Abstract
We sought to examine whether dietary intakes may affect the relationship between ApoB EcoRI and lipid profile, as well as serum inflammatory markers, in patients with type 2 diabetes (T2DM). This current study consisted of 648 diabetic patients. Dietary intake was calculated by a food frequency questionnaire. Biochemical markers (high-density lipoprotein (HDL), total cholesterol (TC), LDL, TG, CRP, IL-18, PGF2α) were measured based on standard protocols. Genotyping of the Apo-B polymorphisms (rs1042031) was conducted by the PCR–RFLP method. The gene-diet interactions were evaluated using GLMs. In comparison to GG homozygotes, A-allele carriers with above the median -CHO intake (≥ 54 percent of total energy) had considerably greater TC and PGF2a concentrations. Furthermore, as compared to GG homozygotes, A-allele carriers with above the median protein intake (≥ 14 percent of total energy) had higher serum levels of TG (P = 0.001), CRP (P = 0.02), TG/HDL (P = 0.005), and LDL/HDL (P = 0.04) ratios. Moreover, A-allele carriers with above the median total fat intake (≥ 35 percent of total calories) had significantly higher TC level (P = 0.04) and LDL/HDL (P = 0.04) ratios compared to GG homozygotes. Furthermore, when compared to GG homozygotes, A-allele carriers who consumed above the median cholesterol (> 196 mg) had greater TG (P = 0.04), TG/HDL (P = 0.01) ratio, and IL-18 (P = 0.02). Furthermore, diabetic patients with the GA, AA genotype who consume above the median cholesterol had lower ghrelin levels (P = 0.01). In terms of LDL/HDL ratio, ApoB EcoRI and dietary intakes of specific fatty acids (≥ 9 percent for SFA and ≥ 12 percent for MUFA) had significant interaction. LDL/HDL ratio is greater in A-allele carriers with above the median SFA intake (P = 0.04), also when they consumed above the median MUFA this association was inverse (P = 0.04). Our study showed that plasma lipid levels in participants carrying the (AA or AG) genotype were found to be more responsive to increasing the percentage of energy derived from dietary fat, CHO, protein, SFA, and cholesterol consumption. Therefore, patients with a higher genetic susceptibility (AA or AG) seemed to have greater metabolic markers with a higher percentage of macronutrient consumption. Also, ApoB EcoRI correlations with metabolic markers might be attenuated with above the median MUFA consumption.
Similar content being viewed by others
Introduction
Type 2 diabetes mellitus (T2DM) is now one of the most common non-communicable diseases in the world. This is due to the fact that T2DM is recognized as one of the leading causes of death worldwide1. According to current trends, the number of fatalities caused by T2DM is expected to rise from 285 million adults in 2012 to over 439 million individuals by 20302,3. In diabetes mellitus, dyslipidemia is one of the most important risk factors for cardiovascular disease4. Indeed, T2DM is correlated to altered lipid profiles, including raised TG and/or low HDL, a condition known as metabolic dyslipidemia5,6.
Findings indicate that genetic differences have a key role in the prevention and treatment of a variety of chronic diseases, including diabetes and dyslipidemia7,8,9,10. Genetic variables are known to be independent predictors for both dyslipidemia and diabetes11,12. As a result, it is critical to find the genes that cause T2DM-related metabolic problems and to develop novel treatments13,14. Particularly, the gene encoding APOB is one of the most widely anticipated genes that could alter lipid metabolism in T2DM15. The Apo-B polymorphism EcoRI (rs1042031) has been found to alter total and LDL cholesterol levels16,17.
Diet plays a significant impact in the initiation and progression of T2DM-related chronic diseases18,19. Additionally, some studies have found an association between dietary macronutrient intake and metabolic diseases such as dyslipidemia and inflammatory indicators20,21. Several recent investigations have found evidence that the response of plasma lipid levels to dietary changes is influenced by genetic factors22.
According to previous research, the ApoB EcoRI (R-) variant reduces the variation in total cholesterol levels between identical twins. Because identical twins had the same genetic code, differences in cholesterol levels between co-twins have to be attributable to a variety of circumstances, including dietary differences23,24,25. Interestingly, multiple studies have suggested that EcoRI genetic diversity interacts with other risk variables such as dietary fatty acid and cholesterol intake, indicating a positive relationship between EcoRI and hypercholesterolemia26.
To the best of the authors' knowledge, there has been no work thus far aimed at evaluating the interaction between EcoRI rs1042031 variant and macronutrient intake on serum levels of lipid and inflammatory markers, leptin and ghrelin. The objectives of this study were to evaluate the association between metabolic markers and EcoRI polymorphism and identify the interaction of EcoRI rs1042031 with the types of dietary macronutrient intake, in regards of metabolic risk factor in T2DM patients.
Methods and materials
Study population
648 patients (252 men and 396 women) were selected for this cross-sectional study from Tehran's referral Diabetes Centers. The present study established in 2019, and the information includes dietary intakes and genetic data collected in previous studies27. Subjects with fasting blood glucose (FBG) ≥ 126 mg/dl, taking diabetic medicines, or subjects fulfilling both criteria were included. Pregnant and lactating women, patients being treated with insulin, and drug-addicted subjects were excluded. The interview-based evaluations were also used to collect general information such as age, gender, work and education status, smoking and alcohol behaviors, lipid-lowering medicine use, T2DM duration, and family history of the disease. All methods were performed in accordance with the Declaration of Helsinki.
Anthropometric and physical activity assessments
Anthropometric values were measured, including weight without shoes on a digital scale with 0.1-kg reliability and height without shoes on 0.1-cm-accuracy height gauges. Weight (kg)/height2 was used to determine the body mass index (BMI) (m). The Tehran University of Medical Sciences Ethics Committee approved the study, and all patients signed an informed consent form before being enrolled. The International Physical Activity Questionnaire (IPAQ) modified version was used to assess physical activity. The IPAQ's reliability and validity have previously been tested among Iranian adults28.
Laboratory tests
All blood samples were obtained at TUMS' nutrition laboratory after 12–14 h of fasting, centrifuged, and stored at − 80 °C. Enzymatic methods were used to quantify serum lipid biomarkers such as HDL, TC, LDL, and TG (Pars Azmun Co., Tehran, Iran). The levels of leptin and ghrelin in the plasma were also determined by the ELISA method (Bioassay Technology Co, China, and Mediagnost, Germany, respectively). Inflammatory markers (CRP, IL-18, and PGF2) were evaluated according to the manufacturer's instructions using enzyme-linked immunosorbent assay kits (R&D Systems, Techne Corporation, Minneapolis, MN).
Genotyping
The salting-out procedure was used to extract genomic DNA. The polymerase chain reaction was used to determine genotyping (PCR–RFLP). The following primers were used to amplify rs1042031: (forward primer: CACTGGGACCTACCAAGAG; reverse primer: CACTGGGACCTACCAAGAG) CTCGAAAGGAAGTGTAATCAC (reverse primer). The reaction included 35 cycles of denaturation at 94 °C for three minutes, followed by 35 cycles of 30 s at 94 °C, 30 s at 56 °C, 40 s at 72 °C, and one cycle of seven minutes at 72 °C and five minutes at 4 °C. PCR-Thermocycler was used to perform the PCR reaction (PEQLAB; GmbH96; Germany).
Dietary assessments
Expert dietitians evaluated dietary intakes using a 147-item, semi-quantitative FFQ29. The frequency of each food item consumed over the previous year was reported by the participants and calculated to grams per day using home measurements. The Iranian Food Composition Table (FCT) and N4 software were used to calculate total energy and dietary elements. The nutritional density approach was also used to adjust this dietary intake for total energy intake. In our study, dietary intakes were categorized into two groups according to the median value, below and above or equal based on our pervious study27.The carbohydrate, protein, total fat, saturated fatty acid, MUFA and cholesterol were categorized into two groups, according to their medians of the population (54, 14, 35, 9,12 percent and 196 mg, respectively).
Statistical analysis
Initially, the Kolmogorov–Smirnov test was applied to determine distribution normality. The 2 test was used to analyze the Hardy–Weinberg equilibrium and categorical variable comparison. Independent Samples T-tests were used to compare quantitative variables between genotypes. The interaction between EcoRI polymorphisms and macronutrient intake on lipid profile, as well as serum inflammatory markers were verified by Generalized Linear Models (GLZM). We used dietary intake groups and genotype categories as a fixed factor and cofounding factors as a covariate then we made an interaction model. The GG genotype has no risk allele and is used as a reference. Moreover, the intake of macronutrients below the median is used as a reference. Biochemical markers (high-density lipoprotein (HDL), total cholesterol (TC), LDL, TG, CRP, IL-18, PGF2α), were response variables, while EcoRI genotypes and each macronutrient intake were considered as factor variables. Model 1: unadjusted; Model 2: adjusted for age, gender, physical activity, alcohol consumption and lipid-lowering medications; Model 3: adjusted for variables in model 2 plus for smoking, total energy intakes and fiber intake. All data were statistically analyzed using SPSS (version 25; SPSS Inc., IL). P value < 0.05 was used to determine statistical significance.
Result
A total of 648 Iranian diabetic patients were categorized based on rs1042031 genotypes and divided into two groups: GA, AA genotypes (n = 145), GG genotype (n = 503). Genotypic and allelic frequencies of EcoRI in men and women are presented in Table 1. Besides, Genotype frequency for dominant homozygote (GG), heterozygote (GA), and recessive homozygote (AA) was 77.7%, 21.1%, and 1.2%, respectively have shown in Table 1. The genotype frequency (P = 0.23) shows no divergence from Hardy–Weinberg Equilibrium (HWE). Furthermore, there was no significant correlation found between the ApoB EcoRI polymorphism and inflammatory markers or dietary intake (P > 0.05). Diabetic patients with GA, AA genotype had more BMI (P = 0.004) and WC (P = 0. 004) compared to GG homozygotes. Furthermore, we revealed a correlation between the ApoB EcoRI polymorphism and serum HDL-C concentration. As a result, plasma HDL-C levels in A-allele carriers were considerably greater than in GG homozygotes. Also, no correlation was found between this polymorphism and other lipid variables (P ≥ 0.05) (Table 2).
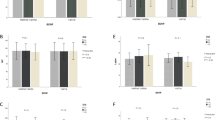
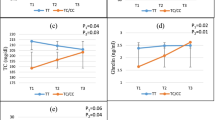
The interactions between our genotype categories (GA, AA/ GG) and median value of total energy intake (54% for CHO, 14% for protein, 35% for total fat, 9% for SFA and 12% for MUFA) and 196 mg for cholesterol on lipid profiles and inflammatory markers are shown in Tables 3 and 4, respectively. Significant results are shown in Fig. 1a–e.
(a) Interaction between the ApoB ECORI and dietary carbohydrate intakes on serum TC, PGF2a level and LDL/HDL rati. P values for the interaction obtained in multivariate models using GLMs. Model 1: unadjusted; Model 2: adjusted for age, gender, physical activity, alcohol consumption and lipid-lowering medications; Model 3: adjusted for variables in model 2 plus for smoking, total energy intakes and fiber intake. Error bar: SEM. (b) Interaction between the ApoB ECORI and dietary protein intakes on serum TG, hs-CRP level, LDL/HDL and TG/HDL ratio. P values for the interaction obtained in multivariate models using GLMs. Model 1: unadjusted; Model 2: adjusted for age, gender, physical activity, alcohol consumption and lipid-lowering medications; model 3: adjusted for variables in model 2 plus for smoking, total energy intakes and fiber intake. Error bar: SEM. (c) Interaction between the ApoB ECORI and total fat intakes on serum TC level and LDL/HDL ratio. P values for the interaction obtained in multivariate models using GLMs. Model 1: unadjusted; Model 2: adjusted for age, gender, physical activity, alcohol consumption and lipid-lowering medications; Model 3: adjusted for variables in model 2 plus for smoking, total energy intakes and fiber intake. Error bar: SEM. (d) Interaction between the ApoB ECORI and cholesterol intakes on serum TG, ghrelin, IL-18 and level and TG/HDL ratio. P values for the interaction obtained in multivariate models using GLMs. Model 1: unadjusted; Model 2: adjusted for age, gender, physical activity, alcohol consumption and lipid-lowering medications; Model 3: adjusted for variables in model 2 plus for smoking, total energy intakes and fiber intake. Error bar: SEM. (e) Interaction between the ApoB ECORI and dietary specific fatty acids intakes containing SFA and MUFA intake on LDL/HDL ratio. P values for the interaction obtained in multivariate models using GLMs. Model 1: unadjusted; Model 2: adjusted for age, gender, physical activity, alcohol consumption and lipid-lowering medications; Model 3: adjusted for variables in model 2 plus for smoking, total energy intakes and fiber intake. Error bar: SEM.
According to our findings on the gene-diet interaction, ApoB EcoRI polymorphism and CHO intake had a significant interaction on serum TC (P = 0.03) and PGF2a level (P = 0.03) in three models. TC and PGF2a concentrations were greatly increased in patients with (AA or AG) genotypes with higher than median of CHO intake (≥ 54% of total energy) compared to the GG genotype. Besides, we observed significant results on LDL/HDL ratio after adjustment for potential confounders in two models (P = 0.04). Therefore, A-allele carriers with higher than median of CHO intake have a higher LDL/HDL ratio compared to GG homozygotes (Fig. 1a).
Also, we revealed that A-allele carriers who consumed a more than median of protein diet (≥ 14 percent of total energy) had greater TG and CRP levels in their plasma, as well as a higher TG/HDL and LDL/HDL ratio, compared to GG genotypes (P = 0.001, P = 0.02, P = 0.005 and P = 0.04 respectively) (Fig. 1b). Moreover, other significant results were detected between ApoB EcoRI and total fat intake on TC level and LDL/HDL ratio in the adjustment model (model 3) (P = 0.04 and P = 0.04 respectively). The A-allele carriers with higher than median intake of total fat (≥ 35% of total energy) revealed significantly increased serum TC level and LDL/HDL ratio compared to GG homozygotes (Fig. 1c).
In crude and two adjusted models, we found a significant interaction between the ApoB EcoRI polymorphism and cholesterol intake on TG, TG/HDL ratio, and IL-18. Therefore, A-allele carriers with above than median cholesterol intake (> 196 mg) have higher TG, TG/HDL ratio, and IL-18 compared to GG homozygotes (P = 0.04, P = 0.01, and P = 0.02 respectively).
Dietary components such as total fat and type of dietary fat, as well as HOMA-IR, are positively related to serum leptin concentrations while serum ghrelin concentrations are negatively related30. In the present study, diabetic patients with GA, AA genotype who consumed higher than median cholesterol have lower ghrelin concentration (P = 0.01) (Fig. 1d).
Finally, we found a considerable interaction between the ApoB EcoRI variant and various fatty acid intakes (≥ 9 percent for SFA, ≥ 12 percent for MUFA) on the LDL/HDL ratio. A-allele carriers who consume above than median of SFA intake have a higher LDL/HDL ratio (P = 0.04), but this relationship is inverted (P = 0.04) when they consume higher than median median of MUFA (Fig. 1e).
Discussion
The ApoB EcoRI polymorphism was found to have a significant relationship with serum HDL-C levels and obesity indices in the current work. A-allele carriers are more sensitive to dyslipidemia and CVD than GG homozygotes because they have a considerably higher general obesity risk than GG genotype17. However, HDL concentration was higher between A –allele carriers than GG genotype. These findings are inconsistent with other study has shown participant with A/A genotype reported considerably greater HDL-C and apoA1 levels, as well as decreased total protein and albumin levels in their plasma compared with GA or GG carriers, however no significant differences in BMI was found17. Based on previous studies, amino acid 4154 is located in the apoB protein's C-terminal domain, which is essential for lipid association. This polymorphism induced amino acid substitution has the potential to have a significant impact on protein function and, as a result, lipid levels31. Our hypothesis is supported by the findings of this investigation, however, it contradicts the findings of a previous study which revealed that no influence of the EcoRI polymorphism on serum lipid levels in healthy subjects32.
Analyzing the interactions between ApoB EcoRI polymorphism and macronutrient intake on lipid profiles showed significant interaction. A-allele carriers with above than median dietary CHO, protein, total fat and SFA intake showed a significantly higher serum LDL/HDL ratio compared to GG homozygotes, also when they consumed higher than median of MUFA this association was inverse. Moreover, we observed that A-allele carriers with a above than median of CHO and total fat had considerably greater serum levels of TC than GG genotype carriers. Furthermore, when protein intake was ≥ 14% of total energy or cholesterol intake more than 196 mg, we observe a significantly higher TG serum level and TG/HDL ratio between A-allele carriers compared to GG genotype. The EcoRI polymorphism influenced the response of plasma TC, LDL, TG, LDL/HDL, and TG/HDL ratios to diet in the current investigation. When AA, AG patients were shifted from a below to above median of fat inrake, or from a below to above median of cholesterol diet, their plasma lipids increased the most. In particular, we found a unique interaction between the ApoB EcoRI polymorphism and macronutrient intake on inflammatory markers and appetite-related hormones in the current study. When cholesterol intake was higher than median intake, patients with AG and AA genotypes displayed significantly higher ghrelin and IL-18, also when consumed than median protein intake they have shown higher CRP. On the other hand, above the median of CHO intake, carriers of A allele were associated with higher PGF2a serum levels than GG genotype.
According to these findings, higher than median of MUFA consumption may reduce the EcoRI rs1042031 associations with cardiometabolic markers with a lower LDL/HDL ratio, but individuals with a higher genetic susceptibility seemed to have an increased chance of cardiometabolic disease, which was related to more than median dietary CHO, protein, total fat, SFA, and cholesterol consumption. Thus, considering adherence to a diet rich in CHO, protein, total fat, SFA and cholesterol can be expected to see the more atherogenic effect in subjects with the AA OR AG genotype.
To the best of our knowledge, this is the first study that shows interaction between dietary intake and ApoB EcoRI on dyslipidemia, inflammatory markers and appetite-related hormones. The EcoRI allele was shown to be more common in the responders than the non-responders between fifty-one participants who were classed as different dietary intakes33. Gylling et al. evaluated the effect of ApoB EcoRI and food on lipid profile in a study similar to ours. In comparison to patients with the GG genotype, carriers of the ApoB EcoRI R + (A) allele showed the most pronounced LDL cholesterol increase and fractional metabolism rate for LDL when shifting from low to high cholesterol and SFA intake34 In line with our findings, Rantala et al. showed that AA individuals had significantly higher total and LDL-cholesterol responses than GG patients in a meta-analysis of seven investigations. Apo-B EcoRI Polymorphisms is responsiveness to diet high in fat contains (45 percent of energy was derived from CHO, 18 percent from protein, and 36 percent from fat and daily cholesterol intake was 420 mg) which is induced a greater increase LDL level in patients carrying the AA genotype compared in someone with the GA or GG genotype, but no significant correlation was found in the low-fat low-cholesterol diet26. Furthermore, Friedlander et al. found that when participants heterozygous for the less common apo B EcoRI allele (AG) consumed a high dietary fat intake, plasma cholesterol and LDL-C levels changed significantly compared to homozygous carriers35. However, some studies have shown no association between EcoRI polymorphism and dietary intake. According to a recent study, a higher intake of wheat- or oat-bran supplementation as dietary fiber did not affect plasma apolipoprotein B-containing lipoprotein concentrations when the ApoB EcoRI polymorphismon was examined36. Another prospective double-blind crossover dietary study found that EcoRI has not fully responded to dietary fat intake. Dietary participants were randomized to drink either a fat- and cholesterol-free liquid supplement or one that contained fat (30–36 g) and cholesterol for three weeks (650–780 mg)37. Friedlander et al. also found no evidence of an association between dietary (low-SFA and high-PUFA) and the apoB EcoRI polymorphism on metabolic indicators38. Except for PUFA intake in our study, other interactions identified for CHO, protein, total fat, SFA, MUFA, and cholesterol of diet on dyslipidemia, inflammatory markers, and appetite-related hormones.
The EcoRI polymorphism's biological function is unclear, hence the mechanism by which it alters metabolic indicators in response to dietary intake is unknown. Variations in the synthesis rate or apo B and all lipoproteins containing apo B catabolism with variable food intake might explain this connection. Previous research has found it difficult to build conclusions about the significance of this polymorphism for cholesterol metabolism since the results are few and contradictory, ranging from no association with cholesterol levels to a strong correlation39,40,41,42,43,44,45,46 to some association with other lipid markers includes TG and VLDL levels40,46,47,48,49. This polymorphism is caused by a single base-pair mutation in the apo B gene's coding area (exon 29), which modifies the amino acid sequence and makes a functional role50. Furthermore, because this polymorphism is found in ApoB's carboxy-terminal tail, it may impact ApoB's ability to connect to LDL receptors51. It's conceivable that this amino acid alteration affects the encoded apo B molecules' LDL receptor affinity45. The apo B EcoRI polymorphism was also connected to cholesterol absorption efficiency in a previous investigation. Cholesterol production and LDL receptor function would be significantly suppressed when cholesterol uptake capacity is high, as it is in the R + patients52,53,54. These findings suggest that common polymorphism of the apo B EcoRI could cause various responses in plasma TC and LDL levels when dietary cholesterol content is significantly changed. It seems these modification was done by changing apo B secretion, LDL receptor affinity and structural stability or combinations of lipoproteins containing apo B with other lipoproteins or different kinds of enzymes such as LP26.
In this term, experimental studies have shown that compared to wild-type (WT) mice on normal diets, Apobec-1 knockout animals (KO) containing apoB100 but not apoB48 effectively make chylomicrons and can absorb dietary fat without obvious deficiency. The frequency of chylomicron secretion was influenced by the migration of apo-B-containing particles into the lumen of the smooth endoplasmic reticulum when dietary TG was low. The rate-limiting process appeared to be the insertion of TG to the apo-B-containing molecules when dietary TG is high (Western diet). In this regard, the findings indicate that apo-B100 construction in chylomicrons is inefficient in the early sTGes of chylomicron assembly in the KO mice, but that intracellular TG accumulation (due to a high-fat diet) surpasses this inefficiency and promotes chylomicron secretion at a higher level. This could also explain why chylomicrons from fat-fed KO mice are 66 percent larger than those from WT mice consuming the same diet. The accumulating TG in the enterocytes may result in bigger TG particles that can be assembled with apo-B100-containing precursors55. Therefore, Another possible pathway by which the EcoRI in this investigation could influence the association between dietary macronutrient intake and lipid profile is at the absorption or chylomicron assembly sTGes in the small intestine.
Furthermore, the amount of hepatic apoB release, and thus LDL density, might also be affected by the amount of fatty acid taken up by peripheral tissues. Some authors observed that decreased fatty acid uptake might lead to an increased source of hepatic TG. The resultant raised levels of fatty acids generated in the liver would lead to increased hepatic secretion of apoB containing lipoproteins56. Genest et al. backed up this hypothesis by experimental study, it has been demonstrated that, normotriglyceridaemic (NTG) hyper-apoB participants remove TGs more slowly following an oral fat load versus healthy subjects57, and that decreased fatty acid uptake by peripherial tissues leads to slower TG synthesis in NTG hyper-apoB individuals' adipocytes58.
Moreover, according to a previous study, decreased ApoB synthesis in the body leads to increased production of leptin and ghrelin, which leads to resistance and obesity59. Furthermore, leptin is responsive to dietary changes, particularly in diabetic patients60. Increased ghrelin levels are linked to lower HDL cholesterol levels and higher LDL cholesterol levels61. Dietary components such as total fat and type of dietary fat, as well as HOMA-IR, are positively related to serum leptin concentrations while serum ghrelin concentrations are negatively related30. BMI, nonHDL, and fibrinogen were found to be strong predictors of inflammatory marker concentrations in a multinomial logistic regression study62. Factors such as lipid metabolism, blood pressure, obesity, glucose metabolism, renal function, and lifestyle had varying effects on the development of the disease through inflammatory factors63.
Therefore, in this study, the pathway through which macronutrients affect the genetic risk provided by EcoRI polymorphisms is unknown; more research is needed to identify such gene-diet treatments in diabetic patients. The Apo B EcoRI polymorphism could be a positive figure for further information on how to manage metabolic disorders in T2DM patients.
Limitations and strengths
Our study is the first study to examine the existence of Apo-B EcoRI gene interaction with macronutrient intake on lipid profile and inflammatory markers in T2DM patients to date. Our study was included all different types of dietary intake. We adjusted to wide range of confounders. Although we provide a novel addition to the literature, some limitations should be considered in the interpretation of this study. It is impossible to argue that our study's cross-sectional design generated any causality; the use of FFQ for dietary assessing, which may have resulted in memory bias. Furthermore, our participants were from the Iranian country which may not be generalized due to racial and regional differences. ApoB EcoRI mutations altered metabolic diseases that responded to dietary changes in our research sample. Although other components might also be implicated and should be investigated further in larger research to identify their probable impact in plasma lipid response to diet and so help to reveal the mechanisms behind differential responsiveness. This may be due to the interaction between unmeasured genetic variation or environmental factors that were different between the healthy individuals and diabetic patients.
Conclusion
Our study showed that plasma lipid concentrations in subjects with the deletion allele (AA or AG) may also be more responsive to an increased percenTGe of energy from dietary fat, CHO, protein, SFA, and cholesterol consumption. Therefore, individuals with greater genetic predisposition (AA or AG) appeared to have higher metabolic markers with higher percenTGe of macronutrient consumption and higher MUFA consumption might attenuate the ApoB EcoRI associations with metabolic markers.
Data availability
The data are not publicly available due to containing private information of participants. Data are however available from the authors upon reasonable request and with permission of Fariba Koohdani.
Abbreviations
- T2DM:
-
Type 2 diabetes mellitus
- HDL:
-
High-density lipoprotein cholesterol
- LDL:
-
Low-density lipoprotein cholesterol
- VLDL:
-
Very low-density lipoprotein
- TG:
-
Triglyceride
- TC:
-
Total cholesterol
- SNP:
-
Single nucleotide polymorphism
- PCR–RFLP:
-
Polymerase chain reaction–restriction fragment length polymorphism
- GLM:
-
Generalized linear model
- BMI:
-
Body mass index
- WC:
-
Waist circumference
References
Zimmet, P., Alberti, K. G. & Shaw, J. Global and societal implications of the diabetes epidemic. Nature 414, 782–787. https://doi.org/10.1038/414782a (2001).
Wild, S., Roglic, G., Green, A., Sicree, R. & King, H. Global prevalence of diabetes: estimates for the year 2000 and projections for 2030. Diabetes Care 27, 1047–1053. https://doi.org/10.2337/diacare.27.5.1047 (2004).
Shaw, J. E., Sicree, R. A. & Zimmet, P. Z. Global estimates of the prevalence of diabetes for 2010 and 2030. Diabetes Res. Clin. Pract. 87, 4–14. https://doi.org/10.1016/j.diabres.2009.10.007 (2010).
Mooradian, A. D. Dyslipidemia in type 2 diabetes mellitus. Nat. Clin. Pract. Endocrinol. Metab. 5, 150–159. https://doi.org/10.1038/ncpendmet1066 (2009).
Taskinen, M.-R. Diabetic dyslipidaemia: from basic research to clinical practice. Diabetologia 46, 733–749 (2003).
Association, A. D. Dyslipidemia management in adults with diabetes. Diabetes Care 27, s68–s71 (2004).
Fuchsberger, C. et al. The genetic architecture of type 2 diabetes. Nature 536, 41–47. https://doi.org/10.1038/nature18642 (2016).
García-Giustiniani, D. & Stein, R. Genetics of dyslipidemia. Arq. Bras. Cardiol. 106, 434–438. https://doi.org/10.5935/abc.20160074 (2016).
McCarthy, M. I. Genomics, type 2 diabetes, and obesity. N. Engl. J. Med. 363, 2339–2350. https://doi.org/10.1056/NEJMra0906948 (2010).
Abaj, F., Sotoudeh, G., Karimi, E., Rafiee, M. & Koohdani, F. Interaction between the dietary indices and PPAR-γ Pro12Ala gene variants on cardiovascular risk factors in patients with type 2 diabetes mellitus. Int. J. Clin. Pract. https://doi.org/10.1111/ijcp.14307 (2021).
Qi, Q., Liang, L., Doria, A., Hu, F. B. & Qi, L. Genetic predisposition to dyslipidemia and type 2 diabetes risk in two prospective cohorts. Diabetes 61, 745–752. https://doi.org/10.2337/db11-1254 (2012).
Miranda-Lora, A. L., Vilchis-Gil, J., Juárez-Comboni, D. B., Cruz, M. & Klünder-Klünder, M. A genetic risk score improves the prediction of type 2 diabetes mellitus in Mexican youths but has lower predictive utility compared with non-genetic factors. Front. Endocrinol. https://doi.org/10.3389/fendo.2021.647864 (2021).
Abaj, F., Rafiee, M. & Koohdani, F. Interaction between CETP polymorphism and dietary insulin index and load in relation to cardiovascular risk factors in diabetic adults. Sci. Rep. 11, 15906. https://doi.org/10.1038/s41598-021-95359-y (2021).
Abaj, F., Rafiee, M. & Koohdani, F. Interaction between dietary total antioxidant capacity and BDNF Val66Met polymorphism on lipid profiles and atherogenic indices among diabetic patients. Sci. Rep. 11, 19108. https://doi.org/10.1038/s41598-021-98663-9 (2021).
Bentzen, J., Poulsen, P., Vaag, A. & Fenger, M. Further studies of the influence of apolipoprotein B alleles on glucose and lipid metabolism. Hum. Biol. 75, 687–703 (2003).
Delghandi, M. et al. DNA polymorphisms of the apolipoprotein B gene (XbaI, EcoRI, and MspI RFLPs) in Norwegians at risk of atherosclerosis and healthy controls. Acta Cardiol. 54, 215–225 (1999).
Hu, P. et al. Effect of apolipoprotein B polymorphism on body mass index, serum protein and lipid profiles in children of Guangxi, China. Ann. Hum. Biol. 36, 411–420. https://doi.org/10.1080/03014460902882475 (2009).
Tucker, K. L. In Present Knowledge in Nutrition 11th edn (eds Marriott, B. P. et al.) 329–345 (Academic Press, 2020).
Sami, W., Ansari, T., Butt, N. S. & Hamid, M. R. A. Effect of diet on type 2 diabetes mellitus: a review. Int. J. Health Sci. 11, 65–71 (2017).
Silveira, B. K. S. et al. Dietary pattern and macronutrients profile on the variation of inflammatory biomarkers: scientific update. Cardiol. Res. Pract. 2018, 4762575. https://doi.org/10.1155/2018/4762575 (2018).
Mizgier, M. et al. Relation between inflammation, oxidative stress, and macronutrient intakes in normal and excessive body weight adolescent girls with clinical features of polycystic ovary syndrome. Nutrients https://doi.org/10.3390/nu13030896 (2021).
Abbey, M. The influence of apolipoprotein polymorphism on the response to dietary fat and cholesterol. Curr. Opin. Lipidol. 3, 12–16 (1992).
Berg, K. DNA polymorphism at the apolipoprotein B locus is associated with lipoprotein level. Clin. Genet. 30, 515–520. https://doi.org/10.1111/j.1399-0004.1986.tb01920.x (1986).
Berg, K. Genetics of coronary heart disease and its risk factors. Ciba Found. Symp. 130, 14–33. https://doi.org/10.1002/9780470513507.ch3 (1987).
Berg, K. Twin studies of coronary heart disease and its risk factors. Acta Genet. Med. Gemellol. 36, 439–453. https://doi.org/10.1017/s0001566000006814 (1987).
Rantala, M., Rantala, T. T., Savolainen, M. J., Friedlander, Y. & Kesäniemi, Y. A. Apolipoprotein B gene polymorphisms and serum lipids: meta-analysis of the role of genetic variation in responsiveness to diet. Am. J. Clin. Nutr. 71(3), 713–724 (2000).
Rafiee, M. et al. The interaction between apolipoprotein B insertion/deletion polymorphism and macronutrient intake on lipid profile and serum leptin and ghrelin levels in type 2 diabetes mellitus patients. Eur. J. Nutr. 58, 1055–1065. https://doi.org/10.1007/s00394-018-1621-5 (2019).
Assessment of Physical Activity of Adolescents in Isfahan. J. Shahrekord Univ. Med. Sci. 3(2), 27–33 (2001).
Esmaillzadeh, A., Mirmiran, P. & Azizi, F. Whole-grain intake and the prevalence of hypertriglyceridemic waist phenotype in Tehranian adults. Am. J. Clin. Nutr. 81, 55–63. https://doi.org/10.1093/ajcn/81.1.55 (2005).
Polak, A. M. et al. The association of serum levels of leptin and ghrelin with the dietary fat content in non-obese women with polycystic ovary syndrome. Nutrients https://doi.org/10.3390/nu12092753 (2020).
Bentzen, J., Jørgensen, T. & Fenger, M. The effect of six polymorphisms in the Apolipoprotein B gene on parameters of lipid metabolism in a Danish population. Clin. Genet. 61, 126–134. https://doi.org/10.1034/j.1399-0004.2002.610207.x (2002).
Saha, N., Tay, J. S. & Humphries, S. E. Apolipoprotein B-gene DNA polymorphisms (XbaI and EcoRI), serum lipids, and apolipoproteins in healthy Chinese. Genet. Epidemiol. 9, 1–10. https://doi.org/10.1002/gepi.1370090103 (1992).
Abbey, M. Apolipoprotein B gene polymorphism associates with plasma cholesterol changes induced by dietary fat and cholesterol. Nutr Metab. Cardiovasc. Dis. 1, 10–12 (1991).
Gylling, H., Kontula, K., Koivisto, U.-M., Miettinen, H. E. & Miettinen, T. A. Polymorphisms of the genes encoding apoproteins A-I, B, C-III, and E and LDL receptor, and cholesterol and LDL metabolism during increased cholesterol intake. Arterioscler. Thromb. Vasc. Biol. 17, 38–44. https://doi.org/10.1161/01.ATV.17.1.38 (1997).
Friedlander, Y., Leitersdorf, E., Vecsler, R., Funke, H. & Kark, J. The contribution of candidate genes to the response of plasma lipids and lipoproteins to dietary challenge. Atherosclerosis 152(1), 239–248 (2000).
Hegele, R. A. et al. Genetic variation associated with differences in the response of plasma apolipoprotein B levels to dietary fibre. Clin. Sci. 85(3), 269–275 (1993).
Abbey, M. et al. Restriction fragment length polymorphism of the apolipoprotein B gene and response to dietary fat and cholesterol. Can. J. Cardiol. 11(Suppl G), 79g–85g (1995).
Friedlander, Y., Berry, E. M., Eisenberg, S., Stein, Y. & Leitersdorf, E. Plasma lipids and lipoproteins response to a dietary challenge: analysis of four candidate genes. Clin. Genet. 47, 1–12. https://doi.org/10.1111/j.1399-0004.1995.tb03913.x (1995).
Talmud, P. J. et al. Apolipoprotein B gene variants are involved in the determination of serum cholesterol levels: a study in normo- and hyperlipidaemic individuals. Atherosclerosis 67, 81–89. https://doi.org/10.1016/0021-9150(87)90267-x (1987).
Peacock, R. et al. Apolipoprotein B gene polymorphisms, lipoproteins and coronary atherosclerosis: a study of young myocardial infarction survivors and healthy population-based individuals. Atherosclerosis 92, 151–164. https://doi.org/10.1016/0021-9150(92)90274-k (1992).
Myant, N. B. et al. Restriction fragment length polymorphisms in the apo B gene in relation to coronary artery disease. Atherosclerosis 77, 193–201. https://doi.org/10.1016/0021-9150(89)90081-6 (1989).
Monsalve, M. V. et al. DNA polymorphisms of the gene for apolipoprotein B in patients with peripheral arterial disease. Atherosclerosis 70, 123–129. https://doi.org/10.1016/0021-9150(88)90106-2 (1988).
Houlston, R. S., Turner, P. R., Lewis, B. & Humphries, S. E. Genetic epidemiology of differences in low-density lipoprotein (LDL) cholesterol concentration: possible involvement of variation at the apolipoprotein B gene locus in LDL kinetics. Genet. Epidemiol. 7, 199–210. https://doi.org/10.1002/gepi.1370070304 (1990).
Hegele, R. A. et al. Apolipoprotein B-gene DNA polymorphisms associated with myocardial infarction. N. Engl. J. Med. 315, 1509–1515. https://doi.org/10.1056/nejm198612113152403 (1986).
Gallagher, J. J. & Myant, N. B. Does the EcoRI polymorphism in the human apolipoprotein B gene affect the binding of low density lipoprotein to the low density lipoprotein receptor?. Arterioscler. Thromb. J. Vasc. Biol. 12, 256–260. https://doi.org/10.1161/01.atv.12.2.256 (1992).
Demant, T. et al. Catabolic rate of low density lipoprotein is influenced by variation in the apolipoprotein B gene. J. Clin. Investig. 82, 797–802. https://doi.org/10.1172/jci113681 (1988).
Tybjaerg-Hansen, A., Nordestgaard, B. G., Gerdes, L. U. & Humphries, S. E. Variation of apolipoprotein B gene is associated with myocardial infarction and lipoprotein levels in Danes. Atherosclerosis 89, 69–81. https://doi.org/10.1016/0021-9150(91)90008-q (1991).
Paulweber, B., Friedl, W., Krempler, F., Humphries, S. E. & Sandhofer, F. Association of DNA polymorphism at the apolipoprotein B gene locus with coronary heart disease and serum very low density lipoprotein levels. Arteriosclerosis (Dallas, Tex.) 10, 17–24. https://doi.org/10.1161/01.atv.10.1.17 (1990).
Aalto-Setälä, K., Kontula, K., Sane, T., Nieminen, M. & Nikkilä, E. DNA polymorphisms of apolipoprotein A-I/C-III and insulin genes in familial hypertriglyceridemia and coronary heart disease. Atherosclerosis 66, 145–152. https://doi.org/10.1016/0021-9150(87)90190-0 (1987).
Shoulders, C. C. et al. Molecular cloning of human LDL apolipoprotein B cDNA. Evidence for more than one gene per haploid genome. Atherosclerosis 58, 277–289. https://doi.org/10.1016/0021-9150(85)90073-5 (1985).
Benn, M. et al. Common and rare alleles in apolipoprotein B contribute to plasma levels of low-density lipoprotein cholesterol in the general population. J. Clin. Endocrinol. Metab. 93, 1038–1045. https://doi.org/10.1210/jc.2007-1365 (2008).
Miettinen, T. A., Gylling, H., Vanhanen, H. & Ollus, A. Cholesterol absorption, elimination, and synthesis related to LDL kinetics during varying fat intake in men with different apoprotein E phenotypes. Arterioscler. Thromb. J. Vasc. Biol. 12, 1044–1052. https://doi.org/10.1161/01.atv.12.9.1044 (1992).
McCombs, R. J., Marcadis, D. E., Ellis, J. & Weinberg, R. B. Attenuated hypercholesterolemic response to a high-cholesterol diet in subjects heterozygous for the apolipoprotein A-IV-2 allele. N. Engl. J. Med. 331, 706–710. https://doi.org/10.1056/nejm199409153311104 (1994).
Gylling, H. & Miettinen, T. A. Cholesterol absorption and synthesis related to low density lipoprotein metabolism during varying cholesterol intake in men with different apoE phenotypes. J. Lipid Res. 33, 1361–1371 (1992).
Kendrick, J. S., Chan, L. & Higgins, J. A. Superior role of apolipoprotein B48 over apolipoprotein B100 in chylomicron assembly and fat absorption: an investigation of apobec-1 knock-out and wild-type mice. Biochem. J. 356(3), 821–827 (2001).
Sniderman, A., Brown, B. G., Stewart, B. F. & Cianflone, K. From familial combined hyperlipidemia to hyperapoB: unravelling the overproduction of hepatic apolipoprotein B. Curr. Opin. Lipidol. 3(2), 137–142 (1992).
Genest, J. et al. Hyperapobetalipoproteinemia. Plasma lipoprotein responses to oral fat load. Arteriosclerosis (Dallas, Tex.) 6, 297–304. https://doi.org/10.1161/01.atv.6.3.297 (1986).
Teng, B., Sniderman, A. D., Soutar, A. K. & Thompson, G. R. Metabolic basis of hyperapobetalipoproteinemia. Turnover of apolipoprotein B in low density lipoprotein and its precursors and subfractions compared with normal and familial hypercholesterolemia. J. Clin. Investig. 77, 663–672. https://doi.org/10.1172/jci112360 (1986).
Haas, M. E., Attie, A. D. & Biddinger, S. B. The regulation of ApoB metabolism by insulin. Trends Endocrinol. Metab. 24, 391–397. https://doi.org/10.1016/j.tem.2013.04.001 (2013).
Iqbal, J., Mascareno, E., Chua, S. & Hussain, M. M. Leptin-mediated differential regulation of microsomal triglyceride transfer protein in the intestine and liver affects plasma lipids. J. Biol. Chem. 295, 4101–4113. https://doi.org/10.1074/jbc.RA119.011881 (2020).
Slama, F. B. et al. Plasma levels of leptin and ghrelin and their correlation with BMI, and circulating lipids and glucose in obese Tunisian women. Asian Biomed. 9, 161–168. https://doi.org/10.5372/1905-7415.0902.382 (2015).
Jovicić, S., Ignjatović, S., Dajak, M., Kangrga, R. & Majkić-Singh, N. Association of lipid and inflammatory markers with C-reactive protein in cardiovascular risk assessment for primary prevention. Clin. Lab. 55, 411–419 (2009).
Lian, Y., Xie, L., Liu, Y. & Tang, F. Metabolic-related markers and inflammatory factors as predictors of dyslipidemia among urban Han Chinese adults. Lipids Health Dis. 18, 167. https://doi.org/10.1186/s12944-019-1109-1 (2019).
Acknowledgements
The authors would like to thank all patients who participate in this study.
Funding
This research was supported by Tehran University of Medical Sciences, Tehran, Iran. (15060).
Author information
Authors and Affiliations
Contributions
F.A. contributed to conception, design, data analyses, data interpretation, editing and manuscript drafting. F.K.: supervised the study. All authors approved the final manuscript for submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abaj, F., Koohdani, F. Macronutrient intake modulates impact of EcoRI polymorphism of ApoB gene on lipid profile and inflammatory markers in patients with type 2 diabetes. Sci Rep 12, 10504 (2022). https://doi.org/10.1038/s41598-022-13330-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-13330-x
- Springer Nature Limited