Abstract
Dickeya fangzhongdai, a bacterial pathogen of taro (Colocasia esculenta), onion (Allium sp.), and several species in the orchid family (Orchidaceae) causes soft rot and bleeding canker diseases. No field-deployable diagnostic tool is available for specific detection of this pathogen in different plant tissues. Therefore, we developed a field-deployable loop-mediated isothermal amplification (LAMP) assay using a unique genomic region, present exclusively in D. fangzhongdai. Multiple genomes of D. fangzhongdai, and other species of Dickeya, Pectobacterium and unrelated genera were used for comparative genomic analyses to identify an exclusive and conserved target sequence from the major facilitator superfamily (MFS) transporter gene region. This gene region had broad detection capability for D. fangzhongdai and thus was used to design primers for endpoint PCR and LAMP assays. In-silico validation showed high specificity with D. fangzhongdai genome sequences available in the NCBI GenBank genome database as well as the in-house sequenced genome. The specificity of the LAMP assay was determined with 96 strains that included all Dickeya species and Pectobacterium species as well as other closely related genera and 5 hosts; no false positives or false negatives were detected. The detection limit of the assay was determined by performing four sensitivity assays with tenfold serially diluted purified genomic DNA of D. fangzhongdai with and without the presence of crude host extract (taro, orchid, and onion). The detection limit for all sensitivity assays was 100 fg (18–20 genome copies) with no negative interference by host crude extracts. The assays were performed by five independent operators (blind test) and on three instruments (Rotor-Gene, thermocycler and dry bath); the assay results were concordant. The assay consistently detected the target pathogen from artificially inoculated and naturally infected host samples. The developed assay is highly specific for D. fangzhongdai and has applications in routine diagnostics, phytosanitary and seed certification programs, and epidemiological studies.
Similar content being viewed by others
Introduction
An emerging bacterial pathogen threatens taro (Colocasia esculenta), onion (Allium sp.), and various species in the orchid family (Orchidaceae). Taro is among the world's essential starch crops needed for food security, especially in Asia, Africa, and the Pacific Islands, with an estimated production value of $4.1 billion dollars in 20161,2,3. In 2017, gross global onion production was valued at $44.7 billion2. Orchids are a growing commodity in global horticultural trade contributing to the economic development of tropical Asia. These crops represent significant value to the agricultural economies of the world, necessitating improved crop protection measures during production and marketing. Recently, Dickeya fangzhongdai was associated with soft rot diseases among these economically important plant species and new occurrences of this pathogen have been reported in several regions4,5,6,7.
Dickeya fangzhongdai is classified within Pectobacteriaceae, a genetically diverse, Gram-negative, facultatively anaerobic bacterial species capable of causing soft rot disease in a range of monocot hosts and also reported to cause bleeding canker disease of pear (Pyrus pyrifolia), a dicot host8,9,10,11,12. Dickeya species may persist in soil, surface water, and groundwater, or as epiphytes and saprophytes13,14,15,16. Infection by Dickeya species begins with attraction, adhesion, and invasion of the host through wounds and natural openings such as stomata. Plant cell wall-degrading enzymes (PCWDE) initiate dissolution of hemicelluloses, a major component of the middle lamella, resulting in further dissolution of cell walls17. Symptoms are not always obvious at initial stages of the disease, and its progress is dependent on the rate of bacterial multiplication as well as environmental conditions such as temperature, humidity, and the availability of oxygen18.
Ideally, regulated pathogens would be detected at borders to prevent disease transmission across cropping areas during trading. Biosecurity protocols for crop management require accurate and robust diagnostic assays to prevent and/or mitigate the spread of economically damaging pathogens19. Diagnostic assays must consider the biological interactions between pathogens, their hosts, and the environment for robust analyte detection and confidence in assay results. Therefore, natural conditions for infection and sample types, i.e., DNA isolation matrix, niche habitat, and pathogen biology, are crucial for the development of reliable diagnostic tools20. Reliable target sensitivity at critically low detection limits is essential for the field application of diagnostic strategies, especially with regards to inherent fluctuation patterns in pathogen abundance over the course of the disease cycle. A robust field implementable diagnostic tool should adequately address inhibitory factors and non-target components of natural isolation matrices to maintain high target sensitivity while also guaranteeing target specificity21.
While endpoint and quantitative PCR are among the few diagnostic techniques currently available for detecting Dickeya spp. and D. fangzhongdai in agricultural food crops22, these methods are impractical for field application. Loop mediated isothermal amplification (LAMP) technology offers a practical solution for field-deployable diagnostics that requires less time and space for point-of-need testing, additional capabilities to laboratories that lack specialized diagnostic equipment23. This diagnostic technology can be implemented globally to supplement biosecurity and disease management programs, especially in regions agriculturally dependent on onion, taro, orchid, and other susceptible crops. LAMP assays are highly sensitive and efficient for detection of nucleic acid sequences and can produce results in thirty minutes or less24,25. As an isothermal reaction, LAMP can facilitate target amplification from crude DNA extracts using only a heating block or water bath23,26. A suite of primers guides the polymerase to create several stem-loop structures of varying lengths, resulting in a high concentration of the target sequence. Several methods are used to visualize the results of a LAMP reaction, ranging from colorimetric indicators to gel electrophoresis, which allows adaptation of this protocol to diverse diagnostician’s needs23,25,27. Multiplex reactions are possible with LAMP due to its compatibility with lateral flow devices28 and fluorescent probes29. Rapid and accurate diagnosis is necessary for efficient disease management, and LAMP results can be determined in less than half an hour25,27. The simplicity, specificity, sensitivity, and speed of LAMP make this method an ideal point-of-need diagnostic tool.
The objective of this study was to develop a highly specific and sensitive LAMP assay to readily test for the target pathogen in field environments. Four target primers and two loop primers were developed to detect a region of the conserved major facilitator superfamily (MFS) transporter gene region unique to D. fangzhongdai and thus detect this pathogen in infected crops. The primers were verified as specific to D. fangzhongdai when tested against closely related species. The developed LAMP assay has potential applications in point-of-need plant disease diagnostics, farm management, disease surveys, and plant biosecurity.
Results
Target selection and primer design
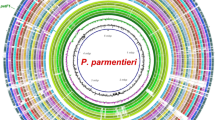
Several potential genomic regions were evaluated for target selection. Regions that were not present in all 13 D. fangzhongdai genomes, including a strain from Hawaii that was not previously reported, were not considered for further LAMP assay development. During endpoint PCR validation tests, 4 regions appeared conserved in D. fangzhongdai alone based on preliminary specificity tests using CFBP8607T (D. fangzhongdai type strain), PL145 (D. fangzhongdai), PL47 (D. zeae), A5307 (D. oryzae), LMG30899 (Dickeya sp.), PL25 (D. dianthicola), and PL84 (D. solani) (data not shown). Later, each region was evaluated in silico using the NCBI genome database. The major facilitator superfamily (MFS) transporter gene region was selected as a highly unique region in D. fangzhongdai and used for development of a specific and robust assay (Fig. 1). This region was not identified in other Dickeya species nor in the closely related genus, Pectobacterium. When tested using an endpoint PCR at an annealing temperature of 57 °C, MFS transporter region primers did not have off-target amplification. In-silico analysis indicated that the 1409-nucleotide long MFS transporter region was not identical in other genomes present in the NCBI GenBank genome database. However, the MFS transporter region was found in all genomes of D. fangzhongdai publicly available as well as one unpublished genome of a D. fangzhongdai strain isolated in Hawaii (Table 1). Results from BLAST database revealed a high identity among all strains of D. fangzhongdai for this region, where query coverage was above 99% and pairwise identity was above 96.5% compared to the genome (NZ_CP025003.1) of the type strain CFBP8607T (Table 1). Endpoint primers and six LAMP primers, designed using the MFS transporter gene, were specific to all D. fangzhongdai genomes available, showing no significant identity with other genomes in the NCBI GenBank database. Endpoint primers have a 100% sequence match to all strains listed in Table 1, and LAMP primers have a 100% sequence match with all genomes obtained from the NCBI GenBank, although 2 SNPs among 6 primers were observed for the strain isolated in Hawaii (Table 2; Fig. 1B). The SNPs did not impact the specificity of the assay (Table 3; Fig. 2).
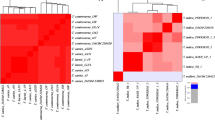
(A) Circular graphic pointing out the location of the target sequence MFS transporter exclusively present in Dickeya fangzhongdai. The ring image illustrates the multiple genome alignment of four D. fangzhongdai strains followed by the other eleven species that currently form Dickeya. The last three outermost rings depict the genomes of three different species within Pectobacterium. The three innermost rings depict the genome coordinates in terms of mega base pairs (Mbp), the GC content (black line in zigzag shape), and the GC skew (zigzag purple +/green −) of the reference genome D. fangzhongdai QZH3. The remaining color-coded rings portray, beginning with the innermost ring, the BLASTn pairwise comparison of D. fangzhongdai QZH3 (NZ_CP031507), D. fangzhongdai ND14b (NZ_CP009460), D. fangzhongdai PA1 (NZ_CP020872), D. fangzhongdai PL145 (unpublished), position of the target gene encoding for a MFS transporter (DYD83_00310, highlighted and labeled in violet color), D. aquatica 174/2T (NZ_LT615367), D. chrysanthemi Ech1591 (NC_012912), D. dadantii NCPPB 2976T (NZ_CM001978), D. dianthicola ME23 (NZ_CP031560), D. lacustris S29T (QNUT00000000), D. oryzae ZYY5T (SULL00000000), D. parazeae Ech586 (NC_013592), D. poaceiphila NCPPB 569T (NZ_CP042220), D. solani IPO 2222 (NZ_CP015137), D. undicola 2B12 (JSYG00000000), D. zeae PL65 (NZ_CP040817), P. parmentieri RNS 08-42-1AT (NZ_CP015749), P. brasiliense 1692 (NZ_CP047495) and P. carotovorum XP-13 (NZ_CP063242). The image was created using the BLAST Ring Image Generator (BRIG) v 0.9540. (B) Alignment of MFS Transporter regions of 13 genomes of D. fangzhongdai and endpoint PCR primers in Geneious Prime. Red represents primers used for endpoint PCR while green represents primers used for LAMP. Gray represents identical regions while all other colors indicate a single nucleotide polymorphism compared to the consensus identity.
Specificity validation of the loop mediated isothermal amplification (LAMP) primer set designed to amplify a unique MFS transporter gene region of the Dickeya fangzhongdai. (A) Sigmoidal amplification curves were produced by seven D. fangzhongdai strains that formed the inclusivity panel: CFBP8607T (positive control) and PL145, PL146, PL147, PL148, PL149, and PL150. No amplification was observed with gDNA from the strains of the exclusivity panel and non-template control (NTC; water). (A,B) Tube 1: CFBP8607T; tubes 2–7: D. fangzhongdai strains, PL145, PL146, PL147, PL148, PL149, and PL150; tubes 8–15: representative strains of the exclusivity panel: LMG27354 (D. aquatica), A6060 (D. dadantii), A6069 (D. zeae), A5572 (D. dianthicola), LMG30903 (D. undicola), A5420 (D. paradisiaca), PL88 (D. solani), PL73 (Pectobacterium carotovorum); and tube N: NTC. All reactions of gDNA template extracted from bacterial strains of the inclusivity panel produced positive results, while all reactions of the exclusivity panel failed to produce amplification. (B) Colorimetric detection of LAMP reaction products under ambient light after the addition of SYBR Green I dye; positive results appear green. (C) Detection of the fluorescent emission from the tubes with positive amplification under UV light.
Specificity assays
The specificity of the LAMP primers was validated using selected inclusivity and exclusivity panels. The inclusivity panel consisted of purified genomic DNA (gDNA) extracted from seven D. fangzhongdai strains, while the exclusivity panel consisted of purified gDNA extracted from 89 non-target bacterial strains of different species (Table 3). The exclusivity panel also included DNA of four plant hosts and one soil sample (Table 3). The assay successfully detected all D. fangzhongdai strains in the inclusivity panel, CFBP8607T and PL145-PL150, while no false positives were obtained from non-target bacterial species, host plants, or soil (Table 3). Based on melting curve results, the single peak result indicated all seven D. fangzhongdai strains presenting positive calls as well (Fig. S1). Results were visualized by real time analysis of sigmoidal amplification curves (Rotor-Gene Q), and colorimetrically after adding SYBR Green I dye30. During amplification, the cycle threshold of positive calls ranged from 7 to 10. After the addition of SYBR Green I dye, tubes with positive reactions turned green under ambient daylight and emitted fluorescence upon excitation with UV light. The results of all seven D. fangzhongdai strains were similar when resolved under the described visualization methods, whereas none of the 94 DNA samples in the exclusivity panel were amplified, evident by real time analysis, the orange appearance under ambient light, and the lack of fluorescence after the addition of SYBR Green dye (Table 3, Fig. 2).
The specificity of the endpoint PCR primer pair that was designed to amplify the MFS transporter gene was also validated using the full LAMP inclusivity panel, which was composed of seven D. fangzhongdai strains and with a subset of the exclusivity panel used to validate the LAMP assay (Fig. 3). The reduced exclusivity panel consisted of 28 bacterial strains of phylogenetically close genera, and of niche- and host sharing-species; the panel included non-target Dickeya species, Pectobacterium, Ralstonia, Xanthomonas, and other Gram-positive bacteria, such as Clavibacter and Rathayibacter species (Fig. 3). All PCR reactions were completed by using the same gDNA sources that were used for the LAMP exclusivity assay. Reactions with D. fangzhongdai gDNA templates yielded the expected amplification product, 421-bp in length, while reactions completed using template DNA from other selected species did not produce any amplification products apparent during resolution of PCR products by agarose gel electrophoresis (Fig. 3).
The specificity validation of the endpoint PCR primer pair designed to specifically detect Dickeya fangzhongdai. A total of 36 reactions including seven D. fangzhongdai: CFBP8607T (positive control, lane 1) and PL145-PL150 (lanes 2–7) from the inclusivity panel used for the validation of the LAMP primer set. The remaining lanes consisted of reactions of gDNA templates extracted from 28 representative strains forming the PCR exclusivity panel (lanes 8–35): A6069, A5420, A6060, PL84, A5572, LMG27354, PL73, CFBP8637, LMG2414, 32F, CFBP8631, LMG17936, LMG30899, LMG30903, CFBP1357, LMG2466, LMG1268, LMG21371, LMG30744, LMG26003, 19170, A2041, A6205, A6219, A1084, A5726, A223, and DP164, and non-template control (NTC, lane 36).
Limit of detection and spiked assays
The limit of detection of the D. fangzhongdai LAMP assay was determined by performing the assay on a tenfold serially diluted preparation of genomic DNA of type strain CFBP8607. The LAMP assay successfully detected the target DNA at a minimum concentration of 100 fg (Fig. 4A–C). To assess possible reaction inhibition by host matrices, three spiked assays (onion, orchid, and taro) were performed by adding 5 µl crude lysate (this 5 µl amount is recommended by the manufacturer when running an assay with plant samples) derived from healthy host tissues to the tenfold serially diluted genomic DNA of D. fangzhongdai23. Assays spiked with any of the three crude host lysates maintained the 100 fg detection limit (Fig. 5). The inclusion of non-template control (NTC; water) in each reaction reported no occurrence of false positive results. Using Rotor-Gene Q, SYBR Green and UV light to validate the results, no discrepancies were found. When comparing assay sensitivity with endpoint PCR, LAMP was 100-fold more sensitive (Figs. 4, 5).
Detection limit determination of loop mediated isothermal amplification (LAMP) assay designed for specific detection of Dickeya fangzhongdai. Purified genomic DNA (CFBP8607T), serially diluted tenfold (from 10 ng to 1 fg) was used per reaction (tubes 1–8), with tube 9 being non-template control (NTC) water. (A) Sigmoidal curves; (B) results after adding SYBR Green I dye to the amplified reaction—a color change from orange to bright green indicates a positive amplification; (C) tubes observed under UV light—fluorescence is indicative of positive amplification. Positive amplification with the LAMP assay was observed for concentrations as little as 100 fg (tube 6). (D) Endpoint sensitivity assay with genomic DNA, serially diluted tenfold (from 10 ng to 1 fg); (E) spiked assay (1 µl of healthy taro corm genomic DNA was added in tenfold serially diluted genomic DNA). In both cases, endpoint PCR was 100-times less sensitive than the LAMP assay.
Detection limit determination of the loop mediated isothermal amplification (LAMP) assay designed for specific detection of Dickeya fangzhongdai in the presence of host matrices. Five µl of crude host lysate was added to purified genomic DNA (CFBP8607), tenfold serially diluted (from 10 ng to 1 fg) (tubes 1 to 8; tube 9 was non-template control). (A) Addition of SYBR Green I dye—a bright green color indicates positive amplification; (B) tubes observed under UV light—fluorescence indicate positive amplification. Amplification was observed for concentrations as little as 100 fg (tube 6), indistinguishable from the limit of detection obtained without the addition of crude host lysate.
Multi-operator blind tests
Five different operators performed multi-operator blind tests with twelve blind samples to confirm reproducibility and robustness of the developed assay. All 12 samples, including the NTC, were tested with the LAMP assay to specifically detect three D. fangzhongdai strains (Fig. 6). All operator results were 100% in agreement and specifically detected D. fangzhongdai (Fig. 6). No false positives or false negatives were detected during the blind test.
Multi-operator blind tests of the Dickeya fangzhongdai specific loop mediated isothermal amplification (LAMP) assay were performed to validate reproducibility and robustness. Twelve representative samples were used: Tubes 1–12: Dickeya lacustris LMG30899, D. dianthicola PL25, D. solani LMG27552, D. zeae A5422, D. fangzhongdai CFBP8607T, D. aquatica LMG27354, D. fangzhongdai PL146, Pantoea sp. A1869, D. fangzhongdai PL148, non-template control (NTC; water), D. chrysanthemi A5641 and D. paradisiaca A5579, respectively.
Multi-device detection of D. fangzhongdai directly from bacterial colonies
Multi-device tests were performed using three different platforms (Qiagen Rotor-Gene G, Bio-Rad thermocycler, and dry bath). Twelve samples were tested, seven were D. fangzhongdai, four were various Dickeya spp. and a non-template control (NTC; water). All seven DNA samples of D. fangzhongdai were validated directly from bacterial colonies. Results were 100% in agreement with the sample correlation (Fig. 7). No false positives or false negatives were detected during the validation test.
Multi-device detection directly from colonies of D. fangzhongdai were validated using three different incubation platforms. Eleven representative samples were tested. Tubes 1–11: D. fangzhongdai strains CFBP8607T, PL145, PL146, PL147, PL148, PL149, PL150, D. aquatica LMG 27354, D. lacustris LMG30899, D. undicola LMG30903, D. oryzae A5410 and non-template negative control (NTC; water), respectively. Three detection methods were used: sigmoidal curves, SYBR Green I dye detected visually, and fluorescence detection under UV light. (A1–A3) Rotor-Gene amplification reaction performed by two independent groups with positive amplification observed by sigmoidal curve and further visualized by adding SYBR Green Dye—bright green color indicative of positive amplification; (B) LAMP amplification using an endpoint thermal cycler and visualized by adding SYBR Green Dye; (C) LAMP amplification using a dry bath (hot plate) and visualized by adding SYBR Green Dye.
Validation using artificially and naturally infected samples
To further determine its viability as a diagnostic test, the LAMP assay was tested with eight strains of D. fangzhongdai by artificially infecting taro corm and orchid stem samples. The LAMP assay accurately determined the presence of D. fangzhongdai in four of the samples inoculated with D. fangzhongdai, and did not cross-react with any samples inoculated with related Dickeya or other microbial species (Figs. 8 and 9). The LAMP assay also accurately determined the presence of D. fangzhongdai in two (confirmed by isolating the bacteria; data not shown) of the six taro corm samples from the field, and no amplification with a healthy taro corm sample (Fig. 10). The addition of SYBR Green dye resulted in a color change from orange to green in all samples containing the LAMP product, indicating DNA amplification and a true positive result (Fig. 10). Furthermore, no color change occurred in the non-D. fangzhongdai infected/inoculated samples, the negative controls, or healthy samples.
Detection of D. fangzhongdai in artificially infected taro samples. (A) Taro slices infected with different Dickeya spp. and a Pectobacterium species; D. fangzhongdai CFBP8607T, and PL145-PL147, D. undicola LMG30903, D. solani LMG25990, D. oryzae A5410, P. carotovorum LMG2404, and control (healthy taro). (B) Sigmoid curve diagram—only D. fangzhongdai infected taro slices were positive, no curve was observed for other Dickeya spp., P. carotovorum LMG2404, NC (negative control; healthy taro) and NTC (non-template control; water). (C) Rotor-Gene amplification reactions were used for visualization of the LAMP product by adding SYBR Green I dye. Green: positive amplification, orange: no amplification. Visualization of SYBR Green I dye results under UV exposure–fluorescence indicates positive amplification. (D) Dry bath amplification reaction was used for visualization of the LAMP product by adding SYBR Green I dye. Visualization of SYBR Green I dye results under UV exposure. 1–2, D. fangzhongdai CFBP8607T (purified gDNA and DNA from infected sample); 3–5, D. fangzhongdai PL145-PL147; 6–9 D. undicola LMG30903, D. solani LMG25990, D. oryzae A5410, P. carotovorum LMG2404; 10–11 healthy taro (NC) and water (NTC).
Detection of Dickeya fangzhongdai in artificially infected orchid samples. (A) Orchid stems infected with different Dickeya spp. and a Pectobacterium species; D. fangzhongdai CFBP8607T and PL145-PL147, D. undicola LMG30903, D. solani LMG25990, D. oryzae A5410, P. carotovorum LMG2404, and Control (healthy orchid). (B) Sigmoid curve diagram—only D. fangzhongdai infected orchid stems were positive, no curve was observed for other Dickeya spp., P. carotovorum LMG2404, NC (negative control; healthy orchid) and NTC (non-template control; water). (C) Rotor-Gene amplification reactions were used for visualization of the LAMP product by adding SYBR Green I dye. Green: positive amplification, orange: no amplification. Visualization of SYBR Green I dye results under UV exposure–fluorescence indicates positive amplification. (D) Dry bath amplification reaction was used for visualization of the LAMP product by adding SYBR Green I dye. Visualization of SYBR Green I dye results under UV exposure. 1–2, D. fangzhongdai CFBP8607T (purified gDNA and DNA from infected sample); 3–5, D. fangzhongdai PL145-PL147; 6–9 D. undicola LMG30903, D. solani LMG25990, D. oryzae A5410, P. carotovorum LMG2404; 10–11 healthy orchid (NC) and water (NTC).
Detection of Dickeya fangzhongdai in naturally infected taro samples. Positive amplification was observed with D. fangzhongdai CFBP8607T (positive control, tube 1) and two D. fangzhongdai-infected taro samples (tubes 5 and 6); no SYBR GREEN color change or fluorescence were observed for non-infected taro samples (tubes 2–4), healthy taro (tube 8) and NTC (non-template control; water; tube 9). No false positives and false negatives were detected.
Discussion
New Dickeya species are continually being described due to the broad natural diversity within the genus. To date, descriptions of 12 Dickeya species have been published and their genomes are available15, including the most recent additions of D. poaceiphila31, D. parazeae32, D. oryzae33 and D. colocasiae16. A previously published LAMP assay, designed to detect Dickeya at the genus level, successfully detected eight known Dickeya spp.34. The LAMP assay developed in the present study utilized a unique genomic region to provide a rapid, and cost-effective diagnostic assay for specific detection of D. fangzhongdai which can be applied easily at point-of-need. The assay was thoroughly validated with all Dickeya and Pectobacterium species and strains of other bacterial species.
A robust diagnostic assay requires stringent selection of a unique and specific genomic region that is well conserved within the target pathogen genomes25,30,35. In silico validation is required to confirm that the target region is present in all strains of the same species and absent in closely related species of Dickeya and Pectobacterium and other species that occupy the same ecological niche21. The unpublished genome of a locally isolated strain of D. fangzhongdai (PL145) was used to increase the breadth of in silico analysis, as the publicly available genomes of the species do not currently include Hawaiian strains (Table 1). Four regions were specific to D. fangzhongdai based on preliminary in silico genome alignment and analyses, and in vitro endpoint PCR tests (data not shown). High band intensity maintained across different strains of D. fangzhongdai and the lack of off-target amplification, even at the relatively low annealing temperature (57 °C), indicated that the MFS transporter was an ideal region for further LAMP assay development. This held true for both sets of primers (endpoint PCR and LAMP assays) designed for this region. LAMP remains an ideal option for diagnostic assay development; it is rapid, comparatively resistant to inhibition, and field deployable20. Results can be visualized in several ways, including colorimetric indicators, turbidity, and gel electrophoresis, which enables LAMP to be used effectively in many laboratory settings25.
The validation of the designed primers with the members of inclusivity and exclusivity panels is mandatory for confirmation of assay specificity25. The positive LAMP results with members of the inclusivity panel shown in Fig. 2; the LAMP assay detected all D. fangzhongdai strains—suggest a broad range detection capability of this assay. In the exclusivity panel, clear negative results occurred for the other species of Dickeya and Pectobacterium (Table 3; Fig. 2). Although two recently described Dickeya species, D. poaceiphila and D. parazeae, were absent from the exclusivity panel, our genomic-informed strategy of selecting the MFS target gene for LAMP development suggests the sequence of the MFS transporter gene is not present in their genomes (Fig. 1A). Hence, the developed LAMP possesses high specificity for distinguishing the species, D. fangzhongdai, from phylogenetically closely related species, plant associated niche-sharing species of additional genera, and common habitat matrices. The detection of a target pathogen can be affected adversely by the presence of plant inhibitors, but inhibitors do not affect all assays (developed using different techniques) equally20,36. A LAMP assay developed for specific detection of a Gram-positive bacterial pathogen, Rathayibacter toxicus, using the OptiGene Master Mix, was less affected by plant inhibitors than an endpoint PCR but more affected than an RPA assay20. The currently described assay showed no effect of plant inhibitors when crude extracts of artificially infected taro, orchid plant materials, and naturally infected taro (prepared using Plant Lysis Kit)25 were tested using the developed LAMP assay (Figs. 8, 9, 10). In contrast to other developed diagnostic methods37,38, this D. fangzhongdai LAMP assay is rapid, requiring only about 15–30 min from sample preparation to detection and does not require sophisticated lab equipment (Fig. 7C).
High sensitivity is a prerequisite for a diagnostic assay that eliminates the probability of false negatives that could have serious negative consequences if a pathogen enters and becomes established in new geographical locations. Therefore, we evaluated the detection limit of the D. fangzhongdai LAMP assay in the presence of three hosts (taro, onion, orchid) matrices. The 5 µl of crude extract of either taro, onion, or orchid was added to preparations of tenfold serially diluted genomic D. fangzhongdai DNA, but no effect of host matrices was observed, and the detection limit remained at 100 fg, which is equivalent to 18–20 genome copies (Figs. 4A–C, 5). These results indicate that the developed assay is highly sensitive, robust, and specific. In comparison, endpoint PCR was performed with the primer set designed using the same unique MFS transporter gene region added to tenfold serially diluted DNA of D. fangzhondai. The target pathogen was detected in all dilutions down to 10 pg (Fig. 4D). An additional spiked assay was performed to assess inhibition by adding 1 µl of plant genomic DNA extracted from healthy taro corm to each PCR reaction mixture containing 1 µl of tenfold serially diluted DNA of D. fangzhondai (Fig. 4E). A 10 pg limit of detection was maintained, but it was 100-fold less sensitive than LAMP detection.
Evaluation of the LAMP reaction by multiple operators on the Rotor-Gene platform provided confidence in detection consistency, robustness, and sensitivity (Fig. 6). Assay performance levels were achieved on multiple operating devices, showing consistency across differing amplification platforms (Fig. 7). Finally, our assay was not inhibited by the presence of plant cellular matrices, indicating that assay functionality is not necessarily dependent on sample purity and suggesting that less-stringent extraction methods may be used. These findings suggest that the LAMP assay can be used with confidence in laboratories as well as at point-of-need.
Methods
Any plant and plant materials used in this research comply with international, national and institutional guidelines.
Target gene selection
Complete genomes of Dickeya species and Pectobacterium species were retrieved from the NCBI GenBank genome database and aligned using the progressiveMauve algorithm in Geneious Prime ver. 2021.2.2. Local collinear blocks (LCB) formed from homologous regions of three strains of Dickeya fangzhongdai (QZH3, PA1, and ND14B), one strain of D. dadantii subsp. dieffenbachiae (NCPPB 2976), and one strain of Pectobacterium atrosepticum (SCRI1043), were sifted for regions having potential for diagnostic development. Potential target regions exclusive to D. fangzhongdai were identified; regions were verified to be unique and universal to the species via NCBI GenBank nucleotide BLAST (https://blast.ncbi.nlm.nih.gov). Potential target genes having high average and median similarity among D. fangzhongdai genomes were considered for further endpoint PCR examination (data not shown). Primers for endpoint PCR were developed using Primer3 v4.1.0 using methods described by Arif and Ochoa-Corona39. Primer specificity was verified using the aforementioned whole genomes of D. fangzhongdai via Geneious Prime, and all available nucleotide sequences in the NCBI database through Primer-BLAST.
LAMP primer design
The veracity of potential target regions were verified by BLAST and twelve publicly available (retrieved from the NCBI GenBank genome database) complete and draft genomes of D. fangzhongdai and one whole genome sequence representative of strains in Hawaii (Mohammad Arif, unpublished information; see Table 1). Dickeya species and Pectobacterium species (Table 3) were used to validate the specificity of the designed primer sets for potential target genes using end-point PCR, and as a result, the major facilitator superfamily (MFS) transporter was selected for LAMP primer development.
Six LAMP primers, forward inner primer, forward outer primer, backward inner primer, backward outer primer, forward loop primer, and backward loop primer, were developed using PrimerExplorer V5 (https://primerexplorer.jp/e/) for the unique region of MFS transporter gene after the specificity of the endpoint primers were validated in vitro. Primer sequences were adjusted manually as necessary to avoid variable regions. All primers used are listed in Table 2.
DNA extraction
Bacterial strains used for this study were streaked from glycerol stocks stored cultures at − 80 °C onto Dextrose Peptone Agar (DPA, peptone 10 g l−1, dextrose 5 g l−1, and agar 17 g l−1) or Nutrient Agar (NA) (BD, Becton Dickinson), and incubated at 26–28 °C for 1 or 2 days depending on the bacterial species. Bacterial gDNA was extracted from a pure culture according to the manufacturer’s instruction (DNeasy Blood and Tissue Kit, Qiagen, Germantown, MD). Plant gDNA was extracted from 100 mg of healthy tissues of orchid (Phalaenopsis sp.), onion (Allium cepa), and taro corm (Colocasia esculenta) using a DNeasy Plant Mini Kit (Qiagen).
Endpoint PCR and LAMP assays
Each LAMP reaction was composed of 15 μl Isothermal Mastermix (Optigene, West Sussex, UK), 2 μl of LAMP primer mix (1.6 μM of Df-FIP and Df-BIP, 0.2 μM of Df-F3 and Df-B3, and 0.4 μM of Df-LF and Df-LB), and 1 μl of template DNA; reactions were brought to a final volume of 25 μl with nuclease free water. LAMP reactions and subsequent melt-curve analyses were conducted using a Rotor-Gene Q (Qiagen). In this study, the D. fangzhongdai type strain CFBP8607T was used as the positive control and template-less reactions were added as negative controls. The amplification temperature of the reaction was set to 65 °C for 20 min. Melting curve analysis took place (99–80 °C) with increments of 0.2 °C. Amplification curves measuring fluorescence were analyzed by Rotor-Gene Q Software 2.3.1.49 to determine positive or negative calls. Melting curve data was used to further validate such results. Results were visually assessed after melting curve analysis by the addition of 3 μl of SYBR Green I (Molecular Probes Inc.) into each tube for result corroboration with fluorescent detections. Positive LAMP results appeared green under natural light and fluorescent during exposure to UV light after the addition of the dye. Negative results appeared orange under natural light, and fluorescence was absent during exposure to UV light.
Endpoint PCR amplification was used to validate the uniqueness of the primer pair designed to amplify a region of the MFS transporter gene. A subset of bacterial strains and plant samples from the LAMP inclusivity and exclusivity panels were selected for the target validation with endpoint PCR. Each 20 μl of endpoint PCR reaction consisted of 10 μl of 2× GoTaq Green Master Mix (Promega), 1 μl of both 5 μM AD-F1 and AD-R1 primers, 1 μl of template DNA, and 7 μl of nuclease-free water. The cycle conditions of the reaction included an initial denaturation of 95 °C for 3 min, followed by 35 cycles of denaturation at 95 °C for 20 s, annealing at 57 °C for 30 s, extension at 72 °C for 20 s, and a final extension at 72 °C for 3 min. PCR amplification of the MFS gene was conducted in a Bio-Rad T100 thermal cycler (Bio-Rad, Hercules, CA), and 15 μl of PCR amplicon was loaded into a 1.5% agarose gel stained with ethidium bromide (Invitrogen, Carlsbad, CA). Gel electrophoresis conditions were 100 V for 45 min. The PCR results were visualized under UV light using a gel documentation system (Bio-Rad Gel Doc-XR+).
Specificity assays
The specificity of the developed LAMP primer set was demonstrated using the extensive and complete inclusivity and exclusivity panels (Table 3). The inclusivity panel consisted of the D. fangzhongdai type strain, CFBP8607T, isolated from pear in China and 6 D. fangzhongdai strains collected from taro in Hawaii. The exclusivity panel, which was composed of 94 non-target gDNA samples, consisted of phylogenetically close bacterial species in the genera Dickeya and Pectobacterium, plant associated niche-sharing bacterial species of additional genera, and gDNA isolated from typical D. fangzhongdai habitat matrices. Habitat matrices consisted of soil and expected host plant species including potato, orchid, and taro. LAMP reactions were prepared according to the specifications described above. The complete inclusivity panel and a subset of the exclusivity panel were also assayed in validation of an endpoint primer set described above.
Sensitivity and spiked sensitivity assays
The limit of the developed LAMP assay’s analyte sensitivity was demonstrated and compared with those of endpoint PCR reactions performed in parallel. The endpoint PCR and LAMP reactions were conducted using the same protocols as previously stated. Sensitivity assays were performed on purified genomic DNA extracted from colonies of the D. fangzhondai type strain, CFBP8607, to determine the limit of LAMP detection. DNA was quantified using a Qubit 4 fluorometer (Thermo Fisher Scientific, Waltham, MA), serially diluted tenfold from 10 ng to 1 fg with nuclease-free water and assessed for both reaction types. The endpoint and LAMP reactions were performed by the addition of 1 μl of diluted DNA into each reaction tube.
Inhibitor resistance and the background impacts of plant tissues on the LAMP assay were demonstrated by performing sensitivity assays on gDNA spiked with plant crude lysates. Crude lysates were prepared from artificially infected orchid (stem), taro (corm), and onion (leaf) tissue; 1 g of tissue from each plant was processed using the Plant Material Lysis Kit (Optigene, Sussex, UK). Five μl of plant lysate was added to each tube with 1 μl serially diluted genomic DNA of D. fangzhondai CFBP8607T (1 fg–10 ng) as used above.
The interference of plant genomic DNA with the accuracy of endpoint PCR analyte detection was examined. One μl of plant genomic DNA extracted from healthy taro corm or orchid leaf was added into each PCR reaction mixture containing 1 µl of tenfold serially diluted genomic DNA of D. fangzhondai (1 fg–10 ng). A negative control lacking bacterial DNA but containing extracted genomic taro or orchid DNA was used to confirm the primer specificity.
Multi-operator blind tests
Multi-operator tests were performed by five independent teams at the University of Hawaii at Manoa to assess the reproducibility of the developed D. fangzhongdai LAMP assay. Each operator performed a blind test with eleven bacterial genomic DNA samples including 3 D. fangzhongdai strains, 7 other Dickeya species, and 1 Pantoea species from the inclusivity and exclusivity panels (Table 3), and a non-template negative control (water). The samples were arranged randomly without bias and undisclosed to the operators. LAMP assays were performed as previously described.
Multi-device detection
The LAMP assay reactions were performed in triplicate to test the efficacy of three different heating devices (Qiagen Rotor-Gene Q, Bio-Rad 100 thermocycler, and dry bath). Eleven fresh bacterial colonies, including D. fangzhongdai strains (CFBP8607T and PL145-PL150) and D. aquatica (LMG27354), D. lacustris (LMG30899), D. undicola (LMG30903), and D. oryzae (A5410) were grown on DPA/NA medium. Colonies were suspended in PCR tubes containing 100 μl of nuclease free water, boiled using the thermocycler at 95 °C for 10 min, and cooled; cooled colonies were centrifuged at full speed for 2 min. One μl of each supernatant was used as a template for LAMP reactions. The thermocycler conditions for LAMP assay were the same as the dry bath, 65 °C for 20 min.
LAMP assay validation using artificially and naturally infected samples
Diagnostic capability and robustness of the developed LAMP assay were demonstrated using artificially infected taro and orchid samples, and naturally infected taro corms. The inoculum of four D. fangzhongdai, A6317 and PL145-147, 3 other species of Dickeya, LMG30903, LMG25990, A5410, and a Pectobacterium sp. LMG2404 were prepared from overnight culture and inoculated onto taro corm and orchid stem slices as follows.
Taro corms and orchid stems were cleaned using tap water and submerged in 0.6% hypochlorite for 3 min followed by three washes with sterile water. A loopful (~ 10 μl) of overnight grown bacterial culture was stabbed inoculated into each taro and orchid slice, and the slices were incubated in petri dishes at room temperature for 12–18 h. A total of 100 mg tissue from each inoculated slice was macerated and used for crude DNA isolation using a Plant Material Lysis Kit (Optigene). Five μl of crude DNA isolated from artificially infected samples were deployed for the LAMP assay. Genomic DNA of D. fangzhongdai and healthy taro corm/orchid stem lysates were treated as positive and negative controls, respectively.
Data availability
The genomes were retrieved from NCBI GenBank database and are available with following accession numbers: D. fangzhongdai QZH3 (NZ_CP031507) https://www.ncbi.nlm.nih.gov/nuccore/NZ_CP031507, D. fangzhongdai ND14b (NZ_CP009460) https://www.ncbi.nlm.nih.gov/nuccore/NZ_CP009460, D. fangzhongdai PA1 (NZ_CP020872) https://www.ncbi.nlm.nih.gov/nuccore/NZ_CP020872, D. aquatica 174/2T (NZ_LT615367) https://www.ncbi.nlm.nih.gov/nuccore/NZ_LT615367, D. chrysanthemi Ech1591 (NC_012912) https://www.ncbi.nlm.nih.gov/nuccore/NC_012912, D. dadantii NCPPB 2976T (NZ_CM001978) https://www.ncbi.nlm.nih.gov/nuccore/NZ_CM001978, D. dianthicola ME23 (NZ_CP031560) https://www.ncbi.nlm.nih.gov/nuccore/NZ_CP031560, D. lacustris S29T (QNUT00000000) https://www.ncbi.nlm.nih.gov/nuccore/QNUT00000000, D. oryzae ZYY5T (SULL00000000) https://www.ncbi.nlm.nih.gov/nuccore/SULL00000000, D. parazeae Ech586 (NC_013592) https://www.ncbi.nlm.nih.gov/nuccore/NC_013592, D. poaceiphila NCPPB 569T (NZ_CP042220) https://www.ncbi.nlm.nih.gov/nuccore/NZ_CP042220, D. solani IPO 2222 (NZ_CP015137) https://www.ncbi.nlm.nih.gov/nuccore/NZ_CP015137, D. undicola 2B12 (JSYG00000000) https://www.ncbi.nlm.nih.gov/nuccore/JSYG00000000, D. zeae PL65 (NZ_CP040817) https://www.ncbi.nlm.nih.gov/nuccore/NZ_CP040817, P. parmentieri RNS 08-42-1AT (NZ_CP015749) https://www.ncbi.nlm.nih.gov/nuccore/NZ_CP015749, P. brasiliense 1692 (NZ_CP047495) https://www.ncbi.nlm.nih.gov/nuccore/NZ_CP047495, and P. carotovorum XP-13 (NZ_CP063242) https://www.ncbi.nlm.nih.gov/nuccore/NZ_CP063242.
References
Miyasaka, S. C., McCulloch, C. E. & Nelson, S. C. Taro germplasm evaluated for resistance to taro leaf blight. HortTech. 22, 838–849 (2012).
FAO-STAT, Food and Agriculture Organization of the United Nations, Value of Agricultural Production. https://www.fao.org/faostat/en/#data/QV/visualize (2019).
Otekunrin, O. A., Sawicka, B., Adeyonu, A. G., Otekunrin, O. A. & Rachoń, L. Cocoyam [Colocasia esculenta (L.) Schott]: Exploring the production, health, and trade potentials in sub-Saharan Africa. Sustainability. 13, 1–19 (2021).
Alič, S., Van Gijsegem, F., Pédron, J., Ravnikar, M. & Dreo, T. Diversity within the novel Dickeya fangzhongdai sp., isolated from infected orchids water and pears. Plant Pathol. 67, 1612–1620 (2018).
Zhou, A. et al. First report of Dickeya fangzhongdai causing soft rot in orchids in Canada. Plant Dis. https://doi.org/10.1094/PDIS-04-21-0771-PDN (2021).
Choi, E. D. et al. First report of bleeding canker of pear tree trunks caused by Dickeya fangzhongdai in Korea. Plant Dis. https://doi.org/10.1094/PDIS-09-20-1948-PDN (2021).
Huang, S. et al. First report of bacterial soft rot disease on taro caused by Dickeya fangzhongdai in China. Plant Dis. https://doi.org/10.1094/PDIS-10-20-2225-PDN (2021).
Tian, Y. et al. Dickeya fangzhongdai sp. nov., a plant pathogenic bacterium isolated from pear trees (Pyrus pyrifolia). Int. J. Syst. Evol. Microbiol. 66, 2831–2835 (2016).
Zhang, J. et al. Genomic analysis of the Phalaenopsis pathogen Dickeya sp. PA1, representing the emerging species Dickeya fangzhongdai 06 Biological Sciences 0604 Genetics. BMC Genomics 19, 1–16 (2018).
Tsai, W. A., Lin, P. R. & Huang, C. J. First report of Dickeya fangzhongdai causing soft rot disease of Welsh onion in Taiwan. Plant Dis. 103, 2665 (2019).
Ma, X., Bonasera, J. M., Asselin, J. A. E., Beer, S. V. & Swingle, B. First report of Dickeya fangzhongdai causing soft rot of onion in New York State. Plant Dis. https://doi.org/10.1094/PDIS-09-19-1940-PDN (2020).
Chen, B. et al. Bleeding canker of pears caused by Dickeya fangzhongdai: Symptoms, etiology, and biology. J. Integr. Agric. 19, 889–897. https://doi.org/10.1016/S2095-3119 (2020).
Hugouvieux-Cotte-Pattat, N., Jacot-des-Combes, C. & Briolay, J. Dickeya lacustris sp. nov., a water-living pectinolytic bacterium isolated from lakes in France. Int. J. Syst. Evol. Microbiol. 69, 721–726 (2019).
Oulghazi, S. et al. Dickeya undicola sp. nov., a novel species for pectinolytic isolates from surface waters in Europe and Asia. Int. J. Syst. Evol. Microbiol. 69, 2440–2444 (2019).
Arif, M., Czajkowski, R. & Chapman, T. Editorial: Genome-wide analyses of Pectobacterium and Dickeya species. Front. Plant Sci. https://doi.org/10.3389/fpls.2022.855262 (2022).
Boluk, G., Dobhal, S., Arizala, D., Alvarez, A. M. & Arif, M. Dickeya colocasiae sp. nov. isolated from wetland taro, Colocasia esculentum. BioRxiv. https://doi.org/10.1101/2022.01.14.476417 (2022).
Lebeau, A. et al. The GacA global regulator is required for the appropriate expression of Erwinia chrysanthemi 3937 pathogenicity genes during plant infection. Environ. Microbiol. 10, 545–559 (2008).
Reverchon, S. & Nasser, W. Dickeya ecology, environment sensing and regulation of virulence programme. Environ. Microbiol. 5, 622–636 (2013).
Mitra, D. Emerging plant diseases: Research status and challenges. Emerg. Plant Pathol. https://doi.org/10.1007/978-981-15-6275-4_1 (2021).
Arif, M., Busot, G. Y., Mann, R., Rodoni, B. & Stack, J. P. Field-deployable recombinase polymerase amplification assay for specific, sensitive and rapid detection of the US select agent and toxigenic bacterium, Rathayibacter toxicus. Biology. https://doi.org/10.3390/biology10070620 (2021).
Boluk, G. et al. Genome-informed recombinase polymerase amplification assay coupled with a lateral flow device for in-field detection of Dickeya species. Plant Dis. https://doi.org/10.1094/PDIS-09-19-1988-RE (2020).
Tian, Y. et al. Real-time PCR assay for detection of Dickeya fangzhongdai causing bleeding canker of pear disease in China. J. Integr. Agric. 19(4), 898–905 (2020).
Ocenar, J. et al. Development of a robust, field-deployable loop-mediated isothermal amplification (LAMP) assay for specific detection of potato pathogen Dickeya dianthicola targeting a unique genomic region. PLoS ONE 14(6), e0218868 (2019).
Notomi, T. et al. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 28, e63–e63 (2000).
Domingo, R. et al. Genome-informed loop-mediated isothermal amplification assay for specific detection of Pectobacterium parmentieri in infected potato tissues and soil. Sci. Rep. 11, 21948 (2021).
Kogovšek, P. et al. Rapid loop-mediated isothermal amplification assays for grapevine yellows phytoplasmas on crude leaf-vein homogenate has the same performance as qPCR. Eur. J. Plant Pathol. 148(1), 75–84 (2016).
Salazar, A., Ochoa-Corona, F. M., Olson, J. D., Babu, B. & Paret, M. Probing loop-mediated isothermal amplification (LAMP) targeting two gene-fragments of rose rosette virus. PLoS ONE 16(11), e0256510 (2021).
Nurul Najian, A. B., Engku Nur Syafirah, E. A. R., Ismail, N., Mohamed, M. & Yean, C. Y. Development of multiplex loop-mediated isothermal amplification (M-LAMP) label-based gold nanoparticles lateral flow dipstick biosensor for detection of pathogenic leptospira. Anal. Chim. Acta. 903, 142–148 (2015).
Becherer, L. et al. Simplified real-time multiplex detection of loop-mediated isothermal amplification using novel mediator displacement probes with universal reporters. Anal. Chem. 90(7), 4741–4748 (2018).
Dobhal, S. et al. Comparative genomics reveals signature regions used to develop a robust and sensitive multiplex TaqMan real-time qPCR assay to detect the genus Dickeya and Dickeya dianthicola. J. Appl. Microbiol. https://doi.org/10.1111/jam.14579 (2020).
Hugouvieux-Cotte-Pattat, N., Brochier-Armanet, C., Flandrois, J. P. & Reverchon, S. Dickeya poaceiphila sp. nov., a plant-pathogenic bacterium isolated from sugarcane (Saccharum officinarum). Int. J. Syst. Evol. Microbiol. 70, 4508–4514 (2020).
Hugouvieux-Cotte-Pattat, N. & Van Gijsegem, F. Diversity within the Dickeya zeae complex, identification of Dickeya zeae and Dickeya oryzae members, proposal of the novel species Dickeya parazeae sp. nov. Int. J. Syst. Evol. Microbiol. 71, 5059 (2021).
Wang, X. et al. Dickeya oryzae sp. nov., isolated from the roots of rice. Int. J. Syst. Evol. Microbiol. 70(7), 4171–4178 (2020).
Yasuhara-Bell, J. H., Marrero, G., Arif, M., de Silva, A. & Alvarez, A. Development of a Loop-Mediated Isothermal Amplification (LAMP) assay for the detection of Dickeya spp. Phytopathology 107, 1339–1345 (2017).
Larrea-Sarmiento, A., Stack, J. P., Alvarez, A. M. & Arif, M. Multiplex recombinase polymerase amplification assay developed using unique genomic regions for rapid on-site detection of genus Clavibacter and C. nebraskensis. Sci. Rep. 11, 12017 (2021).
Dobhal, S., Olsen, J., Arif, M., Garcia-Suarez, J. A. & Ochoa-Corona, F. M. A simplified strategy for sensitive detection of Rose rosette virus compatible with three RT-PCR chemistries. J. Virol. Methods. 232, 47–56 (2016).
Pritchard, L. et al. Detection of phytopathogens of the genus Dickeya using a PCR primer prediction pipeline for draft bacterial genome sequences. Plant. Pathol. 62(3), 587–596 (2013).
Van der Wolf, J. M. et al. Dickeya solani sp. nov., a pectinolytic plant pathogenic bacterium isolated from potato (Solanum tuberosum). Int. J. Syst. Evol. Microbiol. 64, 768–774 (2014).
Arif, M. & Ochoa-Corona, F. M. Comparative assessment of 5′ A/T-rich overhang sequences with optimal and sub-optimal primers to increase PCR yields and sensitivity. Mol. Biotechnol. 55(1), 17–26 (2013).
Alikhan, N. F., Petty, N. K., Zakour, N. L. B. & Beatson, S. A. BLAST Ring Image Generator (BRIG): Simple prokaryote genome comparisons. BMC Genom. 12, 1–10 (2011).
Hugouvieux-Cotte-Pattat, N., Jacot-des-Combes, C., Briolay, J. & Pritchard, L. Proposal for the creation of a new genus Musicola gen. nov., reclassification of Dickeya paradisiaca (Samson et al. 2005) as Musicola paradisiaca comb. nov. and description of a new species Musicola keenii sp. nov. Int. J. Syst. Evol. Microbiol. 71(10), 005037 (2021).
Acknowledgements
This work was supported by the USDA-ARS Agreement no. 58-2040-9-011, Systems Approaches to Improve Production and Quality of Specialty Crops Grown in the U.S. Pacific Basin; sub-project: Genome Informed Next Generation Detection Protocols for Pests and Pathogens of Specialty Crops in Hawaii. USDA National Institute of Food and Agriculture Hatch project 9038H and the Barry and Barbara Brennan Endowment also supported this work. We also thank Drs. Teresita Amore and Gamze Boluk for providing orchid plants and isolating D. fangzhongdai, respectively. The bacterial strains used in this study were stored and maintained by the National Science Foundation funded project (NSF-CSBR grant no. DBI-1561663). This research paper is the outcome of the course “PEPS/MBBE 627 Molecular Diagnostics: Principles and Practices”. The mention of trade names or commercial products in this publication does not imply recommendation or endorsement by the University of Hawaii.
Author information
Authors and Affiliations
Contributions
M.A. conceived and designed the study and reviewed the first draft; A.D., R.W., S.B., S.C., F.U., A.S., N.R., D.L.K., B.B. and T.P. performed the experiment, compiled the data and wrote the first draft of the manuscript; S.D. provided training, supervised the students for the lab experiment and helped in compiling data; D.A. helped in lab experiments, in silico analyses and created BRIG image; D.K. helped in lab experiments; F.O.C., E.A., J.O., J.P.B., D.M.J., J.F., J.P.S. and A.M.A. revised the manuscript and provided ideas and support for the final submission; all authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
DeLude, A., Wells, R., Boomla, S. et al. Loop-mediated isothermal amplification (LAMP) assay for specific and rapid detection of Dickeya fangzhongdai targeting a unique genomic region. Sci Rep 12, 19193 (2022). https://doi.org/10.1038/s41598-022-22023-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-22023-4
- Springer Nature Limited