Abstract
Congenital sensorineural deafness (CSD) has been reported to affect up to 30% of Dalmatian dogs world-wide and while unilaterally deaf dogs can live a close to normal life, dogs suffering bilateral deafness are frequently euthanized. Extreme-white coat patterning as encoded by the gene Melanocyte Inducing Transcription Factor (MITF) has long been postulated as the major risk factor for CSD in the Dalmatian breed. While attempts to identify causative risk variants associated with CSD have been numerous, no genome-wide association study has positively identified MITF as a risk locus for either bilateral or unilateral deafness in the Dalmatian breed to date. In this study, we identified an association with CSD on CFA20 in the vicinity of MITF within Australian Dalmatian dogs. Although not genome-wide significant, the association signal was validated by reanalysing publicly available data and merging the wider data resource with the local data to improve statistical power. The merged data, representing three major global populations of Dalmatian dogs, enabled us to identify a single, well-defined genome-wide significant risk haplotype for CSD. The haplotype was formed by three genome-wide significant associated markers (BICF2G630233852T>C, BICF2G630233861T>C, BICF2G630233888G>A) on CFA20 with 62% of bilaterally deaf dogs homozygous for the risk haplotype (CCA), while 30% of bilaterally deaf and 45% of hearing dogs carried one copy of the risk haplotype. Animals homozygous or heterozygous for the low-risk haplotype were less likely to be unilaterally deaf. While the association between the risk haplotype and deafness is incomplete, animals homozygous for the risk haplotype were 10-times more likely to be bilaterally deaf. Although the underlying causative variants are yet to be discovered, results from this study can now assist with reducing deafness in Dalmatian dogs.
Similar content being viewed by others
Introduction
Congenital sensorineural deafness (CSD, OMIA #000259-9615) in dogs presents a major challenge to breeders and owners, and frequently results in the euthanasia of bilaterally deaf dogs. The prevalence of CSD varies among dog breeds, with the highest prevalence reported in Dalmatians1,2,3,4,5,6,7. Strict selection of breeding animals has decreased the prevalence of CSD in Dalmatians in some countries, however, around one in three Dalmatians world-wide is still affected by hearing loss in either one or both ears8,9,10,11,12. The clinical and histopathological aspects of CSD have been thoroughly examined, and two main types have been described: the neuroepithelial type, and the cochleo-saccular type13,14,15. One difference in the phenotypic expression of the two types is that the neuroepithelial type is usually expressed bilaterally and the cochleo-saccular type can be either unilateral or bilateral16,17. While disease progression varies, deafness in both types is caused by the degeneration of hair cells, the stria vascularis and the vestibular membrane16,17,18. Current findings indicate that the absence of melanocytes in the inner ear is the main underlying predisposing factor and that the cochleo-saccular type is associated with skin and coat pigmentation5,6,12,18,19,20,21.
In the dog, extreme-white coat patterning caused by the S locus and encoded by the gene Melanocyte Inducing Transcription Factor (MITF) has long been postulated as a major risk factor for CSD1,22,23,24,25. The accepted implication has been that extreme-white dogs are equally at risk for CSD by genotype, and that differential phenotypic expression is random. Extreme-white is a breed-hallmark trait of the Dalmatian dog breed, although varying expression of patching and blue eyes suggest that genetic variation exists at the MITF locus26. Dalmatians expressing pigmented patches on their head and/or body appear to have a reduced deafness risk, while the risk increases with the presence of one or two blue eyes1,4,5,8,12,16. Given our current understanding of the phenotypic effects of the gene MITF, both observations support the potential role of MITF in the variable expressivity of CSD.
Despite the clear phenotypic description of CSD, the high prevalence in some breeds, and numerous efforts to uncover the genetic basis of canine deafness, few candidates for causative variants have been identified so far, with none in the Dalmatian dog27,28,29,30,31. Attempts to identify causative risk variants associated with CSD in Dalmatian dogs have been numerous25,30,32,33,34,35,36,37,38. Most recently, a genome-wide association study (GWAS) that included 304 Dalmatian dogs with sensory phenotypes identified strong associations with bilateral deafness on CFA23, CFA30, CFA37 and CFA3839. In contrast, an earlier GWAS identified a range of association signals on alternative chromosomes (CFA6, CFA14, CFA27, CFA29 and CFA31)35. Interestingly, none of the GWAS conducted to date in the Dalmatian breed have positively identified MITF as a risk locus for either bilateral or unilateral deafness. There is inconsistency of associated loci between studies on the same breed, and this suggests that current analyses are either underpowered; the trait is truly complex; or that other factors such as differing phenotyping approaches and population effects are hampering the analyses. Furthermore, studies commonly apply brainstem auditory evoked response (BAER) testing to assess the clinical expression of deafness. BAER results can be obtained using a variety of equipment types and operators. Animals may be awake or sedated when the hearing is assessed. Varying clinical phenotyping conditions and techniques may lead to inaccurate clinical assessments that might particularly affect unilateral versus normal phenotypes, since these animals remain responsive to sound.
Two canine reference assemblies (Broad/canFam2, hereafter referred to as canFam2 and Broad CanFam3.1/canFam3, hereafter referred to as canFam3) include a structural mis-assembly of the MITF gene. This mis-assembly not only affects the placement of exon 1 of the MITF-A isoform but also the gene orientation relative to surrounding genes40. Analysis of the region is further complicated by the existence of retrotransposed pseudogenes of the MITF-M and MITF-A isoforms, with the MITF-A retrocopy located on the Y chromosome40,41. Array markers in recent commercial genotyping arrays (e.g. Illumina Canine 220 K array) include erroneous variants from the MITF-A pseudogene, including the presence of two putative mis-sense mutations (rs851676581 and rs851603213)40. While these factors together hamper the assessment of linkage-disequilibrium (LD) and haplotype throughout the MITF region, there is also the possibility that the retrotransposon itself or other genes located in the genomic region identified may affect the deafness phenotype40.
In this study, a GWAS of CSD using Australian Dalmatian dogs with well characterised hearing phenotypes is performed. Results are validated with public-domain data of independent Dalmatian populations from the UK and the US to improve statistical power to overcome random error associated with population effects and phenotyping.
Results
Genome-wide association analysis
The analysis considered 145 Dalmatians (86 hearing, 33 unilateral deaf and 26 bilateral deaf) from Australia, with hearing phenotypes available. After quality filtering, a total of 78,830 variants were included in the analysis. No individual marker reached genome-wide significance after multiple-test correction by Bonferroni (Table S2). The most significantly associated marker was BICF2G630233861 T>C (CFA20:22045960) Praw = 8.44E − 06 (canFam3), with three of the top 10 markers located near or within the MITF gene. Of the remaining top 10 markers, one was located on CFA9, four on CFA34, one on CFA36 and one on CFA39. None of the top associated markers from this analysis validated previously identified associated loci (Table 1).
Putative functional variant detection
An approximately 300 kilobase region on CFA20 harbouring MITF was assessed using whole-genome sequence data of seven Australian Dalmatian dogs (5 bilateral deaf 2 hearing). The bioinformatic analysis identified a total of 1728 variants on CFA20 (UU_Cfam_GSD_1.0/canFam4 hereafter referred to as canFam4), including the most significantly associated markers previously identified by GWAS (Table S3). Manual inspection of the sequence alignments identified two SINE insertions; one at CFA20:22025737 (canFam4), previously identified by Karlsson et al.26, and one at CFA20:22168470 (canFam4), novel to this analysis. The second SINE (Fig. S2) potentially disrupts the 5’ UTR of two MITF transcripts (transcripts MITF.10 and MITF.14). Of the identified variants, 1014 were single nucleotide variants (SNVs) and 714 were insertion/deletions (INDEL). After focusing on variants near or within MITF transcripts annotated as expressed in the canine in the canFam4 reference genome, and the exclusion of variants unique to dogs carrying the MITF retrotransposon as determined by heterozygosity for rs851676581 and rs851603213, a total of 26 variants (22 SNVs, two repeat-element insertions, and one length polymorphism) remained for consideration (Table 2). The SINE element insertion described by Karlsson et al.26 was monomorphic as was the newly described allele at rs853013202 CFA20:22022763–22022766AAA>T (canFam4), which is located in the PAX3/BRN2 binding site in the MITF-M promoter. Four of the five deaf Dalmatians were homozygous for the alternate allele across all 24 polymorphic variants assessed, while the two hearing Dalmatians were either heterozygous or homozygous for the reference allele. One deaf Dalmatian (Dog 80) shared genotypes of nine variants with the deaf dogs but shared genotypes of 15 variants with hearing dogs. The genomic region homozygous in all deaf dogs includes three variants located in the promotor of MITF-M, six variants in the 5’-UTRs of two dog-specific transcripts (MITF.10 and MITF.14), and seven variants within the transcribed portions of long non-coding RNAs. A further nine variants identified in the vicinity of MITF.1 first exon were poorly genotyped due to high GC content affecting genome sequence coverage. These are reported in Table S3 and are in the range CFA20: 22205233–22207639 (canFam4).
Genome-wide re-analysis of publicly available data
A total of 304 Dalmatians sourced from the UK (N = 120) and the USA (N = 184) were subjected to a quantitative analysis using the first component of variance from multi-dimensional scaling as a covariate to account for population stratification, and a second GWAS applying the Cochran-Mantel Haenszel (CMH) test for stratified case–control data. After filtering, 125,209 variants and 304 dogs passed quality control and were included in subsequent analyses (Table S4). While none of the markers reached genome-wide significance in the genome-wide quantitative association analysis, six of the top 10 associated markers were located on CFA 20 (Table 1). All six markers were in close proximity to MITF. The remaining four markers were located on CFA17 (BICF2P1152155, BICF2G630202319, BICF2P522035) and CFA34 (BICF2P764018). The most significantly associated marker from the genome-wide quantitative association analysis was located at CFA20:23,191,697 (p = 2.4 × 10–5) (canFam3). Similarly, five of the top 10 markers from the genome-wide association analysis applying the CMH test were located on CFA20 (TIGRP2P272624_rs8804915, BICF2G630233852, BICF2G630233861, BICF2G630233888, BICF2G630234028), with the remaining markers on CFA1 (BICF2P814549), CFA7 (BICF2S24318175, BICF2G63086882), and CFA24 (BICF2P594756, BICF2P1033323) (Table 1). All associated markers on CFA20 were located within or near MITF, with two of the markers identified also amongst the top 10 associated markers in the quantitative analysis (CFA20:22031342 and CFA20:22116909) (canFam3). Interestingly, none of the top associated markers identified in previously published studies could be confirmed among the significant markers.
Expanded meta-analysis
Merging the dataset of UK/US Dalmatians with the Australian dataset resulted in a total of 443 dogs with hearing phenotypes, representing three major global populations. Three dogs were represented in more than one cohort (Australian and UK/US), therefore one instance for each dog was coded as missing. After filtering, 79,316 variants remained for the linear association analysis and 92,972 variants for the CMH analysis. The genomic inflation factors were 1.07 and 1.32 respectively. The GWAS revealed a significant quantitative trait locus on CFA20, with all top 10 associated markers located on CFA20, spanning a genomic region of approximately 3 megabases (Table 3). The three most significantly associated SNVs were within or in close proximity to MITF (BICF2G630233852, Praw = 1.04 × 10−8, Pgenome = 0.0008; BICF2G630233861, Praw = 1.04 × 10−8, Pgenome = 0.001; and BICF2G630233888, Praw = 1.62 × 10−8, Pgenome = 0.001) (Fig. 1, Table S5) (canFam3). When the same dataset was subjected to the CMH test, four of the top 10 associated SNVs were located on CFA8 between 55,535,050 and 56,016,460 bp (canFam3). The remaining six SNVs were located on CFA20, covering the same genomic interval as identified in the quantitative analysis. Three of the top 10 SNVs were identified in both analyses, with all three SNVs located on CFA20 in close proximity to MITF.
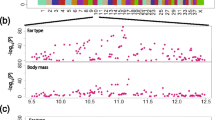
Genome-wide quantitative association meta-analysis of congenital sensorineural deafness in Dalmatian dogs. Negative log of probabilities for SNV markers based on quantitative trait association of congenital sensorineural deafness in 441 Dalmatian dogs sourced from three world continents. Animals were classified as either bilateral deaf, unilateral deaf or hearing based on recorded hearing testing results. Three markers associated with genome-wide significance are circled and their genome-wide significance reported. The Q–Q plot for the analysis is shown as an embedded image. The median Chi-squared value is (Lambda = 1.07551).
None of the markers previously associated with deafness in Dalmatian dogs ranked amongst the highest associated SNVs in this meta-analysis (Table 1). Across all analyses performed in this study, the locus on CFA20 spanning the genomic region between BICF2G630233852 and BICF2G630233888 was consistently identified as significantly associated with deafness in Dalmatians, with one SNV consistently identified as the most significantly associated marker (BICF2G630233852, CFA20:22031342, Praw = 1.04 × 10−8, Pgenome = 0.0008).
MITF risk haplotype analysis
Genotypes for the three genome-wide significant associated markers on CFA20 (BICF2G630233852T>C, BICF2G630233861T>C, BICF2G630233888G>A) were extracted from all 442 Dalmatian dogs, resulting in 53 dogs homozygous for the canFam4 reference haplotype (TTG), 189 dogs homozygous for an alternative haplotype (CAA) and 200 dogs heterozygous (other) (Table S7). Of the 442 dogs, 114 were bilateral deaf, 110 were unilateral deaf and 218 were classified as hearing. When hearing status was assessed based on haplotype, a decreased risk of clinical deafness was observed in animals carrying the canFam4 reference low-risk haplotype (TTG) (Fig. 2A). The same trend was observed when the Australian cohort was analysed separately, with a tenfold increase of deafness in animals homozygous for the alternative haplotype (CCA) compared with animals homozygous for the low-risk haplotype (TTG) (Fig. 2B). While change in the proportion of unilateral deaf dogs was not significantly associated with the risk haplotype, the presence of at least one high-risk allele showed a general trend of reduced unilateral deafness with only 19% of unilateral deaf dogs homozygous for the low-risk haplotype, compared to 27% homozygous for the high risk haplotype (Fig. 2). Using quantitative phenotypes, the association P-values for the high-risk (CCA) and low-risk (TTG) haplotypes, are 2.09 × 10−07 and 1.73 × 10−07 respectively. The probability for the haplotype CCA under quantitative association analysis (bilateral deaf score 2, unilateral scored as 1.5, control scored as 1) was 3.02 × 10−08. Haplotype by phenotype combinations are tabulated for the individual geographic cohorts (Table S8).
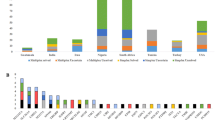
Deafness associated haplotype risk analysis. Deafness associated haplotype risk analysis using three genome-wide significant associated markers on CFA20 [BICF2G630233852T>C, BICF2G630233861T>C, BICF2G630233888G>A]. Animal were categorised as either homozygous risk haplotype, homozygous low-risk haplotype or other, with other including animals heterozygous for the risk/low-risk haplotype as well as other rare haplotype combinations. (A) shows the allocation of dogs (%) using the entire cohort of 442 dogs and (B) shows the allocation of dogs (%) using the Australian cohort of 139 dogs only. Animals classified as bilateral deaf are indicated in dark orange, unilateral deaf animals in yellow and hearing animals in green. Percentages of animals according to deafness status are included in the chart.
Association between pseudogene and deafness phenotype
A total of 14 markers in strong LD with the MITF retrotransposon were extracted from 89 animals genotyped on the 220 K genotyping array. The marker set included five markers on CFA19, seven markers on CFA20 and two markers on CFA32 (Table S6). While none of the 14 markers achieved genome-wide significance in the quantitative association, eight had a significant unadjusted quantitative association with CSD (P < 0.05). When hearing status was assessed against the presence or absence of the pseudogene, the proportion of hearing animals was increased among animals carrying the retrotransposon (Fig. 3).
Relationship between MITF retrotransposon and deafness. Segregation of deafness phenotype according to presence or absence of MITF retrotransposon among animals genotyped using the Illumina 220 K array. Animals classified as bilateral deaf are indicated in dark orange, unilateral deaf animals in yellow and hearing animals in green.
Discussion
White coat patterning caused by the S locus has long been established as a major risk factor for CSD in dogs1,3,4,20. In addition to the overall increase of white areas on the body, the risk of deafness increases in animals lacking facial pigmentation and with one or two blue irises1,3,4,6,7,10,25,30. Complex segregation analyses indicated that while CSD in Dalmatians is clearly heritable, the phenotype most likely follows a dichotomous or trichotomous mode of inheritance12. As a result, the MITF gene, which encodes the S locus, has been intensively studied but no causative mutation underlying CSD has been identified to date26,38. Using data from Dalmatian dogs representing three major global populations and applying multiple analyses enabled us to identify a significant association of CSD on CFA20, in the vicinity of MITF. More importantly, we identified a single, well-defined genome-wide significant risk haplotype for CSD in Dalmatian dogs.
The analysis of well characterised Australian Dalmatian dogs identified multiple array markers on CFA20 among the most associated markers with CSD. This finding was confirmed after combining all datasets available and re-analysing the entire Dalmatian data cohort with adjustment for stratification. This demonstrates that the association signal we have observed is consistent across populations and not unique to Australian-sourced animals. Utilising this population-level analysis enabled the identification of a strongly associated haplotype comprising three co-located markers (BICF2G630233852, BICF2G630233861, BICF2G630233888) that surround exon 1 of the MITF-A isoform (MITF.1 Uppsala University GSD1.0)42 according to the canFam4 genome assembly. Animals homozygous for the non-reference haplotype had a significantly increased risk of deafness. The identified risk haplotype increases bilateral deafness risk with allele dosage in an additive manner (Fig. 2A) altering bilateral deafness frequency from 38% in animals homozygous for the risk allele to 11.7% in animals homozygous for the low-risk haplotype. By contrast, the effect on unilateral deafness is less dramatic with a reduction of only 8% from homozygous high risk to homozygous low risk (27% down to 19%).
A weak association at the MITF locus, and in fact the same region and haplotype as identified in our analyses, was previously identified by Stritzel et al.38. The same research group later confirmed the weak association at the MITF locus in a mapping experiment35. Interestingly, despite a larger cohort size, Hayward et al.39 did not identify MITF as a major risk locus for deafness in Dalmatians. It is likely that a combination of the reference assembly issues raised by Brancalion et al.40 acting to obscure signal in this vicinity, along with heterozygosity of the canFam3 reference dog through the critical region (as evidenced by all three markers of the major haplotype comprising SNV discovered in the boxer assembly), together hampered phasing of haplotypes43. Furthermore, accurate phenotyping is one of the most critical components when conducting genetic studies. Phenotyping errors due to different equipment, operators or sedation methods used, may result in animals being improperly allocated to affection status groups. While not always under the control of the researcher, these errors reduce the power to detect associations44,45. The signal at MITF identified as a leading association in the Australian data is most likely due to most animals being assessed by a single veterinary neurologist using high-end BAER testing equipment. Such testing is expensive, and many breeders opt to have testing carried out at local clinics that have access to other BAER testing equipment operated by less experienced practitioners. Sampling errors in phenotypes generated by the other equipment or other operators with less experience cannot be discounted as a reason for low signal detection across all studies.
As the canFam3 reference assembly includes the risk variant at BICF2G630233888, it is possible that the risk haplotype identified in our study may confer deafness risk in other breeds exhibiting extreme white coat phenotypes, and perhaps particularly, the boxer. This conclusion is supported by a recent study on deafness in Australian Cattle Dogs, which also identified an association between the MITF locus and deafness46. While the most associated SNV in Australian Cattle Dogs differs from the one identified in Dalmatians, the quantitative trait locus identified in these two breeds overlaps. It is possible that genetic selection for the low-risk genotype will be a useful selection metric to reduce ongoing deafness risk when BAER testing is difficult or expensive to obtain, but this remains to be tested in a planned multi-generational study.
In addition to issues with the reference assembly, accurate assessment of MITF has further been impaired by the recently identified MITF retrotransposon on the canine Y chromosome (MITFY)40,41. The accumulation of retrotransposon-specific variants has caused a sex based genotyping bias, which has likely hampered the identification of associations between MITF and CSD41. Considering that regulatory variants or epistatic interactions among remote loci or compound within-locus variant interactions may affect the complex phenotype in an unpredictable manner, previous analyses may have been underpowered to detect the signal with statistical significance. Despite the accumulation of variants and the introduction of a premature stop codon, there is evidence that the MITF retrocopy on the Y chromosome is expressed41. While the exact function of the resulting protein is yet to be investigated, it has been established that many miRNA binding sites are preserved in the expressed transcript. Investigations of the role of miRNAs in the regulation of melanogenesis have indicated that depending on coat colour, miRNA expression profiles differ significantly, and it has been established that the predicted targets of differentially expressed genes are involved in pigmentation47,48,49. This not only indicates a potential role of the MITF retrotransposon on pigmentation and pigmentation-related disorders but supports our finding that the presence of the retrotransposon potentially modifies the risk of CSD (Fig. S1).
When putative functional variants were assessed across the associated region on CFA20, we identified a haplotype that was shared by all sequenced (n = 5) bilateral deaf Dalmatians. The risk-haplotype variants include a previously described length polymorphism in the promotor region of MITF-M at CFA20:22,022,802 (canFam4). This length polymorphism has been identified as a key regulator for white spotting in dogs, with longer versions of the polymorphism resulting in reduced promotor activity50. We were surprised to observe that deaf animals with the high-risk haplotype have a shorter version of the length polymorphism allele 32b (11C7A2G12A) compared to hearing Dalmatians, which were allele 35b (11C10A2G12A) and the newly described variant of length allele 34 (13C7A2G12A) using the descriptive terminology of Karlsson et al.26 and Baranowska Körberg et al.50. The allele-length 34 variant was also observed in a heterozygous form in the discordant deaf Dalmatian (sample identifier = 80). Variants that were unique to sequenced Dalmatians that harboured the MITF retrotransposon (including individual 80) were excluded from our assessment of functional variants. Thus, it is possible that individual 80 is a compound heterozygote for deleterious mutations on two different haplotype backgrounds, and that an alternative risk mutation may occur on the length 34 haplotype background but was not captured due to our attempt to exclude retrotransposon variants. Segregation of two risk alleles on alternate risk haplotypes will increase the complexity of the locus and may play a role in suppressing GWAS mapping signals from the wider MITF locus. The presence of potential alternative risk variants on other haplotypes may be the reason that the protective haplotype was associated with stronger significance than the risk haplotype in the meta-analysis.
The extreme white spotting phenotype that is characteristic of all Dalmatians has been observed to result from significantly lower MITF-M promotor activity. This is likely affected by a SINE insertion polymorphism affecting MITF-M located at CFA20:22,025,737 (canFam4) first described by Karlsson et al.26,50. This variant was monomorphic (SINE present) among our sequenced dogs. However, low levels of bilateral deafness among animals with the low-risk haplotype suggest that while the extreme white encoded by the SINE element may partly affect deafness risk, the risk is modified by additional variants in the MITF region in a complex manner. New variants of interest include the newly identified variant at rs853013202 CFA20:22022763–22022766AAA>T (canFam4) and variants surrounding the MITF.16 isoform. While not polymorphic in the five Dalmatians analysed, the rs853013202 variant is located in the PAX3/BRN2 binding site of the MITF-M promotor. Both BRN2 and PAX3 influence MITF expression51,52, and it is plausible that this variant could influence the phenotypic expression of CSD. In contrast, variants surrounding exon 1 of the MITF.16 isoform were unique to deaf Dalmatians. This MITF isoform has only been described in the dog, and to fully understand the potential impact of these and other variants on melanocyte development, further functional analyses are required.
The MITF-M transcript is exclusively expressed in melanocytes derived from the neural crest and retinal pigment epithelium, and expression at these sites is pivotal for all melanogenic processes53,54. Melanocytes within the auditory and vestibular system of the inner ear play a pivotal role in maintaining auditory and vestibular function in mammals. Located in the stria vascularis, melanocytes are necessary for the generation of the endocochlear potential and decreases in this potential have been linked to age-related hearing loss in humans55. Reduced pigmentation of skin, coat and eye has been associated with inner ear disfunction in a range of mammals, including humans, dog, cat, and horse1,4,56,57. Based on the histological changes observed in the inner ear, deafness in the Dalmatian breed is most likely of the cochleo-saccular type5,6,12,18,19,20,21. Current results indicate that the degeneration of the stria vascularis in the cochleo-saccular type is caused by the absence of melanocytes in the stria vascularis58,59,60. Considering that our results suggest a protective function of the MITFY retrocopy, it is plausible that expressed MITFY transcripts recover at least part of the lost activity and stimulate melanocyte development.
As only incomplete data on iris colour was available for Dalmatians included in this analysis, iris colour was not considered in this study, although presence of blue or part-blue eyes was annotated on the USYD neurology samples. In these animals two observations of iris dilution were in pups with normal hearing. Previous research suggests that variants in MITF are not only associated with CSD in Dalmatians but also with blue irises, and that the presence of one or two blue irises increases the risk of deafness16,38. While studies from humans indicate that variants in MITF are not involved in determining normal eye colour, some disease-associated variants in MITF have been linked to blue irises61. The fact that not all disease associated MITF variants result in blue irises in humans could indicate that other variants in LD with the disease-causing MITF variant are responsible for blue irises, that depending on the location of the MITF variant, iris specific regulatory mechanisms are impacted, or that complex interactions among variants at the locus play a role.
The analysis reported in this study finds that variants in the vicinity of MITF increase deafness risk relative to wild-type alleles. We note that the penetrance of the risk haplotype for the deafness trait is modest with only 38% of dogs homozygous for the risk haplotype being phenotyped as clinically deaf in the combined data resource. Importantly, 12% of dogs homozygous for the low-risk haplotype were also tested as deaf (Table S8) with the majority of the 12% (five of six observations) being in the UK cohort of dogs. The UK cohort had an unusually low record of unilateral deaf dogs that may mean that some were classified along with bilateral deaf dogs as “deaf”. One deaf Australian dog that was homozygous TTG had other health issues (seizures) that may suggest a different disease phenotype. The combined CMH analysis (Table 3) detected genome-wide significant association on CFA8 (BICF2S23510367). These findings suggest that other loci or additional variants at this locus likely contribute to deafness in this breed.
Understanding the genetics of CSD in Dalmatian dogs has presented itself as a true challenge to numerous researchers. While many questions remain, we believe that results from this study open the door for genetic testing. In combination with currently applied selection criteria for breeding animals, selecting animals with the protective MITF haplotype will assist with reducing the number of deaf Dalmatians. Despite this new genetic tool, further studies investigating the underlying mechanisms and identifying functional variants will be crucial to eliminate CSD from this breed.
Materials and methods
Samples and genotyping
Blood or saliva samples of a total of 145 Dalmatians, comprising 86 hearing, 33 unilateral deaf and 26 bilateral deaf individuals were collected in Australia under University of Sydney Animal Ethics Committee (Approval numbers: N00/9-2009/3/5109; N00/10-2012/3/5837; and 2015/902) (Table S1). All experiments were performed in accordance with the recommendations of the Australian Code of the Care and Use of Animals for Scientific Purposes and reporting follows the recommendations in the ARRIVE guidelines. Genomic DNA was extracted from Performagene PG-100 buccal swabs (DNA Genotek, Canada) or EDTA whole-blood samples using the PureLink Genomic DNA Mini Kit (Invitrogen-Thermo Fisher Scientific, Waltham, Massachusetts, United States) following the manufacturers’ instructions. DNA quantity and quality was assessed using a NanoDrop2000 Spectrophotometer (Thermo Fisher, Waltham, Massachusetts, United States). DNA from blood spots applied to Whatman FTA cards was extracted by the genotyping provider (Neogen Inc, Nebraska USA). DNA samples of 139 individuals were genotyped using the Illumina Canine HD genotyping array (either 170 K or 220 K format) (Neogen Inc, Nebraska USA) and whole-genome sequences were generated for six animals (four bilateral deaf and two hearing) using 100 base paired-end Truseq libraries (Ramaciotti Centre for Functional Genomics) at 5–10× coverage on a HiSeq 2000. One bilateral deaf dog (Dog 3360) was sequenced on an Illumina XTen by the Kinghorn Cancer Centre using 100 base paired-end reads and a PCR-free library with approximate 30-fold genome coverage. Genotypes for array markers (canFam3) were extracted from alignments for the first six dogs with whole-genome sequencing data using Arraymaker62.
Phenotyping
Hearing status of Australian Dalmatian dogs was determined with brainstem auditory evoked response (BAER) testing either performed at the University of Sydney Veterinary Teaching Hospital Sydney following routine veterinary protocols or BAER results provided by the owner (referred to as “owner-reported”). Eye colour was noted in dogs tested at the University of Sydney Veterinary Teaching Hospital Sydney but otherwise not explicitly recorded. Attributes of Australian dogs, including deafness status, phenotyping location, and genotyping or sequencing strategy are reported in Table S1. Characteristics of publicly available samples are as previously recorded39.
Genome-wide association analysis
A GWAS of Australian data (N = 145) was conducted using quantitative association analysis in Plink (v1.90 64-bit)63. Data were coded to a quantitative phenotype score: control 1 (bilateral hearing), unilateral 1.5, deaf 2. Variants with a genotyping rate of < 0.1, strong deviation from Hardy–Weinberg expectation (p < 0.005), and a minor allele frequency (MAF) of < 0.1 were removed from further analysis. Genome-wide significance was ascertained by Bonferroni probability using the –adjust option in Plink63, with p-values < 0.05 considered as genome-wide significant. Top associated variants identified in this analysis were assessed against previously published associated markers.
Putative functional variant detection
To assess for the presence of potentially functional variants, all variants were called in the associated region on CFA20 using whole-genome sequence data from seven Australian Dalmatian dogs. Paired sequencing reads were processed relative to the canFam4 reference assembly using standard pipelines including the use of the Burrows-Wheeler Aligner and samtools mpileup for variant detection64,65,66, to take advantage of improvements to the reference assembly throughout the MITF region42. Putative functional variants were called by visualising the regional variant call file (vcf) as a custom track on the UCSC browser and making use of the Variant Annotation Integrator tool67,68. Where necessary, the Integrative Genomics Viewer (IGV)69 was used to visualize variants and sequence alignments relative to the Uppsala GSD 1.0 gene annotations track on the University of California Santa Cruz genome browser for canFam4. Variants unique to dogs that carried the MITF retrotransposon (as ascertained by heterozygosity for missense variants rs851676581 and rs851603213) were excluded from further consideration40. Remaining variants were manually assessed for potential functional implications, concentrating on variants located near, or within functionally relevant MITF transcripts. Manual inspection in the IGV was used to identify putative repeat element insertions within the same regions. Where available, minor allele frequencies for the identified variants were assessed using two published variant resources representing 772 (two Dalmatian) and 590 (two Dalmatian) dogs and canids70,71 after first using the University of Santa Cruz genome browser liftover tool to convert the variant coordinates from canFam4 to canFam3.
Genome-wide re-analysis of UK-USA data
Publicly available UK-USA Dalmatian genotyping array data (N = 304)39 were re-analysed using a quantitative analysis with the first component of multi-dimensional scaling fitted as a covariate to account for the geographic origins of the samples. Variants with a MAF < 0.05, a genotyping rate of < 0.1 and strong deviation from Hardy–Weinberg expectation (p < 0.005) were excluded from the analysis. Bilateral deaf dogs were coded as 2, unilateral as 1.5 and bilateral hearing as 1. Subsequently, the same dataset was subjected to a case–control CMH test for stratified data after using the cluster function in Plink to allocate samples to two predominantly geographical clusters. Deaf and unilateral dogs (N = 170) were coded as cases and bilateral hearing dogs were coded as controls (N = 134). Genome-wide significance was ascertained by Bonferroni probability using the adjusted option in Plink63, with adjusted p-values of < 0.05 considered genome-wide significant.
Expanded meta-analysis
Publicly available genotyping data were merged with array data of 139 Australian Dalmatians with hearing phenotypes. A linear association analysis was performed using Plink63 excluding genotypes with a MAF < 0.05, a genotyping rate < 0.1, and strong deviation from Hardy–Weinberg expectation (p < 0.005). Data were corrected for population stratification by fitting the first and second principal components (C1 and C2) from multi-dimensional scaling analysis as co-variates in the analysis. All data were recoded to a quantitative phenotype (scores: control 1, unilateral 1.5, deaf 2). Using identical filtering criteria for variants, the same expanded cohort of dogs was subjected to a case–control analysis using a CMH test for stratified data. Deaf and unilateral dogs (N = 224) were coded as cases and hearing dogs as controls (N = 216). The cluster function in Plink63 was used to allocate samples to three predominantly geographical clusters. Top associated variants were assessed against previously published associated markers and associated markers from the Australian cohort and vice versa.
Deafness associated risk haplotype analysis
The three most associated markers on CFA20 (BICF2G630233861, BICF2G630233888, BICF2G630233852) were used to perform a haplotype risk analysis. Marker information and corresponding haplotypes were extracted from arrays for all 443 dogs using Plink and a quantitative haplotype risk analysis performed. Deaf animals were scored as 2, unilateral deaf as 1.5 and hearing as 1. Animal hearing status was assessed against haplotypes with animals grouped as either homozygous for the reference haplotype, homozygous for the alternative haplotype, or heterozygous. The cohort of 139 Australian Dalmatians was analysed separately.
Deafness association of pseudogene
Dogs were assessed for the presence of the MITF retrotransposon as described before40. Information on sex was not available for all dogs included in this study, therefore markers in strong LD with the MITF retrotransposon were extracted and a quantitative association analysis was performed using Plink63 to assess the potential involvement of the MITF pseudogene on hearing. As relevant markers are only present on the 220 K genotyping array, dogs genotyped on the 170 K array were excluded from this analysis.
Ethics declaration
Recommendations from the Australian Code of the Care and Use of Animals for Scientific Purposes were strictly adhered to throughout this study. Research was conducted at The University of Sydney, under the Animal Ethics Committee approval no.: Approval numbers: N00/9-2009/3/5109; N00/10-2012/3/5837; and 2015/902. BAER testing was performed at The University Veterinary Teaching Hospital Camperdown following routine veterinary protocols. Reporting in this manuscript is in accordance with the ARRIVE guidelines.
Data availability
Whole genome sequencing data connected with this project are available via the European Nucleotide Archive https://www.ebi.ac.uk/ena/browser/view/PRJEB53145.
References
Strain, G. M. Deafness prevalence and pigmentation and gender associations in dog breeds at risk. Vet. J. 167, 23–32. https://doi.org/10.1016/s1090-0233(03)00104-7 (2004).
Wood, J. L. & Lakhani, K. H. Prevalence and prevention of deafness in the Dalmatian-assessing the effect of parental hearing status and gender using ordinary logistic and generalized random litter effect models. Vet. J. 154, 121–133. https://doi.org/10.1016/s1090-0233(97)80050-0 (1997).
Platt, S., Freeman, J., di Stefani, A., Wieczorek, L. & Henley, W. Prevalence of unilateral and bilateral deafness in border collies and association with phenotype. J. Vet. Intern. Med. 20, 1355–1362. https://doi.org/10.1892/0891-6640(2006)20[1355:pouabd]2.0.co;2 (2006).
De Risio, L. et al. Prevalence, heritability and genetic correlations of congenital sensorineural deafness and pigmentation phenotypes in the Border Collie. Vet. J. 188, 286–290. https://doi.org/10.1016/j.tvjl.2010.05.012 (2011).
De Risio, L., Freeman, J. & Lewis, T. Prevalence, heritability and genetic correlations of congenital sensorineural deafness and coat pigmentation phenotype in the English bull terrier. BMC Vet. Res. 12, 146. https://doi.org/10.1186/s12917-016-0777-6 (2016).
Marsh, O., Freeman, J., Pollard, D. & De Risio, L. Congenital sensorineural deafness in Australian Cattle dogs in the UK: Prevalence and association with phenotype. Vet. J. 274, 105711. https://doi.org/10.1016/j.tvjl.2021.105711 (2021).
Strain, G. M. Congenital sensorineural deafness in Dogo Argentino dogs: Prevalence and phenotype associations. Vet. Rec. 188, e299. https://doi.org/10.1002/vetr.299 (2021).
Lewis, T., Freeman, J. & De Risio, L. Decline in prevalence of congenital sensorineural deafness in Dalmatian dogs in the United Kingdom. J. Vet. Intern. Med. 34, 1524–1531. https://doi.org/10.1111/jvim.15776 (2020).
Famula, T. R., Oberbauer, A. M. & Sousa, C. A. A threshold model analysis of deafness in Dalmatians. Mamm. Genome 7, 650–653. https://doi.org/10.1007/s003359900199 (1996).
Famula, T. R., Oberbauer, A. M. & Sousa, C. A. Complex segregation analysis of deafness in Dalmatians. Am. J. Vet. Res. 61, 550–553. https://doi.org/10.2460/ajvr.2000.61.550 (2000).
Famula, T. R., Oberbauer, A. M. & Williams, D. C. Gender effects in hearing loss in Dalmatians. Prev. Vet. Med. 48, 15–24. https://doi.org/10.1016/s0167-5877(00)00183-5 (2001).
Cargill, E. J., Famula, T. R., Strain, G. M. & Murphy, K. E. Heritability and segregation analysis of deafness in US Dalmatians. Genetics 166, 1385–1393. https://doi.org/10.1534/genetics.166.3.1385 (2004).
Coppens, A. G., Kiss, R., Heizmann, C. W., Deltenre, P. & Poncelet, L. An original inner ear neuroepithelial degeneration in a deaf Rottweiler puppy. Hear Res. 161, 65–71. https://doi.org/10.1016/s0378-5955(01)00354-9 (2001).
Coppens, A. G., Gilbert-Gregory, S., Steinberg, S. A., Heizmann, C. & Poncelet, L. Inner ear histopathology in “nervous Pointer dogs” with severe hearing loss. Hear Res. 200, 51–62. https://doi.org/10.1016/j.heares.2004.08.019 (2005).
Steel, K. P. & Bock, G. R. Hereditary inner-ear abnormalities in animals. Relationships with human abnormalities. Arch Otolaryngol. 109, 22–29. https://doi.org/10.1001/archotol.1983.00800150026005 (1983).
Strain, G. M. Canine deafness. Vet. Clin. N. Am. Small Anim. Pract. 42, 1209–1224. https://doi.org/10.1016/j.cvsm.2012.08.010 (2012).
Sampaio, A. L. et al. Histopathological morphometric study of cochleosaccular dysplasia in Dalmatian dogs. Int. J. Pediatr. Otorhinolaryngol. 74, 934–938. https://doi.org/10.1016/j.ijporl.2010.05.020 (2010).
Hiraide, F. & Paparella, M. M. Histopathology of the temporal bones of deaf dogs. Auris Nasus Larynx 15, 97–104. https://doi.org/10.1016/s0385-8146(88)80014-2 (1988).
Igarashi, M., Alford, B. R., Cohn, A. M., Saito, R. & Watanabe, T. Inner ear anomalies in dogs. Ann. Otol. Rhinol. Laryngol. 81, 249–255. https://doi.org/10.1177/000348947208100211 (1972).
Sommerlad, S. F. et al. Prevalence of congenital hereditary sensorineural deafness in Australian Cattle Dogs and associations with coat characteristics and sex. BMC Vet. Res. 8, 202. https://doi.org/10.1186/1746-6148-8-202 (2012).
Strain, G. M. Congenital deafness and its recognition. Vet. Clin. N. Am. Small Anim. Pract. 29, 895–907. https://doi.org/10.1016/s0195-5616(99)50079-x (1999).
Strain, G. M. The genetics of deafness in domestic animals. Front. Vet. Sci. 2, 29. https://doi.org/10.3389/fvets.2015.00029 (2015).
Rak, S. G. & Distl, O. Congenital sensorineural deafness in dogs: A molecular genetic approach toward unravelling the responsible genes. Vet. J. 169, 188–196. https://doi.org/10.1016/j.tvjl.2004.01.015 (2005).
Strain, G. M. et al. Brainstem auditory-evoked potential assessment of congenital deafness in Dalmatians: Associations with phenotypic markers. J. Vet. Intern. Med. 6, 175–182. https://doi.org/10.1111/j.1939-1676.1992.tb00333.x (1992).
Juraschko, K., Meyer-Lindenberg, A., Nolte, I. & Distl, O. A regressive model analysis of congenital sensorineural deafness in German Dalmatian dogs. Mamm. Genome 14, 547–554. https://doi.org/10.1007/s00335-002-3033-2 (2003).
Karlsson, E. K. et al. Efficient mapping of mendelian traits in dogs through genome-wide association. Nat. Genet. 39, 1321–1328. https://doi.org/10.1038/ng.2007.10 (2007).
Xu, F., Shan, S., Sommerlad, S., Seddon, J. M. & Brenig, B. A missense mutation in the KLF7 gene is a potential candidate variant for congenital deafness in Australian stumpy tail cattle dogs. Genes (Basel). https://doi.org/10.3390/genes12040467 (2021).
Guevar, J., Olby, N. J., Meurs, K. M., Yost, O. & Friedenberg, S. G. Deafness and vestibular dysfunction in a Doberman Pinscher puppy associated with a mutation in the PTPRQ gene. J. Vet. Intern. Med. 32, 665–669. https://doi.org/10.1111/jvim.15060 (2018).
Webb, A. A., Ruhe, A. L. & Neff, M. W. A missense mutation in MYO7A is associated with bilateral deafness and vestibular dysfunction in the Doberman pinscher breed. Can. J. Vet. Res. 83, 142–148 (2019).
Juraschko, K., Meyer-Lindenberg, A., Nolte, I. & Distl, O. Analysis of systematic effects on congenital sensorineural deafness in German Dalmatian dogs. Vet. J. 166, 164–169. https://doi.org/10.1016/s1090-0233(02)00256-3 (2003).
Hytönen, M. K. et al. Missense variant in LOXHD1 is associated with canine nonsyndromic hearing loss. Hum. Genet. 140, 1611–1618. https://doi.org/10.1007/s00439-021-02286-z (2021).
Blum, M. & Distl, O. Phenotypic trends and breeding values for canine congenital sensorineural deafness in Dalmatian dogs. Berl. Munch. Tierarztl. Wochenschr. 127, 70–76 (2014).
Mieskes, K. & Distl, O. Elimination of TMC1 and TMIE as candidates for hereditary non-syndromic deafness in Dalmatian dogs. Anim. Genet. 37, 519–521. https://doi.org/10.1111/j.1365-2052.2006.01497.x (2006).
Mieskes, K. & Distl, O. Evaluation of ESPN, MYO3A, SLC26A5 and USH1C as candidates for hereditary non-syndromic deafness (congenital sensorineural deafness) in Dalmatian dogs. Anim. Genet. 38, 533–534. https://doi.org/10.1111/j.1365-2052.2007.01625.x (2007).
Kluth, S. & Distl, O. Congenital sensorineural deafness in dalmatian dogs associated with quantitative trait loci. PLoS One 8, e80642 (2013). https://doi.org/10.1371/journal.pone.0080642
Vasiliadis, D., Metzger, J. & Distl, O. Demographic assessment of the Dalmatian dog—Effective population size, linkage disequilibrium and inbreeding coefficients. Canine Med. Genet. 7, 3. https://doi.org/10.1186/s40575-020-00082-y (2020).
Stritzel, S., Wöhlke, A. & Distl, O. Elimination of SILV as a candidate for congenital sensorineural deafness in Dalmatian dogs. Anim. Genet. 38, 662–663. https://doi.org/10.1111/j.1365-2052.2007.01660.x (2007).
Stritzel, S., Wöhlke, A. & Distl, O. A role of the microphthalmia-associated transcription factor in congenital sensorineural deafness and eye pigmentation in Dalmatian dogs. J. Anim. Breed. Genet. 126, 59–62. https://doi.org/10.1111/j.1439-0388.2008.00761.x (2009).
Hayward, J. J. et al. A genome-wide association study of deafness in three canine breeds. PLoS ONE 15, e0232900. https://doi.org/10.1371/journal.pone.0232900 (2020).
Brancalion, L., Haase, B., Willet, C. E. & Wade, C. M. Sequence variants of the canine melanocyte inducing transcription factor (MITF) locus reveal a common MITF-A processed pseudogene. Anim. Genet. https://doi.org/10.1111/age.13106 (2021).
Tsai, K. L., Evans, J. M., Noorai, R. E., Starr-Moss, A. N. & Clark, L. A. Novel Y chromosome retrocopies in canids revealed through a genome-wide association study for sex. Genes (Basel). https://doi.org/10.3390/genes10040320 (2019).
Wang, C. et al. A novel canine reference genome resolves genomic architecture and uncovers transcript complexity. Commun. Biol. 4, 185. https://doi.org/10.1038/s42003-021-01698-x (2021).
Lindblad-Toh, K. et al. Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature 438, 803–819. https://doi.org/10.1038/nature04338 (2005).
Liao, J. et al. Impact of measurement error on testing genetic association with quantitative traits. PLoS ONE 9, e87044. https://doi.org/10.1371/journal.pone.0087044 (2014).
Barendse, W. The effect of measurement error of phenotypes on genome wide association studies. BMC Genomics 12, 232. https://doi.org/10.1186/1471-2164-12-232 (2011).
Seddon, J. M. et al. Deafness in Australian Cattle Dogs associated to QTL on chromosome 20 in genome-wide association study analyses. Anim. Genet. https://doi.org/10.1111/age.13115 (2021).
Tian, X. et al. Identification and characterization of microRNAs in white and brown alpaca skin. BMC Genom. 13, 555. https://doi.org/10.1186/1471-2164-13-555 (2012).
Zhu, Z. et al. MicroRNA-25 functions in regulation of pigmentation by targeting the transcription factor MITF in Alpaca (Lama pacos) skin melanocytes. Domest Anim. Endocrinol. 38, 200–209. https://doi.org/10.1016/j.domaniend.2009.10.004 (2010).
Yang, S. et al. Identification of a novel microRNA important for melanogenesis in alpaca (Vicugna pacos). J Anim. Sci. 93, 1622–1631. https://doi.org/10.2527/jas.2014-8404 (2015).
Baranowska Körberg, I. et al. A simple repeat polymorphism in the MITF-M promoter is a key regulator of white spotting in dogs. PLoS ONE 9, e104363. https://doi.org/10.1371/journal.pone.0104363 (2014).
Thurber, A. E. et al. Inverse expression states of the BRN2 and MITF transcription factors in melanoma spheres and tumour xenografts regulate the NOTCH pathway. Oncogene 30, 3036–3048. https://doi.org/10.1038/onc.2011.33 (2011).
Goodall, J. et al. Brn-2 represses microphthalmia-associated transcription factor expression and marks a distinct subpopulation of microphthalmia-associated transcription factor-negative melanoma cells. Cancer Res. 68, 7788–7794. https://doi.org/10.1158/0008-5472.CAN-08-1053 (2008).
Maruotti, J., Thein, T., Zack, D. J. & Esumi, N. MITF-M, a “melanocyte-specific” isoform, is expressed in the adult retinal pigment epithelium. Pigment. Cell Melanoma Res. 25, 641–644. https://doi.org/10.1111/j.1755-148X.2012.01033.x (2012).
Pogenberg, V. et al. Restricted leucine zipper dimerization and specificity of DNA recognition of the melanocyte master regulator MITF. Genes Dev. 26, 2647–2658. https://doi.org/10.1101/gad.198192.112 (2012).
Gratton, M. A., Schmiedt, R. A. & Schulte, B. A. Age-related decreases in endocochlear potential are associated with vascular abnormalities in the stria vascularis. Hear Res. 94, 116–124. https://doi.org/10.1016/0378-5955(96)00011-1 (1996).
Bergsma, D. R. & Brown, K. S. White fur, blue eyes, and deafness in the domestic cat. J. Hered. 62, 171–185. https://doi.org/10.1093/jhered/62.3.171 (1971).
Hauswirth, R. et al. Mutations in MITF and PAX3 cause “splashed white” and other white spotting phenotypes in horses. PLoS Genet. 8, e1002653. https://doi.org/10.1371/journal.pgen.1002653 (2012).
Araki, S. et al. Degeneration of the stria vascularis during development in melanocyte-deficient mutant rats (Ws/Ws rats). Eur. Arch Otorhinolaryngol. 259, 309–315. https://doi.org/10.1007/s00405-002-0480-z (2002).
Sugiura, A. & Hilding, D. A. Cochleo-saccular degeneration in Hedlund white mink. Acta Otolaryngol. 69, 126–137. https://doi.org/10.3109/00016487009123345 (1970).
Du, Y. et al. Degeneration of saccular hair cells caused by MITF gene mutation. Neural Dev. 14, 1. https://doi.org/10.1186/s13064-019-0126-0 (2019).
Sturm, R. A. & Larsson, M. Genetics of human iris colour and patterns. Pigment. Cell Melanoma Res. 22, 544–562. https://doi.org/10.1111/j.1755-148X.2009.00606.x (2009).
Willet, C. E., Haase, B., Charleston, M. A. & Wade, C. M. Simple, rapid and accurate genotyping-by-sequencing from aligned whole genomes with ArrayMaker. Bioinformatics 31, 599–601. https://doi.org/10.1093/bioinformatics/btu691 (2015).
Purcell, S. et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am J. Hum. Genet. 81, 559–575. https://doi.org/10.1086/519795 (2007).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760. https://doi.org/10.1093/bioinformatics/btp324 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079. https://doi.org/10.1093/bioinformatics/btp352 (2009).
Hoeppner, M. P. et al. An improved canine genome and a comprehensive catalogue of coding genes and non-coding transcripts. PLoS ONE 9, e91172. https://doi.org/10.1371/journal.pone.0091172 (2014).
Chew, T., Willet, C. E., Haase, B. & Wade, C. M. Genomic characterization of external morphology traits in kelpies does not support common ancestry with the Australian Dingo. Genes (Basel). https://doi.org/10.3390/genes10050337 (2019).
Hinrichs, A. S. et al. UCSC data integrator and variant annotation integrator. Bioinformatics 32, 1430–1432. https://doi.org/10.1093/bioinformatics/btv766 (2016).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): High-performance genomics data visualization and exploration. Brief Bioinform. 14, 178–192. https://doi.org/10.1093/bib/bbs017 (2013).
Plassais, J. et al. Whole genome sequencing of canids reveals genomic regions under selection and variants influencing morphology. Nat. Commun. 10, 1489. https://doi.org/10.1038/s41467-019-09373-w (2019).
Jagannathan, V., Drögemüller, C., Leeb, T., (Dog Biomedical Variant Database Consortium). A comprehensive biomedical variant catalogue based on whole genome sequences of 582 dogs and eight wolves. Anim. Genet. 50, 695–704. https://doi.org/10.1111/age.12834 (2019).
Acknowledgements
The authors would like to thank the owners of animals included in this study for providing samples and information on the hearing status. We gratefully acknowledge the assistance of the Canine Health Foundation (02157-MOU), The University of Sydney, and the Dalmatian Club of NSW who have provided financial or in-kind support to this work. This project was conducted with technical assistance from students and colleagues: Diane van Rooy, Laura Bunbury-Cruikshank, Hamutal Mazrier, Jennifer Liu, Rhiarn Hoban and Simone Carter. The authors acknowledge the Sydney Informatics Hub, a Core Research Facility, of the University of Sydney for access to the High-Performance Computing cluster ‘Artemis’.
Author information
Authors and Affiliations
Contributions
C.M.W., C.E.W., G.S. and T.C. analysed the data. B.H. and C.M.W. conceptualised the project, contributed to the progression of the aims and wrote the manuscript. G.C. performed BAER testing of animals assessed at the University Veterinary Teaching Hospital Sydney.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Haase, B., Willet, C.E., Chew, T. et al. De-novo and genome-wide meta-analyses identify a risk haplotype for congenital sensorineural deafness in Dalmatian dogs. Sci Rep 12, 15439 (2022). https://doi.org/10.1038/s41598-022-19535-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-19535-4
- Springer Nature Limited