Abstract
Rice blast is a detrimental rice disease caused by the fungus Magnaporthe oryzae. Here, we identified a resistance gene from the rice cultivar Fuhui 2663 which is resistant to the rice blast isolate KJ201. Through isolated population analyses and sequencing approaches, the candidate gene was traced to chromosome 12. With the use of a map-based cloning strategy, the resistance gene was ultimately mapped to an 80-kb resistance locus region containing the Pita gene. Candidate gene prediction and cDNA sequencing indicated that the target resistance gene in Fuhui 2663 was allelic to Pita, thus being referred to as Pita-Fuhui2663 hereafter. Further analysis showed that the Fuhui 2663 protein had one amino acid change: Ala (A) residue 918 in Pita-Fuhui2663 was replaced by Ser (S) in Pita-S, leading to a significant change in the 3D structure of the Pita-S protein. CRISPR/Cas9 knockout experiments confirmed that Pita-Fuhui2663 is responsible for the resistance phenotype of Fuhui 2663. Importantly, Pita-Fuhui2663 did not affect the main agronomic traits of the variety compared to the Pita gene as verified by knockout experiments, indicative of potential applications of Pita-Fuhui2663 in broader breeding programs. Furthermore, a Pita-Fuhui2663-dCAPS molecular marker with good specificity and high efficiency was developed to facilitate rice breeding for resistance to this devastating disease.
Similar content being viewed by others
Introduction
Rice (Oryza sativa L.) is one of the most important food crops worldwide. Rice blast, a fungal disease caused by the filamentous fungus Magnaporthe oryzae (synonym Pyricularia oryzae), is one of the most damaging diseases in rice production, leading to a significant yield reduction and considerably affecting the quality of rice1,2. The cloning and functional analysis of rice blast resistance genes and the study of M. oryzae effector proteins to explore the molecular mechanism of the interactions between rice and M. oryzae will increase the disease resistance of rice3. At present, more than 100 blast resistance genes have been identified, of which 37 have been cloned4. Most of these genes are dominant, and most of them, such as Pita, Pi-1, Pi25, Pigm, and Pia, encode resistance proteins. They can specifically recognize effectors and trigger a series of defense responses to inhibit the growth of pathogen, known as effector-triggered immunity (ETI)1,5,6,7,8. However, there are also disease-resistance genes that do not encode NBS-LRR. For example, Pid2, located on chromosome 6 and encoding a receptor-like kinase, is a constitutively expressed single-copy dominant gene9. Pi21 encodes a protein with five proline-rich regions, and its resistance properties rely on the inactivation of protein function10. Ptr encodes an atypical broad-spectrum resistance protein containing four Armadillo repeat regions and may be a novel E3 ligase11.
A cluster of blast R genes on chromosome 12 of rice, including Pita, Pita2, and Ptr, has been effectively used in the breeding of blast resistant varieties worldwide12,13, suggesting that there may be signal recognition and transduction mechanisms among these R genes triggering rice immunity15. The Pita gene was cloned near the centromere on chromosome 12 of rice and encodes a 928-amino-acid cytoplasmic membrane receptor protein containing NBS-LRR domains14. AVR-Pita is the avirulent gene corresponding to Pita, which encodes a neutral zinc metalloprotease15. Pita was the first rice blast resistance protein confirmed to interact directly with an avirulent protein of the pathogen through its LRR domain, leading to resistance response16. Compared with Pita, Pita2 shows a higher level and wider spectrum of disease resistance. All rice varieties containing Pita2 have been reported to also contain Pita14,17,18. Ptr is a recently identified broad-spectrum rice blast resistance gene that is located physically close to the Pita region in the chromosome to the Pita region. Ptr may be specifically involved in Pita/Pita2-mediated resistance to rice blast and play a role in Pita-mediated signal recognition16. Further research has shown that Ptr is required for the function of Pita. The resistance mediated by Pita and Pita2 was eradicated by the loss of Ptr, suggesting that the integrity of the Ptr gene product is key for pathogen signal transduction. Ptr was significant to in the broad-spectrum resistance of the Pita gene complex. Additionally, a Ptr mutant failed to recognize both AVR-Pita and Pita2-specific AVR genes, and all rice varieties carrying Pita2 contained the same Ptr haplotype. It was preliminarily inferred that Ptr and Pita2 may be the same gene, and Ptr was located about 210 kb upstream of Pita11. Recent research showed that Ptr and Pita2 were alleles. The loss of function of Pita2 but not Pita eliminated the specificity to some AvrPita containing isolates19.
Pita was the second blast resistance gene cloned in rice14. Due to close linkage with other resistance genes, Pita-containing varieties often have a broad blast resistance spectrum and thus have been popularized and well cultivated around the world12,14,20,21. The AVR-Pita176 protein binds directly to the Pi-ta LRD region inside the plant cell to initiate a Pi-ta-mediated defense response9,19. Intriguingly, a naturally occurring susceptible allele always encodes a protein that harbors a single amino acid substitution from alanine to serine at position 918 (A918S) of the Pita resistance protein14,22. The mutation of this residue has been shown to disrupt the interaction between Pita and its avirulent protein in a yeast system and an in vitro binding assay, indicating the importance of Ala at position 918 for the function of Pita12.
Here, we have identified a blast resistance gene in Fuhui 2663, an Indica rice cultivar resistant to blast fungus, and designated Pita-Fuhui2663 was based on nucleotide polymorphism. Pita-Fuhui2663 encodes an NBS-LRR protein containing an amino acid change, A918S, compared with the sequence of the Pita-S protein. Using CRISPR/Cas9 genome editing technology, we knocked out Pita-Fuhui2663 in Fuhui 2663 and found that the knockout mutants lost its resistance, indicating that Pita-Fuhui2663 confers blast resistance. Moreover, the analysis of knockout lines showed that the Pita-Fuhui2663 gene did not affect the main agronomic traits of rice. We also developed a functional molecular marker, Pita-Fuhui2663-dCAPS, for Pita-Fuhui2663. These results suggest that Pita-Fuhui2663 has good application prospects in rice blast resistance breeding.
Materials and methods
Statement
The current study complies with relevant institutional, national, and international guidelines and legislation for experimental research and field studies on plants (either cultivated or wild), including the collection of plant materials.
Plant materials
The Indica rice CO39 and Japonica Lijiangxintuanheigu (LTH) were kept in the Rice Research Institute, Fujian Academy of Agricultural Sciences (Fuzhou, Fujian, China). Fuhui 2663, an Indica restorer line with good quality and blast resistance, was selected and bred by the Rice Research Institute. The F1 plants, generated from Minghui63 as female and Jiafuzhan as male, were back crossed with Minghui63 to produce the BC1F1 generation, and these lines were self-interbred for 9 generations, and a stable line was obtained, named as Fuhui 2663. Fuhui 2663 has been applied for a protected new national plant variety in 2021.
In the summer of 2019, Fuhui 2663 was used as a donor to cross with LTH and CO39. F1 seeds were sown in the spring and F2 seeds were harvested at the Sanya Experimental Station in Hainan Province, China. Fuhui 2663, and the two knockout lines (Pita-Fuhui2663-KO-1 and Pita-Fuhui2663-KO-2) in the background of Fuhui 2663 were planted in the summer of 2021 at the Experimental Station in Fujian Province, China.
Magnaporthe oryzae isolation
The M. oryzae isolate KJ201 was provided by the Fujian Academy of Agricultural Sciences. It was isolated by single-spore separation and stored on sterilized rice stems in a − 20 °C freezer.
Inoculation and resistance evaluation of rice blast
The seeds were planted on petri dishes and, after 5 days, the uniformly sprouting seedlings were moved to small pots containing nutrient-rich soil. Each pot was planted with 12 seedlings, and three replicates were set up. The rice seedlings with 3–4 leaves were moved to an inoculation chamber with high humidity, and the spore suspension of the M. oryzae isolate KJ201 was sprayed on the plants with a spray gun. The inoculated plants were transferred to a plastic box covered with a wet sponge and cultured in a dark room (95–100% relative humidity (RH), 25 °C) for 24 h, and then transferred to a greenhouse at 25–28 °C for 6 days23. Grades 0–3 corresponded to resistant, and grades 4–9 corresponded to susceptible. The disease response was detected 1 week after inoculation, using susceptible cultivar CO39 or LTH as a control and the disease-resistant cultivar Fuhui 2663 as a positive control.
Polymerase chain reaction amplification and marker detection
DNA was extracted from frozen leaves of rice by using the cetyltrimethylammonium bromide method24 with minor modifications. DNA polymerase chain reaction (PCR) amplification was performed as previously described28, amplification was performed using the following programme: 5 min at 94 °C; 35 cycles of 1 min at 94 °C, 40 s at 59 °C (for Indel) or 55 °C (for SSR) and 2 min at 72 °C; and a final extension of 5 min at 72 °C.
Bulked segregant analysis
Bulked segregant analysis was used to search for markers linked to target genes. The leaf DNA of 20 susceptible plants randomly selected from the F2 population (Fuhui 2663 × LTH) was used to construct the susceptible DNA pool, and the leaf DNA of 20 resistant plants randomly selected from the F2 population (Fuhui 2663 × LTH) was used to construct the resistant DNA pool. SSR markers distributed in the rice genome were used for linkage detection, and DNAs extracted from LTH and Fuhui 2663 were used as control. The band type of the marker linked to the susceptible gene was the same as that of LTH.
Primary mapping of the Pita-Fuhui2663 gene
The localization population was constructed by crossing Fuhui 2663 (Indica) with LTH (Japonica). The preliminary mapping of Pita-Fuhui2663 was randomly selected from 45 susceptible plants in all the F2 population (Fuhui 2663 × LTH), and a total of 536 susceptible plants were screened out for fine mapping in all the F2 population (Fuhui 2663 × LTH). MAPMAKER 3.025 was used for linkage analysis of Pita-Fuhui2663 loci and SSR markers, as reported by Rahman26,27.
Fine mapping of the Pita-Fuhui2663 gene
The bioinformatics analysis of the cultivar Nipponbare, published by the International Rice Genome Sequencing Project (IRGSP, http://rgp.dna.affrc.go.jp/IRGSP/index.html), was carried out to construct physical maps of target genes. The clones were anchored to the target gene linkage markers, and sequences were compared using BLAST (http://www.ncbi.nlm.nih.gov/blast/bl2seq/b12.html).
Bioinformatics analysis
Candidate genes were predicted on the basis of existing sequence annotation databases (http://rice.plantbiology.msu.edu/; http://www.tigr.org/). Rice plant DNA and amino acid sequences were used for complete comparison using Clustal X version 1.81. The design of the endonuclease site was based on the available database (http://helix.wustl.edu/dcaps/). The spatial structures of the Pita-Fuhui2663 and Pita-S proteins were predicted according to Robetta (https://robetta.bakerlab.org/submit.php).
Sequencing of candidate genes
Primers were designed according to the predicted full-length sequence of candidate genes. Generally, one primer was designed according to the 3’UTR region of each candidate gene, and then another primer was designed according to the 5’UTR region. The Fuhui 2663 genes was sequenced according to the designed primers.
Targeted mutagenesis of Pita-Fuhui2663 with CRISPR/Cas9
The Pita-Fuhui2663 gene in Fuhui 2663 was targeted with a gRNA spacer that spanned the two exons of the gene. The highly specific gRNA spacer sequences (Supplementary Table 1) were designed using the CRISPR-plant database and website28. Upon transformation, the regenerated plants were analyzed for genome editing mutations in target genes. Individual strains were selected from transgenic CRISPR-edited cell lines for the sequencing of specific mutations in PCR products29. About 300 rice seeds of Fuhui 2663 were subjected to mutation, about 150 seeds were recovered after antibiotic screening, about 20 positive seedlings were obtained by PCR after rooting, and then 5 mutants were obtained by sequencing, and finally 2 stable independent homozygous knockout lines (Pita-Fuhui2663-KO-1 and Pita-Fuhui2663-KO-2) were obtained. The primers for CRISPR/Cas9 used in this study are shown in Supplementary Table 1.
Declarations
The current study complies with relevant institutional, national, and international guidelines and legislation for experimental research and field studies on plants (either cultivated or wild), including the collection of plant materials.
Results
Genetic analysis of the resistance gene
To analyze the blast resistance of Fuhui 2663 in the laboratory, 2-week-old plants growing in the greenhouse were inoculated with blast fungus KJ201 isolate. Using susceptible rice CO39 and LTH as controls, Fuhui 2663 was found to have strong resistance to blast KJ201 (Fig. 1 and Supplementary Fig. 1).
Fuhui 2663 is resistant to KJ201 under laboratory conditions. Plants of Fuhui 2663 and CO39 at the 3–4-leaf stage were transferred to an inoculation chamber under high humidity, and 25 mL of spore suspension of M. oryzae isolate KJ201 was sprayed with an air compressor. The inoculated plants were transferred to a plastic box covered with a wet sponge, cultured in a dark room (95–100% RH, 25 °C) for 24 h, and then transferred to a greenhouse at 25–28 °C for 6 days, after which photographs were taken.
For genetic analysis of Fuhui 2663, the resistance donor Fuhui 2663 was crossbred with susceptible parents CO39 and LTH. A total of 36 F1 individuals (the Fuhui 2663 × CO39 and the Fuhui 2663 × LTH) showed a resistance phenotype against the M. oryzae isolate KJ201. The segregation of resistant (R) and susceptible (S) progenies in the F2 population fitted a 3:1 ratio (the Fuhui 2663 × CO39 F2 population, 412 R: 139 S, v2 = 0.305; the Fuhui 2663 × LTH F2 population, 382 R:124 S, v2 = 0.34, Table 1). The R/S ratio showed that Fuhui 2663 contained a dominant resistance gene.
Mapping of the resistance gene using molecular markers
To further determine the gene responsible for the resistance phenotype of Fuhui 2663, 506 SSR markers were obtained from the rice molecular map, and 317 primer pairs of markers showed polymorphisms between Fuhui 2663 and CO39. These 317 pairs of primer markers were used for linkage analysis of the DNA pools from 20 resistant plants or 20 susceptible plants in the F2 population (Fuhui 2663 × LTH). Each primer pair was used for the detection of 4 DNA samples (Fuhui 2663, CO39, the pool of 20 resistant plants, and the pool of 20 susceptible plants). When using indel-12-4 and indel-12-7 as primers, the test results showed that the size of the PCR product of Fuhui 2663 was the same as that of the DNA pool of 20 resistant plants, while CO39 was the same as that of the DNA pool from 20 susceptible plants. Therefore, we speculated that the Indel-12-4 and Indel-12-7 markers may be linked to the resistance gene.
To preliminarily locate this resistance gene, 45 susceptible individual plants were used for further validation, and the results showed that Indel-12-4 and Indel-12-7 were linked to the resistance locus. Therefore, the resistance gene was located between the markers Indel-12-4 and Indel-12-7 on chromosome 12, with an estimated physical distance of about 8.9 Mb (Fig. 2a).
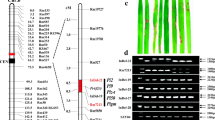
Genetic and physical maps of the Pita-Fuhui2663 gene. (a) Primary mapping of the resistance gene; the Pita-Fuhui2663 gene was identified between markers Indel-12-4 and Indel-12-7. (b) Further mapping of the resistance gene; the Pita-Fuhui2663 gene was identified between markers RM27860 and RM27960. (c) Fine mapping of the resistance gene; the Pita-Fuhui2663 gene was identified between markers Indel-11 and Indel-12. (d) High-resolution mapping of the resistance gene; the Pita-Fuhui2663 gene was eventually mapped to an 80-kb region between markers Indel-26 and Indel-33, and the number of recombinations between the marker and the target gene is shown on the linkage map. (e) There were eight annotated genes in the 80-kb region.
Fine mapping of the resistance gene
To map the resistance gene to a smaller region, 536 susceptible individual plants were identified from the F2 population (Fuhui 2663 × LTH). Further mapping was performed by using published markers (http://archive.gramene.org/markers/), and six SSR polymorphisms (RM27728, RM27766, RM27830, RM27860, RM27960 and RM27983) were selected from 18 markers between Indel-12-4 and Indel-12-7. Using these six markers, the resistance gene was found to be located between the molecular markers RM27860 and RM27960 with a physical distance of 3.28 Mb (Fig. 2b and Supplementary Table 2).
To further localize the resistance gene, five Indel polymorphisms were selected from 20 new markers between RM27860 and RM27960. Indel marker development referred to the insertion or deletion of nucleotide fragments of different sizes at the same site of the genome between related species or different individuals of the same species, and Indel markers were designed on the basis of published rice genome sequences, and the polymorphisms between Fuhui 2663 and CO39 were predicted by comparing the sequences of Nipponbare (http://rgp.dna.affrc.go.jp/) and 93–11 (http://rice.genomics.org.cn/). The resistance gene was found to be located between the molecular markers Indel-11 and Indel-12 on chromosome 12, and the physical distance between the two markers was 353 kb (Fig. 2c and Supplementary Table 2).
For further fine mapping, four polymorphic Indels were selected from 15 new Indel markers (Supplementary Table 2). Six markers (Indel-11, Indel-22, Indel-26, Indel-30, Indel-33, and Indel-12) were used for recombinant screening, and 12, 5, 2, 0, 1, and four recombinant plants were detected, respectively. Therefore, the resistance gene was precisely located in the estimated 80 kb region between Indel-26 and Indel-33 (Fig. 2d).
Candidate genes in the 80 kb region
According to the existing sequence annotation databases (http://rice.plantbiology.msu.edu/; http://www.tigr.org/), there were eight candidate genes in the 80 kb region identified (Fig. 2e), and all genes had a corresponding full-length cDNA. LOC_Os12g18320, LOC_Os12g18330, LOC_Os12g18340, LOC_Os12g18350 and LOC_Os12g18400 encode retrotransposon proteins; LOC_Os12g18360 encodes an NBS-LRR protein; LOC_Os12g18390 encodes a centromere-specific protein; LOC_Os12g18400 encodes a mitochondrial iron-regulated protein (MIR); and LOC_Os12g18410 encodes a retrotransposon protein and also the MIR protein.
Sequence analyses of the resistance gene
Further analysis revealed that LOC_Os12g18360, encoding an NBS-LRR protein, was the Pita gene in this locus14. To see if Pita was involved in the resistance phenotypes, we next sequenced Pita in Fuhui 2663 and CO39. Sequencing revealed a base substitution that resulted in the mutation of amino acid 918 from alanine (A) to serine (S) (Fig. 3), we speculated that this mutation led to the loss of resistance. These results indicated that this resistance gene in Fuhui 2663 was most likely allelic to Pita. Therefore, we named this gene Pita-Fuhui2663. According to the position of Ptr, Ptr was located about 210 kb upstream of Pita-Fuhui2663 and Pita (Supplementary Fig. 2).
Because alanine is a nonpolar amino acid and serine is a polar amino acid, this substitution can be expected to alter the structure of the Pita-S protein. By simulating the spatial structure of Pita-Fuhui2663 and Pita-S based on the Robettamodel program, we observed a structural change between the Pita-Fuhui2663 and Pita-S proteins in the LRR domain (Fig. 4), speculating that Ala918 is an important site for Pita-Fuhui2663, and the mutation to Ser918 may partly affect the function of the Pita-S protein.
Spatial structures of the Pita-Fuhui2663 and Pita-S proteins. There are significant structural changes in the LRR domain based on the Robettamodel program, and the 918th amino acid is substituted from alanine (Pita-Fuhui2663) to serine (Pita-S), which is likely to affect Pita-S protein function. The gray square indicates the site of structural changes.
Pita-Fuhui2663 is responsible for the resistance of Fuhui 2663 to blast
To confirm that Pita-Fuhui2663 is responsible for blast resistance, we investigated whether gene knockout of Pita-Fuhui2663 in the resistant cultivar Fuhui 2663 resulted in a susceptible phenotype. To this end, a sequence-specific guide RNA (gRNA) for the second exon of Pita-Fuhui2663 was designed using the CRISPR/Cas9 gene-editing system. Two plants (Pita-Fuhui2663-KO-1 and Pita-Fuhui2663-KO-2) were obtained from two independent events and were found to have a 1 bp insertion or deletion within a targeted site based on sequencing (Fig. 5a, Supplementary Table 3), affect their amino acid sequence changes (Supplementary Fig. 3).
Pita-Fuhui2663 knockout lines showed enhanced susceptibility to KJ201. (a) Two independent lines (designated Pita-Fuhui2663-KO-1 and Pita-Fuhui2663-KO-2) were generated using the CRISPR/Cas9 system and verified by sequencing. (b) Inoculation of rice blast fungus showed that the two knockout lines generated by CRISPR/Cas9 were susceptible to KJ201, while their parent Fuhui2663 was resistant to KJ201. Leaves were photographed 6 days post-infection with M. oryzae isolate KJ201. (c) Lesion numbers per cm2 on the rice leaves (M ± SD, n > 6 leaves) after inoculation with blast fungus as in (b). **denotes P < 0.01.
We then inoculated the two homozygous mutants with KJ201, and both lines were completely susceptible to KJ201 (Fig. 5b,c). Therefore, we concluded that the targeted mutation of Pita-Fuhui2663 in the resistance of Fuhui 2663 led to disease susceptibility to KJ201, demonstrating that Pita-Fuhui2663 is responsible for blast resistance in Fuhui 2663.
Main agronomic characteristics of Fuhui 2663 and the two knockout lines
We next monitored several key agronomic traits in Fuhui 2663 and the two knockout lines (Pita-Fuhui2663-KO-1 and Pita-Fuhui2663-KO-2) to assess their potential in rice breeding applications. We compared yield-related traits, including plant height, panicle length, number of effective panicles, spikelets per panicle, seed setting rate, thousand-grain weight, and grain length and width. The results showed that there was no significant difference in these main agronomic characteristics between Fuhui 2663 and the two knockout lines (Table 2), suggesting that Pita-Fuhui2663 mainly played a role in rice blast resistance and did not affect other agronomic traits, likely having good breeding value and potential in the future.
Development of a functional marker for the Pita-Fuhui2663 gene
Considering the importance of the Pita-Fuhui2663 gene and the deficiency of existing markers, we developed a Pita-Fuhui2663-dCAPS molecular marker using cloned Pita-Fuhui2663 cDNA sequences for efficient marker-assisted selection (Supplementary Table 1). There is a single base substitution between Pita-Fuhui2663 and Pita-S. According to this difference, we found an EcoP15I enzyme digestion site. The susceptible gene Pita-S could be specifically recognized by the endonuclease EcoP15I, and a 98 bp fragment could be amplified, and Lane 2 showed the amplified target fragment of 98 bp from Pita-S, while the resistance Pita-Fuhui2663 could not, and could only be amplified to a 123 bp fragment, where Lane 1 showed the amplified target fragment of 123 bp from Pita-Fuhui2663 (Supplementary Fig. 4).
Discussion
Genetic and functional analysis of the Pita-Fuhui2663 gene
Although Pita had been cloned, how this residue contributes to the association between Pita and AVR-Pita is unclear. Here, we simulated the spatial structure of Pita-Fuhui2663 and its susceptible variant Pita-S (A918S) and found a structural change in the LRR domain between Pita-Fuhui2663 and Pita-S. In addition, the two knockout mutants also showed different protein structural changes (Fig. 4). Considering the polarity difference between these two amino acids, Pita-S might show a change in the hydrophobic environment provided by Ala, hindering protein interactions. Meanwhile, abundant blast R genes are present not only within but also between species. Within the same rapidly evolving gene family, R genes can exhibit an effector response to develop resistance to rapidly evolving fungal pathogens. This was included in a proposed unique mechanism called "restricted differentiation", in which R genes and pathogen effectors can only follow a limited evolutionary path to improve fitness30. However, further research should be conducted to uncover the molecular machinery.
Analysis of the application prospects of the Pita-Fuhui2663 gene
Fuhui 2663 is a new restorer line bred by our team that harbors excellent characteristics for the paddy field, with a good leaf shape, high seed setting rate, and considerable resistance to blast disease. In this study, we used fine mapping to determine Pita-Fuhui2663 as the single dominant gene that confers Fuhui 2663 blast resistance and performed the functional identification of this gene through the use of the CRISPR/Cas9 system. Previously, a subset of rice core accessions in the United States was collected to evaluate the relationship between blast resistance and yield-related components, revealing that rice genomes with Pita correlated with lighter seed weights31. Recent studies have shown that susceptible Pita has an important effect on rice yield, and different Pita allelic mutants have different effects on rice yield32. However, our data showed that, except for disease susceptibility, the two Pita-Fuhui2663 knockout lines based on Fuhui 2663 showed no significant differences in other agronomic traits, including thousand-grain weight. These suggest that the presence of Pita-Fuhui2663 has little impact on the growth and development of Fuhui 2663. We speculate that one or more unknown factors may be expressed specifically in the genome of Fuhui 2663 to balance its defense and growth. Such a synergistic mechanism has been confirmed in some other blast resistance proteins. For instance, PigmR dimerization for resistance was attenuated competitively by its homolog PigmS, resulting in the suppression of immune responses and yield cost1. Therefore, elucidating the association between Pita-Fuhui2663-induced immunity and seed production in Fuhui 2663 warrants further studies.
In order to further analyze the distribution of Pita-Fuhui2663, we performed SNP (Single nucleotide polymorphisms) calling and haplotype (Hap) analysis of the 3000 sequenced rice genomes available in the CNCGB and CAAS databases33 and found 25 Haps for the Pita-Fuhui2663 gene, including 9 Haps among more than 15 rice resource materials (Supplementary Table 4). Further analysis revealed that Hap 1 and Hap 9, a total of 703 materials contained Pita-Fuhui2663, while the other Haps were different from Pita-Fuhui2663. Therefore, of the 3,000 sequenced rice genomes, there were relatively few Haps containing Pita-Fuhui2663.
Marker-assisted selection is an important method to increase the efficiency of rice resistance breeding34. Jia et al.35 initially analyzed natural variation at the Pita locus and developed a set of dominant markers based on a PCR method. Afterward, Wang et al.36 further optimized the Pita marker system to two pairs of dominant markers, according to a polymorphic site in the intron (GCC to CTAT), which have been widely used in rice breeding. Here, we used the dCAPS method to develop a Pita-Fuhui2663 functional marker using the restriction endonuclease EcoP15I. This marker depends on the crucial polymorphic site (A918S in proteins) in the second exon of Pita-Fuhui2663. It was accordingly used to identify the Pita-Fuhui2663 resistance locus in our tests as a complement to the Pita-Fuhui2663 marker system. Thus, to breed a new hybrid rice variety, breeders can transfer Pita-Fuhui2663 into both restorer and sterile lines using molecular marker-assisted selection.
In conclusion, Fuhui 2663, with excellent comprehensive traits, contains the Pita-Fuhui2663 resistance gene, and we believe that this rice variety will have great potential in future scientific research and breeding applications.
Conclusions
In this study, using a map-based cloning strategy, the Pita-Fuhui2663 gene from rice cultivar Fuhui 2663 was mapped to an 80-kb resistance locus region that contained the Pita gene. Then, CRISPR/Cas9 knockout experiments confirmed that Pita-Fuhui2663 is responsible for the resistance phenotype of Fuhui 2663. Importantly, Pita-Fuhui2663 did not affect the main agronomic traits of the variety compared to the Pita gene as verified by knockout experiments, indicative of potential applications of Pita-Fuhui2663 in broader breeding programs. Finally, a Pita-Fuhui2663-dCAPS molecular marker with good specificity and high efficiency was developed, the application of which will facilitate combating this devastating disease.
Data availability
All data is included in this publication as figures, tables and supplements.
References
Deng, Y. W. et al. Epigenetic regulation of antagonistic receptors confers rice blast resistance with yield balance. Science 355, 962–965 (2017).
Madden, L. V. & Wheelis, M. T. The threat of plant pathogens as weapons against U.S. crops. Annu. Rev. Phytopathol. 41, 155–176 (2003).
Das, A. et al. A novel blast resistance gene, Pi54rh cloned from wild species of rice, Oryza rhizomatis confers broad spectrum resistance to Magnaporthe oryzae. Funct. Integr. Genomic 12, 215–228 (2012).
Wang, B. H., Ebbole, D. J. & Wang, Z. H. The arms race between Magnaporthe oryzae and rice: Diversity and interaction of Avr and R genes. J. Integr. Agric. 16, 2746–2760 (2017).
Hua, L. X. et al. The isolation of Pi1, an allele at the Pik locus which confers broad spectrum resistance to rice blast. Theor. Appl. Genet. 125, 1047–1055 (2012).
Qu, S. H. et al. The broad-spectrum blast resistance gene Pi9 encodes a nucleotide-binding site-leucine-rich repeat protein and is a member of a multigene family in rice. Genetics 172, 1901–1914 (2006).
Chen, J. et al. A Pid3 allele from rice cultivar Gumei2 confers resistance to Magnaporthe oryzae. J. Genet. Genomics 38, 209–216 (2011).
Hutin, M. et al. Ectopic activation of the rice NLR heteropair RGA4/RGA5 confers resistance to bacterial blight and bacterial leaf streak diseases. Plant J. 88, 43–55 (2016).
Chen, X. W. et al. A B-lectin receptor kinase gene conferring rice blast resistance. Plant J. 46, 794–804 (2006).
Fukuoka, S. et al. Loss of function of a proline-containing protein confers durable disease resistance in rice. Science 325, 998–1001 (2009).
Zhao, H. J. et al. The rice blast resistance gene Ptr encodes an atypical protein required for broad-spectrum disease resistance. Nat. Commun. 9, 2039 (2018).
Jia, Y. et al. Rice Pi-ta gene confers resistance to the major pathotypes of the rice blast fungus in the United States. Phytopathology 94, 296–301 (2004).
Jia, Y. Artificial introgression of a large chromosome fragment around the rice blast resistance gene Pi-ta in backcross progeny and several elite rice cultivars. Heredity (Edinb) 103, 355–356 (2009).
Bryan, G. T. et al. A single amino acid difference distinguishes resistant and susceptible alleles of the rice blast resistance gene Pi-ta. Plant Cell 12, 2033–2046 (2000).
Orbach, M. J., Farrall, L., Sweigard, J. A., Chumley, F. G. & Valent, B. A telomeric avirulence gene determines efficacy for the rice blast resistance gene. Plant Cell 12, 2019–2032 (2000).
Jia, Y. L., McAdams, S. A., Bryan, G. T., Hershey, H. P. & Valent, B. Direct interaction of resistance gene and avirulence gene products confers rice blast resistance. EMBO J. 19, 4004–4014 (2000).
Kiyosawa, S. Gene analysis of blast resistance in exotic varieties of rice. JPN. Agr. Res. Q. 6, 8–15 (1971).
Kiyosawa, S. An attempt of classification of world’s rice varieties based on reaction pattern to blast fungus strains. Bull. Natl. Inst. Agrobiol. Resour. 2, 13–39 (1986).
Meng, X. et al. The broad-spectrum rice blast resistance (R) gene Pita2 encodes a novel R protein unique from Pita. Rice 13, 19. https://doi.org/10.1186/s12284-020-00377-5 (2020).
Ma, J., Sun, Y., Yang, Y., Li, J. & Minghui, X. Distribution of Pita gene for rice blast resistance in the rice cultivars (lines) from China Japan and Korea. Mol. Plant Breed. 18, 459–465 (2020).
Shi, K. et al. Distribution of two blast resistance genes Pita and Pib in major rice cultivars in China. J. Plant Genet. Resour. 10, 21–26 (2009).
Jia, Y., Bryan, G., Farrall, L. & Valent, B. Natural variation at the Pi-ta rice blast resistance locus. Phytopathology 93, 1452–1459 (2003).
Yang, D. W. et al. Identification and application of the Pigm-1 gene in rice disease resistance breeding. Plant Biol. (Stuttg) 22, 1022–1029 (2020).
Murray, M. G. & Thompson, W. F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids. Res. 8, 4321–4325 (1980).
Lander, E. S. et al. Mapmaker: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1, 174–181 (1987).
Liu, H. R. & Meng, J. L. MapDraw: A microsoft excel macro for drawing genetic linkage maps based on given genetic linkage data. Hereditas (Beijing) 25, 317–321 (2003).
Rahman, M. L. et al. Identification of QTLs for some agronomic traits in rice using an introgression line from Oryza minuta. Mol. Cells 24, 16–26 (2007).
Xie, K. B., Zhang, J. W. & Yang, Y. N. Genome-wide prediction of highly specific guide RNA spacers for CRISPR-Cas9-mediated genome editing in model plants and major crops. Mol. Plant 7, 923–926 (2014).
Ma, X. L., Chen, L. T., Zhu, Q. L., Chen, Y. L. & Liu, Y. G. Rapid decoding of sequence-specific nuclease-induced heterozygous and biallelic mutations by direct sequencing of PCR products. Mol. Plant 8, 1285–1287 (2015).
Yang, S. H. et al. Rapidly evolving r genes in diverse grass species confer resistance to rice blast disease. P. Natl. Acad. Sci. USA 110, 18572–18577 (2013).
Wang, X., Jia, M. H., Ghai, P., Lee, F. N. & Jia, Y. Genome-wide association of rice blast disease resistance and yield-related components of rice. Mol. Plant Microbe Interact. 28, 1383–1392 (2015).
Liu, J. et al. Fitness benefits play a vital role in the retention of the Pi-ta susceptible alleles. Genetics. 220(4), iyac019 (2022).
Li, Z. K. et al. The 3,000 rice genomes project. Gigascience 3, 7 (2014).
Xu, Y. & Crouch, J. H. Marker-assisted selection in plant breeding: From publications to practice. Crop Sci. 48, 391–407 (2008).
Jia, Y., Wang, Z. & Pratibha, S. Development of dominant rice blast resistance Pi-ta gene markers. Crop Sci. 42, 2145–2149 (2002).
Wang, Z., Jia, Y., Wu, D. & Xia, Y. Molecular markers-assisted selection of the rice blast resistance gene Pi-ta. Acta Agron. Sin. 30, 1259–1265 (2004).
Acknowledgements
We thank Dr. Dagang Tian (Fujian Academy of Agricultural Sciences) for providing the M. oryzae isolate KJ201.
Funding
This work was supported by the Fujian Provincial Natural Science Foundation (No. 2021J01471 and 2022J05076), the Free Exploration Project (ZYTS202202), the Special Fund for Agro-scientific Research in the Public Interest of Fujian Province (No. 2020R1023003), Major Science and Technology Projects of Fujian Province (No. 2020NZ08016), Science and Technology Innovation Team (No. CXTD2021001), 5511 Collaborative Engineering Project (No. XTCXGC2021001), the 100 Talent Plans of Fujian Province and Rice Molecular Breeding Science and Technology Innovation Team (CXTD2021005-3).
Author information
Authors and Affiliations
Contributions
D.Y. conceived and designed the research. F.H. and N.H. carried out most experiments. D.Y., N.H., M.Y., Y.Z. wrote the manuscript, Q.Q.L. revised the manuscript and coordinated submission. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
He, N., Huang, F., Yu, M. et al. Analysis of a rice blast resistance gene Pita-Fuhui2663 and development of selection marker. Sci Rep 12, 14917 (2022). https://doi.org/10.1038/s41598-022-19004-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-19004-y
- Springer Nature Limited
This article is cited by
-
The unconventional resistance protein PTR recognizes the Magnaporthe oryzae effector AVR-Pita in an allele-specific manner
Nature Plants (2024)
-
Allelic variation in rice blast resistance: a pathway to sustainable disease management
Molecular Biology Reports (2024)
-
CRISPR-Cas System, a Possible “Savior” of Rice Threatened by Climate Change: An Updated Review
Rice (2023)