Abstract
Annual cues in the environment result in physiological changes that allow organisms to time reproduction during periods of optimal resource availability. Understanding how circadian rhythm genes sense these environmental cues and stimulate the appropriate physiological changes in response is important for determining the adaptability of species, especially in the advent of changing climate. A first step involves characterizing the environmental correlates of natural variation in these genes. Band-rumped and Leach’s storm-petrels (Hydrobates spp.) are pelagic seabirds that breed across a wide range of latitudes. Importantly, some populations have undergone allochronic divergence, in which sympatric populations use the same breeding sites at different times of year. We investigated the relationship between variation in key functional regions of four genes that play an integral role in the cellular clock mechanism—Clock, Bmal1, Cry2 and Per2—with both breeding season and absolute latitude in these two species complexes. We discovered that allele frequencies in two genes, Clock and Bmal1, differed between seasonal populations in one archipelago, and also correlated with absolute latitude of breeding colonies. These results indicate that variation in these circadian rhythm genes may be involved in allochronic speciation, as well as adaptation to photoperiod at breeding locations.
Similar content being viewed by others
Introduction
Importance of circadian rhythms
An organism’s fitness depends directly on its ability to synchronize important life history events with optimal resource availability1. In seasonal locations, annual cues in the environment result in physiological changes that allow organisms to time reproduction during periods of food abundance2,3. Photoperiod, or the number of hours of daylight during a twenty-four-hour period, varies based on time of year and latitude. Specifically, as latitude increases, photoperiod displays greater annual fluctuation. In some organisms, photoperiod has greater influence on breeding time than do other environmental cues such as temperature or precipitation4,5. Changes in photoperiod directly regulate circadian and circannual rhythms in many temperate species6,7. These rhythms modulate daily and annual changes in physiology and behaviour in response to environmental cues, and without them, organisms have reduced survival of offspring3,8.
The clock machinery is highly complex, and multiple circadian rhythm genes are involved in sensing changes in photoperiod and stimulating appropriate physiological changes (e.g., for breeding). In this study we focus on Clock, Per, Bmal1, and Cry, as these genes transcribe protein products that form a significant part of the primary circadian clock mechanism in vertebrates9,10 (Table 1). The central role of these genes is to synchronize the internal circadian rhythm with external environmental cues2,11. The mechanism consists of positive and negative transcription/translation feedback loops that are expressed in many tissues, such as the suprachiasmatic nucleus of the anterior hypothalamus, mediobasal hypothalamus and pineal gland11,12,13 (Fig. 1). CLOCK and BMAL1 form a heterodimer in the cell nucleus in the presence of light, which binds to E-box regulatory elements in Per and Cry genes, activating transcription and translation into PER and CRY. Once PER and CRY have accumulated to a threshold level, they bind to casein kinase 1δ/ε to form a heterodimer, enter the nucleus, and mediate the displacement of CLOCK-BMAL1. As PER and CRY degrade over time, CLOCK and BMAL1 reform their complex and the cycle repeats itself. The CLOCK-BMAL1 heterodimer is involved in at least two additional regulatory loops promoting the transcription of other important circadian gene products (e.g., REV-ERBα and REV-ERBβ), which ultimately also affect the formation of CLOCK-BMAL114. These feedback loops create rhythmic oscillations in gene expression that manifest as daily or seasonal changes in behaviour and physiology. A mutation in any of these genes can have pleiotropic effects, potentially causing molecular, physiological or behavioural arrhythmicity that may lead to reduced fitness [reviewed in8,11,13]. For example, mutations in several circadian genes reduce fertility in Drosophila [reviewed in 8]; and female blue tits (Cyanistes caeruleus) with fewer glutamine repeats fledge a higher number of offspring15.
Core circadian gene mechanism. The dashed line represents the nuclear membrane. CLOCK and BMAL1 form a heterodimer in the cell nucleus in the presence of light, which binds to E-box regulatory elements in Per and Cry genes, as well as to other Clock Controlled Genes (CCG), causing transcription of PER and CRY (positive symbol). Once PER and CRY have accumulated to a threshold level, they form a heterodimer that enters the nucleus to repress CLOCK-BMAL1 (negative symbol). As PER and CRY degrade over time, CLOCK and BMAL1 can reform their complex, causing transcription of PER and CRY once again as the cycle repeats itself (positive symbol).
Variation in circadian genes with breeding time and latitude
Circadian gene variation has been associated with differences in breeding time in both laboratory and wild populations. For example, in Bmal1 knockout mice, irregular secretions of gonadotropin-releasing hormone resulted in smaller ovaries, smaller uteri, and delayed puberty22,36,37. Additionally, within wild populations of blue tits and Chinook salmon (Oncorhynchus tshawytscha), individuals who bred earlier in the season had Clock alleles with shorter polyQ regions than individuals breeding later15,38,39,40. Variation in circadian gene allele frequencies have also been associated with latitude, suggesting that ecological parameters (e.g. photoperiod) that vary with latitude select for different alleles. For example, Drosophila exhibit a latitudinal cline in the number of Thr-Gly coding repeats in their Per2 sequence41,42; and the cryptochrome gene is highly differentiated between high and low latitude Drosophila populations in Australia43. Although the relationship between latitude and individual genes from the core clock mechanism has been studied previously, the relationship between latitude and combined genes has not yet been defined.
Storm-petrel species complexes as a study system
We explored the association between circadian gene allele variation with breeding season and latitude in the band-rumped and Leach’s storm-petrel species complexes (Hydrobates spp.; Procellariiformes: Hydrobatidae). Storm-petrels are small, pelagic seabirds that breed across a wide latitudinal range in both the North Atlantic and North Pacific oceans. These tube-nosed seabirds are generally philopatric, returning to their native breeding sites annually to reproduce. Multiple colonies have two sympatric allochronic populations, often breeding in the same nest sites in different seasons: one in winter and one in summer, with little to no overlap in breeding time. Seasonal populations have arisen independently within several archipelagoes, and represent a range of divergences from genetically similar populations, through genetically differentiated races, to reproductively isolated species in both complexes44,45,46,47. In this study we tested whether key functional regions of four core circadian genes vary (i) between populations with seasonal differences in breeding time, and/or (ii) between populations (including seasonal races and species) at different latitudes. Evidence from previous studies of natural populations of tits, salmon and Drosophila revealed that the number of glutamine repeats in the Clock gene correlates with both latitude and timing of breeding onset, possibly due to slower degradation of the longer glutamine tails (above). We therefore predicted that allele length would increase with breeding latitude in storm-petrels. We also predicted that Clock alleles would be longer in seasonal populations that initiate breeding when daylength is increasing (spring). Although the mechanisms of action of the other three genes are less well known in birds, we hypothesized that they also would vary by latitude and breeding season, changing phase, binding specificity or photo-entrainment as in other species.
Methods
Sample collection and preparation
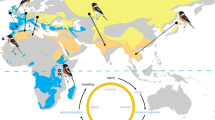
Blood samples were collected from individual breeding band-rumped and Leach’s storm-petrels from throughout most of the species’ ranges, including seasonal populations (Fig. 2)46,47. All birds were handled under approved Animal Use Protocols, and were released immediately after sampling. The summer breeding season was classified as months experiencing increasing temperature and longer daylengths, and the winter breeding season as months experiencing decreasing temperature and shorter daylengths. Difficulties capturing birds mean that sample sizes at some colonies are small and variation may be underestimated. Blood samples were digested using proteinase-K, RNA contamination was removed using RNase, and DNA was extracted and cleaned using the phenol–chloroform method followed by ethanol precipitation48.
Sampling locations of (a) band-rumped, and (b) Leach’s storm-petrel species complexes (Hydrobates spp.). Filled circles indicate summer-breeding colonies, empty circles indicate winter-breeding colonies, and circles with a cross indicate allochronic populations. Boxes represent zoomed-in areas corresponding to plots to the right. The map was created in R using the ggmap package50.
Amplification and sequencing of circadian genes
Polymerase chain reaction (PCR) was performed for key functional portions of each of four core circadian clock genes. For Clock, forward (5'-TTT TCT CAA GGT CAG CAG CTT GT-3') and reverse (5'- CTG TAG GAA CTG TTG (C/T)GG (G/T)TG CTG-3') primers were previously designed to amplify a region of the gene with functional variation in polyQ repeats in avian species16. PCRs were performed in 10 μl reaction volumes, consisting of 5 μl Multiplex Mix (Qiagen, Mississauga, Ontario), 1.0 μl DNA, and 0.15 μl Dye 4—labeled M13 forward and reverse primers. A Biometra T-gradient Thermocycler (Biometrik Analytik, Goettingen, Germany) was used to conduct an initial denaturation at 95 °C for 15 min, followed by 35 cycles of 95 °C for 30 s, 55 °C for 45 s, and 72 °C for 45 s, and a final elongation phase of 72 °C for 3 min.
For Bmal1, forward (5’-CTA AAG TCA GCA TTT GAA AC-3’) and reverse (5’-CTC ACG TTG CTT GCC ACT AC-3’) primers were designed using the full genome of the Leach’s storm-petrel51 to amplify the 93 bp of exon 15. For Per2, each individual was genotyped for a 102 bp region corresponding to nucleotides 2536 to 2638 using previously designed forward (5’-TCT GGT AAA TCA AGT GGT CCY CCA GT-3’) and reverse (5’-TTC TAA TTC AGG TTG TGG CTT TTT GTC-3’) primers. Both these fragments were previously examined in barn owls (Tyto alba), chickens (Gallus gallus), and quails (Coturnix japonica), as these regions contain casein kinase 1e (CK1e) phosphorylation sites important for normal circadian function25. For Cry2, forward (5’-CCA GTC CTC CCC TGC TCG TG-3’) and reverse (5’-GAA ATG CCA GAC TCA CCC TG-3’) primers were designed from the Leach’s storm-petrel full genome to amplify 210 bp of exon 7, which is known to contain multiple flavin adenine dinucleotide (FAD)-binding sites52. PCRs were performed in 25 μl reaction volumes, consisting of 12.5 μl Multiplex Mix (Qiagen, Mississauga, Ontario), 0.04 μM forward primer, 0.04 μM reverse primer, and 2 μl DNA. PCRs were run as above but with annealing temperatures of 58 °C for Bmal1, 50 °C for Per2, and 60 °C for Cry2.
PCR products were electrophoresed through 2% agarose gels to confirm amplification. For Clock, 10 amplifications from Praia spring (Monteiro’s storm-petrel, Hydrobates monteiroi) and fall (band-rumped storm-petrel, H. castro) populations were sequenced by Genome Quebec (McGill University, Montreal, Quebec). The resulting sequences were aligned using BioEdit (version 7.0.953) to confirm amplification of the intended target. A Beckman-Coulter CEQ 8000™ Genetic Analysis System (Core Genotyping Facility, Department of Biology, Queen’s University) was then used to screen for variation in allele lengths in all individuals. The Beckman-Coulter CE 8000 Genetic Analysis software (Beckman Coulter, Inc., Fullerton, CA) was used to score the resulting chromatograms. Clock allele lengths and sequences of 10 individuals were compared to determine the corresponding number of polyQ repeats based on allele length. Amplified products for all other genes were sequenced at Genome Quebec and aligned using Geneious54. Basic Local Alignment Search Tool (BLAST) searches on GenBank were used to confirm that the target genes were correctly amplified55. After base calls on the chromatograms were confirmed by eye, individual sequences were compared for variation.
Previously published data on microsatellite variation was used as a presumptively neutral variation to compare against variation in circadian genes44,47,56.
Data analysis
To test for significant population differentiation within each gene or combination of genes, population genetic analyses were performed in ARLEQUIN57. Population classifications were determined by both breeding time and geographic location. First, Bmal1 (total sample sizes (n): band-rumped storm-petrel n = 296, Leach’s storm-petrel n = 300), Cry2 (band-rumped storm-petrel n = 298, Leach’s storm-petrel n = 232), and Per2 (band-rumped storm-petrel n = 240, Leach’s storm-petrel n = 140) sequences were phased using the program DnaSP v5.10.158 set to 1000 iterations and 100 burn-in. Then, ARLEQUIN was used to test genotypic data for deviations from Hardy–Weinberg proportions (HWP). ARLEQUIN was used to estimate Wright’s FST, \(\phi\) ST for Bmal1, Cry2 and Per2, and Slatkin’s derivation of Wright’s RST for Clock59 between sampling sites and to test their statistical significance (10,100 permutations). Finally, ARLEQUIN was used to test for deviations from linkage equilibrium between pairs of circadian genes (10,100 permutations). Tests for deviations from linkage equilibrium were performed by grouping individuals from different colonies into genetic populations based on nonsignificant FST values and geographic proximity. For tests with greater than 20 comparisons, p-values were adjusted using the False Discovery Rate method and assessed for statistical significance using α = 0.0560. A Principal Components Analysis (PCA) was also performed on all samples that had sequences for all four genes (Clock, Bmal1, Cry2, and Per2) to help identify associations between haplotypes from different genes.
To test whether a significant relationship between circadian gene variation and breeding time or latitude exists, regressions were performed in R using the stats package49. For the Bmal1 and Cry2 sequences, two logistic regressions were run to determine whether the probability of having the most common allele can be predicted by latitude. Likelihood ratio tests and Akaike’s Information Criterion (AIC) of multiple generalized linear models were used to determine which variable(s) (ocean, species, breeding season and/or latitude), or interactions between variables, most accurately predicted presence/absence of specific alleles. Linear regressions were performed between latitude and Clock allele length, controlling for ocean in band-rumped storm-petrels. Due to non-normality of the residuals, confirmed by the Shapiro–Wilk test (band-rumped storm-petrels: W = 0.84, p < 0.001 n Atlantic = 724, n Pacific = 172; Leach’s storm-petrels: W = 0.81, p < 0.001, n = 366), estimates (intercept and slope) and measures of significance (p-values and 95% confidence intervals) of the model were obtained using 1000 bootstrap permutations (bootstrapped R2 values denoted with * in Results). Finally, the mean polyQ repeat number for each population was calculated and significant differences between allochronic populations were assessed using a Student’s t-test in R. While the use of mean allele length may obscure some features of allelic variation, it is deemed appropriate given the co-dominant nature of alleles at the Clock locus44,61.
Finally, to test whether significant correlations between circadian rhythm genes and latitude are better explained by geographic distance, Mantel tests62 were performed in R using the ape package. Genetic distances were estimated using Slatkin’s linearized pairwise FST values, and geographical distance was calculated as Euclidean distance between sampling sites based on latitude and longitude. Individuals from sympatric seasonal populations that were genetically indistinct based on Wright’s FST were grouped together for this analysis. For the band-rumped storm-petrels, samples from the Azores summer-breeding population and Cape Verde were removed from this analysis since they are considered separate species (Monteiro’s storm-petrel, and Cape Verde storm-petrel [H. jabejabe], respectively). Mantel tests were run using Pearson’s correlation for 10,000 permutations, controlling for either ocean (Atlantic or Pacific) or season (summer or winter). All tests were assessed for statistical significance using α = 0.05.
Ethical approval
All samples were collected under Queen’s University University Animal Care Committee approved Animal Use Protocols, and birds were released immediately after handling (i.e. no experiments were conducted). All laboratory work was conducted in accordance with Queen’s University safety and biohazard regulations. All relevant Animal Research: Reporting of In Vivo Experiments (ARRIVE) guidelines were followed. Fieldwork was conducted under permits appropriate to each collection site. All methods were carried out in accordance with relevant guidelines and regulations.
Results
Circadian gene variation
Comparison with published avian sequences on GenBank confirmed that the focal fragments for all circadian genes were amplified correctly (see Table 2 for summaries of variation). Per was invariant across all samples for both species complexes. For Clock, all alleles differed by multiples of 3 bp and comparison of allele lengths and sequences indicated that variation in Clock alleles is due to the number of glutamine repeats.
Significant deviations from linkage equilibrium within genetic populations were found between Bmal1 and Clock in band-rumped storm-petrels (average r2 = 0.00–0.05, p < 0.01), and between all pairs of genes tested in Leach’s storm-petrels (Bmal1 ~ Clock average r2 = 0.00–0.06, p = 0.01, n = 276; Cry2 ~ Clock average r2 = 0.00–0.07, p < 0.01, n = 216; Cry2 ~ Bmal1 average r2 = 0.00–0.22, p < 0.01). A PCA comparing clock gene haplotypes also indicated a potential correlation between Cry2 and Clock variation in band-rumped storm-petrels, but not with the other clock genes and not in Leach’s storm-petrels (Supplementary Figure S1). Results from the linkage disequilibrium tests likely differed from the PCA for the Leach’s storm-petrels as the linkage disequilibrium tests looked for a relationship in each genetic population individually, whereas the PCA grouped all data together and only used samples that had been successfully sequenced for all four clock genes.
Circadian gene variation and breeding season
Among allochronic band-rumped storm-petrel populations, only Azores summer and winter had estimates of FST significantly different from 0 based on Clock allele variation (FST = 0. 29, p < 0.01; Supplementary Table S14). Mean Clock polyQ lengths did not differ significantly between allochronic populations (t = 1.64, df = 893 p = 0.10). In allochronic populations of Leach’s storm-petrels on Guadalupe, pairwise estimates of FST did not differ from 0 (pairwise FST = 0.00, p = 0.57; Supplementary Tables S15, S16) and mean lengths of Clock alleles did not differ (t = 0.83, df = 71, p = 0.41).
Estimates of FST based on Bmal1 allele frequencies did not differ from 0 between band-rumped storm-petrel or Leach’s storm-petrel populations separated by breeding time (Supplementary Tables S17–S20).
Pairwise estimates of FST for Cry2 were significantly different from 0 between summer and winter breeding band-rumped storm-petrel populations in the Azores (FST = 0.49, p < 0.01). In the logistic regression model, breeding season was a significant predictor of Cry2 alleles, with winter-breeding birds 3.17 times more likely to have allele 1 (eβwinter = 3.17, 95% CI = 1.80–5.59, df = 2, n = 298, p < 0.01; Fig. 3). No other estimates of FST differed from 0 between seasonal populations in either species complex (all p > 0.05; Supplementary Tables S20–S24).
Relationship between Clock allele length and absolute latitude in (a) Atlantic band-rumped storm-petrels, (b) Pacific band-rumped storm-petrels, and (c) Leach’s storm-petrels. Points represent the average allele length for each population at a given absolute latitude and season, and vertical lines represent standard errors. Solid slanted lines represent the linear regression lines, while dashed slanted lines represent the 95% bootstrapped confidence intervals.
Circadian gene variation and latitude
Clock allele length correlated significantly with absolute latitude (R2* = 0.18 ± 0.06 for Atlantic band-rumped storm-petrels, R2* = 0.28 ± 0.10 for Pacific band-rumped storm-petrels, and R2* = 0.41 ± 0.07 for Leach’s storm-petrels, all p* < 0.01, Fig. 3). Mantel tests also found a significant correlation between genetic and geographic distance in Atlantic band-rumped storm-petrel populations (r = 0.97, p = 0.03, 10,000 permutations). According to pairwise estimates of Slatkin’s linearized FST, all higher latitude populations of Leach’s storm-petrels (Japan, Aleutians, Aiktak, British Colombia) except the Midun colony were significantly differentiated from lower latitude populations (San Benito, Guadalupe; FST = 0.52–3.38, all p < 0.01; Supplementary Table S15). A Mantel test for Leach’s storm-petrels found no significant correlation between genetic and geographic distance (r = 0.30, p = 0.09, 10,000 permutations).
Latitudinal variation was found in Bmal1 for band-rumped storm-petrels. Based on two logistic regressions controlling for ocean, the probability of having allele 1 was significantly greater at higher absolute latitudes (eβ = 1.21, 95% CI = 1.11–1.31, df = 3, p < 0.01), whereas the probability of having allele 2 was significantly greater at lower latitudes (eβ = 0.80, 95% CI = 0.71 –0.90, df = 3, p < 0.01; Fig. 4; Supplementary Figure S2). The logistic regression was performed controlling for ocean because this was a significant predictor with latitude, according to likelihood ratio tests comparing all potential predictors. These results are consistent with the pairwise Slatkin’s linearized FST estimates indicating that the South Atlantic colonies have greater differentiation from colonies elsewhere (FST = 0.20–1.45, all p < 0.01; Supplementary Table S17). For Leach’s storm-petrels, the likelihood ratio test indicated that logistic regressions including absolute latitude as a predictor were not significantly better than the null model (χ2 = 4.29, df = 2, p = 0.12). Further, none of the pairwise estimates of Slatkin’s linearized FST for Leach’s storm-petrels were significant (Supplementary Tables S19, S20). No significant correlations were found between genetic and geographic distance using Mantel tests for either species complex (Atlantic band-rumped storm-petrels: r = 0.84, p = 0.07; Leach’s storm-petrels: r = -0.01, p = 0.44, 10,000 permutations each).
Relationship between frequency of Bmal1 haplotypes and absolute latitude for (a) band-rumped storm-petrels, and (b) Leach’s storm-petrels. Each point represents whether an individual from a population has the most common allele for that species complex or not, and the colour of the point depicts whether that individual originates from a summer- or winter-breeding population. Solid lines represent the logistic regression curves for the most common haplotype in each species complex, controlling for ocean. Dashed lines represent the standard error surrounding each logistic regression curve.
Similar to Clock and Bmal1, Cry2 variation also varied with latitude. When controlling for season, the frequency of allele 1 in band-rumped storm-petrels and Leach’s storm-petrels was significantly greater at higher absolute latitudes (band-rumped storm-petrels: eβ = 1.07, 95% CI = 1.04–1.09, df = 3, p < 0.01; Leach’s storm-petrels: eβ = 1.05, 95% CI = 1.02–1.09, df = 3, p < 0.01; Fig. 3; Supplementary Tables S4, S5). The frequency of allele 2 in band-rumped storm-petrels was significantly lower at higher absolute latitudes (band-rumped storm-petrels: eβ = 0.96, 95% CI = 0.94–0.98, df = 3, p < 0.01; Fig. 5). No significant correlations between genetic and geographic distance were found for either species complex from the Mantel’s tests (band-rumped storm-petrels: r = 0.53, p = 0.08; Leach’s storm-petrels: r = -0.03, p = 0.50, 10,000 permutations each).
Relationship between frequency of Cry2 haplotypes and absolute latitude for (a) allele 1 in band-rumped storm-petrels, (b) allele 2 in band-rumped storm-petrels and (c) allele 1 in Leach’s storm-petrels. Each point represents whether an individual from a population has the most common allele for that species complex or not, and the colour of the point depicts whether that individual originates from a summer- or winter-breeding population. Solid lines represent the logistic regression curves for the most common haplotype in each species complex, controlling for breeding season. Dashed lines represent the standard error surrounding each logistic regression curve.
Discussion
In this study we aimed to characterize the relationship between variation in four core circadian rhythm genes and both breeding season and absolute latitude. To investigate this relationship, we explored variation in key functional regions in Clock, Bmal1, Cry2 and Per2 in two seabird species complexes containing populations that span a wide range of latitudes. Importantly, some of these populations have undergone allochronic divergence and even speciation within several archipelagos. We ultimately determined that circadian gene alleles (i) differ between seasonal populations within one but not all archipelagos, and (ii) vary among populations spanning different latitudes.
Circadian gene variability in band-rumped and Leach’s storm-petrels
Variation in PolyQ length of Clock alleles
Although no new Clock alleles were detected in band-rumped or Leach’s storm-petrels compared to other species that have been studied, storm-petrels showed a greater total number of alleles at the Clock locus than do most other avian species studied to date, including the barn swallow (Hirundo rustica)63,64, bluethroat (Luscinia svecica)38, great tit (Parus major)65, and multiple species of tree swallow (Tachycineta bicolor, T. thalassina, T. albilinae, T. leucorrhoa, and T. meyeni)66.
The South Atlantic (St. Helena and Ascension Islands) populations of band-rumped storm-petrel and the San Benito Leach’s storm-petrel population deviated from HWP based on Clock allele frequencies. All these populations had heterozygote deficiencies, which can be due to a sampling artifact or population effects such as selection and non-random mating. In the case of the South Atlantic band-rumped storm-petrels, samples from two seasonal populations were grouped together (Supplementary Table S1), and thus the heterozygote deficiency could be due to the Wahlund effect, as neither population deviates from HWP when considered separately (Supplementary Table S2). San Benito comprises three small islands, and samples used in this analysis were taken from West, Central, and East Benito. Although the Wahlund effect is plausible for this group, neither a principal components analysis nor pairwise estimates of FST using SNPs from reduced representation sequencing detected genetic differentiation between islands (Supplementary Figure S3; Supplementary Table S25). In addition, none of six microsatellite loci analyzed in a previous study for Leach’s storm-petrels from San Benito deviated from HWP47, further indicating a lack of population substructure or inbreeding. Therefore, deviation of Clock genotypes from HWP in the San Benito population is more likely explained by purifying selection, selection against heterozygotes, or linkage disequilibrium (see below).
Variation in Bmal1
The two synonymous single nucleotide polymorphisms (SNPs) in Bmal1 in band-rumped storm-petrels involve purine substitutions within adenine and guanine. Although synonymous, these mutations could influence the thermodynamic stability of the mRNA secondary structure, consequently affecting the speed of mRNA translation67. Degradation of BMAL1 is an important step in the negative feedback loop maintaining behavioural and physiological rhythms27, and a change in the speed of translation could alter the circadian phenotype.
In the Leach’s storm-petrels, the two non-synonymous mutations involved one heterozygous individual from Semidi Island with an aspartic acid in place of a glycine, and one homozygous individual from the Guadalupe winter-breeding population with a proline in place of a serine. Serine residues, unlike proline, are susceptible to phosphorylation68, thus the replacement of serine with proline results in the loss of potential phosphorylation sites. In mice, phosphorylation on serine 17 in BMAL1 results in decreased production of DNA binding protein, ultimately hindering transcription and reducing the efficacy of the circadian output69. If this mutation at nucleotide 73 in Bmal1 exon 15 does indeed eliminate a phosphorylation site, then the Guadalupe individual may experience circadian arrhythmicity, consequently impacting its breeding phenology. Even though this mutation was only found in one of the sampled individuals, it may be prevalent in the population since the individual was homozygous at the site of the SNP (Supplementary Table S7).
Variation in Cry2
Despite the high diversity of Cry2 alleles in band-rumped, and especially Leach’s storm-petrels (Table 2), only one mutation was non-synonymous, resulting in an amino acid change from isoleucine to phenylalanine. These two amino acids have similar biochemical properties and thus the mutation likely does not have a significant effect on protein function. The Cry2 exon sequenced in this study centres on the FAD-binding site in the N-terminus, as this region is likely involved in blue light photo-sensing and therefore may be directly involved in entraining circadian rhythms70,71. The paucity of amino acid substitutions despite the high diversity of alleles is therefore not surprising given the conserved nature of the FAD-binding site and its important function70. Whether this region is highly variable in other avian species is uncertain, as the Cry2 gene sequence is relatively unstudied in birds.
Cry2 sequence variation has been studied extensively in mice, insects and plants, but findings from those studies may not provide much insight into Cry2 function in the avian clock. For instance, many circadian genes are expressed differently across tissues in non-mammalian vertebrates72. In fact, Renthlei and colleagues73 discovered that CRY2 expression does not oscillate in the hypothalamus, pineal or intestine in tree sparrows. The pineal gland is thought to be vital to both circadian and circannual rhythms in some migratory songbirds74,75,76,77,78 (but see79) so the lack of a CRY2 oscillation may indicate that this gene plays a different role in the clock mechanism in birds compared to mammals. Ultimately, further research is required to explore fully the involvement of Cry2 in the avian circadian clock mechanism and how its function differs from the mammalian circadian mechanism.
Variation in Per2
No variation in the Per2 gene was detected across the 10 band-rumped storm-petrel populations or the seven Leach’s storm-petrel populations. The lack of variation is surprising given that both band-rumped and Leach’s storm-petrels show geographic variation at microsatellites, mitochondrial DNA, and other candidate genes44,80,81, and could indicate strong purifying selection on the Per2 gene. Similarly, Fidler and Gwinner25 found that Per2 was highly conserved among 12 day- and night-active avian species, and that the predicted translation products were identical.
Linkage disequilibrium
Linkage disequilibrium was found between Clock and Bmal1 in band-rumped storm-petrels and between all combinations of Bmal1, Clock, and Cry2 in Leach’s storm-petrels, indicating that particular allele combinations may offer a selective advantage. In addition, a PCA across all samples for allele variation in the three variable clock gene fragments in this study indicated a possible relationship between Clock and Cry2 in the band-rumped storm-petrels (Supplementary Figure S1). Further, the alleles that correlate together for Clock and Cry2 both tend to occur in higher latitudes.
The non-random association between Clock and Bmal1 alleles for both storm-petrel species complexes is noteworthy, considering the relationship between the protein products of these two genes. Although the genes are located on different chromosomes82,83 and thus not physically linked, their protein products bind together to form the CLOCK-BMAL1 heterodimer, so the genes may have evolved together to preserve binding compatibility and/or enhance binding specificity. Research in mice determined that when CLOCK and BMAL1 bind together, the complex forms unique interfaces that interact with the Per and Cry genes, as well as with the PER-CRY heterodimer84. Mutations that affect these interfaces can lead to structural instability of the CLOCK-BMAL1 heterodimer, reduced transcriptional activity, and arrhythmic circadian oscillations84.
Photoperiod-related selection can also drive this pattern of linkage disequilibrium. All circadian genes in linkage disequilibrium show a latitudinal cline: Clock has a cline in the length of the polyQ tail, whereas Cry2 and Bmal1 have a cline in allele frequencies. However, the relative effects of environmental (e.g. photoperiod) and intrinsic (e.g. binding specificity) pressures are difficult to distinguish. To fully explore whether the binding specificity of protein products differs based on Clock allele length or Bmal1 haplotype, and whether there is a relationship between binding specificity and latitude, further studies could compare levels of binding activity across latitudinally co-distributed variants85.
Finally, linkage disequilibrium can be the result of neutral processes, such as genetic drift. As population size decreases, combinations of alleles can arise due to random effects. Currently, Leach's storm-petrel populations are declining globally, and species breeding on Guadalupe Island are relatively small (2500–10,000 breeding pairs for both Townsend's Hydrobates socorroensis and Ainley's storm-petrels H. cheimomnestes)86. Although most band-rumped storm-petrel population sizes are large, populations that were recently elevated to species status in the Azores (Monteiro's storm-petrel) and Cape Verde (Cape Verde storm-petrel) are small (250–1000 and 15,500–67,500 breeding pairs respectively)86. These small population sizes suggest that Bmal1 and Clock alleles may be inherited together due to chance, rather than because of environmental or intrinsic selection pressures.
Comparison of circadian genes between allochronic populations
Allochronic populations of band-rumped and Leach’s storm-petrels provide an ideal opportunity to explore an association between circadian gene variation and breeding time. Neutral variation differs between summer and winter populations of band-rumped storm-petrels in the Azores and possibly Selvagem, and between seasonal races of Leach’s storm-petrels on Guadalupe Island47,87. In fact, the two seasonal populations of band-rumped storm-petrels in the Azores and Leach’s storm-petrels on Guadalupe Island are considered separate species88. Pairwise estimates of FST for both Clock and Cry2 differed significantly between seasonal populations of band-rumped storm-petrels in the Azores, and breeding season was a significant predictor of Cry2 alleles in the logistic regression model. Otherwise, no consistent patterns were detected based on breeding time. These results contrast with previous studies that found a correlation between the number of Clock polyQ repeats and breeding time in blue tits15 and barn swallows64. In both studies, fewer polyQ repeats were associated with earlier breeding times, and longer polyQ repeats were associated with later breeding times15,64. However, the conserved nature of the Bmal1 exon indicates possible purifying selection between diverging lineages in both band-rumped and Leach’s storm-petrels, and may further highlight the importance of Bmal1 functionality in regulating circadian rhythms.
The lack of a consistent pattern in circadian gene variation between seasonal breeding populations could be due to several factors. Firstly, evaluation of temporal patterns was based on population classification as either summer or winter breeders. This classification may have obscured other influences. For instance, seasonal populations may inhabit different non-breeding ranges and the photoperiod experienced during the non-breeding season may influence when individuals return to colonies to breed. Alternate classification criteria, such as population divergence time, may provide a better explanation for allelic differences between allochronic populations.
Secondly, lack of a consistent seasonal pattern in circadian gene variation may indicate that this variation does not influence breeding season. On a molecular level, determinants of breeding time in birds are complex and not fully understood89. In mammals and birds, photoperiod cues received in the retina signal the circadian clock to produce melatonin. In mammals, melatonin acts as a messenger to stimulate and regulate the gonadal axes89. In birds, however, melatonin production does not appear to stimulate gonads [90, but see91]. Additionally, other circadian genes could have a greater influence on breeding time than the gene fragments chosen in this study. For instance, avian Cry4 is thought to be involved in the light-dependent magnetic compass for migration to the breeding grounds and, consequently, variation in this gene may correlate with breeding time92.
Thirdly, differences in circadian gene variation between allochronic populations may result from neutral processes, as supported by the relatively high divergence in neutral molecular markers between seasonal populations in the Azores. Furthermore, allochronic populations of storm-petrels tend to live close to the equator, where photoperiod is relatively constant; the only significant results were found at the archipelago farthest from the equator (the Azores). The lack of circadian gene variation between allochronic populations breeding at the same latitudes further supports the importance of latitude to circadian gene variation.
Latitudinal variation in circadian genes
Latitudinal cline in polyQ length of Clock alleles
A significant latitudinal cline was detected in both band-rumped and Leach’s storm-petrels in Clock polyQ repeat number. Shorter polyQ repeat regions were more prevalent at lower latitudes, similar to non-migratory blue tits and salmon with variable spawning times38,39,40. The selective advantages of a longer or shorter polyQ tail are unknown, however within-populations, individuals with a greater number of polyQ repeats tend to display delayed phenology (e.g.,93,94,95). Although some studies have attributed differences in allele length to migratory phenotypes, as this would allow additional time to reach the breeding grounds before gonadal maturation, no consistent differences in Clock variation exist between migratory and non-migratory bird species96. For instance, although migratory bluethroats have significantly longer polyQ repeats at more southern latitudes38, an interspecies study comprising 23 trans-Saharan migratory bird species found that species breeding at more northern latitudes had significantly longer polyQ repeats97.
Further, glutamine trinucleotide repeats have a relatively high mutation rate compared to other microsatellites or SNPs, possibly facilitating rapid adaptation to different environmental pressures98,99,100. In fact, some evidence indicates that Clock proteins with shortened glutamine tails have weaker activation of Per and Cry, since the glutamine tail is important for binding to the E-box regulatory sequence that activates their transcription101. This difference in activation could lead to variation in the circadian period length across absolute latitudes102.
The lack of a significant correlation between geographic and genetic distance in microsatellites for the Leach’s storm-petrels47 supports the hypothesis that the latitudinal cline in Clock allele length in these populations is not due to geographic distance alone. In contrast, the significant correlation between geographic and genetic distance in Atlantic band-rumped storm-petrel populations indicates that distance-related genetic processes such as gene flow may be influencing polyQ repeat number, as colonies closer together share similar allele lengths. However, photoperiod also varies with geographic distance to the equator, and therefore could be positively autocorrelated with geographic distance between colonies. Spatial autocorrelation could result in a false conclusion that genetic drift is driving Clock allele length polymorphisms103.
Allelic diversity with latitude in Bmal1, Cry2, and Per2
The lack of diversity in Bmal1 and Cry2 at higher latitudes is surprising given the genetic structure of these two species complexes (Figs. 4 & 5). For instance, Cape Verde populations of band-rumped storm-petrels are considered a separate species87,104, yet these populations, in addition to other populations at higher absolute latitudes (Desertas, Selvagem, Berlengas, Japan, and Hawaii), are fixed for the most common Bmal1 allele. Although more Cry2 than Bmal1 variants exist in samples from higher latitude populations, these populations also exhibit a significantly greater probability of having the Cry2 allele 1. The presence of only a single Bmal1, and possibly Cry2, allele at higher latitudes in our sample could indicate the importance of their functions in regulating breeding time or adjusting the endogenous circadian rhythm to external cues when there is a greater fluctuation in photoperiod. In addition, since photoperiod is less variable at lower latitudes, and selection on the precision of the photoperiodic response is lower in more equatorial birds25, avian populations near the equator may hold greater levels of standing genetic variation.
However, it should be noted that within the Leach's storm-petrel species complex, species and absolute latitude co-vary, as one subspecies exists in higher latitude breeding populations, and the other species and subspecies exist closer to the equator. Thus, the pattern of allelic diversity in circadian genes in Leach's storm-petrels could also be explained by genetic differentiation between species and subspecies.
Conclusion
This study has two main applications: a) understanding the relationship between Clock, Bmal1, Per2, and Cry2 genes and circadian rhythms; and b) understanding mechanisms of allochronic speciation. Firstly, elucidating the relationship between Clock polyQ repeat number and differing Bmal1 or Cry2 variants in the avian clock orthologue is informative when analyzing genetic sequences in multiple species. We found some indications that genetic variation may correlate with breeding time, but stronger evidence that it may play a role in local adaptation to parameters correlated with latitude. Since only parts of the Bmal1, Cry2, and Per2 genes were analyzed, future studies that capture the entire coding region of these genes and expand to other candidate circadian genes, such as Cry1, Cry4 or ADCYAP1104, may provide better insight into how circadian gene variation correlates with latitude and/or breeding time.
Secondly, the Azores band-rumped storm-petrels and the Guadalupe Island Leach’s storm-petrels are some of the few documented examples of allochronic speciation in tetrapods44,46. Additionally, seasonal populations of band-rumped storm-petrels in Desertas, Selvagem, and the Galapagos are all possible examples of incipient speciation, each representing a different stage of speciation. The mechanisms of allochronic speciation, however, are not fully understood. Further research is required to fully understand the potential role of Cry2 in facilitating allochronic speciation, possibly including more individuals from seasonal populations and a larger portion of the gene. The circadian clock mechanism is also highly complex, composed of other genes and regulatory loops that were not covered by this study. Future studies could focus on whether variation in these additional core clock genes is associated with allochronic speciation and whether these genes vary latitudinally. Finally, further research could explore mechanisms of action of these variants, circadian rhythm gene expression and epigenetic variation rather than sequences alone.
Data availability
The datasets used and/or analysed during the current study can be accessed through the Dryad data repository (https://doi.org/10.5061/dryad.dz08kps31) and GenBank (Accession No. OR223820-OR223859).
References
Visser, M. E., Caro, S. P., Oers, K. Van., Schaper, S. V. & Helm, B. Phenology, seasonal timing and circannual rhythms: Towards a unified framework. Philos. Trans. R. Soc. Lond., B, Biol. Sci. 365, 3113–3127. https://doi.org/10.1098/rstb.2010.0111 (2010).
Bradshaw, W. E. & Holzapfel, C. M. Phenotypic evolution and the genetic architecture underlying photoperiodic time measurement. J. Insect Physiol. 47(809–820), 2001. https://doi.org/10.1016/S0022-1910(01)00054-3 (2001).
Thomas, D. W., Blondel, J., Perret, P., Lambrechts, M. M. & Speakman, J. R. Energetic and fitness costs of mismatching resource supply and demand in seasonally breeding birds. Science 291, 2598–2600. https://doi.org/10.1126/science.1057487 (2001).
Lambrechts, M. M., Blondel, J., Maistre, M. & Perret, P. A single response mechanism is responsible for evolutionary adaptive variation in a bird’s laying date. Proc. Natl. Acad. Sci. U.S.A. 94, 5153–5155. https://doi.org/10.1073/pnas.94.10.5153 (1997).
Lambrechts, M. M. & Perret, P. A long photoperiod overrides non-photoperiodic factors in blue tits’ timing of reproduction. Proc. Royal Soc. B 267, 585–588. https://doi.org/10.1098/rspb.2000.1041 (2000).
Dawson, A., King, V. M., Bentley, G. E. & Ball, G. F. Photoperiodic control of seasonality in birds. J. Biol. Rhythms 16, 365–380. https://doi.org/10.1177/074873001129002079 (2001).
Helm, B. et al. Annual rhythms that underlie phenology: Biological time-keeping meets environmental change. Proc. Royal Soc. B 280, 20130016. https://doi.org/10.1098/rspb.2013.0016 (2013).
Yerushalmi, S. & Green, R. M. Evidence for the adaptive significance of circadian rhythms. Ecol. Lett. 1, 970–981. https://doi.org/10.1111/j.1461-0248.2009.01343.x (2009).
Yasuo, S., Watanabe, M., Okabayashi, N., Ebihara, S. & Yoshimura, T. Circadian clock genes and photoperiodism: Comprehensive analysis of clock gene expression in the mediobasal hypothalamus, the suprachiasmatic nucleus, and the pineal gland of Japanese Quail under various light schedules. Endocrinology 144, 3742–3748. https://doi.org/10.1210/en.2003-0435 (2003).
Lowrey, P. L. & Takahashi, J. S. Mammalian circadian biology: Elucidating genome-wide levels of temporal organization. Annu. Rev. Genomics Hum. Genet. 5, 407–441. https://doi.org/10.1146/annurev.genom.5.061903.175925 (2004).
Wingfield, J. C. & Kenagy, G. J. Natural Regulation of Reproductive Cycles. In Vertebrate Endocrinology: Fundamentals and Biomedical Implications (eds Pang, P. K. T. & Schreibman, M. P.) (Academic Press, 1991).
Ko, C. H. & Takahashi, J. S. Molecular components of the mammalian circadian clock. Hum. Mol. Genet. 15, R271–R277. https://doi.org/10.1093/hmg/ddl207 (2006).
Pendergast, J. S., Niswender, K. D. & Yamazaki, S. Tissue-specific function of Period3 in circadian rhythmicity. PLoS ONE 7, e30254. https://doi.org/10.1371/journal.pone.0030254 (2012).
Guillaumond, F., Dardente, H., Giguère, V. & Cermakian, N. Differential control of Bmal1 circadian transcription by REV-ERB and ROR nuclear receptors. J. Biol. Rhythms 20, 391–403. https://doi.org/10.1177/0748730405277232 (2005).
Liedvogel, M., Szulkin, M., Knowles, S. C. L., Wood, M. J. & Sheldon, B. C. Phenotypic correlates of clock gene variation in a wild blue tit population: Evidence for a role in seasonal timing of reproduction. Mol. Ecol. 18, 2444–2456. https://doi.org/10.1111/j.1365-294X.2009.04204.x (2009).
Oishi, K., Fukui, H. & Ishida, N. Rhythmic expression of BMAL1 mRNA is altered in Clock mutant mice: Differential regulation in the suprachiasmatic nucleus and peripheral tissues. Biochem. Biophys. Res. Commun. 268, 164–171. https://doi.org/10.1006/bbrc.1999.2054 (2000).
DeBruyne, J. P. et al. Clock shock: Mouse CLOCK is not required for circadian oscillator function. Neuron 50, 465–477. https://doi.org/10.1016/j.neuron.2006.03.041 (2006).
King, D. P. et al. Positional cloning of the mouse circadian clock gene. Cell 89, 641–653. https://doi.org/10.1016/S0092-8674(00)80245-7 (1997).
Saleem, Q., Anand, A., Jain, S. & Brahmachari, S. The polyglutamine motif is highly conserved at the Clock locus in various organisms and is not polymorphic in humans. Hum. Genet. 109, 136–142. https://doi.org/10.1007/s004390100550 (2001).
Avivi, A. et al. Biological clock in total darkness: The Clock/MOP3 circadian system of the blind subterranean mole rat. Proc. Natl. Acad. Sci. U.S.A. 98, 13751–13756. https://doi.org/10.1073/pnas.181484498 (2001).
Hayasaka, N., LaRue, S. I. & Green, C. B. In vivo disruption of Xenopus CLOCK in the retinal photoreceptor cells abolishes circadian melatonin rhythmicity without affecting its production levels. J. Neurosci. 22, 1600–1607. https://doi.org/10.1523/jneurosci.22-05-01600.2002 (2002).
Alvarez, J. D. et al. The circadian clock protein BMAL1 is necessary for fertility and proper testosterone production in mice. J. Biol. Rhythms 23, 1–19. https://doi.org/10.1177/0748730407311254 (2008).
Dawson, A. Annual gonadal cycles in birds: modeling the effects of photoperiod on seasonal changes in GnRH-1 secretion. Front. Neuroendocrinol. 37, 52–64. https://doi.org/10.1016/j.yfrne.2014.08.004 (2015).
Hickok, J. R. & Tischkau, S. A. In vivo circadian rhythms in gonadotropin-releasing hormone neurons. Neuroendocrinology 91, 110–120. https://doi.org/10.1159/000243163 (2010).
Fidler, A. E. & Gwinner, E. Comparative analysis of Avian BMAL1 and CLOCK protein sequences: A search for features associated with owl nocturnal behaviour. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 136, 861–874. https://doi.org/10.1016/S1096-4959(03)00276-8 (2003).
Albrecht, U. & Ripperger, J. A. Clock Genes. In Encyclopedia of Neuroscience (eds Binder, M. D. et al.) (Springer, 2009).
Reppert, S. M. & Weaver, D. R. Coordination of circadian clocks in mammals. Nature 418, 935–941. https://doi.org/10.1038/nature00965 (2002).
Wilkins, A. K., Barton, P. I. & Tidor, B. The Per2 negative feedback loop sets the period in the mammalian circadian clock mechanism. PLOS Comput. Biol. 3, 2476–2486. https://doi.org/10.1371/journal.pcbi.0030242 (2007).
Carpen, J. D., Archer, S. N., Skene, D. J., Smits, M. & Von Schantz, M. A single-nucleotide polymorphism in the 5′-untranslated region of the hPER2 gene is associated with diurnal preference. J. Sleep Res. 14, 293–297. https://doi.org/10.1111/j.1365-2869.2005.00471.x (2005).
Toh, K. L. et al. An hPer2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 291, 1040–1043. https://doi.org/10.1126/science.1057499 (2001).
Eide, E. J. & Virshup, D. M. Casein kinase I: Another cog in the circadian clockworks. Chronobiol. Int. 18, 389–398. https://doi.org/10.1081/CBI-100103963 (2001).
Hirano, A. et al. A cryptochrome 2 mutation yields advanced sleep phase in humans. Elife 5, 1–21. https://doi.org/10.7554/eLife.16695 (2016).
Yu, X., Liu, H., Klejnot, J. & Lin, C. The Cryptochrome blue light receptors. Arabidopsis Book 8, e0135. https://doi.org/10.1199/tab.0135 (2010).
Vaidya, A. T. et al. Flavin reduction activates Drosophila cryptochrome. Proc. Natl. Acad. Sci. U.S.A. 110, 20455–20460. https://doi.org/10.1073/pnas.1313336110 (2013).
Hirano, A., Braas, D., Fu, Y.-H. & Ptacek, L. J. FAD regulates CRYPTOCHROME protein stability and circadian clock in mice. Cell Rep. 19(2), 255–266. https://doi.org/10.1016/j.celrep.2017.03.041 (2017).
Boden, M. J. & Kennaway, D. J. Reproductive consequences of circadian dysfunction: fertility in the Bmal1 null mouse. Reprod. Fertil. Dev. 16, 280. https://doi.org/10.1071/srb04abs280 (2004).
Boden, M. J. & Kennaway, D. J. Reproduction in the arrhythmic Bmal1 knockout mouse. Reprod. Fertil. Dev. 17, 126–126. https://doi.org/10.1071/SRB05Abs297 (2005).
Johnsen, A. et al. Avian Clock gene polymorphism: evidence for a latitudinal cline in allele frequencies. Mol. Ecol. 16, 4867–4880. https://doi.org/10.1111/j.1365-294X.2007.03552.x (2007).
O’Malley, K. G. & Banks, M. A. A latitudinal cline in the Chinook salmon (Oncorhynchus tshawytscha) Clock gene: Evidence for selection on PolyQ length variants. Proc. Royal Soc. B 275, 2813–2821. https://doi.org/10.1098/rspb.2008.0524 (2008).
O’Malley, K. G., Ford, M. J. & Hard, J. J. Clock polymorphism in Pacific salmon: Evidence for variable selection along a latitudinal gradient. Proc. Royal Soc. B 277, 3703–3714. https://doi.org/10.1098/rspb.2010.0762 (2010).
Kyriacou, C. P., Peixoto, A. A. & Costa, R. A cline in the Drosophila melanogaster period gene in Australia: Neither down nor under. J. Evol. Biol. 20, 1649–1651. https://doi.org/10.1111/j.1420-9101.2007.01352.x (2007).
Sawyer, L. A. et al. The period gene Thr-Gly polymorphism in Australian and African Drosophila melanogaster populations: Implications for selection. Genetics 174, 465–480. https://doi.org/10.1534/genetics.106.058792 (2006).
Kolaczkowski, B., Kern, A. D., Holloway, A. K. & Begun, D. J. Genomic differentiation between temperate and tropical Australian populations of Drosophila melanogaster. Genetics 187, 245–260. https://doi.org/10.1534/genetics.110.123059 (2011).
Friesen, V. L. et al. Sympatric speciation by allochrony in a seabird. Proc. Natl. Acad. Sci. U.S.A. 104, 18589–18594. https://doi.org/10.1073/pnas.0700446104 (2007).
Bolton, M. et al. Monteiro’s Storm-petrel Oceanodroma monteiroi: A new species from the Azores. Ibis 150, 717–727. https://doi.org/10.1111/j.1474-919X.2008.00854.x (2008).
Taylor, R. S. & Friesen, V. L. The role of allochrony in speciation. Mol. Ecol. 26, 3330–3342. https://doi.org/10.1111/mec.14126 (2017).
Taylor, R. S. et al. Sympatric population divergence within a highly pelagic seabird species complex (Hydrobates spp.). J. Avian Biol. https://doi.org/10.1111/jav.01515 (2018).
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Laboratory Press, 1989).
Team, R. C. (2020). R: A language and environment for statistical computing (3.6.2). R Foundation for Statistical Computing, Vienna, Austria. http://www.r-project.org/index.html
Kahle, D. & Wickham, H. ggmap: Spatial Visualization with ggplot2. R J. 5, 144–161 (2013).
Sin, S. Y. W., Hoover, B. A., Nevitt, G. A. & Edwards, S. V. Demographic history, not mating system, explains signatures of inbreeding and inbreeding depression in a large outbred population. Am. Nat. 197, 658–676. https://doi.org/10.1086/714079 (2021).
Bailey, M. J., Chong, N. W., Xiong, J. & Cassone, V. M. Chickens’ Cry2: Molecular analysis of an avian cryptochrome in retinal and pineal photoreceptors. FEBS Lett. 513, 169–174. https://doi.org/10.1016/S0014-5793(02)02276-7 (2002).
Hall, T. A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98 (1999).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649. https://doi.org/10.1093/bioinformatics/bts199 (2012).
Benson, D. A. et al. GenBank. Nucleic Acids Res. 41, D36–D42. https://doi.org/10.1093/nar/gks1195 (2013).
Deane, P. What Traits Predispose the Band-Rumped Storm-Petrel (Oceanodroma castro) to Ecological Speciation in the Absence of Physical Barriers to Gene Flow? (Queen’s University, 2011).
Excoffier, L., Laval, G. & Schneider, S. Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evol. Bioinform. 1, 117693430500100. https://doi.org/10.1177/117693430500100003 (2005).
Librado, P. & Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452. https://doi.org/10.1093/bioinformatics/btp187 (2009).
Slatkin, M. A measure of population subdivision based on microsatellite allele frequencies. Genetics 139, 457–462. https://doi.org/10.1093/genetics/139.1.457 (1995).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. JSTOR 57, 289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x (1995).
Gekakis, N. et al. Role of the CLOCK protein in the mammalian circadian mechanism. Science 280, 1564–1569. https://doi.org/10.1126/science.280.5369.1564 (1998).
Mantel, N. The detection of disease clustering and a generalized regression approach. Cancer Res. 27, 209–220 (1967).
Dor, R. et al. Low variation in the polymorphic Clock gene poly-Q region despite population genetic structure across barn swallow (Hirundo rustica) populations. PloS one 6, e28843. https://doi.org/10.1371/journal.pone.0028843 (2011).
Caprioli, M. et al. Clock gene variation is associated with breeding phenology and maybe under directional selection in the migratory barn swallow. PloS One 7, e35140. https://doi.org/10.1371/journal.pone.0035140 (2012).
Liedvogel, M. & Sheldon, B. C. Low variability and absence of phenotypic correlates of Clock gene variation in a great tit Parus major population. J. Avian Biol. 41, 543–550. https://doi.org/10.1111/j.1600-048X.2010.05055.x (2010).
Dor, R. et al. Clock gene variation in Tachycineta swallows. Ecol. Evol. 2, 95–105. https://doi.org/10.1002/ece3.73 (2012).
Chamary, J. V. & Hurst, L. D. Evidence for selection on synonymous mutations affecting stability of mRNA secondary structure in mammals. Genome Biol. https://doi.org/10.1186/gb-2005-6-9-r75 (2005).
Blom, N., Gammeltoft, S. & Brunak, S. Sequence and structure-based prediction of eukaryotic protein phosphorylation sites. J. Mol. Biol. 294, 1351–1362. https://doi.org/10.1006/jmbi.1999.3310 (1999).
Sahar, S., Zocchi, L., Kinoshita, C., Borrelli, E. & Sassone-Corsi, P. Regulation of BMAL1 protein stability and circadian function by GSK3β-mediated phosphorylation. PLoS One 5(1), e8561. https://doi.org/10.1371/journal.pone.0008561 (2010).
Sancar, A. Structure and function of DNA photolyase and cryptochrome blue-light photoreceptors. Chem. Rev. 103, 2203–2238. https://doi.org/10.1021/cr0204348 (2003).
Hsu, D. S. et al. Putative human blue-light photoreceptors hCRY1 and hCRY2 are flavoproteins. Biochemistry 35, 13871–13877. https://doi.org/10.1021/bi962209o (1996).
Kumar, V., Singh, B. P. & Rani, S. The bird clock: A complex, multi-oscillatory and highly diversified system. Biol. Rhythm Res. 35, 121–144. https://doi.org/10.1080/09291010412331313287 (2004).
Renthlei, Z., Gurumayum, T., Borah, B. K. & Trivedi, A. K. Daily expression of clock genes in central and peripheral tissues of tree sparrow (Passer montanus). Chronobiol. Int. 36, 110–121. https://doi.org/10.1080/07420528.2018.1523185 (2019).
Gaston, S. & Menaker, M. Pineal function: The biological clock in the sparrow?. Science 160, 1125–1127. https://doi.org/10.1126/science.160.3832.1125 (1968).
Kumar, V. & Gwinner, E. Pinealectomy shortens resynchronisation times of house sparrow (Passer domesticus) circadian rhythms. Naturwissenschaften 92, 419–422. https://doi.org/10.1007/s00114-005-0009-6 (2005).
McMillan, J. P. Pinealectomy abolishes the circadian rhythm of migratory restlessness. J. Comp. Physiol. 79, 105–112. https://doi.org/10.1007/BF00697766 (1972).
Rani, S., Singh, S. & Kumar, V. Photoperiodism, pineal clock and seasonal reproduction in the Indian Weaver Bird (Ploceus philippinus). J. Ornithol. 148, 601–610. https://doi.org/10.1007/s10336-007-0236-z (2007).
Rani, S., Singh, S., Malik, S., Singh, J. & Kumar, V. Synchronization of Indian weaver bird circadian rhythms to food and light zeitgebers: Role of pineal. Chronobiol. Int. 26, 653–665. https://doi.org/10.1080/07420520902926009 (2009).
Trivedi, A. K., Malik, S., Rani, S. & Kumar, V. Pinealectomy abolishes circadian behavior and interferes with circadian clock gene oscillations in brain and liver but not retina in a migratory songbird. Physiol. Behav. 156, 156–163. https://doi.org/10.1016/j.physbeh.2016.01.019 (2016).
Smith, A. L., Monteiro, L., Hasegawa, O. & Friesen, V. L. Global phylogeography of the band-rumped storm-petrel (Oceanodroma castro; Procellariiformes: Hydrobatidae). Mol. Phylogenet. Evol. 43, 755–773. https://doi.org/10.1016/j.ympev.2007.02.012 (2007).
Sun, Z., Gómez-Díaz, E., Bailie, A. & Friesen, V. Isolation and characterization of microsatellite loci for storm-petrels. Mol. Ecol. Resour. 9, 913–915. https://doi.org/10.1111/j.1755-0998.2008.02487.x (2009).
Noakes, M. A., Campbell, M. T. & Van Hest, B. J. The chicken CLOCK gene maps to chromosome 4. Anim. Genet. 31, 344–344. https://doi.org/10.1111/j.1365-2052.2000.00666.pp.x (2000).
Gene (NCBI)[Internet]. Bethesda (MD): National Library of Medicine (US), National Center for Biotechnology Information; [1988] –. Accession No. 374115, ARNTL aryl hydrocarbon receptor nuclear translocator like [ Gallus gallus (chicken) ]; [cited 2022 Sept 09]. Available from: https://www.ncbi.nlm.nih.gov/gene/374115
Huang, N. et al. Crystal structure of the heterodimeric CLOCK:BMAL1 transcriptional activator complex. Science 337, 189–194. https://doi.org/10.1126/science.1222804 (2012).
Gillessen, M., Kwak, P. B. & Tamayo, A. A simple method to measure CLOCK-BMAL1 DNA binding activity in tissue and cell extracts. F1000Research 6, 1316. https://doi.org/10.12688/f1000research.11685.2 (2017).
BirdLife International. IUCN Red List for birds. http://www.birdlife.org (2021).
Taylor, R. S. et al. Cryptic species and independent origins of allochronic populations within a seabird species complex (Hydrobates spp.). Mol. Phylogenet. Evol. 139, 106552. https://doi.org/10.1016/j.ympev.2019.106552 (2019).
AOU Classification Committee. North and Middle America. - Proposal Set 2016-C. http://checklist.aou.org/assets/proposals/PDF/2016-C.pdf (2016).
Yasuo, S. & Yoshimura, T. Comparative analysis of the molecular basis of photoperiodic signal transduction in vertebrates. Integr. Comp. Biol. 49, 507–518. https://doi.org/10.1093/icb/icp011 (2009).
Bentley, G. E. Unraveling the enigma: The role of melatonin in seasonal processes in birds. Microsc. Res. Tech. 53, 63–71. https://doi.org/10.1002/jemt.1069 (2001).
Yoshimura, T. et al. Molecular analysis of avian circadian clock genes. Mol. Brain Res. 78, 207–215. https://doi.org/10.1016/S0169-328X(00)00091-7 (2000).
Pinzon-Rodriguez, A., Bensch, S. & Muheim, R. Expression patterns of cryptochrome genes in avian retina suggest involvement of Cry4 in light-dependent magnetoreception. J. R. Soc. Interface 15, 20180058. https://doi.org/10.1098/rsif.2018.0058 (2018).
Bazzi, G. et al. Clock gene polymorphism and scheduling of migration: A geolocator study of the barn swallow Hirundo rustica. Sci. Rep. 5, 12443. https://doi.org/10.1038/srep12443 (2015).
Bourret, A. & Garant, D. Candidate gene–environment interactions and their relationships with timing of breeding in a wild bird population. Ecol. Evol. 5, 3628–3641. https://doi.org/10.1002/ece3.1630 (2015).
Saino, N. et al. Polymorphism at the Clock gene predicts phenology of long-distance migration in birds. Mol. Ecol. 24, 1758–1773. https://doi.org/10.1111/mec.13159 (2015).
Lugo Ramos, J. S., Delmore, K. E. & Liedvogel, M. Candidate genes for migration do not distinguish migratory and non-migratory birds. J. Comp. Physiol. A 203, 383–397. https://doi.org/10.1007/s00359-017-1184-6 (2017).
Bazzi, G. et al. Clock gene polymorphism, migratory behaviour and geographic distribution: A comparative study of trans-Saharan migratory birds. Mol. Ecol. 25, 6077–6091. https://doi.org/10.1111/mec.13913 (2016).
Gemayel, R., Vinces, M. D., Legendre, M. & Verstrepen, K. J. Variable Tandem repeats accelerate evolution of coding and regulatory sequences. Annu. Rev. Genet. 44, 445–477. https://doi.org/10.1146/annurev-genet-072610-155046 (2010).
Gemayel, R., Cho, J., Boeynaems, S. & Verstrepen, K. J. Beyond junk-variable tandem repeats as facilitators of rapid evolution of regulatory and coding sequences. Genes 3, 461–480. https://doi.org/10.3390/genes3030461 (2012).
Haerty, W. & Golding, G. B. Low-complexity sequences and single amino acid repeats: Not just “junk” peptide sequences. Genome 53, 753–762. https://doi.org/10.1139/G10-063 (2010).
Darlington, T. K. et al. Closing the circadian loop: CLOCK-induced transcription of its own inhibitors per and tim. Science 280, 1599–1603. https://doi.org/10.1126/science.280.5369.1599 (1998).
Legendre, P. Spatial autocorrelation: Trouble or new paradigm?. Ecology 74, 1659–1673. https://doi.org/10.2307/1939924 (1993).
Sangster, G. et al. Taxonomic recommendations for British birds: eighth report. Ibis 154, 874–883. https://doi.org/10.1111/j.1474-919X.2012.01273.x (2012).
Sauve, D., Dale, C. A., Tigano, A., Ratcliffe, L. M. & Friesen, V. L. Do candidate genes for migration and behavior explain migratory variation in bluebirds (Sialia spp)?. The Wilson J. Ornithol. 132, 820–829. https://doi.org/10.1676/19-00120 (2021).
Acknowledgements
Thank you to L. Colston-Nepali, B. Harkness, D. Sauve and A. Tigano, and for help with laboratory techniques and data analyses. T. Anker-Nilssen, T. Aarvak, M. Bolton, R. Furness, E. Goméz-Diaz, B. Keitt, L. Monteiro, V. Neves, J. Piatt, R. Pitman, J. Pitocchelli, P. Quillfeldt, A. Smith and Phil Unitt (SDNHM tissue collection) helped with sample collections. DNA sequencing was provided by the Genome Quebec Innovation Centre, and funding was provided by the Natural Science and Engineering Research Council (Discovery Grant to VLF), APEX Resource Management Solutions and Summer Work Experience Program (Queen’s University).
Author information
Authors and Affiliations
Contributions
K.B., T.B., E.E.C., H.G.D., R.S.T, & V.L.F. conceived and designed the experiments; Y.B.-G., P.D., V.L.F., J.F.M., & R.S.T. collected samples; D.A., K.B., T.B., E.E.C., H.G.D., B.A.S.H., & A.M. performed the experimental work and contributed to data analysis; K.B. performed the majority of the data analysis with significant contributions from H.G.D., R.S.T., & V.L.F.; K.B. drafted the manuscript and structured figures. All coauthors revised and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Birchard, K., Driver, H.G., Ademidun, D. et al. Circadian gene variation in relation to breeding season and latitude in allochronic populations of two pelagic seabird species complexes. Sci Rep 13, 13692 (2023). https://doi.org/10.1038/s41598-023-40702-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-40702-8
- Springer Nature Limited