Abstract
Radiation injury from medical, accidental, or intentional sources can induce acute and long-term hepatic dysregulation, fibrosis, and cancer. This long-term hepatic dysregulation decreases quality of life and may lead to death. Our goal in this study is to determine acute changes in biological pathways and discover potential RNA biomarkers predictive of radiation injury. We performed whole transcriptome microarray analysis of mouse liver tissue (C57BL/6 J) 48 h after whole-body irradiation with 1, 2, 4, 8, and 12 Gray to identify significant expression changes in mRNAs, lncRNAs, and miRNAs, We also validated changes in specific RNAs through qRT-PCR. We used Ingenuity Pathway Analysis (IPA) to identify pathways associated with gene expression changes. We observed significant dysregulation of multiple mRNAs across all doses. In contrast, miRNA dysregulation was observed upwards of 2 Gray. The most significantly upregulated mRNAs function as tumor suppressors: Cdkn1a, Phlda3, and Eda2r. The most significantly downregulated mRNAs were involved in hemoglobin synthesis, inflammation, and mitochondrial function including multiple members of Hbb and Hba. The most significantly upregulated miRNA included: miR-34a-5p, miR-3102-5p, and miR-3960, while miR-342-3p, miR-142a-3p, and miR-223-3p were most significantly downregulated. IPA predicted activation of cell cycle checkpoint control pathways and inhibition of pathways relevant to inflammation and erythropoietin. Clarifying expression of mRNA, miRNA and lncRNA at a short time point (48 h) offers insight into potential biomarkers, including radiation markers shared across organs and animal models. This information, once validated in human models, can aid in development of bio-dosimetry biomarkers, and furthers our understanding of acute pathway dysregulation.
Similar content being viewed by others
Introduction
Radiation exposure from medical, accidental, or intentional events causes direct damage to DNA and production of reactive oxygen species (ROS), inducing further damage. Radiation injury to normal tissue may cause cell death and depletion of specific cell types, chronic redox stress leading to long-term dysfunction, and mutagenesis leading to oncogenesis1,2. Previous studies from atomic bomb survivors highlighted the impact of radiation on fatty liver development, long term liver dysfunction and cancer development3,4.
Toxicity and dysfunction in the liver have been reported extensively in the literature, with one study reporting that 70% (14/20) of patients displayed hepatic dysfunction as measured by dysregulation of liver transaminases and alkaline phosphatase levels after radiation therapy5,6. Development of fatty liver disease and insulin resistance has been observed at an increased rate in patients who received radiation therapy and in atomic bomb survivors4,7. Hepatic steatosis, the accumulation of fat in the liver, has also been reported in both rat and rabbit livers between two and six weeks after radiation injury, with greater severity of steatosis being associated with animal death8,9,10. Data from mini pigs indicated that 14 Gy localized radiation to the liver caused alterations in hepatocystolic function11. One study showed serum triglyceride and cholesterol levels increased in the livers of rats who received 6 Gy of radiation12.
Radiation-induced liver disease (RILD) is a dose limiting factor in radiation therapy to the abdomen that can develop in patients weeks to months after cessation of treatment13. The progression to RILD begins with tissue damage and endothelial cell death, which leads to inflammation and increased expression of cytokines as the liver attempts to repair itself14,15. The repair process includes proliferation of hepatocellular progenitors and myofibroblastic hepatic stellate cell transformation which can lead to fibrosis of the liver and RILD16. RILD symptoms include increased levels of liver enzymes including alkaline phosphatase, ascites, abdominal pain, and destruction of central veins. This destruction decreases oxygen delivery and causes tissue dysfunction, which can lead to death.17,18.
Understanding early alterations in transcription after radiation injury will provide new insights into strategies to prevent and mitigate normal liver damage. Further, while tools for radiation biodosimetry exist such as analysis of chromosomal damage, these assays to determine radiation dose may require a high level of technical experience, have low throughput, and can display low intercomparison accuracy19. Further research to develop a high throughput, accurate, and non-invasive method for radiation biodosimetry is ongoing. Prior research by others and from our own laboratory have highlighted the potential utility of messenger RNA (mRNA), microRNA (miRNA), and long non-coding RNA (lncRNA) as biomarkers for radiation injury and radiation sensitivity20,21,22.
Long non-coding RNA (lncRNA), RNA strands over 200 nucleotides that do not code for a protein, have previously been proposed as biomarkers for cancer, cardiovascular disorders, and other diseases23,24. Research from our lab and others have highlighted the radiation response of multiple lncRNA. Some of these lncRNA are otherwise uncharacterized, while others, such as Trp53cor1 and Dino, are relatively well known25,26,27,28. MicroRNA (miRNA), non-coding RNA of roughly 22 nucleotides, exert their functions by hybridizing to complementary sequences in the 3’ UTRs of mRNAs, which lead to either RNA degradation or translation inhibition29. In addition, miRNA are stable in biofluids, making them a potentially useful prognostic and diagnostic indicator of radiation damage through minimally invasive blood draws30,31. Research to discover lncRNA-miRNA-mRNA networks and altered pathways after radiation in the lung has provided insights into early signs of dysfunction and potential biomarkers32.
We observed alterations in lncRNA and miRNA after radiation injury, which demonstrate their potential utility as part of a panel of RNA biomarkers to help determine radiation dose and potential pathogenesis28,33. This study will outline radiation induced dose dependent, liver specific, gene expression level changes after whole body irradiation (WBI) of mice. Understanding how radiation alters mRNA, lncRNA, and miRNA expression at 48 h will aid in identifying and predicting organ damage at both short- and long-term time points. It will also aid in developing strategies for mitigation of liver injury.
Results
Microarray analysis indicates an overall decrease in mRNA expression in liver after irradiation
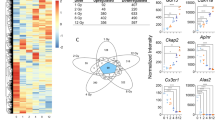
In total, analysis of microarray data (|log2FC|> 1; p value < 0.05) indicated that 2483 genes were shown to be differentially expressed between unirradiated control samples and at least one dose for WBI mice (Fig. 1A). Furthermore, microarray analysis from mouse liver tissue indicated that, at each dose of radiation, more mRNAs were downregulated than upregulated (Fig. 1B). Across all doses, 35 genes were dysregulated compared to control samples. In contrast, 25, 101, 118, 840, and 423 were exclusively expressed at 1, 2, 4, 8, and 12 Gy, respectively (Fig. 1C). Among all doses, the most significantly upregulated genes included Cdkn1a, Eda2r, and Phlda3, and the most significantly downregulated genes included Hba-a2, Serpina9, and Ms4a1 (Fig. 1D). In addition to Hba-a2, we also observed significant downregulation of Hba-a1, Hbb-b1, Hbb-b2, Hbb-bt at all doses of radiation (Supplemental Fig. 1). These upregulated genes (Cdkn1a, Eda2r and Phlda3), are known for their role in cell cycle arrest, apoptosis and tumor suppression, while the most downregulated genes (Hba-a2, Serpina9, and Ms4a1) are associated with hemoglobin synthesis, mitochondrial function and B cell differentiation (Table 1)34,35,36,37,38,39,40,41,42,43. Supplemental Table S1 lists all genes dysregulated by radiation dose focusing on p value and log change. Further analysis of this data indicated that 1023 probes displayed a significant up or downregulation across all doses (Supplemental Table S1), with the top 20 and bottom 20 presented (Supplemental Table S2).
Radiation-induced gene expression profiles in mouse liver tissue. Whole genome microarray analysis was performed on all samples. A linear model was fit to each probe to evaluate differential expression of irradiated samples compared to controls. Criteria of (log2 fold change ((log2FC) > 1 and Benajmini-Hochberg adjusted (B-H) p value < 0.05) relative to controls were used to determine significance and differential expression. (A) Heatmap displays expression patterns, represented by z-score, of all differentially expressed mRNAs across all doses and controls. (B) Venn diagram shows dose distribution and overlap of differentially expressed mRNAs across all doses. (C) The number of down-regulated versus up-regulated mRNAs at each dose are shown in the table. (D) Examples of significant linearly up- and down-regulated mRNAs are shown to display the dose response to radiation in liver tissue samples. Asterisk (*) indicates statistical significance.
We sought to divide gene dysregulation into low (1 and 2 Gy), middle (4 Gy) and high (8 and 12 Gy) dose clusters (Fig. 1C). The low dose cluster had only one gene dysregulated in both 1 Gy and 2 Gy, Set, a nuclear proto-oncogene associated with DNA repair44 (Fig. 1C, Supplemental Table S1). In contrast, 8 Gy and 12 Gy share 273 genes which are not shared with the lower doses.
Ingenuity pathway analysis of dysregulated genes highlights downregulation of immune response and increased cell cycle arrest
A canonical pathway analysis was performed with differentially expressed mRNA using IPA. The top 30 canonical pathways dysregulated by radiation and relevant to normal liver are presented in Fig. 2A (log2FC > 1, B-H p < 0.05) and the top 30 most dysregulated functions are in Fig. 2B. The only pathways altered at 1 Gy radiation were relevant to mitosis and erythropoietin signaling. Multiple pathways relevant to immune response were downregulated starting at 2 Gy radiation (Fig. 2A). Similarly, pathways relevant to senescence and cell cycle arrest were recruited starting at 2 Gy. Surprisingly, the hepatic fibrosis signaling pathway was downregulated at 4 Gy and continued to show downregulation at 8 and 12 Gy (Fig. 2A). Cell survival and cell migration pathways were downregulated as radiation dose increased (Fig. 2B).
Predicted canonical pathway dysregulation in mouse liver samples based on all differentially expressed mRNAs. IPA was used to perform pathway analysis on all differentially expressed mRNAs to predict pathway involvement, independent of the target relationship with differentially expressed miRNAs. (A) Displays the top 30 most significantly dysregulated pathways (B-H p value < 0.05). A positive z-score indicates predicted activation of the pathway based on gene expression and a negative z-score indicates predicted deactivation of the pathway based on gene expression. Pathways are hierarchically clustered by z-score. (B) Displays top 30 most dysregulated functions (B-H p value < 0.05).
We then divided these doses into low (1 and 2 Gy), middle (4 Gy), and high (8 and 12 Gy). We focused on the functions that were most severely activated based on Z-score from Fig. 2 to examine specific genes in the pathway or function. Our goal was to locate potential biomarkers which may be used to predict subsequent pathway dysfunction to aid in medical decision making. The observation of genes going from the high (Supplemental Fig. S2C) to middle (Supplemental Fig. S2B) to low (Supplemental Fig. S2A) are as follows. In 8 and 12 Gy samples, several downstream pathways relevant to inflammation were strongly upregulated. We present genes that IPA has associated with Inflammation of Body Cavity pathway based on 8 Gy (Supplemental Fig. S2C). Genes only dysregulated at 8 and 12 Gy include Epha2, Il1r1, Lipin1, Cd40, Irf5, Gatm, Soat1, and Zbp1. In 4 Gy samples, Apoptosis pathways were most strongly activated (Supplemental Fig. S2B). Genes only dysregulated at 4 Gy include Ncam1, Brca2, Grb10, Ins1, Rcan2, Six4, Psme4 and Wsb1. Oddly, while senescence pathways did not appear significant at 1 Gy within the comparison analysis, they were some of the most strongly activated pathways for 1 Gy when observed individually, with a Z-score greater than 2 for Senescence of Cells (Supplemental Fig. S2A). Within this senescence pathway most genes were also upregulated by all higher doses. Two genes were specific to a single dose. Rad9b, a checkpoint control protein, was only significant at 2 Gy45. Wnt16, which was previously shown to protect cartilage by inhibiting excessive WNT signaling in a mouse model, was only significant at 1 Gy46.
Significant dysregulation is observed across solute carrier families as well as phase I and phase II metabolism genes in the liver
The liver’s role in detoxification and drug metabolism are well studied. We wanted to clarify how radiation impacts influx, phase I and phase II metabolism, and efflux of metabolites in the liver. We observed a decrease in solute carrier organic anion family (SLCO) genes, Slco1a1, Slco2a1, Slco2b1 (Fig. 3A). We also observed decreased expression of solute carrier family (Slc) genes Slc4a1, Slc5a1, Slc6a20a, Slc13a2, and Slc14a1. Interestingly, families of solute carriers did not show consistent up or downregulation. While Slc16a6 decreased in expression after radiation, Slc16a5, Slc6a21, and Slc16a7 showed significant increases in expression by 8 Gy. Similarly, Slc22 family members Slc22a5, Slc22a27, Slc22a29 also showed increased expression, with Slc22a27 and Slc22a29 showing statistical significance at 4 Gy while Slc22a5 showed significance only at 12 Gy. In contrast Slc22a14 showed significantly decreased expression at 8 Gy. Slc25a25 showed significantly increased expression only at 4 Gy while Slc25a35 was significantly decreased at 8 and 12 Gy. Abca8a was significantly decreased at 2, 8 and 12 Gy but not 4 Gy. While Abcd2 was only upregulated significantly at 8 Gy. Slc35b1 was decreased at 4 and 12 Gy while Slc35g2 was only decreased at 8 Gy.
Drug and fat metabolism are dysregulated by radiation injury. Heatmaps of solute carriers (SLC) (A), phase I and phase II metabolism genes (B), and fat metabolism (C) are displayed. Only genes which were shown to be significantly altered ((log2FC > 1), p < 0.01) in at least one dose are displayed. White indicates control samples. Red indicates upregulation compared to control while blue indicates downregulation compared to control.
Due to the role of cyp450 genes in phase I metabolism we chose to study their expression across radiation doses (Fig. 3B). We separated genes in the heatmaps between genes relevant to transport (Fig. 3A), from genes relevant to metabolism of xenobiotics and biosynthesis (Fig. 3B) from genes relevant to fat metabolism (Fig. 3C). We observed dysregulation of several phase II metabolism genes including Sult2a4, Gsta2, and Mgst3. Cyp26a1 showed a statistically significant decrease in expression at 12 Gy. Cyp11b1 was significantly decreased at 2 and 4 Gy. Cyp26b1 is significantly decreased only at 1 Gy. Interestingly, Cyp2u1 was significantly decreased at 8 and 12 Gy while Cyp2j9 was significantly increased at 8 and 12 Gy. Cyp46a1 and the phase II metabolism gene Gsta2 were only significant at 8 Gy. The phase II gene Sult2a4 showed significantly increased expression at 4 and 8 Gy. Mgst3 was significantly increased at 4 Gy. Schematic of radiation induced changes to transporters, Phase I and Phase II genes in liver are presented and further explained in the discussion (Supplemental Fig. S3A).
Since we observed changes to both active and passive transporters relevant to lipid metabolism and because there is a known link between fatty liver development and radiation, we chose to further study genes relevant to these pathways. Acaca shows a significant increase in expression at 2 Gy before decreasing to normal expression (Fig. 3C). Acacb showed decreased expression at 8 and 12 Gy. Acad members 9 and 11, Cpt1c, and Hadhb showed significant increases at 12 Gy. Acot family members 3, 4, and 5 showed significantly increased expression from 4 to 12 Gy. Acsm2, Acss1, and Acss2 were all downregulated as dose increased. The genes Cebpd, Srebf1, Lpin1, Lpin2, Plin4, and Plin5 showed increased expression with increasing dose. The Fabp family members 3, 4, 5, and 12 decreased with increasing radiation doses. Scd1 decreased with increased radiation dose, while Scd3 increased in expression with radiation dose.
In sum, we see a decrease in fatty acid uptake into the liver and a decrease in peroxisomal fatty acid oxidation as radiation doses increase. We also observe increased triacylglycerol synthesis and maintenance after 8 and 12 Gy doses of radiation as depicted in Supplemental Fig. S3B. Hepatic steatosis has previously been associated with RILD and death in animal models. Understanding the genes within this pathway are useful in developing biomarkers to predict liver metabolism dysfunction and greater liver damage.
Increased dose of radiation caused increases in miRNA dysregulation
A separate whole genome analysis was performed on miRNA, (Fig. 4A). No miRNAs were significantly dysregulated at 1 Gy, while 12 miRNAs were dysregulated at 12 Gy (Fig. 4B). From 2 to 12 Gy, miRNA-34a-5p was significantly upregulated. In both 2 Gy and 4 Gy animals, only miR-34a-5p was upregulated. At 8 Gy miR-34a-5p, miR-3102-5p, 466n-3p and miR-302a5p were dysregulated. At 12 Gy miR-34a-5p, miR-3102-5p, miR-142a-5p, miR-142a-3p, miR-342-3p and miR-3960 were dysregulated (Fig. 4C). Notably, miR-466n-3p and miR-302a-5p were only dysregulated at 8 Gy. The most upregulated miRNA included miR-34a-5p, miR-3102-5p, and miR-3960, and the most downregulated included miR-142a-3p, miR-342, and miR-223-3p (Fig. 4D). These downregulated markers were only decreased at 12 Gy. Due to the low number of differentially expressed miRNA, all probes featuring a linear trend upwards or downwards are shown (Supplemental Table S3).
Radiation-induced microRNA expression profiles in mouse liver tissue. Microarray analysis was performed for all samples, and a linear model was fit to each miRNA probe to assess differential expression of irradiated samples compared to controls. Criteria of |log2FC|> 1 and B-H p value < 0.05 relative to controls were used to determine significance and differential expression. (A) Heatmap displays expression patterns, represented by z-score, of all differentially expressed miRNAs across all doses and controls. (B) The number of down-regulated versus up-regulated miRNAs at each dose are shown in the table. (C) Venn diagram shows dose distribution and overlap of differentially expressed miRNAs across all doses. (D) Examples of significant linearly up- and down-regulated miRNAs are shown to display the dose response to radiation in liver tissue samples. Asterisk (*) indicates statistical significance.
Few lncRNA showed a continuous dysregulation in expression after WBI
To examine the relationship between lncRNA and WBI, we filtered whole genome data to only include probes associated with lncRNA. Both discrete and continuous differential expression of lncRNA is depicted in heatmap (Fig. 5A). Some lncRNA display non-monotonic expression with increasing doses of radiation, notably A_30_P01028589 and A_30_P01019037. (Fig. 5A). Discretely dysregulated lncRNA between any dose and 0 Gy samples are shown in Fig. 5B,C. The probes of interest were Trp53cor1, Snhg15, chr1:163528200-163528398_F and chr1:163508244-163586072_F, (Fig. 5D). No lncRNA were significant at 1 Gy (Fig. 5B,C). By 2 Gy, Trp53cor1 had increased to a statistically significant degree and remained upregulated for the higher doses. By 8 Gy, 5 lncRNA including Snhg15 were significantly upregulated. At 12 Gy there were 8 lncRNA which became statistically upregulated, including chr1:163528200-163528398_F and chr1:163508244-163586072_F. All lncRNA which show a significant linear dose response are presented in Supplemental Table S4.
Radiation-induced long non-coding RNA expression profiles in mouse liver tissue. (A) Heatmap displays expression patterns, of all differentially expressed lncRNAs across all doses and controls. A linear model was fit to each lncRNA probe to assess differential expression of irradiated compared to control samples using criteria of |log2FC|> 1 and B-H p value < 0.05. (B) The table shows the number of discrete down- versus up-regulated lncRNAs at each dose. (C) Venn diagram shows dose distribution and overlap of differentially expressed lncRNAs across all doses. (D) Examples of significant linearly up- and down-regulated lncRNAs are shown to display the dose response of lncRNAs to radiation in liver tissue samples. Asterisk (*) indicates statistical significance.
Significant genes found in microarray are validated and consistent across multiple strains
To ensure that significant gene expression changes in response to radiation were not specific to the C57BL/6 J mouse strain, an identical PCR analysis was conducted on C3H mice. The top upregulated genes (Cdkn1a, Eda2r, and Phdla3), as well as the most significantly downregulated gene (Hba-a2), were cross examined at 0, 1, 2, 4, and 8 Gy radiation dosages (Supplemental Fig. S4). Both C57BL/6 J and C3H showed significant upregulation in Cdkn1a at 2, 4, and 8 Gy. With regards to Phlda3, C3H began showing significant upregulation at 8 Gy, whereas Phlda3 expression in C57BL/6 J increased significantly at 2, 4, and 8 Gy. Both strains showed significant downregulation of Hba-a2 beginning at 1 Gy. The consistency in gene regulation post-radiation across multiple strains reinforces the promise of using biomarkers for radiation bio-dosimetry.
In contrast, non-coding RNA Trp53cor1 and miR-34a did not display consistency between C57BL/6 J and C3H mice (Supplemental Fig. S5). Trp53cor1 was statistically significant at 1, 2, and 8 Gy in C57BL/6 J, but was only significant at 4 Gy in C3H. Dino was statistically significant at 2, 4, and 8 Gy in both C57BL/6 J and C3H mice. The miR-34a was significant at 4 and 8 Gy in C57BL/6 J mice, but only reached significance at 4 Gy for C3H mice.
Discussion
Radiation induced liver disease (RILD) limits the application of radiotherapy for the treatment of liver cancers47,48,49. Rescuing the normal liver function and prevention of long-term radiation toxicity demands understanding of the molecular changes induced by radiation in normal liver cells which could then be evaluated for therapeutic interventions.
We observed that some genes in our study have not previously been associated with radiation or any functions, including NR_045989, Gm45941, Gm41572, and Gm39334 (Supplemental Table S2, S5). In addition, some RNA have only received superficial attention for their role in radiation response including miR-8110 (Supplemental Table S4)50. Further information on their roles may lead to as yet undiscovered opportunities for mitigation or utilization as biomarkers for pathology. We recognize that not all genes showed linear dysregulation; Sult2a4 showed increased expression at 2, 4, and 8 Gy and decreased expression at 12 Gy. This non-monotonic response has previously been observed in other animal and human radiation research51,52. We focus the discussion on previously reported RNA and RNA relevant to specific pathway dysfunctions as those may serve as biomarkers of injury and of targets for injury mitigation.
WBI dysregulates genes relevant to hemoglobin synthesis, radiation stress response, immune response and cell cycle arrest in the liver
Multiple hemoglobin family gene members, such as Hbb-b1, Hbb-b2, Hbb-bt, Hba-a1, and Hba-a2, were downregulated in the mouse liver after increasing doses of WBI. Increased levels of free hemoglobin in serum were linked to higher levels of hepatic steatosis, non-alcoholic fatty liver disease, and other metabolic disorders of the liver in males in human population studies53,54,55. Prior data shows hepatocytes synthesize hemoglobin to decrease oxidative stress56. This observed downregulation in our data may indicate damage to the hepatocytes.
The most dose–response upregulated genes were Cdkn1a, Phlda3, and Eda2r, which are regulated by p53 to induce cell cycle arrest or apoptosis57. All three have previously been highlighted as predictive markers of radiation injury in murine models38,58,59,60,61. Cdkn1a has previously been reported as a marker of irradiation in cancer patients undergoing WBI and in murine models from our lab and others25,35,62.
The liver has known functions in metabolism and xenobiotic detoxification. This detoxification includes an immunological response to viruses, bacteria, and other potential pathogens63. Among the most downregulated genes after radiation were two involved in B cell differentiation and activation: Serpina9 and Ms4a164,65. Ms4a1 was one of the most downregulated genes in the blood of male prostate cancer patients suffering from fatigue after radiation therapy39. Serpina9, also known as centerin and GCET1, is found in germinal centers of B cells though its function is not well understood65. While these genes are associated with B cell differentiation, it is possible they also have as yet unelucidated roles in other cell types.
Liver transport and detoxification are compromised after radiation injury
As shown in Fig. 3B, the Slc gene family codes for membrane proteins which allow passive, symport and antiport transport of amino acids, lipids, glucose, anions and cations across the membrane66. These solute carriers may be found on the cellular membrane or the membrane of various organelles and impact drug absorption67. Some are highly substrate specific, while others will transport a range of substrates across. For brevity, the roles of many of these Slc are noted here67,68,69,70,71,72,73,74,75,76.
Highlighted are the Slc most relevant to glucose and fatty acid metabolism. Slc5a1 also called sodium-glucose cotransporter 1 (SGLT1) uses the sodium electrochemical gradient to move glucose into cells77. Slc16a6 transports ketone bodies, while Slc16a7 transports pyruvate, lactate, and ketone bodies70,71. Interestingly, some Slc transporters may act indirectly to modify metabolism, Slc22a14 has previously been shown to indirectly impact triglyceride storage and fatty acid oxidation in a mouse model78.
Only two active transporters were dysregulated. Both Abcd2 and Abca8 are ATP Binding Cassette family members relevant to lipid metabolism79,80. While Very low density lipoprotein receptor (Vldlr) mediates lipid uptake and accumulation81,82. These changes may give insight into energy production pathways within the liver after radiation injury and potential biomarkers to understand liver damage and response over time.
Briefly, Cyp2u1, Cyp2j9, Cyp4f18 and Cyp11b1 all function in arachidonic acid and cholesterol metabolism83,84,85,86,87. Cyp26a1 and Cyp26b1 regulate retinol88. Modification of these pathways has implications for inflammation and response to reactive oxygen species which are produced after radiation injury. Cyp2u1 and Cyp46a1 function in lipid storage and mitochondrial function, with downregulation associated with increased triglyceride synthesis and hepatic lipid droplet formation84,89,90 Radiation-induced dysregulation of cytochrome p450 expression can impact inflammation and metabolism leading to long term dysfunction if unrepaired. Gsta2, Sult2, and Nqo1 aid in detoxification and efflux of metabolites and xenobiotics91,92,93,94. Dysregulation of these pathways has implications for efficacy of medications given post injury. In knowing that the liver has a reduced capacity to metabolize certain medications, clinicians must modify medication doses to avoid secondary toxic effects caused by this differential rate of medication metabolism.
Liver energy homeostasis and lipid storage are dysregulated by radiation
Prior literature has shown that radiation induces increased triglyceride storage, lipid metabolism dysfunction, and mitochondrial dysfunction in the liver10,95,96. These studies led us to interrogate the impact of radiation on lipid metabolism in our mouse model. Briefly, in Fig. 4B, Acaca, Acacb, Acss1, Acss2, Scd1, Scd3, Plin4, Plin5, Srebf1, Lipin1, and Lipin2 have functional roles in lipogenesis and triacylglycerol maintenance97,98,99,100,101,102.
Carnitine palmitoyl transferase 1 (Cpt1), Acyl-CoA dehydrogenase (Acad) and the mitochondrial trifunctional protein beta subunit (Hadhb) are known to bring fatty acids into the mitochondria, and perform needed steps for fatty acid oxidation103,104,105. Overall, we see a decrease in fatty acid uptake into the liver and a decrease in peroxisomal fatty acid oxidation as radiation doses increase. We also observe increased triacylglycerol synthesis and maintenance after 8 and 12 Gy doses of radiation. Prior research in rats indicate that selective 25 Gy radiation to the liver induced fat accumulation at 48 h post radiation106.
Most significantly dysregulated miRNAs play role in glucose metabolism and inflammation
Interestingly, while IPA did not implicate glucose metabolism as a dysregulated canonical pathway after radiation, multiple significantly altered miRNA affect glucose metabolism. Prior research into dysregulation of miR-34a-5p, miR-3102-5p, and miR-142-3p demonstrate that their dysregulation encourages insulin resistance and decreased glucose metabolism107,108,109 While we did not see significant dysregulation of rate limiting genes in glucose oxidation. We do see dysregulation of Slc5a1, a protein co-transporter of glucose and sodium (Fig. 3A).
Dysregulation of lncRNA may have implications for hemoglobin synthesis, and proliferation
While several lncRNA were shown to be dysregulated by radiation in our study, few have received even a name, and their function in normal tissue are poorly understood110,111. Trp53cor1 knockdown in combination with radiation produced decreased apoptosis in mouse embryonic fibroblasts112. We have previously observed upregulation of this lncRNA and Dino in whole mouse blood and in mouse heart after radiation injury28.
There was overlap in gene expression after radiation between Gottingen minipigs and mice indicating potential biomarkers
One limitation of our study is that we chose an early time point to observe genetic dysregulation. While we did not observe death at our short time point in the present mouse study, the anticipated LD50/60 for WBI of C57BL/6 J mice is approximately 7.69–7.81 Gy113. We have chosen to compare our early changes in our mice to our long-term study of Gottingen minipigs114. In that study, minipigs were followed for 45 days post-radiation. Animals were grouped as survivors and decedents depending on whether they survived till 45 days. We identified survival-predictive RNA biomarkers of liver injury in these minipigs. Interestingly, Serum Amyloid A2 (Saa2) was upregulated in the liver of decedent mini pigs. We observed a similar significant upregulation of Saa2 in our mouse liver for mice irradiated at 8 and 12 Gy compared to controls (Supplemental Table S1). Similarly, Gdf15 was upregulated in non-surviving pigs compared to survivors and controls. Gdf15 was upregulated in mice at 12 Gy compared to controls (Supplemental Table S1). Several metabolism markers were downregulated in non-survivors compared to survivors including Phosphoglycerate dehydrogenase (Phgdh), Acss2, and Scd1. Phgdh and Acss2 were only downregulated in mice at 8 Gy compared to controls, while Scd1 was downregulated in mice at 8 and 12 Gy compared to controls (Supplemental Table S1). This similarity in gene expression across species suggests potential for these genes to be used as part of an RNA panel of radiation biomarkers.
RNA biomarkers must be carefully selected
Some RNAs may be good general indicators of radiation exposure while others may be more organ specific and suggest targets for mitigation. The use of miR-34a is an example: differences in age, diabetes status and radiation are all known to impact miR-34a expression115,116,117. Similarly, pathophysiological differences between human and animal model response to radiation highlight potential obstacles in developing an RNA biomarker panel. While animal models provide useful initial information, radiation-induced liver disease in the mouse does not present with veno-occlusive lesions, which is a hallmark in human patients with RILD17,118. To validate the RNA and pathway dysregulation observed in this study and to create useful RNA panels for human patients, our lab will next focus on 3D cultures using multiple human primary cell types together to recapitulate normal human organ response to radiation damage.
This experiment was an initial study to demonstrate the potential utility of lncRNA-miRNA-mRNA in a biomarker panel to determine radiation biodosimetry and elucidate dysregulated pathways to aid in clinical triage and medical decision making. Other types of RNA including piRNA, tsRNA and rsRNAs are receiving attention119,120 and should be explored for their utility in building these panels as well.
Future directions
Understanding the expected patterns of radiation-induced early gene dysregulation in the liver and the dose–response pattern will be useful for diagnosing and mitigating RILD following whole body exposure. It may aid in clinical management of radiotherapy patients. With potential metabolic targets of injury indicated, including dose–response relationships, ongoing studies will address potential mitigators.
Methods
Total body irradiation of mice and sample collection
Six- to eight-week-old female C57BL/6 J and C3H mice received whole body irradiation (WBI) with x-rays using the Small Animal Radiation Research Platform (SARRP Xstrahl Ltd.). Mice were placed in plastic containers and exposed to a single surface dose of 1, 2, 4, 8, or 12 Gy at a dose rate of 1.05 Gy/min. Control mice (0 Gy) were placed in the same type of plastic container and sham irradiated. Three animals per dose were included in the study. Livers of irradiated and control animals were harvested 48 h after WBI. Organs were snap frozen in liquid nitrogen and stored at − 80 °C until processed for RNA isolation. The experimental protocol was approved by a New York University (NYU) Langone Medical Center under an approved IACUC protocol as part of a collaborative study. Tissue collection validation studies were performed at the National Cancer Institute, Radiation Oncology branch using a Pantak x-ray source at a dose rate of 2.28 Gy/min and conducted in accordance with the principles and procedures outlined in the NIH Guide for the Care and Use of Animals and procedures. All methods are reported in accordance with ARRIVE guidelines (https://arriveguidelines.org).
RNA isolation
Samples were bathed in liquid nitrogen and pulverized into a fine powder using a mortar and pestle. Approximately 100 µg of powdered sample was lysed with 700 µl of QIAzol lysis buffer (Cat # 79306, QIAGEN) and homogenized by passing the solution through QIAshredder spin columns (Cat # 79654, QIAGEN). RNA isolation was performed using standard miRNeasy mini kit (Cat # 217004, QIAGEN) according to the manufacturer’s protocol. Quality and quantity of the RNA samples were assessed using a DeNovix DS-11 nanodrop spectrophotometer (DeNovix, DE, US) and Agilent Bioanalyzer with the RNA6000 Nano Lab Chip (Agilent Technologies, Santa Clara, CA).
Microarray analysis
Microarray analysis was performed for sham animals (0 Gy) and 1 Gy, 2 Gy, 4 Gy, 8 Gy, and 12 Gy irradiated animals. Quality assessments and microarray experiments were completed as previously reported33. Samples were hybridized to Agilent Mouse GE 8×60 K v2 arrays for mRNA expression analysis and to Agilent Mouse miRNA 8×60 K v21.0 arrays (Design ID 070155) for miRNA expression analysis. Slides were washed and scanned on an Agilent SureScan Microarray Scanner. Expression values were extracted using Agilent Feature Extraction software.
Real time RT-PCR analysis of RNAs
Individual qRT-PCR reactions using RT2 qPCR primer assays along with RT2 First Strand Synthesis kit and RT2 SYBR Green qPCR Master Mix (QIAGEN) were performed. The following RNA primers were purchased from Qiagen, gene globe IDs are included for mRNA and assay IDs for non-coding RNA: Cdkn1a (PPM02901B-200), Eda2r (PPM32677A-200), Phlda3 (PPM28194A-200), Hba-a2 (PPM69448A-200), miR-34a (YP00204486), Trp53cor1 (LPM12776A), Dino26(FP- GCAATGGTGTGCCTGACTAT; RP- TTCTGGCTTCCCAGAG), and Rplp0 (PPM03561B). QRT-PCR analysis was performed on select miRNA, lncRNA, and mRNA to validate results and determine cross-strain accuracy as C3H mouse liver was also tested for RNA expression. C3H organ RNA extraction and qRT-PCR analysis were the same as explained above. Relative expression was calculated as: 2−dCt where dCt = Ct [test gene] − Ct Rplp028.
Statistical analysis
Analysis of mRNA and miRNA data was performed using R statistical software and the Bioconductor Linear Model for Microarray Analysis (LIMMA) package in R121. Background correction and normalization were performed in R using the normal-exponential correction method and quantile normalization between arrays122. Only probes with intensities above background on at least one array were kept in the dataset for analysis. Transcripts with multiple probes were averaged such that the final set reflected best estimates of transcript level expression. A linear model was fit to each probe to assess differential expression for pair-wise dose comparisons within the liver-tissue samples. This method employed an empirical Bayes smoothing approach that results in more stable model estimates by using information on variance from the whole probe set, despite the small number of arrays. Models were developed for each of the pair-wise comparisons between each dose (1, 2, 4, 8, and 12 Gy) and the control probes (0 Gy), and resulting probes were filtered using log2 fold change and adjusted p value thresholds (|log2FC|> 1, adjusted p value < 0.05)123. Additionally, a nested interaction model was fit for each probe to examine dose within tissue as a linear (continuous) trend. Each model yielded the main effects for the liver tissue and dose within the liver tissue. Probes were filtered using the nested dose coefficients with log fold change and adjusted p value thresholds (|log2FC|> 1, adjusted p value < 0.05).
To identify potential interactions, paired analysis was conducted to evaluate correlative relationships between pairs of differentially expressed mRNA and miRNA probes. mRNA and miRNA probes were paired using shared target transcript Ensembl IDs124. Probes that could not be mapped or paired were excluded. Transcripts for miRNA probes were identified using an Agilent microarray gene dataset and the TargetScan database; transcripts for mRNA probes were identified using an Agilent microarray gene dataset125. Transcript-miRNA pairs with a TargetScan context++ score above − 1 were excluded. Probe pairs with differentially expressed miRNA and mRNA probes were identified within the liver tissue for continuous dose contrast models. Pearson correlation coefficients of miRNA and mRNA expression across all experiments were calculated and plotted for the differentially expressed probe pairs.
Ingenuity pathway analysis
Both core and comparison analyses were performed in IPA (QIAGEN Inc., https://www.qiagenbioinformatics.com/products/ingenuitypathway-analysis). Pathways and function terms that satisfied an absolute z-score > 2 and p value < 0.01 were predicted to be altered based on the gene expression data.
Ethics approval and consent to participate
The experimental protocol was approved by a New York University (NYU) Langone Medical Center under an approved IACUC protocol as part of a collaborative study.
Data availability
Data is available at NCBI GEO #GSE202586.
Abbreviations
- RT:
-
Radiotherapy
- IR:
-
Ionizing radiation
- miRNAs:
-
MicroRNAs
- lncRNAs:
-
Long non-coding RNAs
- mRNAs:
-
Messenger RNAs
- WBI:
-
Whole body irradiation
- RILD:
-
Radiation induced liver disease
References
Montay-Gruel, P., Meziani, L., Yakkala, C. & Vozenin, M. Expanding the therapeutic index of radiation therapy by normal tissue protection. Br. J. Radiol. 92(1093), 20180008 (2019).
Kim, J., Jenrow, K. & Brown, S. Mechanisms of radiation-induced normal tissue toxicity and implications for future clinical trials. Radiat. Oncol. J. 32(3), 103–115 (2014).
Preston, D. L. et al. Solid cancer incidence in atomic bomb survivors: 1958–1998. Radiat. Res. 168(1), 1–64 (2007).
Akahoshi, M. et al. Effects of radiation on fatty liver and metabolic coronary risk factors among atomic bomb survivors in Nagasaki. Hypertens. Res. 26(12), 965–970 (2003).
Tanaka, H., Hayashi, S., Ohtakara, K. & Hoshi, H. Hepatic dysfunction after radiotherapy for primary gastric lymphoma. J. Radiat. Res. 54(1), 92–97 (2013).
Klein, J. et al. Stereotactic body radiotherapy: An effective local treatment modality for hepatocellular carcinoma. Fut. Oncol. 10(14), 2227–2241 (2014).
Tomita, Y. et al. High incidence of fatty liver and insulin resistance in long-term adult survivors of childhood SCT. Bone Marrow Transpl. 46(3), 416–425 (2011).
Guha, C., Sharma, A., Gupta, S., Alfieri, A., Gorla, G., Gagandeep, S. et al. Amelioration of Radiation-induced Liver Damage in Partially Hepatectomized Rats by Hepatocyte Transplantation | Cancer Research [Internet]. (1999) [cited 2022 Feb 4]. Available from: https://cancerres.aacrjournals.org/content/59/23/5871#sec-2
Jiang, L. et al. Proteomic analysis of radiation-induced acute liver damage in a rabbit model. Dose Response. 17(4), 1559325819889508 (2019).
Zhu, W., Zhang, X., Yu, M., Lin, B. & Yu, C. Radiation-induced liver injury and hepatocyte senescence. Cell Death Discov. 7(1), 1–9 (2021).
Kjærgaard, K. et al. Hepatic regeneration following radiation-induced liver injury is associated with increased hepatobiliary secretion measured by PET in Göttingen minipigs. Sci. Rep. 10(1), 1–10 (2020).
Radwan, R. R. & Hasan, H. F. Pioglitazone ameliorates hepatic damage in irradiated rats via regulating anti-inflammatory and antifibrogenic signalling pathways. Free Radic. Res. 53(7), 748–757. https://doi.org/10.1080/1071576220191624742 (2019).
Huang, Y. et al. Clinical parameters for predicting radiation-induced liver disease after intrahepatic reirradiation for hepatocellular carcinoma. Radiat. Oncol. 11(1), 1–9 (2016).
Chen, Y. X. et al. Mesenchymal stem cell–conditioned medium prevents radiation-induced liver injury by inhibiting inflammation and protecting sinusoidal endothelial cells. J. Radiat. Res. 56(4), 700–708 (2015).
Toesca, D. A. S. et al. Strategies for prediction and mitigation of radiation-induced liver toxicity. J. Radiat. Res. 59(Suppl_1), i40–i49 (2018).
Wang, S. et al. Potential role of hedgehog pathway in liver response to radiation. PLoS ONE 8(9), e74141 (2013).
Kim, J. & Jung, Y. Radiation-induced liver disease: Current understanding and future perspectives. In Experimental and Molecular Medicine Vol. 49 e359 (Nature Publishing Group, 2017).
Liang, S. X. et al. Radiation-induced liver disease in three-dimensional conformal radiation therapy for primary liver carcinoma: The risk factors and hepatic radiation tolerance. Int. J. Radiat. Oncol. Biol. Phys. 65(2), 426–434 (2006).
Rothkamm, K. et al. Comparison of established and emerging biodosimetry assays. Radiat. Res. 180(2), 111–119 (2013).
Podralska, M. et al. Non-coding RNAs in cancer radiosensitivity: MicroRNAs and lncrnas as regulators of radiation-induced signaling pathways. Cancers (Basel). 12(6), 1–27 (2020).
Lee, KF. Chen, YC., Hsu, PW., Liu, IY., Wu, LS. MicroRNA expression profiling altered by variant dosage of radiation exposure. Biomed. Res. Int. 2014 (2014).
May, J. M., Bylicky, M., Chopra, S., Coleman, C. N. & Aryankalayil, M. J. Long and short non-coding RNA and radiation response: A review. Transl. Res. 233, 162–179 (2021).
John Liu, S. et al. CRISPRi-based radiation modifier screen identifies long non-coding RNA therapeutic targets in glioma. Genome Biol. 21(1), 1–18 (2020).
Viereck, J. & Thum, T. Circulating noncoding RNAs as biomarkers of cardiovascular disease and injury. Circ. Res. 120(2), 381–399 (2017).
Aryankalayil, M. J. et al. Analysis of lncRNA-miRNA-mRNA expression pattern in heart tissue after total body radiation in a mouse model. J. Transl. Med. 19(1), 336 (2021).
Schmitt, A. M. et al. An inducible long noncoding RNA amplifies DNA damage signaling. Nat. Genet. 48(11), 1370 (2016).
Beer, L. et al. Ionizing radiation regulates long non-coding RNAs in human peripheral blood mononuclear cells. J. Radiat. Res. 58(2), 201–209 (2017).
Aryankalayil, M. J. et al. Radiation-induced long noncoding rnas in a mouse model after whole-body irradiation. Radiat. Res. 189(3), 251–263 (2018).
Fendler, W. et al. Evolutionarily conserved serum microRNAs predict radiation-induced fatality in nonhuman primates. Sci. Transl. Med. 9(379), 1–12 (2017).
Glinge, C. et al. Stability of circulating blood-based microRNAs – pre-analytic methodological considerations. PLoS ONE 12(2), e0167969 (2017).
Balzano, F. et al. miRNA stability in frozen plasma samples. Molecules 20(10), 19030–19040 (2015).
Li, Y. et al. Identification of lncRNA, MicroRNA, and mRNA-Associated CeRNA network of radiation-induced lung injury in a mice model. Dose Response. 17(4), 1559325819891012 (2019).
Aryankalayil, M. J. et al. Microarray analysis of miRNA expression profiles following whole body irradiation in a mouse model. Biomarkers 23(7), 689–703 (2018).
Price, J. G. et al. CDKN1A regulates Langerhans cell survival and promotes Treg cell generation upon exposure to ionizing irradiation. Nat Immunol. 16(10), 1060–1068 (2015).
Ghandhi, S. A., Smilenov, L., Shuryak, I., Pujol-Canadell, M. & Amundson, S. A. Discordant gene responses to radiation in humans and mice and the role of hematopoietically humanized mice in the search for radiation biomarkers. Sci. Rep. 9(1), 1–13 (2019).
Himburg, H. et al. A molecular profile of the endothelial cell response to ionizing radiation. Radiat. Res. 186(2), 141–152 (2016).
van de Vosse, E., van Dissel, J. T. & Ottenhoff, T. H. Genetic deficiencies of innate immune signalling in human infectious disease. Lancet Infect Dis. 9(11), 688–698 (2009).
Kawase, T. et al. PH domain-only protein PHLDA3 is a p53-regulated repressor of Akt. Cell 136(3), 535–550 (2009).
Hsiao, C., Reddy, S., Chen, M. & Saligan, L. Genomic profile of fatigued men receiving localized radiation therapy. Biol. Res. Nurs. 18(3), 281–289 (2016).
Sun, Z. et al. An RNA-seq-based expression profiling of radiation-induced esophageal injury in a rat model. Dose Response 17(2), 1559325819843373 (2019).
Azimzadeh, O. et al. Chronic occupational exposure to ionizing radiation induces alterations in the structure and metabolism of the heart: A proteomic analysis of human formalin-fixed paraffin-embedded (FFPE) cardiac tissue. Int J Mol Sci. 21(18), 1–21 (2020).
Roudkenar, M. H. et al. Gene expression profiles in mouse liver cells after exposure to different types of radiation. J. Radiat. Res. 49(1), 29–40 (2008).
Pervin, M., Unno, K., Konishi, T. & Nakamura, Y. L-arginine exerts excellent anti-stress effects on stress-induced shortened lifespan, cognitive decline and depression. Int. J. Mol. Sci. 22(2), 508 (2021).
Kalousi, A. et al. The nuclear oncogene SET controls DNA repair by KAP1 and HP1 retention to chromatin. Cell Rep. 11(1), 149–163 (2015).
Pérez-Castro, A. J. & Freire, R. Rad9B responds to nucleolar stress through ATR and JNK signalling, and delays the G1–S transition. J. Cell Sci. 125(5), 1152–1164 (2012).
Nalesso, G. et al. WNT16 antagonises excessive canonical WNT activation and protects cartilage in osteoarthritis. Ann. Rheum. Dis. 76(1), 218–226 (2017).
Khozouz, R. F., Huq, S. Z. & Perry, M. C. Radiation-induced liver disease. J. Clin. Oncol. 26(29), 4844–4845 (2008).
Lee, I. J., Seong, J., Shim, S. J. & Han, K. H. Radiotherapeutic parameters predictive of liver complications induced by liver tumor radiotherapy. Int. J. Radiat. Oncol. Biol. Phys. 73(1), 154–158 (2009).
Lawrence, T. S. et al. Hepatic toxicity resulting from cancer treatment. Int. J. Radiat. Oncol. Biol. Phys. 31(5), 1237–1248 (1995).
Chiba, M. et al. Serum miR-375-3p increase in mice exposed to a high dose of ionizing radiation. Sci. Rep. 8(1), 1–11 (2018).
Macaeva, E., Mysara, M., De Vos, W. H., Baatout, S. & Quintens, R. Gene expression-based biodosimetry for radiological incidents: Assessment of dose and time after radiation exposure. Int. J. Radiat. Biol. 95(1), 64–75 (2019).
Lindeman, L. C. et al. Gamma radiation induces locus specific changes to histone modification enrichment in zebrafish and Atlantic salmon. PLoS ONE 14(2), e0212123 (2019).
Juárez-Hernández, E. et al. Association between serum hemoglobin levels and non alcoholic fatty liver disease in a mexican population. Ann. Hepatol. 17(4), 577–584 (2018).
Chung, G. et al. Associations between hemoglobin concentrations and the development of incidental metabolic syndrome or nonalcoholic fatty liver disease. Dig Liver Dis. 49(1), 57–62 (2017).
Giorgio, V. et al. Elevated hemoglobin level is associated with advanced fibrosis in pediatric nonalcoholic fatty liver disease. J. Pediatr. Gastroenterol. Nutr. 65(2), 150–155 (2017).
Liu, W., Baker, S., Baker, R., Nowak, N. & Zhu, L. Upregulation of hemoglobin expression by oxidative stress in hepatocytes and its implication in nonalcoholic steatohepatitis. PLoS ONE 6(9), e24363 (2011).
Brosh, R. et al. p53-dependent transcriptional regulation of EDA2R and its involvement in chemotherapy-induced hair loss. FEBS Lett. 584(11), 2473–2477 (2010).
Broustas, C. et al. Impact of neutron exposure on global gene expression in a human peripheral blood model. Radiat. Res. 187(4), 433–440 (2017).
Han, C., Lim, S., Koo, J., Kim, W. & Kim, S. PHLDA3 overexpression in hepatocytes by endoplasmic reticulum stress via IRE1-Xbp1s pathway expedites liver injury. Gut 65(8), 1377–1388 (2016).
Moding, E. J. et al. An extra copy of p53 suppresses development of spontaneous Kras-driven but not radiation-induced cancer. JCI Insight 1(10), 86698 (2016).
Sproull, M., Shankavaram, U. & Camphausen, K. Novel murine biomarkers of radiation exposure using an aptamer-based proteomic technology. Front. Pharmacol. 26(12), 943 (2021).
Filiano, A. N. et al. Gene expression analysis in radiotherapy patients and C57BL/6 mice as a measure of exposure to ionizing radiation. Radiat. Res. 176(1), 49–61 (2011).
Robinson, M., Harmon, C. & O’Farrelly, C. Liver immunology and its role in inflammation and homeostasis. Cell Mol. Immunol. 13(3), 267–276 (2016).
Pavlasova, G. & Mraz, M. The regulation and function of CD20: an “enigma” of B-cell biology and targeted therapy. Haematologica 105(6), 1494–1506 (2020).
Paterson, M. A., Hosking, P. S. & Coughlin, P. B. Expression of the serpin centerin defines a germinal center phenotype in B-cell lymphomas. Am. J. Clin. Pathol. 130(1), 117–126 (2008).
Lin, L., Yee, S. W., Kim, R. B. & Giacomini, K. M. SLC transporters as therapeutic targets: Emerging opportunities. In Nature Reviews Drug Discovery Vol. 14 543–560 (Nature Publishing Group, 2015).
He, L., Vasiliou, K. & Nebert, D. W. Analysis and update of the human solute carrier (SLC) gene superfamily. Hum. Genomics. 3(2), 195 (2009).
Romero, M. F., Chen, A. P., Parker, M. D. & Boron, W. F. The SLC4 family of bicarbonate ([…formula…]) Transporters. Mol. Aspects Med. 34(2–3), 159 (2013).
Pramod, A. B., Foster, J., Carvelli, L. & Henry, L. K. SLC6 transporters: Structure, function, regulation, disease association and therapeutics. Mol. Aspects Med. 34(2–3), 197 (2013).
Halestrap, A. P. The SLC16 gene family – Structure, role and regulation in health and disease. Mol. Aspects Med. 34(2–3), 337–349 (2013).
Felmlee, M. A., Jones, R. S., Rodriguez-Cruz, V., Follman, K. E. & Morris, M. E. Monocarboxylate transporters (SLC16): Function, regulation, and role in health and disease. Pharmacol. Rev. 72(2), 466–485 (2020).
Palmieri, F. The mitochondrial transporter family SLC25: Identification, properties and physiopathology. Mol. Aspects Med. 34(2–3), 465–484 (2013).
Ruprecht, J. J. & Kunji, E. R. S. The SLC25 mitochondrial carrier family: Structure and mechanism. Trends Biochem. Sci. 45(3), 244–258 (2020).
Nigam, S. K. The SLC22 transporter family: A paradigm for the impact of drug transporters on metabolic pathways, signaling, and disease. Annu. Rev. Pharmacol. Toxicol. 6(58), 663 (2018).
Volk, C. OCTs, OATs, and OCTNs: structure and function of the polyspecific organic ion transporters of the SLC22 family. Wiley Interdiscip. Rev. Membr. Transp. Signal. 3(1), 1–13. https://doi.org/10.1002/wmts.100 (2014).
Juraszek, B. & Nałąecz, K. A. SLC22A5 (OCTN2) carnitine transporter—indispensable for cell metabolism, a Jekyll and hyde of human cancer. Molecules 25(1), 14 (2020).
Song, P., Onishi, A., Koepsell, H. & Vallon, V. Sodium glucose cotransporter SGLT1 as a therapeutic target in diabetes mellitus. Expert Opin. Ther. Targets. 20(9), 1109 (2016).
Kuang, W. et al. SLC22A14 is a mitochondrial riboflavin transporter required for sperm oxidative phosphorylation and male fertility. Cell Rep. 35(3), 109025 (2021).
Park, S. et al. Suppression of ABCD2 dysregulates lipid metabolism via dysregulation of miR-141: ACSL4 in human osteoarthritis. Cell Biochem. Funct. 36(7), 366–376 (2018).
Wang, C. Y. et al. A novel nonsense mutation of ABCA8 in a han-Chinese family with ASCVD Leads to the reduction of HDL-c levels. Front Genet. 15(11), 755 (2020).
Oshio, Y. et al. Very low-density lipoprotein receptor increases in a liver-specific manner due to protein deficiency but does not affect fatty liver in mice. Sci. Rep. 11(1), 1–11 (2021).
Gao, Y. et al. Upregulation of hepatic VLDLR via PPARα is required for the triglyceride-lowering effect of fenofibrate. J. Lipid Res. 55(8), 1622 (2014).
Qu, W. et al. Cytochrome P450 CYP2J9, a new mouse arachidonic acid ω-1 hydroxylase predominantly expressed in brain. J. Biol. Chem. 276(27), 25467–25479 (2001).
Chuang, S. S. et al. CYP2U1, a novel human thymus- and brain-specific cytochrome P450, Catalyzes ω- and (ω-1)-hydroxylation of fatty acids. J. Biol. Chem. 279(8), 6305–6314 (2004).
Ellero, S. et al. Xenobiotic-metabolizing cytochromes P450 in human white adipose tissue: Expression and induction. Drug Metab. Dispos. 38(4), 679–686 (2010).
Christmas, P. et al. Cytochrome P-450 4F18 Is the Leukotriene B4 ω-1/ω-2 Hydroxylase in Mouse Polymorphonuclear Leukocytes: Identification as the functional orthologue of human polymorphonuclear leukocyte CYP4F3A in the down-regulation of responses to LTB4. J. Biol. Chem. 281(11), 7189–7196 (2006).
Kometani, M. et al. Cortisol overproduction results from DNA methylation of CYP11B1 in hypercortisolemia. Sci. Rep. 7(1), 1–9 (2017).
Topletz, A. R. et al. Comparison of the function and expression of CYP26A1 and CYP26B1, the two retinoic acid hydroxylases. Biochem. Pharmacol. 83(1), 149 (2012).
Skulas-Ray, A. C. et al. Omega-3 fatty acids for the management of hypertriglyceridemia: A science advisory from the American Heart Association. Circulation 140(12), E673–E691 (2019).
Si, Z. et al. CYP46A1 as a new regulator of lipid metabolism through CRISPR-based whole-genome screening. FASEB J. 34(10), 13776–13791. https://doi.org/10.1096/fj.202001067R (2020).
Yang, Y. et al. Role of glutathione S-transferases in protection against lipid peroxidation overexpression of hGSTA2–2 in K562 cells protects against hydrogen peroxide-induced apoptosis and inhibits JNK and caspase 3 activation. J. Biol. Chem. 276(22), 19220–19230 (2001).
Gamage, N. et al. Human sulfotransferases and their role in chemical metabolism. Toxicol. Sci. 90(1), 5–22 (2006).
Oh, E. T. & Park, H. J. Implications of NQO1 in cancer therapy. BMB Rep. 48(11), 609–617 (2015).
Ross, D. & Siegel, D. Functions of NQO1 in cellular protection and CoQ10 metabolism and its potential role as a redox sensitive molecular switch. Front. Physiol. 8, 595 (2017).
Wang, S. et al. Potential role of hedgehog pathway in liver response to radiation. PLoS ONE 8(9), e74141 (2013).
Lysek-Gladysinska, M. et al. Long-term effects of low-dose mouse liver irradiation involve ultrastructural and biochemical changes in hepatocytes that depend on lipid metabolism. Radiat. Environ. Biophys. 57(2), 123–132 (2018).
Kim, C. W. et al. Acetyl CoA carboxylase inhibition reduces hepatic steatosis but elevates plasma triglycerides in mice and humans: A bedside to bench investigation. Cell Metab. 26(2), 394 (2017).
Burhans, M. S. et al. Hepatic oleate regulates adipose tissue lipogenesis and fatty acid oxidation. J. Lipid Res. 56(2), 304–318 (2015).
Wang, C. et al. Perilipin 5 improves hepatic lipotoxicity by inhibiting lipolysis. Hepatology 61(3), 870–882 (2015).
Wang, H. et al. Mutations in SREBF1, encoding sterol regulatory element binding transcription factor 1, cause autosomal-dominant IFAP syndrome. Am. J. Hum. Genet. 107(1), 34–45 (2020).
Moslehi, A. & Hamidi-Zad, Z. Role of SREBPs in liver diseases: A mini-review. J Clin Transl Hepatol. 6(3), 332 (2018).
Ferguson, P. J. & El-Shanti, H. Majeed syndrome: A review of the clinical, genetic and immunologic features. Biomolecules 11(3), 1–16 (2021).
Gao, X. F. et al. Enhanced susceptibility of Cpt1c knockout mice to glucose intolerance induced by a high-fat diet involves elevated hepatic gluconeogenesis and decreased skeletal muscle glucose uptake. Diabetologia 52(5), 912–920 (2009).
Leslie, N. et al. Neonatal multiorgan failure due to ACAD9 mutation and complex I deficiency with mitochondrial hyperplasia in liver, cardiac myocytes, skeletal muscle, and renal tubules. Hum. Pathol. 1(49), 27–32 (2016).
Hong, Y. B. et al. A compound heterozygous mutation in HADHB gene causes an axonal Charcot-Marie-tooth disease. BMC Med. Genet. 14(1), 1–8. https://doi.org/10.1186/1471-2350-14-125 (2013).
Martius, G. et al. Hepatic fat accumulation and regulation of FAT/CD36: An effect of hepatic irradiation. Int. J. Clin. Exp. Pathol. 7(8), 5379 (2014).
Chartoumpekis, D. V. et al. Differential EXPRESSION of MicroRNAs in adipose tissue after long-term high-fat diet-induced obesity in mice. PLoS ONE 7(4), e34872. https://doi.org/10.1371/journal.pone.0034872 (2012).
Su, T. et al. MiR-34a-5p and miR-452–5p: The novel regulators of pancreatic endocrine dysfunction in diabetic Zucker rats? Int. J. Med. Sci. 18(14), 3171 (2021).
Hung, Y. H. et al. Acute suppression of insulin resistance-associated hepatic miR-29 in vivo improves glycemic control in adult mice. Physiol. Genom. 51(8), 379–389. https://doi.org/10.1152/physiolgenomics.00037.2019 (2019).
Kobayashi, M. et al. Iron-heme-Bach1 axis is involved in erythroblast adaptation to iron deficiency. Haematologica 102(3), 454–465 (2017).
Recio, L. et al. Differential expression of long noncoding RNAs in the livers of female B6C3F1 mice exposed to the carcinogen furan. Toxicol. Sci. 135(2), 369–379 (2013).
Huarte, M. et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142(3), 409–419 (2010).
Plett, P. et al. Establishing a murine model of the hematopoietic syndrome of the acute radiation syndrome. Health Phys. 103(4), 343–355 (2012).
Chopra, S. et al. Gene expression profiles from heart, lung and liver samples of total-body-irradiated minipigs: Implications for predicting radiation-induced tissue toxicity. Radiat. Res. 194(4), 411–430 (2020).
Halimi, M. et al. Human serum miR-34a as an indicator of exposure to ionizing radiation. Radiat. Environ. Biophys. 55(4), 423–429 (2016).
Liu, C. et al. MiR-34a in Age and tissue related radio-sensitivity and serum miR-34a as a novel indicator of radiation injury. Int. J. Biol. Sci. 7(2), 221 (2011).
Shen, Y. et al. miR-34a and miR-125b are upregulated in peripheral blood mononuclear cells from patients with type 2 diabetes mellitus. Exp. Ther. Med. 14(6), 5589–5596. https://doi.org/10.3892/etm.2017.5254/abstract (2017).
Yannam, G. R. et al. A nonhuman primate model of human radiation-induced venocclusive liver disease and hepatocyte injury. Int. J. Radiat. Oncol. Biol. Phys. 88(2), 404–411 (2014).
Shi, J. et al. PANDORA-seq expands the repertoire of regulatory small RNAs by overcoming RNA modifications. Nat. Cell Biol. 23(4), 424 (2021).
Shi, J., Zhou, T. & Chen, Q. Exploring the expanding universe of small RNAs. Nat. Cell Biol. 24(4), 415–423 (2022).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucl. Acids Res. 43(7), e47 (2015).
Ritchie, M. E. et al. A comparison of background correction methods for two-colour microarrays. Bioinformatics 23(20), 2700–2707 (2007).
Smyth, G. K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3(1), (2004).
Aken, B.L., Ayling, S., Barrell, D., Clarke, L., Curwen, V., Fairley, S., et al. The Ensembl gene annotation system. Database 2016 (2016).
Agarwal, V., Bell, G. W., Nam, J. W. & Bartel, D. P. Predicting effective microRNA target sites in mammalian mRNAs. Elife 4, e05005 (2015).
Acknowledgements
This study was supported by the NIH Intramural Research Program, National Cancer Institute, Center for Cancer Research and National Institute of Allergy and Infectious Diseases (IAA no. NRC-13028). and from the Radiation and Nuclear Countermeasures Program, #Y2-OD-0332-01 NIAID. We thank Dr. Eric Bernhard for his expert advice in editing the manuscript.
Funding
This work was supported by NIAID (IAA #NRC-13028).
Author information
Authors and Affiliations
Contributions
Conceptualization: M.J.A. and C.N.C.; Methodology: M.J.A., M.A.B., S.M.,J.M.M., M.S., L.M.; investigation, M.J.A., M.A.B., S.M., C.V.-B., J.D.; writing – original draft, M.J.A., M.A.B., S.C., S.M., A.S.; writing – review and editing, M.J.A., M.A.B., C.N.C, S.M., S.C.K.,M.K.S., and C.N.C.; funding acquisition, M.J.A., C.N.C.; supervision, M.J.A., M.A.B. and C.N.C. Consent for publication: All the authors have read and approved the manuscript in all respects for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Aryankalayil, M.J., Bylicky, M.A., Martello, S. et al. Microarray analysis identifies coding and non-coding RNA markers of liver injury in whole body irradiated mice. Sci Rep 13, 200 (2023). https://doi.org/10.1038/s41598-022-26784-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-26784-w
- Springer Nature Limited