Abstract
In neuroblastoma, MYCN amplification and 11q-deletion are important, although incomplete, markers of high-risk disease. It is therefore relevant to characterize additional alterations that can function as prognostic and/or predictive markers. Using SNP-microarrays, a group of neuroblastoma patients showing amplification of one or multiple 12q loci was identified. Two loci containing CDK4 and MDM2 were commonly co-amplified, although amplification of either locus in the absence of the other was observed. Pharmacological inhibition of CDK4/6 with ribociclib or abemaciclib decreased proliferation in a broad set of neuroblastoma cell lines, including CDK4/MDM2-amplified, whereas MDM2 inhibition by Nutlin-3a was only effective in p53wild-type cells. Combined CDK4/MDM2 targeting had an additive effect in p53wild-type cell lines, while no or negative additive effect was observed in p53mutated cells. Most 12q-amplified primary tumors were of abdominal origin, including those of intrarenal origin initially suspected of being Wilms’ tumor. An atypical metastatic pattern was also observed with low degree of bone marrow involvement, favoring other sites such as the lungs. Here we present detailed biological data of an aggressive neuroblastoma subgroup hallmarked by 12q amplification and atypical clinical presentation for which our in vitro studies indicate that CDK4 and/or MDM2 inhibition also could be beneficial.
Similar content being viewed by others
Introduction
Neuroblastoma (NB) is a pediatric cancer that affects 1 in 8000 children and, despite its relative rarity, is a leading cause of cancer-related death in childhood. It derives from immature cells of the sympathetic nervous system originating from the neural crest. The primary tumors commonly manifest in the abdomen along sympathetic ganglia or in the medullary region of the adrenal gland. Over half of these patients already have metastasis at diagnosis, the most common sites being bone marrow (70%), cortical bone (56%), regional and distant lymph nodes (31%), liver (30%), and intracranial and orbital sites (18%), while rarer locations are lung (3%) or central nervous system (0.6%)1. The clinical behavior varies greatly, ranging from spontaneous regression or maturation to aggressive tumor growth with fatal outcome despite intensive multimodal therapy. Poor prognosis is especially apparent among patients with high-risk neuroblastoma (HR-NB), with less than 50% 5-year survival2,3.
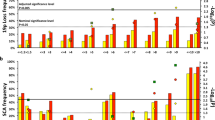
The clinical heterogeneity of this disease is also reflected at the molecular level, at which distinct genomic features of the tumor are critical for stratification and treatment decision. Whereas there is a relative paucity of recurrent single-nucleotide variants (SNVs)4,5,6, persistent copy number alterations (CNAs) are common, so analysis of somatic chromosomal aberrations is crucial7. Tumors with good prognosis are generally near triploid and display aneuploidy with whole chromosome gains and losses (i.e., numerical chromosomal aberrations, NCAs). High-risk tumors with poor prognosis are predominantly near di- or tetraploid and are associated with recurrent segmental chromosomal aberrations (SCAs), such as deletion of chromosome arm 11q or amplification of the MYCN oncogene (MNA)7. Other genomic features linked to adverse outcome include 1p-deletion, 1q-gain, distal 6q-deletion, 19q loss, and 17q-gain7,8,9,10,11. Further genomic alterations related to poor prognosis are those associated with telomere maintenance, such as aberrations in ATRX or TERT genes, as well as genetic alterations in the RAS-MAPK pathway, oncogenic mutations of ALK, or low-frequency mutations in TP5312,13.
MNA is a well-defined biomarker present in 25–30% of all NB, and MNA analysis has been mandatory in clinical practice since the 1980s due to its prognostic and diagnostic value14,15. Apart from the MYCN locus amplification, a focal amplification of the ALK gene has been detected in 1–2% of the tumors, almost exclusively in association with MYCN amplification16. In addition to MYCN and ALK, other amplification regions have been described, although they are infrequent, including various chromosome segments such as 7q21, 11q13, and 16q2210,17. The oncogenic contribution of these amplicons is still unclear, emphasizing the need for further characterization of these regions. We and others have previously also reported high-grade amplification of chromosomal regions 12q13.3–14.1 and 12q15 in a small subset of tumors, sometimes in combination with gain of the chromosomal region 11q13 that includes the CCND1 gene encoding Cyclin D7,10,17,18,19,20,21,22. Amplification of chromosome 12 regions and over-expression of both MDM2 and CDK4 are frequently detected in lipo- and osteosarcoma, in which this amplification is associated with poor prognosis23,24,25,26,27. This has also been reported in other cancers including bladder28, breast29, brain30, and lung31 cancers and melanomas31. Among the different malignancies displaying 12q amplification, two regions mentioned before located at 12q13.3–14.1 and 12q15 are commonly co-amplified. These loci are centered on two genes with a seminal role in cancer development: cyclin-dependent kinase 4 (CDK4), a key regulator of cell cycle progression, and mouse double minute 2 (MDM2), a proto-oncogene involved in p53 regulation. However, cases in which these amplicons appear independent of each other have also been reported23,28,32.
To further elucidate the extent, frequency, and role of the amplification of chromosomal regions in 12q in NBs, we present a detailed analysis of the genetic and clinical features of this subset of NB cases. Furthermore, through cell-based assays, we demonstrate that the pharmacological inhibition of MDM2 and CDK4 is feasible and that these inhibitors decrease proliferation. Therefore, MDM2 and/or CDK4 inhibition could provide an alternative targeted therapy in this patient group.
Results
Analysis of copy number and amplicons
Among genomic profiles derived from SNP microarrays of NB tumors from 436 Swedish patients, we identified nine patients with tumors displaying structural rearrangements and high-grade amplification involving chromosomal regions in the 12q chromosome arm. Eight additional patients with 12q-amplified NB tumors from Poland, Austria, Norway, and Spain as well as two 12q-amplified cell lines were included in the study. Although the genomic profiles of the individual cases differed in the number of amplified regions in chromosome 12, and in the size and junction sites of these amplicons, two loci in 12q13–14 and 12q15 were consistently co-amplified in 16 out of the 19 NB cases (Fig. 1 and Supplemental Fig. 1).
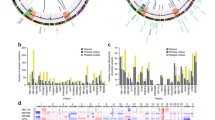
Genomic findings regarding the tumors with 12q amplification. (A) Whole-genome copy number profiles generated from WGS for three NB tumors with 12q amplification and (B) copy number profiles for chromosome 12, 57–72 Mb from the p-terminal with corresponding amplification peaks. (C) Amplified regions at 12q14 in relation to genomic position and genes with the shortest region of overlap (SRO), indicated in blue, based on amplicon boundaries of all primary tumors and cell lines analyzed. (D) Amplified regions at 12q15 in relation to genomic position and genes with the SRO indicated in blue.
The most proximal of the two regions (12q14.1) was amplified in 18 out of the 19 cases and centered on the CDK4 gene. The shortest region of overlap (SRO) of the amplified loci at 12q14.1 was located at Chr12: 58.135–58.270 Mb (defined by 73R6 and AuGJ), harboring the genes TSPAN31, CDK4, MARCH9, CYP27B1, METTL1, METTLE21B, TSFM, AVIL, and CTDSP2 (Fig. 1C). The more distal region at 12q15 was amplified in 17 out of the 19 cases, mainly centered on MDM2, although one of the 12q15 amplicons started in FRS2 intron 1 located distally to MDM2. The SRO of this more distal amplified region (12q15) was located at Chr12: 69,166–69,270 Mb (defined by AuGJ and SpHM) and contained the genes NR_073494, MDM2, and CPM (partial) (Fig. 1D). The fibroblast growth factor receptor substrate 2 (FRS2) gene is located distal to MDM2 at 69.86–69.97 Mb within the 12q15 chromosomal region. The complete FRS2 gene was included among the 12q15 amplicons in the tumors of 14 cases. To summarize, co-amplification of CDK4 and MDM2 was seen in 15 cases (including the two cell lines), amplification of CDK4 in the absence of MDM2 was seen in three cases, and MDM2 amplification in the absence of CDK4 was seen in one case (Table 1).
Furthermore, among this group of 12q-amplified NBs, concomitantly amplified loci included MYCN (seven cases including the two cell lines), multiple loci in 1p (one case), ALK (one case), and CCND1 (cell line LS) (Supplementary Fig. 1). In addition to the CCND1 amplification seen in the cell line, CNAs involving CCND1 were observed in four cases in which three tumors displayed a small focal gain and one tumor a large 11q gain. Segmental gain of 17q was seen in ten cases, whereas whole chromosome 17 gain was seen in five cases (Supplementary Fig. 1).
Expression analysis and survival analysis
We hypothesized that the amplified regions could harbor one or a few oncogenes that drive tumor progression and that the oncogenic effect is dose dependent, meaning that amplification leads to increased mRNA levels. Among the genes analyzed in the two 12q regions, 12 had over 20-fold higher expression in the 12q-amplified group compared with the 12q non-amplified tumors, whereas an additional 12 genes had at least two-fold higher expression compared with the non-amplified tumors. A total of 25 out of 28 analyzed genes displayed statistically significant differences in expression levels (Table 2).
Kaplan–Meier analysis of the Swedish cohort (n = 36) indicates that high expression levels of CDK4, CYP27B1, and TSFM are associated with worse outcome, although not at a statistically significant level. Analysis of two datasets consisting of 88 and 498 NB tumors, publicly available in the R2 database, indicates that high expression of these three genes is associated with low overall survival, with the strongest association between adverse outcome and high expression levels for CDK4 (p = 9.1 × 10–8 and p = 2 × 10–20, respectively) (Supplementary Table 2).
Among the five 12q-amplified tumors included for gene expression analysis, one (i.e., 17E2) did not include MDM2 in the 12q15 amplicon. This sample displayed MDM2 expression at levels similar to those of non-amplified NB tumors with different genomic profiles (e.g., numerical only, MNA, and 11q-deleted) (Supplementary Fig. 3).
Clinical features
As shown in Table 1, out of 17 patient cases for which detailed clinical data were available, seven had localized disease and ten had metastatic spread at time of diagnosis. The 12q-amplified NBs were all of abdominal origin, of which six had a renal location, and not surprisingly initially aroused suspicion of Wilms’ tumor, exemplified by CT images of NB case 45R1 (Fig. 2A–C) and a Wilms’ tumor from a patient of similar age (Fig. 2D–E). A subset of NB tumor hallmarked by 12q-amplification showed atypical large cell morphology compared to other genomic subgroups of NB showing conventional type of nuclei (Fig. 2F–H).
Clinical presentation of 12q-amplified tumors. (A,B) CT images of patient 45R1 showing a large primary NB tumor adjacent the upper part of the right kidney and (C) metastases of the lungs as indicated by arrows. (D,E) Corresponding images of a Wilms’ tumor with comparable growth pattern in a child at the similar age. Hematoxylin–eosin staining showing: (F) conventional poorly differentiated neuroblastoma with uniform nuclei and incipient resetting, patient 17E2, (G) large cell neuroblastoma with partly spindling cells and atypical mitoses, patient 45R1 and (H) undifferentiated large and anaplastic cell neuroblastoma with coarse nuclear chromatin, prominent nucleoli and atypical mitoses, patient 38R3. (I) Kaplan–Meier overall survival analysis show upon poor outcome among NB patients with 12q amplification in comparison to NB patients of different genomic subgroups.
Moreover, the metastatic pattern showed statistically significant difference from what is common for high-risk NB, with a very low incidence of bone marrow involvement (4/10 = 40%) in our cohort as compared to the HR-NBL1/SIOPEN study reporting bone marrow involvement in 76.7% of included cases (375/489)33 (p = 0.015, Fisher’s exact test). Instead, lung metastases were favored (6/10 = 60%), that is an uncommon feature reported by the international neuroblastoma risk group project to be present in 3.6% of all NB (100/2808)34 (p > 0.00001, Fisher´s exact test). Kaplan–Meier analysis of 270 NB cases show that patients with 12q-amplified NB has extraordinary poor outcome in comparison to patients with tumors of other genomic subgroups (i.e. MNA, 11q-deleted, numerical only, 17q-gained or other segmental, as described previously7) (Fig. 2H). Among the patients with 12q-amplification, two of six patients with localized disease show long-term survival (> 300 months) while nine out of ten with metastasized disease had fatal outcome.
Mutational analysis
Whole-genome sequencing (WGS; n = 3) and whole exome sequencing (WES; n = 5) were performed in tumors with 12q amplification from eight patients. Somatic calling for the four tumors for which constitutional DNA was available for sequencing (WGS: n = 3, WES: n = 1) detected an average of 18 nonsynonymous mutations (range 7–29; Supplementary Table 3). For tumors sequenced without matched normal controls (WES, n = 4), we focused on rare nonsynonymous variants in 574 genes with established connections to cancer (Supplementary Table 1). Using this approach, an average of 11 variants remained (range 7–19) (Supplementary Table 3) among the “tumor only” cases. Among these variants, we did not detect any novel nonsynonymous SNVs within the amplified regions at chromosome 12, except a rare INHBE variant seen in one patient. No variants affecting ALK, PHOX2B, NRAS, KRAS, or TP53 were identified in the investigated tumor or germline DNA. However, three rare SNVs in ATRX were detected among the eight 12-amp tumors analyzed by sequencing. Alterations in genes connected to telomere maintenance (i.e., TERT and ATRX) have recently been shown to be recurrent in non-MYCN-amplified NBs, which are also connected to adverse outcome12. The ATRX SNVs included a novel nonsense variant in exon 13 (NM_000489.5; c.3979G > T; p.E1327*) in case SpHM and a novel missense variant in exon 30 (NM_000489.5; c.6327T > G; p.S2147R) in case AuHH, which is predicted to be damaging by altering the protein function. Moreover, an uncommon hemizygous ATRX missense variant (rs200709847) was detected in 34R0 (Supplementary Table 3). Furthermore, a somatic missense mutation in case (59R9) was detected in SIN3A, a gene recently associated with the repression of TERT gene expression35.
In addition to sequence analysis, re-examination of array data showed intergenic ATRX deletion in one case (BG12), while segmental alterations in the proximity of the TERT locus were detected in three cases: 16R0, SpAI, and in the NGP cell line. The breakpoints for the segmental alterations in the two non-MYCN-amplified tumors were located approximately 100 kb proximal to the TERT transcriptional starting point, while the corresponding breakpoint in NGP was located 1 Mb from TERT.
CNAs identified by WGS analysis of three primary tumors were in complete concordance with SNP microarray profiles (Fig. 1A and Supplementary Fig. 1), including amplified regions. WGS also showed extensive structural rearrangements between and within amplified regions (Fig. 3 and Supplementary Table 4).
Mutational spectra and structural variants in 12q-amplified tumors. Circos plots showing structural variants, CNAs, and somatic SNVs. Copy number plots calculated based on the coverage ratio between tumor and corresponding normal tissue are shown on the inner circle, with gain of genomic material indicated in red and loss of genomic material indicated in blue. The lines within the inner circle indicate structural variants within and between chromosomes, while genes affected by somatic SNVs are shown outside the outer circle.
TP53 status in NB cell lines
Prior to treatment with Nutlin-3a, ribociclib, and abemaciclib in NB cell lines, different genomic aspects of the cell lines were characterized by SNP microarray analysis (Supplementary Fig. 2) together with mutational validation of previously reported TP53 mutations36,37,38,39 by Sanger sequencing or by RT-PCR. The four cell lines (KELLY, SK-N-BE(2), SK-N-DZ, and SK-N-FI) displayed pathogenic homozygous missense mutations, rendering p53 inactive, whereas NGP has a TP53 heterozygous mutation which allows a wild-type function of p53 (Supplementary Fig. 4A). RT-PCR verified the deletion of exons 10 and 11 of TP53 that causes a protein truncation of the p53 C-terminus in SK-N-AS (Supplementary Fig. 4B). Relevant genetic alterations detected in the NB cell lines used in this study are summarized in Supplementary Fig. 4C.
Treatment response to Nutlin-3a, ribociclib, and abemaciclib
To study whether cells with the co-amplification of MDM2 and CDK4 can benefit from inhibition directed toward these specific targets, we performed viability studies using MDM2i and CDK4/6i in NB cell lines. For inhibition of MDM2, Nutlin-3a (a selective antagonist of p53-MDM2 binding) was used, while two highly selective inhibitors, ribociclib and abemaciclib, were used for CDK4 inhibition. MDM2 inhibition showed that five out of ten cell lines were sensitive to Nutlin-3a treatment with 50% cell viability reduction already present after 24 h, while the remaining cell lines were resistant to Nutlin-3a even after 72 h of treatment (Fig. 4A, upper panels). The drug concentrations that cause a 50% reduction in cell viability (IC50 values) already produce a significant difference in the case of Nutlin-3a treatment between p53-mutated and p53wild-type (p53wt) cell lines after 24 h of treatment (Fig. 4B, upper panel).
Neuroblastoma cell line response to MDM2i or CDK4/6i. (A) Normalized cell viability after treatment with Nutlin-3a (upper panels), ribociclib (middle panels), and abemaciclib (lower panels) for 24, 48, and 72 h. (B) IC50 values calculated after 24, 48, and 72 h of treatment. The bars represent the 95% CI for each IC50 value with statistical significance indicated as follows: *p < 0.01, **p < 0.001, *** p < 0.0001, and ****p < 0.00001.
The response of Nutlin-3a was highly dependent on TP53 mutational status, with treatment-resistant cell lines having homozygous mutated/deleted TP53 while sensitive cell lines had at least one wild-type copy of TP53, such as NGP, which has a heterozygous TP53 mutation (Supplementary Fig. 4). Regarding CDK4/6i, nine out of ten cell lines displayed a significant response to CDK4/6 inhibition with ribociclib (Fig. 4A, middle panels) or abemaciclib (Fig. 4A, lower panels). Of all investigated cell lines, only SK-N-FI displayed resistance to both ribociclib and abemaciclib treatment (Fig. 4A). No significant difference in IC50 values and response was detected between two CDK4/MDM2-amplified cell lines and non-amplified cell lines after treating the cells with CDK4i (Fig. 4A,B, middle and lower panels). CDK4/MDM2 amplification did not have any significant impact on treatment response in comparison with other TP53wt cell lines after MDM2i treatment (Fig. 4A,B, upper panels). No obvious response pattern in relation to the MYCN amplification and inhibition of CDK4 or MDM2 could be observed. For the detailed response of each treatment and cell line, see Supplementary Fig. 5, while details related to IC50 values for each cell line, treatment, and time point can be found in Supplementary Fig. 6.
Synergy effect of combinatorial CDK4/6 and MDM2 inhibition
Combination treatment with MDM2i and CDK4/6i did not result in a synergetic effect, furthermore the synergistic pattern was not conclusive, for this reason, the overall synergy score was used in order to interpret the results. The combination treatment induced a slight reduction in proliferation compared with single-agent treatment in the cell lines with p53wt, showing a positive score in the overall synergy score (Fig. 5, Table 3). In more detail, the p53wt NB cell lines displayed a synergetic area greater than 10, meaning that the response is over 10% greater than expected at certain drug concentrations. Given the calculated overall synergy scores and the score for the most synergetic area, we can conclude that the combination of MDM2i + CDK4/6i has a positive additive effect in p53wt NB cell lines. No significant difference in drug combination response was seen in LS and NGP, MDM2/CDK4-co-amplified cell lines, as compared with other p53wt cell lines (Fig. 5, Table 3).
Inhibition dose–response matrix and synergy score heatmaps for representative neuroblastoma cell lines. Dose–response matrix and synergy score heatmaps generated by SynergyFinder for combinational treatment with Nutlin-3a and ribociclib and with Nutlin-3a and abemaciclib for the cell lines LS (CDK4/MDM2-amp, p53wt, MNA), SK-N-SH (p53wt, MNA), SK-N-BE (p53mut, MNA), and SK-N-AS (p53mut). Dose–response matrix and synergy score heatmaps for all used cell lines can be found in Supplementary Fig. 5.
For the p53mutated (p53mut) cell lines, the synergy score of three cell lines (KELLY, SK-N-BE(2), and SK-N-DZ) is close to 0, meaning that there is no difference between the single and combination treatments. These cell lines experienced a slight additive effect at specific concentrations of the combined drugs. The remaining two p53mut cell lines (SK-N-AS and SK-N-FI) presented an antagonistic additive overview synergy score having a negative score in the most synergetic area. SK-N-AS displayed a later response to CDK4/6i with abemaciclib than did the rest of the cell lines, while SK-N-FI showed no significant response to CDK4/6i or MDM2i after 72 h of treatment (Fig. 4A,B).
Protein levels of the CDK4/6 and MDM2 pathways
Downstream proteins of the CDK4/6 and MDM2 pathways were analyzed to verify the effect of the inhibitors used for each cell line at its respective IC50 value. CDK4/6 inhibition by both abemaciclib and ribociclib led to a marked decrease in RB phosphorylation levels at Ser780 in most used cell lines, except for the CDK4 and MDM2 co-amplified cell lines (LS and NGP), which have a higher expression than that of the experimental control. Regarding the phosphorylation of RB at Ser807/811, a similar pattern was found, except for KELLY, a cell line that had no changes in the expression at all. In the case of NGP, a decrease in pRB Ser807/811 has been observed (Fig. 6 and Supplementary Fig. 8). CDK4/6i also evoked increased protein levels of CDK4 (in all cell lines) and CDK6 (in all cell lines except for SH-SY5Y, LS, and NGP). Interestingly, CDK4/6i also led to increased levels of PUMA, a downstream target of p53, in four out of the five p53mut cell lines (KELLY, SK-N-DZ, SK-N-BE(2), and SK-N-AS).
Effect on proteins of the CDK4 and MDM2 pathway after CDK4/6 or MDM2 inhibition. The cells were treated with abemaciclib (Abe), ribociclib (Rib), and Nutlin-3a (N3a) at IC50 values for 72 h before protein extraction. This figure is a representation of the protein expression using different blots. The blots are delineated to clarify that belong to different gels or to different parts of the same gel. Each blot was normalized using KU80 as control. Exposure time for each protein analysis was the same for all cell lines.
MDM2 inhibition with Nutlin-3a led to increased levels of p21 and MDM2 in all p53wt cell lines. MDM2i also produced an overexpression of PUMA in all p53wt cell lines as well as in KELLY and SK-N-DZ. We also detected that, after Nutlin-3a treatment, all cell lines (with the exception of SK-N-BE(2)) experienced an increase in p53 (Fig. 6).
Discussion
In this study we investigated the amplification of two common regions at 12q13.3–14.1 and 12q15 detected in a subgroup of NB tumors. These two regions at chromosome 12 presented co-amplification in 15 out of 19 analyzed NB samples, and gene expression analysis showed that most of the genes in the amplicons were over-expressed compared with other NB tumors (Table 2). In other tumor types displaying the co-amplification of 12q13.3–14.1 and 12q15, CDK4 and MDM2 are often reported as possible targets for the respective regions25,28,29,30, and in our cohort of 12q-amplified cases, amplification of CDK4 was seen in 18 out of 19 samples, while amplification of MDM2 was seen in 16 out of 19 samples (Fig. 1 and Supplementary Fig. 1). Fibroblast growth factor receptor substrate 2 (FRS2), also implicated as an oncogenic driver in the 12q15 locus, was amplified in 14 out of the 19 cases with 12q amplification (Fig. 1D). The emergence of gene amplification is still not fully understood but likely involves a multistep process starting with an initial catastrophic event causing DNA breaks followed by retention of cells for which the alteration provide a selective advantage relative its environment, for review see Matsui et al.40. Our previous analyses of NB show that the MYCN amplicon occasionally is accompanied by several additional amplification peaks spread over chromosome region 2p41. The amplified regions are rarely larger that 3–4 Mb and suggests that carrying very large amplified segment provides (e.g. > 10 Mb) is a proliferative disadvantage for the cell. This could be due the risk of including less advantageous genes present in the larger amplicon or the replicative burden of carrying large amplification. Among the investigated cases with 12q-amplification, no one displayed one single, large continuous amplified segment including CDK4, MDM2 and FRS2. Instead, the tumors most often presented two distinct amplification peaks; one centered at CDK4 and another centered at MDM2 (and FRS2). The specificity of this complex, yet highly specific amplification of these genes, suggests a very strong combined tumorigenic driver effect based on cooperative function.
Interestingly, among the 12q-amplified NB samples, five also had focal gain or amplification at 11q involving CCND1 encoding cyclin D1. Cyclin D1 interacts with CDK4/6 and has a crucial role in G1/S transition by regulating RB phosphorylation42. Hence, CCND1 gain could limit the biological consequences of saturation that might occur with endogenous cyclin D expression levels.
Aberrant telomere maintenance in NB is promoted by alterations of ATRX or TERT or through MYCN-driven TERT re-expression and is linked to poor prognosis in NB12,43. Among the 19 cases in our cohort, segmental variants associated with the ATRX gene were found in two patient samples, while segmental rearrangements in the proximity of the TERT loci were detected in three cases. Among the eight patient samples investigated by WES/WGS, three had rare SNVs in ATRX, including a truncated mutation (Supplementary Table 3). Apart from ATRX, few somatic SNVs with obvious relevance for cancer were detected by WES/WGS with an overall low mutational load, which is concordant with previous studies4,5,6,44. The sequenced tumors lacked pathogenic variants in ALK, PHOX2B, TP53, or genes of the RAS-MAPK signaling pathway, which have previously been connected to NB, as well as variants in WT1, TRIM28, FBXW7, NYNRIN, or KDM3B, genes previously linked to Wilms’ tumor45.
The adverse outcome seen for most patients with 12q-amplified tumors (Fig. 2I) indicates that new therapies are needed to improve treatment of this group of NB patients. Both MDM2 and CDK4 have been identified as prioritized targets in NB46 in ongoing pediatric trials of the CDK4/6 inhibitors abemaciclib (LY2835219) and palbociclib (PD0332991) (ClinicalTrials.gov: NCT02255461, NCT02644460), while a trial of ribociclib (LEE011) has recently been completed47. MDM2 targeting agents such as HDM201 (Novartis) are being evaluated in NB (ClinicalTrials.gov: NCT02780128), while RG7112 (Hoffman-La Roche) is in trial in adult solid cancer (ClinicalTrials.gov: NCT00559533). It has been suggested that combinatorial therapies may increase the efficacy in patients with dysregulation of both MDM2 and CDK4. Synergy between CDK4/6 and MDM2 inhibitors has been observed in preclinical studies in liposarcoma48, melanoma,49 and breast cancer50, although contradictory results have been reported regarding the synergistic effects of a combined MDM2 and CDK4 targeting strategy in sarcoma48,51.
CDK4 and/or MDM2 inhibition, either as single-drug or combined treatment, showed that most investigated cell lines were sensitive to CDK4i, although SK-N-F1 displayed resistance to both ribociclib and abemaciclib (Fig. 4A,B, middle and lower panels), possibly due to high expression of multidrug resistance protein 1 (MDR1). Sensitivity to MDM2 inhibition with Nutlin-3a was only seen in p53wt cell lines (Fig. 4A,B, upper panels). The two cell lines with co-amplification of MDM2 and CDK4 displayed sensitivity to both MDM2i and CDK4/6i at levels like those of other p53wt cell lines (Table 3), indicating that directed therapy could be beneficial, despite the amplification and overexpression of the respective target genes. The combination of MDM2i and CDK4i did not present synergetic effects concordant with a recent study of combined MDM2/CDK4i in NB52. According to the overall synergy score, a positive additive effect was found in p53wt cell lines, including the MDM2/CDK4-amplified cell lines, even though non synergetic effect was detected. No positive or even antagonistic effect was seen when combining CDK6i and MDM2i in p53mut cell lines due to lack of response to Nutlin-3a in these cells (Table 3). However, as less than 2% of NB patients have tumors with p53 mutations at the time of diagnosis53,54, this indicates that a large group of NB patients may benefit from MDM2i, and possibly in combination with CDK4/6i in some specific cases. However, the broad sensitivity of MDM2i and CDK4i in investigated cell lines show that MDM2- and CDK4-amplification per se are not reliable biomarkers for response or non-response. Furthermore, with the rapid development of novel small molecule inhibitors or other therapies follows exponential increase of possible drug combinations. This requires additional pre-clinical studies using organoids, PDXs or other model systems in order to provide a strong biological rational for appropriate drug combinations before advancement to human subjects. If proceeding, and given NB being a rare neoplasm, a tumor-agnostic approach would likely be more feasible to establish critical safety and efficacy factors of dosing, timing and, sequencing of regimes for combination treatment.
Clinically, the degree of malignant behavior varied between tumors with 12q amplification, ranging from a localized, triploid tumor with a favorable outcome after surgical treatment only, to primary metastatic disease with a rapid demise. Interestingly, atypical clinical features in some of those tumors causing fatal progression included; non-involvement of bone or bone marrow (the major site of metastatic disease in NB), intra renal primary site and, presence of an inferior cava tumor thrombus and lung metastasis, suggesting an advanced Wilms’ tumor (WT). The deceptive morphology initially also favored the idea of this tumor. The amplifications in the mentioned regions might promote clinical features related to metastatic homing and tumor invasion that mimic WT. The co-amplification of CDK4/MDM2 has also been noted to change the location and clinical features of rare cases of inflammatory myofibroblastic tumors, causing them to be mistaken for gastric cancer with local invasion of the spleen and diaphragm55. Predilection for lung metastasis have been confirmed in several malignancies such as melanoma, sarcoma, kidney- and breast cancer, in which the preferential site for metastasizing also is subtype dependent56. Examination of gene signatures in breast cancer has shown that whereas the signature for bone metastasis was mainly differentiated by cell surface- and secretory proteins57, the signature for lung metastasis correlated with markers for poor prognosis, were less specific for environment and instead promoted aggressive growth and invasiveness. If corresponding gene signature pattern also is present in NB is to our knowledge unknown. NB has also been reported to display a remarkable plasticity relating to the neoplastic origin of NB58,59,60,61,62,63 that also allows transdifferentiation, with switching between adrenergic- and mesenchymal state. If 12q-amplification has effect on NB plasticity or cell state and whether this in turn has effect on infiltration and metastatic homing is not fully investigated. Ultimately, additional studies of disseminating NB cells, their capability to passage organ specific barriers and collaboration with the microenvironment are required to elucidate the specific molecular mechanisms triggered by 12q-amplification that change invasive growth and preferential site for metastatic colonization in NB.
In summary, we present a subgroup of NB characterized by amplification at chromosome 12q with subsequent overexpression of most genes within the amplified regions. Atypical features were observed associated with this subgroup that might correlate to a cell cycle deregulation phenotype given the persistent amplification of CDK4, which is further potentiated by MDM2 as well as FRS2 amplification in some cases. This study provides new information about the possibility of using MDM2i and CDK4/6i drug combination in p53wt NBs, including tumors with MDM2 and CDK4 amplification, although the effect of MDM2 inhibition is highly dependent on the presence of functional p53 activity.
Materials and methods
Sample collection, patients, and ethical statement
All experiments were performed in accordance with the approved guidelines and regulations. Tumor tissue was collected after written or verbal informed consent was obtained from parents/guardians according to ethical permits approved by the local ethics committees of Karolinska Institute and Karolinska University Hospital, the Experimental Research Ethics Committee of the University of Valencia/INCLIVA and the Spanish Society of Pediatric Hematology and Oncology, the University of Vienna, Wroclaw Medical University, and Oslo University Hospital. Genomic DNA was extracted from fresh frozen tumor or blood using the DNeasy Blood & Tissue kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. The patients’ clinical and genetic backgrounds are described in Table 1.
Morphology
The handling of specimen followed recommendations in standard protocol. Fresh material was immediately taken for biologic research and the fixed tumors were treated according to conventional pathology practice to diagnose neuroblastic tumors. Classification followed the International neuroblastoma Pathology Classification (INPC)64.
All tumor samples in the present paper had unfavorable histology; undifferentiated, poorly differentiated or anaplastic/large cell.
SNP microarray analysis
The Swedish samples (n = 436) were screened using Affymetrix SNP microarrays (Thermo Fisher Scientific, Waltham, MA) as described earlier7,44. In addition, 12 NB cell lines were also screened using SNP microarray analysis: IMR32, KELLY, NB69, SH-SY5Y, SK-N-AS, SK-N-BE(2), SK-N-DZ, SK-N-FI, SK-N-SH (ECACC, HPA Culture Collections, Salisbury, UK), SH-EP (ATCC, Manassas, VA), as well as the two 12q-amplified cell lines LS and NGP (kindly provided by Prof. Manfred Schwab, DKFZ Heidelberg, Germany). There were nine patients with tumors displaying high-grade 12q-amplification in the Swedish cohort. In addition to these samples, a selection of patients from Norway (n = 1), Spain (n = 3), Austria (n = 2), and Poland (n = 2) with tumors sharing a similar 12q-amplification pattern were also included (Supplementary Fig. 1). In total, 19 cases with 12q-amplification were included in the study (tumor material from 17 patients and two cell lines). For primary data analysis, GDAS software (Thermo Fisher Scientific) was used, while genomic profiles and amplicon boundaries were determined using either the Chromosome Analysis Suite (ChAS 3.3; Thermo Fisher Scientific) or the Copy Number Analyzer for Affymetrix GeneChip Mapping arrays (CNAG 3.0; Genome Laboratory, Tokyo, Japan; www.genome.umin.jp).
Massive parallel sequencing
WGS was performed on from three patients with tumors displaying 12q amplification, as evaluated from SNP microarrays. DNA from tumor material and matched constitutional DNA were subjected to sequencing and bioinformatical handling as described previously52 Briefly, sequencing was performed on Illumina instrumentation (Illumina, San Diego, CA) at Clinical Genomics, SciLife Laboratories, Stockholm, Sweden for average read depths of 62, 32, and 74 for tumor samples 45R1, 59R9, and 73R6, respectively, and of 30, 36, and 36 for constitutional DNA, respectively. Mapping, realignment around indels, and variant calling were carried out using the Sentieon suite of bioinformatics tools (Sentieon Inc., Mountain View, CA) (Sentieon version v201808.03). The Canvas tool (version 1.38.0.1554)53 was used to call CNAs while structural variants were called using the Manta tool (version 1.1.1)54, with filtering based on germline variation, artifacts caused by problematic regions or, presence in the SweGen Variant Frequency dataset (https://swefreq.nbis.se/) or in our in-house set of normal controls. Patient specific origin of tumor-normal pairs was verified using a previously developed Python script that calculate the shared fraction from 400,000 SNPs65. In addition to WGS, whole exome sequencing (WES) was performed on NB tumors from five additional patients together with constitutional DNA from one of these (17E2) using the SureSelect All Exon V5 50 Mb kit (Agilent, Santa Clara, CA) according to the supplier’s protocol. Thereafter, pair-end sequencing was performed on a HiSeq2500 sequencing system (Illumina) targeting an average of 50–60× raw read depth (GATC, Konstanz, Germany; Otogenetics, Atlanta, GA). Read trimming and mapping to the human reference genome were done using the CLC Genomics workbench (Qiagen, Aahus, Denmark).
Only high-quality called SNVs with a minimum of 10% variant allele frequency and a total read coverage of ten were considered for further analysis and evaluation. All synonymous variants or variants in non-coding regions were excluded except those affecting canonical splice sites. For further filtration, only variants present in 574 genes with established connections to cancer (listed in Supplementary Table 1) were kept when analyzing variants from tumors sequenced without corresponding normal tissue. The remaining variants were assessed manually using the Integrative Genomics Viewer (IGV)66. All genomic positions are given according to the human reference genome hg19/GRCh37, which was also used for mapping both WGS and WES.
Gene expression in the 12q amplicon regions
Total RNA was isolated from 36 tumors and two cell lines (LS and NGP) using ToTALLY RNA (Ambion/Thermo Fisher Scientific) prior cDNA synthesizing using the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA). cDNA corresponding to 250–500 ng of total RNA was loaded in triplicate onto TLDA cards (Applied Biosystems) to determine the expression levels of 31 genes (Table 2) located within the two common amplicon regions at 12q, together with housekeeping genes SDHA, HPRT1, GAPDH, GUSB, and UBC as endogenous controls. Gene expression levels were calculated using the delta delta C(t) method67 with the geometrical mean of the five endogenous controls. Fold change was calculated comparing 12q-amplified tumors (n = 5) and 12q non-amplified tumors (n = 31). Mean expression levels were calculated and compared gene-by-gene by two-tailed independent-sample t-test.
Kaplan–Meier plots were generated for each assay with median expression value as cutoff. Overall survival (OS) was defined as the time between diagnosis and death of any cause at follow-up. The OS of the patients was compared with gene expression levels using a log rank test, with a P-value less than 0.05 considered significant. These data were then compared with two publicly available expression datasets in the R2 database (R2: Genomics Analysis and Visualization Platform, http://r2.amc.nl): the Versteeg (n = 88) data derived from GeneChip Human Genome U133 Plus 2.0 Array (Affymetrix) and the SEQC dataset (n = 498) derived from transcriptome sequencing.
Cell culture, drug treatment, and viability assay
All NB cell lines were cultured under standard practice without antibiotics and routinely tested for mycoplasma. The cell line identity was verified by SNP microarray analysis (Supplementary Fig. 2) while TP53 mutational status was confirmed by Sanger sequencing and RT-PCR.
Ten NB cell lines with different genetic backgrounds68,69 were used for MDM2 and CDK4 inhibition: KELLY, SK-N-BE(2), SK-N-DZ, SK-N-AS, SK-N-FI, SK-N-SH, SH-SY5Y, NB69, LS, and NGP. The cell lines were plated, 24 h before treatment, in 96-well plates at a density of 5000–10,000 cells/well depending on the doubling time of each cell line. The cells were treated with the selective MDM2 antagonist drug Nutlin-3a (Selleckchem, Houston, TX) and/or the selective CDK4/6 inhibitors abemaciclib (MedChemExpress, Monmouth Junction, NJ) or ribociclib (MedChemExpress, Monmouth Junction, NJ). Concentrations of the tested inhibitors ranged from 20 to 10,000 nM (two-fold dilutions); DMSO was used as a control in an amount corresponding to the levels present in the highest used drug concentrations. The cell viability was measured by means of fluorescence using Presto Blue SH cell viability reagent (Thermo Fisher Scientific). Cell proliferation was monitored for 24, 48, and 72 h. The experiments were carried out in six replicates at each concentration and repeated at least three times. The concentrations that caused 50% inhibition of cell growth (IC50) were calculated using GraphPad Prism, version 8.4.3 (GraphPad Software, San Diego, CA).
Drug combination assay
To study a possible synergy effect on cell proliferation after dual MDM2 and CDK4/6 inhibition, cells were treated for 72 h with either (a) Nutlin-3a combined with ribociclib or (b) Nutlin-3a combined with abemaciclib. The concentration ranges were from 0 to 6000 nM for ribociclib and from 0 to 2000 nM for Nutlin-3a and abemaciclib. The viability was normalized as the percentage of inhibition prior to calculating the synergy score for drug combination responses, which were calculated based on the HSA reference model using SynergyFinder70. The derived synergy score data can be directly interpreted as the proportion of the cellular response associated with the drug interaction. As synergy scores near 0 have limited confidence, synergy scores between − 10 and 10 are considered to indicate additive interaction, with positive and negative score values denoting synergy and antagonism, respectively: synergy scores larger than 10 are considered to indicate synergetic interaction, whereas synergy scores below − 10 are considered antagonistic interaction70.
Western blotting
Proteins from cells treated for 72 h with corresponding IC50 concentration for respective cell lines and control cells were extracted using RIPA lysis buffer supplemented with proteinase and phosphatase inhibitors. Protein amount was quantified using the Pierce BCA Protein assay kit (Thermo Fisher Scientific). The same amount of total protein sample (20 µg) was separated from each sample using SDS-PAGE and transferred onto 0.2-µm PVDF membrane. In order to optimize the number of proteins analyzed from a single membrane the blots were cut in multiple pieces according to size. Uncropped blot images are presented in Supplementary Fig. 9. The blots were probed overnight at 4 °C with the following antibodies: CDK4 (sc-56277), CDK6 (sc-7961), MDM2 (sc-965), p53 (sc-126), PUMA (sc-374223), and RB (sc-102) from Santa Cruz Biotechnology (Dallas, TX); p-RBSer807/811 (#702097) from Thermo Fisher Scientific; and p-RBSer780 (#9307), p21 (#2947), and KU80 (#2573) from Cell Signaling Technology (Danvers, MA). Protein levels were quantified using ImageJ software60 with normalization against KU80.
Statistical analysis
The Kaplan–Meier estimates are based on data generated from 270 neuroblastoma cases with available clinical outcome and the analysis was performed using the IBM SPSS statistics for Windows, version 25 while the other statistical analyses were performed using GraphPad Prism, version 8.4.3 (GraphPad Software). The IC50 values and corresponding 95% confidence intervals (95% CIs) were calculated using dose–response sigmoidal curve with variable slope analysis (the log [inhibitor] vs. normalized response–variable slope). The difference in the IC50 values of two groups (p53mut vs. p53wt) was analyzed using Student’s t-test. P values < 0.05 were considered statistically significant.
References
DuBois, S. G. et al. Metastatic sites in stage IV and IVS neuroblastoma correlate with age, tumor biology, and survival. J. Pediatr. Hematol. Oncol. 21, 181–189 (1999).
Valteau-Couanet, D. et al. Long-term results of the combination of the N7 induction chemotherapy and the busulfan-melphalan high dose chemotherapy. Pediatr. Blood Cancer 61, 977–981. https://doi.org/10.1002/pbc.24713 (2014).
Pearson, A. D. et al. High-dose rapid and standard induction chemotherapy for patients aged over 1 year with stage 4 neuroblastoma: A randomised trial. Lancet Oncol. 9, 247–256. https://doi.org/10.1016/S1470-2045(08)70069-X (2008).
Pugh, T. J. et al. The genetic landscape of high-risk neuroblastoma. Nat. Genet. 45, 279–284. https://doi.org/10.1038/ng.2529 (2013).
Pandey, G. K. et al. The risk-associated long noncoding RNA NBAT-1 controls neuroblastoma progression by regulating cell proliferation and neuronal differentiation. Cancer Cell 26, 722–737. https://doi.org/10.1016/j.ccell.2014.09.014 (2014).
Molenaar, J. J. et al. Sequencing of neuroblastoma identifies chromothripsis and defects in neuritogenesis genes. Nature 483, 589–593. https://doi.org/10.1038/nature10910 (2012).
Caren, H. et al. High-risk neuroblastoma tumors with 11q-deletion display a poor prognostic, chromosome instability phenotype with later onset. Proc. Natl. Acad. Sci. U. S. A. 107, 4323–4328. https://doi.org/10.1073/pnas.0910684107 (2010).
Caron, H. Allelic loss of chromosome 1 and additional chromosome 17 material are both unfavourable prognostic markers in neuroblastoma. Med. Pediatr. Oncol. 24, 215–221 (1995).
Ognibene, M., Morini, M., Garaventa, A., Podesta, M. & Pezzolo, A. Identification of a minimal region of loss on chromosome 6q27 associated with poor survival of high-risk neuroblastoma patients. Cancer Biol. Ther. 21, 391–399. https://doi.org/10.1080/15384047.2019.1704122 (2020).
Depuydt, P. et al. Genomic amplifications and distal 6q loss: Novel markers for poor survival in high-risk neuroblastoma patients. J. Natl. Cancer Inst. 110, 1084–1093. https://doi.org/10.1093/jnci/djy022 (2018).
Abbasi, M. R. et al. Impact of disseminated neuroblastoma cells on the identification of the relapse-seeding clone. Clin. Cancer Res. 23, 4224–4232. https://doi.org/10.1158/1078-0432.CCR-16-2082 (2017).
Ackermann, S. et al. A mechanistic classification of clinical phenotypes in neuroblastoma. Science 362, 1165–1170. https://doi.org/10.1126/science.aat6768 (2018).
Eleveld, T. F. et al. Relapsed neuroblastomas show frequent RAS-MAPK pathway mutations. Nat. Genet. 47, 864–871. https://doi.org/10.1038/ng.3333 (2015).
Brodeur, G. M., Seeger, R. C., Schwab, M., Varmus, H. E. & Bishop, J. M. Amplification of N-myc in untreated human neuroblastomas correlates with advanced disease stage. Science 224, 1121–1124. https://doi.org/10.1126/science.6719137 (1984).
Seeger, R. C. et al. Association of multiple copies of the N-myc oncogene with rapid progression of neuroblastomas. N. Engl. J. Med. 313, 1111–1116. https://doi.org/10.1056/NEJM198510313131802 (1985).
De Brouwer, S. et al. Meta-analysis of neuroblastomas reveals a skewed ALK mutation spectrum in tumors with MYCN amplification. Clin. Cancer Res. 16, 4353–4362. https://doi.org/10.1158/1078-0432.CCR-09-2660 (2010).
Guimier, A. et al. Clinical characteristics and outcome of patients with neuroblastoma presenting genomic amplification of loci other than MYCN. PLoS One 9, e101990. https://doi.org/10.1371/journal.pone.0101990 (2014).
Caren, H. et al. High-resolution array copy number analyses for detection of deletion, gain, amplification and copy-neutral LOH in primary neuroblastoma tumors: Four cases of homozygous deletions of the CDKN2A gene. BMC Genom. 9, 353. https://doi.org/10.1186/1471-2164-9-353 (2008).
Molenaar, J. J., van Sluis, P., Boon, K., Versteeg, R. & Caron, H. N. Rearrangements and increased expression of cyclin D1 (CCND1) in neuroblastoma. Genes Chromosomes Cancer 36, 242–249. https://doi.org/10.1002/gcc.10166 (2003).
Mosse, Y. P. et al. Neuroblastomas have distinct genomic DNA profiles that predict clinical phenotype and regional gene expression. Genes Chromosomes Cancer 46, 936–949. https://doi.org/10.1002/gcc.20477 (2007).
Amoroso, L. et al. Genomic coamplification of CDK4/MDM2/FRS2 is associated with very poor prognosis and atypical clinical features in neuroblastoma patients. Genes Chromosomes Cancer 59, 277–285. https://doi.org/10.1002/gcc.22827 (2020).
Ambros, I. M., Brunner, C., Abbasi, R., Frech, C. & Ambros, P. F. Ultra-high density SNParray in neuroblastoma molecular diagnostics. Front. Oncol. 4, 202. https://doi.org/10.3389/fonc.2014.00202 (2014).
Italiano, A. et al. HMGA2 is the partner of MDM2 in well-differentiated and dedifferentiated liposarcomas whereas CDK4 belongs to a distinct inconsistent amplicon. Int. J. Cancer 122, 2233–2241. https://doi.org/10.1002/ijc.23380 (2008).
Hostein, I. et al. Evaluation of MDM2 and CDK4 amplification by real-time PCR on paraffin wax-embedded material: A potential tool for the diagnosis of atypical lipomatous tumours/well-differentiated liposarcomas. J. Pathol. 202, 95–102. https://doi.org/10.1002/path.1495 (2004).
Mejia-Guerrero, S. et al. Characterization of the 12q15 MDM2 and 12q13-14 CDK4 amplicons and clinical correlations in osteosarcoma. Genes Chromosomes Cancer 49, 518–525. https://doi.org/10.1002/gcc.20761 (2010).
Gisselsson, D. et al. Differentially amplified chromosome 12 sequences in low- and high-grade osteosarcoma. Genes Chromosomes Cancer 33, 133–140 (2002).
Ricciotti, R. W. et al. High amplification levels of MDM2 and CDK4 correlate with poor outcome in patients with dedifferentiated liposarcoma: A cytogenomic microarray analysis of 47 cases. Cancer Genet. 218–219, 69–80. https://doi.org/10.1016/j.cancergen.2017.09.005 (2017).
Simon, R. et al. Amplification pattern of 12q13-q15 genes (MDM2, CDK4, GLI) in urinary bladder cancer. Oncogene 21, 2476–2483. https://doi.org/10.1038/sj.onc.1205304 (2002).
An, H. X., Beckmann, M. W., Reifenberger, G., Bender, H. G. & Niederacher, D. Gene amplification and overexpression of CDK4 in sporadic breast carcinomas is associated with high tumor cell proliferation. Am. J. Pathol. 154, 113–118. https://doi.org/10.1016/S0002-9440(10)65257-1 (1999).
Reifenberger, G. et al. Refined mapping of 12q13-q15 amplicons in human malignant gliomas suggests CDK4/SAS and MDM2 as independent amplification targets. Can. Res. 56, 5141–5145 (1996).
Wikman, H. et al. CDK4 is a probable target gene in a novel amplicon at 12q13.3–q14.1 in lung cancer. Genes Chromosomes Cancer 42, 193–199. https://doi.org/10.1002/gcc.20122 (2005).
Muthusamy, V. et al. Amplification of CDK4 and MDM2 in malignant melanoma. Genes Chromosomes Cancer 45, 447–454. https://doi.org/10.1002/gcc.20310 (2006).
Ladenstein, R. et al. Busulfan and melphalan versus carboplatin, etoposide, and melphalan as high-dose chemotherapy for high-risk neuroblastoma (HR-NBL1/SIOPEN): An international, randomised, multi-arm, open-label, phase 3 trial. Lancet Oncol. 18, 500–514. https://doi.org/10.1016/S1470-2045(17)30070-0 (2017).
Dubois, S. G. et al. Lung metastases in neuroblastoma at initial diagnosis: A report from the International Neuroblastoma Risk Group (INRG) project. Pediatr. Blood Cancer 51, 589–592. https://doi.org/10.1002/pbc.21684 (2008).
Bui, C. B. et al. ARID1A-SIN3A drives retinoic acid-induced neuroblastoma differentiation by transcriptional repression of TERT. Mol. Carcinog. https://doi.org/10.1002/mc.23091 (2019).
Nakamura, Y. et al. Functional characterization of a new p53 mutant generated by homozygous deletion in a neuroblastoma cell line. Biochem. Biophys. Res. Commun. 354, 892–898. https://doi.org/10.1016/j.bbrc.2007.01.057 (2007).
Goldschneider, D. et al. Expression of C-terminal deleted p53 isoforms in neuroblastoma. Nucleic Acids Res. 34, 5603–5612. https://doi.org/10.1093/nar/gkl619 (2006).
Mlakar, V. et al. PRIMA-1(MET)-induced neuroblastoma cell death is modulated by p53 and mycn through glutathione level. J. Exp. Clin. Cancer Res. 38, 69. https://doi.org/10.1186/s13046-019-1066-6 (2019).
Drummond, C. J. et al. TP53 mutant MDM2-amplified cell lines selected for resistance to MDM2-p53 binding antagonists retain sensitivity to ionizing radiation. Oncotarget 7, 46203–46218. https://doi.org/10.18632/oncotarget.10073 (2016).
Matsui, A., Ihara, T., Suda, H., Mikami, H. & Semba, K. Gene amplification: Mechanisms and involvement in cancer. Biomol. Concepts 4, 567–582. https://doi.org/10.1515/bmc-2013-0026 (2013).
Caren, H., Abel, F., Kogner, P. & Martinsson, T. High incidence of DNA mutations and gene amplifications of the ALK gene in advanced sporadic neuroblastoma tumours. Biochem. J. 416, 153–159. https://doi.org/10.1042/bj20081834 (2008).
Malumbres, M. & Barbacid, M. To cycle or not to cycle: A critical decision in cancer. Nat. Rev. Cancer 1, 222–231. https://doi.org/10.1038/35106065 (2001).
Hartlieb, S. A. et al. Alternative lengthening of telomeres in childhood neuroblastoma from genome to proteome. Nat. Commun. 12, 1269. https://doi.org/10.1038/s41467-021-21247-8 (2021).
Fransson, S. et al. Estimation of copy number aberrations: Comparison of exome sequencing data with SNP microarrays identifies homozygous deletions of 19q13.2 and iin neuroblastoma. Int. J. Oncol. 48, 1103–1116. https://doi.org/10.3892/ijo.2016.3349 (2016).
Mahamdallie, S. et al. Identification of new Wilms tumour predisposition genes: An exome sequencing study. Lancet. Child Adolesc. Health 3, 322–331. https://doi.org/10.1016/S2352-4642(19)30018-5 (2019).
Moreno, L. et al. Accelerating drug development for neuroblastoma—New drug development strategy: An innovative therapies for children with cancer, European Network for Cancer Research in Children and Adolescents and International Society of Paediatric Oncology Europe Neuroblastoma project. Expert Opin. Drug Discov. 12, 801–811. https://doi.org/10.1080/17460441.2017.1340269 (2017).
Geoerger, B. et al. A phase I study of the CDK4/6 inhibitor ribociclib (LEE011) in pediatric patients with malignant rhabdoid tumors, neuroblastoma, and other solid tumors. Clin. Cancer Res. 23, 2433–2441. https://doi.org/10.1158/1078-0432.CCR-16-2898 (2017).
Laroche-Clary, A. et al. Combined targeting of MDM2 and CDK4 is synergistic in dedifferentiated liposarcomas. J. Hematol. Oncol. 10, 123. https://doi.org/10.1186/s13045-017-0482-3 (2017).
Vilgelm, A. E. et al. MDM2 antagonists overcome intrinsic resistance to CDK4/6 inhibition by inducing p21. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aav7171 (2019).
Portman, N. et al. MDM2 inhibition in combination with endocrine therapy and CDK4/6 inhibition for the treatment of ER-positive breast cancer. Breast Cancer Res. 22, 87. https://doi.org/10.1186/s13058-020-01318-2 (2020).
Sriraman, A., Dickmanns, A., Najafova, Z., Johnsen, S. A. & Dobbelstein, M. CDK4 inhibition diminishes p53 activation by MDM2 antagonists. Cell Death Dis. 9, 918. https://doi.org/10.1038/s41419-018-0968-0 (2018).
Schubert, N. A. et al. Combined targeting of the p53 and pRb pathway in neuroblastoma does not lead to synergistic responses. Eur. J. Cancer 142, 1–9. https://doi.org/10.1016/j.ejca.2020.10.009 (2020).
Tweddle, D. A. et al. The p53 pathway and its inactivation in neuroblastoma. Cancer Lett. 197, 93–98. https://doi.org/10.1016/s0304-3835(03)00088-0 (2003).
Carr-Wilkinson, J. et al. High frequency of p53/MDM2/p14ARF pathway abnormalities in relapsed neuroblastoma. Clin. Cancer Res. 16, 1108–1118. https://doi.org/10.1158/1078-0432.CCR-09-1865 (2010).
He, J. et al. Double amplifications of CDK4 and MDM2 in a gastric inflammatory myofibroblastic tumor mimicking cancer with local invasion of the spleen and diaphragm. Cancer Biol. Ther. https://doi.org/10.1080/15384047.2018.1480290 (2018).
Wu, Q. et al. Breast cancer subtypes predict the preferential site of distant metastases: A SEER based study. Oncotarget 8, 27990–27996. https://doi.org/10.18632/oncotarget.15856 (2017).
Kang, Y. et al. A multigenic program mediating breast cancer metastasis to bone. Cancer Cell 3, 537–549. https://doi.org/10.1016/s1535-6108(03)00132-6 (2003).
Furlan, A. et al. Multipotent peripheral glial cells generate neuroendocrine cells of the adrenal medulla. Science https://doi.org/10.1126/science.aal3753 (2017).
Bedoya-Reina, O. C. et al. Single-nuclei transcriptomes from human adrenal gland reveal distinct cellular identities of low and high-risk neuroblastoma tumors. Nat. Commun. 12, 5309. https://doi.org/10.1038/s41467-021-24870-7 (2021).
Dong, R. et al. Single-cell characterization of malignant phenotypes and developmental trajectories of adrenal neuroblastoma. Cancer Cell 38, 716-733 e716. https://doi.org/10.1016/j.ccell.2020.08.014 (2020).
Hanemaaijer, E. S. et al. Single-cell atlas of developing murine adrenal gland reveals relation of Schwann cell precursor signature to neuroblastoma phenotype. Proc. Natl. Acad. Sci. U. S. A. https://doi.org/10.1073/pnas.2022350118 (2021).
Jansky, S. et al. Single-cell transcriptomic analyses provide insights into the developmental origins of neuroblastoma. Nat. Genet. 53, 683–693. https://doi.org/10.1038/s41588-021-00806-1 (2021).
Kameneva, P. et al. Single-cell transcriptomics of human embryos identifies multiple sympathoblast lineages with potential implications for neuroblastoma origin. Nat. Genet. 53, 694–706. https://doi.org/10.1038/s41588-021-00818-x (2021).
Shimada, H. et al. Terminology and morphologic criteria of neuroblastic tumors: Recommendations by the International Neuroblastoma Pathology Committee. Cancer 86, 349–363 (1999).
Fransson, S. et al. Whole-genome sequencing of recurrent neuroblastoma reveals somatic mutations that affect key players in cancer progression and telomere maintenance. Sci. Rep. 10, 22432. https://doi.org/10.1038/s41598-020-78370-7 (2020).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26. https://doi.org/10.1038/nbt.1754 (2011).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108 (2008).
Kryh, H. et al. MYCN amplicon junctions as tumor-specific targets for minimal residual disease detection in neuroblastoma. Int. J. Oncol. 39, 1063–1071. https://doi.org/10.3892/ijo.2011.1120 (2011).
Umapathy, G. et al. MEK inhibitor trametinib does not prevent the growth of anaplastic lymphoma kinase (ALK)-addicted neuroblastomas. Sci. Signal. https://doi.org/10.1126/scisignal.aam7550 (2017).
Ianevski, A., He, L., Aittokallio, T. & Tang, J. SynergyFinder: A web application for analyzing drug combination dose-response matrix data. Bioinformatics 33, 2413–2415. https://doi.org/10.1093/bioinformatics/btx162 (2017).
Acknowledgements
We would like to posthumously acknowledge Fredrik Hedborg, M.D, Ph.D., of Uppsala University, Uppsala, Sweden for helping initiate this study. We also want to thank Prof. Manfred Schwab, Heidelberg, Germany for providing us with the NB cell lines LS and NGP, and the Genomics and Bioinformatics Core Facility platforms at Sahlgrenska Academy, University of Gothenburg, Gothenburg, Sweden for access to and assistance with instrumentation and analysis. We also thank Vanja Börjesson from the Bioinformatics Core Facility at Gothenburg University for generating the Circos plots. This work was supported by grants from the Swedish Childhood Cancer Foundation (SF—NBCNSPDHEL10/021, NCp2015-0061, TJ2014-0064; TM—2016-147, 2017-166; PK-2017-0122), the Swedish Cancer Foundation (TM—18-1711; PK-19-0566), the Nilsson-Ehle Foundation (HK—29211), the Swedish state under the LUA/ALF agreement (ALFGBG-447171), a project grant from the Laboratory Division of Sahlgrenska University Hospital, the Lions Cancer Fund West, Assar Gabrielssons Foundation, the Sahlgrenska University Hospital Foundation, the Swedish Research Council (TM/SF—2014-3031), and the Swedish Foundation for Strategic Research (TM/PK—RB13-0204) as well as by the EC H2020 project PRIMAGE (#826494), the Vienna Science and Technology Fund WWTF (#LS18-111) and the St. Anna Kinderkrebsforschung.
Funding
Open access funding provided by University of Gothenburg.
Author information
Authors and Affiliations
Contributions
A.M.M., H.K.Ö., J.G., P.K., T.M., and S.F. conceived and designed the experiments as well as the ensuing performance, analysis, and interpretation. A.D., N.J., and T.M. interpreted the SNP microarray data. A.P.B., I.M.A., I.Ø., B.S., R.L., R.N., M.U., S.T.M., P.F.A., L.G.M., G.J., and P.K. helped with patient care, tissue sampling, and clinical information. T.M., S.F., and P.K. formulated strategy and directed the research. A.M.M., H.K.Ö., P.K., S.F., and T.M. drafted the manuscript; all authors edited and approved the final version of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Martinez-Monleon, A., Kryh Öberg, H., Gaarder, J. et al. Amplification of CDK4 and MDM2: a detailed study of a high-risk neuroblastoma subgroup. Sci Rep 12, 12420 (2022). https://doi.org/10.1038/s41598-022-16455-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-16455-1
- Springer Nature Limited