Abstract
Periodontitis is one of the main frequent intraoral diseases. Pathogenesis triggers are the immune responses with pro-inflammatory cytokines production and non-coding RNAs expression. The purpose of the present study was to evaluate the involvement of selected miRNAs in various stages of periodontitis and their relationship with the levels of inflammatory mediators in gingival crevicular fluid (GCF). For this study, 36 subjects (21 with periodontal disease, 15 healthy controls) were selected with an age mean of 59.1 ± 3.7 years. Clinical parameters included plaque index, gingival index, sulcus bleeding index, pocket depth, and clinical attachment level. The GCF samples were taken using capillary paper. The levels of miRNAs in GCF were estimated using a Real-Time PCR and TNFα and IL-6 levels were assessed by enzyme-linked immunosorbent assay (ELISA). The results indicated that the miRNA-103a-3p, miRNA-23a-3p, miRNA-15a-5p, and miRNA-223-3p were significantly upregulated with respect to healthy controls. Significant differences were observed for miRNA-23a-3p, miRNA-103a-3p and miRNA-423-5p levels in accord with the disease stages. Inflammatory mediators evaluated in GCF correlate well with the clinical parameters and the severity of the periodontal disease. miRNAs can represent biomarkers of disease stage and can be investigated as a possible therapeutic target, as well as levels of TNFα and IL-6 may drive the disease progression by acting as prognostic markers.
Similar content being viewed by others
Introduction
The frequent physical and mechanical stress of tissue in the periodontal region and the highly dynamic remodeling determines the development of periodontitis, one of the main chronic inflammatory diseases affecting more than 50% of the world population1. Periodontitis is influenced by the interplay between subgingival microbial dysbiosis, periodontal tissue inflammation, tissue destruction, and genetic alterations, with an imbalanced host response2. A new periodontitis classification developed by a consensus conference between American Academy of Periodontology (AAP) and European Academy of Periodontology (EFP) has been adopted, providing an assessment of disease grading (A,B,C) and staging (I, II, III, IV). Staging depends on the severity of the disease at presentation and its management complexity, while grading provides additional information on the biological characteristics of the disease, including an analysis of periodontitis progression, based on patient history (Tables 1, 2)3.
The stage I represents the early stages of attachment loss; the stage II represents established periodontitis in which periodontal examinations can identify damages at the tooth support. Stage III of periodontitis produces significant damage to the attachment apparatus and tooth loss may occur, in the absence of advanced treatment; this stage is characterized by the presence of deep periodontal lesions and presence of deep intra-bony defects. Stage IV causes significant damage to periodontal support and may cause significant tooth loss, resulting in loss of masticatory function4.
The relationship between immune system and periodontal disease has been highlighted in the regulation of the pathogenetic mechanism of periodontitis, both in the initial stages and in the progression toward a chronic condition. The pathogens responsible for the initiation of periodontal disease recall the mediators of innate immunity, neutrophils, and macrophages at the damage site, producing cytokines and other inflammatory products5. When there is no resolution of the lesion, the specific immune response is activated with the intervention of lymphocytes, macrophages, and dendritic cells. The majority of literature confirm that neutrophils are hyperactive in periodontitis, in particular in severe, early onset forms, at the same time investigators have observed reduced neutrophil functions6. The dysregulation of inflammation and immune pathways, responsible for tissue damage, progression and chronicization of periodontal disease, is highly linked to genetic and epigenetic alterations.
Being epigenetic modulators, the microRNAs (miRNAs), short sequences (19–24 nucleotides in length) of non-coding RNAs, interact with the 3′ untranslated regions (3′ UTR) of target messenger RNAs (mRNA) causing mRNA degradation, translational suppression6 and interfere with the post-transcriptional expression of multiple target genes playing an important regulator role of the immune response. Indeed, miRNAs are responsible for neutrophils’ activity control and their migration to the inflammatory site, as they are involved in the regulation of mRNA sequence stability as well as the inflammatory mediator’s production7,8.
In vivo and in vitro studies have defined the miRNAs as determinants of periodontitis’ pathogenesis, as promoters of microbial persistence and as deregulators of the innate and adaptive immune response which become ineffective against pathogens and lead to the worst prognosis of the disease8,9,10. Moreover, miRNAs expression is also related to osteogenesis and osteoclastogenesis, mainly related to osteoclasts’ activity in bone-resorption and the osteoblasts’ proliferation and differentiation, both necessary for bone tissue homeostasis11,12,13.
Accumulating evidence suggests that the differential expression of miRNAs in gingival tissues and fluids can be causally linked to periodontitis and be promising candidates as potential disease biomarkers14,15.
Based on their characteristics in terms of expression stability and profiles, observed in serum and plasma, saliva and more recently also in the gingival crevicular fluid (GCF), miRNAs are emerging as tools for the detection of alterations of the oral cavity and of systemic diseases14,15. These biofluids can be easily and quickly collected with a minimally invasive procedure, bestowing great potential for the diagnostic and prognostic value of periodontal disease.
Recent investigations on inflammatory response and bone tissue homeostasis have reported a significant increase in salivary expression levels of miR-146a, miR-155 and miR-223, inhibiting osteoclastogenesis and driving the nuclear factor kappa-light-chain-enhancer of activated B cells (NF-kB) pathway activation, in patients affected by periodontitis16,17,18. Moreover, in chronic periodontitis, it has been confirmed a relationship between salivary miRNAs expression and periodontitis' pathological processes, with the highlighting of the contribution of miRNAs (miR-142-3p, miR-146a, miR-155, miR-203, and miR-223) involved in the regulation of bacterial infections, inflammation, and immune response16,19,20.
MiR-223, in particular, has been reported as key regulator of the innate immune responses mostly in association with the ability of the myeloid lineage differentiation21, inflammatory response modulation and infection development22.
Similarities were reported between miR-223 and miRNAs 15 and 23 based on inflammatory target pathways regulation23,24.
Among the miRNAs overexpressed during inflammation, the miR-103 is reported to target tumor necrosis factor (TNF)α, interleukin (IL)-17, IL-1β and the pathway NF-kB25. These miRNAs are also related to the osteoclast development process and bone metabolism, such as the miRNA 42326.
Furthermore, literature evidence suggests an association of miRNAs with inflammatory cytokines such as IL-1α, IL-1β, TNFα, IL-6 and IL-8, which are known to play an active role in periodontal tissue disease and whose unregulated production appears to be involved in chronic leukocyte recruitment and promoting tissue destruction5.
Thus, the aim of this study was to explore for the first time the involvement of miR-15a-5p, miR-23a-3p, miR223-3p, miR-103a-3p, miR-423-5p and TNFα, IL-6 production in periodontal disease pathological and healthy GCF.
Results
Demographic and periodontal parameters
The demographic characteristics of the enrolled subjects are summarized in Table 3. The mean age of participants was not significantly different in the periodontal disease group in comparison with the healthy controls (HC). Also, gender distribution, Body Mass Index (BMI), and lifestyle habits were not statistically different within and between the groups.
As the more accurate indicators of the healthy periodontal support structure around a tooth, clinical attachment level (CAL) and maximum probing depth (MPD) have been used to define the disease stage. The clinical attachment level was measured with probe from cemento-enamel junction (CEJ) to the bottom of periodontal pocket. The maximum probing depth was measured from the gingival margin to the bottom of the gingival sulcus/ pocket. As shown in Table 4, patients were grouped into 3 disease stages: mild which correspond to grade II (8 patients), moderate as grade III (6 patients) and severe as grade IV of new periodontitis classification (7 patients).
TNFα and IL-6 quantification in periodontitis and HC groups
To define the inflammatory condition in patients with periodontitis, TNFα and IL-6 levels were quantified by ELISA assay in patients’ GCF. As reported in Table 5, a significant and progressive increase was observed in relation to the disease stage. Significant higher levels of both cytokines were observed in periodontitis patients at mild, moderate and severe stages with respect to HC (p < 0.001), underlying the involvement of these inflammatory cytokines in the pathogenesis and progression of periodontitis.
miRNAs expression levels in GCF in periodontitis and HC groups
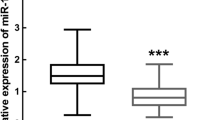
The expression levels of miRNAs 103a-3p, miR423-5p, miR23a-3p, miR15a-5p, and miR223-3p in periodontitis patients and HC were analyzed by Real-time PCR. Figure 1 shows that miR103a-3p, miR23a-3p, miR15a-5p, and miR223-3p were significantly upregulated in GCF collected in the periodontitis group in comparison with the HC, while miR423-5p showed no difference in fold change.
Relative expression levels of miRNAs detected in GCF of patients with periodontitis compared to HC. Data are reported as fold change ± C.I. with respect to HC, equal to 1. The statistical significances shown are **p < 0.01; ***p < 0.001. Data were analyzed using Stata version 15 and GraphPad Prism 6.
miRNAs expression levels in periodontitis patients with different disease stages
To better understand the distribution of miRNAs expression in patients with periodontitis at different severity, we analyzed their expression levels based on disease stages. In detail, although an increase in expression levels was observed for miR-15a-5p and miR-223-3p in comparison with HC, there were no significant differences between disease stages (Fig. 2a, b). The expression levels of miR-23a-3pshowed a fold change of 11.32 in patients with mild periodontitis compared to HC, while, in moderate and severe periodontal disease, the expression levels were 1.9 and 1.4-fold respectively. Furthermore, in our patients, miR-23a-3p was significantly increased in mild periodontitis patients also in comparison with moderate and severe periodontitis patients (p < 0.001) (Fig. 2c).
Boxplots of relative expression levels of (a) miR-15a-5p, (b) miR-223-3p, (c) miR-23a-3p, (d) miR-103a-3p and (e) miR-423-5p in periodontal disease patients GCF sample, at mild, moderate and severe disease stages. Data are reported as fold change ± C.I. with respect to HC, equal to 1. The statistical significances shown are *p < 0.05; **p < 0.01; ***p < 0.001. Data were analyzed using Stata version 15 and GraphPad Prism 6.
About miR-103a-3p, increasing expression levels were observed in relation to the different disease stages, with significant differences between mild and severe stages (p = 0.023), with a variation of 1.27-fold (Fig. 2d). Similarly, miR-423-5p showed an up-regulated expression in relation to disease severity, with a significant increase in severe periodontal disease patients, equal to 3.68-fold, in relation to patients with periodontitis in mild (0.39-fold) and moderate stage (0.45-fold) (Fig. 2e).
The distribution of differential miRNAs expression was confirmed by the radar plot analysis. As shown in Fig. 3, in patients with mild periodontitis, a higher distribution area was reported for miR-23a-3p, supporting the involvement in the early phases of periodontal disease. Moreover, with a lower join area, miR-23a-3p was present also in patients with moderate disease. Furthermore, a major distribution of miR-103a-3p was observed in patients with moderate and severe periodontitis, while in patients with severe disease, we observed a higher distribution of miR-423-5p.
Correlation analysis of miRNAs expression levels and inflammatory mediators
The analysis of the correlation of miRNAs with inflammatory responses in patients with periodontitis pointed out that the cytokines levels were significantly correlated between them and with the disease stage, confirming the increased release in parallel with the disease worsening. Results shown in Table 4 indicate a positive correlation between GCF levels of miR-103a-3p and TNFα in patients with moderate (r = 0.418, p < 0.001) and severe (r = 0.555, p < 0.01) periodontal disease. Moreover, we observed a positive correlation between miR-423-5p and cytokines in patients at severe disease stage (TNFα: r = 0.584, p < 0.01; IL-6: r = 0.526, p < 0.05), in accordance with the up-regulated miRNAs expression and higher inflammatory cytokines levels. Moreover, a negative correlation between miR-23a-3p and TNFα (r = -0.891, p < 0.01) and IL-6 (r = − 0.503, p < 0.05) was observed in the early stage of periodontal disease (Table 6).
Finally, genes were retrieved by GeneCards (https://www.genecards.org/) database27 using “inflammation and bone metabolism” as keyword. As reported in Table 7 the intersection of TNFα, IL-6, miR-103a-3p, miR-423-5p, miR-23a-3p, miR-15a-5p and miR-223-3p were portrayed by the GeneCards Inferred Functionality Score and the relevance score.
Discussion
Periodontitis, one of the top ten highly prevalent diseases worldwide28, presents an amplified inflammatory immune response as a hallmark. The prevalent inflammatory microenvironment, affecting the periodontium and compromising tooth-supporting apparatuses (gingiva, cementum, periodontal ligament, and alveolar bone), negatively influences the general state of health, leading to the persistence of chronic low-grade inflammation for lengthened periods and increasing the risk for cardiovascular, cerebrovascular and respiratory diseases, with a higher probability for metabolic diseases development29,30,31. Much of the tissue damage that occurs during inflammation can be attributed to TNFα and IL-6. The balance of TNFα and IL-6 mirrors the degree of the inflammatory burden contributing to periodontal inflammation32.
TNFα, a pro-inflammatory cytokine released by macrophages, is known for its extensive role in periodontitis-mediated bone loss. Increased concentration of TNFα observed in periodontitis correlates closely with tissue destruction and immune response33. Meanwhile, IL-6, a multifunctional cytokine, has several biological activities, including B-lymphocyte differentiation, T-lymphocyte proliferation, and the stimulation of immunoglobulin (Ig) secretion by B-lymphocytes34. Particularly, IL-6 induces bone resorption by itself and in conjunction with other bone-resorbing agents35.
Our results showed that TNFα and IL-6 production levels were significantly increased in patients with periodontal disease in comparison with HC. Additionally, significant differences in TNFα and IL-6 levels were found between patients in accord with the mild, moderate, and severe periodontitis stages. These data agree with previous findings showing significantly elevated salivary concentrations of IL-6 and TNFα in periodontitis patients36,37,38,39,40,41 and suggest that IL-6 and TNFα GCF levels may have the potential to distinguish different phases of periodontitis.
The miRNAs play a role in the production of pro-inflammatory cytokines by both a “direct” and “indirect” modulation, via the targeting activators or repressors. MiRNAs in body fluids have been reported as high stable molecules with resistance to degradation, suggesting great potential as biomarkers, also because they represent one of the most abundant classes of molecules involved in the regulation of gene expression16.
Despite significant advances in miRNAs involvement in periodontal disease, the regulatory mechanisms are still to be investigated, also due to heterogeneity of the miRNAs’ expression.
Taking in mind that the inflammation in periodontal disease often results in alveolar bone loss and disruption of connective tissue homeostasis42,43,44, we have evaluated in GCF the expression of miR-15a-5p, miR-23a-3p, miR-223-3p, miR-103a-3p, and miR-423-5p, involved in the regulation of innate immune response, cell metabolism, and inflammation, in patients with periodontal disease at three different disease stages (mild, moderate and severe).
The expression of selected miRNAs was up-regulated in patients with respect to HC, in agreement with previous observations12,45.
In accord with its role as a positive regulator of the inflammatory microenvironment and negative regulator of osteogenic differentiation in various cell types46,47, we have observed a high expression for miR-23a-3p in mild periodontal disease, probably acting as a starter for the bone metabolism alteration. Furthermore, miR-103a-3p, related to inflammation and bone metabolism targeting IL-1, prostaglandin E2 (PGE2), and TNFα, was up-regulated in periodontal inflamed tissue disease48,49. In our patients, miR-103a-3p was increased mostly in the moderate disease level. About miR-423-5p, participating in osteoclast metabolisms26, a down-regulation in mild and moderate disease stages was observed with respect to HC, although an over-expression in severe disease stage patients was detected. These data are in accordance with miR-423-5p involvement in osteoclastogenesis, which represents a critical step in periodontal disease physiopathology50, characterized by an active inflammatory state, with high levels of TNFα, needed for osteoclastic cellular maturation. Furthermore, it has been observed that miR-423-5p has a better positive relation with TNFα and IL-6, with the ability in NFkB pathway up-regulation. Referring to miR-223-3p, Bauernfeind et al. suggested its antimicrobial effect against periodontal pathogens51 and regulatory role of inflammation through neutrophil recruitment in chronic inflammatory sites. miR-223-3p is also involved in the differentiation of several immune cells, particularly macrophages, by influencing their activation patterns52. Therefore, has been reported that miR-223-3p is up-regulated in periodontal disease, to regulate tissue homeostasis, to control osteoblast differentiation, and to avoid alveolar bone loss, which are emblems of periodontitis53. In our study groups, although a slight increase in miR-223-3p expression levels has been observed in periodontal disease patients in comparison with HC, no significant differences between the three disease stages were observed, demonstrating a non-decisive role in the regulation of disease progression. miR-15a-5p is reported to be closely related to several diseases, mediating the inflammatory process, but its role is not plenty investigated in periodontal disease. Luan et al. found the up-regulation of miR-15a-5p in both humans and mice affected by periodontitis7. In our patients, although not significantly, miR-15a-5p expression showed a reduced trend during disease progression in parallel with the increase of TNFα and IL-6 levels in GCF. Our results are encouraging, suggesting a different involvement of selected miRNAs in relation to the disease stages. The strong correlation found between miRNAs expression and inflammatory cytokine levels in relation to disease severity supports the deepening of their role as disease markers. Also, it is very interesting that this can be explored in GCF, supporting its high diagnostic, prognostic and therapeutic potential. However, their clinical application still needs to be defined and investigated.
Materials and methods
Patient selection and clinical periodontal parameter examination
Fifteen periodontally HC (aged 57–63 years) and twenty-one patients with periodontal disease (aged 56–61 years), all non-smokers, were referred to the Dental Clinic of the Department of Innovative Technologies in Medicine and Dentistry, “G. d’Annunzio” University, Chieti-Pescara, Italy, and participated in this study.
The inclusion criteria were as follows:
-
Patients between 18 and 75 years
-
Patients with periodontal disease
-
Need of periodontal therapy
The exclusion criteria were as follows:
-
Poor oral hygiene
-
Uncontrolled diabetes mellitus
-
Immune diseases
-
Smoking
-
Bruxism
Every patient and healthy individual enrolled in the investigation provided written informed consent. In addition, the research related to human use has complied with all the relevant national regulations, in accordance with the tenets of the Helsinki Declaration, and has been approved by the experimental research ethics committee of the “G. d’Annunzio” University (254/2019). Diagnosis and classification of the periodontal disease were done according to the new classification, defined in the context of the 2017 World Workshop3, based on clinical and radiographic data. CAL and MPD were measured as previously reported54, to define the disease stages in individual patients. Healthy individuals assumed as the control group did not present any sign or symptom compatible with periodontal disease.
Gingival crevicular fluid samples collection
The GCF sample was taken from a single-rooted tooth, due to its easy access and to avoid errors associated with GCF sample collection, both in patients with periodontitis and HC. Before placing the absorbing paper strip within the sulcus, the supragingival plaque was removed and relative isolation was performed using cotton rolls and aspiration to prevent saliva contamination. Subsequently, the filter paper strip Periopaper (Oraflow, New York, NY, USA) was placed in the gingival sulcus until resistance was felt and left in this position for 30 s following the routine procedure. The filter paper strip was introduced in a sterile tube and stored at − 80 °C, until the subsequent analysis. Meanwhile, the demographic and habit data of the patients enrolled in both groups have been collected on specific Case Report Forms.
RNA extraction and quantification from gingival crevicular fluid
Periopapers were incubated in phosphate-saline buffer (PBS) solution pH 7 for 30 min at room temperature. To recover all the solution contained in the strip, each sample was centrifuged at 500 rpm for 10 min. Cell-free total RNA (including miRNAs) was isolated from 200 µL of PBS solution containing the biological material from the Periopaper using a miRNeasy Serum/Plasma kit (Qiagen, Hilden, Germany). RNA was eluted with 25 µL of RNAse-free water. Total RNA concentrations were quantified using NanoDrop ND 2000 UV-spectrophotometer (Thermo Scientific, Delaware, USA).
Quantitation of miRNAs by real-time PCR
Reverse transcription (RT) reactions were performed using the miRCURY LNA RT Kit (QIAGEN, Hilden, Germany). Real-time PCR reactions were performed in triplicate, in 10µL reaction volumes, using miRCURY LNA SYBR Green PCR Kit (QIAGEN, Hilden, Germany). miRNAs gene expressions were evaluated for the following sequences:
hsa-miR-23a-3p: 5′AUCACAUUGCCAGGGAUUUCC.
hsa-miR-423-5p: 5′UGAGGGGCAGAGAGCGAGACUUU.
hsa-miR-15a-5p: 5′UAGCAGCACAUAAUGGUUUGUG.
hsa-miR-223-3p: 5′UGUCAGUUUGUCAAAUACCCCA.
has-miR-103a-3p: 5′AGCAGCAUUGUACAGGGCUAUGA.
UniSp6: 5′CUAGUCCGAUCUAAGUCUUCGA.
using UniSp6 as an endogenous control gene. Real-time PCR was carried out on CFX Real-Time PCR Detection Systems (Bio-Rad, Hercules, California, USA), programmed as follows: 50 °C for 2 min, 95 °C for 10 min followed by 45 cycles of 95 °C for 15 s and 60 °C for 1 min. The relative gene expression was calculated in the periodontal disease group vs the HC according to the formula 2−∆∆Ct55. The calculations were performed in MS Excel and graphs were drawn using GraphPad Prism 6 (GraphPad, La Jolla, CA, USA).
ELISA assay
The human TNFα and IL-6 concentrations were measured by ‘sandwich’ ELISA (Millipore, Merck KGaA, Germany) on the GCF sample, according to the manufacturer’s instructions. The absorbance of each well was detected with a spectrophotometer, allowing for the generation of a standard curve and subsequent determination of protein concentration. The range of the standard curves was 1.37–12,000 pg/mL for IL-6 and 24.58–6000 pg/mL for TNFα.
Statistical analysis
The mean was taken as the measurement of the main tendency, and the SD was taken as the dispersion measurement for the statistical analysis of the results. Qualitative variables were reported as frequency and percentage. The normal distribution of data was tested by the Kolmogorov–Smirnov test. Student’s t-test was performed to compare baseline variables between study groups. Radar plot analysis for the distribution of differential expression patterns of miRNAs was evaluated. Pearson’s correlation analysis was calculated to assess the relationship between cytokine levels and miRNAs gene expression in patients. Statistically significant was set at p-value < 0.05. Data analysis was performed with Stata version 15 (StataCorp LLC, College Station, TX, USA) and GraphPad Prism 6 Software, version 6.01, 2012.
Ethical approval and consent to participate
This study conformed to the provisions of the Declaration of Helsinki. The protocol was approved by the Ethics Review Committee of G.d’Annunzio University, Chieti-Pescara (Protocol code 254/2019). Informed consent was obtained from all participants.
Strengths and limitations of the study
Our data underline the importance of crevicular fluid as a possible diagnostic tool and means to monitor the status of patients affected by periodontal disease. On the other hand, the relatively small number of studied samples limits the possibility of generalizing the obtained considerations and of using these data to define the diagnostic sensitivity and specificity of the studied miRNAs. The evaluation of miRNAs could be accompanied by the analysis of the proteome, transcriptome and microbiome, in order to obtain a complete picture of their role in periodontal disease.
Data availability
The data supporting the results of this article are included within the article and can be required to the corresponding authors.
References
Kassebaum, N. J. et al. Global, regional, and national prevalence, incidence, and disability-adjusted life years for oral conditions for 195 countries, 1990–2015: A systematic analysis for the global burden of diseases, injuries, and risk factors. J. Dental Res. 96(4), 380–387 (2017).
Han, P., Bartold, P. M. & Ivanovski, S. The emerging role of small extracellular vesicles in saliva and gingival crevicular fluid as diagnostics for periodontitis. J. Periodontal. Res. 57(1), 219–231 (2022).
Papapanou, P. N. et al. Periodontitis: Consensus report of workgroup 2 of the 2017 world workshop on the classification of periodontal and peri-implant diseases and conditions. J. Periodontol. 89(1), S173–S182 (2018).
Tonetti, M. S., Greenwell, H. & Kornman, K. S. Staging and grading of periodontitis: Framework and proposal of a new classification and case definition. J. Periodontol. 89(1), S159–S172 (2018).
Ramadan, D. E., Hariyani, N., Indrawati, R., Ridwan, R. D. & Diyatri, I. Cytokines and chemokines in periodontitis. Eur. J. Dentistry 14(3), 483–495 (2020).
Shaddox, L. M. et al. LPS-induced inflammatory response after therapy of aggressive periodontitis. J. Dent. Res. 92(8), 702–708 (2013).
Luan, X. et al. MicroRNAs and immunity in periodontal health and disease. Int. J. Oral Sci. 10(3), 24 (2018).
Kornman, K. S. Mapping the pathogenesis of periodontitis: A new look. J. Periodontol. 79(8 Suppl), 1560–1568 (2008).
Olsen, I., Singhrao, S. K. & Osmundsen, H. Periodontitis, pathogenesis and progression: miRNA-mediated cellular responses to Porphyromonas gingivalis. J. Oral Microbiol. 9(1), 133 (2017).
Essandoh, K., Li, Y., Huo, J. & Fan, G. C. MiRNA-mediated macrophage polarization and its potential role in the regulation of inflammatory response. Shock 46(2), 122–131 (2016).
Singh, R. P. et al. The role of miRNA in inflammation and autoimmunity. Autoimmun. Rev. 12(12), 1160–1165 (2013).
Stoecklin-Wasmer, C. et al. MicroRNAs and their target genes in gingival tissues. J. Dent. Res. 91(10), 934–940 (2012).
Venugopal, P. et al. Differential expression of microRNAs let-7a, miR-125b, miR-100, and miR-21 and interaction with NF-kB pathway genes in periodontitis pathogenesis. J. Cell. Physiol. 233(8), 5877–5884 (2018).
Kim, S. H., Lee, S. Y., Lee, Y. M. & Lee, Y. K. MicroRNAs as biomarkers for dental diseases. Singap. Dental J. 36, 18–22 (2015).
Kebschull, M. & Papapanou, P. N. Mini but mighty: microRNAs in the pathobiology of periodontal disease. Periodontol. 69(1), 201–220 (2015).
Schmalz, G. et al. MicroRNAs as salivary markers for periodontal diseases: A new diagnostic approach?. Biomed. Res. Int. 2016, 1027525 (2016).
Wu, P., Feng, J. & Wang, W. Expression of miR-155 and miR-146a in the saliva of patients with periodontitis and its clinical value. Am. J. Transl. Res. 13(6), 6670–6677 (2021).
Sipert, C. R. et al. MicroRNA-146a and microRNA-155 show tissue-dependent expression in dental pulp, gingival and periodontal ligament fibroblasts in vitro. J. Oral Sci. 56(2), 157–164 (2014).
Moffatt, C. E. & Lamont, R. J. Porphyromonas gingivalis induction of microRNA-203 expression controls suppressor of cytokine signaling 3 in gingival epithelial cells. Infect. Immun. 79(7), 2632–2637 (2011).
Bahn, J. H. et al. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin. Chem. 61(1), 221–230 (2015).
Aziz, F. The emerging role of miR-223 as novel potential diagnostic and therapeutic target for inflammatory disorders. Cell Immunol. 303, 1–6 (2016).
Haneklaus, M., Gerlic, M., O’Neill, L. A. & Masters, S. L. miR-223: Infection, inflammation and cancer. J. Intern. Med. 274(3), 215–226 (2013).
Fenoglio, C. et al. Decreased circulating miRNA levels in patients with primary progressive multiple sclerosis. Mult. Scler. 19(14), 1938–1942 (2013).
Ma, X. et al. Expression, regulation and function of MicroRNAs in multiple sclerosis. Int. J. Med. Sci. 11(8), 810–818 (2014).
Chen, L. et al. Exosomal miR-103-3p from LPS-activated THP-1 macrophage contributes to the activation of hepatic stellate cells. FASEB J. 34(4), 5178–5192 (2016).
Yang, S. et al. miR-148b-3p, miR-337-5p and miR-423-5p expression in alveolar ridge atrophy and their roles in the proliferation and apoptosis of OMMSCs. Exp. Ther. Med. 16(6), 5334–5342 (2018).
Safran, M. et al. GeneCards Version 3: The human gene integrator. Database (Oxf.) 2010, 20 (2010).
Preshaw, P. M. et al. Treatment of periodontitis reduces systemic inflammation in type 2 diabetes. J. Clin. Periodontol. 47, 737–746 (2020).
Lim, G. et al. Periodontal health and systemic conditions. Dentistry J. 8(4), 130 (2020).
Winning, L. & Linden, G. Periodontitis and systemic disease. BDJ Team 2, 15163 (2015).
Martínez-García, M. & Hernández-Lemus, E. periodontal inflammation and systemic diseases: An overview. Front. Physiol. 12, 709438 (2021).
Cardoso, E. M., Reis, C. & Manzanares-Céspedes, M. C. Chronic periodontitis, inflammatory cytokines, and interrelationship with other chronic diseases. Postgrad. Med. 130, 98–104. https://doi.org/10.1080/00325481.2018.1396876 (2018).
Singh, P., Gupta, N. D., Bey, A. & Khan, S. Salivary TNF-alpha: A potential marker of periodontal destruction. J. Indian Soc. Periodontol. 18(3), 306–310 (2014).
Hirano, T., Akira, S., Taga, T. & Kishimoto, T. Biological and clinical aspects of interleukin 6. Immunol. Today 11(12), 443–449 (1990).
Ishimi, Y. et al. IL-6 is produced by osteoblasts and induces bone resorption. J. Immunol. 145(10), 3297–3303 (1990).
Taba, M. Jr., Jin, Q., Sugai, J. V. & Giannobile, W. V. Current concepts in periodontal bioengineering. Orthod. Craniofac. Res. 8(4), 292–302 (2005).
Miller, C. S., King, C. P., Langub, M. C., Kryscio, R. J. & Thomas, M. V. Salivary biomarkers of existing periodontal disease: A cross-sectional study. J. Am. Dent. Assoc. 137(3), 322–329 (2006).
Scannapieco, F. A., Bush, R. B. & Paju, S. Associations between periodontal disease and risk for nosocomial bacterial pneumonia and chronic obstructive pulmonary disease. A systematic review. Ann. Periodontol. 8(1), 54–69 (2003).
Frodge, B. D., Ebersole, J. L., Kryscio, R. J., Thomas, M. V. & Miller, C. S. Bone remodeling biomarkers of periodontal disease in saliva. J. Periodontol. 79(10), 1913–1919 (2008).
Giannobile, W. V. et al. Saliva as a diagnostic tool for periodontal disease: Current state and future directions. Periodontol. 2000(50), 52–64 (2009).
Machado, V. et al. IL-6 and TNF-α salivary levels according to the periodontal status in Portuguese pregnant women. PeerJ 6, e4710 (2018).
Darveau, R. P. Periodontitis: A polymicrobial disruption of host homeostasis. Nat. Rev. Microbiol. 8(7), 481–490 (2010).
Bartold, P. M. & Van Dyke, T. E. An appraisal of the role of specific bacteria in the initial pathogenesis of periodontitis. J. Clin. Periodontal. 46, 6–11 (2019).
Zhou, Z. et al. Inflammation has synergistic effect with nicotine in periodontitis by up-regulating the expression of α7 nAchR via phosphorylated gsk-3β. J. Cell. Mol. Med. 24, 2663–2676 (2020).
Kapoor, P., Chowdhry, A., Bagga, D. K., Bhargava, D. & Aishwarya, S. MicroRNAs in oral fluids (saliva and gingival crevicular fluid) as biomarkers in orthodontics: Systematic review and integrated bioinformatic analysis. Progress Orthodont. 22(1), 31 (2021).
Costantini, E. et al. Improved osteogenic differentiation by extremely low electromagnetic field exposure: possible application for bone engineering. Histochem. Cell Biol. https://doi.org/10.1007/s00418-022-02126-9 (2022).
Donati, S., Ciuffi, S., Palmini, G. & Brandi, M. L. Circulating miRNAs: A new opportunity in bone fragility. Biomolecules 10, 927 (2020).
Perri, R., Nares, S., Zhang, S., Barros, S. P. & Offenbacher, S. MicroRNA modulation in obesity and periodontitis. J. Dent. Res. 91(1), 33–38 (2012).
Santonocito, S., Polizzi, A., Palazzo, G. & Isola, G. The emerging role of microRNA in periodontitis: Pathophysiology, clinical potential and future molecular perspectives. Int. J. Mol. Sci. 22(11), 5456 (2021).
AlQranei, M. S. & Chellaiah, M. A. Osteoclastogenesis in periodontal diseases: Possible mediators and mechanisms. J. Oral Biosci. 62(2), 123–130 (2020).
Bauernfeind, F. et al. NLRP3 inflammasome activity is negatively controlled by miR-223. J. Immunol. 189(8), 4175–4181 (2012).
Aziz, F. The emerging role of miR-223 as novel potential diagnostic and therapeutic target for inflammatory disorders. Cell. Immunol. 303, 1–6 (2016).
Bellavia, D. et al. Deregulated miRNAs in bone health: Epigenetic roles in osteoporosis. Bone 122, 52–75 (2019).
Costantini, E. et al. Evaluation of salivary cytokines and vitamin D levels in periodontopathic patients. Int. J. Mol. Sci. 21(8), 2669 (2020).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)). Methods 25, 402–440 (2001).
Funding
This research was funded by MIUR funds.
Author information
Authors and Affiliations
Contributions
E.C., M.R. and B.S. initiated, designed and supervised the project and wrote the article. E.C., L.A., B.S. and P.D.G. performed the experiments and contributed to writing the article. P.D.G. and M.R. analyzed and validated the obtained data. M.R., R.M., G.M. and S.C. revised the manuscript and performed the English editing. All authors reviewed the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Costantini, E., Sinjari, B., Di Giovanni, P. et al. TNFα, IL-6, miR-103a-3p, miR-423-5p, miR-23a-3p, miR-15a-5p and miR-223-3p in the crevicular fluid of periodontopathic patients correlate with each other and at different stages of the disease. Sci Rep 13, 126 (2023). https://doi.org/10.1038/s41598-022-26421-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-26421-6
- Springer Nature Limited
This article is cited by
-
MicroRNAs Function in Dental Stem Cells as a Promising Biomarker and Therapeutic Target for Dental Diseases
Molecular Diagnosis & Therapy (2023)