Abstract
Corneal guttae, which are the abnormal growth of extracellular matrix in the corneal endothelium, are observed in specular images as black droplets that occlude the endothelial cells. To estimate the corneal parameters (endothelial cell density [ECD], coefficient of variation [CV], and hexagonality [HEX]), we propose a new deep learning method that includes a novel attention mechanism (named fNLA), which helps to infer the cell edges in the occluded areas. The approach first derives the cell edges, then infers the well-detected cells, and finally employs a postprocessing method to fix mistakes. This results in a binary segmentation from which the corneal parameters are estimated. We analyzed 1203 images (500 contained guttae) obtained with a Topcon SP-1P microscope. To generate the ground truth, we performed manual segmentation in all images. Several networks were evaluated (UNet, ResUNeXt, DenseUNets, UNet++, etc.) and we found that DenseUNets with fNLA provided the lowest error: a mean absolute error of 23.16 [cells/mm\(^{2}\)] in ECD, 1.28 [%] in CV, and 3.13 [%] in HEX. Compared with Topcon’s built-in software, our error was 3–6 times smaller. Overall, our approach handled notably well the cells affected by guttae, detecting cell edges partially occluded by small guttae and discarding large areas covered by extensive guttae.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
Fuchs endothelial dystrophy (FED) is a corneal disease characterized by an increase in thickness of the Descemet’s membrane, a large deposition of extracellular matrix in the corneal endothelium (referred as guttae), and a progressive loss of endothelial cells1. The guttae, which are condensation of collagen growing from the Descemet’s membrane (gutta is the Latin word for ‘droplet’, being guttae the plural form and guttata the adjective form), appear disperse in the early stages of the disease, but as FED progresses, guttae become abundant and visible in the entire endothelium2. This is accompanied by a great loss of endothelial cells that is irreversible due to the lack of cell regeneration. Eventually, the endothelium is not able to support corneal deturgescence, thereby leading to corneal edema, poor visual acuity, and finally the necessity for corneal transplantation to restore vision. FED is the most common cause for the transplantation of the corneal endothelium worldwide3. This disease usually appears in people of 40–60 years of age and it has a slow clinical progression (10–20 years)4. Since many patients with cornea guttata do not progress to FED, it is difficult to assess its incidence and prevalence. Nevertheless, it is estimated that around 4% of the population over 40 years of age have corneal guttae5.
The most common staging system used to describe FED has four levels1,4: in stage 1, guttae appear in the central cornea and they are non-confluent; in stage 2, guttae start to coalesce and spread towards the peripheral cornea, while endothelial cell loss accelerates; in stage 3, stromal edema is present, which might cause the formation of epithelial and subepithelial bullae; and in stage 4, the cornea has become opaque due to chronic edema and visual acuity is critically compromised. Conversely, central guttae can be observed in the elderly without the presence of edema, and the existence of only peripheral guttae is related to a condition called ‘Hassall-Henle bodies’ and does not lead to edema1. An endothelial cell density (ECD) between 400 to 700 cells/mm\(^{2}\) tends to result in corneal decompensation6. Thus, the correct estimation of ECD, along with the other two main endothelial parameters (polymegethism or coefficient of variation in cell size [CV], and pleomorphism or hexagonality [HEX]), is a valuable tool for the diagnosis and monitoring of this dystrophy.
The endothelium can be easily imaged with specular microscopy, a noninvasive, noncontact method that relies on the reflection of light from the interface between the endothelium and aqueous humor7. However, it is crucial to have a smooth, regular endothelial surface to provide good-quality images. In this respect, the guttae push the endothelial cells out of the specular plane and, thus, these appear as ‘black droplets’ in the specular images (Fig. 1). If the guttae are non-confluent and have a relatively small size (similar or less than the average cell size), a human observer could probably infer the cell tessellation in the area. However, manual annotations are very tedious and time-consuming, and there are currently no automatic methods that can perform such inference. In fact, many recent studies have shown that today’s commercial methods provided by the microscope manufacturers to automatically segment the endothelial images depict low reliability, even for healthy corneas8,9,10,11,12.
In recent years, several new approaches to estimate the corneal endothelial parameters have been proposed. Up to 2018, the methods were based on image processing techniques and classic machine learning13,14,15,16,17, but from 2018 onward, a significantly large number of new approaches based on deep learning have been presented18,19,20,21,22,23,24,25,26,27,28,29,30,31,32. Overall, these methods have shown a good performance. However, it is worth noting a few details: (i) some methods (mainly from the pre-deep learning era) evaluated the estimation of the three corneal parameters but the images were from healthy eyes and there was either a manual selection of the region of interest (ROI) or the cells were visible in the whole image14,15,16,17,18,21,23; (ii) some of the first deep learning methods simply evaluated the capacity of neural networks to perform an accurate segmentation, either in healthy19,22 or unhealthy corneas26, without estimating any corneal parameter, which avoids the non-trivial problem of refining the raw segmentation for the purpose of obtaining an accurate biomarker estimation; (iii) other publications only focused in estimating ECD (or the number of cells) in healthy cases29,30 and images with guttae20,27,31, some including a method to select the ROI (although this part is often unclear); and (iv) our previous work is, to the best of our knowledge, the only fully-automatic method to estimate all three parameters in all types of images (heavily blurred24,25 and also with some guttae28). Among the publications dealing with guttae, a quick visual inspection is enough to perceive the inaccurate segmentation around the guttae, where partial cells occluded by the guttae are included as full cells27,31; in contrast, our previous work28 has shown better results but still failed in cases of very advanced disease. Therefore, the accurate segmentation of images in the presence of guttae is still an unsolved problem.
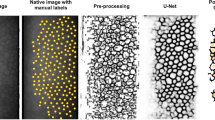
In this study, we present a new approach that automatically segments endothelial images with guttae and estimates the endothelial parameters. The proposed method contains three subprocesses (Fig. 1): (i) a convolutional neural network (CNN) that infers the cell edges in the image (named CNN-Edge, yielding an ‘edge image’); (ii) another CNN model that infers the cells that can be fully identified (named CNN-Body, yielding a [cell] ‘body image’); and (iii) an image processing model based on watershed that refines the edge images, uses the body images to select the well-detected cells, and extracts the corneal parameters (named postprocessing). A comparative analysis is performed to determine what type of deep learning model (UNet33, ResUNeXt34, DenseUNet35, and UNet+/++36) is more appropriate for this task. We also present a new attention mechanism, named feedback non-local attention (fNLA), which can be plugged to any of the aforementioned networks. Different versions of the attention mechanism are tested, and we compare it to a well-know ‘attention UNet’37. This framework is evaluated against our previous approach28, named CNN-ROI (Fig. 2), which is retrained with the dataset of this study. A variation of CNN-Body, named CNN-Blob (Fig. 2), which infers the overall area of interest instead of independent cells, is also evaluated. Finally, these frameworks are compared against the manual annotations and the estimates provided by the microscope’s software.
Flowchart: CNN-Edge detects the cell edges from the specular image, CNN-Body infers the cell bodies that are well detected, and the postprocessing refines the edge images and applies the ROI from the body images to provide the final segmentation. The final segmentation (edges in red, vertices in yellow, non-ROI area in blue) is dilated and superimposed onto the specular image for illustrative purposes.
(a) Illustrative specular image with guttae (grade 4) and (f) its manual annotation (the edges, in yellow, are dilated for illustrative purposes; non-ROI area in blue). For this image, the targets in the different networks are: (b) CNN-Edge, (c) CNN-Body, (d) CNN-Blob, and (e) CNN-ROI. The outcome of the networks during inference (DenseUNet fNLA-Mul) are: (g) CNN-Edge, (h) CNN-Body, (i) CNN-Blob, and (j) CNN-ROI. Red and green arrows indicate two peculiarities of the targets (to be discussed later).
Results
Evaluation metrics
The metrics to evaluate the CNN output images were accuracy, the Sørensen-Dice coefficient (DICE), and modified Hausdorff distance (MHD)38. MHD is a good metric to evaluate the edge images because it measures how close the detected edges are from the manual annotations, whereas DICE can better evaluate the performance for the body images since it measures the overlap of the region of interest (ROI). The evaluation metrics for the corneal parameters were mean absolute error (MAE) and mean absolute percentage error (MAPE). To assess the statistical significance between methods, we used the paired Wilcoxon test to compare the MAPE after assigning a 100% error if no parameter estimate was produced. To assess the clinical statistical significance of our method against the gold standard, we used the Kruskal-Wallis H test. All tests used a statistical significance established at \(\alpha =0.05\).
Comparison between types of CNNs
Quantitatively (Table 1), the basic UNet, ResUNeXt, and DenseUNet seemed to provide similar performance, with slightly better results for DenseUNet (more noticeable if we look at the MAE of the corneal parameters). However, the differences between those networks were discernible in the qualitative results (Fig. 3): in the edge images, UNet and ResUNeXt provided rather binary results and were very conservative in the areas with large guttae (Fig. 3b,c, central area), whereas DenseUNet provided more probabilistic outputs and was able to extend the edge inference into those guttae areas, reaching beyond the manual annotations (Fig. 3d). In the case of the body images, DenseUNet showed a similar probabilistic behavior (Fig. 3), whereas ResUNeXt had significant problems and the lowest performance (Table 1). Thus, DenseUNet was the network to further investigate.
One key point for the success of the edge inference within the guttae areas was the selective training procedure, where some of the images of the batch were sampled from specific guttae subsets. In fact, if the DenseUNets were trained without any stratified subsampling (random selection), such edge inferences did not appear (Fig. 3e). Another interesting observation was that the use of the multiscale networks (+ and ++) had no impact on the edge images but provided a clear improvement in the body images for all types of networks (Table 1), although this did not necessarily result in more accurate parameter estimations.
(a) Illustrative specular image with guttae (grade 5) and its manual annotation (edges in yellow; non-ROI area in blue). For the different networks (indicated on top), the inference of this image for the CNN-Edge (top image) and CNN-Body (bottom image). Topcon’s software did not detect a single cell in this image.
Selection of attention mechanism
In 2018, Oktay et al.37 proposed an attention mechanism in UNets, which modifies the output of a transition block by using the tensors from the lower resolution stage. In our experiments, we tested several variations of this mechanism, but none provided a clear improvement. The best design (Table 1, most like Oktay’s proposal) gave sharper edge images in complicated areas but it occasionally provided poor results for the body images (Fig. 3f).
In this work, we developed a different attention mechanism that would exploit the non-local dependencies between features of different resolution stages while simultaneously bringing back the attention information from the lowest stages back to the highest ones. Quantitatively, the improvement was subtle but perceptible in the basic DenseUNet: the MHD in the edge images was substantially smaller in all cases, and all metrics in the body images improved (Table 1). Qualitatively, they all provided sharper edge images (there were less double-edge artifacts within guttae areas, Fig. 3), and the inference in the body images was more probabilistic (cells within guttae areas appeared with lower intensity instead of just pitch black). Thus, our attention mechanism seemed to correctly use the surrounding information to determine, in a probabilistic manner, whether a cell should be discarded. With regards to the type of aggregation at the end of the attention block, the concatenative and the multiplicative type provided similar results; in contrast, the additive type had the lowest performance in all the experiments. Therefore, we chose the multiplicative type because it uses fewer parameters and provides the overall smallest MAE in the corneal biomarkers (Table 1). On the other hand, the use of fNLA in the multiscale versions +/++ did not yield a clear benefit.
Postprocessing tuning
The postprocessing had two hyperparameters to tune: a threshold to discard weak edges (‘edge threshold’), and a threshold to discard superpixels from the final segmentation (‘body threshold’). We performed a combined grid search and found that, for the edge threshold: (i) a value around 0.1 minimized the error for CV regardless of the body threshold; (ii) for HEX, the optimal value was 0.1 for a body threshold of 0.5 or lower, but it shifted towards 0.2 as we increased the body threshold towards 1; and (iii) for ECD, a higher threshold (0.2–0.3) was better but the differences were very small. Overall, the purpose of the edge threshold is to simply discard false edges created during the postprocessing (watershed). In that respect, a value of 0.1 seemed to be a reasonable choice.
As for the body threshold, we observed that a value of 0.5 (which is the most intuitive choice) yielded the lowest errors in the three corneal parameters for the networks without attention, but it shifted towards 0.75 if we employed networks with fNLA, although the error differences were very small. However, visual inspection showed that no major mistakes were produced with a threshold of 0.5 (Fig. 4, cells with brighter green). We believe that a lower threshold sometimes includes cells that were not annotated in the gold standard, therefore the differences do not indicate segmentation mistakes but mainly register the so-called ‘dissimilarity due to cell variability’ (i.e. the estimated parameters can vary if a different set of cells are used to estimate them).
Finally, the postprocessing computed the hexagonality by using the cell vertices (vertex method) instead of counting the neighboring cells (neighbor method, which inevitably discards the cells in the periphery). For those peripheral cells, it was possible to detect their vertices if a portion of the peripheral edges was visible. This was true for most cases, although sometimes the vertex detection was faulty (Fig. 4IV-e,f). Nevertheless, the MAE in our HEX estimations was 3.14 [%] with the vertex method and 4.13 [%] with the neighbor method. Thus, the vertex method was better even when there were some mistakes in the detection of peripheral vertices.
Eight images with guttae (two cases per grade): (a,b) grade 3, (c,d) grade 4, (e,f) grade 5, (g,h) grade 6. (I) The specular image; the output of the (II) CNN-Edge and (III) CNN-Body for the DenseUNet fNLA-Mul; (IV) the final segmentation (edges in red, vertices in yellow, non-ROI in blue, and the detected cells in two tonalities of green: brighter green for the cells whose body-intensity was between 0.50–0.75, and darker green for the cells whose body-intensity was between 0.75–1.00), and (V) the gold standard (the annotated edges in yellow; non-ROI in blue). On the right, the estimated parameters (gold standard values in parenthesis). Topcon’s software detected 45, 34, 0, 38, 6, 0, 0, and 0 cells (a–h).
Comparison between frameworks
The performance of Topcon’s algorithm was considerably inferior (Table 2), particularly for the images with guttae, where it failed to detect any cell in 30% of the images (as in Fig. 4c,f,g,h) and it only detected one third of the cells (on average) in the remaining images.
Regarding the previous approach CNN-ROI28, the network had no problems to accurately infer the targets (Fig. 2j). However, the question was whether such targets are optimal to subsequently identify the well-detected cells. The error analysis indicated that the CNN-ROI method detected approximately 10–12 more cells per image, but there were a few segmentation mistakes among those cells, and the estimation errors were significantly worse than the CNN-Body approach (Table 2). The paired Wilcoxon test indicated a statistically significant difference between the estimates of approaches CNN-Body and CNN-ROI (\(P<0.0001\), all biomarkers).
As for the CNN-Blob, it detected the same number of cells as CNN-Body (Table 2) and the estimation errors were virtually the same (slightly worse for CNN-Blob in the images with guttae; Table 2). The paired Wilcoxon test indicated no statistically significant difference between the two estimates (\(P=0.69\), \(P=0.13\), and \(P=0.69\), for the ECD, CV, and HEX, respectively). Nevertheless, it is easier for a human observer to interpret what cells are well detected in the CNN-Body output (Fig. 2h) than in the CNN-Blob output (Fig. 2i).
Statistical analysis
The distributions of the estimated parameters from the CNN-Body method and the gold standard passed the Levene’s test for homogeneity, but they did not pass the Shapiro-Wilk normality test. Thus, the Kruskal-Wallis H test was performed and it showed no statistically significant difference between the manual and our automatic assessments for ECD (\(P=0.81\)) and CV (\(P=0.74\)), but it did for HEX (\(P<0.001\)). To further assess the estimates, we performed a Bland-Altman analysis, and it showed that more than 95% of the estimates were within the 95% limit of agreement for all parameters: 96.5% for ECD, 96.6% for CV, and 96.7% for HEX.
Error analysis
We plotted the error of the corneal parameters as a function of the number of cells (Fig. 5) and we fitted two exponentials to the mean and SD of the error using the least-squares method. The error estimates showed a normal distribution along the y-axis for the three parameters, which allowed us to assume that the area within two SDs covered approximately the 95% of the error. Images with and without guttae where plotted with different colors Fig. 5. This evaluation showed that, for images with high number of cells, the error is very low regardless of the presence of guttae; in fact, guttae is a critical factor only for images with very low number of cells. Overall, we can conclude that (i) the number of cells was the main variable to predict the reliability of the estimation, with a clear decrease in the error spread as more cells were detected; (ii) the unreliable cases were images with guttae and less than 25 cells; (iii) HEX required more cells to reduce the error spread, and (iv) there was a notable underestimation in HEX for the images with less than 100 cells.
Error of our estimates of (a) ECD, (b) CV, and (c) HEX for the subsets of images without guttae (black dots, 702 images) and with guttae (colored circles, 483 images), displayed as a function of the number of cells. The y-axis indicates the error computed as the difference between the estimates and the gold standard. The mean (solid line) and two SDs (dashed lines) of the error function were modeled with exponentials.
Average value of the biomarkers from the manual annotations—(a) number of cells, (b) ECD, (c) CV, and (d) HEX—and the standard deviation of the estimated error for the different subsets of graded guttae images. The error SD allows to visualize how close the estimations are to the gold standard values.
Clinical analysis
The number of cells necessary to estimate ECD with high reliability is widely accepted as 75 cells39. At that point, our estimated ECD error was approximately 0 ± 35 [cells/mm\(^2\)] (mean ± SD). At 25 cells, it was 0 ± 45 [cells/mm\(^2\)]. This is expected to be much lower than the uncertainty generated by the “cell variability”. Doughty et al.39 evaluated that uncertainty (in manual annotations) and concluded that estimating ECD using 75 cells entails to assume an uncertainty (of 1 SD) of ±2% (or ±70 cells/mm\(^2\), which is twice than our method’s error), whereas the uncertainty at 25 cells increases to ±10% (or ±350 cells/mm\(^2\), more than seven times larger than our method’s error). Nevertheless, both elements (uncertainty and error) should be taken into consideration when evaluating the reliability of the estimations. Regarding CV and HEX, there are no studies in the literature about the effect of the cell variability to allow us to make a comparison.
We also observed that the number of visible cells and ECD decrease acutely as the amount of guttae increases (Fig. 6), whereas neither CV nor HEX showed a significant change. While it is generally accepted that a CV of less than 30% and a HEX of greater than 60% is usually a sign of a healthy endothelium28, this assumption seems to be untrue for the current dataset. Thus, our results seem to suggest that guttae does not have an impact on neither the cell size variation nor the hexagonality of the visible cells (those that are not actually affected by extensive guttae).
Overall, our proposed method has a very low error spread for the three biomarkers in images graded with mild or moderate guttae (Fig. 6, up to grade 4). For the cases with severe guttae (grade 5–6), the main problem is the low number of detected cells (in grade 6, the average number of cells per image was 13 ± 10), which translates into a large estimation uncertainty, particularly for CV and HEX (Fig. 6). As shown in Fig. 4g,h, we did not observe major segmentation mistakes in those cases, but the low number of visible cells makes the biomarker estimation unreliable.
Discussion
We have presented a fully automatic method for the estimation of the corneal endothelium parameters from specular microscopy images that contain guttae. This is the first time that such images have been solved properly in the literature.
The main factor to achieve such accuracy was the way annotations were performed. We observed that, if the edges hidden by the guttae could be manually delineated, the networks could learn to replicate such pattern. Since endothelial cell edges usually appear as straight lines, the extension of partially occluded edges is sufficient to infer the hidden tessellation if the guttae are small enough. In fact, we observed that our model went beyond what our annotator could do. For example, the guttae indicated by the green arrow in Fig. 2 was too large to be certain about the hidden edges, yet the network provided a segmentation that seemed highly probable. However, further experiments would be needed to assert the trustworthiness of such inference; as a suggestion, the artificial generation of black spots in the images (mimicking the guttae’s visuals) could be a way to evaluate such hypothesis, although that would raise questions regarding the experiment’s reliability.
Our proposed attention mechanism (fNLA) moderately improved the performance in both types of CNNs (CNN-Edge and CNN-Body). Since the basic DenseUNet already achieved an excellent performance, the improvement provided by our attention block was modest. Nevertheless, we believe that the feedback-attention path depicted in our proposed network could yield a good performance in other types of segmentation problems, particularly the ones that infer areas instead of edges.
One interesting behavior of this framework is that, while CNN-Edge provides good inference beyond the manual annotations, CNN-Body is rather conservative. This is because the targets of CNN-Body were based on the original annotations and, thus, any inference by CNN-Edge that surpasses the manual annotations are not considered in the target of CNN-Body. While this might seem a negative quality, the results show that our framework detects practically the same number of cells as the manual annotations and barely any segmentation mistake is observed even in the most difficult cases (Fig. 4). Therefore, we believe this approach is preferred to more daring ones, like CNN-ROI (Table 2).
Overall, our estimates agreed very well with the gold standard and they were significantly better than the ones provided by the instrument’s software, demonstrating the ability of this artificial intelligence framework to accurately estimate the endothelial parameters from images with guttae.
Materials and methods
Datasets
Two datasets were employed in this work. The first dataset came from a clinical study concerning the implantation of a Baerveldt glaucoma implant in the Rotterdam Eye Hospital (Rotterdam, the Netherlands). The clinical study contained 7975 images from 204 patients (average age 66±10 years), who were imaged with a specular microscope (Topcon SP-1P, Topcon Co., Tokyo, Japan) before the device implantation and at 3, 6, 12, and 24 months after, in both the central (CE) and the peripheral supratemporal (ST) cornea. The protocol required five specular images to be taken in each area for each visit, although it was sometimes difficult to reach that number of gradable images (specifically, an average of 4.7 CE images and 3.6 ST images per visit were acquired). Retrospectively, we observed that 15 patients had clear signs of FED stage two: they all had guttae in both CE and ST cornea, with a larger amount in CE and a clear increase in ST during the two-year follow-up. From this subset of patients, 193 images were collected. Furthermore, we observed that another 81 patients had small amounts of non-confluent guttae in the CE cornea (either FED stage one or due to normal aging), and 227 images were collected from these cases. In total, 420 images with presence of guttae were selected from this study. In addition, 400 images from other patients in the study without signs of guttae were selected to build a balanced dataset.
The second dataset came from another clinical study in the Rotterdam Eye Hospital regarding the transplantation of the cornea (ultrathin Descemet Stripping Automated Endothelial Keratoplasty, UT-DSAEK). This dataset contained 383 images of the central cornea from 41 eyes (41 patients, average age 73±7 years) and they were acquired at 1, 3, 6, and 12 months after surgery with the same specular microscope Topcon SP-1P. The included population for the study were patients over 18 years old with FED planned for keratoplasty. Among these patients, FED reappeared in one of them. Another 13 patients showed a small amount of non-confluent, stable guttae during the one-year follow-up. In total, 80 images out of the 383 showed some guttae (all images were included in the present work).
Altogether, the combined dataset contained 1203 images, in which 500 images depicted guttae in various magnitudes. The images covered an area of approximately 0.25 mm \(\times\) 0.55 mm and were saved as 8-bit grayscale images of 240 \(\times\) 528 pixels. All images were manually segmented to create the gold standard, using the open-source image manipulation program GIMP (version 2.10). Furthermore, we collected the endothelial parameters provided by the microscope (Topcon SP-1P performed this with the software IMAGEnet i-base, version 1.32).
Ethics approval and consent to participate
Data was collected in accordance with the principles of the Declaration of Helsinki (October, 2013). Signed informed consent was obtained from all subjects. Participants gave consent to publish the data. Approval was obtained from the Medical Ethical Committee of the Erasmus Medical Center, Rotterdam, The Netherlands (MEC-2014-573). Trial registration for the Baerveldt study: NTR4946, registered 06/01/2015, URL https://www.trialregister.nl/trial/4823. Trial registration for the UTDSAEK study: NTR4945, registered on 15-12-2014, URL https://www.trialregister.nl/trial/4805.
Grading the dataset
The 500 images with guttae were graded based on their complexity to segment them, taking two metrics into account: the amount of guttae and blur present in the image. For both metrics, images were given a value of 1 (mild), 2 (moderate), or 3 (severe), the final grade being the sum of both values. As a result, there were 134 images with low complexity (grades 1–2), 235 images with medium complexity (grades 3–4), and 131 images with high complexity (grades 5–6).
Targets and frameworks
CNN-Edge is the core of the method. If the specular image had good quality (high contrast) and with cells visible in the whole image, the resulting edge image would probably be so well inferred that a simple thresholding and skeletonization would suffice to obtain the binary segmentation. However, these issues (low contrast, blurred areas, and guttae) are present in the current images. In contrast, CNN-Body has the goal of providing a ROI image to discard areas masked by extensive guttae or blurriness.
These CNNs are trained independently and they all have the same design; thus, they are simply trained with different inputs and targets. To create the targets, we make use of the manual annotations (i.e. gold standard), which are binary images where value 1 indicates a cell edge (edges are 8-connected-pixel lines of 1 pixel width), value 0 represents a full cell body, and any area to discard (including partial cells) is given a value 0.5. If a blurred or guttae area is so small that the cell edges could be inferred by observing the surroundings, the edges are annotated instead (Fig. 2f). For all the annotated cells, we identify their vertices; this allows computing the parameter hexagonality from all cells and not only the inner cells (in the latter, HEX is computed by counting the neighboring cells, thus the cells in the periphery of the segmentation are not considered; this way of computing HEX was used in previous publications23,24,25,28,40 and it is how Topcon’s built-in software computes it). Therefore, HEX is now defined as the percentage of cells that have six vertices.
The target of the CNN-Edge only contains the cell edges from the gold standard images, which have been convolved with a 7 \(\times\) 7 isotropic unnormalized Gaussian filter of standard deviation (SD) 1 (Fig. 2b). This provides a continuous target with thicker edges, which proved to deliver better results than binary targets23.
The target of the CNN-Body only contains the full cell bodies from the gold standard images, and partial cells are discarded either because they are partially occluded by large guttae or they are at the border of the image (Fig. 2c). The same probabilistic transformation is applied here. Alternatively, we also evaluated whether a target that also includes the edges between the cell bodies (Fig. 2d) was a better approach (named CNN-Blob).
This framework is similar to our previous approach28, where a model named CNN-ROI, whose input is the edge image, provides a binary map indicating the ROI (Fig. 2j). To create its target, the annotator would simply draw the rough area from the edge images (Fig. 2g) that they would choose as trustworthy, creating a binary target (Fig. 2e).
Backbone of the network
The proposed CNN has five resolution stages (Fig. 7a). We tested three designs depending on the connections of the convolutional layers within each node: consecutively (as in UNet33), with residual connections (ResUNeXt34), and with dense connections (DenseUNet35). In addition, we also explored two multiscale designs (commonly refered as + [plus] and ++ [plusplus]36) in the aforementioned networks: UNet+ (Fig. 7d), UNet++, ResNeXt+, ResNeXt++, DenseUnet+, and DenseUnet++. Our ++ designs differentiates from + ones in that the former use feature addition from all previous transition blocks of the same resolution stage. In total, nine basic networks were tested (the code for all cases can be found in our GitHub).
A schematic representation of the (a) UNet backbone and (d) its equivalent UNet+; a simplified representation of adding fNLA/sNLA blocks to the (b) UNet and (e) UNet+, where a red arrow indicates self-attention (sNLA) if it returns to the same node or feedback attention (fNLA) otherwise; (c) a schematic overview of how the fNLA/sNLA blocks are added to the UNet backbone for the three deepest resolution scales; and (f) a detailed description of an fNLA block with multiplicative aggregation: the feature maps are shown as the shape of their tensors, \(X^{a,b}\) is the tensor to be transformed, \(Y^{a+1,b}\) is the tensor from the lower resolution scale used for attention, the blue boxes (\(\phi\) and g) indicate a 2 \(\times\) 2 transpose convolution with strides 2, the red boxes (\(\theta\) and \(\Omega\)) indicate a 1 \(\times\) 1 convolution, \(\oplus\) denotes matrix multiplication, and \(\odot\) denotes element-wise multiplication. In the case of sNLA, \(\phi\) and g are also a 1 \(\times\) 1 convolution with \(X^{a,b}\) as input. In attention terminology44, Q is for query, K for key, V for value.
Here, we briefly describe the network that provided the best performance: the DenseUNet. In this network, the dense nodes have 4, 8, 12, 16, or 20 convolutional blocks, depending on the resolution stage, with a growth rate (GR) of 5 feature maps per convolutional layer. Each convolutional block has a compression layer, Conv(1 \(\times\) 1, 4\(\cdot\)GR maps) + BRN41 + ELU42, and a growth layer, Conv(3 \(\times\) 3, GR maps) + BRN + ELU, followed by Dropout(20%) + Concatenation (with block’s input), except the first dense block, which lacks the compression layer and dropout43 (all nodes in the first resolution stage lack dropout). The transition layer (short-connections) has Conv(1 \(\times\) 1, \(\alpha\) maps) + BRN + ELU, the downsampling layer has Conv(2 \(\times\) 2, strides 2, \(\alpha\) maps) + BRN + ELU, and the upsampling layer has ConvTranspose(2 \(\times\) 2, strides 2, \(\alpha\) maps) + BRN + ELU, being \(\alpha\) = (number of blocks in previous dense block) \(\times\) GR/2. The output of the last dense block, \(X^{04}\), uses a transition layer with 2 maps to provide the output of the network.
The attention mechanism
The core of the attention block (Fig. 7f) is inspired by Wang et al.’s non-local attention method44, which also resembles a scaled dot-product attention block45,46. Wang et al.’s design, which is employed in the deepest stages of a classification network, proposes a self-attention mechanism where (among other differences) Q, K, and V in Fig. 7f are the same tensor. In our case, the attention block is added at the end of each dense block and it is named feedback non-local attention (fNLA, Fig. 7f) or self-non-local attention (sNLA), depending where it is placed within the network (Fig. 7c): if there exists a tensor from a lower resolution stage, the attention mechanism makes use of it (fNLA), but in the absence of such lower tensor, a self-attention operation is performed (sNLA, where \(\phi\) and g in Fig. 7f become a 1 \(\times\) 1 convolution with the same input \(X^{a,b}\)). In the DenseUNet, the nodes of the encoder use fNLA (except the deepest node), and the nodes of the decoder use sNLA (Fig. 7b); in the DenseUNet+/++, only the largest decoder use sNLA (Fig. 7e).
We also explored three types of attention mechanism depending on the type of aggregation at the end of the block: (a) the default case (Fig. 7f) uses multiplicative aggregation, where a single attention map (from \(\Omega\)) is sigmoid activated and then element-wise multiplied to all input feature maps; (b) in the case of concatenative aggregation, \(\Omega\) is simply an ELU activation, whose output maps (C/8 maps) are concatenated to the input features; (c) in case of additive aggregation, \(\Omega\) becomes a 1 \(\times\) 1 convolution with ReLU activation and C output feature maps, which are then summed to the input tensor. For comparison, Wang et al.’s model44 is an sNLA with additive aggregation.
Intuitively, an sNLA mechanism computes the response at a specific point in the feature maps as the weighted sum of the features at all positions. In fNLA, the attention block maps the input tensor against the output features from the lower dense block, thus allowing the attention mechanism to use information created further ahead in the network. Moreover, the feedback path created in the encoder allows to propagate the attention features from the deepest dense block back to the first block. While endothelial specular images do not possess such long-range dependencies (features separated by a distance of 3–4 cells do not seem to be correlated), this attention operation might be useful in the presence of large blurred areas (such as guttae). In this work, we explored the use of this attention mechanism for DenseUNet and the multiscale versions DenseUNet+/++.
Description of the postprocessing
The postprocessing aims to fix any edge discontinuity. Here, we have improved a process that was first described in a previous publication23. The steps are:
(I) We estimate the average cell size (l) by using Fourier analysis in the edge image23. As we proved in previous work25, this estimation is simple, extremely robust, and accurate.
(II) As new step, we add a perimeter to the edge image with intensity 0.5. This closes any partial cell in touch with the border.
(III) We smooth the edge image with a Gaussian filter whose standard deviation is \(\sigma =k_{\sigma }l\), being \(k_{\sigma }=0.2\) (this parameter was derived in a previous publication40). This fixes any edge discontinuity.
(IV) We apply classic watershed47 to the smoothed edge image, which detects weak edges. This provides a binary segmentation where edges are 8-connected-pixel lines of 1 pixel width.
(V) In another new step, we identify every edge and vertex in the segmentation. The vertices are the branch points of the segmentation, and the edges are the set of 8-connected positive pixels whose endpoints are constrained to vertices17. We set 2 pixels as the minimum length for an edge (edges of only 1 pixel are fused with the vertices at its endpoints to become a single vertex of 3 pixels). For every edge, we check its mean intensity (from the edge images) to discard the weak ones: if it is lower than 0.1 (threshold evaluated in the “Results” section), the edge is discarded. However, we make a distinction when removing edges: if the edge is internal, all pixels of the edge are completely removed and, thus, the vertex-pixels at the end of that edge become edge-pixels; in contrast, if the edge is in contact with the non-ROI area, we only remove a few pixels in the middle of the edge in order to preserve the vertices. This is relevant because we use the vertices to determine the HEX.
(VI) In the updated binary segmentation, every superpixel in contact with the border of the image is discarded. For the remaining superpixels, we checked the Body/ROI image to determine whether to keep or discard them. If using the body/blob image, a superpixel is included if the average intensity of their pixels is above 0.5. If using the ROI image, a superpixel is included if at least 85% of its area is within the ROI.
(VII) In the final segmentation, the parameters are estimated.
Implementation
To evaluate the networks, a ten-fold cross-validation was performed (with all images from one eye in the same fold). All networks were implemented in Tensorflow 2.4.1 on a single NVIDIA V100 GPU with 32GB of memory. A training batch size of 15 images was employed, where six images were sampled from a specific guttae subgroup (two images from each complexity level). The dimensions of the DenseUNet were selected such that it could fit within the GPU memory; to build the multiscale DenseUNet+/++, we reduced the GR to 4 so that they could still fit in memory while having a similar number of parameters. In that respect, ResUNeXt+/++ and UNet+/++ had one less convolutional block in each resolution stage than the ResUNeXt and UNet (except the lowest stage).
The network hyperparameters were categorical cross-entropy as loss function, Nadam optimizer48, initial learning rate (lr) of 0.001, no early stop, 200 epochs for CNN-Edge (learning decay of 0.99, considering that \(lr_{epoch-i}=lr \cdot lr_{decay}^i\)) and 100 epochs for CNN-Body, CNN-Blob, and CNN-ROI (learning decay of 0.97). For data augmentation, images were randomly flipped left-right and up-down, and elastic deformations were employed. The specular images were only normalized to have a range between 0–1 instead of 0–255. The networks were programmed in Python 3.7, and the parameter estimation and statistical analyses were done in Matlab 2020a (MathWorks, Natick, MA).
Data availability
The datasets used during the current study are available from the corresponding author on reasonable request.
Code availability
Code for the networks and their weights are available in our GitHub repository at https://github.com/jpviguerasguillen/feedback-non-local-attention-fNLA.
References
Elhalis, H., Azizi, B. & Jurkunas, U. V. Fuchs endothelial corneal dystrophy. Ocul. Surf. 8(4), 173–184 (2010).
McLaren, J. W., Bachman, L. A., Kane, K. M. & Patel, S. V. Objective assessment of the corneal endothelium in Fuchs’ endothelial dystrophy. Investig. Ophthalmol. Vis. Sci. 55(2), 1184–1190 (2014).
Gain, P. et al. Global survey of corneal transplantation and eye banking. JAMA Ophthalmol. 134(2), 167–176 (2016).
Adamis, A. P., Filatov, V., Tripathi, B. J. & Tripathi, R. C. Fuchs’ endothelial dystrophy of the cornea. Surv. Ophthalmol. 38(2), 149–168 (1993).
Moshirfar, M., Somani, A. N., Vaidyanathan, U. & Patel, B. C. Fuchs Endothelial Dystrophy. https://www.ncbi.nlm.nih.gov/books/NBK545248/ (StatPearls Publishing, 2021).
Foster, C. S., Azar, D. T. & Dohlman, C. H Smolin and Thoft’s the Cornea: Scientific Foundations & Clinical Practice 4th edn, 46–48 (Lippincott Williams & Wilkins, 2004).
McCarey, B. E., Edelhauser, H. F. & Lynn, M. J. Review of corneal endothelial specular microscopy for FDA clinical trials of refractive procedures, surgical devices, and new intraocular drugs and solutions. Cornea 27(1), 1–16 (2008).
Huang, J. et al. Comparison of noncontact specular and confocal microscopy for evaluation of corneal endothelium. Eye Contact Lens 44, S144–S150 (2018).
Price, M. O., Fairchild, K. M. & Price, F. W. Jr. Comparison of manual and automated endothelial cell density analysis in normal eyes and DSEK eyes. Cornea 23(5), 567–873 (2013).
Luft, N., Hirnschall, N., Schuschitz, S., Draschl, P. & Findl, O. Comparison of 4 specular microscopes in healthy eyes and eyes with cornea guttata or corneal grafts. Cornea 34(4), 381–386 (2015).
Gasser, L., Reinhard, T. & Böhringer, D. Comparison of corneal endothelial cell measurements by two non-contact specular microscopes. BMC Ophthalmol. 15, 87 (2015).
Kitzmann, A. S. et al. Comparison of corneal endothelial cell images using a noncontact specular microscope and the confoscan 3 confocal microscope. Investig. Ophthalmol. Vis. Sci. 24(8), 980–984 (2004).
Piórkowski, A. & Gronkowska-Serafin, J. Towards precise segmentation of corneal endothelial cells. In IWBBIO 2015, LNCS, Vol. 9043, 240–249 (Granada, Spain, 2015).
Selig, B., Vermeer, K. A., Rieger, B., Hillenaar, T. & Luengo Hendriks, C. L. Fully automatic evaluation of the corneal endothelium from in vivo confocal microscopy. BMC Med. Imaging 15, 13 (2015).
Scarpa, F. & Ruggeri, A. Development of a reliable automated algorithm for the morphometric analysis of human corneal endothelium. Cornea 35(9), 1222–1228 (2016).
Al-Fahdawi, S. et al. A fully automated cell segmentation and morphometric parameter system for quantifying corneal endothelial cell morphology. Comput. Methods Programs Biomed. 160, 11–23 (2018).
Vigueras-Guillén, J. P. et al. Corneal endothelial cell segmentation by classifier-driven merging of oversegmented images. IEEE Trans. Med. Imaging 37(10), 2278–2289 (2018).
Fabijańska, A. Segmentation of corneal endothelium images using a U-Net-based convolutional neural network. Artif. Intell. Med. 88, 1–13 (2018).
Nurzynska, K. Deep learning as a tool for automatic segmentation of corneal endothelium images. Symmetry 10(3), 60 (2018).
Daniel, M. C. et al. Automated segmentation of the corneal endothelium in a large set of ‘real-world’ specular microscopy images using the U-net architecture. Nat. Sci. Rep. 9, 4752 (2019).
Fabijańska, A. Automatic segmentation of corneal endothelial cells from microscopy images. Biomed. Signal Process. Control 47, 145–148 (2019).
Kolluru, C. et al. Machine learning for segmenting cells in corneal endothelium images. In Proceedings of SPIE, Vol. 10950, 109504G (San Diego, CA, USA, 2019).
Vigueras-Guillén, J. P. et al. Fully convolutional architecture vs sliding-window CNN for corneal endothelium cell segmentation. BMC Biomed. Eng. 1, 4 (2019).
Vigueras-Guillén, J. P., Lemij, H. G., van Rooij, J., Vermeer, K. A. & van Vliet, L. J. Automatic detection of the region of interest in corneal endothelium images using dense convolutional neural networks. In Proceedings o SPIE, Medical Imaging, Vol. 10949, p. 1094931 (San Diego, CA, USA, 2019).
Vigueras-Guillén, J. P., van Rooij, J., Lemij, H. G., Vermeer, K. A. & van Vliet, L. J. Convolutional neural network-based regression for biomarker estimation in corneal endothelium microscopy images. In 41st Conference Proceedings of IEEE Engineering in Medicine and Biology Society (EMBC), 876–881 (Berlin, Germany, 2019).
Joseph, N. et al. Quantitative and qualitative evaluation of deep learning automatic segmentations of corneal endothelial cell images of reduced image quality obtained following cornea transplant. J. Med. Imaging 7(1), 014503 (2020).
Sierra, J. S., et al. Automated corneal endothelium image segmentation in the presence of cornea guttata via convolutional neural networks. In Proceedings of SPIE, Applications of Machine Learning, vol. 11511, 115110H, online (2020).
Vigueras-Guillén, J. P. et al. Deep learning for assessing the corneal endothelium from specular microscopy images up to 1 year after ultrathin-DSAEK surgery. Transl. Vis. Sci. Technol. 9(2), 49 (2020).
Karmakar, R., Nooshabadi, S. & Eghrari, A. An automatic approach for cell detection and segmentation of corneal endothelium in specular microscope. Graefes Arch. Clin. Exp. Ophthalmol. 260, 1215–1224 (2021).
Kucharski, A. & Fabijańska, A. CNN-watershed: A watershed transform with predicted markers for corneal endothelium image segmentation. Biomed. Signal Process. Control 68, 102805 (2021).
Shilpashree, P. S., Suresh, K. V., Sudhir, R. R. & Srinivas, S. P. Automated image segmentation of the corneal endothelium in patients with Fuchs dystrophy. Transl. Vis. Sci. Technol. 10(13), 27 (2021).
Herrera-Pereda, R., Taboada Crispi, A., Babin, D., Philips, W. & Holsbach Costa, M. A review on digital image processing techniques for in-vivo confocal images of the cornea. Med. Image Anal. 73, 102188 (2021).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: Convolutional networks for biomedical image segmentation. In: Medical Image Computing and Computer-Assisted Intervention (MICCAI), LNCS, Vol. 9351, 234–241 (Munich, Germany, 2015).
Xie, S., Girshick, R., Dollár, P., Tu, Z. & He, K. Aggregated residual transformations for deep neural networks. In IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 5987–5995 (Honolulu, HI, USA, 2017).
Jégou, S., Drozdzal, M., Vazquez, D., Romero, A. & Bengio, Y. The one hundred layers tiramisu: Fully convolutional DenseNets for semantic segmentation. In IEEE Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), 1175–1183 (Honolulu, HI, USA, 2017).
Zhou, Z., Siddiquee, M. M. R., Tajbakhsh, N. & Liang, J. UNet++: Redesigning skip connections to exploit multiscale features in image segmentation. IEEE Trans. Med. Imaging 39(6), 1856–1867 (2020).
Oktay, O. et al. Attention U-Net: Learning where to look for the pancreas. In 1st Conference on Medical Imaging Deep Learning (MIDL) (Amsterdam, The Netherlands, 2018).
Dubuisson, M.-P. & Jain, A. K. A modified Hausdorff distance for object matching. In Proceedings of 12th International Conference on Pattern Recognition (ICPR) 566–568 (Jerusalem, Israel, 1994).
Doughty, M. J., Müller, A. & Zaman, M. L. Assessment of the reliability of human corneal endothelial cell-density estimates using a noncontact specular microscope. Cornea 19(2), 148–158 (2000).
Vigueras-Guillén, J. P. et al. Improved accuracy and robustness of a corneal endothelial cell segmentation method based on merging superpixels. In 15th International Conference Image Analysis and Recognition (ICIAR), LNCS, Vol. 10882, 631–638 (Póvoa de Varzim, Portugal, 2018).
Ioffe, S. Batch renormalization: Towards reducing minibatch dependence in batch-normalized models. In Proceedings of 31st Conference on Neural Information Processing Systems (NeurIPS) (Long Beach, NY, USA, 2017).
Clevert, D.-A., Unterthiner, T. & Hochreiter, S. Fast and accurate deep network learning by exponential linear units (ELUs). In International Conference on Learning Representations (ICLR) (San Juan, Puerto Rico, 2016).
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: A simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Wang, X., Girshick, R., Gupta, A. & He, K. Non-local neural networks. In IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 7794–7803 (Salt Lake City, UT, USA, 2018).
Vaswani, A., et al. Attention is all you need. In Proceedings of 31st Conference on Neural Information Processing Systems (NeurIPS) (Long Beach, NY, USA, 2017).
Srinivas, A. et al. Bottleneck transformers for visual recognition. In IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 16514–16524 (Nashville, TN, USA, 2021).
Beucher, S. & Meyer, F. The morphological approach to segmentation: The watershed transformation. In Mathematical Morphology in Image Processing, 1st edn, 433–481 (Marcel Dekker Inc., 1993).
Dozat, T. Incorporating Nesterov momentum into Adam. In International Conference on Learning Representations (ICLR) Workshop 2013–2016 (San Juan, Puerto Rico, 2016).
Acknowledgements
This work was supported by the Dutch Organization for Health Research and Healthcare Innovation (ZonMw, The Hague, The Netherlands) under Grants 842005004 and 842005007, and by the Combined Ophthalmic Research Rotterdam (CORR, Rotterdam, The Netherlands) under grant no. 2.1.0. The authors thank Angela Engel, Caroline Jordaan, and Annemiek Krijnen for their contribution in acquiring the images.
Author information
Authors and Affiliations
Contributions
All authors contributed to the present study. J.P.V.G., B.T.H.D., J.R., and K.A.V. conceived the idea. J.R. and H.G.L. designed and conducted the clinical studies. E.I. coordinated the acquisition of the data for the Baerveldt study. J.P.V.G. wrote the manuscript and performed the statistical analyses guided by L.J.V. and K.A.V. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vigueras-Guillén, J.P., van Rooij, J., van Dooren, B.T.H. et al. DenseUNets with feedback non-local attention for the segmentation of specular microscopy images of the corneal endothelium with guttae. Sci Rep 12, 14035 (2022). https://doi.org/10.1038/s41598-022-18180-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-18180-1
- Springer Nature Limited