger National Park, South Africa. Front. Microbiol. 7, 1638 (2016).
Neilson, J. W. et al. Life at the hyperarid margin: Novel bacterial diversity in arid soils of the Atacama Desert, Chile. Extremophiles 16, 553–566 (2012).
de Vos, P. et al. The Firmicutes. Bergey’s Manual of Systematic Bacteriology. (Springer, 2009).
Brockett, B. F. T., Prescott, C. E. & Grayston, S. J. Soil moisture is the major factor influencing microbial community structure and enzyme activities across seven biogeoclimatic zones in western Canada. Soil Biol. Biochem. 44, 9–20 (2012).
Yang, Y., Dou, Y. & An, S. Testing association between soil bacterial diversity and soil carbon storage on the Loess Plateau. Sci. Total Environ. 626, 48–58 (2018).
Rajaniemi, T. K. & Allison, V. J. Abiotic conditions and plant cover differentially affect microbial biomass and community composition on dune gradients. Soil Biol. Biochem. 41, 102–109 (2009).
Novoa, A., Rodríguez, R., Richardson, D. & González, L. Soil quality: A key factor in understanding plant invasion? The case of Carpobrotus edulis (L.) N.E.Br. Biol. Invasions 16, 429–443 (2014).
Penfield, S. Seed dormancy and germination. Curr. Biol. 27, R874–R878 (2017).
Tielbörger, K. & Prasse, R. Do seeds sense each other? Testing for density-dependent germination in desert perennial plants. Oikos 118, 792–800 (2009).
Renne, I. J. et al. Eavesdropping in plants: delayed germination via biochemical recognition. J. Ecol. 102, 86–94 (2014).
Yannelli, F. A., Novoa, A., Lorenzo, P., Rodríguez, J. & Le Roux, J. J. No evidence for novel weapons: biochemical recognition modulates early ontogenetic processes in native species and invasive acacias. Biol. Invasions 22, 549–562 (2020).
Al-Wakeel, S. A. M., Gabr, M. A., Hamid, A. A. & Abu-El-Soud, W. M. Allelopathic effects of Acacia nilotica leaf residue on Pisum sativum L. Allelopath. J. 19, 411 (2007).
Scholes, M. C., Scholes, R. J., Otter, L. B. & Woghiren, A. J. Biogeochemistry: The cycling of elements. in The Kruger Experience: Ecology and Management of Savanna Heterogeneity (eds. du Toit, J. T., Rogers, K. H. & Biggs, H. C.) 130–148 (Island Press, 2003).
Kyalangalilwa, B., Boatwright, J. S., Daru, B. H., Maurin, O. & van der Bank, M. Phylogenetic position and revised classification of Acacia s.l. (Fabaceae: Mimosoideae) in Africa, including new combinations in Vachellia and Senegalia. Bot. J. Linn. Soc. 172, 500–523 (2013).
van Wyk, B. & van Wyk, P. Field Guide to Trees of Southern Africa. (Struik Nature, 2013).
Coates Palgrave, K. & Coates Palgrave, M. Palgrave’s Trees of Southern Africa. (Struik Publishers, 2002).
Novoa, A., Kumschick, S., Richardson, D. M., Rouget, M. & Wilson, J. R. U. Native range size and growth form in Cactaceae predict invasiveness and impact. NeoBiota 30, 75–90 (2016).
Allen, S. E. Chemical Analysis of Ecological Materials. (Blackwell Scientific Publications, 1989).
Tabatabai, M. A. & Bremner, J. M. Use of p-nitrophenyl phosphate for assay of soil phosphatase activity. Soil Biol. Biochem. 1, 301–307 (1969).
Kandeler, E. & Gerber, H. Short-term assay of soil urease activity using colorimetric determination of ammonium. Biol. Fertil. Soils 6, 68–72 (1988).
Allison, S. D. & Vitousek, P. M. Extracellular enzyme activities and carbon chemistry as drivers of tropical plant litter decomposition. Biotropica 36, 285–296 (2004).
German, D. P., Chacon, S. S. & Allison, S. D. Substrate concentration and enzyme allocation can affect rates of microbial decomposition. Ecology 92, 1471–1480 (2011).
Lane, D. J. et al. Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc. Natl. Acad. Sci. 82, 6955–6959 (1985).
Tringe, S. G. & Hugenholtz, P. A renaissance for the pioneering 16S rRNA gene. Curr. Opin. Microbiol. 11, 442–446 (2008).
Bukin, Y. S. et al. The effect of 16s rRNA region choice on bacterial community metabarcoding results. Sci. Data 6, 1–14 (2019).
Chakravorty, S., Helb, D., Burday, M., Connell, N. & Alland, D. A detailed analysis of 16S ribosomal RNA gene segments for the diagnosis of pathogenic bacteria. J. Microbiol. Methods 69, 330–339 (2007).
Beckers, B. et al. Performance of 16s rDNA primer pairs in the study of rhizosphere and endosphere bacterial microbiomes in metabarcoding studies. Front. Microbiol. 7, 1–15 (2016).
Thijs, S. et al. Comparative evaluation of four bacteria-specific primer pairs for 16S rRNA gene surveys. Front. Microbiol. 8, 1–15 (2017).
Schloss, P. D., & Westcott, S. L. Assessing and improving methods used in operational taxonomic unit-based approaches for 16S rRNA gene sequence analysis. Appl. Environ. Microbiol. 77(10), 3219–3226 (2011).
Schloss, P. D. et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
McMurdie, P. J. & Holmes, S. Waste not, want not: Why rarefying microbiome data is inadmissible. PLoS Comput. Biol. 10, e1003531 (2014).
Weiss, S. et al. Normalization and microbial differential abundance strategies depend upon data characteristics. Microbiome 5, 1–18 (2017).
de Cárcer, D. A., Denman, S. E., McSweeney, C. & Morrison, M. Evaluation of subsampling-based normalization strategies for tagged high-throughput sequencing data sets from gut microbiomes. Appl. Environ. Microbiol. 77, 8795–8798 (2011).
Chiapusio, G., Sánchez, A. M., Reigosa, M. J., González, L. & Pellissier, F. Do germination indices adequately reflect allelochemical effects on the germination process?. J. Chem. Ecol. 23, 2445–2453 (1997).
Oksanen, J. F. et al. vegan: Community Ecology Package. R package version 2.3-3. (2016).
Jost, L. Entropy and diversity. Oikos 113, 363–375 (2006).
Jost, L. The relation between evenness and diversity. Diversity 2, 207–232 (2010).
Jost, L. Partitioning diversity into independent alpha and beta components. Ecology 88, 2427–2439 (2007).
Charney, N. & Record, S. vegetarian: Jost Diversity Measures for Community Data. R package version 1.2. (2012).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral. Ecol. 26, 32–46 (2001).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, 1–18 (2011).
Clarke, K. R. Non-parametric multivariate analyses of changes in community structure. Aust. J. Ecol. 18, 117–143 (1993).
Acknowledgements
This work was supported by the Czech Science Foundation (project no. 19–13142S, and EXPRO no. 19–28807X); Czech Academy of Sciences (long-term research development project RVO 67985939); the DST-NRF Centre of Excellence for Invasion Biology; Faculty of Science and Department of Biological Sciences, Macquarie University; and the South African National Department of Environment Affairs through its funding of the South African National Biodiversity Institute’s Invasive Species Programme.
Author information
Authors and Affiliations
Contributions
A.N. and L.C.F. conceived the research; A.N., L.C.F. and J.J.L.R. designed and performed the experiments; A.N. and J.H.K. analysed the data and drafted the manuscript, which was critically reviewed by all authors; and all authors approved the final version of the manuscript for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Novoa, A., Foxcroft, L.C., Keet, JH. et al. The invasive cactus Opuntia stricta creates fertility islands in African savannas and benefits from those created by native trees. Sci Rep 11, 20748 (2021). https://doi.org/10.1038/s41598-021-99857-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-99857-x
- Springer Nature Limited
This article is cited by
-
Assessing the level of compliance with alien plant regulations in a large African protected area
Biological Invasions (2022)