Abstract
The plant pathogenic bacterium Pseudomonas syringae pv. tomato DC3000 (Pst DC3000) has become a paradigm to investigate plant-bacteria interactions due to its ability to cause disease in the model plant Arabidopsis thaliana. Pst DC3000 uses the type III secretion system to deliver type III secreted effectors (T3SEs) directly into the plant cytoplasm. Pst DC3000 T3SEs contribute to pathogenicity by suppressing plant defense responses and targeting plant’s physiological processes. Although the complete repertoire of effectors encoded in the Pst DC3000 genome have been identified, the specific function for most of them remains to be elucidated. Among those effectors, the mitochondrial-localized T3E HopG1, suppresses plant defense responses and promotes the development of disease symptoms. Here, we show that HopG1 triggers necrotic cell death that enables the growth of adapted and non-adapted pathogens. We further showed that HopG1 interacts with the plant immunity-related protein AtNHR2B and that AtNHR2B attenuates HopG1- virulence functions. These results highlight the importance of HopG1 as a multi-faceted protein and uncover its interplay with AtNHR2B.
Similar content being viewed by others
Introduction
Pseudomonas syringae is a plant pathogenic Gram-negative bacterium that causes diseases in a wide range of plants. Due to this broad host range, the species has been divided into more than 50 pathovars (pv), each pathovar designation based on their host of isolation1. Among those pathovars, Pseudomonas syringae pv. tomato DC3000 (Pst DC3000), the causal agent of bacterial speck on tomato, has become a model pathogen to understand bacterial pathogenicity towards plants because it can also cause disease in model plants. Wild type Pst DC3000 causes disease in the model plant Arabidopsis thaliana, and the Pst DC3000 hopQ1-1 mutant is able to cause disease in Nicotiana benthamiana1,2,3,4. The pathogenicity of Pst DC3000 is mostly due to the type III secretion system (T3SS), a complex of proteins encoded by the Hypersensitive Response and Pathogenicity/ Hypersensitive Response and Conserved (Hrp/Hrc) genes5. Hrp/Hrc- encoded proteins assemble an apparatus spanning the inner and outer bacterial membranes that enables the bacterium to deliver bacterial proteins (effectors), directly into the host cytoplasm6,7,8.
The genome of Pst DC3000 encodes 28 type III effectors (T3Es) that are delivered into plant cells9,10,11, where acting together interfere with plant immune responses to facilitate bacterial parasitism12,13,14. The plant immune responses include two main branches: (1) Pathogen-Associated Molecular Patterns-Triggered Immunity (PTI) that recognizes conserved features in pathogens known as Pathogen-Associated Molecular Patterns (PAMPs) through surface-localized Pattern Recognition Receptors (PRRs)15,16,17, and (2) Effector-Triggered Immunity (ETI), that recognizes pathogen effector molecules by R (resistance) proteins15,16,18. An outcome of ETI is the elicitation of the hypersensitive response (HR), a type of localized programmed cell death that restricts pathogen proliferation and tissue damage19,20,21.
Pst DC3000 T3Es are functionally redundant, as mutations in individual effectors do not have a significant impact on pathogenicity, yet all the effectors are required for full pathogenicity in tomato and Arabidopsis22. Interestingly, deletion of the T3E HopQ1-1 expands the host range of Pst DC3000 to N. benthamiana4. Remarkably, only eight T3Es effectors: AvrPtoB, HopM1, AvrE, HopE1, HopG1, HopAM1-1, HopAA1-1 and HopN1 make up the minimal functional set of effectors needed for Pst DC3000 to be virulent in N. benthamiana23. Among those effectors, HopG1 was previously implicated in the suppression of PTI and ETI responses24,25,26. Further research showed that HopG1 localizes to mitochondria24, where it interacts with the mitochondrial-localized kinesin motor protein to modulate actin cytoskeleton and promote the development of chlorotic symptoms27.
We previously identified AtNHR2B (Arabidopsis thaliana nonhost resistance 2B) as a protein that functions in plant immunity28,29,30,31. Atnhr2b mutant plants are immunocompromised, and as a result, they are susceptible to pathogens that normally do not cause disease in wild-type plants, called non-adapted pathogens31. We further showed by live-cell imaging that AtNHR2B tagged with the green fluorescent protein (GFP) localizes to chloroplasts and compartments of the endomembrane system31, and through proteomics approaches, we also showed that AtNHR2B interacts with proteins localized to chloroplasts, mitochondria and nucleus32.
Here, we show that the Pst DC3000 T3E HopG1 interacts with AtNHR2B and that AtNHR2B interferes with HopG1 virulence functions. We also expand the range of functions for HopG1 with the discovery that transient or stable expression of HopG1 in N. benthamiana and Arabidopsis, respectively, triggers necrotic cell death that enables the growth of adapted and non-adapted pathogens. This necrotic cell death is a virulence function that confers advantage to the pathogen, rather than an HR-type of programmed cell death that confers resistance to the plant. These results highlight the importance of HopG1 as a multifaceted protein and uncover the interplay between HopG1 and AtNHR2B.
Results
Adapted pathogens changed the pattern of localization and abundance of AtNHR2B-GFP
We previously showed that AtNHR2B is induced by pathogens and pathogen-derived elicitors, and that AtNHR2B-GFP transiently expressed in N. benthamiana localizes to the cytoplasm and to highly dynamic structures (punctae) reminiscent of subcellular compartments of the endomembrane system31. AtNHR2B is induced by non-adapted bacterial pathogens (that are unable to cause disease in a particular host), as well as, adapted bacterial pathogens (that cause disease in a specific host); however, we do not know if pathogens’ lifestyle (non-adapted vs adapted) have an effect on the function of AtNHR2B. Since AtNHR2B is an immune-related protein, we hypothesized that adapted pathogens could target this protein for parasitism. To start testing that hypothesis, we evaluated the localization of AtNHR2B-GFP by transient expression in N. benthamiana. N. benthamiana plants transiently expressing AtNHR2B-GFP were mock-treated with water, inoculated with P. syringae pv. tabaci (Pstab) (adapted pathogen of N. benthamiana), or with Pst DC3000 (non-adapted pathogen of N. benthamiana). Mock-treated plants and plants inoculated with Pst DC3000 showed the expected localization to cytoplasm and punctae. Interestingly, after inoculation with Pstab, the signal from AtNHR2B-GFP was limited to the cytoplasm (Fig. 1a).
Adapted pathogens interfere with AtNHR2B-GFP localization and protein abundance. (a) The host adapted pathogen Pstab alters AtNHR2B-GFP localization. N. benthamiana plants transiently expressing AtNHR2B-GFP were infiltrated with water (mock), or inoculated with either the adapted pathogen Pstab or the non-adapted pathogen Pst DC3000 at 1 × 106 CFU/ml. Treated leaf samples were collected after 24 hpi and imaged by laser scanning confocal microscopy. Confocal images were taken at ×20 magnification and using an excitation and emission wavelengths of 488 nm and 500–535 nm, respectively. White arrows show punctate bodies. Scale bar = 10 µm. (b) The host adapted pathogen Pst DC3000 alters AtNHR2B-GFP localization in Arabidopsis. Arabidopsis transgenic plants expressing AtNHR2B-GFP were mock-treated or inoculated with the adapted pathogen Pst DC3000 and the non-adapted pathogen Pstab at 1 × 106 CFU/ml. Treated leaf samples were collected after 24 hpi and imaged by laser scanning confocal microscopy. Confocal images were taken at ×20 magnification and using an excitation and emission wavelengths of 488 nm and 500–535 nm, respectively. White arrows show punctate bodies. Scale bar = 10 µm. (c) The host adapted pathogen Pstab alters AtNHR2B-GFP protein abundance in tobacco plants. N. benthamiana plants transiently expressing AtNHR2B-GFP were infiltrated with water (mock) or inoculated with either the adapted pathogen Pstab or the non-adapted pathogen Pst DC3000 as described for (a). Treated leaf samples were collected after 24 hpi to evaluate protein abundance by Western blot using anti-GFP and anti-actin antibodies. Protein quantification was done by ImageJ software (http://rsb.info.nih.gov/ij/), using actin signal for normalization. Number below the protein bands represent the AtNHR2-GFP/Actin ratios. (d) The host adapted pathogen Pst DC3000 alters AtNHR2B-GFP protein abundance in Arabidopsis plants. A. thaliana transgenic plants expressing AtNHR2B-GFP were infiltrated with water (mock), or inoculated with either the adapted pathogen Pst DC3000 or the non-adapted pathogen Pstab as described for (b). Treated leaf samples were collected after 24 hpi to quantify protein abundance by Western blot using anti-GFP and anti-Actin antibodies. Protein quantification was done as described for (c).
The change in AtNHR2B-GFP pattern of localization in N. benthamiana associated with inoculation with the adapted pathogen was validated in transgenic Arabidopsis plants expressing AtNHR2B-GFP from its native promoter31. In such plants, the fluorescent signal was very low in mock-treated plants, but significantly increased upon inoculation with Pst DC3000 and Pstab. Similar to the results obtained in N. benthamiana, the pattern of AtNHR2B-GFP localization differed depending on the pathogen used for the inoculation; inoculation with the non-adapted pathogen of A. thaliana, Pstab, caused the localization of AtNHR2B-GFP to cytoplasm and small punctae as previously described31 (Fig. 1b). However, inoculation with the adapted pathogen of A. thaliana, Pst DC3000, changed the pattern of AtNHR2B-GFP localization and the fluorescence signal was detected in the cytoplasm only and appeared diffused and distorted (Fig. 1b).
To evaluate if the diffusion of the fluorescence signal was due to adapted pathogens interfering with protein abundance, we collected leaves from N. benthamiana and Arabidopsis plants expressing AtNHR2B-GFP that were mock-treated, or inoculated with their corresponding adapted and non-adapted pathogens. Collected leaves were used for protein extraction and quantitative Western blot analysis using anti-GFP antibodies to detect AtNHR2B-GFP and anti-actin antibodies for normalization. N. benthamiana expressing AtNHR2B-GFP and mock-treated, showed low levels of AtNHR2B-GFP. However, inoculation with both adapted (Pstab) and non-adapted (Pst DC3000) pathogens increased AtNHR2B-GFP concentration in comparison with mock-treated plants, but the amount of AtNHR2B-GFP was reduced by 50% in plant samples treated with Pstab in comparison with plants treated with Pst DC3000 (Fig. 1c, Supplementary Fig. 1). In a similar way, AtNHR2B-GFP transgenic Arabidopsis lines did not accumulate any protein after mock treatment and showed ~ 40% less AtNHR2B-GFP after inoculation with the adapted pathogen Pst DC3000 than plants inoculated with Pstab (Fig. 1d. Supplementary Fig. 2). Altogether, these results revealed that adapted pathogens alter AtNHR2B-GFP protein localization and abundance.
The Pst DC3000 mutant lacking HopG1 is able to grow better in the Atnhr2b mutant background
The finding that the localization and protein abundance of AtNHR2B is altered by adapted pathogens, prompted us to investigate whether AtNHR2B could be a target for Pst DC3000 T3E. The finding that HopG1 localizes to mitochondria24,27, together with our previous result that AtNHR2B interacts with proteins localized to mitochondria32, led us to hypothesize that HopG1 could target AtNHR2B. To test that hypothesis, we initially evaluated the growth of wild-type Pst DC3000 and the Pst DC3000 ΔhopG1 mutant in wild-type Col-0 and in Atnhr2b mutant plants at 3 days post-inoculation (dpi) (Fig. 2). As an adapted pathogen of Arabidopsis, the wild-type strain Pst DC3000 is able to grow in wild-type Col-0 to 107 CFU/cm2 at 3 dpi. In contrast, the growth of the Pst DC3000ΔhopG1 was ~ 10-fold lower than the growth of wild-type Pst DC3000 in wild-type Col-0. In the Atnhr2b mutant plants, the growth of the wild-type Pst DC3000 was equivalent to its growth in wild-type Col-0 plants. Interestingly, the Pst DC3000ΔhopG1 mutant grew to higher levels (~ 107 CFU/cm2) in Atnhr2b mutant plants, and those levels were equivalent to the growth of Pst DC3000 in wild-type Col-0. These results suggest that the Atnhr2b mutation restores the growth defect in the PstDC3000ΔhopG1 mutant supporting a functional relationship between HopG1 and AtNHR2B.
The growth defect in Pst DC3000 hopG1 deletion mutant is restored in Atnhr2b mutant Arabidopsis plants. Wild-type Col-0 and the Atnhr2b mutant were syringe-inoculated with the wild-type Pst DC3000 and the Pst DC3000 Δ hopG1 mutant at a concentration of 1 × 104 CFU/ml. Inoculated leaves were collected at 0 and 3 dpi to enumerate bacterial populations. Bars represent means and standard deviation for Pst DC3000 and ΔhopG1 bacterial populations in Col-0 and Atnhr2b plants for four replications. Different letters above the bars represent statistically significant difference in bacterial populations of wild-type Pst DC3000 and the Pst DC3000 Δ hopG1 in Col-0 versus their respective growth in the Atnhr2b mutant plants using a one-way ANOVA and Tukey’s multiple comparison test with P-value ≤ 0.05. The experiments were repeated three times with consistent results.
The Pst DC3000 effector HopG1 interacts with AtNHR2B
To start dissecting the functional relationship between HopG1 and AtNHR2B, we tested the physical interaction between AtNHR2B and HopG1 with the yeast two- hybrid system using HopG1 as a bait and AtNHR2B as a prey. Yeast co-transformed with pDEST32::HopG1 and pDEST22::AtNHR2B grew on Triple Drop Out (TDO) media (-leu,-trp,-his) containing 15 mM 3-AT indicating the transcriptional activation of histidine biosynthetic genes, as a result of the interaction between HopG1 and AtNHR2B. That interaction is not the result of auto-activation, because yeast transformed with the empty vector pDEST32 and pDEST22::AtNHR2B did not grow on TDO + 15 mM 3-AT (Fig. 3a).
HopG1 interacts with AtNHR2B. (a) HopG1 interacts with AtNHR2B in yeast. Yeast strain MaV203 was co-transformed with pDEST32::HopG1 and pDEST22:: AtNHR2B, or pDEST32 and pDEST22::AtNHR2B and transformants were isolated in on Double Drop Out (DDO, SD/-Leu/-Trp) media and transferred to Triple Drop Out (TDO, SD/-His/-Leu/-Trp) + 15 mM 3-Amino-1,2,4-Triazole. All the experiments were repeated three times and shown similar results. (b) HopG1 and AtNHR2B-GFP interact in planta. HopG1-FLAG and AtNHR2B-GFP, or HopG1-FLAG and 35S-GFP combinations were transiently co-expressed in 5-week-old N. benthamiana plants. Infiltrated leaves were harvested for protein extraction followed by immunoprecipitation using GFP Nanobody/VHH coupled to agarose beads. Immunoprecipitated samples were separated by SDS-PAGE electrophoresis and transferred to a Polyvinylidene difluoride (PVDF) membrane for Western Blot analysis using anti-FLAG and anti-GFP antibodies. (c) Bimolecular fluorescence complementation analysis shows the interaction between HopG1 and AtNHR2B in planta. HopG1 fused to the C-terminal encoding half of EYFP (HopG1- cEYFP), was transiently co-expressed in N. benthamiana with AtNHR2B fused to the N-terminal encoding half of EYFP (nEYFP-AtNHR2B), or with the truncated version AtNHR2B Δ1–195 fused to the N-terminal encoding half (nEFYP-AtNHR2B Δ1–195). At 3 dpi, infiltrated leaves were imaged by laser scanning confocal microscopy using excitation and emission wavelengths of 488 nm and 500–535 nm, respectively. Confocal images were taken at ×63 magnification. Scale bars = 50 µm.
To evaluate the HopG1/AtNHR2B interaction in the appropriate biological context, we transiently co-expressed HopG1 fused to the FLAG epitope (HopG1-FLAG) with AtNHR2B-GFP for co-immunoprecipitation. HopG1-FLAG and free GFP were also co-infiltrated and used as control. Immunoprecipitation of AtNHR2B-GFP using GFP Nanobody/VHH coupled to agarose beads (GFP-Trap® Agarose, Chromotek), co-immunoprecipitated HopG1-FLAG as detected by Western Blot using Anti-FLAG antibodies (Fig. 3b, Supplementary Fig. 3). In contrast, immunoprecipitation of free GFP did not co-immunoprecipitate HopG1-FLAG, demonstrating that the physical interaction between HopG1 and AtNHR2B also occur in planta and it is not an artifact of the GFP tag. We further evaluated the in-situ interaction of AtNHR2B and HopG1 by co-expressing AtNHR2B fused to the N-terminal half encoding domain of the enhanced yellow fluorescent protein gene (nEYFP), and the HopG1 fused to the C-terminal half encoding domain of EYFP (cEYFP) for Bimolecular fluorescence complementation (BiFC) assays33. The in-situ interaction between AtNHR2B and HopG1 was revealed by the restoration of the yellow fluorescence and shown to occur in the cytoplasm; importantly, amino acids 1–195 in AtNHR2B are important for the interaction as deletion of the region encoding those amino acids abolishes the interaction with HopG1 (Fig. 3c).
HopG1 expression causes virulence-related cell death in N. benthamiana that is attenuated by AtNHR2B
To further understand how the interaction between HopG1 and AtNHR2B in planta was related with the function of HopG1, we transiently co-expressed HopG1-FLAG with free GFP or with AtNHR2B-GFP in N. benthamiana. At 4 days after infiltration, the section of the leaf expressing HopG1-FLAG alone (Supplementary Fig. 4a), or co-expressed with GFP (Fig. 4a) showed extensive cell death, but that cell death was reduced in the section of the leaf co-expressing HopG1-FLAG and AtNHR2B-GFP (Fig. 4a, Supplementary Fig. 4a). Similar results were obtained when transiently expressing in N. benthamiana another version of HopG1 epitope-tagged with HA (HopG1-HA), and when transiently co-expressing AtNHR2B-GFP with HopG1-HA (Supplementary Fig. 4b). To further understand how AtNHR2B reduced the cell death phenotype caused by HopG1, we evaluated HopG1 protein abundance by Western blot. Tissues co-expressing HopG1-FLAG with AtNHR2B-GFP had reduced levels of HopG1-FLAG in comparison with tissues co-expressing HopG1-FLAG with GFP (Fig. 4b, Supplementary Fig. 5). Similar results were obtained when transiently expressing in N. benthamiana HopG1-HA alone or in combination with AtNHR2B-GFP (Supplementary Fig. 6). Co-expression of AtNHR2B-GFP with HopG1-FLAG did not alter the AtNHR2B-GFP protein abundance (Supplementary Fig. 7). Collectively, these results indicate that HopG1 transiently expressed in N. benthamiana induces cell death that is attenuated by AtNHR2B degrading HopG1.
Transient expression of HopG1 in N. benthamiana triggers cell death that is attenuated by AtNHR2B. (a) Phenotypic evaluation of cell death in N. benthamiana. HopG1-FLAG was transiently co- expressed in N. benthamiana with AtNHR2B-GFP or with GFP alone. At 24 hpi, 20 μM DEX was sprayed to induce the expression of HopG1-FLAG. Cell death was evaluated at 4 dpi. (b) AtNHR2B alters the abundance of HopG1 protein. HopG1-FLAG was transiently co- expressed in N. benthamiana with AtNHR2B-GFP or with GFP alone. At 24 hpi, 20 μM DEX was sprayed to induce the expression of HopG1-FLAG. Infiltrated leaves were collected at 72 hpi for protein extraction and Western Blot analysis using anti-FLAG antibodies. Ponceau S staining shows equal protein loading. (c) Transient expression of HopG1-FLAG enhances Pstab growth in N. benthamiana. Wild-type N. benthamiana and N. benthamiana plants transiently expressing AtNHR2B-GFP, HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG, were syringe-inoculated with the adapted bacterial pathogen Pstab at a concentration of 1 × 104 CFU/ml. HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG expressing leaves were sprayed with 20 μM DEX 24 h prior to Pstab inoculation. Inoculated leaves were collected at 0, 1, 2 and 3 dpi to enumerate bacterial populations. Bars represent means and standard deviation for bacterial growth in wild-type N. benthamiana and in N. benthamiana plants transiently expressing AtNHR2B-GFP, HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG. Different letters above bars represent statistically significant difference in the growth of Pstab in wild-type N. benthamiana and N. benthamiana plants transiently expressing the aforementioned constructs, using a two-way ANOVA and Tukey’s multiple comparison test with P-value ≤ 0.05. The experiments were repeated three times with consistent results.
Since the cell death phenotype can be an HR defense response or necrosis associated with virulence, we resolved between those two alternatives by inoculating N. benthamiana plants transiently expressing HopG1-FLAG with Pstab to evaluate its growth at 1, 2 and 3 dpi. For these experiments, we took advantage of a version of Pstab expressing the fluorescent protein GFP-TIR34 (Pstab-GFP-TIR), to be able to discriminate Pstab from the A. tumefaciens strains used for transient expression. As an adapted pathogen of N. benthamiana, Pstab-GFP-TIR progressively grew in wild-type N. benthamiana and reached more than 108 CFU/cm2 at 3 dpi (Fig. 4c). In contrast, N. benthamiana plants that were transiently expressing AtNHR2B-GFP, HopG1-FLAG or co-expressing AtNHR2B-GFP and HopG1-FLAG had significantly lower bacterial populations, in comparison with wild-type plants, likely as a result of Agrobacterium-induced PTI (Fig. 4c). Interestingly, among transiently-expressing plants, those expressing HopG1-FLAG showed the highest levels of Pstab-GFP-TIR populations at 1, 2 and 3 dpi. Moreover, while Pstab-GFP-TIR populations at 1 dpi were equivalent between plants transiently expressing AtNHR2B-GFP alone or in combination with HopG1-FLAG, at 2 and 3 dpi Pstab-GFP-TIR populations in plants co-expressing AtNHR2B-GFP and HopG1-FLAG were intermediate when compared with populations in plants expressing either AtNHR2B-GFP or HopG1-FLAG alone (Fig. 4c). These results are consistent with the reduction in cell death phenotype and HopG1-FLAG accumulation in N. benthamiana plants co-expressing AtNHR2B-GFP with HopG1-FLAG. Taken together, these results show that HopG1 triggers plant cell death that is not an HR, but either contributes to plant susceptibility by suppressing PTI, enhances pathogen’ virulence or both. Moreover, this virulence activity of HopG1-inducing cell death that promotes pathogen multiplication, is actively attenuated by AtNHR2B.
Arabidopsis transgenic plants expressing HopG1-FLAG induce mitochondrial ROS-related cell death that is attenuated by AtNHR2B
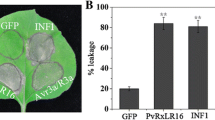
To further validate the results observed in N. benthamiana, we obtained transgenic Arabidopsis lines expressing HopG1-FLAG under dexamethasone inducible promoter, and further crossed them with transgenic lines expressing AtNHR2B-GFP. Wild-type Col-0 plants and transgenic plants expressing HopG1-FLAG, AtNHR2B-GFP and HopG1-FLAG/AtNHR2B-GFP were inoculated with the non-adapted pathogen Pstab or mock-treated with water to evaluate cell death after staining with Trypan Blue35. Inoculation with Pstab triggered cell death in transgenic plants expressing HopG1-FLAG alone or in combination with AtNHR2B-GFP, whereas, no cell death was observed in wild-type Col-0 plants, nor in plants expressing AtNHR2B-GFP alone. Importantly, less cell death was observed in transgenic plants co-expressing HopG1-FLAG with AtNHR2B-GFP, suggesting that AtNHR2B-GFP also attenuates the HopG1-triggered cell death in Arabidopsis (Fig. 5a). We confirmed that to be the case, by using the same plants’ genotypes and pathogen inoculation conditions to quantify cell death by electrolyte leakage at 5-, 10-, and 20-h post-inoculation (hpi). The results showed that as early at 10 hpi, plants expressing HopG1-FLAG have the highest levels of electrolyte leakage that were significantly different from the electrolyte leakage levels in Col-0 plants, as well as in plants expressing AtNHR2B-GFP and plants co-expressing HopG1-FLAG and AtNHR2B-GFP (Fig. 5b).
Transgenic expression of HopG1 in Arabidopsis causes cell death that is attenuated by AtNHR2B. (a) Cell death phenotype in HopG1 expressing plants is reduced by expression of AtNHR2B. Five-week-old wild-type Col-0, AtNHR2B-GFP, HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG plants were syringe-inoculated with Pstab at a concentration of 1 × 106 CFU/mL or infiltrated with water (mock). HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG plants were sprayed with 20 μM DEX 24 h prior bacterial inoculation. At 24 hpi, treated leaves were detached and stained with 0.05% trypan blue. Images were taken on a light microscope using bright field. (b) Ion leakage is enhanced in plants expressing HopG1. Five-week-old wild-type Col-0 and transgenic plants expressing AtNHR2B-GFP, HopG1-FLAG, and AtNHR2B-GFP/HopG1-FLAG were syringe-inoculated with Pstab at a concentration of 1 × 106 CFU/ml to evaluate electrolyte leakage at 5, 10, 20 and 24 hpi. HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG were sprayed with 20 μM DEX 24 h prior to bacterial inoculation. Bars represent means and standard deviation of conductivity values (µS/cm). Different letters above bars represent statistically significant difference in electrolyte leakage among plant genotypes using a one-way ANOVA and Fisher’s multiple comparison test with P-value ≤ 0.05. (c) Arabidopsis plants expressing HopG1-FLAG generate higher levels of ROS from mitochondrial origin. Wild-type Col-0 and transgenic plants expressing HopG1-FLAG, AtNHR2B-GFP and AtNHR2B-GFP/HopG1-FLAG were flood-inoculated with the non-adapted bacterial pathogen Pstab at a concentration of 1 × 106 CFU/ml, or infiltrated with water (mock). HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG were sprayed with 20 μM DEX 24 h prior bacterial inoculation. Leaf disks were incubated with MitoTracker Red CM-H2XRos and mitochondrial ROS fluorescence was measured using excitation/emission wavelengths of 570 nm and 535 nm, respectively.
Because cell death phenotypes are regulated by reactive oxygen species (ROS), and mitochondria are one of the source of ROS, it was necessary to evaluate how HopG1 contributes to the production of ROS in the mitochondria. For that purpose, a mitochondria-specific ROS sensor, MitoTracker Red CM-H2XRos (ThermoFisher Scientific, Waltham, MA) was used to evaluate mitochondrial ROS produced after pathogen infection. Inoculation of Pstab triggered an accumulation of mitochondrial ROS in all plant genotypes. However, the levels of accumulation varied between genotypes. Overall, the lowest levels of mitochondrial ROS were observed in wild-type Col-0 and moderate levels were found in transgenic plants expressing AtNHR2B-GFP and co-expressing HopG1-FLAG and AtNH2B-GFP. Remarkably, at 3 min of the assay, the levels of mitochondrial ROS in plants expressing HopG1-FLAG started to increase and those levels peaked at 7 min when they were ~ 3-fold higher than in wild-type Col-0 and ~ 1.5-fold higher than in plants expressing AtNHR2B-GFP or co-expressing AtNHR2B-GFP and HopG1-FLAG (Fig. 5c, Supplementary Table S1). These results showed that the HopG1-mediated cell death is associated with the production of reactive oxygen species in the mitochondria where HopG1 localizes.
HopG1 interferes with AtNHR2B function to promote disease
Our results demonstrating an interaction between HopG1 and AtNHR2B and a possible interplay between both proteins, prompted us to further investigate the antagonism between HopG1 and AtNHR2B. HopG1 was previously shown to interfere with callose deposition when HopG1-HA expressing plants were inoculated with the Pst DC3000 hrcC mutant24. In our assays, we used Pstab that also triggers PTI in Arabidopsis as demonstrated by the callose deposits in the wild-type Col-0 plants and in transgenic Arabidopsis plants expressing AtNHR2B-GFP (Fig. 6a). As previously reported, transgenic plants expressing HopG1-FLAG were devoid of callose deposits and were comparable to mock-treated plants. Similarly, plants co-expressing HopG1-FLAG and AtNHR2B-GFP were also devoid of callose deposits suggesting that AtNHR2B is unable to interfere with the PTI-suppressing activities of HopG1 (Fig. 6a).
Transgenic expression of HopG1 interferes with AtNHR2B-dependent plant defense responses. (a) Plants expressing HopG1 are deficient in callose deposition. Five-week-old wild-type Col-0, AtNHR2B-GFP, HopG1-FLAG, and AtNHR2B-GFP/HopG1-FLAG were syringe-inoculated with the non-adapted bacterial pathogen Pstab at a concentration of 1 × 5/ml. HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG were sprayed with 20 μM DEX 24 h prior to bacterial inoculation. Inoculated leaves were detached at 24 hpi and stained with 5% aniline blue staining to evaluate callose deposition. Images were taken using a confocal microscope under DAPI filter. Scale bar = 20 μm. (b) Transgenic expression of HopG1- contributes to bacterial growth. Wild-type Col-0, AtNHR2B-GFP, HopG1-FLAG, and AtNHR2B-GFP/HopG1-FLAG were syringe inoculated with Pstab at a concentration of 1 × 106 CFU/ml. HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG were sprayed with 20 μM DEX 24 h prior bacterial inoculation. Leaf samples were collected at 0 and 3 dpi. Bars represent means of CFU/cm2. Different letters above bars represent statistically significant difference in bacterial growth among plant genotypes using a one-way ANOVA and Tukey’s multiple comparison test with P-value ≤ 0.05. All above experiments were repeated three times with similar results.
Consistent with the activities of HopG1 suppressing PTI and the inability of AtNHR2B to interfere with HopG1-mediated PTI suppression, transgenic plants expressing HopG1-FLAG alone or in combination with AtNHR2B-GFP supported 10-fold higher growth of Pstab at 3 dpi in comparison with Pstab growth in Col-0, or in plants expressing AtNHR2B-GFP (Fig. 6b). Taken together, these results demonstrate that in Arabidopsis, HopG1 interferes with plant defense responses even when AtNHR2B-GFP is overexpressed. Moreover, the increased bacterial growth and lack of callose deposition, suggest that similar to the results obtained in N. benthamiana, the HopG1-mediated cell death is not related to the HR but to a virulence mechanism related to HopG1 function.
Discussion
Pst DC3000 deploys a plethora of effectors into the plant cell to interfere with plant defense responses, alter cellular processes and promote bacterial parasitism6,7. The Pst DC3000 T3E HopG1 appears to be of paramount importance in the pathogenicity of Pst DC3000 by being one of five effectors constituting the minimal repertoire that makes Pst DC3000ΔhopQ1-1 pathogenic in N. benthamiana4,23. Previous studies also demonstrated that HopG1 suppresses the HR in N. benthamiana based on experimental data showing that only the Pst DC3000 hopG1 mutant triggered the HR, whereas the wild type Pst DC3000, or the Pst DC3000 hopG1 (phopG1) complementation strain did not26. The elicitation of the HR by the Pst DC3000 hopG1 mutant, suggests that the absence of HopG1 uncovered the HR eliciting activity of other effectors. In contrast, the presence of HopG1 in wild type Pst DC3000, and in the Pst DC3000 hopG1 (phopG1) complementation strain, suppresses the HR. Furthermore, transient expression of the cell death inducer BAX136 in N. benthamiana triggered cell death, as expected, but this cell death was not observed when BAX1 was transiently co-expressed with HopG126.
In addition to suppressing the HR, transgenic expression of HopG1-HA in Arabidopsis also led to suppression of PTI, observed as a reduction in callose deposition after infiltration with the Flg 21 peptide, or after inoculation with the non-pathogen Pseudomonas fluorescens24. We confirmed these results by showing that Arabidopsis plants expressing HopG1-FLAG and inoculated with the non-adapted pathogen Pstab are also devoid of callose deposits.
Other previous observations on the virulence function of HopG1, revealed its role in the remodeling of the cytoskeleton demonstrated to occur through the interaction of HopG1 with the mitochondrial-localized kinesin motor protein27. The mitochondrial localization of HopG1 previously led to the hypothesis that HopG1 alters mitochondrial function, and in support of that hypothesis, Arabidopsis plants expressing HopG1-HA showed reduced oxygen consumption and enhanced ROS levels24. However, that study used the ROS sensitive probe H2DCFDA (2′-7′-dichlorodihydrofluorescein), that does not discriminate among the multiple sources of ROS. In this study, we used a mitochondria-specific fluorogenic ROS sensor and defined more precisely that the higher levels of ROS in HopG1-FLAG transgenic lines are from mitochondrial origin. Thus, our results provide stronger evidence that HopG1-FLAG expression and localization to mitochondria actually induces the production of mitochondrial ROS that likely activate a cell death program.
In this study, we uncovered that indeed HopG1 induces necrotic cell death by transient expression in N. benthamiana and by stable expression in Arabidopsis. Previously, Agrobacterium-mediated transient expression of Pst DC3000 HopG1 in 59 plant accessions that included N. benthamiana, revealed that this effector caused cell death in a limited number of plants (ca 5)37. Such cell death phenotype was explained as a recognition by the plant (HR) given that the effector was from a non-adapted pathogen36. However, a Pst DC3000 strain deleted of all effectors and only harboring HopG1 failed to elicit the HR in N. benthamiana38, demonstrating that HopG1 by itself is unable to cause the HR. Our data supports the latter findings and further demonstrates that actually, the cell-death phenotype observed is the result of the virulence function of HopG1.
The cell death phenotypes that effectors trigger can be difficult to assess as they might correspond to completely opposite outcomes: the HR, a plant defense response that restricts pathogen proliferation, or necrosis, a virulence function of the pathogen that sustains pathogen proliferation. Thus, to resolve if the HopG1-dependent cell death was HR or virulence-related necrosis, we inoculated the HopG1-FLAG-expressing N. benthamiana plants with its adapted pathogen Pstab to evaluate bacterial growth as evidence of bacterial virulence. We showed that plants expressing HopG1-FLAG sustained higher levels of Pstab in comparison with plants not expressing HopG1-FLAG.Similarly, transgenic Arabidopsis plants expressing HopG1-FLAG enabled the growth of Pstab which is a non-adapted pathogen in Arabidopsis. Because Pstab do not encode HopG139, and HopG1 in very important in the pathogenesis of Pst DC3000, these results demonstrate that transient or stable expression of HopG1-FLAG in plants supplements the deficiency of HopG1 in Pstab making it a better pathogen in both its host plant as well as in a non-host plant.
We also uncovered a functional relationship between HopG1 and AtNHR2B based on the following lines of evidence: (1) Atnhr2b mutation restores the growth deficiency of the Pst DC3000 ΔhopG1 mutant; (2) HopG1 directly interacts with AtNHR2B in yeast and in planta; (3) AtNHR2B suppresses the necrotic cell death induced by HopG1 and (4) AtNHR2B degrades HopG1. The results indicate that HopG1 and AtNHR2B counteract each other with HopG1 inducing necrotic cell death and AtNHR2B attenuating HopG1-induced necrosis. This interplay is clear with the results showing that in N. benthamiana transient expression of HopG1-FLAG enhances the virulence of Pstab increasing its populations, whereas co-expression of HopG1-FLAG and AtNHR2B-GFP in N. benthamiana effectively reduces Pstab populations. Intriguingly, in Arabidopsis, HopG1-mediated cell-death related phenotypes are attenuated by AtNHR2B, while callose deposition and Pstab growth are not. Altogether, these results would indicate a specific function of AtNHR2B attenuating necrotic cell death possibly through targeted degradation of HopG1. More studies are needed to fully understand the interplay between HopG1 and AtNHR2B.
Collectively, our results and the results of others highlight that HopG1 is a multi-faceted protein that can suppress the HR, interfere with plant defense responses and cause necrotic cell death associated with chlorotic symptoms. All these phenotypes associated with HopG1 highlight its function early in infection, suppressing immune responses, and later in infection triggering necrotic cell death. The functions of HopG1 are consistent with the hemi-biotrophic lifestyle of Pst DC3000 that combines an early biotrophic phase suppressing cell death responses with a late necrotrophic phase inducing cell death.
Materials and methods
Bacterial strains
Wild-type Pstab, Pst DC3000 and Pst DC3000ΔhopG1 mutant were grown on King’s B (KB) medium supplemented with rifampicin (25 μg/ml). Agrobacterium tumefaciens strains were grown in Luria‐Bertani (LB) medium supplemented with rifampicin (25 μg/ml) and kanamycin (50 µg/ml).
Pstab-GFP-TIR was generated by introducing the plasmid pGFP-TIR34 into Pstab by electroporation. Pstab-GFP-TIR was grown on KB medium supplemented with gentamycin (20 µg/ml). All the strains were grown at 28 °C. Bacterial strains used in this study are listed in Supplementary Table S2.
Plant materials and growth conditions
Arabidopsis thaliana seeds were planted in soil for two weeks, transplanted to individual pots and grown for four more weeks. Plants were grown in growth chambers at 21 °C with an 8/16 h light/dark cycles. Nicotiana benthamiana seeds were sown in soil, transplanted after 2 weeks and grown for 4 weeks under growth chamber conditions at 25 °C with a 10/14 h light/dark cycle.
Transgenic lines expressing HopG1-FLAG under the expression of the glucocorticoid promoter were obtained from Dr. Jim Alfano (University of Nebraska, Lincoln). HopG1-FLAG plants were crossed with lines expressing AtNHR2B-GFP to generate AtNHR2B-GFP/HopG1-FLAG transgenic lines.
This research was conducted in compliance with all relevant institutional, national, and international guidelines and legislation for handling of plant materials.
Plasmid constructs
pENTR/SD:HopG138 was transferred into the yeast bait vector pDEST32 through an LR reaction of Gateway Cloning (Thermo Fisher Scientific, Waltham, MA) to generate a transcriptional fusion to the GAL4-DNA binding domain. AtNHR2B in the entry vector pDONR201 was cloned into the yeast prey vector pDEST22 to generate a fusion to the GAL4 activation domain.
Transient expression in N. benthamiana
Agrobacterium tumefaciens GV2260 harboring constructs of interest were induced and infiltrated into fully expanded leaves of 3-week old N. benthamiana plants using a needle-less syringe as previously described31. Infiltrated leaves were used to visualize protein localization or in situ protein–protein interaction by laser scanning fluorescence microscopy, or to identify proteins of interest by Western blot.
Western blot
Nicotiana benthamiana transiently expressing proteins of interest were collected in liquid nitrogen. Tissue was ground in liquid nitrogen and homogenized with protein extraction buffer (50 mM Hepes-pH 7.5, 250 mM sucrose, 10 mM EDTA, 5% glycerol, 50 mM sodium pyrophosphate, 25 mM sodium fluoride, 1 mM sodium molybdate, 3 Mm DTT, 1 mM PMSF and plant protease inhibitor cocktail) (Thermo Scientific)40. Arabidopsis plants expressing AtNHR2B-GFP were harvested in liquid nitrogen and homogenized with protein extraction buffer (100 mM Tris–HCl, pH 7.5, 150 mM NaCl, 1 mM EDTA, 10 mM MgCl2 0.20% NP40, 0.1% SDS, 5 mM DTT, 10% glycerol, 1 mM phenylmethylsulfonyl fluoride (PMSF) and plant protease inhibitor cocktail) (Thermo Scientific).
The concentrations of protein in the supernatant were determined by using the Bio-Rad protein assay reagent (Bio-Rad). Samples, typically 30–50 mg, were separated on 10% acrylamide containing SDS-PAGE gels (mini protean; Bio-Rad) and transferred to polyvinylidene fluoride membrane (GE Healthcare). Membranes were incubated with appropriate antibodies: anti-GFP (1:2,000 dilution; Invitrogen), Anti-FLAG (A5892, Sigma) and anti-HA antibody (H6908, Sigma), were used at dilution of 1:1,000. Rabbit antisera (GE Healthcare) was used as secondary antibody with 1:5,000 dilution. Chemiluminescent detection was done by using the ImageQuant™ LS-500 image system (GE Healthcare).
Live-cell imaging
Nicotiana benthamiana plants transiently expressing AtNHR2B-GFP, co-expressing AtNHR2B-nEYFP with HopG1-cEYFP, or AtNHR2BΔ1-195nEYP with HopG1-cEYFP were used to evaluate AtNHR2B-GFP localization, or AtNHR2B/HopG1 interaction by bimolecular fluorescence complementation (BiFC), respectively. GFP and YFP signals were imaged on a Leica SP2 or Leica Stellaris 8 Laser Scanning Confocal Microscope (Leica Microsystems, Buffalo Grove, IL) at 48hpi. Arabidopsis plants expressing AtNHR2B-GFP were also evaluated by laser scanning fluorescence microscopy. GFP and YFP fluorescence was imaged using excitation wavelength of 488 nm and an emission wavelength of 500 to 535 nm.
Bacterial inoculation into Arabidopsis thaliana plants
Five-week-old Arabidopsis thaliana plants, genotypes wild-type Col-0, Atnhr2b mutant, or expressing AtNHR2B-GFP, HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG were syringe-inoculated with PstDC3000, PstDC3000 (ΔhopG1) or Pstab at different concentrations depending on the experiment.
Bacterial multiplication assays
Leaf disks (0.5 cm2) from inoculated plants were collected, serially-diluted and plated as previously described31. Each experiment was repeated three times.
Yeast two-hybrid assay
The yeast strain Mav203 was grown in YPDA at 30 °C overnight with constant shaking. The OD600 of overnight grown cultures was measured, diluted to an OD600 of 0.4 in a final volume of 50 ml of YPAD and grown for additional 3 h. Yeast cells in mid-log phase were co-transformed with pDEST32/pDEST22::AtNHR2B or pDEST32::HopG1/ pDEST22::AtNHR2B using the Frozen-EZ Yeast Transformation II Kit (Zymo Research, Irvine, CA). Transformed yeast cells were plated on Double Drop Out (DDO, SD/-Leu/-Trp) selection plates and grown at 30 °C for 4 days. Single colonies were picked from the plates and cultured in 15 mL DDO broth at 30 °C overnight. The overnight culture was diluted to an OD600 of 0.2 and plated on triple dropout medium (TDO, SD/-His/-Leu/-Trp) containing 15 mM 3-Amino-1,2,4-Triazole (3-AT) and grown at 30 °C for 4 days.
Co-immunoprecipitation
Four-week-old N. benthamiana plants were co-infiltrated with two strains of A. tumefaciens strains harboring AtNHR2B-GFP and HopG1-FLAG, or A. tumefaciens strains harboring AtNHR2B-GFP and 35S-GFP. Infiltrated leaves were collected at 2 dpi and tissue, ground in liquid nitrogen and homogenized in 1 mL of co-immunoprecipitation extraction buffer (100 mM Tris–HCl, pH 7.5, 150 mM NaCl, 1 mM EDTA, 10 mM MgCl2, 10% Glycerol, 0.2% Nonidet P-40, 1 mM PMSF, 5 mM DTT, 1X Proteinase inhibitor cocktail (Sigma Aldrich, St. Louis, MO). Protein extracts were incubated for 30 min on ice and centrifuged at 4 °C for 30 min at 13,000 rpm. Supernatants containing extracted proteins were collected in a pre-chilled 1 ml Eppendorf tube. Protein concentration was measured by Bradford Assay (BioRad, Hercules, CA) and protein expression in input samples was confirmed by Western blot as described above.
One milligram of total protein extract was mixed with 25 µl of GFP Nanobody/VHH coupled to agarose beads (GFP-Trap® Agarose, Chromotek) and incubated for 3 h and 30 min at 4 °C with end to end rocking. After incubation, protein complexes bound to beads were washed once with 1 × TBS buffer (50 mM Tris–HCl, 150 mM NaCl, pH 7.5) and twice with 1 × TBS buffer (50 mM Tris–HCl, 500 mM NaCl, pH 7.5). Protein complexes bound to the beads were eluted in 2 × SDS protein loading buffer, loaded and ran into an SDS-PAGE gel and transferred to PVDF membranes. Proteins were detected by Western Blot as described above.
Cell death assay
Five-week old wild-type Col-0, AtNHR2B-GFP, HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG plants were syringe-inoculated with Pstab at OD600 = 0.02 (1 × 106 CFU/ml). Control plants were inoculated with water only. At 24 hpi, six to nine inoculated leaves were detached from six independent plants for each genotype. Collected leaves were stained with 0.05% trypan blue for 45 min at room temperature and washed twice with PBS35. Images were taken on a light microscope using bright field.
Callose deposition
Five-week old wild-type Col-0, AtNHR2B-GFP, HopG1-FLAG and HopG1-FLAG/AtNHR2B-GFP were sprayed with 20 µM dexamethasone to induce expression of HopG1-FLAG. At 24 h after dexamethasone treatment, plants were syringe-inoculated with Pstab at OD600 = 0.02 (1 × 106 CFU/ml). Ten leaves from six independent plants for each genotype and inoculated with Pstab or infiltrated with water were detached after 24 hpi and stained with 5% aniline blue to visualize callose deposits. Images were taken by Nikon 90i upright scanning laser confocal microscope (Nikon) using a DAPI (4′,6-diamidino-2-phenylindole) filter with excitation wavelength of 405 nm and an emission wavelength of 450–510 nm.
Electrolyte leakage assay
Five- week old wild-type Col-0, AtNHR2B-GFP, HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG plants were induced by dexamethasone prior to inoculation with Pstab at OD600 = 0.02 (1 × 106 CFU/ml). Twelve leaves were collected immediately after infiltration and 0.5 cm2 leaf disks were excise and rinsed with 50 ml for 30 min, to wash ions released during disc excision. After 30 min, water was replaced with 12 ml of fresh deionized water and conductivity was measured at 5, 10, and 20 hpi using the Orion Star A215 conductivity cell (013005MD) (Thermo Fisher Scientific).
Mitochondrial ROS production
Five-week old wild-type Col-0, AtNHR2B-GFP, HopG1-FLAG and AtNHR2B-GFP/HopG1-FLAG plants grown on soil were sprayed with a 20 µM dexamethasone solution supplemented with 0.01% silwet, 12 h before inoculation. Leaf disks cut out using a 1.2 cm2 core-borer and harvested tissue was transferred to clear bottom plates for fluorometric analysis and submerged in Pstab inoculum at a final concentration of 1 × 107 CFU/ml. For mock-treatment, plants were submerged in water. Two hours after inoculation with either Pstab mock-treatment, MitoTracker Red CM-H2XRos (ThermoFisher Scientific, Waltham, MA) was added at a final concentration of 0.005 mM and incubated for 10 min before taking the first reading. Fluorescence was measured with an excitation wavelength of 570 nm and an emission wavelength of 535 nm on BioTek luminescence microplate reader.
References
Xin, X. F., Kvitko, B. & He, S. Y. Pseudomonas syringae: What it takes to be a pathogen. Nat. Rev. Microbiol. 16, 316–328. https://doi.org/10.1038/nrmicro.2018.17 (2018).
Xin, X. F. & He, S. Y. Pseudomonas syringae pv. tomato DC3000: A model pathogen for probing disease susceptibility and hormone signaling in plants. Annu. Rev. Phytopathol. 51, 473–498. https://doi.org/10.1146/annurev-phyto-082712-102321 (2013).
Wei, H. L. & Collmer, A. Defining essential processes in plant pathogenesis with Pseudomonas syringae pv. tomato DC3000 disarmed polymutants and a subset of key type III effectors. Mol. Plant Pathol. 19, 1779–1794. https://doi.org/10.1111/mpp.12655 (2018).
Wei, C. F. et al. A Pseudomonas syringae pv. tomato DC3000 mutant lacking the type III effector HopQ1-1 is able to cause disease in the model plant Nicotiana benthamiana. Plant J. 51, 32–46. https://doi.org/10.1111/j.1365-313X.2007.03126.x (2007).
Collmer, A. et al. Pseudomonas syringae Hrp type III secretion system and effector proteins. Proc. Natl. Acad. Sci. USA 97, 8770–8777. https://doi.org/10.1073/pnas.97.16.8770 (2000).
Lindeberg, M., Cunnac, S. & Collmer, A. Pseudomonas syringae type III effector repertoires: Last words in endless arguments. Trends Microbiol. 20, 199–208. https://doi.org/10.1016/j.tim.2012.01.003 (2012).
Cunnac, S., Lindeberg, M. & Collmer, A. Pseudomonas syringae type III secretion system effectors: Repertoires in search of functions. Curr. Opin. Microbiol. 12, 53–60. https://doi.org/10.1016/j.mib.2008.12.003 (2009).
Buttner, D. & He, S. Y. Type III protein secretion in plant pathogenic bacteria. Plant Physiol. 150, 1656–1664. https://doi.org/10.1104/pp.109.139089 (2009).
Schechter, L. M. et al. Multiple approaches to a complete inventory of Pseudomonas syringae pv. tomato DC3000 type III secretion system effector proteins. Mol. Plant Microbe Interact 19, 1180–1192 (2006).
Petnicki-Ocwieja, T. et al. Genomewide identification of proteins secreted by the Hrp type III protein secretion system of Pseudomonas syringae pv. tomato DC3000. Proc. Natl. Acad. Sci. USA 99, 7652–7657 (2002).
Lindeberg, M. et al. Closing the circle on the discovery of genes encoding Hrp regulon members and type III secretion system effectors in the genomes of three model Pseudomonas syringae strains. Mol. Plant–Microbe Interact. MPMI 19, 1151–1158. https://doi.org/10.1094/MPMI-19-1151 (2006).
Toruno, T. Y., Stergiopoulos, I. & Coaker, G. Plant–pathogen effectors: Cellular probes interfering with plant defenses in spatial and temporal manners. Annu. Rev. Phytopathol. 54, 419–441. https://doi.org/10.1146/annurev-phyto-080615-100204 (2016).
Macho, A. P. Subversion of plant cellular functions by bacterial type-III effectors: Beyond suppression of immunity. New Phytol. 210, 51–57. https://doi.org/10.1111/nph.13605 (2016).
Wilton, M. & Desveaux, D. Lessons learned from type III effector transgenic plants. Plant Signal Behav. 5, 746–748. https://doi.org/10.1073/pnas.0904739107 (2010).
Jones, J. D. & Dangl, J. L. The plant immune system. Nature 444, 323–329. https://doi.org/10.1038/nature05286 (2006).
Chisholm, S. T., Coaker, G., Day, B. & Staskawicz, B. J. Host-microbe interactions: Shaping the evolution of the plant immune response. Cell 124, 803–814. https://doi.org/10.1016/j.cell.2006.02.008 (2006).
Zipfel, C. et al. Bacterial disease resistance in Arabidopsis through flagellin perception. Nature 428, 764–767 (2004).
Cui, H., Tsuda, K. & Parker, J. E. Effector-triggered immunity: From pathogen perception to robust defense. Annu. Rev. Plant Biol. 66, 487–511. https://doi.org/10.1146/annurev-arplant-050213-040012 (2015).
Van Doorn, W. et al. Morphological classification of plant cell deaths. Cell Death Differ. 18, 1241–1246 (2011).
Dangl, J. L. & Jones, J. D. Plant pathogens and integrated defence responses to infection. Nature 411, 826–833 (2001).
Van Der Biezen, E. A. & Jones, J. D. Plant disease-resistance proteins and the gene-for-gene concept. Trends Biochem. Sci. 23, 454–456 (1998).
Kvitko, B. H. et al. Deletions in the repertoire of Pseudomonas syringae pv. tomato DC3000 type III secretion effector genes reveal functional overlap among effectors. PLoS Pathog. 5, e1000388 (2009).
Cunnac, S. et al. Genetic disassembly and combinatorial reassembly identify a minimal functional repertoire of type III effectors in Pseudomonas syringae. Proc. Natl. Acad. Sci. USA 108, 2975–2980. https://doi.org/10.1073/pnas.1013031108 (2011).
Block, A. et al. The Pseudomonas syringae type III effector HopG1 targets mitochondria, alters plant development and suppresses plant innate immunity. Cell. Microbiol. 12, 318–330. https://doi.org/10.1111/j.1462-5822.2009.01396.x (2010).
Guo, M., Tian, F., Wamboldt, Y. & Alfano, J. R. The majority of the type III effector inventory of Pseudomonas syringae pv. tomato DC3000 can suppress plant immunity. Mol. Plant Microbe Interact 22, 1069–1080 (2009).
Jamir, Y. et al. Identification of Pseudomonas syringae type III effectors that can suppress programmed cell death in plants and yeast. Plant J. 37, 554–565 (2004).
Shimono, M. et al. The Pseudomonas syringae type III effector HopG1 induces actin remodeling to promote symptom development and susceptibility during infection. Plant Physiol. 171, 2239–2255. https://doi.org/10.1104/pp.16.01593 (2016).
Heath, M. C. Nonhost resistance and nonspecific plant defenses. Curr Opin Plant Biol 3, 315–319 (2000).
Panstruga, R. & Moscou, M. J. What is the molecular basis of nonhost resistance?. Mol. Plant Microbe Interact 33, 1253–1264. https://doi.org/10.1094/MPMI-06-20-0161-CR (2020).
Senthil-Kumar, M. & Mysore, K. S. Nonhost resistance against bacterial pathogens: Retrospectives and prospects. Annu. Rev. Phytopathol. 51, 407–427. https://doi.org/10.1146/annurev-phyto-082712-102319 (2013).
Singh, R. et al. Two chloroplast-localized proteins: AtNHR2A and AtNHR2B, contribute to callose deposition during nonhost disease resistance in Arabidopsis. Mol. Plant Microbe Interact 31, 1280–1290. https://doi.org/10.1094/MPMI-04-18-0094-R (2018).
Singh, R. et al. The arabidopsis proteins AtNHR2A and AtNHR2B are multi-functional proteins integrating plant immunity with other biological processes. Front. Plant Sci. 11, 232. https://doi.org/10.3389/fpls.2020.00232 (2020).
Martin, K. et al. Transient expression in Nicotiana benthamiana fluorescent marker lines provides enhanced definition of protein localization, movement and interactions in planta. Plant J. 59, 150–162. https://doi.org/10.1111/j.1365-313X.2009.03850.x (2009).
Miller, W. G. & Lindow, S. E. An improved GFP cloning cassette designed for prokaryotic transcriptional fusions. Gene 191, 149–153. https://doi.org/10.1016/s0378-1119(97)00051-6 (1997).
Kabbage, M., Williams, B. & Dickman, M. B. Cell death control: The interplay of apoptosis and autophagy in the pathogenicity of Sclerotinia sclerotiorum. PLoS Pathog. 9, 1. https://doi.org/10.1371/journal.ppat.1003287 (2013).
Baek, D. et al. Bax-induced cell death of Arabidopsis is meditated through reactive oxygen-dependent and -independent processes. Plant Mol. Biol. 56, 15–27. https://doi.org/10.1007/s11103-004-3096-4 (2004).
Wroblewski, T. et al. Comparative large-scale analysis of interactions between several crop species and the effector repertoires from multiple pathovars of Pseudomonas and Ralstonia. Plant Physiol. 150, 1733–1749. https://doi.org/10.1104/pp.109.140251 (2009).
Wei, H. L., Zhang, W. & Collmer, A. Modular study of the type III effector repertoire in Pseudomonas syringae pv. tomato DC3000 reveals a matrix of effector interplay in pathogenesis. Cell Rep. 23, 1630–1638. https://doi.org/10.1016/j.celrep.2018.04.037 (2018).
Baltrus, D. A. et al. Dynamic evolution of pathogenicity revealed by sequencing and comparative genomics of 19 Pseudomonas syringae isolates. PLoS Pathog. 7, e1002132. https://doi.org/10.1371/journal.ppat.1002132 (2011).
Heese, A. et al. The receptor-like kinase SERK3/BAK1 is a central regulator of innate immunity in plants. Proc. Natl. Acad. Sci. USA 104, 12217–12222. https://doi.org/10.1073/pnas.0705306104 (2007).
Kvitko, B. H. et al. Deletions in the repertoire of Pseudomonas syringae pv. tomato DC3000 type III secretion effector genes reveal functional overlap among effectors. PLoS Pathog. 5, 16. https://doi.org/10.1371/journal.ppat.1000388 (2009).
Acknowledgements
We thank Gitta Coaker and James M. Elmore for providing the HopG1-FLAG and HopG1-HA constructs, Jim Alfano for providing the HopG1-FLAG seeds provided and Brian Kvitko for providing the Pst DC3000 ΔhopG1 mutant. We also thank Maria Emilia Aldana for providing assistance with statistical analyses.
Funding
This work was supported by the National Science Foundation CAREER award Grant (IOS # 1842970) and by Equipment Grants Program, award #2021-05017, from the US Department of Agriculture, National Institute of Food and Agriculture.
Author information
Authors and Affiliations
Contributions
C.M.R., R.S., C.R.P. conceived and designed the experiments. Performed the experiments: C.R.P., R.S., R.C., P.R.L. performed experiments. C.M.R., C.R.P., R.S., R.C. analyzed the data. C.M.R., C.R.P. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rodríguez-Puerto, C., Chakraborty, R., Singh, R. et al. The Pseudomonas syringae type III effector HopG1 triggers necrotic cell death that is attenuated by AtNHR2B. Sci Rep 12, 5388 (2022). https://doi.org/10.1038/s41598-022-09335-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-09335-1
- Springer Nature Limited
This article is cited by
-
Rational design of ROS scavenging and fluorescent gold nanoparticles to deliver siRNA to improve plant resistance to Pseudomonas syringae
Journal of Nanobiotechnology (2024)
-
Maintenance of tRNA and elongation factors supports T3SS proteins translational elongations in pathogenic bacteria during nutrient starvation
Cell & Bioscience (2022)