Abstract
Recent outbreaks of Zika virus (ZIKV) infection have highlighted the need for a better understanding of ZIKV-specific immune responses. The ZIKV envelope glycoprotein (EZIKV) is the most abundant protein on the virus surface and it is the main target of the protective immune response. EZIKV protein contains the central domain (EDI), a dimerization domain containing the fusion peptide (EDII), and a domain that binds to the cell surface receptor (EDIII). In this study, we performed a systematic comparison of the specific immune response induced by different EZIKV recombinant proteins (EZIKV, EDI/IIZIKV or EDIIIZIKV) in two mice strains. Immunization induced high titers of E-specific antibodies which recognized ZIKV-infected cells and neutralized the virus. Furthermore, immunization with EZIKV, EDI/IIZIKV and EDIIIZIKV proteins induced specific IFNγ-producing cells and polyfunctional CD4+ and CD8+ T cells. Finally, we identified 4 peptides present in the envelope protein (E1–20, E51–70, E351–370 and E361–380), capable of inducing a cellular immune response to the H-2Kd and H-2Kb haplotypes. In summary, our work provides a detailed assessment of the immune responses induced after immunization with different regions of the ZIKV envelope protein.
Similar content being viewed by others
Introduction
Research on the immune response to the Zika virus (ZIKV), a mosquito-borne flavivirus, has increased after recent outbreaks1,2,3. Unlike other flaviviruses, ZIKV transmission was also observed through non-vector transmission (sexual, transfusional, and vertical)4,5. ZIKV infection in pregnant women has been associated with congenital malformations (brain calcification, microcephaly, and spontaneous abortion), characterizing the congenital Zika syndrome (CZS)6,7,8,9, while in adults it is associated with Guillain-Barré syndrome (GBS)10,11. The rapid global spread of ZIKV and the suspected association with serious neurological implications have led to the urgent need for an effective vaccine and specific treatment against the virus. Even with scientific efforts, little is known about the ZIKV-specific immune response and there are still no licensed therapeutic or prophylactic vaccines against ZIKV.
ZIKV has an 11 kb positive-sense single-stranded RNA (ssRNA) genome that encodes a single polyprotein that is cleaved into three structural proteins (Capsid (C), Premembrane/Membrane (prM/M) and Envelope (E)) and seven non-structural proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B, and NS5) involved in virus replication and assembly12,13. The envelope protein mediates viral assembly, binding to cell receptors and is essential for the subsequent fusion of the membrane involved in virus entry into the target cell14. Similar to other flaviviruses, the ZIKV E protein contains three distinct domains: the central domain (EDI), the domain responsible for dimerization that contains the fusion peptide (EDII), and the domain that binds to the cell surface receptor (EDIII)15.
Regarding the humoral immune response induced by flaviviruses, several studies have shown that protein E is highly immunogenic. Furthermore, protein E is the main target of several ZIKV-specific neutralizing antibodies16,17 that also confer protection in animal models of infection18,19. Several E-specific monoclonal antibodies (mAbs) inhibited ZIKV infection19,20,21,22. In addition, passive transfer of E-specific mAb reduced vertical transmission and mortality in mice17. Although antibodies are the main correlates of protection against ZIKV infection, T cell immunity also plays an important role in controlling virus replication23. It has already been demonstrated that the absence of CD8+ T cells during ZIKV infection is capable of increasing mortality in mice24. CD4+ T cells also participate in the generation of protective immunity, since their depletion reduced the induction of anti-ZIKV antibodies25,26 and CD8+ T cell responses27.
In this study, we investigated the induction of humoral and cellular immune responses after immunization of BALB/c and C57Bl/6 mice with the ZIKV-envelope protein (EZIKV) and its domains (EDI/IIZIKV and EDIIIZIKV). We observed that immunization with EDI/IIZIKV and EDIIIZIKV proteins induced high titers of specific antibodies, which recognized ZIKV-infected cells and neutralized the virus. In addition, immunization with the proteins was able to induce specific IFNγ-producing cells and polyfunctional CD4+ and CD8+ T cell responses. We also mapped immunodominant epitopes in ZIKV-envelope region, and identified four peptides (E1–20, E51–70, E351–370 and E361–380) capable of inducing specific T cells to the H-2Kd and H-2Kb haplotypes.
Results
Production of recombinant envelope proteins
After alignment of the 69 ZIKV isolates we observed 96.75% of homology among the amino acid sequences. Additionally, across the different isolates, it was possible to identify 13 different sequences with at least one amino acid mutation corresponding to 18.9% of the ZIKV isolates. The artificial gene corresponding to the consensus sequence of the ZIKV envelope was synthetized (Supplementary Table 1) and cloned to produce the entire ectodomain of the envelope recombinant protein (EZIKV) (amino acids 291–690) and its domains EDI/IIZIKV (aa 291–600) and EDIIIZIKV (aa 601–690) (Supplementary Fig. 1a). The recombinant EZIKV, EDI/IIZIKV and EDIIIZIKV were purified by affinity chromatography (Supplementary Fig. 1b) and further recognized by anti-His tag (Supplementary Fig. 1c). We also performed a Dot Blot with the expressed proteins using an anti-flavivirus 4G2 monoclonal antibody, which recognizes a conformational epitope present in the E protein domain II fusion loop. The 4G2 specifically recognized EZIKV and EDI/IIZIKV proteins, but not the EDIIIZIKV domain, suggesting that the expressed proteins retained the correct conformation (Supplementary Fig. 1d).
Immunization with ZIKV envelope proteins induces a potent specific humoral immune response in BALB/c and C57Bl/6 mice
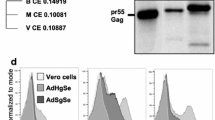
Next, we assessed the immunogenicity of the subunit vaccines in BALB/c and C57Bl/6 mice. For this purpose, mice received two doses with an equimolar amount of EZIKV, EDI/IIZIKV or EDIIIZIKV administered subcutaneously in the presence of adjuvant poly (I:C) (Fig. 1a). Fourteen days after each dose, sera were tested for reactivity against ZIKV envelope proteins. Sera from all groups immunized with the recombinant proteins presented antigen-specific antibodies in both BALB/c (Fig. 1b) and C57Bl/6 (Fig. 1c) strains. Furthermore, after the boost, the antibody titers increased significantly. A head-to-head comparison after the second dose revealed that BALB/c mice immunized with EZIKV or EDIIIZIKV presented slightly higher antibody titers when compared to animals that received EDI/IIZIKV (Supplementary Fig. 2a). However, no significant differences were observed between antibody titers in C57Bl/6 mice that received EZIKV, EDI/IIZIKV or EDIIIZIKV. The humoral response against different domains was also analyzed by ELISA (Supplementary Fig. 2b). Anti-EDI/IIZIKV and anti-EDIIIZIKV humoral immune responses showed high specificity, while anti-EZIKV antibodies reacted with EDI/IIZIKV and EDIIIZIKV proteins. In addition, an immunofluorescence assay (IFA) showed that sera from mice immunized with different ZIKV envelope proteins recognized ZIKV-infected cells (Fig. 1d). In contrast, the control group immunized with only poly (I:C) did not produce specific antibodies against the ZIKV envelope proteins (Fig. 1b,c) and was unable to recognize ZIKV-infected cells (Fig. 1d).
Specific humoral immune response elicited after immunization with EZIKV, EDI/IIZIKV or EDIIIZIKV recombinant protein in two different mouse strains. (a) Immunization Strategy (created with BioRender.com). BALB/c or C57Bl/6 mice (n = 3 control groups and n = 4 experimental groups) were immunized subcutaneously twice with equimolar amounts of the EZIKV, EDI/IIZIKV or EDIIIZIKV combined with 50 μg poly(I:C). Control groups received only poly(I:C). Mice were bled 14 days after each dose to evaluate humoral response. (b, c) Total specific-IgG antibody titers on a logarithm scale (Log10) in the sera of (b) BALB/c or (c) C57Bl/6 mice. Empty symbols represent pre boost serum and filled symbols represent post boost serum. Post boost sera was inactivated in order to assess their ability to (d) recognize ZIKV-infected Vero cells (MOI = 0.1) or (e) neutralize ZIKV. (d) For IFA assay, mouse serum and goat anti-mouse IgG conjugated with fluorescein isothiocyanate was used as primary and secondary antibodies respectively. (e) For PRNT, sera were incubated with 100 PFU of ZIKV and the neutralization capacity were represented by 50% of viral neutralization (NT50). Data represent mean ± SEM of 4 independent experiments. Statistical significance was measured by One-way ANOVA followed by Tukey’s post hoc test, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Immunization with different envelope proteins induces neutralizing antibodies against ZIKV infection
To evaluate the quality of the antibodies generated, we performed standard plaque reduction neutralization testing (PRNT). In BALB/c mice, we observed that the serum of all immunized groups reduced ZIKV infection (Fig. 1e). However, we can observe that the antibody titers that promoted 50% of viral neutralization (NT50) were higher in the group immunized with EZIKV or EDIIIZIKV in the presence of poly (I:C). The lowest NT50 values were observed in the groups immunized with EDI/IIZIKV. For C57Bl/6 mice, we observed a similar profile, but immunization with EDIIIZIKV led to a higher production of neutralizing antibodies when compared to immunization with EZIKV. In contrast, the naïve or adjuvant group did not display significant neutralizing antibodies against the virus. So far, our data demonstrate that the EDIII component induces the most robust humoral immune response against ZIKV.
Subunit vaccines induced IFNγ-producing cells against recombinant ZIKV envelope proteins in different mouse strains
Next, we evaluated whether immunization with EZIKV, EDI/IIZIKV and EDIIIZIKV mixed with poly (I:C) in BALB/c and C57Bl/6 mice would induce cellular-mediated immunity. Splenocytes harvested fifteen days after boost were incubated with recombinant ZIKV envelope proteins to assess specific cytokine production. Figure 2 shows IFNγ-producing cells by ELISpot. In BALB/c (Fig. 2a) or C57Bl/6 (Fig. 2b) splenocytes, we observed that the group that received EZIKV + poly (I:C) presented IFNγ-producing cells when stimulated with all recombinant ZIKV proteins. On the contrary, splenocytes from mice immunized with EDI/IIZIKV + poly (I:C) or EDIIIZIKV + poly (I:C) only induced IFNγ-producing cells against EZIKV and EDI/IIZIKV or against EZIKV and EDIIIZIKV, respectively. We also observed a lower number of IFNγ-producing cells in splenocytes from BALB/c mice stimulated with EDI/IIZIKV when compared to the entire protein or domain III (Fig. 2a). Moreover, BALB/c mice immunized with adjuvanted EDI/IIZIKV or EDIIIZIKV presented higher number of IFNγ-producing cells against EZIKV when compared to the correlated domain used in immunization (Fig. 2a). Furthermore, the administration of poly (I:C) alone did not induce IFNγ-producing cells.
Specific IFNγ-producing cells after immunization with ZIKV-envelope proteins. Analysis of the specific cellular immune response after immunization of (a) BALB/c or (b) C57Bl/6 mice as described in Fig. 1a. Fifteen days after the boost, the splenocytes were cultured in the presence of equimolar amount of recombinant envelope proteins for 18 h to evaluate the number of IFN-γ producing cells by ELISpot assay. SFU: spot forming units. Statistical significance was measured by Two-way ANOVA followed by Tukey’s post hoc test, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Data represent mean ± SEM of 4 independent experiments.
T-cell epitope coverage
To evaluate the breadth of T cell responses and map the coverage of ZIKV-derived epitopes, a total of 39 peptides (20 amino acids overlapping 12-mer) were synthetized comprising the ZIKV E protein sequence (aa 291–690) conserved among 69 ZIKV isolates (GenBank accession numbers available at Supplementary Table 1). An optimized matrix containing 10 peptide pools was generated using Deconvolute This! Software (Supplementary Table 2)28. Splenocytes from BALB/c immunized with EZIKV + poly (I:C) presented IFNγ-producing cells to peptides presented in pools 1, 5, 6 and 8 (Fig. 3a). In addition to those covered by the BALB/c strain, C57Bl6 mice that received EZIKV + poly (I:C) also showed an IFNγ response against pool 4 (Fig. 3b). BALB/c mice immunized with EDI/IIZIKV + poly (I:C) presented a higher number of IFNγ-producing cells when stimulated by pools 1, 6, and 8 (Fig. 3a) but also against pools 2, 4, 7 and 10. On the other hand, IFNγ response by the same subunit vaccine in C57Bl/6 mice was exclusively directed to peptides present in pools 1, 6, and 8 (Fig. 3b). In both mouse strains, immunization with EDIIIZIKV + poly (I:C) induced an IFNγ-response against pools 5 and 8 (Fig. 3a,b).
Mapping of T cell epitopes after immunization with recombinant ZIKV envelope proteins. Analysis of the specific cellular immune response after immunization of (a, c) BALB/c or (b, d) C57Bl/6 mice as described in Fig. 1a. Fifteen after the second dose, the spleen of each animal was removed and the splenocytes were cultured in the presence of 10 mg/mL of the (a, b) pool of ZIKV peptides or (c, d) individual peptides to evaluate the number of IFNγ-producing cells by ELISpot assay. SFU: spot forming units. Statistical significance was measured by Two-way ANOVA followed by Tukey’s post hoc test, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Data represent mean ± SEM of 3 independent experiments.
From those results, we selected a total of 9 out of 39 potential peptides for further evaluation. ZIKV peptides from ectodomain I/II: (ZIKV 1 (E1–20), ZIKV 2 (E11–30), ZIKV 5 (E41–60), ZIKV 6 (E51–70), ZIKV 25 (E241–260) and ZIKV 27 (E261–280)) and ectodomain III: (ZIKV 33 (E321–340), ZIKV 36 (E351–370) and ZIKV 37 (E361–380)) were individually tested (Fig. 3c,d). Notably, immunization with EZIKV + poly (I:C) induced IFNγ-producing cells against two peptides present in domain I/II (ZIKV 1 (E1–20) and ZIKV 6 (E51–70)) and two present in domain III (ZIKV 36 (E351–370) and ZIKV 37 (E361–380)) in both BALB/c (Fig. 3c) and C57Bl/c (Fig. 3d) mice, albeit with a higher magnitude in C57Bl/6 mice. Furthermore, the same peptides induced responses after immunization with the respective domains. In mice immunized with EDI/IIZIKV + poly (I:C) or EDIIIZIKV + poly (I:C), the response was mainly directed against ZIKV 1 (E1–20) and ZIKV 6 (E51–70) or ZIKV 36 (E351–370) and ZIKV 37 (E361–380), respectively. The most immunogenic peptides in the ZIKV envelope amino acid sequence, ZIKV 1 (E1–20), ZIKV 6 (E51–70), ZIKV 36 (E351–370) and ZIKV 37 (E361–380) are represented in Supplementary Fig. 3a.
Subunit vaccines induce specific T cells that proliferate and produce pro-inflammatory cytokines
Next, we sought to evaluate whether immunization with different the subunit vaccines induce Env-specific CD4+ and CD8+ T cells able to proliferate (Fig. 4) or produce IFNγ and/or TNFα (Figs. 5 and 6) (representative gating strategies in Supplementary Fig. 3b). In BALB/c mice, immunization with EZIKV induced proliferation of CD4+ (Fig. 4a) and CD8+ (Fig. 4b) T cells against the proteins EZIKV, EDI/IIZIKV and EDIIIZIKV, as well as against the peptides present in domain I/II (ZIKV 1 (E1–20) and ZIKV 6 (E51–70)) and domain III (ZIKV 36 (E351–370) and ZIKV 37 (E361–380)). The proliferation of CD4+ and CD8+ T cells in mice immunized with the domains (EDI/IIZIKV and EDIIIZIKV) was specific, i.e., targeted to the EZIKV protein and to the domain used in immunization. The same phenomenon was observed for CD4+ (Fig. 4c) and CD8+ (Fig. 4d) T cell proliferation in C57BL/6 mice. It is worth mentioning that the frequency of proliferating CD8+ T cells was higher in C57Bl/6 (Fig. 4d) when compared to the BALB/c strain (Fig. 4b).
Immunization with recombinant ZIKV envelope proteins induces proliferation of specific CD4+ and CD8+ T cells. Analysis of CD4+ and CD8 + T cell proliferation after immunization of (a, b) BALB/c or (b, d) C57Bl/6 mice as described in Fig. 1a. Fifteen days after the second dose, the spleen of each animal was removed and the splenocytes were labeled with CFSE (1.25 μM) and cultured in the presence of equimolar amounts of recombinant proteins or 5 μg/mL of the individual peptides for 5 days. After labeling with fluorochrome-conjugated anti-CD3, -CD4 and -CD8, cells were analyzed by flow cytometry (representative gating strategies shown in Supplementary Fig. 3b). Initially, a gate was performed on CD3+ cells (T lymphocytes), followed by gates on CD4+ and CD8+ populations. Within the two T cells subpopulations (a, c) CD3+CD4+ and (b, d) CD3+CD8+, the decrease in CFSE fluorescence intensity was evaluated. The frequency of cell proliferation was calculated by subtracting the values from the culture of unstimulated cells. Statistical significance was measured by Two-way ANOVA followed by Tukey’s post hoc test, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Data represent mean ± SEM of 2 independent experiments.
Immunization with recombinant ZIKV envelope proteins induces polyfunctional CD4+ T cells. Analysis of polyfunctional cells after immunization of (a) BALB/c or (b) C57Bl/6 mice as described in Fig. 1a. Fifteen days after the second dose, the spleen of each animal was removed and cultured in the presence of equimolar amounts of recombinant proteins or 5 μg/mL of the individual peptides. For the detection of cytokine-producing T cells, the cells were restimulated on the 4th day for 12 h in the presence of recombinant proteins, anti-CD28 and brefeldin A. Cells were stained with anti-CD3 and -CD4, then permeabilized and labeled for intracellular cytokines. After selecting the T cell populations that produce cytokines (representative gating strategies are shown in Supplementary Fig. 3b), a Boolean combination was created using the FlowJo software to determine the frequency of each response based on all possible combinations of CD4+ T cell cytokine producers. Heatmap was used to determine the frequency of CD4+ T cells that produce IFNγ and TNFα when stimulated with the different proteins (columns). The frequency of CD4 + T cells that produce cytokines was calculated by subtracting the values from the unstimulated cell culture.
Immunization with recombinant ZIKV envelope proteins induces polyfunctional CD8+ T cells. Analysis of polyfunctional cells after immunization of (a) BALB/c or (b) C57Bl/6 mice as described in Fig. 1a. Fifteen days after the second dose, the spleen of each animal was removed and cultured in the presence of equimolar amounts of recombinant proteins or 5 μg/mL of the individual peptides. For the detection of cytokine-producing T cells, the cells were restimulated on the 4th day for 12 h in the presence of recombinant proteins, anti-CD28 and brefeldin A. Cells were stained with anti-CD3 and -CD8, then permeabilized and labeled for intracellular cytokines. After determining the populations of T cells that produce cytokines (representative gating strategies are shown in Supplementary Fig. 3b, a Boolean combination was created using the FlowJo software to determine the frequency of each response based on all possible combinations of CD8+ T cells cytokine producers. Heatmap was used to determine the frequency of CD8+ T cells that produce IFNγ and TNFα when stimulated with the different proteins (columns). The frequency of cells that produce cytokines was calculated by subtracting the values from the unstimulated cell culture.
Subsequently, we analyzed the cytokine profile of specific T lymphocytes. In both BALB/c (Figs. 5a, 6a and Fig. S4a) and C57Bl/6 (Figs. 5b, 6b and Fig. S4b) mice, immunization with EZIKV induced CD4+ and CD8+ T cells able to produce IFNγ and TNFα alone or simultaneously against all the stimuli (EZIKV, EDI/IIZIKV, EDIIIZIKV recombinant proteins; peptides ZIKV 1 (E1–20), ZIKV 6 (E51–70), ZIKV 36 (E351–370) and ZIKV 37 (E361–380)). As expected, immunization with the EDI/IIZIKV and EDIIIZIKV domains was able to induce a polyfunctional response only against the recombinant protein EZIKV or the specific domains used in the immunization. In contrast, the group immunized with the adjuvant alone did not induce a significant frequency of CD4+ and CD8+ T cells that proliferated or produced cytokines. These data suggest that immunization with ZIKV envelope proteins was able to induce specific T cell responses. Furthermore, this result suggests the ability of ZIKV 1 (E1–20), ZIKV 6 (E51–70), ZIKV 36 (E351–370) and ZIKV 37 (E361–380) peptides to bind to H-2Kd and H-2Kb haplotypes and induce T cell immunity.
Discussion
Recent outbreaks of ZIKV and the possible sequelae due to neurological morbidity in newborns and adults1,3 led to substantial changes in public health policies. There is no vaccine or treatment against the virus, highlighting the need for a better understanding of specific immunity. The ZIKV envelope protein is essential for virus entry into cells and is the main antigen that triggers host immune responses14. Here, we evaluated the ability of the recombinant protein EZIKV and its domains (EDI/IIZIKV and EDIIIZIKV) to induce humoral and cellular immune responses in two different mouse strains. C57Bl/6 and BALB/c mice were immunized with equimolar amounts of the recombinant proteins in the presence of the adjuvant poly (I:C).
The development of antibodies is considered fundamental against viral infections29. In flaviviruses, E protein, prM, and NS1 are the main targets of the antibody response30. We observed that the proteins EZIKV, EDI/IIZIKV and EDIIIZIKV were highly immunogenic, inducing specific antibodies that neutralized the virus. However, antibodies induced after boost with EZIKV or EDIIIZIKV displayed greater neutralizing capacity when compared to anti-EDI/IIZIKV antibodies. Previous studies demonstrate that domain III of the envelope protein is the main target for neutralizing antibodies 20,21,31,32,33,34,35,36. In addition, domain III does not induce antibody-dependent enhancement (ADE) and protected mice after challenge with ZIKV33,35. Furthermore, we observed that antibodies generated against immunization with the recombinant protein EZIKV were able to recognize the EDI/IIZIKV and EDIIIZIKV domains, with higher titers against domain I/II when compared to domain III. Sera from animals immunized with EDI/IIZIKV or EDIIIZIKV specifically recognized the same administered protein as well as the entire EZIKV protein but were unable to recognize the distinct domain. Within the E-specific response, the relative proportion of DI/II versus DIII antibodies showed considerable variability in serological studies with West Nile and Dengue (DENV) viruses, with an overall prevalence of antibodies against domains I/II34,37,38,39,40.
During ZIKV infection, although there is a prevalence of antibodies against domains I/II that reach a peak during the beginning of the infection and fall over time, antibodies generated against domain III are more neutralizing and persist for a long period of time41. Here, sera from EDIIIZIKV-immunized mice presented greater ability to neutralize ZIKV than those induced after immunization with EZIKV or EDI/IIZIKV in the C57Bl/6 strain. However, for BALB/c mice, sera from EZIKV or EDIIIZIKV mice showed the same neutralization capacity. In fact, protective anti-ZIKV antibody titers have already been observed in BALB/c mice after immunization with a truncated EZIKV protein 42. Likewise, C57BL/6 mice immunized with EDIIIZIKV showed protective humoral immunity35. Yang et al. demonstrated that immunization with virus-like particles (VLP) containing EDIIIZIKV in the presence of poly (I:C) induced a strong humoral response in C57BL/6 mice36.
Several studies demonstrated the important role of the cellular immune response against flaviviruses. The absence of CD8+ T cells during ZIKV infection increases mortality in mice24. DENV-specific CD8+ T cells induce cross-protection against ZIKV infection, including during pregnancy27. CD8+ T cells were shown to be essential to control yellow fever virus (YFV) and ZIKV infection in mice deficient in B lymphocytes43,44. CD4+ T cells also participate in the generation of protective immunity, as their depletion reduced the generation of anti-ZIKV antibodies25,26 and CD8+ T cell responses27. Recently, CD4+ T cells and IFNγ signaling have been shown to play a central role in protection during Zika virus infection45. Transfer experiments revealed that CD4+ T cells are required to protect against lethal challenge by ZIKV46. Furthermore, in a murine model of neuroinvasive ZIKV infection, the absence of CD4+ T cells leads to more neurological sequelae and increased viral titers in the central nervous system46. Indeed, the presence of polyfunctional CD4+ T cell responses is also implicated in protection against Japanese encephalitis virus (JEV) infection47, and is a hallmark after effective YFV vaccination48,49. We observed that immunization with EZIKV, EDI/IIZIKV and EDIIIZIKV proteins induced specific IFNγ-producing cells and polyfunctional CD4+ and CD8+ T cell responses.
Several studies have been carried out to identify immunodominant epitopes recognized by CD4+ and CD8+ T cells during flavivirus infections, particularly in DENV infection50. In DENV-infected patients and participants vaccinated with a live attenuated tetravalent vaccine, T cell epitopes were mapped in several regions of the DENV proteome, although CD8+ T cells preferentially recognized NS3, NS5, and NS4b regions, while CD4+ T cells tended to recognize structural proteins and NS151,52,53,54,55. Similarly, the same profile was detected in JEV infection47 and YFV vaccination49,56. In patients infected with ZIKV, CD4+ T cells target structural and nonstructural proteins in equal proportions, while CD8+ T cells preferentially focus on structural proteins24,50. In the context of previous exposure to DENV54, the CD8+ T cell response is modulated towards nonstructural proteins. A study with a DENV-naïve/ZIKV-infected patient, CD4+ and CD8+ T cell responses target preferentially NS2A and envelope proteins, respectively57. Furthermore, murine models have been instrumental not only in understanding the role of cellular immunity during flavivirus infections, but also in determining the epitopes recognized by T cells. Immunization of AG129 mice (Ifnar1−/−, Ifngr1−/−) and human HLA class II transgenic mice revealed that CD4+ T cell responses were directed to NS1, NS3, NS5 and envelope proteins58. A similar profile of CD8+ T cell responses was detected in infected C57BL/6 mice24. In this work, we identified four peptides present in the envelope region of the virus (ZIKV1(E1–20), ZIKV6(E51–70), ZIKV36(E351–370) and ZIKV37(E361–380)), capable of inducing a cellular immune response to the H-2Kd and H-2Kb haplotypes. Previous work58 mapped different ZIKV-immunodominant epitopes in HLA class II transgenic mice after immunization and the peptide ZIKV1(E1–20), the same we mapped (Supplementary Fig. 3a), was presented by HLA-DR1, -DR4, and DR1501 and -DQ8. Also, epitope mapping was performed during ZIKV infection in H-2b mice. Five CD8 immunodominant peptides were mapped24: two within peptide ZIKV1 (E1–20) (IGVSNRDFV and SNRDFVEGM), two in ZIKV6 (E51–70) region (TTVSNMAEV and RSYCYEASI) and one in the ZIKV36 (E351–370) region (MAVDMQTLTPV). In addition, one CD4-immunodominant peptide25 was also mapped in the middle of ZIKV36 (E351–370) and ZIKV37 (E361–380) peptide regions (PVGRLITANPVITES) (Supplementary Fig. 3a).
Collectively, our results demonstrate that EZIKV, EDI/IIZIKV, and EDIIIZIKV proteins are highly antigenic and immunogenic, inducing specific humoral and cellular immune responses. Furthermore, epitope mapping during immunization allowed the identification of immunodominant epitopes. In summary, our work provides a detailed assessment of the post-immunization immune response in different strains of mice mapping T-specific recognition regions in the envelope protein of ZIKV. These findings could help to better understand the immune response against ZIKV and add valuable information for future vaccine design.
Materials and methods
Production of optimized EZIKV, EDI/IIZIKV and EDIIIZIKV sequences
The alignment of ZIKV envelope (EZIKV) sequence (aa 291–690 of the ZIKV polyprotein) was generated using 69 Brazilian ZIKV sequences by the software ClustalW (GenBank accession numbers are available at Supplementary Table 1). The codon optimized gene was synthesized (GenScript, NJ) and cloned into the pET21a vector (pET21a-EZIKV). Then, the EDI/IIZIKV ectodomain (aa 291–600) was amplified by PCR (primers sense 5′-GGGCTAGCATTCGTTGCATCG-3′ and anti-sense 5′-CCCTCGAGCGCGGTGCACAGGCTGTA-3′; and EDIIIZIKV (aa 601–690) (primers sense 5′-GGGCTAGCGCGTTCACCTTTACCAAAATT-3′ and antisense 5′-GGCTCGAGCCAGTGGTGGGT-3′) using Phusion High Fidelity DNA Polymerase (New England Biolabs) as recommended by the manufacturer. The PCR product was cloned into the pJET1.2/blunt vector (Thermo Fisher Scientific), digested with endonucleases NheI and XhoI (New England Biolabs). We purified the digested fragment using PureLink Quick Plasmid DNA kit (Invitrogen) and then cloned in the pET21a vector using T4 DNA ligase enzyme (New England Biolabs).
Expression and purification of EZIKV, EDI/IIZIKV, and EDIIIZIKV proteins
EZIKV, EDI/IIZIKV and EDIIIZIKV recombinant proteins were expressed as monomers as previously described59. Briefly, transformed E. coli BL21 (DE3) RIL strain was cultured at 37 °C under agitation (200 rpm). After addition of 0.01 mM isopropyl β-d-1 thiogalactopyranoside (IPTG, Sigma) the bacterial pellet was suspended and lysed in a high-pressure system. The recombinant proteins were purified using a nickel affinity chromatography Ni-Sepharose histidine-tagged resin (GE Healthcare) as recommended by the manufacturer. Analysis of purified recombinant EZIKV, EDI/IIZIKV and EDIIIZIKV proteins were performed by electrophoresis using 15% SDS-PAGE gel under reducing conditions.
Western blot
Purified recombinant EZIKV, EDI/IIZIKV and EDIIIZIKV proteins (500 ng) were submitted to 15% SDS-PAGE electrophoresis under reducing conditions and then transferred to a nitrocellulose membrane (Hybond-C extra nitrocellulose—GE Healthcare). Next, membranes were blocked with PBS containing Tween 20 (PBST) (0.05% v/v), non-fat milk (5% w/v) and BSA (2.5% w/v), overnight at 4 °C. After each step, the membranes were washed 3 times with PBST. Then, the membranes were incubated with anti-his 6 × tag (1:5000—Thermo Fisher Scientific) for two hours at room temperature (rt). Nitrocellulose membranes were then incubated with HRP-labeled goat anti-mouse IgG (1:5000; KPL) at rt for 1 h. We used a chemiluminescence kit (ECL kit, GE Healthcare) to develop the reaction as recommended by manufacturer’s instructions and analyzed by Alliance 4.7 software (Uvitec; Cambridge).
Dot blot
Purified recombinant proteins EZIKV, EDI/IIZIKV, EDIIIZIKV and BSA (Bovine Serum Albumin) were added on nitrocellulose membrane (1 μγ) in total volume of 10 µL. After the membranes were completely dry, the following steps were performed as described above with just a minor modification. The primary antibody used was the anti-flavivirus 4G2 (1 μg/mL).
Mice and immunization
Female BALB/c or C57Bl/6 mice (6- to 8-weeks-old) were bred at Centro de Desenvolvimento de Modelos Experimentais para Medicina e Biologia (CEDEME)—UNIFESP. All mice were housed at Division of Immunology—UNIFESP. The experiments were approved by the UNIFESP Institutional Animal Care and Use Committee (IACUC) (protocol number #2020100418), in accordance with the recommendations of the Federal Law 11.794 (2008) and the Guide for the Care and Use of Laboratory Animals of the Brazilian National Council of Animal Experimentation (CONCEA). Mice were immunized with two doses, fifteen days apart, with equimolar amounts of EZIKV (10 μg), EDI/IIZIKV (7.78 μg) or EDIIIZIKV (2.44 μg) in the presence of poly (I:C) adjuvant (50 μg; Invivogen) in a total volume of 100 μL at the base of the tail (subcutaneously). The mice were bled by submandibular vein after each dose and were euthanized fifteen days after the second dose.
Measurement of ZIKV-specific antibodies
For ELISA, 96-well plates (high binding, Costar) were coated at rt overnight with 250 ng/well of EZIKV, EDI/IIZIKV or EDIIIZIKV diluted in 50 μL/well of PBS 1x. After each step the plates were washed with PBS Tween 20 (PBST) (0.02% v/v). Then, the plates were blocked for 2 h at rt with 150 μL of PBST, BSA (1% w/v) and non-fat milk (5% w/v). Next, serum from mice immunized were serially diluted and 100μL were applied to each well for 2 h at rt. Plates were then incubated with horseradish peroxidase-labeled goat anti-mouse IgG (1:10,000; KPL) for 2 h at rt. The enzymatic reaction was developed with 1 mg/mL of o-phenylenediamine (OPD, Sigma) diluted in phosphate–citrate buffer (0.2 M Na2PO4 and 0.2 M C6H807), pH 5, containing 0.03% (v/v) hydrogen peroxide and was stopped with 4 N H2SO4. We used a ELISA reader (EnSpire Multimode Plate Reader; PerkinElmer) to read plates at 492 nm (OD492). The antibody titer was determined by the highest dilution of serum that presented an OD492nm between 0.1 and 0.2.
Plaque Reduction Neutralization Test (PRNT)
A ZIKV isolate from Brazil (ZIKVBR), described by Cugola et al.60, was amplified in Vero E6 cells (ATCC CRL-1586) in complete MEM medium (supplemented with 10% FBS and 1% penicillin/streptomycin (GIBCO)) for 96 h. For the neutralization assay, 1 × 105 Vero CCL-81 cells (ATCC CCL-81) were plated in 24-well plates (Costar) in complete MEM medium and incubated overnight at 37 °C with 5% CO2. The following day, serum samples from immunized mice were previously inactivated for 30 min at 56 °C and incubated in the presence of 100 Plaque Forming Units (PFU) of ZIKV. Then, serum samples were serially diluted in 2% MEM medium (containing 2% FBS, 1% penicillin/streptomycin (GIBCO)) and then incubated with 100 PFU of ZIKV per well, for 1 h at 37 °C with 5% CO2. In addition, we added a dose test (DT)—which corresponds to 100 PFU; DT50 (50 PFU), mock (cell only), serum from non-immunized control mouse and from ZIKV-infected patient. Then, cells were incubated with a mixture containing the serum-virus for 3 h at 37 °C. Subsequently, the cells were overlayed with MEM medium with CMC (1.6% carboxymethylcellulose (CMC, Sigma) containing 10% FBS, 1% penicillin/streptomycin, 0.05% Amphotericin B (Fungizone, Gibco)) and incubated at 37 °C. After 4 days, the medium with CMC was completely removed and washed twice with PBS1X. Cells were then fixed with 4% paraformaldehyde (Sigma), stained with crystal violet (0.2%, Sigma) for half an hour and the excess dye was removed with distilled water.
Immunofluorescence assay (IFA)
The immunofluorescence assay was performed as described previously59. Briefly, 1 × 104 ZIKV-infected Vero cells were added to a multi-well glass slides in a MOI 0.1 at rt for 1 h. Cells were then fixed with acetone 80% solution (v/v) and incubated at − 20 °C for 30 min. After each step, wells were washed 3× with PBS 1X. Following incubation for 30 min with primary antibody (mouse immune sera, 1:500), goat anti-mouse IgG conjugated with FITC (1:750; Sigma) was added for 30 min. Immunofluorescence assay was performed using fluorescence microscopy (Olympus BX21) and the images were captured by CellSens software.
Splenocyte isolation
After euthanasia (2 weeks after the last dose) the spleens were aseptically removed, and ammonium chloride potassium (ACK) was used to lyse red blood cells. Splenocytes were resuspended in RPMI medium supplemented with 10% of fetal bovine serum, 40 μg/mL of gentamicin, 1% v/v vitamin solution, 2 mM l-glutamine, 1% v/v non-essential aminoacids solution, 1 mM sodium pyruvate, 1% penicillin/streptomycin and 5 × 10–5 M of 2-mercaptoethanol (all from Gibco).
Peptides
A peptide library (39 peptides) comprising the ZIKV envelope protein consensus sequence was synthesized (GenScript USA Inc) with purity more than 75% (20 amino acids overlapping 12-mer). Peptides were resuspended in DMSO (10 mg/mL) and stored at − 20 °C and organized into an optimized matrix (Supplementary Table 2) using DeconvoluteThis! Software as described previously28.
ELISpot assay
IFNγ producing cells were assessed using IFNγ ELISpot Ready-SET-Go! Kit (eBiosciences) as recommended by the manufacturer instructions. Briefly, ELISpot plates (MAIPS 4510, Millipore) were coated with IFNγ-capture antibody. After washes and blocking, splenocytes (3 × 105 cells) were added and incubated with pooled or individual peptides (10 μg/mL); equimolar amounts of recombinant proteins (EZIKV, EDI/IIZIKV, EDIIIZIKV) or R10 (negative control). We used the AID ELISpot Reader System (Autoimmun Diagnostika GmbH, Germany) to count the number of spots. The number of IFNγ producing cells from stimulated wells were subtracted from the non-stimulated wells.
T cell proliferation and cytokine production
To assess ZIKV-specific T cell proliferation, isolated splenocytes were labeled with carboxyfluorescein succinimidyl ester (CFSE), as previously described59. Briefly, splenocytes were labeled with 1.25 μM of CFSE (Molecular Probes), pouring the tube every two minutes for 10 min at 37 °C. Cells were then washed, resuspended and cultured for 5 days in the presence of the different stimuli. After 5 days, cells were first washed with buffer containing PBS with 0.5% BSA and 2 mM EDTA (FACS Buffer) and then stained with anti-mouse CD3-APCCy7 (clone 145-2C11), CD4-Pacific Blue (clone RM4-5) and CD8-APC (clone 53–6.7). For specific intracellular cytokine detection, splenocytes were cultured with the same antigens in the presence of anti-CD28 (2 μg/mL, BD Pharmigen) for second activation signal and Brefeldin A GolgiPlug™ (BD Pharmigen) for protein transport inhibition. Next, cells were washed with FACS buffer and stained with anti-mouse CD3-APCCy7 (clone 145-2C11), CD4-PerCP (clone RM4-5) and CD8-Pacific Blue (clone 53–6.7). Cells were then fixed and permeabilized using Cytofix/Cytoperm™ kit (BD Pharmigen) and washed with Perm/Wash buffer (BD Pharmigen). Cells were stained with anti-mouse TNFα-PECy7 (clone MP6-XT22) and IFNγ-APC (clone XMG1.2). All antibodies used for flow cytometry were from BD Pharmingen. The samples were acquired using the FACSCanto II flow cytometry (BD Biosciences) and analyzed using FlowJo software (Tree Star). To allow proper compensation, unstained and all single-color controls were performed. The frequency of proliferating cells was calculated by subtracting the values from unstimulated cells.
Data analysis
Data normality tests were performed in GraphPad Prism including Shapiro–Wilk and D’Agostino-Pearson omnibus tests. Statistical significance (p-values) was calculated by One-way or Two-way ANOVA followed by Tukey honestly significantly different (HSD) post hoc test. Statistical analysis and graphical representation were performed using GraphPad Prism version 9.0 software.
Ethics statement
This study was carried out in compliance with the recommendations of the Federal Law 11.794 (2008), the Guide for the Care and Use of Laboratory Animals of the Brazilian National Council of Animal Experimentation (CONCEA) and the ARRIVE guidelines (https://arriveguidelines.org). The protocol (number 2020100418) was approved by the UNIFESP Animal Care and Use Committee (IACUC).
References
Gatherer, D. & Kohl, A. Zika virus: A previously slow pandemic spreads rapidly through the Americas. J. Gen. Virol. 97, 269–273. https://doi.org/10.1099/jgv.0.000381 (2016).
Dick, G. W., Kitchen, S. F. & Haddow, A. J. Zika virus. I. Isolations and serological specificity. Trans. R. Soc. Trop. Med. Hyg. 46, 509–520 (1952).
WHO. ZIKA virus, microcephaly, Guillain–Barré syndrome. in (ed. Report, S.) (World Health Organization, 2017).
Brooks, J. T. et al. Update: Interim guidance for prevention of sexual transmission of ZIKA Virus—United States, July 2016. MMWR Morb. Mortal Wkly. Rep. 65, 745–747. https://doi.org/10.15585/mmwr.mm6529e2 (2016).
Collins, M. H. & Waggoner, J. J. Detecting vertical ZIKA transmission: emerging diagnostic approaches for an emerged flavivirus. ACS Infect. Dis. 5, 1055–1069. https://doi.org/10.1021/acsinfecdis.9b00003 (2019).
de Araujo, T. V. B. et al. Association between microcephaly, Zika virus infection, and other risk factors in Brazil: Final report of a case-control study. Lancet Infect. Dis. 18, 328–336. https://doi.org/10.1016/S1473-3099(17)30727-2 (2018).
Franca, G. V. et al. Congenital Zika virus syndrome in Brazil: A case series of the first 1501 livebirths with complete investigation. Lancet 388, 891–897. https://doi.org/10.1016/S0140-6736(16)30902-3 (2016).
Brasil, P. et al. Zika virus infection in pregnant women in Rio de Janeiro. N. Engl. J. Med. 375, 2321–2334. https://doi.org/10.1056/NEJMoa1602412 (2016).
Kleber de Oliveira, W. et al. Increase in reported prevalence of microcephaly in infants born to women living in areas with confirmed ZIKA virus transmission during the first trimester of pregnancy—Brazil. MMWR Morb. Mortal. Wkly. Rep. 65, 242–247. https://doi.org/10.15585/mmwr.mm6509e2 (2016).
Cao-Lormeau, V. M. et al. Guillain–Barre Syndrome outbreak associated with Zika virus infection in French Polynesia: A case–control study. Lancet 387, 1531–1539. https://doi.org/10.1016/S0140-6736(16)00562-6 (2016).
Styczynski, A. R. et al. Increased rates of Guillain–Barre syndrome associated with Zika virus outbreak in the Salvador metropolitan area, Brazil. PLoS Negl. Trop. Dis. 11, e0005869. https://doi.org/10.1371/journal.pntd.0005869 (2017).
Sirohi, D. et al. The 3.8 A resolution cryo-EM structure of Zika virus. Science 352, 467–470. https://doi.org/10.1126/science.aaf5316 (2016).
Faye, O. et al. Molecular evolution of Zika virus during its emergence in the 20(th) century. PLoS Negl. Trop. Dis. 8, e2636. https://doi.org/10.1371/journal.pntd.0002636 (2014).
Dai, L. et al. Structures of the Zika virus envelope protein and its complex with a flavivirus broadly protective antibody. Cell Host Microbe 19, 696–704. https://doi.org/10.1016/j.chom.2016.04.013 (2016).
Whitehead, S. S., Blaney, J. E., Durbin, A. P. & Murphy, B. R. Prospects for a dengue virus vaccine. Nat. Rev. Microbiol. 5, 518–528. https://doi.org/10.1038/nrmicro1690 (2007).
Stettler, K. et al. Specificity, cross-reactivity, and function of antibodies elicited by Zika virus infection. Science 353, 823–826. https://doi.org/10.1126/science.aaf8505 (2016).
Sapparapu, G. et al. Neutralizing human antibodies prevent Zika virus replication and fetal disease in mice. Nature 540, 443–447. https://doi.org/10.1038/nature20564 (2016).
Larocca, R. A. et al. Vaccine protection against Zika virus from Brazil. Nature https://doi.org/10.1038/nature18952 (2016).
Zhao, H. et al. Structural basis of Zika virus-specific antibody protection. Cell 166, 1016–1027. https://doi.org/10.1016/j.cell.2016.07.020 (2016).
Wang, Q. et al. Molecular determinants of human neutralizing antibodies isolated from a patient infected with Zika virus. Sci. Transl. Med. 8, 369ra179. https://doi.org/10.1126/scitranslmed.aai8336 (2016).
Robbiani, D. F. et al. Recurrent potent human neutralizing antibodies to Zika Virus in Brazil and Mexico. Cell 169, 597-609.e511. https://doi.org/10.1016/j.cell.2017.04.024 (2017).
Li, C. et al. A single injection of human neutralizing antibody protects against Zika virus infection and microcephaly in developing mouse embryos. Cell Rep. 23, 1424–1434. https://doi.org/10.1016/j.celrep.2018.04.005 (2018).
Lima, N. S., Rolland, M., Modjarrad, K. & Trautmann, L. T cell immunity and Zika virus vaccine development. Trends Immunol. 38, 594–605. https://doi.org/10.1016/j.it.2017.05.004 (2017).
Elong Ngono, A. et al. Mapping and role of the CD8+ T cell response during primary Zika Virus infection in mice. Cell Host Microbe 21, 35–46. https://doi.org/10.1016/j.chom.2016.12.010 (2017).
Elong Ngono, A. et al. CD4+ T cells promote humoral immunity and viral control during Zika virus infection. PLoS Pathog. 15, e1007474. https://doi.org/10.1371/journal.ppat.1007474 (2019).
Scott, J. M. et al. Cellular and humoral immunity protect against vaginal Zika virus infection in mice. J. Virol. https://doi.org/10.1128/JVI.00038-18 (2018).
Regla-Nava, J. A. et al. Cross-reactive dengue virus-specific CD8(+) T cells protect against Zika virus during pregnancy. Nat. Commun. 9, 3042. https://doi.org/10.1038/s41467-018-05458-0 (2018).
Precopio, M. L. et al. Optimizing peptide matrices for identifying T-cell antigens. Cytometry A 73, 1071–1078. https://doi.org/10.1002/cyto.a.20646 (2008).
Murin, C. D., Wilson, I. A. & Ward, A. B. Antibody responses to viral infections: A structural perspective across three different enveloped viruses. Nat. Microbiol. 4, 734–747. https://doi.org/10.1038/s41564-019-0392-y (2019).
Rey, F. A., Stiasny, K., Vaney, M. C., Dellarole, M. & Heinz, F. X. The bright and the dark side of human antibody responses to flaviviruses: Lessons for vaccine design. EMBO Rep. 19, 206–224. https://doi.org/10.15252/embr.201745302 (2018).
Crill, W. D. & Roehrig, J. T. Monoclonal antibodies that bind to domain III of dengue virus E glycoprotein are the most efficient blockers of virus adsorption to Vero cells. J. Virol. 75, 7769–7773. https://doi.org/10.1128/JVI.75.16.7769-7773.2001 (2001).
Gromowski, G. D. & Barrett, A. D. Characterization of an antigenic site that contains a dominant, type-specific neutralization determinant on the envelope protein domain III (ED3) of dengue 2 virus. Virology 366, 349–360. https://doi.org/10.1016/j.virol.2007.05.042 (2007).
Tai, W. et al. Critical neutralizing fragment of Zika virus EDIII elicits cross-neutralization and protection against divergent Zika viruses. Emerg. Microbes Infect. 7, 7. https://doi.org/10.1038/s41426-017-0007-8 (2018).
Wahala, W. M., Kraus, A. A., Haymore, L. B., Accavitti-Loper, M. A. & de Silva, A. M. Dengue virus neutralization by human immune sera: Role of envelope protein domain III-reactive antibody. Virology 392, 103–113. https://doi.org/10.1016/j.virol.2009.06.037 (2009).
Yang, M., Dent, M., Lai, H., Sun, H. & Chen, Q. Immunization of Zika virus envelope protein domain III induces specific and neutralizing immune responses against Zika virus. Vaccine 35, 4287–4294. https://doi.org/10.1016/j.vaccine.2017.04.052 (2017).
Yang, M., Lai, H., Sun, H. & Chen, Q. Virus-like particles that display Zika virus envelope protein domain III induce potent neutralizing immune responses in mice. Sci. Rep. 7, 7679. https://doi.org/10.1038/s41598-017-08247-9 (2017).
Lai, C. Y. et al. Antibodies to envelope glycoprotein of dengue virus during the natural course of infection are predominantly cross-reactive and recognize epitopes containing highly conserved residues at the fusion loop of domain II. J. Virol. 82, 6631–6643. https://doi.org/10.1128/JVI.00316-08 (2008).
Beltramello, M. et al. The human immune response to Dengue virus is dominated by highly cross-reactive antibodies endowed with neutralizing and enhancing activity. Cell Host Microbe 8, 271–283. https://doi.org/10.1016/j.chom.2010.08.007 (2010).
Crill, W. D., Hughes, H. R., Delorey, M. J. & Chang, G. J. Humoral immune responses of dengue fever patients using epitope-specific serotype-2 virus-like particle antigens. PLoS ONE 4, e4991. https://doi.org/10.1371/journal.pone.0004991 (2009).
Oliphant, T. et al. Induction of epitope-specific neutralizing antibodies against West Nile virus. J. Virol. 81, 11828–11839. https://doi.org/10.1128/JVI.00643-07 (2007).
Yu, L. et al. Delineating antibody recognition against Zika virus during natural infection. JCI Insight. https://doi.org/10.1172/jci.insight.93042 (2017).
Han, J. F. et al. Immunization with truncated envelope protein of Zika virus induces protective immune response in mice. Sci. Rep. 7, 10047. https://doi.org/10.1038/s41598-017-10595-5 (2017).
Bassi, M. R. et al. CD8+ T cells complement antibodies in protecting against yellow fever virus. J. Immunol. 194, 1141–1153. https://doi.org/10.4049/jimmunol.1402605 (2015).
Nazerai, L. et al. A new in vivo model to study protective immunity to Zika virus infection in mice with intact type i interferon signaling. Front. Immunol. 9, 593. https://doi.org/10.3389/fimmu.2018.00593 (2018).
Lucas, C. G. O. et al. Critical role of CD4(+) T cells and IFNgamma signaling in antibody-mediated resistance to Zika virus infection. Nat. Commun. 9, 3136. https://doi.org/10.1038/s41467-018-05519-4 (2018).
Hassert, M. et al. CD4+T cells mediate protection against Zika associated severe disease in a mouse model of infection. PLoS Pathog. 14, e1007237. https://doi.org/10.1371/journal.ppat.1007237 (2018).
Turtle, L. et al. Human T cell responses to Japanese encephalitis virus in health and disease. J. Exp. Med. 213, 1331–1352. https://doi.org/10.1084/jem.20151517 (2016).
Gaucher, D. et al. Yellow fever vaccine induces integrated multilineage and polyfunctional immune responses. J. Exp. Med. 205, 3119–3131. https://doi.org/10.1084/jem.20082292 (2008).
Akondy, R. S. et al. The yellow fever virus vaccine induces a broad and polyfunctional human memory CD8+ T cell response. J. Immunol. 183, 7919–7930. https://doi.org/10.4049/jimmunol.0803903 (2009).
Slon Campos, J. L., Mongkolsapaya, J. & Screaton, G. R. The immune response against flaviviruses. Nat. Immunol. 19, 1189–1198. https://doi.org/10.1038/s41590-018-0210-3 (2018).
Duangchinda, T. et al. Immunodominant T-cell responses to dengue virus NS3 are associated with DHF. Proc. Natl. Acad. Sci. USA 107, 16922–16927. https://doi.org/10.1073/pnas.1010867107 (2010).
Rivino, L. et al. Differential targeting of viral components by CD4+ versus CD8+ T lymphocytes in dengue virus infection. J. Virol. 87, 2693–2706. https://doi.org/10.1128/JVI.02675-12 (2013).
Weiskopf, D. et al. The human CD8+ T cell responses induced by a live attenuated tetravalent dengue vaccine are directed against highly conserved epitopes. J. Virol. 89, 120–128. https://doi.org/10.1128/JVI.02129-14 (2015).
Grifoni, A. et al. Prior dengue virus exposure shapes T cell immunity to Zika virus in humans. J. Virol. https://doi.org/10.1128/JVI.01469-17 (2017).
Grifoni, A. et al. Global assessment of dengue virus-specific CD4(+) T cell responses in dengue-endemic areas. Front. Immunol. 8, 1309. https://doi.org/10.3389/fimmu.2017.01309 (2017).
James, E. A. et al. Yellow fever vaccination elicits broad functional CD4+ T cell responses that recognize structural and nonstructural proteins. J. Virol. 87, 12794–12804. https://doi.org/10.1128/JVI.01160-13 (2013).
Ricciardi, M. J. et al. Ontogeny of the B- and T-cell response in a primary Zika virus infection of a dengue-naive individual during the 2016 outbreak in Miami, FL. PLoS Negl. Trop. Dis. 11, e0006000. https://doi.org/10.1371/journal.pntd.0006000 (2017).
Reynolds, C. J. et al. T cell immunity to Zika virus targets immunodominant epitopes that show cross-reactivity with other Flaviviruses. Sci. Rep. 8, 672. https://doi.org/10.1038/s41598-017-18781-1 (2018).
Amaral, M. P. et al. Homologous prime-boost with Zika virus envelope protein and poly (I:C) induces robust specific humoral and cellular immune responses. Vaccine 38, 3653–3664. https://doi.org/10.1016/j.vaccine.2020.03.037 (2020).
Cugola, F. R. et al. The Brazilian Zika virus strain causes birth defects in experimental models. Nature 534, 267–271. https://doi.org/10.1038/nature18296 (2016).
Acknowledgements
We thank Mr. Geová Santos for assistance in the animal facility.
Funding
This research was supported by São Paulo Research Foundation (FAPESP, Grants number 2017/17471-7, 2021/13004-0 and 2018/05320-7), the Brazilian National Research Council (CNPq)/Institute for Investigation in Immunology (CNPq, Grant 465434/2014-2) and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES, Finance Code 001). V.A.S.L and F.C.C received fellowship from FAPESP. H.F.S.S and J.A.M received fellowship from CAPES. J.S.A received fellowship from CNPq, S.B.B. and D.S.R received fellowships from FAPESP and CNPq.
Author information
Authors and Affiliations
Contributions
V.A.S.L.: investigation, methodology, formal analysis, validation, writing—original draft, visualization. J.S.A.: investigation, methodology, visualization. H.F.S.S., F.C.C., J.A.M.: investigation. R.M.A.: resources. S.B.B.: conceptualization, methodology, resources, writing—review & editing. D.S.R.: conceptualization, methodology, resources, writing—original draft, writing—review & editing, visualization, supervision, project administration, funding acquisition.
Corresponding author
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lunardelli, V.A.S., de Souza Apostolico, J., Souza, H.F.S. et al. ZIKV-envelope proteins induce specific humoral and cellular immunity in distinct mice strains. Sci Rep 12, 15733 (2022). https://doi.org/10.1038/s41598-022-20183-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-20183-x
- Springer Nature Limited
This article is cited by
-
Advances in vaccines: revolutionizing disease prevention
Scientific Reports (2023)