Abstract
Oncocytic thyroid cancer is characterized by the aberrant accumulation of abnormal mitochondria in the cytoplasm and a defect in oxidative phosphorylation. We performed metabolomics analysis to compare metabolic reprogramming among the oncocytic and non-oncocytic thyroid cancer cell lines XTC.UC1 and TPC1, respectively, and a normal thyroid cell line Nthy-ori 3-1. We found that although XTC.UC1 cells exhibit higher glucose uptake than TPC1 cells, the glycolytic intermediates are not only utilized to generate end-products of glycolysis, but also diverted to branching pathways such as lipid metabolism and the serine synthesis pathway. Glutamine is preferentially used to produce glutathione to reduce oxidative stress in XTC.UC1 cells, rather than to generate α-ketoglutarate for anaplerotic flux into the TCA cycle. Thus, growth, survival and redox homeostasis of XTC.UC1 cells rely more on both glucose and glutamine than do TPC1 cells. Furthermore, XTC.UC1 cells contained higher amounts of intracellular amino acids which is due to higher expression of the amino acid transporter ASCT2 and enhanced autophagy, thus providing the building blocks for macromolecules and energy production. These metabolic alterations are required for oncocytic cancer cells to compensate their defective mitochondrial function and to alleviate excess oxidative stress.
Similar content being viewed by others
Introduction
Thyroid oncocytic carcinoma, also known as Hürthle cell carcinoma, has a unique phenotype that is characterized by its enlarged oxyphilic cytoplasm where mitochondria are aberrantly accumulated1,2. In the most but not all of these cancers, mitochondria are dysfunctional due to missense or less frequently disruptive mutations in the mitochondrial DNA (mtDNA) coding for complex I and/or III of the electron transport chain (ETC), which is essential for oxidative phosphorylation (OXPHOS)3,4,5,6, although the mere presence of such mtDNA mutations does not necessarily mean tumors that the tumors are oncocytic7. In addition to a compensatory increase in mitochondrial biogenesis8,9,10, we have recently demonstrated that the oncocytic phenotype (that is, the accumulation of mitochondria) is also caused by defective expression of mitochondria-eating protein (MIEAP)11, an indispensable molecule for non-canonical mitophagy (see12 for a review). Thus, this type of tumor must totally rely on glycolysis for energy production by shifting from OXPHOS to aerobic glycolysis (called the Warburg effect)13. It has been documented that metabolic reprogramming like this is a hallmark of cancer14. Indeed it has previously been shown that glucose in the culture medium is indispensable for in vitro growth/survival and ATP production of the oncocytic thyroid cancer cell line XTC.UC115,16. XTC.UC1 cells have also been shown to exhibit a lower basal oxygen consumption rate (OCR) and a higher basal extracellular acidification rate (ECAR)17,18. Furthermore, higher expression of glycolysis-related proteins, glucose transporter 1, hexokinase II, carbonic anhydrase IX and monocarboxylate transporter 4, has been demonstrated in oncocytic thyroid cancer tissues compared to non-oncocytic cancer tissues19. Although energy production by glycolysis is inefficient from an energetic perspective, it fulfills the biomass requirements for rapid cell proliferation.
However, there have been contradictory data reported; for example, the expression levels of lactate dehydrogenase (LDH) have been shown to be lower in oncocytic thyroid tumors than non-oncocytic tumors10. Similar data have also been reported in renal and pituitary oncocytomas, compared to normal kidney tissues and non-oncocytic pituitary adenomas, respectively, showing diminished glycolysis with low glucose 6-phosphate, low to normal lactate and low LDH levels20,21. Normalization of mitochondrial complex I function by gene transfer in osteosarcoma cells with impaired complex I function has been shown to increase LDH expression, indicating that a functional complex I is essential for the induction of glycolysis22.
These controversial data prompted us to perform metabolomics analyses and compare the metabolic statuses of the oncocytic thyroid cancer cell line XTC.UC1, the non-oncocytic papillary cancer cell line TPC1, and the immortalized normal thyroid cell line Nthy-ori 3-1; the latter two are widely used to compare metabolic states with XTC.UC1 cells15,18. Metabolomics is an emerging omics technology that is useful for systematically quantifying metabolites in biological samples. We also generated a mtDNA-defective TPC1 cell line (ρ0 cells) to see if the phenotypic differences between XTC.UC1 and TPC1 cells are due to differences in mitochondrial function.
Results
Seahorse analysis
First, we used a Seahorse XF-96 extracellular flux analyzer to compare OCR among XTC.UC1, TPC1, TPC1 ρ0 and Nthy-ori 3-1 cells (Fig. 1). TPC1 and Nthy-ori 3-1 cells exhibited comparably normal patterns of basal and maximal OCR in response to treatment with mitochondrial complex inhibitors, while TPC1 ρ0 cells showed extremely low basal OCR and no response to treatments, confirming normal and completely abolished mitochondrial function in the parental TPC1 and TPC1 ρ0 cells, respectively. XTC.UC1 cells exhibited an intermediate pattern, that is, lower basal OCR and partial response to mitochondrial inhibitors, data consistent with Addie et al.23, indicating the ETC in XTC.UC1 cells being partially impaired. In contrast, another group has reported a slightly different data on mitochondrial function in XTC.UC1 cells, indicating almost complete abolishment of mitochondrial function18.
Mitochondrial oxygen consumption rates in Nthy-ori 3-1, XTC.UC1, TPC1 and TPC1 ρ0 cells. The oxygen consumption rates were measured at the basal levels and after 2 μM oligomycin (O), 0.25 μM phenylhydrazone (P) and 0.5 μM rotenone/antimycin (R/A) treatments as described in the “Materials and methods”. Blue, Nthy-ori 3-1; yellow, XTC.UC1; red with solid lines, TPC1; red with broken lines, TPC1 ρ0 cells. Data are presented as means ± SE.
Metabolomics analysis
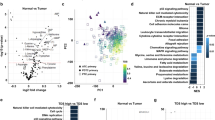
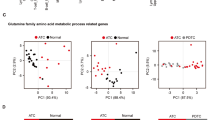
We performed C-SCOPE-based metabolomics analysis to clarify the differences in intracellular metabolites among XTC.UC1, TPC1 and Nthy-ori 3-1 cells. C-SCOPE can analyze water-soluble charged metabolites, including the glycolytic products, intermediates of the TCA cycle, as well as amino acids. All the data are illustrated in Supplementary Fig. S1 and the raw data are shown in Supplementary Table S1. The heatmap for hierarchical cluster analysis of normalized metabolic intensities, which provides a global picture of the relative amounts of metabolites, showed clear separations among the three cell lines, as did the principal component analysis (Supplementary Fig. S2A,B). Analysis of metabolite changes among the three cell lines revealed that the relative abundance of 33 metabolites was significantly (p < 0.01) altered between Nthy-ori 3-1 and TPC1 cells (19 upregulated), 45 were altered between Nthy-ori 3-1 and XTC.UC1 cells (37 upregulated) and 35 were altered between TPC1 and XTC.UC1 cells (29 upregulated). We then focused on some intriguing differences, which are summarized in Fig. 2. Regarding the glycolysis pathway, the concentration of glucose-6-phosphate was highest in XTC.UC1, indicating the highest glucose uptake. However, the concentrations of the downstream metabolites, such as glyceraldehyde 3-phosphate (G3P) and 3-phosphoglyceric acid (3PG), decreased gradually in a stepwise manner, presumably because the former was converted to dihydroxyacetone phosphate and then to glycerol 3-phosphate, and the latter to serine. G3P is thought to be used for lipid metabolism (see below), and 3PG for serine synthesis and one-carbon metabolism. Thus, substantial amounts of the metabolic intermediates in the glycolysis pathway were diverted to the production of other biomolecules in XTC.UC1 cells. Consistently, lactate was significantly (p < 0.05) but only marginally higher in XTC.UC1 than in the other two cell line. The Warburg effect was not manifest in TPC1 cells, because there were comparable glucose 6-phosphate and lactate levels between TPC1 and Nthy-ori 3-1 cells, which is consistent with a previous study17.
Representative data on glycolysis, the TCA cycle, lipid metabolism, glutathione synthesis and methionine metabolism. Blue, Nthy-ori 3-1; yellow, XTC.UC1; red with solid lines, TPC1; red with broken lines, TPC1 ρ0 cells. *, p < 0.01 vs. Nthy-ori 3-1; #, p < 0.01 vs. Nthy-ori 3-1; $, p < 0.01 vs. TPC1 cells. [Reproduced with permission from Human Metabolome Technologies, Inc., Tsuruoka, Japan with some modifications].
Turning to the TCA cycle, despite comparable levels of acetyl-CoA among three cell lines, citrate was lowest in XTC.UC1, indicating a block of glucose entry into the TCA cycle due to impaired OXPHOS. Citrate was likely to be transferred to the cytoplasm and converted back to acetyl-CoA and oxaloacetate, the former of which was then metabolized to malonyl CoA and, together with glycerol 3-phosphate (see above), was involved in fatty acid synthesis.
Low TCA cycle intermediates are usually replenished. For example, glutamine can be converted to glutamic acid and then to α-ketoglutaric acid (αKG), which is incorporated into the TCA cycle as an anaplerotic influx. However, in XTC.UC1 cells, high glutamine and glutamic acid but low αKG suggested little if any influx of glutamic acid into the TCA cycle. Instead, glutamic acid was converted to glutathione (GSH). Other amino acids required for GSH synthesis, cysteine and glycine, were also higher in XTC.UC1 than in TPC1 cells (see below).
Interestingly, succinic acid was higher, despite lower citrate and αKG in XTC.UC1 cells. Succinyl-CoA can be generated, as another anaplerotic flux, from branched chain amino acids (valine and isoleucine), threonine and methionine24, all of which were very high in XTC.UC1 cells (see below). The reaction of succinyl-CoA to succinic acid catalyzed by succinyl-CoA ligase is accompanied by ADP/GDP to ATP/GTP conversion25, called a mitochondrial substrate level phosphorylation (mtSLP), contributing to high-energy molecule production.
Furthermore, the reactions from methionine to S-adenosylmethionine (SAM) and then to S-adenosylhomocysteine (SAH) appeared to be accelerated in XTC.UC1 cells. SAM is a methyl donor for both DNA and histone methylation reactions that influence the epigenetic control of gene expression26,27. Indeed we have recently found extensive methylation of the MIEAP gene in XTC.UC1 cells11. In addition, together with serine (and glycine), methionine is involved in one-carbon metabolism27.
Glycolysis and OXPHOS dependency
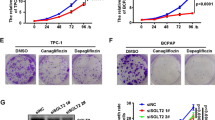
As expected from the metabolomics analysis showing the glycolytic phenotype of XTC.UC1 cells, glucose uptake was higher in XTC.UC1 cells than in TPC1 cells (Fig. 3A). In cell number measurements, cell number approximately doubled in glucose-containing medium and increased by ~ 40% in glucose-free medium in TPC1 cells in 48 h. However, in XTC.UC1 cells, although their growth pattern in glucose-containing medium was similar to that of TPC1, their cell number was reduced by ~ 40% in 48 h (Fig. 3B). Morphologically, cell death was clearly observed in XTC.UC1 cells at 24 and 48 h. To exclude the possibility that the different vulnerabilities between TPC1 and XTC.UC1 cells were due to different intrinsic phenotypes of the 2 cell types, the same experiment was performed with TPC1 ρ0 cells. Although glucose uptake was not elevated (Fig. 3A), this cell line showed a cell survival pattern similar to that of XTC.UC1 cells (Fig. 3B), clearly indicating that the difference in cell survival between TPC1 and XTC.UC1 cells shown in Fig. 3B was due to different functioning of mitochondria. Similarly, oxamate, an LDH inhibitor, killed more XTC.UC1 cells than TPC1 cells (Fig. 3C). These data are consistent with the glycolytic phenotype of XTC.UC1 cells. Indeed almost complete suppression of lactate production with oxamate was confirmed (Fig. 3D). In contrast, since TPC1 cells depend more on the TCA cycle and the ETC for energy production than XTC.UC1 cells, it was predictable that atovaquone, a complex III inhibitor of the ETC, killed more TPC1 cells than XTC.UC1 cells (Fig. 3E).
Glucose uptake (A), relative cell numbers in cells cultured in the presence and absence of glucose (B), with oxamate (C) or atovaquone (E), and the effect of oxamate on lactate production (D). Glucose uptake, cell number measurements and lactate measurements were performed as described in the “Materials and methods”; briefly, in (C–E), the cells were treated with 10–60 mM oxamate or 5–20 μM atovaquone for 48 h. In (B), red circles with solid lines, TPC1/glucose ( +); red triangles with solid lines, TPC1/glucose free; red circles with broken lines, TPC1 ρ0 cells/glucose ( +); red triangles with broken lines, TPC1 ρ0 cells/glucose free; yellow circles, XTC.UC1/glucose ( +); yellow triangles, XTC.UC1/glucose free. In (C,E): red, TPC1; yellow, XTC.UC1 cells. Data are presented as means ± SE. In (B): * and **, p < 0.01 and 0.05, respectively, vs. 0 h; in (A) and (B–D): *, p < 0.01; **, p < 0.05.
Glutamine dependency
The metabolomics data indicate the preferential use of glutamine for glutathione synthesis, not for the αKG supplementation, in XTC.UC1 cells. Thus, glutamine depletion showed little effect on TPC1 growth, but decreased cell number of XTC.UC1 by ~ 75% in 48 h (Fig. 4A). Again, the similar survival pattern between XTC.UC1 and TPC1 ρ0 cells indicates that the the difference in cell survival between the 2 cell lines was due to different function of mitochondria. In XTC.UC1 cells cultured in glutamine free medium, glutathione production was significantly decreased and ROS production was elevated, while cell death and ROS overproduction were cancelled by NAC, a ROS scavenger (Fig. 4B–D), demonstrating that elevated ROS was responsible for cell death by glutamine depletion.
Effects of glutamine depletion on cell survival in XTC.UC1, TPC1 and TPC1 ρ0 cells (A), and on GSH and ROS productions and cell survival in XTC.UC1 cells treated with/without NAC (B–D). GSH and ROS measurements and cell number measurements were performed as described in the “Materials and methods”; briefly, the cells were cultured in glutamine-containing or free medium for 48 h in (B), and in glutamine-containing or free medium with/without 4 mM NAC for 48 h in (C,D). In (A): red circles with solid lines, TPC1/ glutamine (+); red triangles with solid lines, TPC1/ glutamine free; red circles with broken lines, TPC1 ρ0 cells/ glutamine (+); red triangles with broken lines, TPC1 ρ0 cells/ glutamine free; yellow circles, XTC.UC1/ glutamine (+); yellow triangles, XTC.UC1/ glutamine free. Data are presented as means ± SE. In (A): * and **, p < 0.01 and 0.05, respectively, vs. 0 h; in (B–D): *, p < 0.01.
Amino acid metabolism
As shown in Fig. 5A, most amino acid concentrations were markedly higher in XTC.UC1 cells as compared to the other two cell lines. Higher intracellular amino acid contents are likely due to higher amino acid uptake from the culture medium and/or amino acid recycling by autophagy28. Among the many amino acid transporters so far identified, SLC7A5 (LAT1) and SLC1A5 (ASCT2) are frequently overexpressed in cancer cells29. RT-PCR analysis of mRNAs for these two amino acid transporters showed higher (> twofold) expression of ASCT2 in XTC.UC1 cells than in TPC1 cells (Fig. 5B). However, since ASCT2 transports only neutral, nonpolar amino acids and not others30, the data shown in Fig. 5A cannot be solely explained by higher ASCT2 expression. Therefore, autophagic activity was then compared between XTC.UC1 and TPC1 cells. Higher amounts of the processed LC3-II relative to the unprocessed LC3-I indicated enhanced autophagic activity in XTC.UC1 cells as compared to the comparable levels of LC3-I and LC3-II in TPC1 cells (Fig. 5C and Supplementary Fig. S3). Indeed inhibition of autophagy with chloroquine significantly decreased amino acid levels in XTC.UC1 cells (Fig. 5D,E). It should be noted here that XTC.UC1 cells have defective mitophagy due to a loss-of-function mutation in PARK218 and defective expression of MIEAP11. Therefore, mitochondria cannot be efficiently eliminated in this cell line, although autophagic flux is increased.
Intracellular amino acid concentrations in Nthy-ori 3-1, TPC1 and XTC.UC1 cells (A); mRNA expression of amino acid transporters LAT1 and ASCT2 (B); autophagy activities in XTC.UC1 and TPC1 cells (B,C); and the effect of the autophagy inhibitor chloroquine on autophagic activity and intracellular amino acid levels in XTC.UC1 cells (D,E). The results in (A) are from Supplementary Fig. S1, and RT-PCR for LAT1 and ASCT2, mRNAs, WB for LC3, p62 and β-actin and intracellular amino acid measurements were performed as described in the “Materials and methods”. In A, blue, Nthy-ori 3-1; yellow, XTC.UC1; red, TPC1 cells. Data are presented as means ± SE (n = 3 for amino acid levels) or range (n = 2 for RT-PCR). *, p < 0.01 vs. Nthy-ori 3-1; #, p < 0.01 vs. Nthy-ori 3-1; $, p < 0.01 vs. TPC1 cells. In WB in (C,D), the blots were cropped, and uncropped images are shown in Supplementary Fig. S4A,B.
Discussion
In the present study, we first compared the metabolic profiles among oncocytic and non-oncocytic thyroid cancer cell lines, XTC.UC1 and TPC1, and a normal thyroid cell line, Nthy-ori 3-1. Although glucose uptake was indeed higher in XTC.UC1 cells than in TPC1 cells, an enhanced glycolytic pathway not only generates the end-products of glycolysis (pyruvate, lactate and ATP) but also provides the metabolic intermediates for lipid metabolism and the serine synthetic pathway (SSP) in XTC.UC1 cells. Thus, although previous studies15,16 have demonstrated that XTC.UC1 cells largely rely on glycolysis for ATP production and cell survival, and our current results show that XTC.UC1 cells and TPC1 ρ0 cells are more vulnerable to glucose depletion and the LDH inhibitor oxamate than are TPC1 cells, ATP production by the glycolytic pathway may not be improved as much by higher glucose uptake. Lower ATP production by glycolysis is likely compensated by the much faster reactions of glycolysis than OXPHOS, but it is known that glycolysis intermediates are actively deviated from energy production to the provision of precursors necessary for macromolecule biosynthesis and cell proliferation in cancer cells31. Even in non-oncocytic thyroid cancers, the importance of lipid and serine metabolism has been reported32,33,34, and the expression levels of SSP-related enzymes have been shown to be high, especially in thyroid cancers with mutant BRAF34. Our data indicate that this type of metabolic reprogramming is more notable in oncocytic thyroid cancers, which is consistent with a previous article reporting the induction of the SSP by a complex I deficit35. It has also been reported that enzymes involved in the SSP and one-carbon metabolism are upregulated by the accumulation of OXPHOS defects in colon epithelial cells of Lgr5-creER;Apc f/f; PolgA mut/mut mice, in which the accumulation of mtDNA mutations are accelerated35.
High concentrations of succinate and some amino acids that can be converted to succinyl-CoA (valine, isoleucine, threonine and methionine) imply that glutamine-independent, anaplerotic mtSLP appears to contribute to ATP production to some extent. Further studies are necessary to clarify this issue.
The lower levels of citrate and αKG in XTC.UC1 cells make sense, because impaired OXPHOS hampers the entry of glucose metabolites into the TCA cycle. Lower levels of the TCA cycle intermediates are usually replenished, and glutaminolysis is the most important anaplerotic pathway as mentioned above. αKG is used either to generate succinate and ATP/GTP from succinyl-CoA and ADP/GDP (mtSLP)25 and then pyruvate and lactate through malate36 (collectively called the oxidative TCA cycle)28,37, or to generate citrate for fatty acid synthesis (called reductive carboxylation)38, especially in the TCA cycle or OXPHOS-defective cancer cells or cancer cells treated with metformin (a complex I inhibitor)23,39,40,41. Alternatively, glutamine is used to synthesize nucleotides42 or glutathione23. Our data showing low αKG and high glutathione indicate that glutamine is preferentially used for glutathione synthesis, not for the anaplerotic pathway into the TCA cycle, in XTC.UC1 cells. This may be at least in part attributed to higher ROS production in XTC.UC1 cells11,15. Indeed we showed that this ROS-suppressing pathway is critical for cell survival. High glutathione synthesis has also been demonstrated in renal oncocytoma with high intracellular ROS levels20,43,44. The higher concentration of fumarate observed in XTC.UC1 cells (Fig. 2) has also been reported to participate in redox homeostasis by activating glutathione peroxidase 1, which eliminates ROS with glutathione45.
Higher concentrations of intracellular amino acids have been reported in cancer cells in general46, and glycolytic intermediates supply cancer cells with the amino acids required for fast cellular proliferation47. We revealed here that both higher expression of the amino acid transporter ASCT2 and augmented autophagic activity contributed to higher amounts of amino acids in XTC.UC1 cells, although overexpression of LAT1 and ASCT2 have previously been reported even in non-oncocytic thyroid cancer cells and are correlated with poor patient prognosis48,49,50,51. A previous study has revealed higher autophagic activity in XTC.UC1 than in Nthy-ori 3-1 cells, which compensates dysfunctional mitophagy by the PARK2 mutation18. As mentioned above, these amino acids are likely used for glutathione synthesis, TCA cycle anaplerosis, methylation, etc., in XTC.UC1 cells.
While we were performing this research project, Addie et al.23 published a metabolomic analysis of thyroid cancer cell lines with or without “near-homozygous genome” including XTC.UC1 cells (lower complex I and II activities), FTC-236 cells (lower complex I, II and IV activities), and SW579 and BHP2-7 cells (both having normal OXPHOS). A near-homozygous genome is a chromosomal abnormality frequently observed in oncocytoma3,52, and BHP2-7 cells are genetically identical to TPC1 cells53. Although it is very difficult to interpret their data because each cell line exhibited unique metabolic characteristics. When compared the data on XTC.UC1 and TPC1 (BHP2-7) cells between our study and theirs, there are some inconsistent data between 2 studies. For examples, although the concentrations of many amino acids were commonly increased in XTC.UC1 compared to TPC1 (BHP2-7) cells in both studies, those of asparagine and arginine were declined in ours, and glutamine, serine and tryptophan were reduced in theirs. Furthermore, lactate was higher in ours but lower in theirs, and citrate was lower in ours but higher in theirs in XTC.UC1 compared to TPC1 (BHP2-7) cells (citrate; see below).
The reasons for these discrepancies are unknown; they may represent heterogeneity within these cell lines among different laboratories. Some characteristics of XTC.UC1 and/or TPC1 cells might have changed during a long period of in vitro cell culture. Indeed there is one difference in genotyping data of XTC.UC1 cells between Cellosaurus and ours (Supplementary Fig. S2).
In conclusion, our data demonstrate that XTC.UC1 cells exhibit higher glycolytic activity than TPC1 cells, but the glycolytic intermediates are utilized not only for the completion of glycolysis but also for supplementation of branching pathways such as lipid metabolism and the SSP. Growth, survival and redox homeostasis of XTC.UC1 cells depend more on both glucose and glutamine than in TPC1 cells. Finally, higher intracellular amino acid contents, which are due to higher glycolytic activity, and enhanced cellular uptake and autophagy, provide the building blocks for macromolecule and energy production. These metabolic alterations are needed for oncocytic cancer cells to compensate for their defective mitochondrial function and to alleviate excess oxidative stress.
Materials and methods
Cell lines used in this study
The oncocytic thyroid cancer cell line XTC.UC1 (kindly provided by prof. Porcelli AM, Bologna University, Italy)54, the non-oncocytic papillary thyroid cancer cell line TPC1 (kindly provided by prof. Fagin JA, Sloan Kettering Cancer Center, NY, USA)55 and the immortalized normal thyroid cell line Nthy-ori 3-1 (from the Health Protection Agency Culture Collections, Salisbury, UK)56 were used in this study. TPC1 cells have the RET-PTC1 chromosome translocation. XTC.UC1 cells have mutations in the ND1, cytochrome b15 and p53 (a homozygous 451C > A, corresponding to Pro151Thr) genes57 but no point mutations in BRAF or RAS, and no chromosomal translocation in RET, NTRK1 or ALK (data not shown). The authenticity of TPC1 was confirmed by short tandem repeat profiling previously58, and those of XTC.UC1 and Nthy-ori 3-1 were examined in this study and found to be the same as those in the Cellosaurus except for one focus in XTC.UC1 (FGA) (Supplementary Table S2) (National Institute of Biomedicine Innovation, Ibaraki, Japan).
XTC.UC1 cells were cultured in DMEM (08459-64, Nacalai Tesque Inc., Kyoto, Japan) with 10% FBS and antibiotics, TPC1 cells in RPMI 1640 (30264-85, Nacalai Tesque) with 5% FBS and antibiotics and Nthy-ori 3-1 cells in RPMI 1640 with 10% FBS and antibiotics. For glucose-free culture, RPMI 1640 (No Glucose) with L-glutamine (09892-15, Nacalai Tesque) and DMEM (No Glucose) with L-glutamine (09891-25, Nacalai Tesque) were used, and for glutamine-free culture, RPMI 1640 without L-Gln (05176-25, Nacalai Tesque) and DMEM (4.5 g/L glucose) without L-Gln (08488-55, Nacalai Tesque) were used. None of media contained pyruvate. Cell culture was performed at 37 °C in a humidified incubator with 5% CO2.
Metabolite measurements
Metabolic extracts were prepared from ~ 5 × 106 cells with methanol containing Internal Standard Solution (H3304–1002, Human Metabolome Technologies (HMT), Inc., Tsuruoka, Japan), transferred into a microfuge tube and centrifuged at 2300×g and 4 °C for 5 min. The upper aqueous layer was then centrifugally filtered through a Millipore 5-kDa cutoff filter at 9100×g at 4 °C for 120 min to remove proteins. The filtrate was centrifugally concentrated and resuspended in 50 μL dH2O, and analyzed using a capillary electrophoresis (CE)-connected ESI-TOFMS and CE-MS/MS system (HMT, CARCINOSCOPE). Cationic compounds were analyzed in the positive mode of CETOFMS and anionic compounds were analyzed in the positive and negative modes of CE-MS/MS. The data were analyzed by MasterHands ver.2.17.1.11 (Keio University, Tsuruoka, Japan) and MassHunter Quantitative Analysis B.06.00 (Agilent Technologies, Santa Clara, CA), and the peak area of each metabolite was calculated. The peak area was then normalized to an internal standard, and the metabolite concentrations were calculated using standard curves. Overall, 116 compounds were measured59,60.
Generation of TPC1 ρ0 cells
TPC1 cells in a 10 cm dish (1 × 106 cells) were cultured for 8 weeks with medium containing 0.5 μg/mL ethidium bromide (315-90051, Nippon Gene, Tokyo, Japan), 50 μg/mL uridine (U3003, Sigma-Aldrich, St Louis, MD, USA) and 1 mM sodium pyruvate (P8574, Sigma-Aldrich). Medium was changed twice a week. After 8 weeks, almost complete elimination of mtDNA was confirmed with PicoGreen staining (P7581, Thermo Fisher, MA, USA)61 and real time PCR62. Supplementary Fig. S4 shows the absence of mtDNA (A) in PicoGreen staining and an extremely lower amount of mtDNA detected by PCR (B) in ethidium bromide-treated cells. The cells were then cultured for another 2 weeks without ethidium bromide, and the absence of mtDNA was again confirmed with PicoGreen staining and PCR. Thereafter these TPC1 ρ0 cells were kept in RPMI 1640 supplemented with 5% FBS, antibiotics 50 μg/mL uridine and 1 mM sodium pyruvate.
To quantify mtDNA by PCR, total DNA was isolated from cells using a DNeasy Blood & Tissue Kit (69504, Qiagen, Valencia, CA) according to the manufacturer’s instructions, and then used for PCR amplification of 12S ribosomal DNA (12S rDNA) encoded by mtDNA and and β-ACTIN encoded by nuclear DNA (nDNA) with TB-Green Premix Ex Taq II (RR820A, Takara Bio, Kusatsu, Japan) and a Thermal Cycler Dice Real-Time System (Takara Bio). Then mtDNA/nDNA ratio was used to estimate the relative mtDNA copy number, i.e., the relative amounts of mitochondria. The primers used were described previously11; 12Sr DNA primers are 5ʹ-TAACCCAAGTCAATAGAAGCC-3ʹ (forward) and 5ʹ-CTAGAGGGATATGAAGCACC-3ʹ (reverse), and human β-ACTIN primers are 5ʹ-GAGCGGGAAATCGTGCGTGAC-3ʹ (forward) and 5ʹ-GGAAGGAAGGCTGGAAGAGTG-3ʹ (reverse). The PCR conditions were 40 cycles of denaturation at 95 °C for 30 s, annealing at 55 °C for 30 s and extension at 72 °C for 30 s. The correct sizes of PCR products were also confirmed with 2% agarose gel electrophoresis (data not shown).
Mitochondrial stress test
The mitochondrial oxygen consumption rate (OCR) was measured using a Seahorse XF-96 extracellular flux analyzer (Seahorse Bioscience, North Billerica, MA, USA). The cells were seeded in an XF-96 plate (0.8 × 104 cells/well) and incubated in regular culture medium with 5% FBS for 24 h. Meanwhile, the sensor cartridge was calibrated with calibration buffer supplied by the manufacturer at 37 °C without CO2. For measurement of OCR, the culture medium was replaced with the XF Base Medium (Seahorse Bioscience) containing 1 mM sodium pyruvate, 10 mM glucose and 2 mM glutamine (all from Seahorse Biosciences). After 1 h culture without CO2, readings were taken at the base line and after the addition of various chemicals such as 2 μM oligomycin, 0.25 μM phenylhydrazone or 0.5 μM rotenone/antimycin (all from Seahorse Bioscience). OCR was automatically calculated and recorded using the sensor cartridge and Seahorse XF-96 software. The plates were saved, and the cell number was determined with a Cell Counting Kit-8 (Dojindo, Kumamoto, Japan) according to the manufacturer’s protocol.
Cell number measurements
The cells in a 96 well plate (3 × 103 cells/well) were treated with atovaquone (5–20 μM; A2545, Tokyo Chemical Industry Co., Ltd., Tokyo, Japan)) or oxamate (10–60 mM; O2751, Sigma-Aldrich) or cultured in glucose- or glutamine-free medium for up to 48 h. The viable cell numbers were then determined with the water-soluble tetrazolium salt (WST) assay using a Cell Counting Kit-8. The plates were read with a 2030 Multilabel Plate Reader ARVO X3 (Perkin Elmer, Branchburg, NJ) at 450 nm.
Glucose uptake measurements
The cells in a 96 well plate (2.5 × 102 TPC1 and 2 × 103 XTC.UC1 cells/well) were incubated in regular culture medium. Five days later, the cell numbers for each cell line was confirmed to be approximately the same. The cells were washed twice with PBS containing Ca+ and Mg+ (PBS(+)), preincubated with KRPH buffer containing 2% BSA for 40 min, and then cultured with 1 mM 2-deoxyglucose for 20 min. After lysing the cells, intracellular glucose contents were measured with a Glucose Uptake Colorimetric Assay Kit (K676-100, BioVision, Inc., Waltham, MA) according to the manufacturer’s protocol. The plates were read with a 2030 Multilabel Plate Reader ARVO X3 (PerkinElmer) at 412 nm.
Lactate measurements
The cells in a 96 well plate (5 × 103 cells/well) were incubated with or without 10 and 50 mM oxamate for 48 h. The culture media were then collected and diluted with H2O accordingly. Lactate concentrations in the media were measured using by a Lactate Assay Kit-WST (L256, Dojindo) according to the manufacturer's instruction. The plate was read with a 2030 Multilabel Plate Reader ARVO X3 (PerkinElmer) at 450 nm.
GSH measurements
The cells in a 96 well plate (2.5 × 103 cells/well) were incubated in regular culture medium or glutamine-free medium with or without 4 mM N-acetyl cysteine (NAC; A7250, Sigma–Aldrich) for 48 h. GSH levels were then measured with a GSH-Glo™ Glutathione Assay Kit (V6911, Promega, Madison, WI) according to the manufacturer's protocol. The plates were read with a 2030 Multilabel Plate Reader ARVO X3 (PerkinElmer) to detect luminescence.
ROS measurements
The cells in a 96-well plate (5 × 104 cells/well) were washed with PBS ( +), incubated with 5 μM 2’,7’-dichlorofluorescin diacetate (C6827, Thermo Fisher Scientific, Waltham, MA) in PBS ( +) for 0.5 h at 37 °C and washed again with PBS ( +). The plates were read with a 2030 Multilabel Plate Reader ARVO X3 (PerkinElmer) at excitation and emission wavelengths of 485 and 528 nm, respectively. The results were expressed as relative values vs. the control.
Intracellular L-amino acid measurements
XTC.UC1 cells in a 10 cm dish were cultured with or without 1–30 μM chloroquine (08660-04, Nacalai Tesque) for 48 h. Then 2 × 106 cells were suspended in 500 μL of L-amino acid assay buffer (K639-100-1, Bio Vision Inc.), completely lysed with a homogenizer and centrifuged. The supernatants obtained were used for amino acid quantitation using an L-Amino Acid Quantitation Kit (BioVision Inc.) according to the manufacturer's instructions. The plates were read with a 2030 Multilabel Plate Reader ARVO X3 (PerkinElmer) at excitation and emission wavelengths of 535 and 587 nm, respectively.
Western blotting
Expression of LC3, p62 and β-actin were determined by immunoblotting with 40 μg cell lysates as described previously63. After sodium dodecyl sulfate polyacrylamide gel electrophoresis and transfer onto a polyvinylidene fluoride membrane (Bio-Rad Laboratories Inc., Tokyo, Japan), the membranes were blotted with the following antibodies and chemiluminescence system (Thermo Fisher Scientific, Rockford, IL). The primary and secondary antibodies used were (i) polyclonal rabbit anti-LC3 (PM036, MBL International, Woburn, MA; 1:1000) and polyclonal goat anti-rabbit horseradish peroxidase (HRP)-conjugated IgG (7074, Cell Signaling Technology, Danvers, MA; 1:1000) for LC3; (ii) polyclonal guinea pig anti-p62 (GP62-C, Progen, Heidelberg, Germany; 1:1000) and polyclonal rabbit anti-guinea pig HRP-conjugated IgG (61-4620, Innovative Research, Novi, MI; 1:1000) for p62; and (iii) monoclonal mouse anti β-actin (C4, Santa Cruz Biotechnology, Dallas, TX; 1:1000) and polyclonal horse anti-mouse HRP-conjugated IgG (7076, Cell Signaling Technology; 1:1000) for β-actin. Protein detection was performed using LAS-3000 (FUJIFILM) and quantified using NIH ImageJ software. A molecular weight Novex® Sharp pre-stained Protein Standard (LC5800, Thermo Fisher Scientific) was run in parallel.
Quantitative real-time RT-PCR
Total RNA was extracted from cells using ISOGEN reagent (311-02501, Nippon Gene, Tokyo, Japan). cDNA was synthesized from 500 ng total RNA with SuperScript III reverse transcriptase (18080-400, Thermo Fisher Scientific) using random hexamers. Quantitative PCR was then carried out using a Thermal Cycler Dice Real-Time System (Takara Bio) using TB Green Premix EX Taq II (RR820A, Takara Bio). The primers used were 5′-ACCATGGTTCTGGTCTCCTG-3′ and 5′-GGGCAAAGAGTAAACCCACA-3′ for ASCT2, and 5′-GGCCTTCATCGCAGTACATC-3′ and 5′-GCCTTCACGCTGTAGCAGTT-3′ for LAT1. The PCR conditions were 40 cycles of denaturation at 95 °C for 15 s, annealing at 55 °C for 15 s, and extension at 72 °C for 20 s. The cycle threshold values, which were determined using second derivative, were used to calculate the normalized expression of the indicated mRNAs using Q-Gene soft-ware using β-actin for normalization. The negative control reaction was done without reverse transcription in parallel. The correct sizes of PCR products were confirmed with 2% agarose gel electrophoresis (data not shown).
Statistical analysis
All data are expressed as means ± SE (n = 3) or ranges (n = 2) and differences between groups were examined for statistical significance using Scheffe's multiple comparison test. A p-value < 0.05 was considered statistically significant. All the experiments were repeated at least twice with essentially the same results.
Data availability
All data generated or analyzed during this study are included in this published article.
References
McFadden, D. G. & Sadow, P. M. Genetics, diagnosis, and management of Hürthle cell thyroid neoplasms. Front. Endocrinol. 12, 696386. https://doi.org/10.3389/fendo.2021.696386 (2021).
Asa, S. L. & Mete, O. Oncocytic change in thyroid pathology. Front. Endocrinol. 12, 678119. https://doi.org/10.3389/fendo.2021.678119 (2021).
Corver, W. E. et al. Near-haploidization significantly associates with oncocytic adrenocortical, thyroid, and parathyroid tumors but not with mitochondrial DNA mutations. Genes Chromosomes Cancer 53, 833–844. https://doi.org/10.1002/gcc.22194 (2014).
Gopal, R. K. et al. Widespread chromosomal losses and mitochondrial DNA alterations as genetic drivers in Hurthle cell carcinoma. Cancer Cell 34, 242-255.e245. https://doi.org/10.1016/j.ccell.2018.06.013 (2018).
Ganly, I. et al. Integrated genomic analysis of Hurthle cell cancer reveals oncogenic drivers, recurrent mitochondrial mutations, and unique chromosomal landscapes. Cancer Cell 34, 256-270.e255. https://doi.org/10.1016/j.ccell.2018.07.002 (2018).
Gasparre, G. et al. Disruptive mitochondrial DNA mutations in complex I subunits are markers of oncocytic phenotype in thyroid tumors. Proc. Natl. Acad. Sci. USA 104, 9001–9006. https://doi.org/10.1073/pnas.0703056104 (2007).
Larman, T. C. et al. Spectrum of somatic mitochondrial mutations in five cancers. Proc. Natl. Acad. Sci. USA 109, 14087–14091. https://doi.org/10.1073/pnas.1211502109 (2012).
Savagner, F. et al. Mitochondrial activity in XTC.UC1 cells derived from thyroid oncocytoma. Thyroid 11, 327–333. https://doi.org/10.1089/10507250152039055 (2001).
Savagner, F. et al. PGC-1-related coactivator and targets are upregulated in thyroid oncocytoma. Biochem. Biophys. Res. Commun. 310, 779–784. https://doi.org/10.1016/j.bbrc.2003.09.076 (2003).
Baris, O. et al. Transcriptional profiling reveals coordinated up-regulation of oxidative metabolism genes in thyroid oncocytic tumors. J. Clin. Endocrinol. Metab. 89, 994–1005. https://doi.org/10.1210/jc.2003-031238 (2004).
Mussazhanova, Z. et al. Causative role for defective expression of mitochondria-eating protein in accumulation of mitochondria in thyroid oncocytic cell tumors. Cancer Sci. 111, 2814–2823. https://doi.org/10.1111/cas.14501 (2020).
Nakamura, Y. & Arakawa, H. Discovery of Mieap-regulated mitochondrial quality control as a new function of tumor suppressor p53. Cancer Sci. 108, 809–817. https://doi.org/10.1111/cas.13208 (2017).
Vander Heiden, M. G. & DeBerardinis, R. J. Understanding the intersections between metabolism and cancer biology. Cell 168, 657–669. https://doi.org/10.1016/j.cell.2016.12.039 (2017).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: The next generation. Cell 144, 646–674. https://doi.org/10.1016/j.cell.2011.02.013 (2011).
Bonora, E. et al. Defective oxidative phosphorylation in thyroid oncocytic carcinoma is associated with pathogenic mitochondrial DNA mutations affecting complexes I and III. Cancer Res. 66, 6087–6096. https://doi.org/10.1158/0008-5472.can-06-0171 (2006).
Porcelli, A. M. et al. The antioxidant function of Bcl-2 preserves cytoskeletal stability of cells with defective respiratory complex I. Cell. Mol. Life Sci. 65, 2943–2951. https://doi.org/10.1007/s00018-008-8300-2 (2008).
Lee, J. et al. A metabolic phenotype based on mitochondrial ribosomal protein expression as a predictor of lymph node metastasis in papillary thyroid carcinoma. Medicine 94, e380. https://doi.org/10.1097/md.0000000000000380 (2015).
Lee, J. et al. Dysregulation of Parkin-mediated mitophagy in thyroid Hurthle cell tumors. Carcinogenesis 36, 1407–1418. https://doi.org/10.1093/carcin/bgv122 (2015).
Kim, H. M. & Koo, J. S. Differential expression of glycolysis-related proteins in follicular neoplasms versus Hürthle cell neoplasms: A retrospective analysis. Dis. Mark. 2017, 6230294. https://doi.org/10.1155/2017/6230294 (2017).
Kürschner, G. et al. Renal oncocytoma characterized by the defective complex I of the respiratory chain boosts the synthesis of the ROS scavenger glutathione. Oncotarget 8, 105882–105904. https://doi.org/10.18632/oncotarget.22413 (2017).
Feng, J. et al. Enhancement of mitochondrial biogenesis and paradoxical inhibition of lactate dehydrogenase mediated by 14–3-3η in oncocytomas. J. Pathol. 245, 361–372. https://doi.org/10.1002/path.5090 (2018).
Calabrese, C. et al. Respiratory complex I is essential to induce a Warburg profile in mitochondria-defective tumor cells. Cancer Metab. 1, 11. https://doi.org/10.1186/2049-3002-1-11 (2013).
Addie, R. D. et al. Metabolic reprogramming related to whole-chromosome instability in models for Hürthle cell carcinoma. Sci. Rep. 10, 9578. https://doi.org/10.1038/s41598-020-66599-1 (2020).
Owen, O. E., Kalhan, S. C. & Hanson, R. W. The key role of anaplerosis and cataplerosis for citric acid cycle function. J. Biol. Chem. 277, 30409–30412. https://doi.org/10.1074/jbc.R200006200 (2002).
Seyfried, T. N., Arismendi-Morillo, G., Mukherjee, P. & Chinopoulos, C. On the Origin of ATP Synthesis in Cancer. iScience 23, 101761. https://doi.org/10.1016/j.isci.2020.101761 (2020).
Mattaini, K. R., Sullivan, M. R. & Vander Heiden, M. G. The importance of serine metabolism in cancer. J. Cell Biol. 214, 249–257. https://doi.org/10.1083/jcb.201604085 (2016).
Ducker, G. S. & Rabinowitz, J. D. One-carbon metabolism in health and disease. Cell Metab. 25, 27–42. https://doi.org/10.1016/j.cmet.2016.08.009 (2017).
Shiratori, R. et al. Glycolytic suppression dramatically changes the intracellular metabolic profile of multiple cancer cell lines in a mitochondrial metabolism-dependent manner. Sci. Rep. 9, 18699. https://doi.org/10.1038/s41598-019-55296-3 (2019).
Saito, Y. & Soga, T. Amino acid transporters as emerging therapeutic targets in cancer. Cancer Sci. 112, 2958–2965. https://doi.org/10.1111/cas.15006 (2021).
Utsunomiya-Tate, N., Endou, H. & Kanai, Y. Cloning and functional characterization of a system ASC-like Na+-dependent neutral amino acid transporter. J. Biol. Chem. 271, 14883–14890. https://doi.org/10.1074/jbc.271.25.14883 (1996).
Abla, H., Sollazzo, M., Gasparre, G., Iommarini, L. & Porcelli, A. M. The multifaceted contribution of α-ketoglutarate to tumor progression: An opportunity to exploit?. Semin. Cell Dev. Biol. 98, 26–33. https://doi.org/10.1016/j.semcdb.2019.05.031 (2020).
Uddin, S. et al. Fatty acid synthase and AKT pathway signaling in a subset of papillary thyroid cancers. J. Clin. Endocrinol. Metab. 93, 4088–4097. https://doi.org/10.1210/jc.2008-0503 (2008).
von Roemeling, C. A. et al. Aberrant lipid metabolism in anaplastic thyroid carcinoma reveals stearoyl CoA desaturase 1 as a novel therapeutic target. J. Clin. Endocrinol. Metab. 100, E697-709. https://doi.org/10.1210/jc.2014-2764 (2015).
Sun, W. Y., Kim, H. M., Jung, W. H. & Koo, J. S. Expression of serine/glycine metabolism-related proteins is different according to the thyroid cancer subtype. J. Transl. Med. 14, 168. https://doi.org/10.1186/s12967-016-0915-8 (2016).
Smith, A. L. et al. Age-associated mitochondrial DNA mutations cause metabolic remodelling that contributes to accelerated intestinal tumorigenesis. Nat. Cancer 1, 976–989. https://doi.org/10.1038/s43018-020-00112-5 (2020).
Moreadith, R. W. & Lehninger, A. L. The pathways of glutamate and glutamine oxidation by tumor cell mitochondria. Role of mitochondrial NAD(P)+-dependent malic enzyme. J. Biol. Chem. 259, 6215–6221 (1984).
DeBerardinis, R. J. et al. Beyond aerobic glycolysis: transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. Proc. Natl. Acad. Sci. USA 104, 19345–19350. https://doi.org/10.1073/pnas.0709747104 (2007).
Scott, D. A. et al. Comparative metabolic flux profiling of melanoma cell lines: Beyond the Warburg effect. J. Biol. Chem. 286, 42626–42634. https://doi.org/10.1074/jbc.M111.282046 (2011).
Chen, Q. et al. Rewiring of glutamine metabolism is a bioenergetic adaptation of human cells with mitochondrial DNA mutations. Cell Metab. 27, 1007-1025.e1005. https://doi.org/10.1016/j.cmet.2018.03.002 (2018).
Mullen, A. R. et al. Reductive carboxylation supports growth in tumour cells with defective mitochondria. Nature 481, 385–388. https://doi.org/10.1038/nature10642 (2012).
Fendt, S. M. et al. Metformin decreases glucose oxidation and increases the dependency of prostate cancer cells on reductive glutamine metabolism. Cancer Res. 73, 4429–4438. https://doi.org/10.1158/0008-5472.Can-13-0080 (2013).
Kodama, M. et al. A shift in glutamine nitrogen metabolism contributes to the malignant progression of cancer. Nat. Commun. 11, 1320. https://doi.org/10.1038/s41467-020-15136-9 (2020).
Priolo, C. et al. Impairment of gamma-glutamyl transferase 1 activity in the metabolic pathogenesis of chromophobe renal cell carcinoma. Proc. Nat. Acad. Sci. USA 115, E6274-e6282. https://doi.org/10.1073/pnas.1710849115 (2018).
Gopal, R. K. et al. Early loss of mitochondrial complex I and rewiring of glutathione metabolism in renal oncocytoma. Proc. Nat. Acad. Sci. USA 115, E6283-e6290. https://doi.org/10.1073/pnas.1711888115 (2018).
Jin, L. et al. Glutamate dehydrogenase 1 signals through antioxidant glutathione peroxidase 1 to regulate redox homeostasis and tumor growth. Cancer Cell 27, 257–270. https://doi.org/10.1016/j.ccell.2014.12.006 (2015).
Hirayama, A. et al. Quantitative metabolome profiling of colon and stomach cancer microenvironment by capillary electrophoresis time-of-flight mass spectrometry. Cancer Res. 69, 4918–4925. https://doi.org/10.1158/0008-5472.Can-08-4806 (2009).
Iommarini, L., Calvaruso, M. A., Kurelac, I., Gasparre, G. & Porcelli, A. M. Complex I impairment in mitochondrial diseases and cancer: parallel roads leading to different outcomes. Int. J. Biochem. Cell Biol. 45, 47–63. https://doi.org/10.1016/j.biocel.2012.05.016 (2013).
Shen, L. et al. Upregulation of the solute carrier family 7 genes is indicative of poor prognosis in papillary thyroid carcinoma. World J. Surg. Oncol. 16, 235. https://doi.org/10.1186/s12957-018-1535-y (2018).
Kim, H. M., Lee, Y. K. & Koo, J. S. Expression of glutamine metabolism-related proteins in thyroid cancer. Oncotarget 7, 53628–53641. https://doi.org/10.18632/oncotarget.10682 (2016).
Häfliger, P. et al. The LAT1 inhibitor JPH203 reduces growth of thyroid carcinoma in a fully immunocompetent mouse model. J. Exp. Clin. Cancer Res. 37, 234. https://doi.org/10.1186/s13046-018-0907-z (2018).
Enomoto, K. et al. A novel therapeutic approach for anaplastic thyroid cancer through inhibition of LAT1. Sci. Rep. 9, 14616. https://doi.org/10.1038/s41598-019-51144-6 (2019).
Corver, W. E. et al. Genome haploidisation with chromosome 7 retention in oncocytic follicular thyroid carcinoma. PloS One 7, e38287. https://doi.org/10.1371/journal.pone.0038287 (2012).
Schweppe, R. E. et al. Deoxyribonucleic acid profiling analysis of 40 human thyroid cancer cell lines reveals cross-contamination resulting in cell line redundancy and misidentification. J. Clin. Endocrinol. Metab. 93, 4331–4341. https://doi.org/10.1210/jc.2008-1102 (2008).
Zielke, A. et al. Establishment of a highly differentiated thyroid cancer cell line of Hurthle cell origin. Thyroid 8, 475–483. https://doi.org/10.1089/thy.1998.8.475 (1998).
Ishizaka, Y., Ushijima, T., Sugimura, T. & Nagao, M. cDNA cloning and characterization of ret activated in a human papillary thyroid carcinoma cell line. Biochem. Biophys. Res. Commun. 168, 402–408. https://doi.org/10.1016/0006-291x(90)92335-w (1990).
Lemoine, N. R. et al. Characterisation of human thyroid epithelial cells immortalised in vitro by simian virus 40 DNA transfection. Br. J. Cancer 60, 897–903. https://doi.org/10.1038/bjc.1989.387 (1989).
Aydemirli, M. D. et al. Targeted treatment options of recurrent radioactive iodine refractory Hürthle cell cancer. Cancers 11, 1. https://doi.org/10.3390/cancers11081185 (2019).
Shimamura, M., Nagayama, Y., Matsuse, M., Yamashita, S. & Mitsutake, N. Analysis of multiple markers for cancer stem-like cells in human thyroid carcinoma cell lines. Endocr. J. 61, 481–490 (2014).
Makinoshima, H. et al. Epidermal growth factor receptor (EGFR) signaling regulates global metabolic pathways in EGFR-mutated lung adenocarcinoma. J. Biol. Chem. 289, 20813–20823. https://doi.org/10.1074/jbc.M114.575464 (2014).
Yoshida, K. et al. Metabolome analysis reveals a diversity of cancer tissues in advanced epithelial ovarian cancer. Cancer Cell. Int. 21, 314. https://doi.org/10.1186/s12935-021-02014-7 (2021).
Ashley, N., Harris, D. & Poulton, J. Detection of mitochondrial DNA depletion in living human cells using PicoGreen staining. Exp. Cell Res. 303, 432–446. https://doi.org/10.1016/j.yexcr.2004.10.013 (2005).
Khozhukhar, N., Spadafora, D., Rodriguez, Y. & Alexeyev, M. Elimination of mitochondrial DNA from mammalian cells. Curr. Protoc. Cell Biol. 78, 20.11.21–20.11.14. https://doi.org/10.1002/cpcb.39 (2018).
Kurashige, T. et al. Basal autophagy deficiency causes thyroid follicular epithelial cell death in mice. Endocrinology 160, 2085–2092. https://doi.org/10.1210/en.2019-00312 (2019).
Acknowledgements
This work was supported in part by a Grants-in-Aid for Scientific Research (19K090282 to YN) from the Ministry of Education, Culture, Sports, Science and Technology, Tokyo, Japan.
Author information
Authors and Affiliations
Contributions
Y.N. and N.M. conceived the experiments; T.K., M.S., K.H., and M.M. conducted the experiments; Y.N. and T.K. analyzed the results; and Y.N. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kurashige, T., Shimamura, M., Hamada, K. et al. Characterization of metabolic reprogramming by metabolomics in the oncocytic thyroid cancer cell line XTC.UC1. Sci Rep 13, 149 (2023). https://doi.org/10.1038/s41598-023-27461-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-27461-2
- Springer Nature Limited
This article is cited by
-
CYB5R3 functions as a tumor suppressor by inducing ER stress-mediated apoptosis in lung cancer cells via the PERK-ATF4 and IRE1α-JNK pathways
Experimental & Molecular Medicine (2024)