Abstract
Mastitis by non-aureus staphylococci (NAS) is a significant issue in dairy buffalo farming. In a herd with subclinical NAS mastitis, we identified Staphylococcus microti as the predominant species. To assess milk protein integrity and investigate potential disease markers, we characterized 12 NAS-positive and 12 healthy quarter milk samples by shotgun peptidomics combining peptide enrichment and high-performance liquid chromatography/tandem mass spectrometry (LC–MS/MS). We observed significant changes in the milk peptidome. Out of 789 total peptides identified in each group, 49 and 44 were unique or increased in NAS-positive and healthy milk, respectively. In NAS-positive milk, the differential peptides belonged mainly to caseins, followed by milk fat globule membrane proteins (MFGMP) and by the immune defense/antimicrobial proteins osteopontin, lactoperoxidase, and serum amyloid A. In healthy milk, these belonged mainly to MFGMP, followed by caseins. In terms of abundance, peptides from MFGMP and immune defense protein were higher in NAS-positive milk, while peptides from caseins were higher in healthy milk. These findings highlight the impact of NAS on buffalo milk quality and mammary gland health, even when clinical signs are not evident, and underscore the need for clarifying the epidemiology and relevance of the different NAS species in this dairy ruminant.
Similar content being viewed by others
Introduction
The water buffalo (Bubalus bubalis) is the second most relevant dairy species after the cow (Bos taurus)1, with over 97 million tons of milk produced each year2. Mastitis caused by an intramammary infection (IMI) is one of the diseases with the highest impact on the economic performance and welfare of dairy animals3. Water buffaloes are generally regarded as less susceptible to mastitis than cows4,5. Still, the real impact of intramammary infections (IMI) may be underestimated due to the higher prevalence of subclinical mastitis and issues with the setting of somatic cell count (SCC) thresholds5,6, which need proper implementation for mastitis monitoring within dairy herd improvement programs.
The main etiologic agents of clinical and subclinical IMI in buffalo are staphylococci5,6. S. aureus is a highly impacting pathogen for clinical severity and ability to spread and persist in the herd, but non-aureus staphylococci (NAS) are most frequently isolated from the milk6,7,8. Moreover, milk NAS in water buffalo have been recently reported as a source of antibiotic resistance9,10,11,12,13.
The relationship between different NAS and mammary gland health is poorly known. Identification of NAS at the species level is seldom carried out in routine milk bacteriology because of analytical cost issues, combined with the sub-optimal performance of traditional biochemical methods5,7. Genotypic identification is also problematic in some cases due to the high similarity between some species14. When possible, NAS identification is carried out by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF-MS)15,16,17. Recently, we detected significant changes in the protein composition of buffalo milk with staphylococcal mastitis8. In that study, we highlighted the need to clarify the role of the different NAS species in this dairy animal and to further investigate the impact of NAS on buffalo milk quality. Shotgun peptidomics is an approach providing an in-depth perspective on the changes occurring in the peptide profile of many dairy products, adding useful information to the proteomic approach18. This method can assess the impact of different conditions by combining the simultaneous identification of thousands of peptides with their quantification in each sample19. Therefore, this approach is ideal for quantitatively investigating the differences in the peptidome of milk from healthy animals compared to that from infected udder quarters with mastitis30.
In this study, we investigated the impact of NAS IMI on the buffalo milk peptidome with a proteomic analysis pipeline entailing peptide enrichment, high-performance liquid chromatography/tandem mass spectrometry, and bioinformatic analysis, taking into account the causative NAS species and the milk SCC in the definition of the sample groups.
Results
Milk somatic cell counts and bacteriology
Staphylococcus microti was identified in all the NAS-positive milk samples with MALDI-TOF-MS scores higher than 2.00 but in two cases (1.93 and 1.95, respectively). Three milk samples showed the growth of a second colony type, identified as Aerococcus viridans in one sample and Streptococcus uberis in two samples. The downstream peptidomic analysis was carried out by classifying the samples according to the combination of the bacteriological and somatic cell count (SCC) information, as detailed in Table 1. The complete data are reported in Supplementary Table 1.
Differential peptidomics
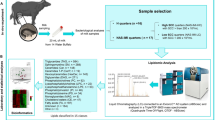
The milk samples listed in Table 1 were subjected to a pipeline entailing peptide enrichment, peptide analysis by high-performance liquid chromatography/tandem mass spectrometry (LC–MS/MS), and bioinformatic analysis to identify differential peptides in the two sample groups. The experimental protocol used in this study is schematically summarized in Fig. 1A.
Shotgun label-free quantitative peptidomic analysis. (A) Overview of the protocols applied for the analysis of peptides in NAS-positive and healthy milk samples (B) Venn diagram of all the peptides identified in milk samples from healthy and NAS-positive buffaloes. Peptides were considered differentially abundant if they were present only in NAS-positiver or Control milk or showed significant Welch t test difference (cut-off at 1% permutation-based False Discovery Rate).
789 and 789 peptides were identified in the NAS-positive and healthy milk samples, respectively, for a total of 833 identified peptides (Fig. 1B).Among the 745 peptides present in both groups, 5 were increased in NAS-positive in comparison to healthy milk (Welch's t test: FDR 0.01). No peptides were found decreased in NAS-positive milk. Overall, the analysis identified 49 peptides which were increased (5) or present only in NAS-positive (44), and 44 peptides which were present only in healthy milk (44). Table 2 reports the number of total peptides and the number of unique and differential peptides identified in NAS-positive and healthy milk samples.
Table 3 details the sequence of all the unique and differential peptides identified in NAS-positive and healthy milk, their originating protein, and the cell location/function based on the UniProtKB protein knowledge base or scientific literature sources20,21,22,23,24,25.
Figure 2A,B illustrate the distribution of all unique and differential peptides identified in NAS-positive and healthy milk in terms of number and abundance, respectively, according to the cell location/function of the originating protein and highlight the different nature of the unique and differential peptides identified in the two sample groups. The number of total and differential peptides identified in the two groups was similar, but their nature in terms of originating proteins differed. In NAS-positive milk, 28 of the 49 peptides (57.14%) belonged to caseins, mainly beta-casein (15, 30,61%), followed by alphaS2 (7, 14.29%) and alphaS1 (6, 12.24%), and 9 peptides belonged to proteins of the milk fat globule membrane (MFGMP) (18.37%). Interestingly, 5 peptides belonged to proteins with immune defense/antimicrobial functions (10.20%), namely osteopontin, lactoperoxidase, and serum amyloid A-3. The peptides belonging to proteins with other locations/functions, including cell/vesicle membrane, nucleus/cytosol, and secreted, were 7 out of 49 (14.29%). Conversely, in healthy milk, most unique peptides (17 out of 38,63%) belonged to MFGMP, and 16 (36.36%) belonged to caseins, mainly beta casein (5, 11.36%), alphaS1 (5, 11.36%), alphaS2 (4, 9.09%), and kappa (2, 4.55%). Only 2 unique peptides (4.54%) belonged to inflammatory/immune defense proteins. The remaining 9 unique peptides belonged to proteins with other locations/functions (20.45%).
In terms of their relative abundance, those derived from casein micelle proteins were higher in healthy milk, while those derived from MFGMP were higher in NAS-positive milk. On the other hand, Immune defense proteins were higher in NAS-positive milk also in terms of relative abundance.
The differential peptides were manually analyzed and classified according to their C-terminal amino acid. As shown in Fig. 3, R at the C-term was considerably less frequent (10.2%) in the peptides unique or more abundant in healthy milk. On the other hand, peptides ending especially with K (44.9%), V (8.16%), and F (4.08%) were more frequent in NAS-positive milk.
Discussion
Based on our findings, the presence of a NAS IMI was associated with changes in the peptide composition of water buffalo milk. The differential peptides identified were derived from proteins with very different functions and localizations. As milk quality and technological properties may be affected, this deserves consideration26.
We detected four differential peptides from serum amyloid A (SAA) in NAS-positive animals. SAA is associated with high SCC and mastitis in bovine cows24,27, being an acute phase protein24 that is overexpressed in milk during mastitis28,29. The mammary gland produces a specific form of SAA, the M-SAA330,31, which can be upregulated by S. aureus lipoteichoic acid32. One differential peptide originating from SAA A-3 (VISNARETIQGITDPLLKGMTRDQVREDSKADQ FANEWGR) was found uniquely in NAS-positive milk, in line with our previous finding of the SAA protein only in the milk of water buffaloes with staphylococcal IMI by shotgun proteomics8. Interestingly, in another shotgun peptidomics study, we detected SAA peptides only in cows with NAS IMI33. Thus, our detection of SAA only in the milk of animals with NAS IMI further supports its diagnostic potential in the dairy buffalo34. Nevertheless, the influence of other physiological variables including parity and stage of lactation on M-SAA levels will have to be assessed35.
Three unique and significantly differential peptides originating from osteopontin were found in NAS-positive milk. This is also in line with our previous peptidomic study on bovine cows33, although we did not identify the intact protein by proteomics in the water buffalo8. Among other biological roles, osteopontin upregulates interferon-gamma and interleukin-12 and downregulates interleukin-10 and plays a role in inducing type I immunity36. In cows, osteopontin peptides have been reported in subclinical mastitis37 and experimental Escherichia coli IMI21,22. As also indicated in a recent review on NAS affecting cows, this further indicates that NAS indeed elicit an inflammatory response in the mammary gland, as confirmed by the increased milk SCC. The present finding may support the hypothesis that NAS provide cross-protection against other mastitis pathogens38 as components of the mammary gland microbiota39,40.
On the other hand, most of the unique peptides found in healthy milk belonged to proteins of the milk fat globule membrane (perilipin 2, butyrophilin, GLYCAM-1, sodium-dependent phosphate cotransporter, annexins, glycoprotein-2)25,41, in line with the observations made by shotgun proteomics8. The predominance of MFG membrane proteins in healthy milk might be related to the high fat content of buffalo milk, and therefore to the higher abundance of these lipid secretion vesicles compared to cow milk. MFG are an important source of nutraceutical components, including membrane proteins, and the possible influence of NAS IMI on their integrity may deserve further consideration concerning nutritional value, product quality, and technological properties26. When looking at the differential distribution of peptides in terms of abundance, we observed that healthy milk was characterized by a higher abundance of casein proteolytic peptides, and NAS-positive milk by a higher abundance of peptides derived from MFG membrane proteins and immune defence proteins. While the first finding might be influenced by the higher abundance of caseins in healthy vs infected milk, the second finding further highlights the impact of NAS IMI on integrity and abundance of MFG membrane proteins and immune defence proteins, respectively, reinforcing the above considerations8.
The distribution of unique and differential peptides based on their C-terminal aminoacid showed a higher frequency of peptides ending with R in healthy milk as opposite to peptides ending especially with K, V, and F, in NAS-positive milk, in line with the observations made by our previous peptidomic work in bovine cows33. According to the MEROPS database, plasmin generates peptides ending with R and K at the C-term, while elastase, cathepsin D and cathepsin G generate peptides ending with V and F at their C-term42. Our results suggest a more intense proteolytic activity by plasmin and endogenous proteases released by inflammatory cells in NAS-positive milk.
The impact of NAS IMI on the buffalo milk peptidome was less intense than observed in cows in our recent work33. However, as mentioned above, many findings were consistent including the presence in NAS-positive milk of peptides derived from osteopontin and SAA, and the different frequency of C-terminal aminoacids in the proteolytic peptides of the two sample groups33.
Concerning the etiologic agent, the identification of S. microti as the predominant species in the milk of water buffaloes with subclinical mastitis is noteworthy as only one study reported its association with mastitis in bovine cows43. S. microti is closely associated with S. rostri and S. muscae, and it has been first isolated from Microtus arvalis, the common vole. Since its description, it has been isolated from rodents/insectivores and a female sandfly43. Therefore, the role of animal vectors might be relevant in this case. Adding to anatomical and physiological characteristics, important differences characterize bubaline cows and bovine cows in terms of animal management, farming practices (housing, feeding, bedding, milking routine), environmental temperature and humidity, and presence of water ponds, and consequently contact with different microbial reservoirs including wild and domestic animals. This may lead to mammary gland exposure and colonization by other NAS species than the bovine dairy cows, as well as to different bacterial loads in the farm environment, and should be carefully considered.
Methods
Animals and milk samples
The study was carried out on quarter milk samples collected from a commercial water buffalo dairy farm located in Campania, Southern Italy, with an increased bulk tank somatic cell count related to NAS IMI. The farm maintained the milking buffaloes in free-stall barns with deep-bedded cubicles with straw. All the animals were fed with a balanced Total Mixed Ration in feed alleys with headlocks. Lactating cows were milked twice a day in a double-10 herringbone parlour milking. The farm was free of brucellosis and paratuberculosis. All the milk samples used for this study were collected within the frame of a diagnostic routine visit for monitoring the health status of the herd. This practice is approved by the Ethical Committee of the University of Milan (Comitato Etico 15.02.16 Parere numero 2/16) “allowing the use, under informed consent of the owners, of the residual volume of samples for studies on metabolic biomarkers”. All methods and procedures were performed in accordance with the relevant institutional guidelines and regulations. The methods described and the results reported were compliant with the ARRIVE guidelines for reporting animal research44.
Milk samples were processed as indicated by the National Mastitis Council45. Before sampling, teats were cleaned with a pre-dipping foam containing lactic acid, and the apex was scrubbed and disinfected with alcohol. The first streams of milk were discharged, and 20 mL was collected aseptically from each quarter into sterile vials. The milk samples were kept at 4 °C until they reached the laboratory (within the day) at the IZS in Portici for bacteriological assays and somatic cell count.
Bacteriological analysis and somatic cell count
Bacteriological cultures were performed according to the National Mastitis Council45. Briefly, 10 µL of milk was spread on blood agar (Trypticase Soy Agar with 5% defibrinated sheep blood) and incubated at 37 °C for 24 h in aerobic conditions. For a preliminary identification of the etiological agent, Gram staining, catalase, and coagulase tests were performed on the colonies observed for all positive cultures, as part of the routine milk analysis. The SCC was measured in milk samples using the Fossomatic (Foss) instrument following the UNI EN ISO 13366-2: 2007 standard and expressed as the number of cells per mL of milk. Two equally populated sample groups (12 samples per group) were defined based on the combined results: NAS-positive (no more than two colony types, bacterial count per colony ≥ 500 colony-forming units per mL (CFU/mL), SCC ≥ 100,000 cells/mL) and healthy (no growth, SCC < 100,000 cells/mL).
MALDI-TOF-MS for bacterial identification
Milk samples growing coagulase-negative staphylococci at the routine milk analysis were sent frozen to the milk quality laboratory at the University of Milan for microbial identification by MALDI-TOF-MS. There, milk samples were thawed at room temperature, and 100 µL of milk was spread on blood agar half-plates to obtain bacterial colonies for MALDI-TOF-MS identification. After incubation for 24 h at 37 °C, the plates were examined for microbial growth. When no more than two different colony types were present, these were counted and assessed by MALDI-TOF-MS for microbial identification as described previously46, with minor modifications. One colony was deposited in duplicate on the target plate using a toothpick, overlaid with 1 µL of α-cyano-4-hydroxycinnamic acid (Bruker Daltonik GmbH, Bremen, Germany) and left to dry. The spectra were acquired with a microFlex™ mass spectrometer (Bruker Daltonik GmbH) in the positive mode, including two spots of Bacterial Test Standard (Bruker Daltonik GmbH) for calibration in each target plate. The obtained spectra were interpreted against the MBT Compass® 4.1 database. Log scores of ≥ 1.7 and ≥ 2.0 were the thresholds for the genus and species level identification, respectively.
Milk sample preparation for peptidomic analysis
The milk samples were processed for peptidomic analysis as described previously33. Briefly, milk was defatted by centrifugation and processed for the depletion of high-molecular-weight proteins on centrifugal filters. The filtrate was precipitated, and peptides were dried, dissolved in 1% (v/v) formic acid and desalted before MS analysis.
Tandem mass spectrometry analysis of peptides
Tandem mass spectrometry analysis of peptides was carried out with duplicate runs for each sample as described previously33. Briefly, LC-ESI-MS/MS analysis was performed on a Dionex UltiMate 3000 directly connected to an Orbitrap Fusion Tribrid Mass Spectrometer (Thermo Fisher Scientific) by a nanoelectrospray ion source. Peptide mixtures were enriched on 75 μmID × 150 mm Acclaim PepMap RSLC C18 column and separated employing the following LC gradient: 4% ACN in 0.1% formic acid for 3 min, 4–28% ACN in 0.1% formic acid for 130 min, 28–40% ACN in 0.1% formic acid for 20 min, 40–95% ACN in 0.1% formic for 2 min and 95–4% ACN in 0.1% formic acid for 3 min at a flow rate of 0.3 μL/min. MS spectra of eluting peptides were collected over an m/z range of 375–1500 using a resolution setting of 120,000, operating in the data-dependent mode to automatically alternate between Orbitrap-MS and Orbitrap-MS/MSacquisition. HCD MS/MS spectra were collected for the 20 most abundant ions in each MS scan using a normalized collision energy of 30%, and an isolation window of 1.7 m/z. Rejection of + 1 and unassigned charge states were enabled.
Database search, peptide identification, and differential analysis
Raw label-free MS/MS files from Thermo Xcalibur software (version 4.1) were analyzed using Proteome Discoverer software (version 2.2, Thermo Fisher Scientific)47 and searched with the Sequest algorithm against the proteome of Bubalus bubalis from NCBI 01-08-2019 and Staphylococcus from Uniprot 18-06-2019. Only peptides with high FDR confidence were considered (FDR 0.01 strict, FDR 0.05 relaxed) to remove false-positive matches. The assigned peptides are filtered by minimal peptide length (6 amino acids) and m/z accuracy (8 ppm). The quality of a match between sequence and observed peaks was provided by a high cross-correlation score (≥ 1.5). PSM confidence was set to High.
Unspecific digestion was chosen, and neither fixed nor variable modifications were set. The resulting peptides and protein hits were further screened accepting only those hits listed as high confidence and with an Xcorr ≥ 1.5. Only peptides present and quantified in 66,6% of the repeats were positively identified and used for statistical analysis. Peptides were considered increased or decreased if they showed a significant Welch t test difference (cut-off at 1% permutation-based FDR) or if they were present with a frequency ≥ 66,6% in either NAS-positive or healthy milk group but less than 66,6% in the other group48. Statistical analysis was performed using the Perseus software (version 1.5.5.3, http://www.biochem.mpg.de/mann/tools/). Peptide sequences were analyzed manually for C-terminal amino acids. The potential proteases generating the cuts were classified based on the MEROPS database42 by evaluating the specificities of the main proteases generating the cuts49. The protein functions reported in Table 3 were retrieved from the UniProtKB protein knowledgebase (http://www.uniprot.org).
Data availability
The data have been deposited to the ProteomeXchange with identifier PXD028793.
References
IDF bulletin—International Dairy Federation. The world dairy situation 2007. Bulletin No. 423/2007 (2007).
Zicarelli, L. Current trends in buffalo milk production. J. Buffalo Sci. 9, 121–132 (2020).
Ruegg, P. L. A 100-Year Review: Mastitis detection, management, and prevention. J. Dairy Sci. 100, 10381–10397 (2017).
Wanasinghe, D. D. Mastitis among buffalos in Sri Lanka. in Proc. First World Buffalo Congr. Cairo, Egypt. 1331–1333 (1985).
Guccione, J. & Ciaramella, P. Mastitis in Mediterranean buffaloes. J. Dairy Vet. Sci. 2, 1–4 (2017).
Moroni, P. et al. Relationships between somatic cell count and intramammary infection in buffaloes. J. Dairy Sci. 89, 998–1003 (2006).
Guccione, J. et al. Clinical outcomes and molecular genotyping of Staphylococcus aureus isolated from milk samples of dairy primiparous Mediterranean buffaloes (Bubalus bubalis). J. Dairy Sci. 97, 7606–7613 (2014).
Pisanu, S. et al. Proteomic changes in the milk of water buffaloes (Bubalus bubalis) with subclinical mastitis due to intramammary infection by Staphylococcus aureus and by non-aureus staphylococci. Sci. Rep. 9, 1–14 (2019).
Aqib, A. I. et al. Antibiotic susceptibilities and prevalence of Methicillin resistant Staphylococcus aureus (MRSA) isolated from bovine milk in Pakistan. Acta Trop. 176, 168–172 (2017).
El-Ashker, M. et al. Microarray-based detection of resistance genes in coagulase-negative staphylococci isolated from cattle and buffalo with mastitis in Egypt. Trop. Anim. Health Prod. 52, 3855–3862 (2020).
Santos, A. D. S. et al. Antimicrobial resistance profile of non-aureus Staphylococci isolates from buffalo, goat and sheep mastitis in the Northeast region of Brazil. J. Dairy Res. 87, 290–294 (2020).
Badua, A. T., Boonyayatra, S., Awaiwanont, N., Gaban, P. B. V. & Mingala, C. N. Antibiotic resistance and genotyping of mecA-positive methicillin-resistant Staphylococcus aureus (MRSA) from milk and nasal carriage of dairy water buffaloes (Bubalus bubalis) in the Philippines. J. Adv. Vet. Anim. Res. 7, 397–406 (2020).
Badua, A. T., Boonyayatra, S., Awaiwanont, N., Gaban, P. B. V. & Mingala, C. N. Methicillin-resistant Staphylococcus aureus (MRSA) associated with mastitis among water buffaloes in the Philippines. Heliyon 6, e05663 (2020).
Lange, C. C. et al. Species-level identification of staphylococci isolated from bovine mastitis in Brazil using partial 16S rRNA sequencing. Vet. Microbiol. 176, 382–388 (2015).
Mahmmod, Y. S., Nonnemann, B., Svennesen, L., Pedersen, K. & Klaas, I. C. Typeability of MALDI-TOF assay for identification of non-aureus staphylococci associated with bovine intramammary infections and teat apex colonization. J. Dairy Sci. 101, 9430–9438 (2018).
Hulland, C., Dufour, S. & Munoz, M. Milk Bacteriological Analysis Using MALDI-TOF Technology (National Mastitis Council, 2018).
Nonnemann, B. et al. Bovine mastitis bacteria resolved by MALDI-TOF mass spectrometry. J. Dairy Sci. 102, 2515–2524 (2019).
Punia, H. et al. Identification and detection of bioactive peptides in milk and dairy products: Remarks about agro-foods. Molecules 25, 3328 (2020).
Aletti, F. et al. Peptidomic analysis of rat plasma: Proteolysis in hemorrhagic shock. Shock 45, 540–554 (2016).
Lu, J. et al. The protein and lipid composition of the membrane of milk fat globules depends on their size. J. Dairy Sci. 99, 4726–4738 (2016).
Wang, K. X. & Denhardt, D. T. Osteopontin: Role in immune regulation and stress responses. Cytokine Growth Factor Rev. 19, 333–345 (2008).
Bissonnette, N., Dudemaine, P. L., Thibault, C. & Robitaille, G. Proteomic analysis and immunodetection of the bovine milk osteopontin isoforms. J. Dairy Sci. 95, 567–579 (2012).
Fong, B. Y., Norris, C. S. & MacGibbon, A. K. H. Protein and lipid composition of bovine milk-fat-globule membrane. Int. Dairy J. 17, 275–288 (2007).
Ceciliani, F., Ceron, J. J., Eckersall, P. D. & Sauerwein, H. Acute phase proteins in ruminants. J. Proteomics 75, 4207–4231 (2012).
Reinhardt, T. A. & Lippolis, J. D. Bovine milk fat globule membrane proteome. J. Dairy Res. 73, 406–416 (2006).
Brijesha, N. & Aparna, H. S. Comprehensive characterization of bioactive peptides from Buffalo (Bubalus bubalis) colostrum and milk fat globule membrane proteins. Food Res. Int. 97, 95–103 (2017).
Isobe, N., Kubota, H., Yamasaki, A. & Yoshimura, Y. Lactoperoxidase activity in milk is correlated with somatic cell count in dairy cows. J. Dairy Sci. 94, 3868–3874 (2011).
Grönlund, U., Hultén, C., Eckersall, P. D., Hogarth, C. & PerssonWaller, K. Haptoglobin and serum amyloid A in milk and serum during acute and chronic experimentally induced Staphylococcus aureus mastitis. J. Dairy Res. 70, 379–386 (2003).
Grönlund, U., Hallén Sandgren, C. & Persson Waller, K. Haptoglobin and serum amyloid A in milk from dairy cows with chronic sub-clinical mastitis. Vet. Res. 36, 191–198 (2005).
McDonald, T. L., Larson, M. A., Mack, D. R. & Weber, A. Elevated extrahepatic expression and secretion of mammary-associated serum amyloid A 3 (M-SAA3) into colostrum. Vet. Immunol. Immunopathol. 83, 203–211 (2001).
Larson, M. A., Weber, A., Weber, A. T. & McDonald, T. L. Differential expression and secretion of bovine serum amyloid A3 (SAA3) by mammary epithelial cells stimulated with prolactin or lipopolysaccharide. Vet. Immunol. Immunopathol. 107, 255–264 (2005).
Weber, A., Weber, A. T., McDonald, T. L. & Larson, M. A. Staphylococcus aureus lipotechoic acid induces differential expression of bovine serum amyloid A3 (SAA3) by mammary epithelial cells: Implications for early diagnosis of mastitis. Vet. Immunol. Immunopathol. 109, 79–83 (2006).
Addis, M. F. et al. Influence of subclinical mastitis and intramammary infection by coagulase-negative staphylococci on the cow milk peptidome. J. Proteomics 226, 103885 (2020).
Singh, M. et al. Estimation of acute phase proteins as early biomarkers of buffalo subclinical mastitis. Asian J. Anim. Vet. Adv. 10, 894–902 (2015).
Pisanu, S. et al. Impact of Staphylococcus aureus infection on the late lactation goat milk proteome: New perspectives for monitoring and understanding mastitis in dairy goats. J. Proteomics 221, 103763 (2020).
Lund, S. A., Giachelli, C. M. & Scatena, M. The role of osteopontin in inflammatory processes. J. Cell Commun. Signal. 3, 311–322 (2009).
Guerrero, A. et al. Peptidomic analysis of healthy and subclinically mastitic bovine milk. Int. Dairy J. 46, 46–52 (2015).
De Buck, J. et al. Non-aureus staphylococci and bovine udder health: Current understanding and knowledge gaps. Front. Vet. Sci. 8, 360 (2021).
Catozzi, C. et al. The microbiota of water buffalo milk during mastitis. PLoS One 12, e0184710 (2017).
Oikonomou, G. et al. Milk microbiota: What are we exactly talking about?. Front. Microbiol. 11, 1–15 (2020).
Pisanu, S. et al. The sheep milk fat globule membrane proteome. J. Proteomics 74, 350–358 (2011).
Rawlings, N. D., Barrett, A. J. & Bateman, A. MEROPS: The database of proteolytic enzymes, their substrates and inhibitors. Nucleic Acids Res. 40, D343–D350 (2012).
Król, J. et al. Isolation of Staphylococcus microti from milk of dairy cows with mastitis. Vet. Microbiol. 182, 163–169 (2016).
du Sert, N. P. et al. The arrive guidelines 2.0: Updated guidelines for reporting animal research. PLoS Biol. 18, e3000410 (2020).
Middleton, J. R., Fox, L. K. & Pighetti, G. Laboratory Handbook on Bovine Mastitis (National Mastitis Council, 2017).
Monistero, V. et al. Genotyping and antimicrobial susceptibility profiling of streptococcus uberis isolated from a clinical bovine mastitis outbreak in a dairy farm. Antibiotics 10, 644 (2021).
Galli, A. et al. Cluster-assembled zirconia substrates promote long-term differentiation and functioning of human islets of Langerhans. Sci. Rep. 8, 1–17 (2018).
Toni, M. et al. Environmental temperature variation affects brain protein expression and cognitive abilities in adult zebrafish (Danio rerio): A proteomic and behavioural study. J. Proteomics 204, 103396 (2019).
Kelly, A. L., O’Flaherty, F. & Fox, P. F. Indigenous proteolytic enzymes in milk: A brief overview of the present state of knowledge. Int. Dairy J. 16, 563–572 (2006).
Acknowledgements
This work was funded by the University of Milan, Grant Piano di Sostegno per la Ricerca, Anno 2018—Linea 2—PSR2018_DIP_026 Milk Lipidomics and Peptidomics. We acknowledge the UNITECH OMICS of the University of Milan for MS data acquisition, and the support of the APC central fund of the University of Milan to M.F.A.
Author information
Authors and Affiliations
Contributions
Conceptualization: F.C., M.F.A., M.A., V.B., R.P., F.T., G.T., E.D.C., D.V. Study coordination: M.F.A., F.C. Collection of samples, G.D.V., G.C. Milk analysis and identification of pathogens: D.V. Formal analysis: M.F.A., E.M.M., M.P. Data curation and visualization: M.F.A., E.M.M. Writing—original draft: MFA. Writing—review and editing: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Addis, M.F., Maffioli, E.M., Penati, M. et al. Peptidomic changes in the milk of water buffaloes (Bubalus bubalis) with intramammary infection by non-aureus staphylococci. Sci Rep 12, 8371 (2022). https://doi.org/10.1038/s41598-022-12297-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-12297-z
- Springer Nature Limited