Abstract
Pancreatic adenocarcinoma (PAAD) has high mortality and a very poor prognosis. Both surgery and chemotherapy have a suboptimal therapeutic effect, and this caused a need to find new approaches such as immunotherapy. Therefore, it is essential to develop a new model to predict patient prognosis and facilitate early intervention. Our study screened out and validated the target molecules based on the TCGA-PAAD dataset. We established the risk signature using univariate and multivariate Cox regression analysis and used GSE62452 and GSE28735 to verify the accuracy and reliability of the model. Expanded application of PAAD-immune-related genes signature (-IRGS) on other datasets was conducted, and the corresponding nomograms were constructed. We also analyzed the correlation between immune-related cells/genes and potential treatments. Our research demonstrated that a high riskscore of PAAD-IRGS in patients with PAAD was correlated with poor overall survival, disease-specific survival and progression free interval. The same results were observed in patients with LIHC. The models constructed were confirmed to be accurate and reliable. We found various correlations between PAAD-IRGS and immune-related cells/genes, and the potential therapeutic agents. These findings indicate that PAAD-IRGS may be a promising indicator for prognosis and of the tumor-immune microenvironment status in PAAD.
Similar content being viewed by others
Introduction
Pancreatic adenocarcinoma (PAAD) is one of the most common carcinomas globally and ranks 6th in cancer-related deaths1. Although considerable progress has been made in diagnosis and treatment2, the 5-year survival rate of PAAD is still less than 10%3. Therefore, there is still a need for new ways to predict patient prognosis and augment early intervention to maximize long-term survival.
The development of high-throughput sequencing has revolutionized DNA and RNA research4 and broadened the scope of research into potential biological progress and mechanisms of human disease5. Several studies have revealed differentially expressed mRNA/miRNA/lncRNA and differentially expressed genes (DEGs) of pancreatic carcinoma in recent years6,7,8,9,10. Although its theoretical value to the diagnosis and prognosis of pancreatic carcinoma has been detailed, the biological mechanisms, clinical significance, and the interaction between DEGs during pancreatic carcinoma tumorigenesis are yet to be explored.
Inflammation mediates and participates in various pathophysiological processes, including classic pathways of infection, immune elimination, tissue repair and regeneration11,12. The current studies put forwards a new point of view that inflammation is tightly associated with tumorigenesis, progression and metastasis of cancer13,14. Tumor risk factors can stimulate an extrinsic inflammatory response, while innate inflammatory response contributes to tumor progression, indicating that a complex network exists in tumor-immune microenvironment. Furthermore, immune-related genes (IRGs), including interleukin (IL)-1015, IL-616, tumor necrosis factor-α (TNF-α)17 and (C-X-C motif) ligand (CXCL) chemokine family18 played a vital role in tumor proliferation, metabolism and metastasis. The occurrence and development of pancreatic cancer are recognized to be closely linked with inflammation. Local and systemic chronic inflammation could elevate the risk of PAAD, and PAAD-related inflammatory infiltration might simultaneously enhance tumor progression and metastasis19. Beyond the mechanism of an imbalance between inflammatory cell infiltration and immunosuppressive phenotype in the tumor-immunity microenvironment, obesity and diabetes are associated with promoting inflammation and inhibiting autophagy to Create a suitable environment for the tumorigenesis of PAAD through oxidative stress and metabolic impairments20.
Due to the interaction between immune-mediated inflammation and tumorigenesis, identifying whether immune response influences the prognosis of cancer patients has become a research hotspot. Quite a few carcinoma prognosis-related biomarkers have been identified and used to create models to predict patient survival21,22,23,24. However, there has not been much regarding IRGs signature for PAAD, let alone an immune-related prognostic model. In this study, we used the Cancer Genome Atlas (TCGA) and the Gene Expression Omnibus (GEO) database to screen out high-risk IRGs and create a novel risk-score signature and nomogram based on the IRGs for predicting the prognosis of PAAD patients. We also identified and comprehensively analyzed potential clinical therapeutic targets. Our findings may highlight the outstanding function of the IRGs signature in predicting PAAD patients’ prognosis and reveal its potential ability to predict the prognosis of patients with liver hepatocellular carcinoma (LIHC).
Materials and methods
Data acquisition and processing
We downloaded the TCGA-PAAD and TCGA-LIHC data sets, including: RNA sequences, raw clinical data and prognostic information, from the TCGA database (https://portal.gdc.cancer.gov/). Data of normal tissues from the GTEx database (https://gtexportal.org/) was obtained for supplementary. The gene expression data were converted to Transcripts per million reads (TPM) format and log2 transformed. Other data was cleaned and batch corrected with clinical information retained. Gene expression profiles and prognostic data of GSE2873525 and GSE6245226 were collected from the GEO database (http://www.ncbi.nlm.nih.gov/geo/) and used as validation datasets.
We obtained complete IRGs names, totaling 2483 from the “Resources-Gene Lists” module of the Immunology Database and Analysis Portal (ImmPort)27 (https://www.immport.org/home).
DEGs & IRGs screening and intersecting
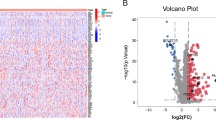
We first conducted a differential gene expression analysis to screen for genes expressed differently between pancreatic tumors and normal tissues, based on the RNA sequence dataset of TCGA-GTEx-PAAD. The log2(Fold Change) (FC) and adjusted p-value (P.adj) were calculated using R. Then, |log2(FC)|> 1 & P.adj < 0.05 was considered as the cut-off criteria for significant DEGs. These were subsequently intersected with the IRGs above. “ggplot2” package of R was used to visualize the performance with volcanoes plot and Venn diagram.
Enrichment analysis for DEGs & IRGs
We performed the Kyoto encyclopedia of genes and genomes (KEGG) pathway and Gene Ontology (GO) enrichment analysis and the results were plotted using “ggplot2” (version 3.3.3) and “clusterProfiler” (version 3.14.3) packages in R28 for the genes of intersection. The settings modes used were: biological process (BP), cellular component (CC) and molecular function (MF) with P.adj < 0.05 were considered statistically significant and output visualized cnetplots respectively.
Construction of PAAD-related IRGs signature (PAAD-IRGS) for prognosis
Based on the gene analysis above, we obtained independent immune-related prognostic risk genes using the Least absolute shrinkage and selection operator (Lasso) regression analysis29, followed by univariate and multivariate Cox regression analysis for further identification. LASSO is a popular algorithm, extensively utilized in medical studies30,31,32,33. Next, the Toil procedure from the university of California Santa Cruz (UCSC) Xena (34 was used to analyze the difference in the expression of the genes identified above in unpaired samples of PAAD. The log2(Transcripts per million (TPM) + 1) for log-scale was used in the assessments. The diagnostic value of these genes was evaluated using receiver operating characteristic (ROC) curves.
After this procedure, the optimal related IRGs were retained to establish the PAAD-IRGS. We compared the expression level of these genes in different pathologic stages and conducted the exclusively related KEGG and GO analysis. According to the expression level (EXP) and multivariate COX regression coefficient β value of the genes, the formula of the immune-related risk score signature is as follows35:
Based on the risk score of each sample, the cohort was divided into two groups (low-risk with 0–50% vs high-risk with 50–100%). The performance of the classifier was assessed using ROC. Finally, we performed survival analyses of overall survival (OS) for single and combined genes using Kaplan–Meier and the log-rank test.
Assessment of PAAD-IRGS and relevant clinical nomogram
The model to predict 1–3 years OS was evaluated using time-dependent ROC and decision curve analysis (DCA). Next, clinicopathologic characteristics of patients from TCGA-PAAD were collected and analyzed using univariate and multivariate COX regression analysis. Based on the clinical risk indicators (CRI) and PAAD-IRGS, we established a nomogram model to predict 1–3 years OS probability in PAAD patients. The nomogram was calibrated and assessed using DCA to verify its accuracy and reliability. The predictive accuracy of classical TMN-stage, PAAD-IRGS, CRI and nomogram were compared using the concordance index (C-Index).
Validation and extended application of PAAD-IRGS
To validate the specificity and precision of PAAD-IRGS, we utilized GSE28735 and GSE62452, which contained sufficient gene expression and prognosis data, to conduct differential expression analysis, survival analysis, diagnostic/prognostic value and applicability of clinical decision evaluation.
For assessing the extended applicability of PAAD-IRGS, considering the disease categories and histological homologies, we selected the TCGA-LIHC (n = 374) for further validation of the model. The difference in expression level of these genes between tumor and normal tissues was compared, and their individual and unified diagnostic ability. According to the standard established above, the LIHC cohort was grouped as low- (0–50%) and high-risk (50–100%) groups. Single-gene and unified signature OS analyses were performed using Kaplan–Meier curves, followed by time-dependent ROC and DCA analysis. Similarly, we established a nomogram model to predict 1–3 years OS probability in LIHC patients, based on the PAAD-IRGS and CRI, obtained from the TCGA-LIHC cohort through univariate and multivariate COX regression analysis. Calibration and DCA were performed to verify the reliability and accuracy of the model. Then the classical TMN-stage, PAAD-IRGS, CRI of TCGA-LIHC, and synthetic nomogram were compared with C-Index to assess their accuracy and clinical value for LIHC.
Furthermore, we expanded the application of PAAD-IRGS to predict 1–3 years disease-specific survival (DSS) and progression-free interval (PFI) of patients in TCGA-PAAD.
Immuno-correlation analysis and drug prediction of PAAD-IRGS
We conducted the PAAD-IRGS risk score correlation analysis with 24 immune-related cells36 in PAAD using the spearman’s test37. Subsequently, survival analysis of several significant immune-related cells was conducted to identify whether they were risk factors of PAAD using tumor immune estimation resource (TIMER), version 2.0 database38,39,40. Then we downloaded the immunophenoscore (IPS)41 data from The Cancer Immunome Atlas (TCIA) database (https://tcia.at/patients), which supports results of comprehensive immunogenomic analysis of next generation sequencing data (NGS) based on TCGA42, for analyzing the correlation between PAAD-IRGS and immune response in PAAD patients.
Relationships between PAAD-IRGS risk score and three kinds of immunomodulators expression in PAAD based on TCGA were explored and visualized with heatmaps, as well as relevant drug prediction accordingly via tumor-immune system interaction database (TISIDB)43 (http://cis.hku.hk/TISIDB/index.php), integrating multiple heterogeneous data. We searched the website with the gene symbol S100P, S100A2 and MMP12 and download relevant information in the "drug" module. Circle map and annotations were performed accordingly.
Analysis of protein expression of the PAAD-IRGS
The human protein atlas (HPA) database44,a spatial map of the human proteome (http://www.proteinatlas.org/humanproteome/pathology) was used to ascertain the physiological and pathological expression data of S100P, S100A2 and MMP12. As supplementary, we used UALCAN (http://ualcan.path.uab.edu/index.html) to conduct protein level analysis of S100P, S100A2 and MMP12 genes. It is a comprehensive and interactive public resource for cancer OMICS data analysis45, provided by the Clinical proteomic tumor analysis consortium (CPTAC) dataset46.
Statistical analysis
All statistical analyses were performed with R (version 3.6.3). Normally distributed variables were analyzed using the t-test and one-way ANOVA test and non-normally distributed variables with nonparametric tests. Log-rank test and Cox regression were used for survival analysis, Pearson’s correlation and spearman’s rank correlation test for correlation analysis. P or P.adj < 0.05 was considered statistically significant. The correlations was defined as follows: 0.00–0.10 (negligible), 0.10–0.39 (weak), 0.40–0.69 (moderate), 0.70–0.89 (strong), 0.90–1.00 (very strong)47.
Results
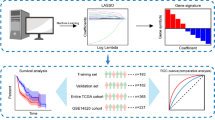
The study design for this work is shown in Fig. 1.
DEGs & IRGs analysis
178 PAAD patients with gene expression and prognostic information and 4 matched adjacent normal samples were included in the training cohort. 25,597 gene IDs were analyzed after removing null values, in which we obtained 539 differentially expressed genes that met the cut-off criterion of |log2(FC)|> 1 & P.adj < 0.05 in PAAD (236 genes up-regulated while 303 down-regulated) (Fig. 2A). Through the intersection of 490 DEGs and 1744 IRGs, 49 differentially expressed IRGs in PAAD were screened out (Fig. 2B).
Screening of differentially expressed genes and immune-related genes related to pancreatic adenocarcinoma. (A) Volcano plot of 539 DEGs; (B) Venn diagram of intersection of DEGs and IRGs. (C) KEGG pathways analysis of genes in DEGs & IRGs; (D) GO analysis of genes in DEGs & IRGs; (E) Volcano plot of 49 genes in DEGs & IRGs. DEGs differentially expressed genes, IRGs immune-related genes, KEGG Kyoto Encyclopedia of Genes and Genomes, GO Gene Ontology.
Enrichment analysis
The KEGG pathways which were most associated with immunity involved in natural killer mediated cytotoxicity (P < 0.001), B cell receptor signaling pathway (P < 0.001) and chemokine signaling pathway (P < 0.05) (Fig. 2C). Specifically, regulation of the immune effector process, cell killing and humoral immune response of the biological process (BP) module (all P < 0.001) were observed to be associated with immunity. So was major histocompatibility complex (MHC) protein binding and cytokine receptor binding of molecular functional (MF) module (Fig. 2D). Gene overlap is highlighted in the volcano plot (Fig. 2E).
Construction and assessment of PAAD-IRGS
We further analyzed the genes identified above to identify the potential diagnostic and prognostic value of IRGs in PAAD. Based on LASSO regression analysis, four prognostic risk biomarkers were identified (high expression of S100P, S100A2, and MMP12 was associated with poor prognosis, while low expression of DEFA5 was associated with better prognosis) (Fig. 3A). S100P, S100A2 and MMP12 were expressed higher in tumor tissues, compared with normal tissues (P < 0.001), while the opposite was true for DEFA5 expression (P < 0.05) (Fig. 3B). The area under curve (AUC) of S100P, S100A2, and MMP12 were 0.971, 0.968, and 0.981, indicating their excellent diagnostic value. However, DEFA5 was considered an inefficient biomarker for diagnosis (AUC = 0.438) (Fig. 3C). Subsequent univariate and multivariate COX regression analyses were conducted on the four genes, excluding DEFA5 (P = 0.164) (Fig. 3D). The model of PAAD-IRGS was finally comprised of S100P, S100A2 and MMP12. We plugged the corresponding regression coefficients into the equation as follows to complete the establishment of PAAD-IRGS: PAAD-IRGS = EXP(S100P) × 0.132 + EXP(S100A2) × 0.098 + EXP(MMP12) × 0.095.
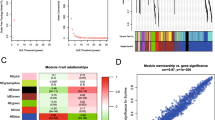
Establishment of IRGs signature (IRGS). (A) Ten-time cross-validation for tuning parameter selection in the Lasso regression model and risk analysis of four immune-related genes in patients with PAAD; (B) differential expression of four IRGs between tumor and normal tissues of patients with PAAD; (C) diagnostic value of four IRGs for patients with PAAD; (D) univariate and multivariate COX analysis of four IRGs is shown in forest map. *P < 0.05, **P < 0.01, ***P < 0.001.
Furthermore, we performed PAAD-IRGS specialized differential expression analysis in different pathology stages. Using the Gene expression profiling interactive analysis (GEPIA) database48, a statistically significant difference in S100P expression was observed in different pathology stages of TCGA-PAAD (P < 0.001) (Fig. 4A). After a single gene correlation analysis of these three genes, we obtained 79 co-correlated genes (Fig. 4B). Based on further enrichment analysis, KEGG pathways seemingly involved in ECM-receptor interaction, regulation of actin cytoskeleton, p53 signaling pathway, focal adhesion and pancreatic cancer, and GO pathway focused on cell-membrane organization and connection (Fig. 4C).
A comprehensive evaluation of IRGS. (A) Expression of 3 signature genes in different pathologic stages of PAAD; (B) Venn diagram of intersection of enrichment analysis of 3 signature genes; (C) GO and KEGG analysis of 3 signature genes; (D) diagnostic value of IRGS in PAAD; (E) single-gene survival analysis of OS was shown in Kaplan–Meier curves respectively; (F) Kaplan–Meier curves show that OS was significantly different between the low- and high-risk groups in TCGA-PAAD. OS overall survival; *P < 0.05, **P < 0.01, ***P < 0.001.
The model showed a better diagnostic capability than individual genes with an AUC of 0.993 (95% confidence interval (CI) = 0.987–0.998) (Fig. 4D). By single gene survival analysis, we observed that patients in S100A2 high-expression group had a worse OS than patients in S100A2 low-expression group (hazard ratio (HR) = 1.62, 95% CI = 1.07–2.46, P = 0.023). However, there is no statistically significant difference between low and high expression groups of S100P or MMP12 (Fig. 4E). Patients in the PAAD-IRGS high-risk score group had a much worse OS than patients in low-risk score group (HR = 2.21, 95% CI = 1.45–3.39, P < 0.001) (Fig. 4F).
Establishment of PAAD-IRGS based prognosis model
A total of 182 TCGA-PAAD patients were included in the prognostic analysis with the baseline characteristics shown in Table 1. Time-dependent ROC analysis was conducted to assess the accuracy of PAAD-IRGS for prediction of OS in PAAD patients. It showed an above average performance of 1 (AUC = 0.679), 2 (AUC = 0.696), and 3 years (AUC = 0.713) (Fig. 5A). DCA showed that model has a good clinical utility (Fig. 5B). T3&T4 stage (P = 0.030), N1 stage (P = 0.004), pathological stage II (P = 0.033), radiation therapy (P = 0.013), primary therapy outcome of PR&CR (P < 0.001), R1&R2 resection (P = 0.028), histological grade G2 (P = 0.047)/G3&G4 (P = 0.008), non-head of pancreas neoplasm (P = 0.004) and PAAD-IRGS (P < 0.001) were significantly correlated with OS. Radiation therapy (HR = 0.437, 95%CI = 0.228–0.835, P = 0.012), primary therapy outcome of PR&CR (HR = 0.547, 95%CI = 0.324–0.923, P = 0.024), R1&R2 resection (HR = 1.896, 95%CI = 1.087–3.308, P = 0.024) and PAAD-IRGS (HR = 2.312, 95%CI = 1.245–4.294,P = 0.008) were independent factors impacting the OS of patients with PAAD (Table 2). Based on the above analysis, the nomogram incorporating PAAD-IRGS and multiple clinicopathological characteristics was plotted (Fig. 5C). Through comparison, the concordance index (C-Index) of TNM-stage, PAAD-IRGS, Nomogram (only clinical indicators), and Nomogram + IRGS was 0.567, 0.639, 0.706 and 0.723 (Table 3), respectively. Additionally, Nomogram calibration curves (Fig. 5D) showed good predictive accuracy of the model and DCA (Fig. 5E).
Evaluation of PAAD-IRGS and establishment and assessment of relevant nomograms. (A) The time-dependent ROC curve of the PAAD-IRGS for predicting 1, 2, and 3-year OS; (B) decision curve analysis for evaluating the PAAD-IRGS; (C) an PAAD-IRGS -based nomogram included with 8 clinical components predicting 1, 2, and 3-year OS of PAAD; (D) nomogram calibration curve for 1, 2, and 3-year. (E) decision curve analysis for evaluating the net benefits of nomogram at 1, 2, and 3 years. *P < 0.05, **P < 0.01, ***P < 0.001.
Validation and extension of PAAD-IRGS
For further validation of the reliability of PAAD-IRGS, we employed two datasets of the GEO database. Differential expression, survival, diagnostic value, prognostic value analysis and DCA were performed in both datasets. The three genes had a higher expression in the tumor tissues than in normal tissues (P < 0.001) of GSE28735 (Fig. 6A). Patients in a high-risk group of PAAD-IRGS had worse OS than that of the low-risk group (HR = 2.35, 95%CI = 1.08–5.14, P = 0.032) (Fig. 6B). Consistent with the results above, although S100P (AUC = 0.929), S100A2 (AUC = 0.764), MMP12 (AUC = 0.828) (Fig. 6C) showed considerable diagnostic values for PAAD respectively, PAAD-IRGS had the optimum diagnostic ability (AUC = 0.943, 95%CI = 0.896–0.991) (Fig. 6D). In addition, time-dependent ROC showed the model had an above-average ability to predict 1—(AUC = 0.671), 2—(AUC = 0.600), and 3—year OSs (AUC = 0.866) (Fig. 6E). The model also had an acceptable net benefit based on DCA (C-Index = 0.644, 95%CI = 0.598–0.690) (Fig. 6F). Similar results of differential expression (P < 0.001) (Fig. 7A) and OS probability (HR = 1.84, 95%CI = 1.02–3.32, P = 0.044) (Fig. 7B) were obtained in GSE62452, as well as the independent diagnostic value of S100P (AUC = 0.865), S100A2 (AUC = 0.745), MMP12 (AUC = 0.811) (Fig. 7C) and all of them combined (AUC = 0.885, 95%CI = 0.828–0.943) (Fig. 7D). The corresponding ROC analysis showed an above-average performance in predicting 1—(AUC = 0.536), 2—(AUC = 0.672), and 3—year prognosis (AUC = 0.861) (Fig. 7E). Although 1-year net benefit of prognostic prediction was not satisfactory, 2- and 3-years showed a much better net benefit (C-Index = 0.580, 95%CI = 0.531–0.629) (Fig. 7F).
Validation of PAAD-IRGS with GSE28735. (A) Expression level of 3 IRGs in GSE28735 cohort; (B) Kaplan–Meier curves show a better OS in the low-risk group than the high-risk group; (C) diagnostic value of 3 IRGs for PAAD patients in GSE28735 cohort; (D) diagnostic value of PAAD-IRGS in GSE28735 cohort; (E) time-dependent ROC curve analysis of the PAAD-IRGS at 1, 2, and 3 years in GSE28735 cohort; (F) decision curve analysis for evaluating the net benefits of PAAD-IRGS in GSE28735 cohort. *P < 0.05, **P < 0.01, ***P < 0.001.
Validation of PAAD-IRGS with GSE62452. (A) Expression level of 3 IRGs in GSE62452 cohort; (B) Kaplan–Meier curves show a better OS in the low-risk group than the high-risk group; (C) diagnostic value of 3 IRGs for PAAD patients in GSE62452 cohort; (D) diagnostic value of PAAD-IRGS in GSE62452 cohort; (E) time-dependent ROC curve analysis of the PAAD-IRGS at 1, 2, and 3 years in GSE62452 cohort; (F) decision curve analysis for evaluating the net benefits of PAAD-IRGS in GSE62452 cohort. *P < 0.05, **P < 0.01, ***P < 0.001.
Hepatobiliary and pancreatic carcinoma were categorized as a unity of clinical disease due to their close anatomical correlation and mutual functional assistance. To verify the universal applicability of the PAAD-IRGS, the TCGA-LIHC data was used to validate the findings. S100A2, S100P and MMP12 were all over expressed in tumor tissues based on paired (P < 0.01) (Fig. 8A) and unpaired expression analysis (P < 0.001) (Fig. 8B). The diagnostic ROC curves also showed their independent and unified diagnostic value for LIHC (S100P: AUC = 0.739; S100A2: AUC = 0.723; MMP12: AUC = 0.773; model: AUC = 0.812, 95%CI = 0.767–0.857) (Fig. 8C,D). LIHC patients had a worse OS in S100P (HR = 1.43, 95% CI = 1.01–2.02, P = 0.44)/S100A2 (HR = 1.81, 95% CI = 1.27–2.57, P = 0.001)/MMP12 (HR = 1.58, 95% CI = 1.11–2.23, P = 0.01) high-expression group (Fig. 8E) and PAAD-IRGS high-risk group (HR = 1.83, 95% CI = 1.29–2.60, P = 0.001) (Fig. 8F). PAAD-IRGS also had a considerable prognostic value for LIHC patients according to ROC analysis (1-year: AUC = 0.651; 2-year: AUC = 0.612; 3-year: AUC = 0.597) (Fig. 8G) and DCA (Fig. 8H). Furthermore, we extracted baseline characteristics of TCGA-LIHC shown in Table 4 and conducted univariate and multivariate COX regression analysis to establish a nomogram based on PAAD-IRGS and multiple clinicopathologic factors (Fig. 8I). T3&T4 stage (P < 0.001), M1 stage (P = 0.017), pathological stage III&IV (P < 0.001), tumor-bearing status (P < 0.001) and PAAD-IRGS (P < 0.001) were significantly correlated with OS. Tumor-bearing status (HR = 1.992, 95%CI = 1.246–3.185, P = 0.004) and PAAD-IRGS (HR = 2.180, 95%CI = 1.180–4.026, P = 0.013) were independent factors impacting the OS of patients with LIHC (Table 5). Nomogram calibration curves (Fig. 8J) showed good predictive accuracy of the model, and DCA (Fig. 8K) confirmed the clinical utility of the nomogram. Consistent with the nomogram of PAAD, the comprehensive nomogram of LIHC showed the best accuracy (C-Index = 0.666, 95%CI = 0.630–0.701) than any other indicator (Table 3).
Validation of PAAD-IRGS with TCGA-LIHC. (A) Paired comparison of 3 IRGs expression levels in TCGA-LIHC; (B) unpaired comparison of 3 IRGs expression levels in TCGA-LIHC by including the relevant normal tissues of the GTEx database as controls; (C) diagnostic value of 3 IRGs for TCGA-LIHC patients; (D) diagnostic value of PAAD-IRGS for TCGA-LIHC patients; (E) single-gene survival analysis of OS for TCGA-LIHC; (F) PAAD-IRGS survival analysis of OS for TCGA-LIHC; (G) time-dependent ROC curve analysis of the PAAD-IRGS at 1, 2, and 3 years for TCGA-LIHC; (H) decision curve analysis for evaluating the net benefits of PAAD-IRGS for TCGA-LIHC; (I) an PAAD-IRGS-based nomogram included with 4 clinical components predicting 1, 2, and 3-year OS of TCGA-LIHC; (J) nomogram calibration curve for 1, 2, and 3-year. (K) Decision curve analysis for evaluating the net benefits of nomogram at 1, 2, and 3 years. *P < 0.05, **P < 0.01, ***P < 0.001.
For the further expanded application of PAAD-IRGS, we found that it performed well in predicting disease-specific survival (DSS) and progression-free interval (PFI) of PAAD patients. Patients in PAAD-IRGS high-risk group had a significantly worse DSS (HR = 2.54, 95%CI = 1.55–4.15, P < 0.001) (Fig. 9A). Time-dependent ROC showed its robust prognostic predictive value (1-year: AUC = 0.730; 2-year: AUC = 0.724; 3-year: AUC = 0.749) and DCA further validated its clinical applicability (C-Index = 0.680, 95%CI = 0.649–0.711) (Fig. 9B,C). We constructed a comprehensive nomogram composed of PAAD-IRGS and clinicopathological factors (Table 6) (Fig. 9D). Its accuracy and efficiency were evaluated (C-Index = 0.775, 95%CI = 0.742–0.808, Table 3) (Fig. 9E,F). Similarly, Patients in PAAD-IRGS high-risk group had a significantly worse PFI (HR = 2.28, 95%CI = 1.53–3.40, P < 0.001) (Fig. 10A). The model had good clinical utility (C-Index = 0.649, 95%CI = 0.618–0.681) (Fig. 10B) and predictive value for prognosis (1-year: AUC = 0.666; 2-year: AUC = 0.723; 3-year: AUC = 0.730) (Fig. 10C). The nomogram based on this model is shown in Fig. 10D using variables summarized in Table 7. The validation analysis results are in Table 3 and Fig. 10E,F (C-Index = 0.742, 95%CI = 0.712–0.771).
Establishment and assessment of PAAD-IRGS-based nomograms for DSS in TCGA-PAAD. (A) Single-gene and IRGs signature Survival analysis of DSS in PAAD; (B) the time-dependent ROC curve of the PAAD-IRGS for predicting 1, 2, and 3-year DSS; (C) decision curve analysis for evaluating the PAAD-IRGS; (D) an PAAD-IRGS-based nomogram included with 8 clinical components predicting 1, 2, and 3-year DSS of PAAD; (E) nomogram calibration curve for 1, 2, and 3-year. (E) Decision curve analysis for evaluating the net benefits of nomogram at 1, 2, and 3 years. DSS, disease-specific survival; *P < 0.05, **P < 0.01, ***P < 0.001.
Establishment and assessment of PAAD-IRGS-based nomograms for PFI in TCGA-PAAD. (A) Single-gene and PAAD-IRGS survival analysis of PFI in PAAD; (B) the time-dependent ROC curve of the PAAD-IRGS for predicting 1, 2, and 3-year PFI; (C) decision curve analysis for evaluating the PAAD-IRGS; (D) an PAAD-IRGS-based nomogram included with 7 clinical components predicting 1, 2, and 3-year PFI of PAAD; (E) nomogram calibration curve for 1, 2, and 3-year. (E) Decision curve analysis for evaluating the net benefits of nomogram at 1, 2, and 3 years. PFI progress free interval; *P < 0.05, **P < 0.01, ***P < 0.001.
Immunity associated analysis of PAAD-IRGS
Tumor-infiltrating immunocytes (TIICs) play an important role in the complex tumor-immune microenvironment and have been shown to influence the progression of various tumors49,50. Thus, we must investigate any relationship between PAAD-IRGS and TIICs in PAAD. We used a lollipop plot to perform the correlation analysis of 24 immune-related cells (Fig. 11A). There was a significant positive correlation between PAAD-IRGS and NK CD56bright cells (r = 0.333, P < 0.001) and Th2 cells (r = 0.367, P < 0.001) and negative correlation with plasmacytoid dendritic cells (pDC) (r = −0.348, P < 0.001) and follicular helper T cell (TFH) (r = -0.344, P < 0.001). However, only B cell, CD4+ T cell and NK cell infiltration levels were correlated with OS of PAAD patients. Patients with high B cell (HR = 0.776, P = 0.0147) or NK cell (HR = 0.788, P = 0.0226) infiltration level had a better OS, while high CD4+ T cell + Th2 cell infiltration level associated with worse OS (HR = 1.36, P = 0.00337) (Fig. 11B). There was no statistically significant difference between high-risk and low-risk groups in patients with PD-1 blocker/CTLA4 blocker/CTLA4&PD-1 blocker or without immune-blocker (Fig. 11C).
Analysis of correlation between PAAD-IRGS and immune-related cells and relevant immunotherapy. (A) Lollipop plot of PAAD-IRGS and immune infiltration cells correlation in TCGA-PAAD; (B) survival analysis of immune-related cells infiltration in PAAD; (C) analysis of immunotherapeutic efficiency based on PAAD-IRGS in TCGA-PAAD. IPS immunephenoscore. *P < 0.05, **P < 0.01, ***P < 0.001.
As a supplement, we conducted correlation analysis between immunomodulators and PAAD-IRGS, which were visualized as heatmaps (Figs. 12A, 13A, 14A). For immune-inhibitors, PAAD-IRGS had positive correlation with TGFB1 (r = 0.372, P < 0.001), LGALS9 (r = 0.674, P < 0.001), IL10RB (r = 0.555, P < 0.001) and CD274 (r = 0.227, P = 0.002), negative correlation with KDR (r = −0.330, P < 0.001), CD160 (r = −0.358, P < 0.001), BTLA (r = −0.224, P = 0.003) and ADORA2A (r = −0.243, P = 0.001) (Fig. 12B). For (MHC) molecule, HLA-B (r = 0.271, P < 0.001), HLA-C (r = 0.229, P = 0.002), B2M (r = 0.482, P < 0.001), HLA-A (r = 0.357, P < 0.001), TAP2 (r = 0.302, P < 0.001), TAPBP (r = 0.330, P < 0.001), HLA-F (r = 0.261, P < 0.001) and TAP1 (r = 0.324, P < 0.001) were positively related with PAAD-IRGS (Fig. 13B). As to immune-stimulators, there were 6 genes negatively related with PAAD-IRGS (Fig. 14B) while 15 genes had a positive correlation (Fig. 14C).
Analysis of correlation between PAAD-IRGS and immuno-stimulators. (A) Immune-stimulator genes—PAAD-IRGS e heatmap; (B) correlation analysis of immune-stimulator genes and PAAD-IRGS (negative). (C) Correlation analysis of immune-stimulator genes and PAAD-IRGS (positive). *P < 0.05, **P < 0.01, ***P < 0.001.
PAAD-IRGS related drugs
TISIDB is a web portal for tumor and immune system interaction, which supports genomics, transcriptomics, and clinical data from TCGA and mechanism, and drug information from public databases. We can only obtain potential drugs associated with PAAD-IRGS, which is demonstrated in a network diagram (Fig. 15). Currently, drugs targeting PAAD-IRGS (S100P, S100A2 and MMP12) remained in the experimental stage, and effective targeted drugs for pancreatic cancer are still in the blank.
Analysis of protein expression of the PAAD-IRGS
We obtained the protein expression pattern of S100P and S100A2 in different cancers based on the HPA database. Expression of S100P in most pancreatic (83.3%) and liver (54.5%) cancers showed moderate to intense cytoplasmic and nuclear staining (Fig.S1A). Immunohistochemistry (IHC) results also confirmed that S100P was highly expressed in PAAD and LIHC than in corresponding normal tissues (Fig.S1B). Although the level of S100A2 protein expression was lower than that of S100P (Fig.S2A), we can still observe the moderate intensity of S100A2 in PAAD and LIHC than in corresponding normal tissues (Fig.S2B). The information on MMP12 in the HPA database was absent, we conducted further verification using the UALCAN database. To be consistent, the protein expression of S100P was higher in PAAD and LIHC than in corresponding normal tissues (P < 0.001) (Fig.S3A), as well as in MMP12 (P < 0.001) (Fig.S3B). Despite the data absent in LIHC, the protein expression of S100A2 was higher in PAAD than in normal tissues (P < 0.01) (Fig.S3C).
Discussions
Although pancreatic cancer is still one of the leading causes of cancer-related death worldwide, some improvements in patient outcomes have been made due to advancements in therapeutics51. Since there are no obvious clinical symptoms in the early stage, pancreatic cancer is usually advanced at diagnosis. Secondly, the high mortality of PAAD seems to be inextricably associated with its suppressed immune microenvironment and significant decrease of T cell infiltration levels in the tumor52. Although immunotherapy has revolutionized the cancer treatment model, PAAD patients rarely respond to these therapies due to poor activation and infiltration of T cells in the tumor-immunity microenvironment (TIME). Recent research has revealed potential epigenetic-transcriptional mechanisms by which tumor cells remodel their TIME and suggested EGFR inhibitors as potential immunotherapy sensitizers in PAAD53. Intra-tumoral IFN-γ-producing Th22 cells were reported to be associated with TNM staging and the worst outcomes in PAAD54. γδ T Cells were also considered to promote pancreatic oncogenesis by restraining αβ T Cells activation55. Each T cell subpopulations secretes different cytokines and chemokines that modulate the immune response in synergistic and opposite ways56. Additionally, expansion of immunosuppressive B cells induced by IL-1β might promote PAAD57, and many extracellular matrix (ECM) components, including collagen, growth factors, cytokines, chemokines, and cancer-associated fibroblast (CAF) play vital role in tumor progression58. All tumor-immunity components in the TIME interact continuously, constructing a complex stroma-tumor crosstalk network. Due to the complexity of tumor-immunity mechanisms, there is still no effective way to predict prognosis in clinical practice. Our study aimed to discover immune-related biomarkers and establish a robust model to predict prognosis in PAAD patients.
The TCGA-PAAD dataset was used to screen for potentially immune-related DEGs , then analyzed for differential expression and intersection. GO and KEGG enrichment analyses were also performed to confirm that the mechanisms involved in these genes were focused on immune-related pathways (Fig. 2D). Furthermore, we narrowed down the results by Lasso regression analysis and obtained three key IRGs finally through the univariate and multivariate Cox regression analysis. The PAAD-IRGS comprised of S100P, S100A2 and MMP12 had an outstanding diagnostic value (Fig. 4D) and accurately predicted the prognosis for PAAD patients (Fig. 5A,B). We specially performed secondary enrichment analysis on PAAD-IRGS, revealing that this model was also associated with pathways of ECM and cell-membrane junction and immune-related pathways (Fig. 4C).
Among the three genes, S100P, a 95-amino-acid protein belonging to the S100 family, was regarded as a promising diagnostic59 and prognostic biomarker60 for pancreatic cancer with a potential mechanism of regulating invasion into the lymphatic endothelial monolayer61, which is consistent with our results. S100A2, another member of the S100 family, was reported as a prognostic biomarker involved in immune infiltration and immunotherapy response prediction in pancreatic cancer62, which matches our findings. Turn to MMP12, as one of the members of the matrix metalloproteinases family, it encodes extracellular matrix participating in the (EMT) which was identified as a strictly programmed shift playing a crucial role in tumor invasion and metastasis63. MMP12 was also revealed to be a potential diagnostic biomarker for pancreatic carcinoma. Its up-regulation was associated with a poor prognosis64. These genes were verified to be closely correlated with different cancers, especially the diagnosis and prognosis of PAAD, a finding we also made in this work. Although many types of diagnostic or prognostic biomarkers, and even to some extent predictive models have been identified in recent studies60,65,66,67,68, we discovered three IRGs with high specificity. We integrated them to establish a novel prognostic model for PAAD. Compared with other models, our model had an extremely remarkable performance on both diagnosis and prognosis prediction in PAAD patients.
In our study, patients in the PAAD-IRGS high-risk group had a significantly worse OS than those in the low-risk group (Fig. 4F), indicating that the PAAD-IRGS score may be an independent risk factor when evaluating the prognosis of PAAD patients. Additionally, time-dependent ROC and DCA results (Fig. 5A,B) showed that PAAD-IRGS had a good performance in prediction prognosis. The nomogram integrating PAAD-IRGS and multiple clinicopathological variables showed better accuracy and reliability than any singular variable (Table 3).
We not only evaluated and validated the PAAD-IRGS by using two datasets of pancreatic cancer from GEO, but also investigated its application to hepatocellular carcinoma. Hepatobiliary and pancreatic diseases are often classified into the same category since they are anatomically and functionally linked. Although the cholangiocarcinoma dataset of TCGA was discarded due to its small sample size, we found that the PAAD-IRGS had excellent diagnostic and prognostic value on LIHC patients. We also combined relevant clinicopathological variables with PAAD-IRGS to construct a comprehensive nomogram model, which showed good accuracy and robustness. Based on the results, we might speculate whether the three genes participated in the oncogenesis, progression and metastasis of LIHC and PAAD partially or collectively. However this needs further exploration. We also looked into using PAAD-IRGS to predict DSS and PFI in patients with PAAD. The results of PAAD-IRGS and the relevant prognostic model were encouraging. Unlike other biomarkers that only had diagnostic value, PAAD-IRGS had the dual capability to predict diagnosis and prognosis with high accuracy. Several multiple-genes prognostic model have been established and reported67,68. Compared with them, our model had outstanding general applicability with high accuracy and stability. As to the miRNA or lncRNA-related signatures65,69,70,71, our PAAD-IRGS was more stable and convinced; Compared with multiple-gene signatures9,68, necroptosis-related gene signature72 and m6A-related gene signature73,74, which had been reported, our PAAD-IRGS was new and more versatile with outstanding performance. It can be well applied to prognostic prediction of multiple cancers with different prognostic parameters. Its good diagnostic ability for various cancer and its relationship with tumor-immunity would make it promising for further research.
There were several limitations of this research to be concerned about. The limitation to this study worth noting include: there may be an effect on the result due to batch effect and differences in sample sizes that are difficult to eliminate completely. Secondly, although the prognostic value of the PAAD-IRGS was evaluated in multiple datasets, large-scale clinical research is still necessary for further validation. Thirdly, we conducted correlation analyses between PAAD-IRGS and immune-related cells/immunomodulators and disclosed some potential immune-related targets. However, the underlying mechanisms and pathways need further investigation and experiment validation.
In conclusion, our study established a novel prognostic model comprised of three genes with high specificity for predicting prognosis in patients with PAAD. This model demonstrated excellent performance in predicting both diagnosis and prognosis. Since PAAD-IRGS can be generalized, it may be a beneficial predictive model in clinical practice.
Data availability
The original contributions presented in the study are included in the article/supplementary material. Further inquiries can be directed to the corresponding author/s. All data and original files in our work are freely available under a ‘Creative Commons BY 4.0’ license. All methods were carried out in accordance with relevant guidelines and regulations.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 71(3), 209–249 (2021).
McGuigan, A. et al. Pancreatic cancer: A review of clinical diagnosis, epidemiology, treatment and outcomes. World J. Gastroenterol. 24(43), 4846–4861 (2018).
Siegel, R. L. et al. Cancer statistics, 2021. CA Cancer J. Clin. 71(1), 7–33. https://doi.org/10.3322/caac.21654 (2021) ((epub 2021 Jan 12. Erratum in: CA Cancer J Clin. 2021 Jul;71(4):359)).
Lo Giudice, C., Pesole, G. & Picardi, E. High-throughput sequencing to detect DNA–RNA changes. Methods Mol. Biol. 2181, 193–212 (2021).
Rego, S. M. & Snyder, M. P. High throughput sequencing and assessing disease risk. Cold Spring Harb. Perspect. Med. 9(1), a026849 (2019).
Almeida, P. P., Cardoso, C. P. & de Freitas, L. M. PDAC-ANN: An artificial neural network to predict pancreatic ductal adenocarcinoma based on gene expression. BMC Cancer 20, 82 (2020).
Yang, C. et al. Evaluation of the diagnostic ability of laminin gene family for pancreatic ductal adenocarcinoma. Aging (Albany NY). 11, 3679–3703 (2019).
Long, N. P. et al. An integrative data mining and omics-based translational model for the identification and validation of oncogenic biomarkers of pancreatic cancer. Cancers (Basel). 11, 155 (2019).
Wu, M. et al. Identification of a nine-gene signature and establishment of a prognostic nomogram predicting overall survival of pancreatic cancer. Front. Oncol. 27(9), 996 (2019).
Wang, W. et al. A novel mRNA-miRNA-lncRNA competing endogenous RNA triple sub-network associated with prognosis of pancreatic cancer. Aging (Albany NY). 11(9), 2610–2627 (2019).
Medzhitov, R. Origin and physiological roles of inflammation. Nature 454(7203), 428–435 (2008).
Karin, M. & Clevers, H. Reparative inflammation takes charge of tissue regeneration. Nature 529(7586), 307–315 (2016).
Singh, N. et al. Inflammation and cancer. Ann. Afr. Med. 18(3), 121–126 (2019).
Taniguchi, K. & Karin, M. NF-κB, inflammation, immunity and cancer: Coming of age. Nat. Rev. Immunol. 18(5), 309–324 (2018).
Ahmad, N. et al. IL-6 and IL-10 are associated with good prognosis in early stage invasive breast cancer patients. Cancer Immunol. Immunother. 67, 537–549 (2018).
Weber, R. et al. IL-6 as a major regulator of MDSC activity and possible target for cancer immunotherapy. Cell Immunol. 359, 104254 (2021).
Cruceriu, D. et al. The dual role of tumor necrosis factor-alpha (TNF-α) in breast cancer: Molecular insights and therapeutic approaches. Cell Oncol. (Dordr). 43(1), 1–18 (2020).
Chen, X. et al. The role of CXCL chemokine family in the development and progression of gastric cancer. Int. J. Clin. Exp. Pathol. 13(3), 484–492 (2020).
Padoan, A., Plebani, M. & Basso, D. Inflammation and pancreatic cancer: Focus on metabolism, cytokines, and immunity. Int. J. Mol. Sci. 20(3), 676 (2019).
Gukovsky, I. et al. Inflammation, autophagy, and obesity: Common features in the pathogenesis of pancreatitis and pancreatic cancer. Gastroenterology 144(6), 1199–209.e4 (2013).
Huang, H. et al. Prognostic value of preoperative systemic immune-inflammation index in patients with cervical cancer. Sci. Rep. 9(1), 3284 (2019).
Shen, S. et al. Development and validation of an immune gene-set based prognostic signature in ovarian cancer. EBioMedicine 40, 318–326 (2019).
Qu, Y. et al. Identification of immune-related genes with prognostic significance in the microenvironment of cutaneous melanoma. Virchows Arch. 478(5), 943–959 (2021).
Zhu, C. et al. Esophageal cancer associated immune genes as biomarkers for predicting outcome in upper gastrointestinal tumors. Front. Genet. 19(12), 707299 (2021).
Zhang, G. et al. Integration of metabolomics and transcriptomics revealed a fatty acid network exerting growth inhibitory effects in human pancreatic cancer. Clin. Cancer Res. 19(18), 4983–4993 (2013).
Yang, S. et al. A novel MIF signaling pathway drives the malignant character of pancreatic cancer by targeting NR3C2. Cancer Res. 76(13), 3838–3850 (2016).
Bhattacharya, S. et al. ImmPort, toward repurposing of open access immunological assay data for translational and clinical research. Sci. Data. 27(5), 180015 (2018).
Yu, G. et al. clusterProfiler: An R package for comparing biological themes among gene clusters. Omics J. Integr. Biol. 16(5), 284–287 (2012).
Alhamzawi, R. & Ali, H. T. M. The Bayesian adaptive lasso regression. Math. Biosci. 303, 75–82 (2018).
Liu, Z. et al. Machine learning-based integration develops an immune-derived lncRNA signature for improving outcomes in colorectal cancer. Nat. Commun. 13(1), 816 (2022).
Liu, Z. et al. Stemness refines the classification of colorectal cancer with stratified prognosis, multi-omics landscape, potential mechanisms, and treatment options. Front. Immunol. 27(13), 828330 (2022).
Liu, Z. et al. Integrative analysis from multi-center studies identities a consensus machine learning-derived lncRNA signature for stage II/III colorectal cancer. EBioMedicine 75, 103750 (2022).
Liu, Z. et al. Development and clinical validation of a novel six-gene signature for accurately predicting the recurrence risk of patients with stage II/III colorectal cancer. Cancer Cell Int. 21(1), 359 (2021).
Vivian, J. et al. Toil enables reproducible, open source, big biomedical data analyses. Nat. Biotechnol. 35(4), 314–316 (2017).
Zeng, J. H. et al. Comprehensive investigation of a novel differentially expressed lncRNA expression profile signature to assess the survival of patients with colorectal adenocarcinoma. Oncotarget 8(10), 16811–16828 (2017).
Bindea, G. et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39(4), 782–795 (2013).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14(1), 1–15 (2013).
Li, T. et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 48(W1), W509–W514 (2020).
Li, T. et al. TIMER: A web server for comprehensive analysis of tumor-infiltrating immune cells. Can. Res. 77(21), e108–e110 (2017).
Li, Bo. et al. Comprehensive analyses of tumor immunity: Implications for cancer immunotherapy. Genome Biol. 17(1), 174 (2016).
Charoentong, P. et al. Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep. 18(1), 248–262 (2017).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165(1), 35–44 (2016).
Ru, B. et al. TISIDB: An integrated repository portal for tumor-immune system interactions. Bioinformatics 19, 210 (2019).
Thul, P. J. & Lindskog, C. The human protein atlas: A spatial map of the human proteome. Protein Sci. 27(1), 233–244 (2018).
Chandrashekar, D. S. et al. UALCAN: A portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia 19(8), 649–658 (2017).
Chen, F. et al. Pan-cancer molecular subtypes revealed by mass-spectrometry-based proteomic characterization of more than 500 human cancers. Nat. Commun. 10(1), 5679 (2019).
Schober, P., Boer, C. & Schwarte, L. A. Correlation coefficients: Appropriate use and interpretation. Anesth. Analg. 126(5), 1763–1768 (2018).
Tang, Z. et al. GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45(W1), W98–W102 (2017).
Zhang, S. et al. Immune infiltration in renal cell carcinoma. Cancer Sci. 110(5), 1564–1572 (2019).
Ye, L. et al. Tumor-infiltrating immune cells act as a marker for prognosis in colorectal cancer. Front. Immunol. 17(10), 2368 (2019).
Tempero, M. A. NCCN guidelines updates: Pancreatic cancer. J. Natl. Compr. Cancer Netw. 17(55), 603–605 (2019).
Morrison, A. H., Byrne, K. T. & Vonderheide, R. H. Immunotherapy and prevention of pancreatic cancer. Trends Cancer 4(6), 418–428 (2018).
Li, J. et al. Epigenetic and transcriptional control of the epidermal growth factor receptor regulates the tumor immune microenvironment in pancreatic cancer. Cancer Discov. 11(3), 736–753 (2021).
Niccolai, E. et al. Intra-tumoral IFN-γ-producing Th22 cells correlate with TNM staging and the worst outcomes in pancreatic cancer. Clin. Sci. (Lond). 130(4), 247–258 (2016).
Daley, D. et al. γδ T cells support pancreatic oncogenesis by restraining αβ T cell activation. Cell 166(6), 1485-1499.e15. https://doi.org/10.1016/j.cell.2016.07.046 (2016) ((epub 2016 Aug 25. Erratum in: Cell. 2020 Nov 12;183(4):1134-1136)).
Ajina, R. & Weiner, L. M. T-cell immunity in pancreatic cancer. Pancreas 49(8), 1014–1023 (2020).
Takahashi, R. et al. Interleukin-1β-induced pancreatitis promotes pancreatic ductal adenocarcinoma via B lymphocyte-mediated immune suppression. Gut 70(2), 330–341 (2021).
Zaghdoudi, S. et al. FAK activity in cancer-associated fibroblasts is a prognostic marker and a druggable key metastatic player in pancreatic cancer. EMBO Mol. Med. 12(11), e12010 (2020).
Hu, H. et al. Diagnostic value of S100P for pancreatic cancer: A meta-analysis. Tumour Biol. 35(10), 9479–9485 (2014).
Zou, W. et al. Up-regulation of S100P predicts the poor long-term survival and construction of prognostic signature for survival and immunotherapy in patients with pancreatic cancer. Bioengineered 12(1), 9006–9020 (2021).
Nakayama, H. et al. S100P regulates the collective invasion of pancreatic cancer cells into the lymphatic endothelial monolayer. Int. J. Oncol. 55(1), 211–222 (2019).
Chen, Y. et al. S100A2 is a prognostic biomarker involved in immune infiltration and predict immunotherapy response in pancreatic cancer. Front. Immunol. 23(12), 758004 (2021).
Lin, H., Yang, B. & Teng, M. T-cell immunoglobulin mucin-3 as a potential inducer of the epithelial-mesenchymal transition in hepatocellular carcinoma. Oncol. Lett. 14(5), 5899–5905 (2017).
Xie, J. et al. Identification of potential diagnostic biomarkers in MMPs for pancreatic carcinoma. Medicine (Baltimore) 100(23), e26135 (2021).
Ishige, F. et al. MIR1246 in body fluids as a biomarker for pancreatic cancer. Sci. Rep. 10(1), 8723 (2020).
Zhao, M. & Dai, R. HIST3H2A is a potential biomarker for pancreatic cancer: A study based on TCGA data. Medicine (Baltimore) 100(46), e27598 (2021).
Liu, B. et al. Construction of a five-gene prognostic model based on immune-related genes for the prediction of survival in pancreatic cancer. Biosci. Rep. 41(7), BSR20204301 (2021).
Jia, Y. et al. Development of a 12-biomarkers-based prognostic model for pancreatic cancer using multi-omics integrated analysis. Acta Biochim. Pol. 67(4), 501–508 (2020).
Shi, X. et al. Three-lncRNA signature is a potential prognostic biomarker for pancreatic adenocarcinoma. Oncotarget 9(36), 24248–24259 (2018).
Qi, B. et al. An immune-related lncRNA signature for the prognosis of pancreatic adenocarcinoma. Aging (Albany NY). 13(14), 18806–18826 (2021).
Yu, Y., Feng, X. & Cang, S. A two-microRNA signature as a diagnostic and prognostic marker of pancreatic adenocarcinoma. Cancer Manag. Res. 13(10), 1507–1515 (2018).
Wu, Z. et al. Novel necroptosis-related gene signature for predicting the prognosis of pancreatic adenocarcinoma. Aging (Albany NY). 14(2), 869–891 (2022).
Tang, R. et al. The role of m6A-related genes in the prognosis and immune microenvironment of pancreatic adenocarcinoma. PeerJ 28(8), e9602 (2020).
Meng, Z. et al. The m6A-related mRNA signature predicts the prognosis of pancreatic cancer patients. Mol. Ther. Oncol. 29(17), 460–470 (2020).
Acknowledgements
Figure 1 was designed using images from Freepik.com (https://www.freepik.com/).
Funding
This study was supported by Ningbo Health Branding Subject Fund (PPXK2018-03).
Author information
Authors and Affiliations
Contributions
Conception and writing, L.D.; Charting and writing, J.M.; data analysis, L.D.; reference acquisition, X.C.C.; comments and suggestions, C.D.L.; manuscript revision, C.J.L. All the authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dai, L., Mugaanyi, J., Cai, X. et al. Pancreatic adenocarcinoma associated immune-gene signature as a novo risk factor for clinical prognosis prediction in hepatocellular carcinoma. Sci Rep 12, 11944 (2022). https://doi.org/10.1038/s41598-022-16155-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-16155-w
- Springer Nature Limited