Abstract
Idiopathic pulmonary fibrosis (IPF) is the most common and serious type of idiopathic interstitial pneumonia, characterized by chronic, progressive, and low survival rates, while unknown disease etiology. Until recently, patients with idiopathic pulmonary fibrosis have a poor prognosis, high mortality, and limited treatment options, due to the lack of effective early diagnostic and prognostic tools. Therefore, we aimed to identify biomarkers for idiopathic pulmonary fibrosis based on multiple machine-learning approaches and to evaluate the role of immune infiltration in the disease. The gene expression profile and its corresponding clinical data of idiopathic pulmonary fibrosis patients were downloaded from Gene Expression Omnibus (GEO) database. Next, the differentially expressed genes (DEGs) with the threshold of FDR < 0.05 and |log2 foldchange (FC)| > 0.585 were analyzed via R package “DESeq2” and GO enrichment and KEGG pathways were run in R software. Then, least absolute shrinkage and selection operator (LASSO) logistic regression, support vector machine-recursive feature elimination (SVM-RFE) and random forest (RF) algorithms were combined to screen the key potential biomarkers of idiopathic pulmonary fibrosis. The diagnostic performance of these biomarkers was evaluated through receiver operating characteristic (ROC) curves. Moreover, the CIBERSORT algorithm was employed to assess the infiltration of immune cells and the relationship between the infiltrating immune cells and the biomarkers. Finally, we sought to understand the potential pathogenic role of the biomarker (SLAIN1) in idiopathic pulmonary fibrosis using a mouse model and cellular model. A total of 3658 differentially expressed genes of idiopathic pulmonary fibrosis were identified, including 2359 upregulated genes and 1299 downregulated genes. FHL2, HPCAL1, RNF182, and SLAIN1 were identified as biomarkers of idiopathic pulmonary fibrosis using LASSO logistic regression, RF, and SVM-RFE algorithms. The ROC curves confirmed the predictive accuracy of these biomarkers both in the training set and test set. Immune cell infiltration analysis suggested that patients with idiopathic pulmonary fibrosis had a higher level of B cells memory, Plasma cells, T cells CD8, T cells follicular helper, T cells regulatory (Tregs), Macrophages M0, and Mast cells resting compared with the control group. Correlation analysis demonstrated that FHL2 was significantly associated with the infiltrating immune cells. qPCR and western blotting analysis suggested that SLAIN1 might be a signature for the diagnosis of idiopathic pulmonary fibrosis. In this study, we identified four potential biomarkers (FHL2, HPCAL1, RNF182, and SLAIN1) and evaluated the potential pathogenic role of SLAIN1 in idiopathic pulmonary fibrosis. These findings may have great significance in guiding the understanding of disease mechanisms and potential therapeutic targets in idiopathic pulmonary fibrosis.
Similar content being viewed by others
Introduction
Idiopathic pulmonary fibrosis (IPF) is a progressive, chronic, fibrotic interstitial lung disease with unknown etiology and poor prognosis1, 2. Idiopathic pulmonary fibrosis is characterized by lung tissue remodeling and scarring, decreased lung function, and decreased quality of life, which eventually leads to respiratory failure3,4,5,6,7. In the elderly, this disease is more prevalent and the average survival rate is only 2–3 years following diagnosis8. Despite the introduction of new antifibrotic therapies, idiopathic pulmonary fibrosis remains a fatal disease with a median survival of 5.7 years and limited treatment options9, 10. Therefore, it is urgent to find biomarkers for the treatment of idiopathic pulmonary fibrosis.
Machine learning is a data analysis method that automatically constructs analytical models, which has been widely used in clinical medicine11. Previous studies showed that machine learning can be used to predict myocardial infarction, identify pathologies, and improve surgical outcomes12. In recent years, machine learning has been widely used in the diagnosis and treatment of many diseases13,14,15. In addition, machine learning algorithms known as random forest (RF) models are made up of many individual decision trees that are constructed iteratively from random subsets of predictor and dependent variables16. The least absolute shrinkage and selection operator (LASSO) logistic regression analysis is a linear regression method with regularization that can be applied to high-dimensional analysis17. Support vector machine-recursive feature elimination (SVM-RFE) can be used to screen the best combinations of variables after modeling different numbers of variables due to its nonlinear discrimination characteristics18. Thus, using machine learning algorithms to identify biomarkers of idiopathic pulmonary fibrosis is of great significance.
In this study, we downloaded the gene expression profile and its corresponding clinical data of idiopathic pulmonary fibrosis patients from the GEO database. Based on this data, the DEGs with the threshold of FDR < 0.05 and |log2 foldchange (FC)|> 0.585 were identified between idiopathic pulmonary fibrosis and normal samples. Then, we combined least absolute shrinkage and selection operator (LASSO) logistic regression, support vector machine-recursive feature elimination (SVM-RFE), and random forest (RF) algorithms to screen out the biomarkers of idiopathic pulmonary fibrosis. The accuracy of these biomarkers was evaluated according to ROC curves. Moreover, we used the CIBERSORT algorithm to assess the infiltration of immune cells and the relationship between the infiltrating immune cells and the biomarkers. Finally, we assessed the potential pathogenic role of the biomarker (SLAIN1) in the development of idiopathic pulmonary fibrosis using mouse and cellular models.
Methods
Animals
C57BL6 mice aged 8 weeks were obtained from Jackson Laboratories, which were kept under specific pathogen-free (SPF) conditions in the animal barrier facility of Jiangxi University of Traditional Chinese Medicine. The study and all procedures were conducted in accordance with the ethical guidelines and regulations approved by the Ethics Committee of Jiangxi University of Traditional Chinese Medicine. We confirm that all methods were performed in accordance with the relevant guidelines and regulations (https://www.nature.com/srep/journal-policies/editorial-policies#experimental-subjects). Upon arrival, mice were acclimatized for 3 days before induction of idiopathic pulmonary fibrosis, and their health and wellbeing were monitored at least daily throughout the experiments. All experiments were conducted and reported according to ARRIVE guidelines (https://arriveguidelines.org/arrive-guidelines).
Cell culture
Human fetal lung fibroblast cell line (HFL1) and the human pulmonary epithelial A549 cell line were purchased from the American Type Culture Collection (ATCC). A549 lung cancer cells were cultured in DMEM medium supplemented with 2mMl glutamine and 10% fetal bovine serum (FBS). HFL1 cells were cultured with RPMI 1640 medium supplemented with 2mMl glutamine and 10% fetal bovine serum (FBS). Both cell lines were incubated in a humidified incubator with 5% CO2 at 37 °C and supplemented with 1% penicillin and 1% streptomycin. TGF-β was used to produce a cellular model of pulmonary fibrosis in A549 and HFL1 for 48 h.
Western blotting analysis
Collected lung tissue samples were cut into pieces, incubated with RIPA lysis buffer on ice for 40 min, and cleared by centrifugation at 13,000 rpm/min for 5 min. Protein concentrations were measured using the BCA Protein Assay kit (Beotime Institute of Biotechnology, Nanjing, China) according to the manufacturer's instructions. Then, 50 μg of protein from each sample was separated by 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to a polyvinylidene fluoride (PVDF) membrane. Membranes were blocked with 5% skim milk or 5% bovine serum albumin (BSA) and incubated overnight at 4 °C with the indicated concentrations of the following primary antibodies from Abcam (Cambridge, UK): Actin (Cat. No. ab179467; dilution, 1:5000; Abcam) and SLAIN-1 Polyclonal antibody (Cat no: 22123-1-AP; dilution, 1:1000; Protein Tech Group, Inc). This was followed by incubation with secondary antibodies (dilution, 1:5000–1:10,000; Abcam) at 37 °C for 1 h. The labeled protein bands were scanned on an HP Scanjet 5500 (Hewlett Packard France, Les Ullis, France). Finally, the relative protein concentration was determined by the gray level of the band with the Quantity One 4.40 software (Bio-Rad Laboratories Inc).
HE staining and Masson staining
Lung tissue was fixed with 4% polyformaldehyde solution for 24 h, dehydrated and paraffin embedded and cut into 4-μm slices. Then, the slices (3 slices per mouse) were stained with hematoxylin and eosin (H&E) to observe histopathological changes in the colon under a microscope. Masson trichrome staining was performed using Masson trichrome staining kit (Beyotime, China). Histological sections were stained with hematoxylin and eosin (H&E) to assess histopathological changes, and Masson trichrome staining was performed following the manufacturer's instructions.
RNA isolation and quantitative real-time PCR (qPCR)
RNA was extracted from lung homogenates or cultured cells using Qiagen RNeasy kits or Trizol (Vazyme) according to the manufacturer's instructions. Reverse transcription was performed using the HiScript○R II Q RT SuperMix Kit (Vazyme, Nanjing, China) as instructed by the manufacturer. SYBR green (Vazyme) and Quantstudio 6 Real-Time PCR System (Applied Biosystems) were used for qPCR. GeneCopoeia provided the primers that were used in this study. The normalizer employed in this study was GAPDH. The following primers were used for qPCR: Human GAPDH: 5′-GGA GCG AGA TCC CTC CAA AAT-3′ and 5′-GGC TGT TGT CAT ACT TCT CAT GG-3′;Human SLAIN1: 5′-CAT CAC CGG GAC AGC TTC AA-3′ and 5′-GAA CGG TTG GAC TCA CAT AGG-3′;Mouse GAPDH: 5′- AGG TCG GTG TGA ACG GAT TTG-3′ and 5′- TGT AGA CCA TGT AGT TGA GGT CA -3′;Mouse Slain1: 5′- ACT GAT GTT CAG ATC ATG GCT CG-3′ and 5′- ACT GCA TGT CCC CTT TTT CCC-3′.
Data acquisition
Data for idiopathic pulmonary fibrosis and healthy specimens were downloaded from the GEO dataset (GEO, https://www.ncbi.nlm.nih.gov/geo/database). The GSE150910 dataset, consisting of 103 idiopathic pulmonary fibrosis samples and 103 normal samples19, was utilized in this study. The clinical features of participants diagnosed with IPF and unaffected controls are summarized in Table 1. Age, gender, ethnicity, and smoking history showed no statistically significant disparities between healthy individuals and IPF patients in this cohort. The GSE110147 dataset contained 22 idiopathic pulmonary fibrosis samples and 11 normal samples20. DEGs are identified using the above GEO dataset.
DEGs screening, data processing, and DEG analysis
To remove batch effects from both datasets, we used the merge and combat functions of SVM to create metadata groups. We identified DEGs with the threshold of FDR < 0.05 and |log2 foldchange (FC)|> 0.585 between idiopathic pulmonary fibrosis and normal samples via package “DESeq2” in R software (version: 4.1.2). Volcano maps and heat maps of DEGs were visualized using the “pheatmap” (Version:1.0.12) and “ggplot2” (Version:3.4.2) software packages. In the volcano plot, DEGs with log2FC < 0 were considered to be down-regulated, while those with log2FC > 0 were considered to be up-regulated21. The heatmap of the expression of biomarkers are visualized using the “pheatmap” packages (Version:1.0.12).
Functional enrichment analysis
To explore the underlying mechanisms of DEGs in idiopathic pulmonary fibrosis, the “clusterProfiler” R package22 was used to perform the Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis23,24,25. Statistical significance was defined as FDR values less than 0.05 for both KEGG and GO enrichment analyses.
Candidate biomarker screening
In this study, three machine learning algorithms were utilized to screen out characteristic genes of idiopathic pulmonary fibrosis, including random forests (RF), least absolute shrinkage and selection operator (LASSO) logistic regression, and support vector machine-recursive feature elimination (SVM-RFE). It is reported that RF26, 27, LASSO logistic regression17, and SVM-RFE28 algorithms were of great significance to identify key biomarkers. In recent years, these three algorithms have been widely used in research to identify diagnostic or prognostic factors29,30,31,32,33,34. The RF algorithm was performed with package “randomForest” in R software (version: × 64 4.1.2); LASSO logistic regression analysis was carried out with package “glmnet” in R software (version: × 64 4.1.2); SVM-RFE algorithm was performed with package “e107” in R software (version: × 64 4.1.2). The potential biomarkers were yielded by intersecting the characteristic genes identified by RF, LASSO logistic regression, and SVM-RFE algorithms. Furthermore, the accuracy of the biomarkers was evaluated by the receiver operating characteristic (ROC) curve in the training set and test set.
Assays of immune cellular patterns in microenvironment
CIBERSORT is a deconvolution algorithm used to calculate the abundance of 22 types of infiltrated immune cells between the idiopathic pulmonary fibrosis group and the control group35. P less than 0.05 was considered statistically significant. Group comparisons were performed using the Wilcoxon rank sum test. The package “ggplot2” was employed to draw a violin plot for visualize the distinction of immune infiltrating cells. Furthermore, the “corrplot” package was used to visualize the correlation heat map of the relationship between 22 immune infiltrating cells.
Correlation analysis between biomarkers and infiltrating immune cells
The relationship of the biomarkers with the levels of immune infiltrating cells was explored using Spearman’s rank correlation analysis in R software (version: × 64 4.1.2). The “ggplot2” package was used to visualize the results. p values < 0.05 were considered statistically significant36.
Statistical analysis
All statistical analyses were performed using the R software 4.1.2 and GraphPadPrism 8. RF analysis was conducted using the “RandomForest” Package, the LASSO Cox regression was undertaken by the “glmnet” package, and the SVM analysis was done by using the “e1071” R package. ROC curves were used to estimate the diagnostic accuracy of the cancer markers. Spearman's rank correlation test was used to establish the significance of correlation between the expression of genes and the infiltration of immune cells. The wilcoxon test identifies the significance of any differences between the two groups. The statistically significant difference was defined as the p value being less than 0.05.
Ethics statement
The animal study was reviewed and approved by the Research Ethics Committee of Jiangxi University of Traditional Chinese Medicine.
Results
Identification of differentially expressed genes in idiopathic pulmonary fibrosis
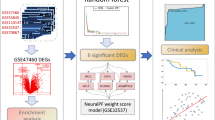
The workflow of this study is illustrated in Fig. 1. A total of 103 normal samples and 103 idiopathic pulmonary fibrosis samples from GSE150910 were used to evaluate the differences between the two samples, including 2359 upregulated genes and 1299 downregulated genes, by identification using the “DESeq2” package (Fig. 2A). The heatmap of these differentially expressed genes was shown in Fig. 2B.
Identification of differentially expressed genes between idiopathic pulmonary fibrosis and normal samples. (A) Volcano plot of the GSE150910 dataset with the cut-off criteria of |log2FC|> 0.585 and FDR < 0.05. (B) Heatmap visualization of the DEGs between idiopathic pulmonary fibrosis and normal samples.
Functional correlation analysis
To further explore the potential biological functions of DEGs of idiopathic pulmonary fibrosis in human, we constructed GO and KEGG enrichment analysis The results of GO enrichment analysis suggested that differentially expressed genes were mainly enriched in external encapsulating structure organization, extracellular matrix organization, and extracellular structure organization in the biological process aspect (Figs. 3A,B). In the aspect of cellular components, these differentially expressed genes were mainly involved in collagen–containing extracellular matrix, motile cilium, and axoneme. In the aspect of molecular function, these differentially expressed genes were mainly gathered in signaling receptor activator activity, receptor ligand activity, and extracellular matrix structural constituent. KEGG pathway enrichment analysis revealed that the differentially expressed genes were mainly enriched in 15 pathways, such as Neuroactive ligand–receptor interaction, Cytokine–cytokine receptor interaction, and Viral protein interaction with cytokine and cytokine receptor (Figs. 3C,D). These results suggest that extracellular matrix plays an important role in idiopathic pulmonary fibrosis.
Identification and assessment of biomarkers
Three validated machine learning algorithms (LASSO, RF, SVM-RFE) were applied to identify key characteristic genes associated with IPF. 46 characteristic genes were identified using LASSO algorithm (Fig. 4A). 60 characteristic genes were screened out using RF algorithm (Fig. 4B). Moreover, 34 characteristic genes were identified as biomarkers based on SVM-RFE algorithm (Fig. 4C). Only the overlapping genes (FHL2, HPCAL1, RNF182 and SLAIN1) were ultimately selected as biomarkers of IPF (Fig. 4D). In addition, the selected biomarkers showed good differential expression in the training set and test set, the expression levels of FHL2 were elevated in the idiopathic pulmonary fibrosis group (Fig. 5A), while the expression level of SLAIN1, HPCAL1, and RNF182 were reduced in the idiopathic pulmonary fibrosis group (Fig. 5B–D). As indicated by the differential analysis in the test set, the results were consistent with the outcomes of the training set (Fig. 6A–D). The heatmap of these four biomarkers in the training set and test set were shown in Fig. 7A–B, which suggested that the expression level of FHL2 was correlated with IPF group positively, the expression level of SLAIN1, HPCAL1, and RNF182 were correlated with IPF group negatively.
Identification of biomarkers of idiopathic pulmonary fibrosis. (A) Characteristic genes selection via LASSO algorithm. (B) Characteristic genes selection via random forest algorithm. (C) Characteristic genes selection via SVM-RFE algorithm. (D) Venn diagram showed the intersection of characteristic genes obtained by the three indicated algorithms. The overlapping characteristic genes represent the biomarkers of idiopathic pulmonary fibrosis.
Diagnostic effectiveness of biomarkers in idiopathic pulmonary fibrosis
To further assess the diagnostic value of the identified genes in idiopathic pulmonary fibrosis, ROC analysis was conducted for the four key genes in both the training and test sets. The results demonstrated that the four diagnostic biomarkers, as identified by the machine learning algorithm, exhibited strong diagnostic capabilities in the training set. FHL2 had an AUC of 0.954 (95% CI 0.924–0.978), HPCAL1 had an AUC of 0.955 (95% CI 0.926–0.979), RNF182 had an AUC of 0.917 (95% CI 0.875–0.955), and SLAIN1 had an AUC of 0.916 (95% CI 0.874–0.954) (Supplementary Fig. 1A–D). Furthermore, the diagnostic effectiveness of these biomarkers was validated in the independent test set, with FHL2 achieving an AUC of 0.926 (95% CI 0.822–0.992), HPCAL1 an AUC of 1.000 (95% CI 1.000–1.000), RNF182 an AUC of 0.946 (95% CI 0.843–1.000), and SLAIN1 an AUC of 1.000 (95% CI: 1.000–1.000) (Supplementary Fig. 2A–D). As illustrated in the figures above, all four genes exhibited strong discriminatory ability for idiopathic pulmonary fibrosis.
Evaluation of SLAIN1 expression in vivo and in vitro
To ensure the robustness of our findings, we initiated our investigation by establishing a murine model and a cellular model. As shown in Fig. 8A, Masson and HE staining revealed that the pulmonary fibrosis in bleomycin-treated mice lung tissues was notably more severe than in the PBS-treated mice lung tissues, thus affirming the successful construction of the mouse model. Subsequently, our attention turned to examining the expression level of SLAIN1, which was found to be significantly lower in the IPF samples compared to the normal samples (Fig. 8B,C).
Validation the role of SLAIN1 in vivo and vitro. (A) Photomicrographs of PBS-treated lung sections and Bleomycin-treated lung sections stained with Masson staining and HE staining, respectively. (B) Quantification of mRNA expression levels of SLAIN1 in the mouse model. (C) Western blot for SLAIN1 in the mouse model. Bleomycin-treated group was the experimental group, and PBS-treated group was the control group. (D) Quantification of SLAIN1 expression level in the A549 cells. (E) Western blot of SLAIN1 in A549 cells over time with fibrosis. (F) Quantification of SLAIN1 expression level in the HFL1 cells. (G) Western blot of SLAIN1 in HFL1 cells over time with fibrosis.
Furthermore, we extended our investigation to evaluation the expression level of SLAIN1 in A549 and HFL1 cells. To replicate the conditions of IPF in vitro, we exposed A549 and HFL1 cells to TGF-β. As depicted in Fig. 8D,F, the mRNA expression level of SLAIN1 in A549 and HFL1 cells was substantially reduced. Moreover, western blotting demonstrated a gradual decrease in the expression level of SLAIN1 in A549 and HFL1 cells (Fig. 8E,G). In summary, our comprehensive examination of SLAIN1 expression both in vivo and in vitro strengthens our understanding of its potential role as a biomarker and its involvement in the pathogenesis of idiopathic pulmonary fibrosis.
Immune infiltration
The infiltration status of 22 types of immune cells between idiopathic pulmonary fibrosis group and control group were assessed with CIBERSORT algorithm. The percentage of the 22 types of immune cells between idiopathic pulmonary fibrosis group and control group was shown in the bar plot (Supplementary Fig. 3A). The correlation of 22 types of immune cells revealed that T cells follicular helper was positively related with Plasma cells (r = 0.41), NK cells resting was positively related with T cells CD4 naive (r = 0.37), whereas T cells follicular helper was negatively related to T cells CD4 memory resting (r =− 0.51), NK cells resting was positively related with T cells follicular helper (r =− 0.39) (Supplementary Fig. 3B). The violin plot of the immune cell infiltration difference demonstrated that patients with idiopathic pulmonary fibrosis had a higher level of B cells memory, Plasma cells, T cells CD8, T cells follicular helper, T cells regulatory (Tregs), Macrophages M0, and Mast cells resting compared with the control group (Supplementary Fig. 3C).
Correlation analysis between biomarkers and immune cells
As indicated from the correlation analysis, SLAIN1 displayed a positive correlation with Eosinophils (r = 0.26, p < 0.001), Monocytes (r = 0.47, p < 0.001), Neutrophils (r = 0.22, p < 0.05), NK cells resting (r = 0.49, p < 0.001), T cells CD4 memory resting (r = 0.45, p < 0.001), and T cells CD4 naïve (r = 0.2, p < 0.05), and a negative correlation with B cells memory (r =− 0.28, p < 0.001), B cells naïve (r =− 0.2, p < 0.05), Macrophages M0 (r =− 0.19, p < 0.05), Mast cells activated (r =− 0.18, p < 0.05), NK cells activated (r =− 0.15, p < 0.05), Plasma cells (r =− 0.53, p < 0.001), T cells CD8(r =− 0.15, p < 0.05), T cells follicular helper (r =− 0.43, p < 0.001), and T cells regulatory (Tregs) (r =− 0.39, p < 0.001) (Supplementary Fig. 4A–J). It can be concluded that SLAIN1 was correlated with immune cells.
Discussion
When the lung sustains injury, the fibrosis process begins and the disease continues to progress37, 38. Idiopathic pulmonary fibrosis is a lethal, progressive fibrosing parenchymal lung disease that affects millions of patients worldwide and is refractory to most treatment options39. Fibrosis may play an important role in the later development of the disease, which may be the result of the interaction between multiple pathogenic factors40. Potentially powerful noninvasive biomarkers can provide critical diagnostic information while also being critical to understanding the course of IPF41. Patients at increased risk for IPF may benefit from clinical trials that allow physicians to select credible fibrosis biomarkers42.
In this study, we downloaded two GSE datasets from the GEO database to identify differentially expressed genes between IPF and normal lung tissues. Next, GO and KEGG analyses were then performed to explore the biological functions of DEGs in IPF. Combined with least absolute shrinkage and selection operator (LASSO) logistic regression, support vector machine-recursive feature elimination (SVM-RFE), and random forest (RF) algorithms, four biomarkers for idiopathic pulmonary fibrosis were screened out. The receiver operating characteristic (ROC) curve was calculated to additionally evaluate the diagnostic accuracy of biomarkers. Moreover, the CIBERSORT algorithm was employed to assess the infiltration of immune cells and the relationship between the infiltrating immune cells and the biomarkers. Finally, we detected the expression level of SLAIN1 to assess its potential role in the pathogenesis of idiopathic pulmonary fibrosis using a mouse model and cellular model.
As for the four biomarkers, previous studies showed that FHL2 has been identified as a biomarker of lung cancer and elevated level of FHL2 exacerbates the outcome of patients with non-small cell lung cancer (NSCLC) and the malignant phenotype in NSCLC cells43. Among visinin-like proteins (VILIPs), HPCAL1 belongs to the neuronal calcium sensor (NCS) family, which is responsible for calcium signaling in neurons44. Innate immune responses triggered by TLRs are inhibited by RNF182 by promoting degradation of p65 via K48-linked ubiquitination45. SLAIN1, indispensable for neuronal sprouting and brain development, significantly contributes to axon elongation by promoting microtubule growth46. Recent findings have suggested that SLAIN1 may play a pivotal role in intellectual disability47. Moreover, both in the training set and test set, the prognostic value of these four biomarkers were higher. Therefore, the model we established has excellent accuracy both in the training set and test set. Taken together, these four biomarkers are expected to be potential targets for the diagnosis of idiopathic pulmonary fibrosis.
Furthermore, we confirmed the expression level of SLAIN1 in the pathogenesis of idiopathic pulmonary fibrosis using both a mouse model. Masson and HE staining showed that the pulmonary fibrosis of bleomycin-treated mice lung tissues was more severe than PBS-treated mice lung tissues, which demonstrated that the mouse model was successfully constructed. The SLAIN1 expression level in A549 and HFL1 cells was significantly lower, which indicated that our model was equally successful in cell validation. Based on the above results, we can affirm that our findings bolster the reliability of the prognostic model.
Despite these advantages, this study has some limitations. The molecular mechanisms underlying the identified biomarkers in idiopathic pulmonary fibrosis (IPF) have not been fully investigated, warranting further experimental exploration. Moreover, as clinical patients were not included in this study, the diagnostic potential of the identified genes for IPF was indirectly assessed. Additional prospective studies are necessary to validate and translate these findings into clinical practice.
Conclusion
In summary, this research successfully pinpointed four promising biomarkers (FHL2, HPCAL1, RNF182, and SLAIN1) and investigated the potential involvement of SLAIN1 in the pathogenesis of idiopathic pulmonary fibrosis. These discoveries hold substantial importance in advancing our comprehension of the disease's mechanisms and identifying potential avenues for therapeutic intervention in idiopathic pulmonary fibrosis.
Data availability
The GSE150910 and GSE110147 datasets used in current study are available in the GEO repository (http://www.ncbi.nlm.nih.gov/geo/). Further inquiries can be directed to the corresponding author.
References
Herazo-Maya, J. D. et al. Peripheral blood mononuclear cell gene expression profiles predict poor outcome in idiopathic pulmonary fibrosis. Sci. Transl. Med. 5(205), 205ra136. https://doi.org/10.1126/scitranslmed.3005964 (2013).
Li, F. J. et al. Citrullinated vimentin mediates development and progression of lung fibrosis. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aba2927 (2021).
Quinn, C., Wisse, A. & Manns, S. T. Clinical course and management of idiopathic pulmonary fibrosis. Multidiscip. Respir. Med. 14(1), 1–9 (2019).
Kreuter, M. et al. The clinical course of idiopathic pulmonary fibrosis and its association to quality of life over time: Longitudinal data from the INSIGHTS-IPF registry. Respir. Res. 20(1), 1–13 (2019).
Kreuter, M. et al. Health related quality of life in patients with idiopathic pulmonary fibrosis in clinical practice: Insights-IPF registry. Respir. Res. 18(1), 1–10 (2017).
Martinez, F. J. et al. Idiopathic pulmonary fibrosis. Nat. Rev. Dis. Primers 3(1), 1–19 (2017).
van Manen, M. J., Geelhoed, J. M., Tak, N. C. & Wijsenbeek, M. S. Optimizing quality of life in patients with idiopathic pulmonary fibrosis. Ther. Adv. Respir. Dis. 11(3), 157–169 (2017).
Sgalla, G., Biffi, A. & Richeldi, L. Idiopathic pulmonary fibrosis: Diagnosis, epidemiology and natural history. Respirology 21(3), 427–437 (2016).
Jaeger, B. et al. Airway basal cells show a dedifferentiated KRT17(high)Phenotype and promote fibrosis in idiopathic pulmonary fibrosis. Nat. Commun. 13(1), 5637. https://doi.org/10.1038/s41467-022-33193-0 (2022).
Raghu, G. et al. An official ATS/ERS/JRS/ALAT statement: Idiopathic pulmonary fibrosis: evidence-based guidelines for diagnosis and management. Am. J. Resp. Crit. Care Med. 183(6), 788–824. https://doi.org/10.1164/rccm.2009-040GL (2011).
Deo, R. C. Machine learning in medicine. Circulation 132(20), 1920–1930. https://doi.org/10.1161/CIRCULATIONAHA.115.001593 (2015).
Rajkomar, A., Dean, J. & Kohane, I. Machine learning in medicine. N. Engl. J. Med. 380(14), 1347–1358 (2019).
Furukawa, T. et al. A comprehensible machine learning tool to differentially diagnose idiopathic pulmonary fibrosis from other chronic interstitial lung diseases. Respirology 27(9), 739–746 (2022).
Choi, Y. et al. Identification of usual interstitial pneumonia pattern using RNA-Seq and machine learning: Challenges and solutions. BMC Genomics 19(2), 147–159 (2018).
Pan, J. et al. Unsupervised machine learning identifies predictive progression markers of IPF. Eur. Radiol. 33(2), 925–935 (2022).
Romulo, C. L. et al. Global state and potential scope of investments in watershed services for large cities. Nat. Commun. 9(1), 4375. https://doi.org/10.1038/s41467-018-06538-x (2018).
Tibshirani, R. Regression shrinkage and selection via the lasso. J. Roy. Stat. Soc.: Ser. B (Methodol.) 58(1), 267–288 (1996).
Wu, L. D. et al. Analysis of potential genetic biomarkers using machine learning methods and immune infiltration regulatory mechanisms underlying atrial fibrillation. BMC Med. Genomics 15(1), 64. https://doi.org/10.1186/s12920-022-01212-0 (2022).
Furusawa, H. et al. Chronic hypersensitivity pneumonitis, an interstitial lung disease with distinct molecular signatures. Am. J. Resp. Crit. Care Med. 202(10), 1430–1444. https://doi.org/10.1164/rccm.202001-0134OC (2020).
Cecchini, M. J., Hosein, K., Howlett, C. J., Joseph, M. & Mura, M. Comprehensive gene expression profiling identifies distinct and overlapping transcriptional profiles in non-specific interstitial pneumonia and idiopathic pulmonary fibrosis. Resp. Res. 19(1), 153. https://doi.org/10.1186/s12931-018-0857-1 (2018).
Feng, H. et al. Identification of significant genes with poor prognosis in ovarian cancer via bioinformatical analysis. J. Ovarian Res. 12(1), 35. https://doi.org/10.1186/s13048-019-0508-2 (2019).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16(5), 284–287. https://doi.org/10.1089/omi.2011.0118 (2012).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28(11), 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51(D1), D587–D592. https://doi.org/10.1093/nar/gkac963 (2023).
Strobl, C., Boulesteix, A. L., Zeileis, A. & Hothorn, T. Bias in random forest variable importance measures: Illustrations, sources and a solution. BMC Bioinform. 8, 25. https://doi.org/10.1186/1471-2105-8-25 (2007).
Wang, H., Yang, F. & Luo, Z. An experimental study of the intrinsic stability of random forest variable importance measures. BMC Bioinform. 17, 60. https://doi.org/10.1186/s12859-016-0900-5 (2016).
Suykens, J. A. & Vandewalle, J. Least squares support vector machine classifiers. Neural Process. Lett. 9(3), 293–300 (1999).
Tang, J. et al. Computational advances of tumor marker selection and sample classification in cancer proteomics. Comput. Struct. Biotechnol. J. 18, 2012–2025 (2020).
Yu, F., Wei, C., Deng, P., Peng, T. & Hu, X. Deep exploration of random forest model boosts the interpretability of machine learning studies of complicated immune responses and lung burden of nanoparticles. Sci. Adv. https://doi.org/10.1126/sciadv.abf4130 (2021).
Ma, X., Su, J., Wang, B. & Jin, X. Identification of characteristic genes in whole blood of intervertebral disc degeneration patients by weighted gene coexpression network analysis (WGCNA). Comput. Math. Meth. Med. https://doi.org/10.1155/2022/6609901 (2022).
Zhang, Y. et al. Identifying discriminative features for diagnosis of Kashin–Beck disease among adolescents. BMC Musculoskelet. Disord. 22(1), 1–10 (2021).
Jubair, S., Alkhateeb, A., Tabl, A. A., Rueda, L. & Ngom, A. A novel approach to identify subtype-specific network biomarkers of breast cancer survivability. Netw. Model. Anal. Health Inform. Bioinform. 9(1), 1–12 (2020).
Wang, H., Lengerich, B. J., Aragam, B. & Xing, E. P. Precision Lasso: accounting for correlations and linear dependencies in high-dimensional genomic data. Bioinformatics 35(7), 1181–1187 (2019).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12(5), 453–457. https://doi.org/10.1038/nmeth.3337 (2015).
Ito, K. & Murphy, D. Application of ggplot2 to pharmacometric graphics. CPT Pharmacom. Syst. Pharmacol. 2, e79. https://doi.org/10.1038/psp.2013.56 (2013).
Moss, B. J., Ryter, S. W. & Rosas, I. O. Pathogenic mechanisms underlying idiopathic pulmonary fibrosis. Annu. Rev. Pathol. 17, 515–546. https://doi.org/10.1146/annurev-pathol-042320-030240 (2022).
Spagnolo, P. et al. Idiopathic pulmonary fibrosis: Disease mechanisms and drug development. Pharmacol. Ther. 222, 107798. https://doi.org/10.1016/j.pharmthera.2020.107798 (2021).
Al-Tamari, H. M. et al. FoxO3 an important player in fibrogenesis and therapeutic target for idiopathic pulmonary fibrosis. EMBO Mol. Med. 10(2), 276–293. https://doi.org/10.15252/emmm.201606261 (2018).
Richeldi, L., Collard, H. R. & Jones, M. G. Idiopathic pulmonary fibrosis. Lancet 389(10082), 1941–1952. https://doi.org/10.1016/S0140-6736(17)30866-8 (2017).
Stainer, A. et al. Molecular biomarkers in idiopathic pulmonary fibrosis: State of the art and future directions. Int. J. Mol. Sci. https://doi.org/10.3390/ijms22126255 (2021).
Maher, T. M. et al. An epithelial biomarker signature for idiopathic pulmonary fibrosis: An analysis from the multicentre PROFILE cohort study. Lancet Respir. Med. 5(12), 946–955. https://doi.org/10.1016/S2213-2600(17)30430-7 (2017).
Li, N., Xu, L., Zhang, J. & Liu, Y. High level of FHL2 exacerbates the outcome of non-small cell lung cancer (NSCLC) patients and the malignant phenotype in NSCLC cells. Int. J. Exp. Pathol. 103(3), 90–101. https://doi.org/10.1111/iep.12436 (2022).
Burgoyne, R. D., Helassa, N., McCue, H. V. & Haynes, L. P. Calcium sensors in neuronal function and dysfunction. Cold Spring Harbor Perspect. Biol. 11(5), a035154 (2019).
Cao, Y., Sun, Y., Chang, H., Sun, X. & Yang, S. The E3 ubiquitin ligase RNF182 inhibits TLR-triggered cytokine production through promoting p65 ubiquitination and degradation. FEBS Lett. 593(22), 3210–3219. https://doi.org/10.1002/1873-3468.13583 (2019).
D’anca, M., Buccellato, F. R., Fenoglio, C. & Galimberti, D. Circular RNAs: Emblematic players of neurogenesis and neurodegeneration. Int. J. Mol. Sci. 23(8), 4134 (2022).
Harripaul, R. et al. Mapping autosomal recessive intellectual disability: Combined microarray and exome sequencing identifies 26 novel candidate genes in 192 consanguineous families. Mol. Psychiatry 23(4), 973–984. https://doi.org/10.1038/mp.2017.60 (2018).
Acknowledgements
The authors thank the contributors to the GEO database for sharing date.
Funding
This work was supported by grants from the National Natural Science Foundation of China (No. 81860826, No.82260912), the Natural Science Foundation of Jiangxi Province (No.20212BAB216061, No.20212BAG70038, No.20192ACBL20023, No.20212BAB206061), Jiangxi University of Chinese Medicine Science and Technology innovation Team Development Program (xiao fa [2022]No.50), Jiangxi Clinical Medical Research Center of Lung Diseases (Gan ke fa she [2019] No. 44), Jiangxi Provincial Key Laboratory of Traditional Chinese Medicine Epidemic Diseases (Jiangxi Traditional Chinese Medicine Science and Education Zi [2021] No. 3),the Effect and Molecular Mechanism of Wenfei Huaxian Decoction on the Exosomes of Lung Mesenchymal Stem Cells (No. 81860826), the New Drug Research of Wenfei Huaxian Decoction on the Treatment of Pulmonary Fibrosis (No. KJD13063), Clinical Study on the Treatment of COVID-19 with Traditional Chinese Medicine (No. 2020YBBGW0003), Provincial Department of Education science and technology research project (No. GJJ190130).
Author information
Authors and Affiliations
Contributions
S.K., L.L., and H.C.: Conceptualization and methodology. S.K., L.M., Z.W. Software and data management. G.Z. M.Q., W.Z., and H.C.: Validation. L.L., Z.W., and L.L.: Reviewed and edited the manuscript. L.L. was the guarantor of this work and therefore had full access to all data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis. All authors listed made substantial, direct, intellectual contributions to this work and approved its publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, Z., Chen, H., Ke, S. et al. Identifying potential biomarkers of idiopathic pulmonary fibrosis through machine learning analysis. Sci Rep 13, 16559 (2023). https://doi.org/10.1038/s41598-023-43834-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-43834-z
- Springer Nature Limited
This article is cited by
-
Single-cell RNA sequencing reveals special basal cells and fibroblasts in idiopathic pulmonary fibrosis
Scientific Reports (2024)