Abstract
The most prevalent primary glomerulonephritis and leading cause of end-stage renal disease worldwide is IgA nephropathy (IgAN). More and more studies are describing urinary microRNA (miRNA) as a non-invasive marker for a variety of renal diseases. We screened candidate miRNAs based on data from three published IgAN urinary sediment miRNAs chips. In separate confirmation and validation cohorts, we included 174 IgAN patients, 100 patients with other nephropathies as disease controls (DC), and 97 normal controls (NC) for quantitative real-time PCR. A total of three candidate miRNAs, miR-16-5p, Let-7g-5p, miR-15a-5p were obtained. In both the confirmation and validation cohorts, these miRNAs levels were considerably higher in the IgAN than in NC, with miR-16-5p significantly higher than in DC. The area under the ROC curve for urinary miR-16-5p levels was 0.73. Correlation analysis suggested that miR-16-5p was positively correlated with endocapillary hypercellularity (r = 0.164 p = 0.031). When miR-16-5p was combined with eGFR, proteinuria and C4, the AUC value for predicting endocapillary hypercellularity was 0.726. By following the renal function of patients with IgAN, the levels of miR-16-5p were noticeably higher in the IgAN progressors than in the non- progressors (p = 0.036). Urinary sediment miR-16-5p can be used as noninvasive biomarkers for the assessment of endocapillary hypercellularity and diagnosis of IgA nephropathy. Furthermore, urinary miR-16-5p may be predictors of renal progression.
Similar content being viewed by others
Introduction
IgA nephropathy (IgAN), the most prevalent primary glomerulonephritis world-wide, causes end-stage renal disease (ESRD) in 30–40% of patients with biopsy-proven IgAN, making it the major reason of ESRD in many areas around the world. In addition, this disease is particularly common in Asian populations, accounting for 40–50% of primary glomerulonephritis1. Renal biopsy is the primary technique for confirming the diagnosis of IgAN, which is an invasive test that is not easily repeated and cannot be performed in many hospitals. Therefore, it is important to find non-invasive biomarkers that can diagnose IgAN.
Endogenous non-coding RNAs known as microRNAs (miRNAs) have a length of 19–25 nucleotides and are significant regulators of gene expression. They achieve this by promoting the breakdown of mRNA and/or blocking its translation. MiRNAs are readily available, relatively stable in form, and not easily degraded2. Urine sediment has the advantages of being non-invasive, painless, and can be repeatedly retained, and has a variety of exfoliated cells, including podocytes, renal tubular epithelial cells, immune cells, and stem cells in urine, which can be used to explore cellular changes in the damaged kidney. As a result, discovering non-invasive indicators of IgAN using urine sediment miRNAs may be a novel method3.
At this stage, most studies on biomarkers related to miRNAs in IgAN have used miRNA microarray for screening, and the obtained target miRNAs was scattered and lack systematic review. Most studies included only IgAN patients and normal controls. The purpose of our present research was to review the results of previous related studies to obtain target miRNAs. Other nephropathies that are not IgAN were also included. To research the correlation between miRNAs and the pathological and clinical indicators of IgAN, as well as any potential mechanisms involved.
Methods
Overall study design
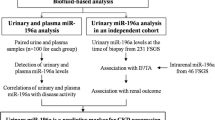
In this research, we reviewed a total of three profiles of urinary sediment miRNAs in the literature on IgAN by Zhi-Yu Duan, Cheuk-Chun Szeto, and Nannan Wang4,5,6, and found that miR-16-5p, Let-7g-5p, and miR-15a-5p were expressed in the same trend in the three miRNAs profiles, all of them were significantly higher in IgAN compared with normal controls, thus identifying the target miRNAs for this study, as shown in Fig. 1. This study included two separate patient cohorts. First, in the development cohort, we included 30 patients with biopsy-proven IgAN from October 2018 to February 2019 and 30 NC as controls for small sample validation. In the validation cohort, a total of 144 patients with IgAN, 100 DC and 67 NC from March 2019 to May 2022 were included for further extended validation.
Enrollment of subjects
Patients who had their initial kidney biopsies at the Department of Nephrology, First Medical Center of Chinese PLA General Hospital from October 2018 to May 2022 were consecutively recruited for this study. A total of 174 cases of IgAN, 63 cases of membranous nephropathy (MN), 8 cases of minimal change disease (MCD), 16 cases of focal segmental glomerular sclerosis (FSGS), and 13 cases of diabetic nephropathy (DN) were included according to the diagnosis confirmed by renal biopsy. We excluded 51 patients, 17 patients because of fewer than 8 glomeruli in light microscopic, 2 patients with ESRD prior to renal biopsy, and we also excluded patients with concurrent diagnosis of two or more renal diseases (n = 4), secondary IgA nephropathy (n = 13) (e.g. hepatitis-associated glomerulonephritis, systemic lupus erythematosus, allergic purpura, psoriasis, dry syndrome), and infections of the gastrointestinal, respiratory, or urinary tracts (n = 15). The normal controls were medical workers and routine physical examination patients from the First Medical Center of Chinese PLA General Hospital who had normal kidney function, normal routine urine results, and no personal or family history of kidney disease or other serious illnesses. At the time of the kidney biopsy, all patients had their demographic and clinical information, including their age, sex, mean arterial pressure (MAP), serum creatinine (Scr), and 24-hours urine protein excretion (UPE) recorded. A formula developed by the Chronic Kidney Disease Epidemiology Collaboration (CKD-EPI) was used to calculate glomerular filtration rate (eGFR)7.
Assessing renal histopathology damage
Two qualified pathologists graded the pathological sections of kidney biopsies from IgAN patients using the Oxford classification scoring system8. (1) Mesangial hypercellularity (M0/1): 50% of mesangial area with more than three cells is M1; (2) Endocapillary hypercellularity (E0/1): absent or present; (3) Segmental glomerulosclerosis (S0/1): absent or present; (4) Tubular atrophy or interstitial fibrosis (T0/1/2): <25% is T0, >25% to 50% is T1, >50% is T2; (5) Cellular or fibrocellular crescent (C0/1/2): C0 for no crescent, C1 for <25%, and C2 for >25%.
Sample preparation
Collect 50–100 ml of urine sample from the patient early in the morning on the day of kidney biopsy, place it in a 4 °C refrigerator for temporary storage after collection, and process it within 6 hours. Before use, the supernatant and urine sediment were divided into 50 ml and 1.5 ml EP tubes, respectively, and kept in a −80 °C refrigerator.
RNA extraction and real-time quantitative PCR
Following the manufacturer's instructions, total RNA from urine sediment was extracted using TRIzol (Invitrogen, USA). Using NanoDrop Onec spectrophotometer (Thermo Fisher Scientific, USA), the acquired RNA's concentration (ng/ul) and purity (A260/280) were evaluated. 500ng RNA was used as the system, using miRcute Plus miRNA First-Strand cDNA Kit (Tiangen Biotech, Beijing, China) and Applied Biosystems VeritiTM 96-well Thermal Cycler (Applied Biosystems, USA) for reverse transcription. The housekeeping gene U6 was employed for miRNA detection9, and all primers were purchased from Tiangen Biotech. Urine sediment hsa-U6, hsa-miR-16-5p, hsa-Let-7g-5p, and hsa-miR-15a-5p were quantified using miRcute Plus miRNA qPCR Kit (SYBR Green) (Tiangen Biotech, Beijing, China) and Bio-Rad cfx96 real-time system (CFX96, Bio-Rad, USA). The relative quantitative ΔΔCt technique was used to determine the variations in each target miRNA's expression levels in the samples.
Clinical follow up
After renal biopsy, the patient's renal function is followed up for at least 12 months. According to the KDIGO guidelines, progressor IgAN patients are those with a yearly drop in eGFR more than 5 ml/min/1.73 m2, and vice versa for non- progressor10.
Statistical analysis
For graphing and plotting, both SPSS 26.0 and GraphPad Prism 9.4.1 for MAC were used. According to the situation, data were presented as mean ± standard deviation or median (inter-quartile range). The 2(−ΔΔCt) relative quantification method was used to compare the expression of miRNAs. Shapiro-Wilk tests were performed on normality analysis. For normally distributed data, the t test or one-way ANOVA were employed. For data which weren't normally distributed, the Mann-Whitney or Kruskal-Wallis rank test was applied. Correlation analysis was performed using the Pearson or Spearman methods to explore the association between miRNAs and clinical and pathological indicators. The sensitivity and specificity of miRNAs for the diagnosis of IgA nephropathy were calculated utilizing receiver operating characteristic (ROC) curves and area under the ROC curves (AUC). P < 0.05 was used to define statistical significance for the results. All computations were performed with two-tailed probabilities.
Ethical approval
The studies involving human participants were reviewed and approved by Research Ethics Committee of Chinese PLA General Hospital (approval number S2018-206-01). The patients/participants provided their written informed consent to participate in this study.
Results
A review of diagnostic markers of miRNAs in urinary sediment of IgA nephropathy
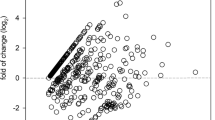
In this study, we reviewed and compared a total of three profiles of miRNAs in the urinary sediment of IgAN by Zhi-Yu Duan and Nannan Wang from the research team of the Department of Nephrology, First Medical Center of Chinese PLA General Hospital, and Cheuk-Chun Szeto from the research team of the Chinese University of Hong Kong, as shown in Table 1. The expression trends of miR-16-5p, Let-7g-5p, and miR-15a-5p were found to be consistent in the three expression profiles, and all were noticeably higher in IgAN than in normal controls with at least a 10 fold-change, according to Table 2.
Clinical and pathological features of patients with IgA nephropathy
We included a total of 174 patients with IgAN, 100 disease control patients and 97 healthy individuals. The clinical and demographic details of the study participants are summarized in Table 3. The pathological features of the IgAN group are listed in Table 4.
Urinary miR-16-5p level in the cohorts
First, in the confirmation cohort, we validated three miRNAs in 30 patients with IgAN and 30 normal patients. Patients with IgAN had significantly greater levels of miR-16-5p expression in urine sediment when compared to the normal controls, P < 0.0001, with a statistically significant difference (Fig. 2A). To further clarify whether these urine sediment markers are specific for the diagnosis of IgAN, we included 100 disease controls including MN, FSGS, and DN in the validation cohort. Urine sediment miR-16-5p differed significantly from both disease controls and normal controls (P = 0.0201,P < 0.0001) , according to Fig. 2B. The expressions of Let-7g-5p and miR-15a-5p in the two cohorts were shown in Supplementary Fig. 1.
(A) Comparison of miR-16-5p expression levels between the IgAN group and normal group in a confirmation cohort. IgAN group, n = 30, Normal control, n = 30. (B) Comparison of miR-16-5p expression levels between the IgAN group and Disease control group and normal control group in a validation cohort. IgAN group, n = 144, Disease control, n = 100, Normal control, n = 67. *, P < 0.05; ****, P < 0.0001.
Diagnostic value of miR-16-5p
ROC curves were used to examine the effectiveness of urine miRNA levels in the diagnosis of IgA nephropathy. The area under the curve (AUC) of miR-16-5p, Let-7g-5p, and miR-15a-5p ROC curves were all statistically significant, miR-16-5p had the best diagnostic value (AUC = 0.73, 95% CI 0.679–0.781, P < 0.0001) (Table. 5). When the optimal cut-off value of miR-16-5p was 0.324, the positive predictive value (PPV) was 63.3% and the negative predictive value (NPV) was 68.4%.
Correlation between miR-16-5p and endocapillary hypercellularity
Through correlation analysis, we found that miR-16-5p had no significant correlation with clinical indicators such as eGFR (r = -0.144 p = 0.058) and proteinuria (r = 0.018 p = 0.811), while in terms of pathological indicators, miR-16-5p in urine sediment was positively correlated with endocapillary hypercellularity (r = 0.164 p = 0.031). Patients with E1 classification of IgAN had considerably greater levels of miR-16-5p in urine sediment than E0 classification[1.013(0.335-5.676) vs 0.512(0.224–1.918)] (Fig. 3A). In addition, endocapillary hypercellularity was significantly negatively correlated with blood complement C4 (r = -0.188 p = 0.013), the blood complement C4 of IgAN with E1 was significantly lower than that of E0 [23.85(18.98–27.55) vs 26.35(22.53–30.83)]. Finally, the complete correlation analysis data of the three miRNAs were shown in supplementary materials (Supplementary Table 1).
Prediction values of miR-16-5p in endocapillary hypercellularity
According to ROC analysis, the AUC values of miR-16-5p, eGFR, proteinuria and C4 for predicting IgAN E1 grade were 0.619, 0.600, 0.480 and 0.363, respectively. miR-16-5p had the highest value for predicting E1 grade alone. When miR-16-5p was combined with eGFR, proteinuria and C4, the AUC value for predicting endocapillary hypercellularity was 0.726, and the predictive value was improved from the previous one.
Correlation between miR-16-5p and renal progression
To confirm the efficacy of urine sediment miRNAs in predicting the development of renal function in IgAN, 122 IgAN patients were followed for a mean of 8.5 (4–18) months, with a total of 54 IgAN followed for >12 months, 14 of whom were progressors, including one patient who developed ESKD with eGFR ≤15 ml/min/1.73m2, and 40 patients in the non- progressors. The level of miR-16-5p was considerably greater in the progressor than in the non-progressor (P = 0.036) (Fig. 3B), suggesting that the higher the baseline miR-16-5p expression, the faster the decline in eGFR.
Discussion
In this study, we identified three miRNA targets that differed significantly in urine levels between IgA nephropathy and normal controls. Among them, urinary miR-16-5p is a potential marker of endocapillary hypercellularity in IgA nephropathy, which has a good diagnostic value and may be a predictor of renal function progression in IgA nephropathy.
Previous research has looked into the differences in miRNA levels in the urine of people with IgAN4,5,6. For example, Wang et al.6 reported that miR-3613-3p expression was significantly downregulated in the urine of IgAN patients and miR-3613-3p and miR-4668-5p correlated with disease severity. MiR-25-3p, miR-144-3p, and miR-486-5p in urinary sediment may be diagnostic markers of IgA nephropathy and are mainly derived from urinary red blood cells. Higher levels of miR-144-3p expression are linked to lower urine protein and better renal function4. Similarly, Liang et al.11reported urinary miR-21 and miR-205 as prospective prognostic markers for the assessment of interstitial tubular damage in IgAN, and urinary miR-204 was reported as a diagnostic tool for IgA nephropathy5. Therefore, we reviewed the urinary sediment miRNA profile published in previous studies on IgAN and found that miR-16-5p, Let-7g-5p, and miR-15a-5p expression trends were consistent in three studies4,5,6, all of which were noticeably increased in patients with IgAN in contrast to normal controls, with a fold change > 10, thus identifying the target miRNAs for this study. The PCR results of the confirmation cohort showed significant differences between IgAN and normal patients, totally supporting the trend of the microarray study discussed above. We then included disease controls for non-IgAN in an independent validation cohort, and the results suggested that urinary miR-16-5p may be a diagnostic biomarker of IgAN. Analysis of the ROC curve revealed good specificity and sensitivity.
In addition to the diagnostic value, we further investigated the clinical significance of these urinary sediment miRNAs. Proteinuria, eGFR and Oxford classification were all closely associated with the progression and prognosis of IgAN12,13,14. In our results, miR-16-5p levels were significantly higher in the urine sediment of patients with E1 in contrast to E0 patients, and blood complement C4 levels were lower in E1 compared with E0 patients, further confirming that endocapillary hypercellularity is a hallmark of active inflammation15 and that endothelial cell proliferation is primarily the result of inflammatory cell infiltration16. Bao et al.17used mesangial cells to stimulate endothelial cells, and their miRNA microarray results showed high expression of miR-16-5p. Therefore, we hypothesized that under galactose-deficient IgA1 stimulation, mesangial cells secreted substances that stimulated endothelial cells to overexpress miR-16-5p. Studies have demonstrated that in patients with IgAN who are not receiving immunosuppressive therapy, endocapillary hypercellularity is a reliable indicator of the rate of renal function loss18. Thus the lack of prognostic value for the E score in other studies is likely the result of immunosuppression-related treatment bias19,20. A number of subsequent studies have provided evidence for immunosuppressive treatment of intracapillary proliferative IgAN21,22. In addition, Let-7g-5p was positively correlated with baseline proteinuria and negatively correlated with eGFR. The higher the expression level of Let-7g-5p, the more its 24-hour urinary protein and the worse its renal function, suggesting that Let-7g-5p may be associated with prognosis. miR-15a-5p was not associated with proteinuria, eGFR, or pathological indicators. The follow-up of renal function in IgAN patients revealed significant differences in miR-16-5p expression levels between the progressor and non- progressor, all of which suggest that urinary sediment miRNAs may be related to the progression and prognosis of IgAN.
Although the biological functions of miR-16-5p, Let-7g-5p, and miR-15a-5p have not been clarified, bioinformatics analysis may elucidate the roles of these miRNAs by predicting hub genes. A total of 133 common target genes of miR-16-5p, Let-7g-5p, and miR-15a-5p were obtained from database miRDB and starBase. By Gene Ontology (GO) enrichment analysis23, the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis24,25 and the Protein-Protein Interaction (PPI) network construction26,27, the hub gene CCND1 may have a significant impact on how these miRNAs regulate the biological process of IgAN (Supplementary Figs. 2–4, unpublished). According to GO analysis, the functions of these miRNAs are mainly enriched in the regulation of catabolic, metabolic and renal water homeostasis. Antidiuretic hormone (AVP) is important for water homeostasis, and it has been shown that copeptin, a reliable and stable alternative to circulating AVP, was found to correlate with disease severity and prognosis in IgAN patients by detecting copeptin28. According to KEGG pathway enrichment analysis, these miRNAs were significantly enriched in FoxO signaling pathway, mTOR signaling pathway, and Wnt signaling pathway. The FoxO signaling pathway has been shown to play an important role in diseases associated with altered B-cell proliferation and/or activation29, while up- and down-regulation of some B cells was indeed found by studies of peripheral blood cell subsets in IgAN30. Activation of the mTOR signaling pathway causes activation of downstream signaling proteins such as p70s6K, leading to extracellular matrix deposition in glomerular mesangial cells, renal tubular and collecting duct epithelial cells, renal fibroblast activation, and renal interstitial fibrosis. Rapamycin, as an inhibitor of the mTOR signaling pathway, can effectively inhibit glomerular mesangial cell proliferation and extracellular matrix secretion by early low-dose application, reduce IgA deposition, and protect renal function31. It has also been demonstrated that excessive Wnt signaling pathway activation contributes to the development of IgAN32. In addition, A critical gene that controls the cell cycle is CCND1, and turning on its expression can encourage cell growth. One study found that aberrant miR-320 expression in IgAN urine and kidney tissue promoted B cell proliferation and increased the expression of CCND133. In a rat model of IgAN, autophagy is inhibited, while the mTOR/S6k1 pathway is activated and CCND1 expression is increased34, all of which provide directions for further studies.
Our study has some shortcomings. First, all samples were from a single center and were Chinese, whereas the prevalence of IgAN and disease characteristics vary considerably among different countries and ethnic groups, and further validation is still needed. Second, in this study, the basic demographic and clinical information (e.g., age, renal function, or proteinuria) of patients with IgAN was not exactly matched to the diseased controls and normal controls, and a strict pairwise analysis was not performed due to differences in the number of subjects. In addition, the cellular origin of miRNAs in urine sediment is not clear. miR-16-5p, Let-7g-5p, and miR-15a-5p were reported to be the top 20 miRNAs in human blood erythrocytes35, so we speculate that these miRNAs may be derived from erythrocytes, but further experiments are still needed to verify them in urine. Finally, how miR-16-5p, Let-7g-5p, and miR-15a-5p act on IgAN via CCND1 remains to be further explored.
In conclusion, in the first part of the study, we reviewed and validated a significant rise in miR-16-5p, Let-7g-5p, and miR-15a-5p in IgAN. Then, we found a significant correlation between miR-16-5p and endocapillary hypercellularity. Finally, we evaluated the effect of miR-16-5p on the progression of IgAN renal function and made some bioinformatic speculations. These findings support the idea that miR-16-5p is a potential marker reflecting endocapillary hypercellularity and predicting IgAN progression.
Data availability
The raw date supporting the conclusions of this article will be made available by the authors, without undue reservation. Correspondence and requests for materials should be addressed to G.-Y.C.
References
Lai, K. N. et al. IgA nephropathy. Nat. Rev. Dis. Prim. 2, 16001. https://doi.org/10.1038/nrdp.2016.1 (2016).
Lu, T. X. & Rothenberg, M. E. Microrna. J. Allergy Clin. Immunol. 141(4), 1202–1207. https://doi.org/10.1016/j.jaci.2017.08.034 (2018).
Lyu, L. L., Feng, Y. & Liu, B. C. Urinary biomarkers for chronic kidney disease with a focus on gene transcript. Chin. Med. J. 130(18), 2251–2256. https://doi.org/10.4103/0366-6999.213965 (2017).
Duan, Z. Y. et al. Selection of urinary sediment miRNAs as specific biomarkers of IgA nephropathy. Sci. Rep. 6, 23498. https://doi.org/10.1038/srep23498 (2016).
Szeto, C. C. et al. Urinary mirna profile for the diagnosis of IgA nephropathy. BMC Nephrol. 20(1), 77. https://doi.org/10.1186/s12882-019-1267-4 (2019).
Wang, N. et al. Profiling and initial validation of urinary microRNAs as biomarkers in IgA nephropathy. PeerJ 3, e990. https://doi.org/10.7717/peerj.990 (2015).
Levey, A. S. et al. A new equation to estimate glomerular filtration rate. Ann. Intern. Med. 150(9), 604–612. https://doi.org/10.7326/0003-4819-150-9-200905050-00006 (2009).
Trimarchi, H. et al. Oxford classification of IgA nephropathy 2016: An update from the Iga Nephropathy Classification Working Group. Kidney Int. 91(5), 1014–1021. https://doi.org/10.1016/j.kint.2017.02.003 (2017).
Duan, Z. Y. et al. U6 can be used as a housekeeping gene for urinary sediment mirna studies of IgA nephropathy. Sci. Rep. 8(1), 10875. https://doi.org/10.1038/s41598-018-29297-7 (2018).
Levin, A. et al. Kidney Disease: Improving Global Outcomes (KDIGO) CKD Work Group. KDIGO 2012 Clinical Practice Guideline for the Evaluation and Management of Chronic Kidney Disease. Kidney Int. Suppl. 3(1), 1–150 (2013).
Liang, S. et al. Urinary sediment miRNAs reflect tubulointerstitial damage and therapeutic response in IgA nephropathy. BMC Nephrol. 18(1), 63. https://doi.org/10.1186/s12882-017-0482-0 (2017).
Canney, M. et al. Quantifying duration of proteinuria remission and association with clinical outcome in IgA nephropathy. J. Am. Soc. Nephrol. 32(2), 436–447. https://doi.org/10.1681/asn.2020030349 (2021).
Inker, L. A. et al. Association of treatment effects on early change in urine protein and treatment effects on GFR slope in IgA nephropathy: An individual participant meta-analysis. Am. J. Kidney Dis. 78(3), 340–9.e1. https://doi.org/10.1053/j.ajkd.2021.03.007 (2021).
Alamartine, E. et al. The use of the Oxford classification of IgA nephropathy to predict renal survival. Clin. J. Am. Soc. Nephrol.: CJASN 6(10), 2384–2388. https://doi.org/10.2215/cjn.01170211 (2011).
Tanaka, S. et al. Secular trends in the incidence of end-stage renal disease and its risk factors in Japanese patients with immunoglobulin a nephropathy. Nephrol. Dial. Transplant. 33(6), 963–971. https://doi.org/10.1093/ndt/gfx223 (2018).
Soares, M. F. et al. Relationship between Renal Cd68(+) infiltrates and the Oxford classification of Iga nephropathy. Histopathology 74(4), 629–637. https://doi.org/10.1111/his.13768 (2019).
Bao, H. et al. Mir-223 downregulation promotes glomerular endothelial cell activation by upregulating importin Α4 and Α5 in Iga nephropathy. Kidney Int. 85(3), 624–635. https://doi.org/10.1038/ki.2013.469 (2014).
Chakera, A. et al. Prognostic value of endocapillary hypercellularity in Iga nephropathy patients with no immunosuppression. J. Nephrol. 29(3), 367–375. https://doi.org/10.1007/s40620-015-0227-8 (2016).
El Karoui, K. et al. Focal segmental glomerulosclerosis plays a major role in the progression of Iga nephropathy. Ii. Light Microscopic and Clinical Studies. Kidney Int. 79(6), 643–654. https://doi.org/10.1038/ki.2010.460 (2011).
El Karoui, K. et al. A clinicopathologic study of thrombotic microangiopathy in Iga nephropathy. J. Am. Soc. Nephrol. 23(1), 137–148. https://doi.org/10.1681/asn.2010111130 (2012).
Shen, X. H. et al. Reversal of active glomerular lesions after immunosuppressive therapy in patients with Iga nephropathy: A repeat-biopsy based observation. J Nephrol 28(4), 441–449. https://doi.org/10.1007/s40620-014-0165-x (2015).
Hou, J. H. et al. Mycophenolate Mofetil combined with prednisone versus full-dose prednisone in Iga nephropathy with active proliferative lesions: A randomized controlled trial. Am. J. Kidney Dis. 69(6), 788–795. https://doi.org/10.1053/j.ajkd.2016.11.027 (2017).
Pomaznoy, M., Ha, B. & Peters, B. Gonet: A tool for interactive gene ontology analysis. BMC Bioinf. 19(1), 470. https://doi.org/10.1186/s12859-018-2533-3 (2018).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Prot. Sci. 28(11), 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. Kegg for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res 51(D1), D587–D592. https://doi.org/10.1093/nar/gkac963 (2023).
Smoot, M. E., Ono, K., Ruscheinski, J., Wang, P.-L. & Ideker, T. Cytoscape 2.8: New features for data integration and network visualization. Bioinformatics 27(3), 431–432. https://doi.org/10.1093/bioinformatics/btq675 (2011).
Franceschini, A. et al. String V9.1: Protein–protein interaction networks, with increased coverage and integration. Nucl. Acids Res. 41, D808–D815. https://doi.org/10.1093/nar/gks1094 (2013).
Zittema, D., van den Brand, J. A., Bakker, S. J., Wetzels, J. F. & Gansevoort, R. T. Copeptin, a surrogate marker for arginine vasopressin, is associated with disease severity and progression in Iga nephropathy patients. Nephrol. Dial. Transplant. 32(1), i146–i153. https://doi.org/10.1093/ndt/gfw391 (2017).
Hinman, R. M. et al. Foxo3–/– mice demonstrate reduced numbers of pre-B and recirculating B cells but normal splenic B cell sub-population distribution. Int. Immunol. 21(7), 831–842. https://doi.org/10.1093/intimm/dxp049 (2009).
Wang, Y. Y. et al. Functional implications of regulatory B cells in human Iga nephropathy. Scand. J. Immunol. 79(1), 51–60. https://doi.org/10.1111/sji.12128 (2014).
Tian, J., Wang, Y., Guo, H. & Li, R. The Akt/Mtor/P70s6k pathway is activated in Iga nephropathy and rapamycin may represent a viable treatment option. Exp. Mol. Pathol. 99(3), 435–440. https://doi.org/10.1016/j.yexmp.2015.08.004 (2015).
Cox, S. N. et al. Altered modulation of Wnt-Beta-catenin and Pi3k/Akt pathways in Iga nephropathy. Kidney Int 78(4), 396–407. https://doi.org/10.1038/ki.2010.138 (2010).
Li, C., Shi, J. & Zhao, Y. Mir-320 promotes B cell proliferation and the production of aberrant glycosylated Iga1 in Iga nephropathy. J. Cell Biochem. 119(6), 4607–4614. https://doi.org/10.1002/jcb.26628 (2018).
Liu, D. et al. Rapamycin enhances repressed autophagy and attenuates aggressive progression in a rat model of Iga nephropathy. Am. J. Nephrol. 45(4), 293–300. https://doi.org/10.1159/000456039 (2017).
Doss, J. F. et al. A comprehensive joint analysis of the long and short Rna transcriptomes of human erythrocytes. BMC Genom. 16, 952. https://doi.org/10.1186/s12864-015-2156-2 (2015).
Funding
This study was supported by the Natural Science Foundation of China (NSFC) (82170686), National Key Research and Development (R&D) Program of China (2018YFA0108803), and the Science and Technology Project of Beijing, China (D181100000118004).
Author information
Authors and Affiliations
Contributions
M.Z., Z.-Y.D., and G.-Y.C. designed the study. G.-Y.C. and X.-M.C. revised and approved the analytical protocol. P.W., Z.-Y.D., Q.-Y.Z., and X.-G.-X.X. collected the samples. P.W., Y.Z., S.-W.D., and J.W. collected the clinical and pathological data. M.Z. was in charge of finishing the experiment and the writing of the manuscript. Z.-Y.D., G.-Y.C., Y.Z., S.-W.D., J.W., and X.-M.C. revised and amended the manuscript. All authors participated in the discussion. All authors contributed to the article and approved the the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, M., Duan, ZY., Zhang, QY. et al. Urinary miR-16-5p can be used as a potential marker of endocapillary hypercellularity in IgA nephropathy. Sci Rep 13, 6048 (2023). https://doi.org/10.1038/s41598-023-32910-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-32910-z
- Springer Nature Limited
This article is cited by
-
Blood and urine biomarkers of disease progression in IgA nephropathy
Biomarker Research (2024)
-
Clinical significance of urinary exosomal microRNAs in patients with IgA nephropathy
Scientific Reports (2023)