Abstract
The existing biomarkers are insufficient for predicting the prognosis of pancreatic ductal adenocarcinoma (PDAC). Intraductal papillary mucinous neoplasm (IPMN) is a precursor to PDAC; therefore, identifying biomarkers from differentially expressed genes (DEGs) of PDAC and IPMN is a new and reliable strategy for predicting the prognosis of PDAC. In this study, four datasets were downloaded from the Gene Expression Omnibus database and standardized using the R package ‘limma.’ A total of 51 IPMN and 81 PDAC samples were analyzed, and 341 DEGs in PDAC and IPMN were identified; DEGs were involved in the extracellular matrix and tumor microenvironment. An acceptable survival prognosis was demonstrated by SDC1 and ITGA2, which were highly expressed during in vitro PDAC cell proliferation, apoptosis, and migration. SDC1high was enriched in interferon alpha (IFN-α) response and ITGA2high was primarily detected in epithelial-mesenchymal transition (EMT), which was verified using western blotting. We concluded that SDC1 and ITGA2 are potential prognostic biomarkers for PDAC associated with IPMN. Downregulation of SDC1 and ITGA2 expression in PDAC occurs via a mechanism involving possible regulation of IFN-α response, EMT, and immunity, which may act as new targets for PDAC therapy.
Similar content being viewed by others
Introduction
Pancreatic cancer (PC) is a type of malignant tumor with one of the highest mortality rates. According to the latest epidemiological data, 64,050 new cases of PC and 50,550 new cancer-related deaths have been recorded worldwide in 20231. Pancreatic ductal adenocarcinoma (PDAC) is the most common type of PC2. The 5-year survival rate of patients with PDAC is less than 10%3. Development of an effective treatment for PC has become a major clinical challenge. For traditional treatments, such as radiotherapy and chemotherapy, early (or late) drug resistance in patients with PDAC is always a concern4. Similarly, immunotherapy is being increasingly used for PDAC treatment, but significant challenges have been encountered because of the special tumor microenvironment (TME), which makes it difficult for immune cells to infiltrate and activate5. The identification of novel prognostic biomarkers will provide a strategy for the prevention of PDAC and identification of newer therapeutic targets.
The forward shift of the treatment threshold for PDAC is currently advocated as an effective and prudent move, and effective prognostic markers can demonstrate the clinical value in disease prevention. With the rapid development of imaging technology, the diagnosis of precancerous lesions associated with PC has become easier, which has focused research efforts on these precancerous lesions. Intraductal papillary mucinous neoplasm (IPMN), a precancerous lesion, is considered the most common type of pancreatic cystic tumor6. First discovered and described by Ohashi in 1982, it is the second most common pancreatic tumor7. IPMN is a rare tumor that originates from the epithelium8. It is a mucin-producing tumor that secretes a large amount of mucus and forms pancreatic cysts, which can gradually become malignant through the ‘hyperplasia-adenoma-cancer’ pathway9,10. There are three main types of IPMN: the main pancreatic duct type (MD-IPMN), branch-duct type (BD-IPMN), and mixed type11. Among them, the incidence of malignancy in MD-IPMN ranges from 57–92%12. Before IPMN develops into PDAC, there must be differences in gene expression between tissues.
The development of bioinformatics analysis and public databases13,14 can facilitate an understanding of the molecular mechanisms underlying IPMN and PDAC. The data of these platforms provide support for searching effective prognostic biomarkers and therapeutic targets. COL10A1, mediated by noncoding RNA, indicates that the prognosis of PC is poor. It involves mechanisms related to immunity15. In addition, we previously established seven immune-related miRNAs, including TNFSF9, TNFRSF9, KIR3DL1, and HAVCR2, as better prognostic models for PDAC16. Wang et al.17 studied miRNAs in the cystic fluid of low-grade benign and highly invasive pancreatic cystic lesions by sequencing and identified that miRNAs, such as miR-216a and miR-217, may be early detection biomarkers for pancreatic cystic lesions developing into PDAC. Early PDAC detection is associated with better survival18. Given the difficulty of diagnosis and treatment of PDAC, the identification of PDAC prognostic markers from precancerous and non-invasive lesions will be a reliable strategy; bioinformatics analysis makes this identification easier. The differentially expressed genes (DEGs) between IPMN and PDAC can be identified using bioinformatics programs19,20. New prognostic biomarkers are identified to determine whether they have predictive value for the survival of PDAC. Thus, identifying survival-related biomarkers of PDAC from these DEGs is a reliable strategy for establishing new prognostic biomarkers.
To identify potential biomarkers and predict the prognosis of patients with PDAC, we conducted bioinformatics analyses to identify DEGs between IPMN and PDAC samples. We then analyzed the DEGs using enrichment analysis to explore the possible mechanisms underlying the transformation from IPMN to PDAC. We further screened hub genes using bioinformatics methods. A survival analysis of the hub genes was performed to screen out the survival-related biomarkers for PDAC. We studied their clinical value based on public data. Furthermore, we verified their effects on the biological behavior of PDAC cells in vitro. Then, we performed the preliminary exploration and verification of the internal mechanisms. Our results indicated that SDC1 and ITGA2 are potential prognostic biomarkers of PDAC.
Results
Identification of DEGs
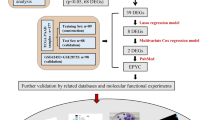
The flowchart is shown in Fig. 1. A total of 132 samples (51 IPMN and 81 PDAC) were included in this study. In total, 341 DEGs were identified (Fig. 2A). As shown in Fig. 2B, 230 genes were upregulated, and 111 were downregulated.
Flow chart of biomarker identification and validation. PDAC Pancreatic ductal adenocarcinoma; IPMN Intraductal papillary mucinous neoplasm; GEO DEGs Differentially expressed genes; GO Gene Ontology; KEGG Kyoto Encyclopedia of Genes and Genomes; PPI Protein–protein interaction; K–M Kaplan–Meier; GSEA Gene set enrichment analysis; ROC Receiver operating characteristics; AUC Area under the curve; qRT-PCR Quantitative real-time polymerase chain reaction.
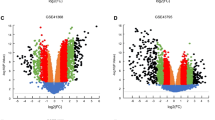
Identification of DEGs between PDAC and IPMN. (A) The heat map illustrates the differential expression of 341 DEGs between IPMN and PDAC samples identified with the thresholds of |log2 fold-change|> 0.5 and adjusted P-value < 0.05. (B) Volcano plots of DEGs. Colors represent the expression level of the genes, as follows: red, upregulated; green, downregulated; and gray, not statistically significant. PDAC Pancreatic ductal adenocarcinoma; IPMN Intraductal papillary mucinous neoplasm; DEGs Differentially expressed genes.
Enrichment analysis results
The results of the GO analysis are shown in Fig. 3A and B. The extracellular matrix (ECM) was particularly emphasized. KEGG analysis showed that these DEGs are mainly involved in pathways such as the PI3K-Akt signaling pathway, protein digestion and absorption, complement and coagulation cascades, focal adhesion, amoebiasis, retinol metabolism, and pancreatic secretion (Fig. 3C,D).
Functional enrichment analysis. (A) GO functional enrichment analysis of the 341 DEGs, the most significant one is marked in red. (B) GO Circle plot; blue represents biological process (BP), yellow represents cellular component (CC), and green represents molecular function (MF). (C) KEGG functional enrichment analysis of the 341 DEGs, the most significant one is marked in red. (D) KEGG Chord plot; the size of the arc represents the correlation between the pathway and pancreatic cancer. Abbreviations: DEGs Differentially expressed genes; GO Gene Ontology; KEGG Kyoto Encyclopedia of Genes and Genomes.
Identification of hub genes
The constructed PPI network consisted of 298 nodes and 1250 edges (Fig. 4A). The top 10 genes were screened using the “CytoHubba” (plugin). The interaction mode between DEGs was determined based on key genes (Fig. 4B,C). The gene and protein names and the degrees of these hub genes are listed in Table S1.
Identification of hub genes through a PPI network diagram. (A) PPI network of 341 DEGs. Different node colors were determined based on the logFC of DEGs; the higher the degree, the darker the color. (B) Results of the cytoHubba topological analysis. (C) Hub genes network module diagram. The top ten hub genes sorted by degree are marked in yellow. Abbreviations: PPI Protein–protein interaction; DEGs Differentially expressed genes.
SDC1 and ITGA2 are potential prognostic biomarkers of PDAC related to IPMN
Among the top 10 hub genes, FN1, MMP9, COL1A1, COL1A2, COL3A1, SDC1, POSTN, and ITGA2 were highly expressed in PC tissues (Fig. 5A). The results showed that high expression of SDC1 (SDC1high) significantly affected the survival of patients with PDAC (HROS: 1.54, 95% CI 1.02–2.34, P = 0.042; HRDSS: 1.62, 95% CI 1.01–2.60, P = 0.044; HRPFI: 1.71, 95% CI 1.15–2.52, P = 0.008; Fig. 5B). Similarly, the high expression of ITGA2 (ITGA2high) can also be a good predictor of poor prognosis in patients with PDAC. (HROS: 1.97, 95% CI 1.28–3.03, P = 0.002; HRDSS: 2.11, 95% CI 1.30–3.44, P = 0.003; HRPFI: 2.13, 95% CI 1.42–3.19, P < 0.001; Fig. 5C). The K–M curves of the other genes are shown in Supplementary Figure S1.
Selection of survival-related biomarkers. (A) Ten hub gene expression differences in pancreatic samples of the TCGA and GTEx datasets (171 normal samples and 179 pancreatic cancer samples). (B, C) K‑M curves and Cox regression between the high-risk and low-risk groups of SDC1 and ITGA2 in the TCGA-PAAD dataset. (D) Time-dependent ROC curve analysis measuring the predictive performance on 1-, 3-, and 5-year OS of SDC1 and ITGA2 in the TCGA-PAAD dataset. K–M Kaplan–Meier; OS Overall survival; ROC Receiver operating characteristic; AUC Area under the curve.
As shown by the ROC curves, SDC1 and ITGA2 had an acceptable predictive effect on survival prognosis, especially in the prediction of a 5-year survival rate. For SDC1high, the AUC1-year was 0.609, AUC3-year was 0.683, and AUC5-year was 0.799. For ITGA2high, the AUC1-year was 0.655, AUC3-year was 0.637, and AUC5-year was 0.766 (Fig. 5D).
Based on these results, 178 patients of the TCGA-PAAD dataset were equally divided into two groups. The cutoff value of the division is the median expression level of SDC1 and ITGA2. In addition, we analyzed the clinicopathological characteristics of the patients according to the high and low expression of SDC1 and ITGA2 (Table 1). The expression of SDC1 and ITGA2 were affected by ‘primary therapy outcome’ and ‘histologic grade’ (P < 0.05). There were also significant differences in ITGA2high and ITGA2low between different ‘T stages’ and ‘pathologic stages’ (P < 0.05). Other clinicopathological characteristics were not affected.
Depletion of SDC1 and ITGA2 suppressed proliferation, induced apoptosis, and impaired migration in PDAC cells in vitro
We further elucidated the role of SDC1 and ITGA2 in the biological behavior of PDAC cell lines. SDC1 and ITGA2 were knocked down in PDAC cells using siRNA targeting the gene sequence. Transfection efficiency was examined using qRT-PCR (Fig. 6A) and western blotting (Fig. 6B). Si-SDC1 #1, si-SDC1 #2, si-ITGA2 #1, and si-ITGA2 #3 showed the best knockdown effects and were selected for further studies. The proliferative capacities of PDAC cells were remarkably suppressed when SDC1 and ITGA2 were downregulated, as illustrated by the CCK-8 assay (Fig. 6C). The results of the EdU assay were similar (Fig. 6D and Figure S2). Knockdown of SDC1 and ITGA2 promoted apoptosis in PDAC cells (Fig. 6E). Next, we found that PDAC cell migration was impaired after SDC1 and ITGA2 knockdown (Fig. 6F).
SiRNA-mediated knockdown of SDC1 and ITGA2 regulates PDAC cell proliferation, apoptosis, and invasion in vitro. (A) BxPC-3 and MIA-PaCa2 were transfected with siRNAs. qRT-PCR was used to detect the transfection efficiency. (B) Si-SDC1 #1, si-SDC1 #2, si-ITGA2 #1, and si-ITGA2 #3 had better knockdown effects and were selected for further studies, which were detected by western blot. (C, D) CCK-8 assay and EdU assay were conducted to examine the BxPC-3 and MIA-PaCa2 cell proliferation viability after the knockdown of SDC1 and ITGA2. (E) Knocking down SDC1 and ITGA2 expression promoted apoptosis in PDAC cells. (F) Migration of BxPC-3 and MIA-PaCa2 was detected after transfection with si-SDC1 and si-ITGA2, respectively, for 0 h and 24 h. These data are representative of three independent experiments with similar results. PDAC Pancreatic ductal adenocarcinoma; qRT-PCR Quantitative real-time polymerase chain reaction; CCK-8 Cell Counting Kit-8. (ns P ≥ 0.05; *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001).
GSEA and immune cell infiltration results
The GSEA results showed that SDC1high was mainly found in E2F targets, G2M checkpoint, glycolysis, IFN-α response, and KRAS signaling in DN. ITGA2high was mainly detected in angiogenesis, apical surface, epithelial–mesenchymal transition (EMT), heme metabolism, and MYC targeting the V1 pathway. Both SDC1low and ITGA2low were enriched in the pancreatic β-cell pathway (Fig. 7A).
There were no significant immune correlations with SDC1. A positive correlation was identified for ITGA2 expression and the levels of B cells (P = 3.87 × 10−3), CD8+ T cells (P = 9.25 × 10−10), macrophages (P = 1.84 × 10−3), neutrophils (P = 1.87 × 10−4), and dendritic cells (P = 2.30 × 10−5), indicating the key role of ITGA2 expression in immune infiltration (Fig. 7B).
Depletion of SDC1 and ITGA2 suppressed EMT, enhanced the expression of pro-inflammatory cytokines, and promoted tumor growth
We used qRT-PCR to detect changes in EMT makers and cytokine makers in BxPC-3 and MIA-PaCa2 cell lines. The results showed that after inhibiting the expression of SDC1 and ITGA2, the EMT process of pancreatic cancer cell lines was inhibited; this was manifested by an increased expression of E-cadherin and decreased expression of N-cadherin, Snail, and Twist (Fig. 8A,B). Subsequently, we observed that after inhibiting the expression of SDC1 and ITGA2, the expression of pro-inflammatory cytokines was upregulated in BxPC-3 and MIA-PaCa2 cell lines, shown by increased transcription levels of IFN-α, IL-1β, IL-6 and TNF-α (Fig. 8C,D).
Depletion of SDC1 and ITGA2 suppressed EMT, enhanced the expression of pro-inflammatory cytokines, and promoted tumor growth. (A) Si-SDC1 and si-ITGA2 inhibited EMT in BxPC-3 cells, as indicated by qRT-PCR; (B) Si-SDC1 and si-ITGA2 inhibited EMT in MIA-PaCa2 cells, as indicated by qRT-PCR; (C) Si-SDC1 and si-ITGA2 upregulated the expression of pro-inflammatory cytokines in BxPC-3 cells; (D) Si-SDC1 and si-ITGA2 upregulated the expression of pro-inflammatory cytokines in MIA-PaCa2 cells; (E) Relative SDC1 expression in tumor-bearing nude mice; (F) Relative ITGA2 expression in tumor-bearing nude mice; (G) Regression curves of relative mRNA expression and tumor volume; (H) Knocking down of SDC1 and ITGA2 inhibited tumor growth. Abbreviations: EMT Epithelial–mesenchymal transition; qRT-PCR Quantitative real-time polymerase chain reaction. (ns P ≥ 0.05; *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001).
Thereafter, we inoculated BxPC-3 and MIA-PaCa2 cell lines into immunodeficient mice to construct a mouse tumorigenic model, and detected the transcription levels of SDC1 and ITGA2, by qRT-PCR, in tumor tissues on the fifth and tenth days after tumorigenesis by qRT-PCR (Fig. 8E,F). The results showed that with the extension of inoculation time of pancreatic cancer cells, the expression levels of SDC1 and ITGA2 in tumor tissues also increased. Regression analysis of tumor tissue volume and expression of SDC1 and ITGA2 showed a positive correlation ed with the expression of SDC1 and ITGA2 (Fig. 8G, Table S4,S4). Finally, we found that after inhibiting SDC1 and ITGA2 expression in BxPC-3 and MIA-PaCa2 cell lines, the tumor size was significantly reduced, showing a potential therapeutic effect (Fig. 8H, Table S5,S5).
Discussion
The current survival rate of patients with PDAC is not ideal. Owing to the lack of effective means to predict the prognosis, it is difficult for doctors and patients to choose the best treatment when PDAC occurs and develops21. There is even evidence that the survival rate of patients with PDAC has further declined22. According to statistics, the ratio of incidence to mortality of pancreatic cancer is 1:0.9. There are increasing concerns that PDAC may gradually rise in the ranking of cancer-related causes of death23. Existing biomarkers, such as KRAS, can predict the prognosis of PDAC, but they are not effective enough24. Thus, there is an urgent need to identify new and effective prediction markers. The progression of precancerous lesions to PDAC is also accelerated by the loss of several tumor suppressors25. IPMN is a precursor to PDAC26; however, there have been few prior studies evaluating the DEGs between IPMN and PDAC. Therefore, in this study, we conducted a bioinformatics analysis and identified the DEGs between IPMN and PDAC.
Based on the results of enrichment analysis, we concluded that these DEGs are closely related to the ECM and TME, which are reflected in cell adhesion, metabolism, and angiogenesis. Survival analysis is a key method for screening prognostic markers, and ROC curves can be used to evaluate this predictive power. After creating a hierarchy of hub genes and performing survival analysis, SDC1 and ITGA2 were finally selected as two survival-related biomarkers of PDAC from the DEGs between PDAC and IPMN.
Syndecan-1 (SDC1) belongs to the syndecan family27. As a member of cell surface transmembrane acetyl heparan sulfate proteoglycans, SDC1 is involved in cell–cell and cell–matrix interactions, cell proliferation, and cell migration. In addition, it affects inflammation, wound-healing process, and tumor progression by controlling the above-mentioned cell functions. SDC1high has varying significance in different kinds of cancer. SDC1high might predict a poor prognosis of breast cancer, but a better prognosis of colorectal cancer28,29. To date, few studies have investigated the role of SDC1 in PC30. In 2005, Juuti et al.31 conducted an experimental study and confirmed that the expression of interstitial SDC1 is an independent prognostic marker of PC, whereas the expression of epithelial SDC1 only predicts a good prognosis in resectable diseases. Notably, Yablecovitch et al.32 showed that serum SDC1 levels were significantly high in patients with PDAC. Our study compared the expression of SDC1 in PDAC and IPMN samples, and we found that the expression of SDC1 in PDAC was higher than that in IPMN. SDC1high indicated poor survival in patients with PDAC. The experimental results in PDAC cells confirmed that depletion of SDC1 can significantly suppress in vitro PDAC cell proliferation, induce cell apoptosis, and impair cell migration. As confirmed by Yao et al.33, SDC1 is the key medium for phagocytosis by PC macrophages. This corroborates the fact that SDC1 regulates macrophage phagocytosis on the cell surface and promotes PDAC cell growth.
Integrin alpha 2 (ITGA2), a subunit of integrins, is overexpressed in malignancies and is associated with cancer progression34, especially reflected in the promotion of malignant behavior in tumor cell biology35. Nones et al.36 reported that ITGA2high predicted a low survival rate in patients with PDAC. Deichmann et al.37 observed that immunohistochemical examination of samples from 105 patients with PDAC showed high ITGA2 expression in 43% of patients. For patients with PDAC who had undergone resection, ITGA2 is a biomarker that could predict their prognosis. Our results supported this conclusion. PD-L1 on the surface of tumor cells can inhibit the anti-tumor activity of CD8*T cells. Overexpression of ITGA2 can activate the STAT3 signal pathway and upregulate the expression of PD-L1, thus promoting the invasion of malignant tumors, which are closely related to immunity38,39,40. Islam et al.41 used bioinformatics to analyze the prognostic role of ITGA2 in PDAC; however, they did not perform experimental validation, making the results less credible. We not only verified the effect of ITGA2 on the biological behavior of PC cells in vitro, but also performed a preliminary exploration of the mechanism. Our results showed that the specific mechanism by which ITGA2high exerts its pro-pancreatic cancer effects might be closely related to the TME. Wu et al.41 evaluated the effects of ITGA2 inhibitors in vivo using a KrasG12D-driven mouse model of PC. Notably, their results showed that when ITGA2 was pharmacologically inhibited, the pre-TME was counteracted, and pancreatic injury was reversed. Zhou et al.42 found that ITGA2 inhibited DNA repair in PC and consequently exerted a radio-sensitizing effect, which may be a novel perspective for the application of ITGA2 as a target.
As a recognizable precursor of PC, the progression of IPMN appears to be markedly immune-tolerant. The immune microenvironment of IPMN is not as severe as pancreatic intraepithelial neoplasia (PanIN), even though immunity is increasingly suppressed during the transformation of IPMN to PDAC43. As shown in the GSEA results, the IFN-α response pathway and KRAS signaling pathway are the two pathways that are significantly enriched when SDC1 is highly expressed. qRT-PCR results showed that the knockdown of SDC1 promoted IFN-α expression, validating the conclusions obtained using GSEA. There is no direct evidence that ITGA2 plays a role in PC metastasis; nonetheless, our experimental study showed that ITGA2 promotes the migration of BxPC-3 and MIA-PaCa2 cells. In addition, we found that ITGA2high signifies EMT based on the GSEA results. Subsequent experimental results confirmed that the knockdown of ITGA2 effectively inhibited EMT. It is well known that EMT activates cancer cell metastasis, precisely because of the characteristics of mesenchymal cells that epithelial cells enhance cell movement and migration44. Increased expression of pro-inflammatory cytokines may be associated with response to immunotherapy and better prognosis. Furthermore, our results showed that ITGA2 expression was positively correlated with the infiltration of multiple immune cells. Both SDC1low and ITGA2low were significantly enriched in pancreatic β cells, and inhibition of their expression may protect β cells from damage. We also confirmed that inhibiting SDC1 and ITGA2 expression makes mouse tumors smaller in vivo. In summary, targeting the SDC1 and ITGA2 pathways alone or in combination with immunotherapy may improve the survival of patients with PDAC.
Most IPMNs never develop into PDAC and can be safely monitored. The treatment of IPMN is sometimes extensive. The current basis for the treatment of IPMN relies on the Fukuoka guidelines, European evidence-based guidelines for cystic neoplasms of the pancreas, International Pancreatic Association guidelines, and expert opinion. Strong scientific evidence related to predicting which IPMNs are at high risk and will develop into aggressive diseases is lacking45. Therefore, identifying biomarkers that can predict the risk of malignant transformation remains a highly meaningful but challenging task46. SDC1 and ITGA2 are among the DEGs identified in IPMN and PDAC, and monitoring their expression is of great value in predicting whether IPMN will develop into PDAC with a poorer prognosis. Further studies are required to confirm their efficacy.
Clinical analysis demonstrated the prognostic value of SDC1 and ITGA2, with high expression, predicting poor survival rates in PDAC. This predictive power was accepted, as demonstrated by the ROC curves. However, studies on prognostic biomarkers for PDAC often lack clinical validation, and our study suffers from this limitation. Nevertheless, in general, our study identified two biomarkers in silico: SDC1 and ITGA2. In addition, we verified their inhibitory effect on the biological behavior of PDAC cells by knocking down their expression in vitro and conducted a preliminary exploration of their mechanisms. Although an in vivo experiment was not performed, our results support the conclusions of this study on SDC1 and ITGA2. We believe that SDC1 and ITGA2 may also be biomarkers for detecting the conversion of IPMN to PDAC, which warrants further study. With the development of multidisciplinary cooperation and technology, the prediction of PC prognosis and the discovery of newer treatments may continue to improve. SDC1 and ITGA2 should be considered in strategies evaluating combinations of targeted therapy and immunotherapy.
Conclusion
We conclude that SDC1 and ITGA2 are potential prognostic biomarkers for PDAC associated with IPMN. The downregulation of SDC1 and ITGA2 expression in PDAC may occur via a mechanism involving regulation of the IFN-α response, EMT, and immunity, which could act as potential new targets for PDAC therapy.
Material and methods
Ethics statement
This study was supervised by the Ethics Committee of Guang’anmen Hospital (IACUC-GAMH-2023-004).
Data collection
The expression profiles of the sequencing datasets GSE26647, GSE63104, GSE16515, and GSE28735 of the Gene Expression Omnibus (GEO) (https://www.ncbi.nlm.nih.gov/geo/)47 were downloaded. They are all publicly available. The specific data information is listed in Table S2.
RNA expression and clinical data are still downloaded in the usual way16. The data of the Cancer Genome Atlas (TCGA) and Genotype-Tissue Expression (GTEx) databases were downloaded from the University of California at Santa Cruz Xena database (https://xenabrowser.net/datapages/). We then transformed the gene expression matrix into log2 (TPM + 1) values for further use48. The clinical information included stage, sex, age, and race49.
Identification of differentially expressed genes in IPMN and PDAC samples
DEGs were filtered using the R package ‘limma’50. A P-value < 0.05 and an absolute log fold-change (FC) > 0.5 for the DEGs were defined as the cutoff. The DEGs were visualized using the R packages. The ‘complexheatmap’ package was used to generate heat maps. The ‘ggplot2’ package was used to draw volcano maps51.
Enrichment analysis of DEGs
Gene Ontology (GO) enrichment analysis was used to reveal the potential biological processes of IPMN-related DEGs in PDAC. We then visualized the results using the R packages ‘Goplot’ and ‘ggplot2′51. Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis52,53 was performed using the R package ‘clusterProfiler’54,55.
Identification of hub genes
In the first step, the protein–protein interaction (PPI) network was constructed via the STRING database (http://string-db.org/)56. In step two, Cytoscape software (3.8.0) was used to visualize the PPI network. In the last step, the “CytoHubba” plugin was applied to identify significant gene clusters sorted by degree. The top ten genes were considered hub genes57.
Selection of survival-related biomarkers
We identified survival-related biomarkers from the top ten hub genes through Kaplan–Meier (K–M) survival analysis. A total of 178 patients with PDAC were divided into two groups (‘High expression’ and ‘Low expression’) based on an automatically generated best cutoff value. Two important indicators were calculated: Hazard ratios (HR) with 95% confidence intervals (CI) and Cox regression P-values.
Clinical analysis of survival-related biomarkers
We compared the expression of survival-related biomarkers in normal and tumor tissue samples from TCGA-PAAD and GTEx datasets58. We then created a receiver operating characteristic (ROC) curve using the ‘SurvivalROC’ R package58. We analyzed their correlation with the clinicopathological features of PC using the clinical information obtained from the TCGA-PAAD dataset.
Molecular mechanism of survival-related biomarkers
Gene set enrichment analysis (GSEA) was performed based on the TCGA-PAAD and HALLMARK databases to investigate the biological functions and potential signaling pathways of survival-related biomarkers59. They were considered significantly enriched if the false discovery rate was < 0.25, P-value was < 0.05, and |LogFC| was > 0.2.
We analyzed the correlations between survival-related biomarkers and immune cells in PDAC tissues using TIMER (https://cistrome.shinyapps.io/timer/).
Cell culture and treatment
The BxPC-3 and MIA-PaCa2 cell lines, derived from human PDAC, were purchased from the China Center for Type Culture Collection (CCTCC, Wuhan, China). The cells were grown in a humidified incubator containing 5% CO2. The complete medium contained Dulbecco’s modified Eagle’s medium (DMEM; HyClone, Cat#SH30022.01, USA), 10% certified fetal bovine serum (FBS; BI, Cat#04–001-1ACS, Israel), and 1% penicillin–streptomycin. The temperature of the incubator was set at 37 °C.
Experimental animals
In accordance to the ARRIVE Guidelines before and during experiments, all methods were performed in accordance with the relevant guidelines and regulations, animals were also housed in compliance with institutional guidelines of Guang’anmen Hospital. Male BALB/c nude mice (4–5 weeks old, average weight 20 g) were purchased from Beijing Vital River Laboratory Animal Technology Co., Ltd. The mice were acclimatized for 1–2 weeks prior to the experiments and were housed in a pathogen-free environment with a temperature of 25 ± 2 °C, humidity of 50 ± 5%, and ad libitum access to food and water.
siRNA transfection
siRNAs were designed and synthesized by RiboBio Co., Ltd. (Guangzhou, China). Opti-MEM (Invitrogen, Cat#31985-070, USA) and Lipofectamine™ 3000 Transfection Reagent (Invitrogen, Cat#L3000-008, USA) were used for the transfection of siRNAs, according to the manufacturer’s instructions. NC represents the negative control group.
Quantitative real-time polymerase chain reaction
Quantitative real-time PCR (qRT-PCR) was performed to verify the interference efficiency of siRNA transfection using the ChamQ SYBR qPCR Master Mix (Vazyme, Cat#Q311-02, Nanjing, China) on an RT-PCR system (CFX96 Touch; Bio-Rad, USA) according to the manufacturer’s protocol. The gene expression was calculated using the 2−ΔΔCT value method, which was performed in triplicate60. The primers used for qRT-PCR were designed by Beijing Qingke Biotechnology Co., Ltd. (Qingdao Project Department, China). Primer sequences for genes and the experimental procedure are listed in Supplementary Table S3.
Western blotting
Western blotting was performed as previously described61. The primary antibodies included those for SDC1 (Cat#10593-1-AP, Proteintech Group, Wuhan, China), ITGA2 (Proteintech Group, Cat#24552-1-AP). GAPDH was used for normalization. The immunoblots were examined with a visible imaging system (Bio-Rad Laboratories, Hercules, CA, USA).
Cell proliferation assay
BxPC-3 and MIA-PaCa2 cells were seeded in 96-well plates (5 × 103 cells/well). Cell Counting Kit-8 (CCK-8) solution (10-μL; Beyotime, Cat#C0038, Shanghai, China) was pipetted into each well at the indicated time points (24, 48, and 72 h). The absorbance at 450 nm was measured using a multifunctional enzyme labeling instrument (Imark-22353, Bio-Rad Laboratories). Cell viability was calculated using GraphPad Prism software. An EdU Cell Proliferation Kit with Alexa Fluor 555 (Beyotime, Cat#C0075L) was used to show the cell viability of different groups clearly.
Cell apoptosis assay
Apoptotic cells were detected using Annexin V-FITC along with the PI solution using a flow cytometry assay. Add the corresponding reagent according to the product instructions (Annexin V-FITC apoptosis detection kit, Vazyme, Cat# A211-02). The number of cells and reagent dose were determined according to experimental conditions. Flow cytometric analysis was performed using a flow cytometer (BD AccuriC6 Plus Flow Cytometer, Franklin Lakes, NJ, USA). Photographs were taken three times randomly.
Wound-healing assay
A wound-healing assay was used to detect the migration of BxPC-3 and MIA-PaCa2 cells. BxPC-3 and MIA-PaCa2 cells were transfected with si-SDC1, si-ITGA2, or NC in 6-well plates and cultured for 48 h until 90% confluence was reached. Wound closure was visualized at 0 and 24 h. The distance between the wound edges was calculated as previously described62.
Animal experiments
The BxPC-3, MIA-PaCa2, si-SDC1-BxPC-3, si-SDC1-MIA-PaCa2, si-ITGA2-BxPC-3, and si-ITGA2-MIA-PaCa2 cell suspensions were added to a 1:3 formulation of Matrigel and PBS. Next, 5 × 106 cells were subcutaneously injected into the back of the right upper limbs of nude mice. The sizes of tumors in these nude mice were measured. Tumor volume (V) was calculated using the equation V = L × W2/2, where L denotes long diameter and W denotes short diameter. The BxPC-3 and MIA-PaCa2 mice were then divided into two groups of six mice each, namely, the low-volume group and high-volume group. The mice in our study were terminated on day 35 and all mice were euthanised at this time point in the form of lethal doses of anaesthesia.
Statistical analysis
Data were expressed as the mean ± SD or number of cases (%). For clinical data, the Wilcoxon test was used to test the statistical differences between multiple variables. For the categorical variables, the chi-squared test or Fisher’s exact test was used. We used R (version 4.2.1) software to conduct the statistical analyses. T-test was used in two-group comparisons to determine the variability of data obtained from in vitro experiments. For multiple groups, the one-way analysis of variance was used. Logistic regression analysis was used to determine the correlation between tumor volume and mRNA expression. GraphPad Prism (V8.2.1) was used to analyze the experimental data. Differences were considered statistically significant when the P-value was < 0.05.1
Data availability
Article/Supplementary Material containing the original contributions is available. Contact the corresponding author for more information.
References
Siegel, R. L., Miller, K. D., Wagle, N. S. & Jemal, A. Cancer statistics, 2023. CA Cancer J. Clin. 73, 17–48. https://doi.org/10.3322/caac.21763 (2023).
Jentzsch, V., Davis, J. A. A. & Djamgoz, M. B. A. Pancreatic cancer (PDAC): Introduction of evidence-based complementary measures into integrative clinical management. Cancers (Basel) 12, 3096. https://doi.org/10.3390/cancers12113096 (2020).
Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. Cancer statistics, 2022. CA Cancer J. Clin. 72, 7–33. https://doi.org/10.3322/caac.21708 (2022).
Zeng, S. et al. Chemoresistance in pancreatic cancer. Int. J. Mol. Sci. 20, 4504. https://doi.org/10.3390/ijms20184504 (2019).
Tan, E. & El-Rayes, B. Pancreatic cancer and immunotherapy: Resistance mechanisms and proposed solutions. J. Gastrointest. Cancer 50, 1–8. https://doi.org/10.1007/s12029-018-0179-z (2019).
Werner, J., Fritz, S. & Buchler, M. W. Intraductal papillary mucinous neoplasms of the pancreas–a surgical disease. Nat. Rev. Gastroenterol. Hepatol. 9, 253–259. https://doi.org/10.1038/nrgastro.2012.31 (2012).
Yamada, S. et al. Comparison of the survival outcomes of pancreatic cancer and intraductal papillary mucinous neoplasms. Pancreas 47, 974–979. https://doi.org/10.1097/MPA.0000000000001110 (2018).
Tanaka, M. et al. International consensus guidelines for management of intraductal papillary mucinous neoplasms and mucinous cystic neoplasms of the pancreas. Pancreatology 6, 17–32. https://doi.org/10.1159/000090023 (2006).
Campbell, N. M., Katz, S. S., Escalon, J. G. & Do, R. K. Imaging patterns of intraductal papillary mucinous neoplasms of the pancreas: An illustrated discussion of the international consensus guidelines for the management of IPMN. Abdom Imaging 40, 663–677. https://doi.org/10.1007/s00261-014-0236-4 (2015).
Pollini, T. et al. The tumour immune microenvironment and microbiome of pancreatic intraductal papillary mucinous neoplasms. Lancet Gastroenterol. Hepatol. 7, 1141–1150. https://doi.org/10.1016/S2468-1253(22)00235-7 (2022).
Machado, N. O., Al Qadhi, H. & Al Wahibi, K. Intraductal papillary mucinous neoplasm of pancreas. N. Am. J. Med. Sci. 7, 160–175. https://doi.org/10.4103/1947-2714.157477 (2015).
Dal Borgo, C. et al. The clinical management of main duct intraductal papillary mucinous neoplasm of the pancreas. Dig. Surg. 36, 104–110. https://doi.org/10.1159/000486869 (2019).
Tomczak, K., Czerwinska, P. & Wiznerowicz, M. The Cancer Genome Atlas (TCGA): An immeasurable source of knowledge. Contemp. Oncol. (Pozn) 19, A68-77. https://doi.org/10.5114/wo.2014.47136 (2015).
Barrett, T. et al. NCBI GEO: Archive for functional genomics data sets–update. Nucleic Acids Res. 41, D991-995. https://doi.org/10.1093/nar/gks1193 (2013).
Liu, Q. et al. Bioinformatics-based analysis: Noncoding RNA-mediated COL10A1 is associated with poor prognosis and immune cell infiltration in pancreatic cancer. J. Healthc. Eng. 2022, 7904982. https://doi.org/10.1155/2022/7904982 (2022).
Shen, Q. et al. An immune-related microRNA signature prognostic model for pancreatic carcinoma and association with immune microenvironment. Sci. Rep. 12, 9123. https://doi.org/10.1038/s41598-022-13045-z (2022).
Wang, J. et al. Next generation sequencing of pancreatic cyst fluid microRNAs from low grade-benign and high grade-invasive lesions. Cancer Lett. 356, 404–409. https://doi.org/10.1016/j.canlet.2014.09.029 (2015).
Okano, K. & Suzuki, Y. Strategies for early detection of resectable pancreatic cancer. World J. Gastroenterol. 20, 11230–11240. https://doi.org/10.3748/wjg.v20.i32.11230 (2014).
Lei, Y. et al. Applications of single-cell sequencing in cancer research: Progress and perspectives. J. Hematol. Oncol. 14, 91. https://doi.org/10.1186/s13045-021-01105-2 (2021).
Lv, G. et al. The application of single-cell sequencing in pancreatic neoplasm: Analysis, diagnosis and treatment. Br. J. Cancer https://doi.org/10.1038/s41416-022-02023-x (2022).
Park, W., Chawla, A. & O’Reilly, E. M. Pancreatic cancer: A review. JAMA 326, 851–862. https://doi.org/10.1001/jama.2021.13027 (2021).
Zeng, H. et al. Changing cancer survival in China during 2003–15: A pooled analysis of 17 population-based cancer registries. Lancet Glob. Health 6, e555–e567. https://doi.org/10.1016/S2214-109X(18)30127-X (2018).
Rahib, L. et al. Projecting cancer incidence and deaths to 2030: The unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 74, 2913–2921. https://doi.org/10.1158/0008-5472.CAN-14-0155 (2014).
Wang, S. et al. The molecular biology of pancreatic adenocarcinoma: Translational challenges and clinical perspectives. Signal Transduct. Target Ther. 6, 249. https://doi.org/10.1038/s41392-021-00659-4 (2021).
Mizrahi, J. D., Surana, R., Valle, J. W. & Shroff, R. T. Pancreatic cancer. Lancet 395, 2008–2020. https://doi.org/10.1016/S0140-6736(20)30974-0 (2020).
Goral, V. Pancreatic cancer: Pathogenesis and diagnosis. Asian Pac. J. Cancer Prev. 16, 5619–5624. https://doi.org/10.7314/apjcp.2015.16.14.5619 (2015).
Woods, A. Syndecans: Transmembrane modulators of adhesion and matrix assembly. J. Clin. Invest. 107, 935–941. https://doi.org/10.1172/JCI12802 (2001).
Mei, Y. et al. Characterization of glucose metabolism in breast cancer to guide clinical therapy. Front. Surg. 9, 973410. https://doi.org/10.3389/fsurg.2022.973410 (2022).
Li, Z. et al. High expression of SDC1 in stromal cells is associated with good prognosis in colorectal cancer. Anticancer Drugs https://doi.org/10.1097/CAD.0000000000001441 (2022).
Betriu, N., Bertran-Mas, J., Andreeva, A. & Semino, C. E. Syndecans and pancreatic ductal adenocarcinoma. Biomolecules 11, 349. https://doi.org/10.3390/biom11030349 (2021).
Juuti, A., Nordling, S., Lundin, J., Louhimo, J. & Haglund, C. Syndecan-1 expression–a novel prognostic marker in pancreatic cancer. Oncology 68, 97–106. https://doi.org/10.1159/000085702 (2005).
Yablecovitch, D. et al. Serum syndecan-1: A novel biomarker for pancreatic ductal adenocarcinoma. Clin. Transl. Gastroenterol. 13, e00473. https://doi.org/10.14309/ctg.0000000000000473 (2022).
Yao, W. et al. Syndecan 1 is a critical mediator of macropinocytosis in pancreatic cancer. Nature 568, 410–414. https://doi.org/10.1038/s41586-019-1062-1 (2019).
Adorno-Cruz, V. & Liu, H. Regulation and functions of integrin alpha2 in cell adhesion and disease. Genes Dis. 6, 16–24. https://doi.org/10.1016/j.gendis.2018.12.003 (2019).
Adorno-Cruz, V. et al. ITGA2 promotes expression of ACLY and CCND1 in enhancing breast cancer stemness and metastasis. Genes Dis. 8, 493–508. https://doi.org/10.1016/j.gendis.2020.01.015 (2021).
Nones, K. et al. Genome-wide DNA methylation patterns in pancreatic ductal adenocarcinoma reveal epigenetic deregulation of SLIT-ROBO, ITGA2 and MET signaling. Int. J. Cancer 135, 1110–1118. https://doi.org/10.1002/ijc.28765 (2014).
Deichmann, S. et al. Overexpression of integrin alpha 2 (ITGA2) correlates with poor survival in patients with pancreatic ductal adenocarcinoma. J. Clin. Pathol. https://doi.org/10.1136/jclinpath-2022-208176 (2022).
Aerts, J. G. & Hegmans, J. P. Tumor-specific cytotoxic T cells are crucial for efficacy of immunomodulatory antibodies in patients with lung cancer. Cancer Res. 73, 2381–2388. https://doi.org/10.1158/0008-5472.CAN-12-3932 (2013).
Villaruz, L. C., Kalyan, A., Zarour, H. & Socinski, M. A. Immunotherapy in lung cancer. Transl. Lung Cancer Res. 3, 2–14. https://doi.org/10.3978/j.issn.2218-6751.2013.10.13 (2014).
Ren, D. et al. Overexpressed ITGA2 promotes malignant tumor aggression by up-regulating PD-L1 expression through the activation of the STAT3 signaling pathway. J. Exp. Clin. Cancer Res. 38, 485. https://doi.org/10.1186/s13046-019-1496-1 (2019).
Ren, T., Xue, X., Wang, X., Zhou, X. & Dang, S. Bioinformatic and experimental analyses of key biomarkers in pancreatic cancer. Exp. Ther. Med. 22, 1359. https://doi.org/10.3892/etm.2021.10794 (2021).
Zhou, C. et al. ITGA2 overexpression inhibits DNA repair and confers sensitivity to radiotherapies in pancreatic cancer. Cancer Lett. 547, 215855. https://doi.org/10.1016/j.canlet.2022.215855 (2022).
Hernandez, S. et al. Diminished immune surveillance during histologic progression of intraductal papillary mucinous neoplasms offers a therapeutic opportunity for cancer interception. Clin. Cancer Res. 28, 1938–1947. https://doi.org/10.1158/1078-0432.CCR-21-2585 (2022).
Weng, Y. et al. Identification of EMT-related alternative splicing event of TMC7 to promote invasion and migration of pancreatic cancer. Front. Immunol. 13, 1089008. https://doi.org/10.3389/fimmu.2022.1089008 (2022).
Fonseca, A. L., Kirkwood, K., Kim, M. P., Maitra, A. & Koay, E. J. Intraductal papillary mucinous neoplasms of the pancreas: Current understanding and future directions for stratification of malignancy risk. Pancreas 47, 272–279. https://doi.org/10.1097/MPA.0000000000000999 (2018).
Khoury, R. E. et al. What is the incidence of malignancy in resected intraductal papillary mucinous neoplasms? An analysis of over 100 US institutions in a single year. Ann. Surg. Oncol. 25, 1746–1751. https://doi.org/10.1245/s10434-018-6425-6 (2018).
Singhania, A. et al. Multitissue transcriptomics delineates the diversity of airway T cell functions in asthma. Am. J. Respir. Cell Mol. Biol. 58, 261–270. https://doi.org/10.1165/rcmb.2017-0162OC (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Wang, Z., Jensen, M. A. & Zenklusen, J. C. A practical guide to the cancer genome atlas (TCGA). Methods Mol. Biol. 1418, 111–141. https://doi.org/10.1007/978-1-4939-3578-9_6 (2016).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47. https://doi.org/10.1093/nar/gkv007 (2015).
Walter, W., Sanchez-Cabo, F. & Ricote, M. GOplot: An R package for visually combining expression data with functional analysis. Bioinformatics 31, 2912–2914. https://doi.org/10.1093/bioinformatics/btv300 (2015).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361. https://doi.org/10.1093/nar/gkw1092 (2017).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51, D587–D592. https://doi.org/10.1093/nar/gkac963 (2023).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16, 284–287. https://doi.org/10.1089/omi.2011.0118 (2012).
Li, E., Luo, T. & Wang, Y. Identification of diagnostic biomarkers in patients with gestational diabetes mellitus based on transcriptome gene expression and methylation correlation analysis. Reprod. Biol. Endocrinol. 17, 112. https://doi.org/10.1186/s12958-019-0556-x (2019).
Szklarczyk, D. et al. STRING v10: Protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447-452. https://doi.org/10.1093/nar/gku1003 (2015).
Chin, C. H. et al. cytoHubba: Identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 8(4), 1–7. https://doi.org/10.1186/1752-0509-8-S4-S11 (2014).
Vivian, J. et al. Toil enables reproducible, open source, big biomedical data analyses. Nat. Biotechnol. 35, 314–316. https://doi.org/10.1038/nbt.3772 (2017).
Wang, Z., Meng, Z. & Chen, C. Screening of potential biomarkers in peripheral blood of patients with depression based on weighted gene co-expression network analysis and machine learning algorithms. Front. Psychiatry 13, 1009911. https://doi.org/10.3389/fpsyt.2022.1009911 (2022).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408. https://doi.org/10.1006/meth.2001.1262 (2001).
Tan, Z. et al. The role of tumor inflammatory microenvironment in lung cancer. Front. Pharmacol. 12, 688625. https://doi.org/10.3389/fphar.2021.688625 (2021).
Martinotti, S. & Ranzato, E. Scratch wound healing assay. Methods Mol. Biol. 2109, 225–229. https://doi.org/10.1007/7651_2019_259 (2020).
Acknowledgements
We sincerely thank the contributors in the TCGA, GTEx, GEO, and Editage (www.editage.cn) for English language editing.
Funding
This study was supported by the Scientific and Technological Innovation Project of China Academy of Chinese Medical Sciences (No. CI2021A01805) and the Natural Science Foundation of Beijing (No. 7222296).
Author information
Authors and Affiliations
Contributions
CL.Z., Q.S., and B.P. developed the study concept and design. CL.Z., Q.S., FD.L, MQ.G., XC.J., and Y.L. performed the initial literature review. CL.Z., Q.S., FD.L, F.Y., MQ.G., XC.J, Y.L., XY.Z., GE.E., and X.P. participated in extracting and analyzing data and experimental studies. CL.Z. and X.P. wrote the first draft. CL.Z., Q.S., FD.L, F.Y., MQ.G., XC.J, Y.L., XY.Z., GE.E., X.P., and B.P. participated in the development and revisions of the manuscript. B.P. obtained funding and provided study supervision. All authors contributed to the writing of the final manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Cl., Shen, Q., Liu, Fd. et al. SDC1 and ITGA2 as novel prognostic biomarkers for PDAC related to IPMN. Sci Rep 13, 18727 (2023). https://doi.org/10.1038/s41598-023-44646-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-44646-x
- Springer Nature Limited
This article is cited by
-
EPYC functions as a novel prognostic biomarker for pancreatic cancer
Scientific Reports (2024)