Abstract
Leaves and flowers of Amorphophallus konjac do not develop simultaneously thus unique features can be elucidated through study of flowering transformation in A. konjac. In this study, transcriptome libraries of A. konjac leaf buds (LB) and flower buds (FB) were constructed followed by high-throughput sequencing. A total of 68,906 unigenes with an average length of 920 bp were obtained after library assembly. Out of these genes, 24,622 unigenes had annotation information. A total of 6859 differentially expressed genes (DEGs) were identified through differential expression analysis using LB as control. Notably, 2415 DEGs were upregulated whereas 4444 DEGs were downregulated in the two transcriptomes. Go and KEGG analysis showed that the DEGs belonged to 44 functional categories and were implicated in 98 metabolic pathways and 38 DEGs involved in plant hormone signal transduction. Several genes were mined that may be involved in A. konjac flower bud differentiation and flower organ development. Eight DEGs were selected for verification of RNA-seq results using qRT-PCR analysis. Two FLOWERING LOCUS T (FT) genes named AkFT1 and AkFT2 were identified though homologous analysis may be the florigen gene implicated in modulation of A. konjac flowering. These genes were significantly upregulated in flower buds compared with the expression levels on leaf buds. Overexpression of AkFT genes though heterologous expression in Arabidopsis showed that the transgenics flowered at a very early stage relative to wild type plants. These findings indicate that AkFT1 and AkFT2 function as regulation genes in A. konjac flowering development and the two genes may present similar functions during flowering transition.
Similar content being viewed by others
Introduction
Amorphophallus konjac is a perennial plant and a member of Araceae family. Approximately 163 species of Amorphophallus plants have been identified and are widely distributed in China, Japan and Southeast Asia1. Amorphophallus corm is characterized by large amounts of glucomannan which is often regarded as salutary dietary polysaccharide used in food, medical, health care and other industries2. Amorphophallus plants are mainly planted in the mountains and hills as a food source and do not compete with cereal crops for land3. Amorphophallus plants are propagated through asexual methods using mini corm or corm cuttings4,5. Asexual propagation is associated with low propagation coefficient, high susceptibility to soft rot and other diseases6. Seed propagation is generally considered a better way for crop reproduction. However, A. konjac like most Amorphophallus species takes more than three years to bloom and they are self-incompatible. Therefore, the seed setting rate in the field is low7,8,9. Furthermore, A. konjac exhibits a unique flower development process. Germination of the corm either results in a vegetative leaf or a reproductive flower thus the leaf and flower do not occur concurrently during its growing years. Study of flowering related genes and exploring the molecular mechanism of flowering is important for developing approaches to shorten the flowering time by manipulating flowering-related genes of A. konjac thus improving the reproductive efficiency.
Molecular biology research of A. konjac is currently limited10,11,12. Lack of reference genome sequence significantly hinders gene mining and molecular breeding of A. konjac. Transcriptome analysis is an easier approach to obtain genetic information or expression profiles of genes. Currently, the flowering mechanism of several flowering plants has been preliminarily analyzed through transcriptome analysis, and high amounts of gene information have been obtained. Comparative transcriptome analysis was used to explore various transcription factor families and metabolic pathways involved in flower development in Cicer arietinum13, Hypericum perforatum L.14. Jatropha curcas15 and Wucai (Brassica campestris L.)16. However, the molecular mechanism of flower formation and flowering related genes in A. konjac has not been elucidated.
FLOWERING LOCUS T (FT) is one of the key genes in the regulation of flowering in plants17. FT is expressed mainly in leaves and is transported via the phloem to the apical meristem tissue of the stem tip to function in Arabidopsis thaliana18. FT can integrate signals from various flowering regulatory pathways, including endogenous factors and environmental conditions, to regulate the timing of flowering in plants19. Previous studies have shown that plant FT genes in A. thaliana, rice, wheat, and maize have a role in regulating flowering time20,21,22. In addition, FT genes can also affect A. thaliana seed development, control bulb formation in onion and regulate potato storage organ formation23,24,25.
In this study, Illumina Hiseq high-throughput sequencing technology was used to sequence the transcriptome of A. konjac leaf buds and flower buds. The findings from differential expression analysis showed that several genes are implicated in flowering. Two flowering related genes highly homologous to FT gene were identified in A. konjac and were named AkFT1 and AkFT2, respectively. Overexpression of the two genes in A. thaliana through heterologous transformation significantly accelerated flowering in the transgenic plants relative to the wild type plants. These results provide a basis for further study on the molecular mechanism of A. konjac flower development process.
Materials and methods
Materials
Amorphophallus konjac K.Koch plants were grown in Sangzhi (Zhangjiajie, China) under natural conditions. RNA was extracted from leaf primordium and flower primordium for transcriptome analysis.
Arabidopsis thaliana ecotype Columbia (sustained in our laboratory) was used as the model plant for gene function analysis. Candidate genes were transformed into Arabidopsis through inflorescence infiltration method. Arabidopsis seeds were surface-sterilized, germinated on a plate containing Murashige and Skoog (MS) medium with 2% (w/v) sucrose and 0.75% (w/v) agar supplemented with 30 mg/L basta to select transgenic plants. Plants were transferred into pots containing a mixture of topsoil and vermiculite (3:1). The plants were then grown in a growth chamber at 25 °C under a 16-h light/8-h dark photoperiod.
Construction, sequencing and analysis of cDNA library
Total RNA was extracted from buds using HiPure HP Plant RNA Kit (Magen, China). cDNA libraries were prepared using NEBNext®Ultra™ RNA Library Prep Kit for Illumina® (NEB, USA). 1 μg RNA per sample was used as input material for preparation of libraries. The libraries were sequenced using an Illumina Hiseq 2000 platform at Biomarker Technologies Co, LTD (Beijing, China) and paired-end reads of 2 × 100 bp were generated.
Raw data in fastq format were firstly processed using in-house Perl scripts. Clean data were obtained by removing reads containing adapters and low-quality reads from the raw data. Transcriptome assembly was performed using Trinity software with min_kmer_cov set to 2 by default and all other parameters set default.
Assembled unigenes were annotated using the following databases: GO (http://www.geneontology.org), KEGG (http://www.genome.jp/kegg/)26, COG (http://www.ncbi.nlm.nih.gov/COG/), KOG (http://www.ncbi.nlm.nih.gov/KOG/), eggNOG4.5 (http://eggnogdb.embl.de/), Swiss-Prot (http://www.uniprot.org/), Pfam (http://pfam.xfam.org/) and nr database (ftp://ftp.ncbi.nih.gov/blast/db/) using BLAST tool with e-value < 1e−5.
Differentially expressed genes in the two libraries were identified using DESeq2 R package (https://bioconductor.org/packages/release/bioc/html/DESeq2.html). p-value corrected by Benjamini–Hochberg method27, “p-value < 0.01 and fold change ≥ 2” were used as the criteria for screening differentially expressed genes (DEGs). GO enrichment analysis was achieved by the topGO R package based on the Kolmogorov–Smirnov test. KEGG pathway enrichment was performed by KOBAS 2.0 software with FDR ≤ 0.05.
Identification and cloning of FT genes from A. konjac
FT genes were retrieved from unigene annotation of the two cDNA libraries derived from A. konjac buds. Total RNA was extracted from A. konjac using HiPure HP Plant RNA Kit (Magen, China). RNA samples were reverse transcribed to cDNA using ReverTra Ace® qPCR RT Master Mix (Toyobo, Japan). Full-length coding sequences of A. konjac FT genes were amplified with Golden Star T6 Super PCR Mix (TsingKe, China). The thermocycling conditions were as follows: denaturation of cDNA at 98 °C for 2 min, 32 cycles of 98 °C for 10 s, 60 °C for 10 s, and 72 °C for 5 s. PCR products were cloned into pClone007 Versatile Simple Vector (TsingKe, China) and sequenced at TsingKe Technologies Co, LTD. Phylogenetic analysis was performed using MEGA 7 software28. Conserved motifs in protein sequence were identified using MEME tool (http://meme-suite.org/) with default parameters29.
Preparation of AkFT1 and AkFT2 overexpression lines
AkFT1 and AkFT2 gene were cloned into pEGAD vector (sustained in our laboratory) at the AgeI and SmaI site using ClonExpress II One Step Cloning Kit (Vazyme, China). Sequencing was performed at TsingKe Technologies Co, LTD to confirm that the genes were cloned successfully. Expression of AkFT1 and AkFT2 gene was modulated by CaMV35S promoter. Arabidopsis thaliana ecotype Columbia plants were transformed using Agrobacterium tumefaciens strain, GV3101 harboring AkFT1 and AkFT2 overexpression vector using the floral dip method30.
A total of 8 individual transgenic lines carrying 35S::AkFT1 and 35S::AkFT2 were established, respectively. Among the two transgenic materials, we selected three independent transgenic lines for analysis, and at least 40 plants were selected for observation and statistical analysis.
Real‑time quantitative and semi-quantitative RT-PCR analysis
cDNA sequences of A. konjac were obtained as described above. AkEF1-α was used as the internal reference gene12 and eight DEGs genes were selected for qRT-PCR analysis. qRT-PCR was performed using ChamQ Universal SYBR qPCR Master Mix (Vazyme, China). The PCR program was: 95 °C for 30 s, then 40 cycles of 95 °C for 10 s and 60 °C for 30 s. All amplification reactions were performed in three replicates. Relative expression of each gene was determined by 2−∆∆CT method31.
Total RNA was extracted from A. thaliana using TRIzol Reagent (CWBIO, China). RNA was then reverse transcribed to obtain cDNA. Further, 2 × Taq Master Mix (Dye Plus) (Vazyme, China) was used for semi-quantitative RT-PCR analysis using the following thermocycling conditions: denaturation of cDNA at 95 °C for 3 min, 28 cycles of 95 °C for 15 s, 60 °C for 15 s, and 72 °C for 30 s. A. thaliana Actin2 (AtACTIN2) gene was used as the control gene. Sequences of the gene-specific primer sets are presented in Table 1.
Ethics approval and consent to participate
We confirm that the collection of plant material did not involve any endangered or protected plant species and declare that the work reported here is consistent with the IUCN Policy Statement on Research Involving Species at Risk of Extinction and the current laws of China.
Results
Transcriptome analysis
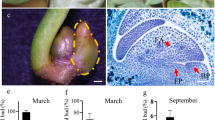
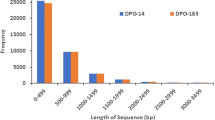
A. konjac is a unique plant with a single petiole and a compound leaf at the top whereby the leaf and flower do not appear concurrently. Therefore, vegetative or reproductive growth of A. konjac is initiated through leaf buds or flower buds (Fig. 1). Two cDNA libraries were constructed from leaf primordium (LB) and flower primordium (FB) of A. konjac to explore gene expression during the development of leaf buds and flower buds, resulting in 48.52 and 47.95 million raw reads in the two cDNA libraries, respectively. After filtering adapters and low-quality reads from the raw data, approximately 47.79 and 47.23 million clean reads were obtained in the two cDNA libraries. Further, all clean reads were de novo assembled using Trinity software32 and 68,906 unigenes were obtained from the two cDNA libraries. The average size of unigenes was 920 bp, and the N50 length of unigenes was 1403 bp (Table 2). About 89.11% of unigenes had a length ranging from 300 to 2000 bp.
Unigenes were analyzed using GO, KEGG, COG, KOG, eggNOG, Swissprot, Pfam and nr databases to identify key functions of the genes. A total of 24,622 unigenes were annotated, representing only one third of the total number of unigenes. The number of unigenes annotated in each database was then determined (Table 3). Notably, 24,246 unigenes were annotated in nr database, accounting for 35.19% of the total number of unigenes, whereas only 6739 unigenes had annotation information in COG database. Functional annotation of genes expressed in leaf buds and flower buds further enriched the gene pool of A. konjac.
Differential expression analysis of genes in leaf buds and flower buds
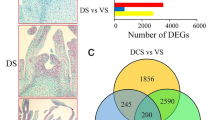
With LB as the control, 6859 significant DEGs were selected with 2415 DEGs were upregulated whereas 4444 DEGs were downregulated (Fig. 2a). Functional annotation was performed on the identified DEGs according to the expression levels of the genes in two libraries. A total of 2908 DEGs had annotation information (Table 4). Analysis showed that the nr database had the highest number of DEGs with annotated information with 2842 DEGs.
GO functional analysis showed that 1381 DEGs were enriched in 44 classes of three major categories (biological process, cell composition and molecular function) (Fig. 2b). Some DEGs (672) were annotated as metabolic processes which was the most representative class under the “biological process” category. Some DEGs (517) were annotated as “cell part”, which was the most significantly enriched term in “cellular component” category. Under “molecular function”, DEGs were mainly involved in “binding” (611) and “catalytic activity” (639) processes. TopGO software was used to explore enrichment of DEGs33. The top 10 GO terms with the significant enrichment of DEGs are presented in Table 5. The top 3 GO terms included DNA integration, structural constituent of ribosome and RNA-dependent DNA biosynthetic process.
Moreover, 862 DEGs were annotated into 98 metabolic pathways in KEGG pathway analysis (Fig. 2c). The most significantly enriched pathways were ribosome (80 DEGs), plant hormone signal transduction (38 DEGs) and phenylpropanoid biosynthesis (38 DEGs). Some DEGs (38) were implicated in plant hormone signal transduction and may be involved in flower bud differentiation and floral organ development of A. konjac, and studies should further explore their functions. These metabolic pathways provide a molecular foundation for studying the specific processes involved in leaf bud and flower bud development of A. konjac. KEGG pathway enrichment analysis showed that the top 20 pathways associated with high number of DEGs included ribosome, phenylpropanoid biosynthesis and the pentose and glucuronate interconversions (Fig. 2d). These metabolic pathways are mainly involved in synthesis of organic matter and energy transfer, indicating that there are some differences in material and energy requirements between leaf bud and flower bud development.
DEGs related to flowering
Genes implicated in gibberellin synthesis or flowering signaling pathway were significantly differentially expressed in the two transcriptomes. c75063.graph_c0 and c82483.graph_c0 are homologous to GAI gene and their expression was downregulated in flower buds. The log2 (fold change) of the two GAI genes was − 6.47 and − 6.70, respectively. c76528.graph_c0 which was homologous to GA20ox gene was highly expressed in flower buds. The log2 (fold change) of GA20ox gene was 8.26. Expression level of c63309.graph_c0 and c74067.graph_c0 which were homologous to GA2ox was lower in flower buds compared with the expression level in leaf bud. The log2 (fold change) of the two GA2ox genes was − 3.77 and − 6.21, respectively. 4 SPL homologous genes were identified from DEGs and their expression level was significantly high in flower buds relative to the expression level in leaf buds. c73015.graph_c0 and c100034.graph_c0 were homologous FT genes and had high expression level in flower buds. The log2 (fold change) of the two FT genes was 4.73 and 7.53, respectively. Nine MADS-box transcription factor genes were identified from DEGs, with 7 upregulated genes and 2 downregulated genes in flower buds. Notably, several DEGs which showed specific expression profile in flower buds were implicated in floral development.
Other phytohormonal-related gene expression differences during flowering of A. konjac
Phytohormones participate in a variety of physiological and biochemical processes, and are involved in regulation of growth and development of plants. Expression patterns of phytohormone biosynthesis and signal transduction related genes in leaf buds and flower buds were analyzed to determine the regulatory effect of other phytohormones except GA on the flowering of A. konjac. Expression of auxin biosynthesis gene, YUCCA4; auxin transporter protein gene, AUX1; auxin responsive genes IAA9, IAA10 and IAA13 was upregulated in flower buds. In addition, expression of cytokinin dehydrogenase gene, CKX5 was upregulated in flower buds compared with the expression level in leaf buds. Cytokinin response factor gene, AHP was downregulated in flower buds relative to leaf buds. ABA receptor gene, PYL4 and ABA biosynthesis related genes such as CCD8B and NCED1 were downregulated in flower buds compared with leaf buds. Ethylene (ETH) biosynthesis related genes including ACS1, ACS3 and ACS9 were upregulated. The gene that encodes the rate-limiting enzyme in Brassinosteroid (BR) biosynthesis, DET2 was upregulated. Jasmonates acid (JA) biosynthesis related gene, 4CLL6, and repressor of JA responses including TIFY9 and TIFY10A were downregulated in flower buds relative to the expression level in leaf buds. Differential expression of these genes implied that these phytohormones play complex and different roles in development of leaf and flower buds. Further studies should explore the potential regulatory mechanisms.
Verification of expression of DEGs related to flowering of A. konjac
Eight DEGs implicated in flowering, of which six genes were upregulated and two genes were downregulated, were used for verification of expression levels obtained from transcriptome analysis. The results of quantitative real time-PCR were in agreement with transcriptome analysis results (Fig. 3), indicating that the transcriptome results were reliable.
Verification of DEGs related to flowering and correlation analysis. (a) Verification of DEGs using qRT-PCR. The qPCR section was plotted with the expression of genes in leaf buds as a control and the relative expression of genes in flower buds, and the RNA-seq section was plotted with the leaf bud transcriptome as a control and the log2 (fold change) of genes. (b) Pearson correlation analysis of the expression of DEGs between qRT-PCR and RNA-Seq.
Identification and genetic analysis of FT genes in A. konjac
Two candidate genes encoding PEBP protein, c73015.graph_c0 and c100034.graph_c0, were selected from the DEGs of leaf buds and flower buds of A. konjac. Specific PCR primers were designed to amplify the two candidate genes, respectively. c73015.graph_c0 comprised an ORF with 555 bp, and prediction showed that it encoded for 184 amino acids. c100034.graph_c0 comprised an ORF with 525 bp encoding 174 amino acids. The deduced protein sequences encoded by the two candidate genes were compared with other functional FT protein sequences, through phylogenetic analysis using MEGA 7 software28 (Fig. 4). The results showed that c73015.graph_c0 clustered with ZCN8, and c100034.graph_c0 clustered with Hd3a and VRN3. This indicates that PEBP protein of A. konjac was evolutionary related to PEBP proteins from monocots such as rice, wheat and maize.
Phylogenetic analysis of FT proteins. Phylogenetic tree of FT homologous proteins using the neighbor-joining method by MEGA 7 (Bootstrap: 1000 replicates). Conservative motif analysis of FT homologous proteins using MEME. The proteins are as follows: A. thaliana AtFT, AAF03936.1; Rice Hd3a, BAO03048.1; Wheat VRN3, ABK32208.1; Glycine max GmFTL3, ACA24487.1; Maize ZCN8, NP_001106247.1; Tomato SP5G, NP_001307981.1; Potato StSP3d, BAV67096.1.
Conserved motifs in these proteins were analyzed using MEME tool29 to further explore the function of the two FT homologous proteins (Fig. 4). The findings showed that the proteins shared motifs, indicating that FT protein structure is highly conserved in plants, and they have similar functions. These two FT homologous proteins are implicated in promoting flowering of A. konjac. Notably, c73015.graph_c0 was named AkFT1 and c100034.graph_c0 was named AkFT2 according to the above results.
Functional analysis of AkFT genes expressed in A. thaliana
Phenotypic and flowering time analyses were performed on WT and transgenic plants (Fig. 5). The flowering statistics of 8 individual transgenic lines are shown in Supplementary Fig. S1 online. Semi quantitative RT-PCR results showed that AkFT1 and AkFT2 high expression levels in transgenic plants, respectively, but not in WT plants (Fig. 5a,b). Rosette leaves of the two types of transgenic plants was smaller and the vegetative growth period was shorter compared with those of WT plants (Fig. 5c,d). Flowering time of WT plants was 25–28 days, and 10–12 rosette leaves were observed during bolting. The number of rosette leaves ranged from 5 to 6 during the bloom period and the flowering time was about 16 days in the 35S::AkFT1 transgenic plants. Moreover, the 35S::AkFT2 transgenic plants took 18–20 days to bloom, and showed 6–7 rosette leaves at this period (Fig. 5e,f). These results indicated that AkFT1 and AkFT2 may play a role in promoting flowering of A. thaliana, respectively.
Overexpression of the AkFT genes in A. thaliana. (a,b) Semi-quantitative PT-PCR analysis of the AkFT genes in WT and transgenic plants. (c,d) Phenotypic observation of flowering in WT and transgenic plants. (e,f) Flowering time statistics in WT and transgenic plants. When the bolting length reached 1 cm, it was recorded as the beginning of bolting. The number of rosette leaves per plant during bolting was counted, n ≥ 40.
Discussion
A. konjac is a unique plant whereby flowers and leaves do not occur simultaneously under natural conditions. The molecular mechanism of flower transition and development in A. konjac has not been fully elucidated. Studies in A. thaliana and other model plants indicate that at least six flowering pathways are involved in molecular control of flowering transition. Plants flower transition is controlled by growth and environmental factors through photoperiod, vernalization, autonomous, thermo sensory, gibberellin and age pathways20,34,35,36,37. These pathways transmit signals that regulate expression of FT, SUPPRESSOR OF OVEREXPRESSION OF CO1 (SOC1), LEAFY (LFY) and other flowering integration factors. These factors then activate the floral meristem identity genes, such as APETALA1(AP1), FRUITFULL(FUL) and other MADS box genes thus promoting floral primordia differentiation and flower organ development38,39. FT in Arabidopsis is synthesized in the leaf during the day then it is transported into meristem and functions as a co-transcription factor19. Therefore, FT is referred as a florigen. Regulation of the duration of light when corm is in the dormancy period can affect flowering in A. bulbifer8. However, corms mainly grow under the soil, and photoperiod may not be the main pathway that affects flowering of A. konjac. Exogenous gibberellin application during corm dormancy accelerates A. muelleri blooming3,40, indicating that gibberellin pathway is implicated in regulating Amorphophallus flowering. In addition, the corm age affects its flowering pattern. In the present study, gene expression profile of A. konjac leaf buds and flower buds was explored by transcriptome analysis and several DEGs were implicated in gibberellin synthesis and signaling.
We obtained 68,906 unigenes from the leaf bud and flower bud transcriptomes, of which only 24,622 unigenes (35.73%) had functional annotation information. The absence of annotation information for some genes may be due to insufficient genome-wide information, limitations of transcriptome analysis, and incomplete information in functional databases. These genes may have unique functions in konjac and deserve further research. A total of 6859 DEGs were identified by comparing the transcriptome of A. konjac leaf buds and flower buds and 2908 DEGs had functional annotation information. Lack of annotation information of several DEGs can be attributed to lack of reference genome and incomplete database information of A. konjac. This indicates that leaf buds and flower buds in A. konjac exhibit different and unique processes and pathways. Several key genes involved in flowering and the possible regulatory pathways in A. konjac were analyzed. Gibberellin homeostasis in plants is achieved by strict regulation of the activities of “activating enzymes” (GA20ox and GA3ox) and “inactivating enzymes” (GA2ox)41,42. GA20ox showed low expression levels in leaf buds, however, its showed high expression levels in flower buds. Two GA2ox genes were significantly downregulated in flower buds compared with the expression level in leaf buds. These results indicated that synthesis and degradation of gibberellin significantly affects flowering of A. konjac. DELLA protein is a negative regulator of gibberellin signaling pathway, and a member of GAI‐RGA‐and‐SCR (GRAS) family43,44. A. thaliana expresses five DELLA proteins, including GAI, RGA (REPRESSOR OF ga1–3), RGL1 (RGA-LIKE 1), RGL2 and RGL345,46. Two GAI homologous genes were downregulated or even not expressed in flower buds, which may slow flowering of A. konjac. SPL genes regulate the flowering time of A. thaliana through DELLA-dependent and DELLA-independent pathways. Notably, interaction between SPL genes and miR156 can affect flowering through the age pathway47,48. Four SPL homologous genes showed high expression level in flower buds, implying that these genes are implicated in promoting the flowering of A. konjac. The expression product of DEHYDRATION-RESPONSIVE ELEMENT-BINDING PROTEIN3 (DREB3) is an AP2/EREBP-type transcription factor and overexpression of DREB3 delays flowering in tobacco49. DREB3 was highly expressed in leaf buds, but showed low expression levels in flower buds, indicating that DREB3 is a negative regulator of flowering. Most floral meristem identity genes and floral organ identity genes are members of MADS-box transcription factor family50,51. Nine MADS-box genes were identified from DEGs with seven upregulated genes and two downregulated genes in flower buds, implying that different MADS-box genes have different expression patterns and may play different roles in flower development. However, further studies should explore the specific mechanism.
In addition to Gibberellin, other phytohormones affect the flowering of A. konjac through complex regulatory mechanisms. Previous studies report that auxin plays a key role in development of inflorescence, flower meristem and flower organs52,53,54. Cytokinins modulate initiation and development of reproductive organs55,56. ETH is involved in regulation of the flowering time of plants57. In the current study, significant differences were observed in the expression profiles of several important genes related to auxin, cytokinins, ABA, ETH, JA and Br biosynthesis and signal transduction in leaf and flower buds. Notably, YUCCA4 (Auxin biosynthesis), CCD8B (ABA biosynthesis), NCED1 (ABA biosynthesis), ACS1 (ETH biosynthesis), ACS3 (ETH biosynthesis), ACS9 (ETH biosynthesis), DET2 (BR biosynthesis), 4CLL6 (JA biosynthesis) were differentially expressed in leaf and flower buds. Differential expression of genes implicated in plant hormone signal transduction affects expression of responsive genes, which may ultimately affect the flowering of A. konjac.
The FT gene encodes Phosphatidyl Ethanolamine-binding Protein (PEBP) protein which is a plant flowering integration factor58,59. Currently, FT genes from different plants have been reported21,22,25,60,61,62. Some plants have several types of FT genes, and their functions are different. Previous studies report that FT can integrate signals from various flowering pathways to promote plant flowering63,64,65,66. FT-like and TERMINAL FLOWER 1(TFL1)-like genes affect several physiological processes in plants, such as seed development and germination in A. thaliana23,67, corm formation of potato68 and bulb development of onion24. In this study, two FT homologous genes were isolated from flower buds of A. konjac and were named AkFT1 and AkFT2. Sequence alignment showed that the two FT proteins are high homologous to several plant FT proteins and belonged to FT-like protein. Phylogenetic analysis showed that these genes were highly related to FT protein expressed in monocotyledons, such as rice, wheat and maize. The two FT homologous proteins shared conserved motifs with several FT proteins, indicating that they may have similar biological functions. AkFT1 and AkFT2 were significantly upregulated in flower buds relative to the expression level in leaf buds. This finding was further confirmed by qRT-PCR analysis, indicating that they may be positive regulators of flowering in A. konjac. Significantly, FT protein is transported via the phloem for a long distance, and finally lead to the formation of flowers in the shoot apex17,69. This unique feature of A. konjac leaf and flower not developing simultaneously, and the detection of AkFT1 and AkFT2 mRNAs in flower buds, suggests that the expression pattern of FT in Konjac differs from that of other plants. Overexpression of AkFT1 and AkFT2 reduced the vegetative growth period of A. thaliana and accelerated flowering compared with the wild type plants. AkFT1 and AkFT2 genes may play a critical role similar to the function of florigen during flowering transition of A. konjac, however, the precise molecular mechanism should be explored further.
In summary, comprehensive gene expression information of leaf buds and flower buds of A. konjac was obtained through transcriptome analysis. These results showed that some genes are differentially expressed during the development of leaf buds and flower buds. Two FT homologous genes (AkFT1 and AkFT2) were identified, which exhibited high expression level in flower buds relative to the expression level in leaf buds. Overexpression of AkFT1 and AkFT2 significantly decreased flowering time of transgenic A. thaliana relative to the wild type plants.
References
Hetterscheid, W. & Bogner, J. Notes on the genus Amorphophallus (Araceae) 10. Revision of the endemic Amorphophallus species of Madagascar. Bot. Jahrb. 121, 1–17 (1999).
Chua, M., Baldwin, T. C., Hocking, T. J. & Chan, K. Traditional uses and potential health benefits of Amorphophallus konjac K. Koch ex N.E.Br. J. Ethnopharmacol. 128, 268–278. https://doi.org/10.1016/j.jep.2010.01.021 (2010).
Santosa, E. et al. Gibberellic acid application causes erratic flowering on young corms of Amorphophallus muelleri Blume (Araceae). Hortic. J. 88(1), 92–99. https://doi.org/10.2503/hortj.UTD-016 (2019).
Zhang, D., Wang, Q. & George, S. S. Mechanism of staggered multiple seedling production from Amorphophallus bulbifer and Amorphophallus muelleri and its application to cultivation in Southeast Asia. Trop. Agric. Dev. 54, 84–90. https://doi.org/10.11248/jsta.54.84 (2010).
Tajuddin, M., Sopandie, D. & Lontoh, A. P. Characteristics of growth, flowering and corm yield of iles-iles (Amorphophallus muelleri) genotypes at third growing period. Biodiversitas 21(2), 570–577. https://doi.org/10.13057/biodiv/d210219 (2020).
Zhao, J., Zhang, D., Srzednicki, G., Borompichaichartkul, C. & Kanlayanarat, S. Morphological and growth characteristics of Amorphophallus muelleri Blume-a commercially important konjac species. Acta Hortic. 875, 501–508. https://doi.org/10.17660/ActaHortic.2010.875.65 (2009).
Sumarwoto, S. Iles-iles (Amorphophallus muelleri Blume); description and other characteristics. Biodiversitas 6(3), 185–190. https://doi.org/10.13057/biodiv/d060310 (2005).
Zhao, J., Zhang, D., Srzednicki, G., Kanlayanarat, S. & Borompichaichartkul, C. Asexual reproduction of Amorphophallus bulbifer by low-cost artificial-induction technique. Acta Hortic. 837, 351–358. https://doi.org/10.17660/ActaHortic.2009.837.47 (2009).
Santosa, E., Lontoh, A. P., Kurniawati, A., Sari, M. & Sugiyama, N. Flower development and its implication for seed production on Amorphophallus muelleri Blume (Araceae). J. Hortik. Indones. 7(2), 65–74. https://doi.org/10.29244/jhi.7.2.65-74 (2016).
Gille, S. et al. Deep sequencing of voodoo lily (Amorphophallus konjac): An approach to identify relevant genes involved in the synthesis of the hemicellulose glucomannan. Planta 234, 515–526. https://doi.org/10.1007/s00425-011-1422-z (2011).
Diao, Y. et al. De novo transcriptome and small RNA analyses of two Amorphophallus species. PLoS One 9, e95428. https://doi.org/10.1371/journal.pone.0095428 (2014).
Wang, K. et al. Cloning and evaluation of reference genes for quantitative real-time PCR analysis in Amorphophallus. PeerJ 5, e3260. https://doi.org/10.7717/peerj.3260 (2017).
Singh, V. K., Garg, R. & Jain, M. A global view of transcriptome dynamics during flower development in chickpea by deep sequencing. Plant Biotechnol. J. 11(6), 691–701. https://doi.org/10.1111/pbi.12059 (2013).
Galla, G., Vogel, H., Sharbel, T. F. & Barcaccia, G. De novo sequencing of the Hypericum perforatum L. flower transcriptome to identify potential genes that are related to plant reproduction sensu lato. BMC Genomics 16(1), 1–22. https://doi.org/10.1186/s12864-015-1439-y (2015).
Xu, G., Huang, J., Yang, Y. & Yao, Y. A. Transcriptome analysis of flower sex differentiation in Jatropha curcas L. using RNA sequencing. PLoS One 11(2), e0145613. https://doi.org/10.1371/journal.pone.0145613 (2016).
Chen, G. et al. Comparative transcriptome analysis between fertile and CMS flower buds in Wucai (Brassica campestris L.). BMC Genomics 19(1), 1–17. https://doi.org/10.1186/s12864-018-5331-4 (2018).
Mathieu, J., Warthmann, N., Küttner, F. & Schmid, M. Export of FT protein from phloem companion cells is sufficient for floral induction in Arabidopsis. Curr. Biol. 17(12), 1055–1060. https://doi.org/10.1016/j.cub.2007.05.009 (2007).
Liu, L. et al. FTIP1 is an essential regulator required for florigen transport. PLoS Biol. 10(4), e1001313. https://doi.org/10.1371/journal.pbio.1001313 (2012).
Putterill, J. & Varkonyi-Gasic, E. FT and florigen long-distance flowering control in plants. Curr. Opin. Plant Biol. 33, 77–82 (2016).
Halliday, K. J., Salter, M. G., Thingnaes, E. & Whitelam, G. C. Phytochrome control of flowering is temperature sensitive and correlates with expression of the floral integrator FT. Plant J. 33, 875–885. https://doi.org/10.1046/j.1365-313X.2003.01674.x (2003).
Kojima, S. et al. Hd3a, a rice ortholog of the Arabidopsis FT gene, promotes transition to flowering downstream of Hd1 under short-day conditions. Plant Cell Physiol. 43(10), 1096–1105. https://doi.org/10.1093/pcp/pcf156 (2002).
Meng, X., Muszynski, M. G. & Danilevskaya, O. N. The FT-like ZCN8 gene functions as a floral activator and is involved in photoperiod sensitivity in maize. Plant Cell 23(3), 942–960. https://doi.org/10.1105/tpc.110.081406 (2011).
Zhang, B., Li, C., Li, Y. & Yu, H. Mobile TERMINAL FLOWER1 determines seed size in Arabidopsis. Nat. Plants 6(9), 1146–1157. https://doi.org/10.1038/s41477-020-0749-5 (2020).
Khokhar, K. M. Environmental and genotypic effects on bulb development in onion—A review. J. Hortic. Sci. Biotechnol. 92(5), 448–454. https://doi.org/10.1080/14620316.2017.1314199 (2017).
Navarro, C. et al. Control of flowering and storage organ formation in potato by FLOWERING LOCUS T. Nature 478(7367), 119–122. https://doi.org/10.1038/nature10431 (2011).
Kanehisa, M., Furumichi, M., Sato, Y., Ishiguro-Watanabe, M. & Tanabe, M. KEGG: Integrating viruses and cellular organisms. Nucleic Acids Res. 49, D545–D551. https://doi.org/10.1093/nar/gkaa970 (2021).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Methodol. 57, 289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x (1995).
Sudhir, K., Glen, S. & Koichiro, T. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33(7), 1870–1874. https://doi.org/10.1093/molbev/msw054 (2016).
Bailey, T. L. et al. MEME Suite: Tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208. https://doi.org/10.1093/nar/gkp335 (2009).
Clough, S. J. & Bent, A. F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743. https://doi.org/10.1046/j.1365-313X.1998.00343.x (1998).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29(9), e45 (2001).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652. https://doi.org/10.1038/nbt.1883 (2011).
Alexa, A. & Rahnenführer, J. Gene set enrichment analysis with topGO. Bioconductor Improv. 27, 1–26 (2009).
Cerdán, P. D. & Chory, J. Regulation of flowering time by light quality. Nature 423, 881–885. https://doi.org/10.1038/nature01636 (2003).
Fornara, F., de Montaigu, A. & Coupland, G. SnapShot: Control of flowering in Arabidopsis. Cell 141(3), 550–552. https://doi.org/10.1016/j.cell.2010.04.024 (2010).
Wang, J. W. Regulation of flowering time by the miR156-mediated age pathway. J. Exp. Bot. 65, 4723–4730. https://doi.org/10.1093/jxb/eru246 (2014).
Bao, S., Hua, C., Shen, L. & Yu, H. New insights into gibberellin signaling in regulating flowering in Arabidopsis. J. Integr. Plant Biol. 62(1), 118–131. https://doi.org/10.1111/jipb.12892 (2020).
Chandler, J. W. Founder cell specification. Trends Plant Sci. 16(11), 607–613. https://doi.org/10.1016/j.tplants.2011.08.005 (2011).
Sri, T., Gupta, B., Tyagi, S. & Singh, A. Homeologs of Brassica SOC1, a central regulator of flowering time, are differentially regulated due to partitioning of evolutionarily conserved transcription factor binding sites in promoters. Mol. Phylogenet. Evol. 147, 106777. https://doi.org/10.1016/j.ympev.2020.106777 (2020).
Santosa, E., Sugiyama, N., Kurniawati, A., Lontoh, A. P. & Sari, M. Variation in floral morphology of agamosporous (Amorphophallus muelleri Blume) in natural and gibberellin induced flowering. J. Appl. Hortic. 20(1), 15–23 (2018).
Yamaguchi, S. Gibberellin metabolism and its regulation. Annu. Rev. Plant Biol. 59, 225–251. https://doi.org/10.1146/annurev.arplant.59.032607.092804 (2008).
Hedden, P. & Thomas, S. G. Gibberellin biosynthesis and its regulation. Biochem. J. 444(1), 11–25. https://doi.org/10.1042/BJ20120245 (2012).
Peng, J. et al. ‘Green revolution’ genes encode mutant gibberellin response modulators. Nature 400(6741), 256–261. https://doi.org/10.1038/22307 (1999).
Ariizumi, T., Lawrence, P. K. & Steber, C. M. The role of two F-box proteins, SLEEPY1 and SNEEZY, in Arabidopsis gibberellin signaling. Plant Physiol. 155(2), 765–775. https://doi.org/10.1104/pp.110.166272 (2011).
Cao, D., Hussain, A., Cheng, H. & Peng, J. Loss of function of four DELLA genes leads to light and gibberellin-independent seed germination in Arabidopsis. Planta 223(1), 105–113. https://doi.org/10.1007/s00425-005-0057-3 (2005).
Galvao, V. C., Horrer, D., Kuettner, F. & Schmid, M. Spatial control of flowering by DELLA proteins in Arabidopsis thaliana. Development 139, 4072–4082. https://doi.org/10.1242/dev.080879 (2012).
Wang, J. W., Czech, B. & Weigel, D. miR156-regulated SPL transcription factors define an endogenous flowering pathway in Arabidopsis thaliana. Cell 138, 738–749. https://doi.org/10.1016/j.cell.2009.06.014 (2009).
Yu, S. et al. Gibberellin regulates the Arabidopsis floral transition through miR156-targeted SQUAMOSA promoter binding-like transcription factors. Plant Cell 24, 3320–3332. https://doi.org/10.1105/tpc.112.101014 (2012).
Lin, P., Shen, C., Chen, H., Yao, X. H. & Lin, J. Improving tobacco freezing tolerance by co-transfer of stress-inducible CbCBF and CbICE53 genes. Biol. Plant. 61(3), 520–528. https://doi.org/10.1007/s10535-016-0687-2 (2017).
Chae, E., Tan, Q. K. G., Hill, T. A. & Irish, V. F. An Arabidopsis F-box protein acts as a transcriptional co-factor to regulate floral development. Development 135(7), 1235–1245. https://doi.org/10.1242/dev.015842 (2008).
Thompson, B. E. et al. bearded-ear encodes a MADS box transcription factor critical for maize floral development. Plant Cell 21(9), 2578–2590. https://doi.org/10.1105/tpc.109.067751 (2009).
Pagnussat, G. C., Alandete-Saez, M., Bowman, J. L. & Sundaresan, V. Auxin-dependent patterning and gamete specification in the Arabidopsis female gametophyte. Science 324(5935), 1684–1689. https://doi.org/10.1126/science.1167324 (2009).
McSteen, P. Auxin and monocot development. Cold Spring Harb. Perspect. Biol. 2(3), a001479. https://doi.org/10.1101/cshperspect.a001479 (2010).
Yoshida, A., Ohmori, Y., Kitano, H., Taguchi-Shiobara, F. & Hirano, H. Y. ABERRANT SPIKELET AND PANICLE1, encoding a TOPLESS-related transcriptional co-repressor, is involved in the regulation of meristem fate in rice. Plant J. 70(2), 327–339. https://doi.org/10.1111/j.1365-313X.2011.04872.x (2012).
Werner, T. & Schmülling, T. Cytokinin action in plant development. Curr. Opin. Plant Biol. 12(5), 527–538. https://doi.org/10.1016/j.pbi.2009.07.002 (2009).
Zhang, L. et al. TaCKX6-D1, the ortholog of rice OsCKX2, is associated with grain weight in hexaploid wheat. New Phytol. 195(3), 574–584. https://doi.org/10.1111/j.1469-8137.2012.04194.x (2012).
Lin, Z., Zhong, S. & Grierson, D. Recent advances in ethylene research. J. Exp. Bot. 60(12), 3311–3336. https://doi.org/10.1093/jxb/erp204 (2009).
Chardon, F. & Damerval, C. Phylogenomic analysis of the PEBP gene family in cereals. J. Mol. Evol. 61(5), 579–590. https://doi.org/10.1007/s00239-004-0179-4 (2005).
Karlgren, A. et al. Evolution of the PEBP gene family in plants: Functional diversification in seed plant evolution. Plant Physiol. 156(4), 1967–1977. https://doi.org/10.1104/pp.111.176206 (2011).
Kardailsky, I. et al. Activation tagging of the floral inducer FT. Science 286(5446), 1962–1965. https://doi.org/10.1126/science.286.5446.1962 (1999).
Yan, L. et al. The wheat and barley vernalization gene VRN3 is an orthologue of FT. Proc. Natl. Acad. Sci. 103(51), 19581–19586. https://doi.org/10.1073/pnas.0607142103 (2006).
Wu, F., Sedivy, E. J., Price, W. B., Haider, W. & Hanzawa, Y. Evolutionary trajectories of duplicated FT homologues and their roles in soybean domestication. Plant J. 90(5), 941–953. https://doi.org/10.1111/tpj.13521 (2017).
Parcy, F. Flowering: A time for integration. Int. J. Dev. Biol. 49(5–6), 585–593. https://doi.org/10.1387/ijdb.041930fp (2004).
Böhlenius, H. et al. CO/FT regulatory module controls timing of flowering and seasonal growth cessation in trees. Science 312(5776), 1040–1043. https://doi.org/10.1126/science.1126038 (2006).
Hisamatsu, T. & King, R. W. The nature of floral signals in Arabidopsis. II. Roles for FLOWERING LOCUS T (FT) and gibberellin. J. Exp. Bot. 59(14), 3821–3829. https://doi.org/10.1093/jxb/ern232 (2008).
Zhu, Y., Klasfeld, S. & Wagner, D. Molecular regulation of plant developmental transitions and plant architecture via PEPB family proteins—An update on mechanism of action. J. Exp. Bot. https://doi.org/10.1093/jxb/eraa598 (2021).
Xi, W., Liu, C., Hou, X. & Yu, H. MOTHER OF FT AND TFL1 regulates seed germination through a negative feedback loop modulating ABA signaling in Arabidopsis. Plant Cell 22(6), 1733–1748. https://doi.org/10.1105/tpc.109.073072 (2010).
Abelenda, J. A., Navarro, C. & Prat, S. From the model to the crop: Genes controlling tuber formation in potato. Curr. Opin. Biotechnol. 22(2), 287–292. https://doi.org/10.1016/j.copbio.2010.11.013 (2011).
Zhu, Y., Liu, L., Shen, L. & Yu, H. NaKR1 regulates long-distance movement of FLOWERING LOCUS T in Arabidopsis. Nat. Plants 2(6), 1–10. https://doi.org/10.1038/NPLANTS.2016.75 (2016).
Acknowledgements
Thanks for Professor Yong-yao Wu provided materials.
Funding
This study was supported by grants from Education Department of Hunan Province (18A104) and the Natural Science Foundation in Hunan Province (2019JJ50239, 2021JJ30325).
Author information
Authors and Affiliations
Contributions
H.G. designed and completed all the research and write the paper; Y.Z., L.H., Y.H., J.C. and H.Z. participate the bioinformation analysis and discussion; X.Z. supervised the project and revised the manuscript; All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gao, H., Zhao, Y., Huang, L. et al. Comparative analysis of buds transcriptome and identification of two florigen gene AkFTs in Amorphophallus konjac. Sci Rep 12, 6782 (2022). https://doi.org/10.1038/s41598-022-10817-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-10817-5
- Springer Nature Limited
This article is cited by
-
Combined Analysis of the Metabolome and Transcriptome Sheds New Light on the Mechanisms of Seed Maturation in Amorphophallus muelleri
Journal of Plant Growth Regulation (2024)
-
Transcriptomic analysis reveals biosynthesis genes and transcription factors related to leaf anthocyanin biosynthesis in Aglaonema commutatum
BMC Genomics (2023)