Abstract
The boot-shaped respiratory complex I (CI) consists of a mitochondrial matrix and membrane domain organized into N-, Q- and P-modules. The N-module is the most distal part of the matrix domain, whereas the Q-module is situated between the N-module and the membrane domain. The proton-pumping P-module is situated in the membrane domain. We explored the effect of aging on the disintegration of CI and its constituent subcomplexes and modules in Drosophila flight muscles. We find that the fully-assembled complex remains largely intact in aged flies. And while the effect of aging on the stability of many Q- and N-module subunits in subcomplexes was stochastic, NDUFS3 was consistently down-regulated in subcomplexes with age. This was associated with an accumulation of many P-module subunits in subcomplexes. The potential significance of these studies is that genetic manipulations aimed at boosting, perhaps, a few CI subunits may suffice to restore the whole CI biosynthesis pathway during muscle aging.
Similar content being viewed by others
Introduction
Aging—defined as a progressive functional decline—is a major risk factor for several debilitating diseases. One of the hallmarks of aging is mitochondrial dysfunction1. Accordingly, identifying interventions that ameliorate or at least delay mitochondrial dysfunction in model organisms is a crucial need that would likely have therapeutic implications for human aging.
Studies in Drosophila have identified several genetic manipulations of mitochondrial proteins that have shown some promise in increasing lifespan. For example, induction of Dynamin-related protein 1 (Drp1), which is a major regulator of mitochondrial fission, during midlife in Drosophila increases lifespan2. Similarly, forced expression of the yeast alternative NADH:ubiquinone oxidoreductase, which catalyzes the transfer of electrons from NADH to ubiquinone via FAD without pumping protons across the mitochondrial inner membrane increases lifespan3,4,5. Further, overexpression of the alternative NADH:ubiquinone oxidoreductase reduces reactive oxygen species (ROS) production and increases several markers of CI activity3,4,5. Paradoxically, a restrained knockdown of several CI proteins also enhances longevity6,7. While the exact mechanisms involved in triggering the increased lifespan caused by mild CI disruption are still being unraveled, compensatory mitochondrial stress signaling cascades seem to contribute prominently. However, the question of whether boosting the expression of individual CI subunits can extend lifespan remains uncharted.
The boot-shaped mitochondrial CI is the most sophisticated component of the oxidative phosphorylation (OXPHOS) system8,9. Human CI has 45 subunits organized into two domains of the complex, dubbed the matrix and membrane domains10. There are three distinct functional modules of CI known as the N-, Q- and P-modules. The N-module contains the flavin mononucleotide (FMN) cofactor which accepts electrons from NADH in the mitochondrial matrix. The Q-module sits between the N-module and the membrane domain and transfers electrons to ubiquinone. The proton-pumping P-module is essentially the membrane domain [reviewed in11,12,13].
Fourteen core subunits form the catalytic centers of CI and are conserved from the ancestral enzyme in bacteria to the eukaryotic enzyme. Seven core subunits (NDUFS1, NDUFS2, NDUFS3, NDUFS7, NDUFS8, NDUFV1 and NDUFV2) are encoded by the nucleus, translated in the cytoplasm, and imported into the mitochondrion, while the other seven are encoded by mitochondrial DNA (mtDNA) and translated in the mitochondrion (i.e., ND1, ND2, ND3, ND4, ND4L, ND5 and ND6). The 31 remaining subunits are referred to as accessory or supernumerary subunits, as they are not directly involved in performing the bioenergetics functions of CI. During CI assembly, specific subcomplexes consisting of a few CI subunits form largely independently of each other and merge in a stereotypic fashion en route to forming the mature complex12,14,15. About a dozen or so CI assembly factors (CIAFs) have been identified. CIAFs are proteins that are usually found in association with specific subcomplexes and assist with the assembly process; but they are subsequently released when assembly is complete. The N-, Q- and P-modules are synthesized from specific subcomplexes or assembly intermediates that can be tracked by immunoblotting or complexome profiling techniques16,17,18,19.
CI function can deteriorate with age via several mechanisms. First, the complex can disintegrate with age, resulting in a reduction in the amount of CI that can be detected in blue native polyacrylamide gels. Second, the complex can be preserved with age but still result in a deterioration of CI function, perhaps, due to increased protein oxidation or other post-translational modifications on subunits. Third, with the identification of several phospholipid molecules tightly associated with several membrane domain subunits, oxidation of these lipid molecules, could also in principle impair CI function. Indeed, there are many other ways by which CI function can deteriorate with age. Consequently, identifying ways to preserve CI function is essential.
Although the alternative NADH:ubiquinone oxidoreductase from yeast can enhance, restore, or substitute for CI function in multiple organisms, there has been limited enthusiasm for overexpressing individual CI subunits as a possible means to improve CI function3,4,5,20,21,22,23,24,25. This is largely due to the large number of CI subunits in eukaryotic organisms, making it seem unlikely that forced expression of one or a few CI subunits or CIAFs could improve or ameliorate CI function with age. However, although the proteasome is comparable to CI as it has many subunits, it has been shown in C. elegans that overexpression of a single 19S proteasome subunit, RPN-6, is sufficient to confer proteotoxic stress resistance and increase longevity. In addition, overexpression of the human ortholog of RPN-6, PSMD11, is also sufficient to increase both proteasome assembly and activity in human embryonic stem cells26,27. As a consequence, we explored the effect of aging on the disintegration or stability of CI and its constituent subcomplexes and modules, with the view that uncovering such information may provide clues for up-regulating specific CI subunits or CIAFs to preserve, restore or enhance CI activity during aging. We find that the fully-assembled complex remains largely intact in 8-week-old flies. Further, aging was associated with an up-regulation of multiple membrane domain subunits—including some mtDNA-encoded subunits—in subcomplexes. The disintegration or stability of a few subunits was variable. Interestingly, however, NDUFS3, a component of the Q-module, was consistently down-regulated in subcomplexes with age at 25 °C; and Q-module biosynthesis was perturbed in flies exposed to mild thermal stress at 27 °C. As a result, we propose that genetic manipulations aimed at enhancing NDUFS3 expression, and perhaps a few of the subunits that showed stochastic expression changes with age, may be crucial in forestalling CI disintegration under stress or during muscle aging.
Results
The amount of fully-assembled CI is not altered during muscle aging or mild thermal stress
In this study, we explored the effect of aging on one of the properties of CI that can impact its function with age—the biogenesis and assembly of CI. We isolated mitochondria from thoraces of adult flies 1 week, 5 weeks and 8 weeks after they eclosed at 25 °C; and solubilized their mitochondrial membranes in digitonin at a ratio of 10 g of digitonin: 3 g of protein. Afterwards, we analyzed the integrity of their OXPHOS complexes using blue native polyacrylamide gel electrophoresis (BN-PAGE) and subsequent Coomassie and silver staining (Fig. S1a,b). Surprisingly, we failed to observe any reproducible changes in the amount of fully-assembled CI or other OXPHOS complexes with age (up to 8 weeks at 25 °C), in mitochondria from adult flight muscles (Fig. S1a,b). We also examined whether raising the adult flies under mild thermal stress at 27 °C—which should accelerate the aging process—would alter the amount of CI assembled. This was accomplished by monitoring CI assembly after eclosure at 25 °C and within 1 h after shifting the flies to 27 °C, as well as 2 and 4 weeks after shifting to 27 °C. Similar to observations made at 25 °C, OXPHOS integrity was normal even after 4 weeks at 27 °C (Fig. S1a,b). These results were corroborated with in-gel CI, complex II (CII), and complex IV (CIV) activities at both 25 °C and 27 °C (Fig. S1c). Taken together, we conclude that the amount of CI assembled remains largely intact in mitochondria from 4- and 8-week aged flight muscles at 27 °C and 25 °C, respectively.
The expression of dNDUFS3 in subcomplexes is down-regulated with age
Previous studies from our research group have revealed that the amount of CI assembled in Drosophila flight muscles as determined by BN-PAGE can sometimes appear normal although de novo biosynthesis of various CI subcomplexes may be impaired16,28. This is most likely due to the fact that once OXPHOS complexes are assembled in Drosophila flight muscles, they tend to be very stable to the point where very little change, if any, in the amount of OXPHOS complexes assembled can be detected by BN-PAGE, even when de novo CI biogenesis is hampered. We decided to exploit this property to ascertain how de novo CI biogenesis is altered in the aging muscle despite the relatively normal amounts of fully-assembled CI.
During CI biogenesis, subcomplexes consisting of a few CI subunits are first synthesized and eventually coalesce with other subcomplexes and CI subunits en route to forming the fully-assembled CI12,14,15. The three modules of CI are synthesized from specific subcomplexes (Figs. 1a and S2a). Using the same convention employed in our previous publications, for simplicity and clarity, we prefix all Drosophila melanogaster orthologs of CI subunits with the letter “d”; their exact CG numbers have been described previously15,16,28,29.
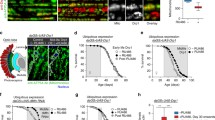
The expression of dNDUFS3 in subcomplexes is down-regulated with age. (a) A depiction of mitochondrial CI showing the matrix and membrane domains oriented almost orthogonally to each other, and the relative positions of the N-, Q-, PP- and PD-modules. A commercially available antibody that detects dNDUFS3 and antibodies we generated against dNDUFS7, dNDUFS8, dNDUFA5 and dNDUFA9 were used to examine the biogenesis of the Q-module. (b–f) Mitochondrial preparations from thoraces dissected from Dmef2-Gal4/w1118 (wt) flies raised at 25 °C or 27 °C for up to 8 weeks and 4 weeks, respectively, were analyzed by BN-PAGE, followed by immunoblotting with the antibodies indicated. The blots were imaged following a short exposure to detect the holoenzyme (fully-assembled CI) and supercomplexes, after which the region corresponding to the holoenzyme and supercomplexes was cut off, and the rest of the blot re-imaged after a longer exposure to detect the subcomplexes. The following antibodies were used: anti-NDUFS3 which detects dNDUFS3 (b), anti-dNDUFS7 (c), anti-dNDUFS8 (d), anti-dNDUFA5 (e) and anti-dNDUFA9 (f). The arrowheads mark the positions of the holoenzyme and the asterisks denote subcomplexes discussed in the text. Initiating and intermediary Q-module subcomplexes in panel (b) are denoted by # and *, respectively. We used the amount of holoenzyme detected by each antibody as a loading control for the subcomplexes. See Figures S2 and S3 for replicates and S7 for full-length gels and blots.
The Q-module consists of dNDUFS2, dNDUFS3, dNDUFS7, dNDUFS8, dNDUFA5 and dNDUFA9. Therefore, we monitored Q-module stability via Western blotting of blue native gels 1 week, 5 weeks, and 8 weeks after eclosure at 25 °C, with anti-NDUFS3 (which detects dNDUFS3), anti-dNDUFS7, anti-dNDUFS8, anti-dNDUFA5 and anti-dNDUFA9 antibodies (Figs. 1b–f and S2b–f). As the amount of the holoenzyme—fully-assembled CI devoid of supercomplexes—is not altered with age, we used the amount of the fully-assembled CI, shown in the short exposure panels, as loading controls for the subcomplexes which are shown in the long exposure panels. We observed an age-dependent decrease in the level of incorporation of dNDUFS3 in both initiating and intermediary subcomplexes of the Q-module at 25 °C (Figs. 1b, S2b and S3). A similar age-dependent decline in the incorporation of dNDUFS3 was observed in the intermediary Q-module subcomplex of samples isolated 2 and 4 weeks after incubation at 27 °C (Figs. 1b, S2b and S3). However, no reproducible pattern of expression was observed for dNDUFS7-containing subcomplexes at either 25 °C or 27 °C (Figs. 1c and S2c). The amount of dNDUFS8 that had incorporated into subcomplexes with age remained relatively uniform at 25 °C but was progressively reduced with age at 27 °C (Figs. 1d and S2d). Similar to dNDUFS7, no consistent pattern of expression was observed for dNDUFA5- or dNDUFA9-containing subcomplexes at either temperature (Figs. 1e,f, S2e,f). Hence, we conclude that while the pattern of expression of different Q-module subunits varies with age, one of the initiating subunits for Q-module synthesis, dNDUFS3, is down-regulated in CI subcomplexes during muscle aging.
Stochastic expression of N-module subunits in subcomplexes with age
Next, we turned our attention to checking how the integrity of the N-module changes with age. N-module integrity was examined by performing Western blots with antibodies that detect dNDUFV1, dNDUFV2, dNDUFV3, dNDUFS4, dNDUFA6, dNDUFA7 and dNDUFA12 (Figs. 2a and S4a). The stabilization or incorporation of both dNDUFV1 and dNDUFV2 was stochastic, as it appeared to be robustly upregulated at various time points, but with no obvious consistency between the replicates at 25 °C or 27 °C (Figs. 2b,c, S4b,c). The level of expression of dNDUFV3 did not show a consistent variation at either 25 °C or 27 °C (Figs. 2d and S4d). The amount of dNDUFS4 in subcomplexes appeared stable at 25 °C but was slightly upregulated at 27 °C (Figs. 2e and S4e). Finally, the extent of incorporation or stability of three additional N-module subunits—dNDUFA6, dNDUFA7 and dNDUFA12—also showed oscillations at various time points at 25 °C and 27 °C (Figs. 2f–h and S4f–h). Hence, we conclude that subcomplexes containing N-module subunits have a stochastic pattern of expression during normal aging at 25 °C or accelerated aging induced by thermal stress at 27 °C.
Stochastic expression of N-module subunits in subcomplexes with age. (a) An illustration of mitochondrial CI depicting the approximate positions of the N-, Q-, PP- and PD-modules. Biogenesis of the N-module was tracked by immunoblotting with antibodies against dNDUFV1, dNDUFV2, dNDUFV3, dNDUFS4, dNDUFA6, dNDUFA7 and dNDUFA12. (b–h) Mitochondrial preparations isolated from thoraces of wt flies at the time points shown and raised at the temperatures indicated were analyzed by BN-PAGE, followed by Western blotting with the antibodies indicated. The antibodies used were anti-dNDUFV1 (b), anti-dNDUFV2 (c), anti-dNDUFV3 (d), anti-dNDUFS4 (e), anti-dNDUFA6 (f), anti-dNDUFA7 (g), and anti-dNDUFA12 (h). The arrowheads mark the positions of the holoenzyme which was used as a loading control for the subcomplexes. The asterisks denote subcomplexes discussed in the text. See Fig. S4 for replicates and S7 for full-length gels and blots.
A variable wave of up-regulation of several PP-module subcomplexes occurs during muscle aging
The membrane domain can be subdivided into a proximal PP-module and a distal PD-module, corresponding to regions of the membrane domain closest to, and more distal from the region where the matrix and membrane domains adjoin, respectively (Figs. 3a and S5a) [reviewed in11,12,13]. The seven mtDNA-encoded CI subunits—dND1, dND2, dND3 dND4, dND4L, dND5 and dND6—are localized to the membrane domain. PP-module mtDNA-encoded CI subunits are dND1, dND2, dND3 dND4L and dND6. To determine how the integrity of the PP-module varies with age, we first examined the extent of incorporation of the mtDNA-encoded subunits of the PP-module. In general, subcomplexes containing the PP-module mtDNA-encoded CI subunits vacillated between a slight increase to remaining largely unchanged at 27 °C with age; however, these subcomplexes remained largely stable at 25 °C (Figs. 3b–f and S5b–f).
A variable wave of up-regulation of several PP-module subcomplexes occurs during muscle aging. (a) A diagram of mitochondrial CI showing the antibodies used to monitor the synthesis of the PP-and PD- modules. (b–l) Mitochondrial preparations obtained from thoraces of wt flies at the time points and temperatures indicated were analyzed by BN-PAGE, followed by Western blotting. The antibodies used were anti-dND1 (b), anti-dND2 (c), anti-dND3 (d), anti-dND4L (e), anti-dND6 (f), anti-dNDUFC2 (g), anti-dNDUFS5 (h), anti-dNDUFA11 (i), anti-dNDUFA13 (j), anti-dNDUFA8 (k), and anti-dNDUFA1 (l). See Fig. S5 for replicates and S7 for full-length gels and blots.
We also examined the stability of some subcomplexes containing nuclear-encoded PP-module subunits such as dNDUFC2, dNDUFS5, dNDUFA11, dNDUFA13, dNDUFA8 and dNDUFA1. The amounts of subcomplexes containing dNDUFC2, dNDUFS5, dNDUFA11 and dNDUFA13 were variable at midlife at 25 °C (4 weeks); nevertheless, by 8 weeks at 25 °C some subcomplexes containing these 4 subunits were generally elevated relative to what was observed at midlife (Figs. 3g–j and S5g–j). Similarly, the amount of dNDUFC2-containing subcomplexes was reduced at 2 weeks at 27 °C but increased at 4 weeks (Figs. 3g and S5g). However, we detected subcomplexes containing dNDUFS5, dNDUFA11 and dNDUFA13 that were increased at both 2 and 4 weeks at 27 °C (Figs. 3h–j and S5h–j). Additionally, we detected dNDUFA8- and dNDUFA1-containing subcomplexes that were also up-regulated after 8 weeks at 25 °C and 4 weeks at 27 °C, respectively (Figs. 3k,l and S5k,l). When considered as a whole, we conclude that a variable wave of upregulation of several PP-module subcomplexes occurs during muscle aging.
Multiple PD-module subcomplexes accumulate during muscle aging
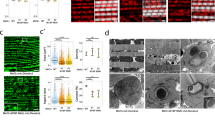
Lastly, we examined the effect of aging on subcomplexes containing PD-module subunits (Fig. 3a). Immunoblotting revealed that some PD-module subcomplexes containing dND5, dNDUFB1, dNDUFB2, dNDUFB3, dNDUFB5, dNDUFB6, dNDUFB8 and dNDUFB9 accumulated with age at both 25 °C and 27 °C (Figs. 4a–h and S6a–h). We also observed that dNDUFB5-, dNDUFB6- and dNDUFB8-containing subcomplexes were particularly up-regulated at 8 weeks and 4 weeks at 25 °C and 27 °C, respectively (Figs. 4e–g and S6e–g). Further, dNDUFB11-containing subcomplexes accumulated with age at 27 °C; but this was not apparent when we analyzed subcomplexes at 25 °C (Figs. 4i and S6i). Subcomplexes containing dNDUFB4, dNDUFB7 and dNDUFB10, however, showed a variable pattern of expression, appearing to be increased at some time points but not in a consistent manner at both 25 °C and 27 °C (Figs. 4j–l and S6j–l). We note that the up-regulation of PD-module subcomplexes was not restricted to the nuclear-encoded subunits as a subcomplex containing dND5 also displayed patterns of upregulation at both 25 °C and 27 °C (Figs. 4a and S6a). On the whole, we conclude that multiple PD-module subcomplexes accumulate during muscle aging.
Multiple PD-module subcomplexes accumulate during muscle aging. (a–l) Mitochondrial preparations from thoraces of wt flies at the time points and temperatures indicated were analyzed by BN-PAGE, followed by immunoblotting. The antibodies used for the immunoblots were anti-dND5 (a), anti-dNDUFB1 (b), anti-dNDUFB2 (c), anti-dNDUFB3 (d), anti-dNDUFB5 (e), anti-dNDUFB6 (f), anti-dNDUFB8 (g), anti-dNDUFB9 (h), anti-dNDUFB11 (i), anti-dNDUFB4 (j), anti-dNDUFB7 (k) and anti-dNDUFB10 (l). See Fig. S6 for replicates and S7 for full-length gels and blots.
Discussion
We have examined how the integrity of the fully-assembled respiratory CI and the level of expression or stability of multiple CI subcomplexes in Drosophila flight muscles changes with increased lifespan. Surprisingly, our analyses of OXPHOS integrity by blue native PAGE and in-gel OXPHOS activity assays revealed that the integrity of the fully-assembled CI is not conspicuously altered with age. Instead, it is the de novo biogenesis of some CI subcomplexes that varies with age, such that flight muscle aging is associated with distinct changes in subcomplexes of different modules of the complex. The age-dependent alterations of different subunits within the same module was not always concordant. As a case in point, there was a decline in the amount of subcomplexes containing dNDUFS3, but the pattern of expression of other Q-module subunits in subcomplexes varied with age. In addition, while the age-dependent subcomplex profiles of many N-module subunits was random, membrane domain subcomplexes generally trended toward up-regulation with age. In this context, we hypothesize that if these observations are reproducible in humans, it will indicate that therapeutic attempts to improve CI function in aged individuals may not necessarily require increasing the expression of all CI subcomplexes but perhaps only the select few that deteriorate during aging.
Interestingly, we also uncovered what appeared to be variable waves of compensatory up-regulation of some N-module subcomplexes. We postulate that this is part of an adaptive mitochondrial stress response triggered, conceivably, as a result of increased ROS formation. RNAi-mediated knockdown of the C. elegans ortholog of NDUFS3, nuo-2, increases lifespan, possibly due to the induction of compensatory mitochondrial stress responses30. Additionally, as dNDUFS3 is one of the subunits that houses Fe–S clusters, degeneration of dNDUFS3 in subcomplexes would likely expose the highly redox-sensitive Fe–S clusters to ROS formation, which in turn, could also cause the activation of ROS-dependent adaptive mitochondrial stress signaling cascades. Undoubtedly, these periodic cycles of induction of N-module subcomplexes during muscle aging would have to be studied in more detail to identify the precise regulatory controls of this phenomenon, as the signaling pathways regulating such events could also be exploited in therapeutic settings to stimulate various aspects of CI biogenesis.
Given that dNDUFS3 is one of the critical subunits that is involved in the initiation of CI biogenesis, it is tempting to speculate on why its expression level in subcomplexes deteriorates so rapidly with age. Possible reasons include a hypersensitivity to ROS-induced protein modifications due to its close proximity to Fe–S clusters. But this cannot be the sole reason as the amounts of several Fe–S cluster-interacting core subunits in subcomplexes (such as dNDUFS7, dNDUFS8, dNDUFV1 and dNDUFV2) did not fall with age, at least not as rapidly as dNDUFS3. These findings further highlight the master regulatory role that dNDUFS3 has in stimulating CI biogenesis, and now, as we report, in the stalling of biogenesis as well.
A stoichiometric imbalance between nuclear- and mtDNA-encoded CI subunits, referred to as mitonuclear protein imbalance, activates the mitochondrial unfolded protein response (UPRmt)31. While we have not examined the level of expression of all nuclear- and mtDNA-encoded CI subunits to assess whether a mitonuclear imbalance exists at the protein translational level to trigger the UPRmt, the robust accumulation of several membrane domain subcomplexes could also produce a membrane domain:matrix domain protein imbalance that may also activate the UPRmt, and possibly other components of the mitochondrial protein quality control machinery. This would have beneficial effects on the whole mitochondrial proteome. In this regard, the dichotomy in subcomplex accumulation between the membrane and matrix domains could elicit a compensatory adaptive response.
We note that although the amount of fully-assembled CI remained relatively unchanged after 8 weeks at 25 °C, it is possible that some of the subcomplexes that accumulate could be re-initiated into the CI biosynthesis pathway at a later time point to replenish degenerating modules or the fully-assembled CI. In this regard, it has been shown in a mammalian cell line that a CI salvage pathway that turns over the N-module at a higher rate than other parts of the complex exists17. This ensures CI homeostasis at a lower bioenergetic cost than if the whole complex had to be degraded and resynthesized. However, we chose not to analyze the period beyond 8 weeks as it would be difficult to examine the impact of other aspects of aging on the CI assembly process. We also note that the apparent stability of the fully-assembled CI could be a reflection of a steady state where individual oxidatively damaged CI subunits are replaced without requiring a replacement of the whole complex as has been described previously32.
To summarize, we have characterized how the CI subcomplex profile changes with age at 25 °C and 27 °C. In general, we find that the deterioration of CI biogenesis observed with increasing age at 25 °C is accelerated at 27 °C. We also find that the rate of decay of subcomplexes is uneven, raising the possibility that forced expression of a few CI subunits such as dNDUFS3, may be all that is necessary to re-initiate the whole CI biogenesis pathway in mid- to late-adult life. We further observed that while many membrane domain subcomplexes accumulate with age, the fate of matrix domain subcomplexes with age was less predictable. Although we have not explored how CIAFs are altered in subcomplexes with age, our findings hint at the possibility that overexpression of a few CIAFs may be able to restore a deficient CI biogenesis pathway during muscle aging.
Methods
Drosophila stocks and husbandry
Drosophila strains were reared in vials containing agar, yeast, molasses and cornmeal medium supplemented with propionic acid and methylparaben in humidified environmental chambers (Forma environmental chambers) on a 12-h:12-h dark:light cycle. Genetic crosses were set up between female flies of the genotype, y w; Dmef2-Gal4 (Ranganayakulu et al.,1996)33, and w1118 males at 25 °C. After the flies eclosed, they were maintained at 25 °C or 27 °C as explained below.
To examine the effect of normal aging on CI biogenesis, flies were aged at 25 °C for 1, 5 and 8 weeks prior to dissection of thoraces and subsequent mitochondria purification. To examine the effect of accelerated aging due to mild thermal stress on CI biogenesis, newly eclosed flies at 25 °C were immediately moved to 27 °C for up to an hour as the first time point used in our blue native immunoblots, and then 2 and 4 weeks prior to mitochondrial isolation from thoraces.
Mitochondria isolation
Mitochondrial purification was performed essentially as described previously28,34. Thoraces were quickly dissected and gently crushed with a dounce homogenizer (10 strokes) in 500 μl of a prechilled mitochondrial isolation buffer consisting of 250 mM sucrose, 0.15 mM MgCl2, 10 mM tris–HCl, pH 7.4, supplemented with Halt Protease inhibitors (Pierce). Tissue homogenates were centrifuged twice at 500g for 5 min at 4 °C to eliminate the cuticle and other insoluble material. Afterwards, the supernatant was recovered and centrifuged at 5000g for 5 min at 4 °C to obtain the mitochondria-enriched pellet, which was washed twice in the mitochondrial isolation buffer and preserved at − 80 °C until further processing.
Blue native polyacrylamide gel electrophoresis (BN-PAGE)
BN-PAGE was performed using NativePAGE gels (Life Technologies) and following the manufacturer’s protocol as described previously28,35.
Silver staining
Silver staining of blue native gels was performed with a SilverXpress staining kit (Life Technologies) as described previously28,35, following the manufacturer’s instructions.
In-gel complex I, II and IV activity
In-gel OXPHOS activities were performed as described previously28,35.
In-gel complex I activity was assessed by incubating the native gels in 0.1 mg/ml NADH, 2.5 mg/ml nitrotetrazolium blue chloride (NTB), 5 mM tris–HCl (pH 7.4) at 25 °C.
In-gel complex II activity was assessed by incubating the native gels in 20 mM sodium succinate, 0.2 mM phenazine methosulfate, 2.5 mg/ml NTB, 5 mM tris–HCl (pH 7.4) at 25 °C.
In-gel complex IV activity was assessed by incubating the native gels in 50 mM sodium phosphate (pH 7.2), 0.05% 3,3’- diaminobenzidine tetrahydrochloride (DAB), 50 μM horse heart cytochrome c at 25 °C.
Generation of peptide polyclonal antibodies
Rabbit polyclonal antibodies recognizing various segments of specific target proteins in Drosophila were generated by Biomatik using the synthetic peptides listed below:
Peptide antigen: Cys-RYWKESQDPKPTDFLSK | Target Protein: dNDUFA6 (CG7712) |
Peptide antigen: ELLTNKGTHEPSFLSP-Cys | Target Protein: dNDUFC2 (CG12400) |
Peptide antigen: NMDERFSRVMYQRDFRLTD-Cys | Target Protein: dNDUFA1 (CG34439) |
Peptide antigen: Cys-PNSRKWSNTELGVPKDGF | Target Protein: dNDUFB2 (CG40472) |
Peptide antigen: GEPYTVPHASTYKVES-Cys | Target Protein: dNDUFB3 (CG10320) |
Peptide antigen: SNEEQEFIKRKHEAT-Cys | Target Protein: dNDUFB4 (CG12859) |
Peptide antigen: THYMKPDVMPGPD-Cys | Target Protein: dNDUFB7 (CG5548) |
Peptide antigen: Cys-EKQYGKPDPKDLGHH | Target Protein: dNDUFB9 (CG9306) |
Peptide antigen: RRHGPVGSGMKEEAAH -Cys | Target Protein: dNDUFB10 (CG8844) |
Peptide antigen: NTSPKKDETITAPTS-Cys | Target Protein: dNDUFB11 (CG6008) |
Western blotting
Western blotting was performed as previously described16. In brief, following the separation of protein complexes on 3–12% precast Bis–Tris Native PAGE gels (Life Technologies), the proteins were transferred to polyvinylidene difluoride (PVDF) membranes (Bio-Rad). Subsequently, the PVDF membrane was blocked in 5% (wt/vol) nonfat dry milk (NFDM) in tris-buffered saline (TBS) for 30 min and incubated in the appropriate primary antibody dissolved in 2% BSA and 0.1% Tween 20 in TBS (TBST) overnight at 4 °C. Following the overnight incubation, the blot was rinsed four times for 10 min each in 0.1% TBST, blocked for 30 min in 5% (wt/vol) NFDM in TBST, and incubated for 2 h at room temperature with the appropriate HRP-conjugated secondary antibody dissolved in 2% BSA and 0.1% TBST. Afterwards, samples were rinsed four times for 10 min each in 0.1% TBST. Immunoreactivity was detected by a SuperSignal West Pico PLUS Chemiluminescent kit (ThermoFisher) and analyzed by a ChemiDoc gel imaging system from Bio-Rad. The primary antibodies used were anti-NDUFS3 (Abcam, ab14711), anti-dNDUFS728, anti-dNDUFS828, anti-dNDUFA529, anti-dNDUFA929, anti-dNDUFV116, anti-dNDUFV229, anti-dNDUFV328, anti-dNDUFS429, anti-dNDUFA6 (generated during this study), anti-dNDUFA729, anti-dNDUFA1228, anti-dNDUFS516, anti-dNDUFC2 (this study), anti-dNDUFA1 (this study), anti-dNDUFA828, anti-dNDUFA1128, anti-dNDUFA1329, anti-dND116, anti-dND216, anti-dND316, anti-dND4L16, anti-dND516, anti-dND616, anti-dNDUFB129, anti-dNDUFB2 (this study), anti-dNDUFB3 (this study), anti-dNDUFB4 (this study), anti-dNDUFB516, anti-dNDUFB616, anti-dNDUFB7 (this study), anti-dNDUFB816, anti-dNDUFB9 (this study), anti-dNDUFB10 (this study), and anti-dNDUFB11 (this study). Secondary antibodies used were goat anti-rabbit horseradish peroxidase (PI31460 from Pierce) and goat anti-mouse horseradish peroxidase (PI31430 from Pierce).
Data availability
All data needed to evaluate the conclusions in the paper are present in the paper and/or the Supplementary Materials.
References
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217. https://doi.org/10.1016/j.cell.2013.05.039 (2013).
Rana, A. et al. Promoting Drp1-mediated mitochondrial fission in midlife prolongs healthy lifespan of Drosophila melanogaster. Nat. Commun. 8, 448. https://doi.org/10.1038/s41467-017-00525-4 (2017).
Sanz, A. et al. Expression of the yeast NADH dehydrogenase Ndi1 in Drosophila confers increased lifespan independently of dietary restriction. Proc. Natl. Acad. Sci. USA 107, 9105–9110. https://doi.org/10.1073/pnas.0911539107 (2010).
Hur, J. H. et al. Increased longevity mediated by yeast NDI1 expression in Drosophila intestinal stem and progenitor cells. Aging (Albany NY) 5, 662–681. https://doi.org/10.18632/aging.100595 (2013).
Bahadorani, S. et al. Neuronal expression of a single-subunit yeast NADH-ubiquinone oxidoreductase (Ndi1) extends Drosophila lifespan. Aging Cell 9, 191–202. https://doi.org/10.1111/j.1474-9726.2010.00546.x (2010).
Copeland, J. M. et al. Extension of Drosophila life span by RNAi of the mitochondrial respiratory chain. Curr. Biol. 19, 1591–1598. https://doi.org/10.1016/j.cub.2009.08.016 (2009).
Owusu-Ansah, E., Song, W. & Perrimon, N. Muscle mitohormesis promotes longevity via systemic repression of insulin signaling. Cell 155, 699–712. https://doi.org/10.1016/j.cell.2013.09.021 (2013).
Fiedorczuk, K. et al. Atomic structure of the entire mammalian mitochondrial complex I. Nature 538, 406–410. https://doi.org/10.1038/nature19794 (2016).
Agip, A. A. et al. Cryo-EM structures of complex I from mouse heart mitochondria in two biochemically defined states. Nat. Struct. Mol. Biol. 25, 548–556. https://doi.org/10.1038/s41594-018-0073-1 (2018).
Agip, A. A., Blaza, J. N., Fedor, J. G. & Hirst, J. Mammalian respiratory complex I through the lens of Cryo-EM. Annu. Rev. Biophys. https://doi.org/10.1146/annurev-biophys-052118-115704 (2019).
Rhooms, S. K., Murari, A., Goparaju, N. S. V., Vilanueva, M. & Owusu-Ansah, E. Insights from Drosophila on mitochondrial complex I. Cell Mol. Life. Sci. https://doi.org/10.1007/s00018-019-03293-0 (2019).
Formosa, L. E., Dibley, M. G., Stroud, D. A. & Ryan, M. T. Building a complex complex: Assembly of mitochondrial respiratory chain complex I. Semin. Cell Dev. Biol. 76, 154–162. https://doi.org/10.1016/j.semcdb.2017.08.011 (2018).
Vartak, R. S., Semwal, M. K. & Bai, Y. An update on complex I assembly: The assembly of players. J. Bioenergy Biomembr. 46, 323–328. https://doi.org/10.1007/s10863-014-9564-x (2014).
Stroud, D. A. et al. Accessory subunits are integral for assembly and function of human mitochondrial complex I. Nature 538, 123–126. https://doi.org/10.1038/nature19754 (2016).
Garcia, C. J., Khajeh, J., Coulanges, E., Chen, E. I. & Owusu-Ansah, E. Regulation of mitochondrial complex I biogenesis in drosophila flight muscles. Cell Rep. 20, 264–278. https://doi.org/10.1016/j.celrep.2017.06.015 (2017).
Murari, A., Rhooms, S. K., Goparaju, N. S., Villanueva, M. & Owusu-Ansah, E. An antibody toolbox to track complex I assembly defines AIF’s mitochondrial function. J Cell Biol https://doi.org/10.1083/jcb.202001071 (2020).
Szczepanowska, K. et al. A salvage pathway maintains highly functional respiratory complex I. Nat. Commun. 11, 1643. https://doi.org/10.1038/s41467-020-15467-7 (2020).
Formosa, L. E. et al. Dissecting the roles of mitochondrial complex I intermediate assembly complex factors in the biogenesis of complex I. Cell Rep. 31, 107541. https://doi.org/10.1016/j.celrep.2020.107541 (2020).
Guerrero-Castillo, S. et al. The assembly pathway of mitochondrial respiratory chain complex I. Cell Metab 25, 128–139. https://doi.org/10.1016/j.cmet.2016.09.002 (2017).
Marella, M., Seo, B. B., Thomas, B. B., Matsuno-Yagi, A. & Yagi, T. Successful amelioration of mitochondrial optic neuropathy using the yeast NDI1 gene in a rat animal model. PLoS ONE 5, e11472. https://doi.org/10.1371/journal.pone.0011472 (2010).
DeCorby, A., Gásková, D., Sayles, L. C. & Lemire, B. D. Expression of Ndi1p, an alternative NADH:ubiquinone oxidoreductase, increases mitochondrial membrane potential in a C. elegans model of mitochondrial disease. Biochim. Biophys. Acta 1767, 1157–1163. https://doi.org/10.1016/j.bbabio.2007.07.003 (2007).
Bai, Y. et al. Lack of complex I activity in human cells carrying a mutation in MtDNA-encoded ND4 subunit is corrected by the Saccharomyces cerevisiae NADH-quinone oxidoreductase (NDI1) gene. J. Biol. Chem. 276, 38808–38813. https://doi.org/10.1074/jbc.M106363200 (2001).
Seo, B. B., Wang, J., Flotte, T. R., Yagi, T. & Matsuno-Yagi, A. Use of the NADH-quinone oxidoreductase (NDI1) gene of Saccharomyces cerevisiae as a possible cure for complex I defects in human cells. J. Biol. Chem. 275, 37774–37778. https://doi.org/10.1074/jbc.M007033200 (2000).
Seo, B. B. et al. Molecular remedy of complex I defects: rotenone-insensitive internal NADH-quinone oxidoreductase of Saccharomyces cerevisiae mitochondria restores the NADH oxidase activity of complex I-deficient mammalian cells. Proc. Natl. Acad. Sci. USA 95, 9167–9171. https://doi.org/10.1073/pnas.95.16.9167 (1998).
McElroy, G. S. et al. NAD+ regeneration rescues lifespan, but not ataxia, in a mouse model of brain mitochondrial complex I dysfunction. Cell Metab 32, 301-308.e306. https://doi.org/10.1016/j.cmet.2020.06.003 (2020).
Vilchez, D. et al. RPN-6 determines C. elegans longevity under proteotoxic stress conditions. Nature 489, 263–268. https://doi.org/10.1038/nature11315 (2012).
Vilchez, D. et al. Increased proteasome activity in human embryonic stem cells is regulated by PSMD11. Nature 489, 304–308. https://doi.org/10.1038/nature11468 (2012).
Murari, A. et al. Dissecting the concordant and disparate roles of NDUFAF3 and NDUFAF4 in mitochondrial complex I biogenesis. Science 24, 102869. https://doi.org/10.1016/j.isci.2021.102869 (2021).
Murari, A. et al. IDH2-mediated regulation of the biogenesis of the oxidative phosphorylation system. Sci. Adv. 8, eabl8716. https://doi.org/10.1126/sciadv.abl8716 (2022).
Dillin, A. et al. Rates of behavior and aging specified by mitochondrial function during development. Science 298, 2398–2401. https://doi.org/10.1126/science.1077780 (2002).
Houtkooper, R. H. et al. Mitonuclear protein imbalance as a conserved longevity mechanism. Nature 497, 451–457. https://doi.org/10.1038/nature12188 (2013).
Lazarou, M., McKenzie, M., Ohtake, A., Thorburn, D. R. & Ryan, M. T. Analysis of the assembly profiles for mitochondrial- and nuclear-DNA-encoded subunits into complex I. Mol. Cell Biol. 27, 4228–4237. https://doi.org/10.1128/MCB.00074-07 (2007).
Ranganayakulu, G., Schulz, R. A. & Olson, E. N. Wingless signaling induces nautilus expression in the ventral mesoderm of the Drosophila embryo. Dev. Biol. 176, 143–148. https://doi.org/10.1006/dbio.1996.9987 (1996).
Rera, M. et al. Modulation of longevity and tissue homeostasis by the Drosophila PGC-1 homolog. Cell Metab. 14, 623–634. https://doi.org/10.1016/j.cmet.2011.09.013 (2011).
Murari, A. & Owusu-Ansah, E. Analyzing the integrity of oxidative phosphorylation complexes in Drosophila flight muscles. STAR Protoc. 2, 101021. https://doi.org/10.1016/j.xpro.2021.101021 (2021).
Acknowledgements
We thank past and present members of the Owusu-Ansah lab for general discussions. We also thank Henry Colecraft, Jeanine D’Armiento, Alexander Galkin, Wes Grueber, Laura Johnston, Andrew Marks, Martin Picard, Liza Pon, Michael Schlame, Eric Schon and Mimi Shirasu-Hiza for fly stocks, reagents, and critical discussions. We acknowledge the Bloomington Drosophila Stock Center for various fly strains. Work in the Owusu-Ansah lab is supported by NIH grants AR077312 (R21) and GM124717 (R35) and a Schaefer Research Scholars award.
Author information
Authors and Affiliations
Contributions
K.F.B.H. performed most of the experiments in Figs. 1, 2, 3 and 4 and Figs. S1, S2, S4, S5, S6 and S7, with some help from B.M. and A.M. A.M. performed the experiments in Fig. S3. K.F.B.H., A.M., B.M. and E.O.A. analyzed and discussed results. E.O.A. conceived and managed the project. E.O.A wrote the manuscript and got feedback from other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hossain, K.F.B., Murari, A., Mishra, B. et al. The membrane domain of respiratory complex I accumulates during muscle aging in Drosophila melanogaster. Sci Rep 12, 22433 (2022). https://doi.org/10.1038/s41598-022-26414-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-26414-5
- Springer Nature Limited