Abstract
Clear cell renal carcinoma (ccRCC) is one of the most common cancers worldwide. In this study, a new model of immune-related genes was developed to predict the overall survival and immunotherapy efficacy in patients with ccRCC. Immune-related genes were obtained from the ImmPort database. Clinical data and transcriptomics of ccRCC samples were downloaded from GSE29609 and The Cancer Genome Atlas. An immune-related gene-based prognostic model (IRGPM) was developed using the least absolute shrinkage and selection operator regression algorithm and multivariate Cox regression. The reliability of the developed models was evaluated by Kaplan–Meier survival curves and time-dependent receiver operating characteristic curves. Furthermore, we constructed a nomogram based on the IRGPM and multiple clinicopathological factors, along with a calibration curve to examine the predictive power of the nomogram. Overall, this study investigated the association of IRGPM with immunotherapeutic efficacy, immune checkpoints, and immune cell infiltration. Eleven IRGs based on 528 ccRCC samples significantly associated with survival were used to construct the IRGPM. Remarkably, the IRGPM, which consists of 11 hub genes (SAA1, IL4, PLAUR, PLXNB3, ANGPTL3, AMH, KLRC2, NR3C2, KL, CSF2, and SEMA3G), was found to predict the survival of ccRCC patients accurately. The calibration curve revealed that the nomogram developed with the IRGPM showed high predictive performance for the survival probability of ccRCC patients. Moreover, the IRGPM subgroups showed different levels of immune checkpoints and immune cell infiltration in patients with ccRCC. IRGPM might be a promising biomarker of immunotherapeutic responses in patients with ccRCC. Overall, the established IRGPM was valuable for predicting survival, reflecting the immunotherapy response and immune microenvironment in patients with ccRCC.
Similar content being viewed by others
Introduction
Renal cell carcinoma (RCC) is one of the most prevalent cancers worldwide, accounting for nearly 90% of all kidney cancers1,2. Its incidence is approximately twice as high in men as in women3. RCC consists of three main histological subtypes with different molecular and genetic characteristics; clear cell renal cell carcinoma (ccRCC) is the most extensive histological subtype (approximately 80%)4. Even in patients with localized ccRCC, surgical resection, which is considered the best treatment choice, is associated with a high rate of recurrence5,6. However, due to the anatomical location of the kidneys early symptoms in many patients with ccRCC are subtle and easily overlooked leading to delayed diagnosis. Consequently, by the time of diagnosis, metastases are already advanced in many cases7,8. Over the past 20 years, there has been limited progress in ccRCC prognosis. It is essential to investigate the mechanisms underlying ccRCC in detail and develop novel therapeutic targets to enhance the survival of patients with ccRCC9,10.
The immune system is closely linked to the prognosis of malignant tumors11. Immuno-oncology, the immune microenvironment, and immune cells are essential factors in tumorigenesis12,13. All cancers can be considered immune to some extent, and immunotherapy exploits the host immune system to produce a specific immune response to identify and eliminate cancer cells, thereby reducing the incidence of tumor metastasis and recurrence14,15. The tumor immune microenvironment (TIME) is integral to immunotherapy and has gradually gained increasing attention. Analysis of the TIME can help improve responsiveness to immunotherapy. Additionally, TIME can be used as a significant prognostic indicator to enhance the efficacy of precise treatments16,17. Considering the vital importance of the TIME, the immediate task is to build a signature of immune-related genes (IRGs) that are closely related to the TIME to explore the IRGs of patients with ccRCC.
Although ccRCC signatures based on IRGs have been identified recently, a more comprehensive and credible indicator is urgently needed. It is possible to predict both survival and immunotherapy in ccRCC patients18,19. Therefore, in this study, an IRG-based prognostic model (IRGPM) was established through cancer bioinformatics and genomics, and its reliability was validated using several datasets. Moreover, the value of IRGPM in the survival of patients with ccRCC, together with its potential predictive role in immunotherapy, was examined. Finally, the IRGPM can guide clinicians on cancer immunotherapy in patients with ccRCC.
Methods
Acquisition of sample information
Clinical information together with RNA-seq data for the samples of ccRCC were gathered from The Cancer Genome Atlas (TCGA) (https://portal.gdc.cancer.gov/) and Gene Expression Omnibus (GEO) (https://www.ncbi.nlm. nih.gov/gds/) and named for the TCGA-KIRC (n = 528) and GSE29609 (n = 39). TCGA-KIRC and GSE29609 are classified as training cohort and validation cohort, separately. Furthermore, 1793 IRGs were acquired from The Immunology Database and Analysis Portal (ImmPort) database (https://www.immport.org/home)20.
Construction of differentially expressed IRGs (DEIRGs)
Differentially expressed genes (DEGs) between ccRCC and normal samples were identified using the R package “edgeR” and “DESeq2” based on the following criterion: false discovery rate (FDR) < 0.05 and |log2 fold-change (FC)|> 1.521,22. In addition, DEIRGs were obtained by analysis of the crossover of IRGs and DEGs. The R package “ggplot2” was employed to visualize the DEIRGs using the Venn diagram and create a volcano plot of DEIRGs23.
Biological function analysis
To study the bio-functions of DEIRGs, we conducted functional enrichment analyses of DEIRGs, including the Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO), with the R package “clusterProfiler”24. The GO terms cover biological process (BP), molecular function (MF) and cellular constituents (CC)25,26,27,28. The signaling pathways of KEGG and GO terms were enriched as measured by FDR < 0.01. This was followed by a visual analysis of the first 10 most critical KEGG signaling pathways and 15 most related GO terms applying the R package “ggplot2”29. For further investigation of the underlying mechanism of DEIRGs, Gene Set Enrichment Analysis (GSEA) was implemented to elucidate the significant biological process with the R package “clusterProfiler”30. P adjust < 0.05 and FDR < 0.25 were regarded as statistically significant.
Signature exploitation and reliability assessment
IRGs related to prognosis were determined, and IRGPM based on the training set was built and then its predictive performance was validated in the GSE29609 cohort. Specifically, a univariate Cox proportional hazard regression analysis was implemented while investigating prognosis-related IRGs to assess the relationship between DEIRGs and overall survival (OS) in training set31. The prognosis-related IRGs were identified using the significance value of P ≤ 0.05. Thereafter, Least Absolute Shrinkage and Selection Operator (LASSO) was employed to selected the most relavant IRGs associated with prognosis via the R package “glmnet” and “survival”32. Lambda. min was chosen as the cutoff. Additionally, the IRGPM, which predicts the prognosis OS of ccRCC patients, was built applying Multivariate Cox proportional hazard regression analysis. In multivariate Cox regression analysis, the risk score of each patient with ccRCC was weighted based on their estimated regression coefficient. Patients were then classified as low- and high-risk groups using the median risk score as a threshold. Kaplan–Meier (K–M) analysis was performed for the comparison of survival between both groups via the R package “survival” to further validate the IRGPM's predictive power. Furthermore, the “time ROC” R package was applied for generating the ROC curve, which illustrates the sensitivity and specificity of IRGPM including 1-, 3-, and 5-year survival33,34.
Relationship between IRGPM and clinical pathological factors
Univariate and multivariate Analysis of OS for clinical-pathological parameters together with IRGPM were implemented in TCGA-KIRC cohort, and GSE29609 through R package “survival”35. In addition, the correlation of IRGPM with different clinic-pathological parameters (sex, histologic state, M stage, N stage, pathologic stage, T stage) was assessed with Mann–Whitney U tests.
Construction of prognostic nomogram
From the IGRPM and the clinical parameters (N-stage, M-stage, T-stage, pathological stage, Histological grade, and Sex), a nomogram was further built that could predict the probability of survival of patients suffering from ccRCC. Furthermore, calibration curves were drawn to check the predictive validity of nomograms, and the agreement between actual and predicted survival performance can also be compared. Nomograms along with its calibration curves were plotted with the R package “rms”.
Evaluation of immune cell infiltration and checkpoint
R package “GSVA” was exploited to examine the immune cell infiltration between low- and high-riskscore groups36,37. The association between the immune checkpoints/immune cells and the model's risk score was assessed through the Mann–Whitney U test, including programmed cell death protein 1 (PD1/PDCD1), cytotoxic T-lymphocyte-associated protein 4 (CTLA4), PDCD1LG2, PD ligand 1 (PDL1/CD274), T cell immunoglobulin and ITIM domain (TIGIT), Sialic Acid Binding Ig Like Lectin 15 (SIGLEC15), lymphocyte activation gene-3 (LAG3) and Hepatitis A virus cellular receptor 2 (HAVCR2).
Effectiveness of immunotherapy
Immunophenoscore (IPS) was employed to predict the immune checkpoint blockades (ICBs) responses in the TCGA-KIRC patients. IPS is calculated based on gene expression of four fundamental components: immunomodulators, MHC molecules, suppressor cells and effector cells, on scale ranging from 0 to 10. ccRCC patients’ IPS was obtained from The Cancer Immunome Atlas (TCIA)38.
Statistical analysis
Statistical analyses were conducted in R software (version 3.6.3), with the Kruskal–Wallis test and Wilcoxon test adopted for non-normally distributed data between multiple and two groups, separately. While the one-way analysis of variance (ANOVA) and unpaired Student’s t-test were separately applied for the normally distributed variables between multiple and two groups. The R package “survminer” was exploited for the Kaplan–Meier survival plots. P < 0.05 illustrated a statistical significance, all statistical P-values were two-sided.
Ethical approval and consent to participate
All data in this study were collected from public data- bases: TCGA and GEO. This article does not contain any studies with patients or animals performed by any of the authors.
Results
Analysis of DEIRGs
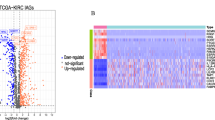
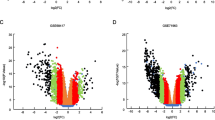
Figure 1 provides a comprehensive flowchart of the study. Analysis of 528 ccRCC samples and 72 normal samples identified 11,498 DEGs. Meanwhile, 679 DEIRGs were obtained from the crossover of 11,498 DEGs and 1793 IRGs, including 559 increased and 120 decreased genes (Fig. 2A,B and Table S1). Functional enrichment analysis revealed that the top three relevant signaling pathways for DEIRGs were cytokines-cytokine receptor interaction, viral protein interaction with cytokines and cytokine receptors, and natural killer cell-mediated cytotoxity. (Fig. 2C and Table S2). Besides, the most enriched terms of BP, CC, and MF are “humoral immune response”, “T cell receptor complex” and “antigen binding”, respectively (Fig. 2D and Table S3). GSEA analysis has indicated strong associations between DEIRGs and oncological pathways such as response chemokine receptors binding chemokines, chemokine signaling pathways and cytokine-cytokine receptor interaction (Fig. 2E and Table S4).
Identification of IRGPM. (A)Volcano plot for differentially expressed genes between normal patients and ccRCC based on TCGA-KIRC cohort. (B) Venn diagram of DEGs and IRGs. DEGs: Differentially expressed genes, IRGs: immune-related genes. (C) KEGG Analysis of the top 10 enrichment pathways. (D) GO enrichment analysis of 679 DEIRGs in ccRCC. BP, CC, and MF indicates molecular function, cellular constituents, and biological process, respectively. DEIRGs indicates differentially expressed immune-related genes. (E) GSEA revealed several significantly enriched oncological signatures. The upper part of the plot shows the enrichment score; the lower part of the plot shows the ranked list metric for the gene set.
Construction of IRGPM
Univariate -Cox regression analysis was carried out for screening the OS-associated DEIRGs of KIRC patients and then 193 prognostic DEIRGs were determined (P < 0.05, Table S5). Subsequently, LASSO Cox analysis was conducted based on the 193 prognostic DEIRGs, and lambda.min was chosen to prevent overfitting during the procedure (Fig. S1). In total, 11 hub genes were converged after tenfold cross-validation, which were SAA1, IL4, PLAUR, PLXNB3, ANGPTL3, AMH, KLRC2, NR3C2, KL, CSF2, and SEMA3G. Then the multivariate Cox analysis was conducted for the establishment of IRGPM. Finally, 11 OS-related DEIRGs were identified and applied to establish a multivariate Cox IRGPM (Fig. S2). The IRGPM’s risk score was calculated via their coefficients of hub genes in the following way: Risk score = (Expression level of SAA1 × 0.037455) + (Expression level of IL4 × 0.493798) + (Expression level of PLAUR × 0.129711) + (Expression level of PLXNB3 × 0.232950) – (Expression level of ANGPTL3 × 0.117101) + (Expression level of AMH × 0.163395) + (Expression level of KLRC2 × 0.212960) – (Expression level of NR3C2 × 0.210899) – (Expression level of KL × 0.666003) + (Expression level of CSF2 × 0.297717) – (Expression level of SEMA3G × 0.408205).
IRGPM predicts survival of ccRCC patients
Patients suffering from ccRCC were classified as high- and low-riskscore groups according to median IRGPM risk score (Fig. 3A and Table S6). In comparison with patients in the low riskscore group, those in the high riskscore group present a marked lower OS (Fig. 3B). The IRGPM was validated in the GSE29609 cohort, which also showed a high performance in predicting the OS (Fig. S3). Moreover, the time-dependent ROC curve illustrates the reliability of IRGPM (Fig. 3C). In the TCGA-KIRC cohort, for 1, 3 and 5-year survival, the area under the curve (AUC) was 0.765, 0.726 and 0.745, respectively, it shows that the constructed IRGPM is useful in monitoring the survival rate.
IRGPM precisely predicts the survival status of ccRCC patients. (A) Distribution of risk scores, expression of 11 OS-related DEIRGs, and survival status for ccRCC patients in low- and high-risk groups based on TCGA-KIRC cohort. “0” indicates alive and “1” indicates dead (B) The Kaplan–Meier curves for overall survival between low- and high-risk groups based on TCGA-KIRC cohort. (C) Analysis of the IRGPM's time-dependent ROC curve for 1-, 3-, and 5- years of survival based on TCGA-KIRC cohort.
IRGPM is significantly associated with disease progression
Analysis of the correlation between IRGPM and multiple clinicopathological parameters was investigated by the Mann–Whitney U test. In the TCGA-KIRC cohort, the high risk score group is positively related to patients with advanced pathologic stage, advanced histologic grade, advanced TNM stage, and male sex (Fig. 4A,B). The findings indicate that IRGPM is associated significantly with diverse clinicopathological factors, with higher riskscore corresponding to poorer clinicopathological status.
IRGPM risk score with different clinicopathological factors in ccRCC patients. (A and B) The association between IRGPM and clinicopathological factors in the TCGA-KIRC cohort. (C) Nomogram model predicting 1-, 3- and 5-year OS for ccRCC patients in TCGA-KIRC cohort. (D) The 1-, 3- and 5-year calibration curve of the nomogram in TCGA-KIRC cohort. The ideal line at 45° indicates a perfect prediction, the blue, red, green lines respectively represent 1‐, 3‐, and 5-year survival.
Moreover, we built a prognostic nomogram according to IRGPM and important clinicopathological parameters, to give a predictable quantitative analysis tool for predicting the patients' survival risk (Fig. 4C). Furthermore, the calibration curves of prognostic nomograms indicated excellent agreement between predicted values and the actual 1, 3 and 5-year survival rates in the TCGA-KIRC cohort (Fig. 4D).
IRGPM predicts the immune cell infiltration of the ccRCC microenvironment
To further reveal the role of IPGRM on the TIME, immune cells infiltrating in ccRCC patients were investigated via the GSVA algorithm. Among the immune cells, the high riskscore group significant positive associated with the proportional numbers of T cells, DC, Cytotoxic cells, B cells, aDC, Th1 cells, Th2 cells, Macrophages, CD8 T cells, Treg and natural killer (NK) CD56bright cells (Fig. 5A and Table S7). Meanwhile, we studied the prognostic value of selected important immune cells, with higher infiltration abundance of CD4+T cells (Fig. 5B), activated memory M0 Macrophage (Fig. 5C), and activated NK cell (Fig. 5D), and Tregs (Fig. 5E) was significantly negatively correlated with OS. In conclusion, IRGPM associated with the infiltration level of the majority of immune cells, showing that the IRGPM could potentially indicate the status of the TIME.
Analysis of immune cell infiltration. (A) The differences in immune cell infiltrations between low- and high-risk groups. The red and blue box represented the IRGPM high- and low-risk groups, respectively. The relationships between OS and (B) T cell CD4+ memory. (C) Macrophage. (D) NK cell and (E) T cell regulatory based on TIMER2.0.
The correlation between IRGPM and immunotherapy response
Furthermore, the potential relationship between the IRGPM and immune checkpoints, including CTLA4, PD1/PDCD1, PDL1/CD274, PDCD1LG2, SIGLEC15, TIGIT, HAVCR2, LAG3, were investigated (Table S8). Figure 6A demonstrates that the low riskscore group was significantly positively correlated to CD274, PDCD1LG2, and HAVCR2, while it was negatively correlated to CTLA-4, TIGIT, LAG3, and PDCD1. The findings exhibited that the low riskscore group is involved in better immunotherapy efficacy of ICBs of CD274, PDCD1LG2, and HAVCR2, and their high expression is related to a better prognosis. The high riskscore group will benefit more from immunotherapy efficacy from ICBs of CTLA-4, TIGIT, LAG3, and PDCD1. IPS was found to be a promising index for evaluating the ICBs therapy response, and higher IPS indicated better immunotherapy response. A higher PD1/PDL1/PDL2 and IPS–CTLA4 blocker score was observed in the high riskscore group (Fig. 6B) The results of immunotherapy response are consistent with immunecheckpoint, a higher PDCD1 and CTLA-4 expression was observed in the high riskscore group.
(A) The expression level of several salient checkpoints (CTLA4, CD274, PDCD1LG2, SIGLEC15, TIGIT, HAVCR2, LAG3, and PDCD1) between low- and high- riskscore groups based on TCGA-KIRC. (B) The difference in IPS between low- and high- riskscore groups. IPS: Immunophenoscore. IPS-A: CTLA4 blocker, IPS-B: PD1/PDL1/PDL2 blocker and IPS-C: CTLA4 and PD1/PDL1/PDL2 blocker.
Discussion
In recent years, immunotherapy has gained popularity as a treatment for various cancers, including ccRCC39,40,41. Studies have reported that the immunological landscape of the ccRCC microenvironment could be a critical prognostic factor that should not be ignored to improve the potential for accurate treatment18,19,42. However, the efficacy of immunotherapy varies among individuals, and only a proportion of patients experience clinical benefits. Therefore, it is crucial to create a robust metric for predicting the survival of ccRCC patients and expand the repertoire of effective cancer immunotherapies.
In recent years, genomics and bioinformatics have made it possible to precisely determine molecular signatures. While some researchers have developed prognostic indicators based on miRNAs, lncRNAs, or mRNA, we believe that an immune gene-based signature is more appropriate for predicting the prognosis in immunotherapy43,44. Eleven optimal IRGs-based models were established as follows: SAA1, IL4, PLAUR, PLXNB3, ANGPTL3, AMH, KLRC2, NR3C2, KL, CSF2, and SEMA3G. Among these 11 IRGs, some have been found to be closely associated with immune cells. SAA1 is involved in the activation of monocytes and macrophages during inflammation45. IL-4 functions as a T-cell-derived B-cell growth factor46. ANGPTL3 plays an active role in the Treg expansion function47. KLRC4 participates in the antitumor immune role of the NK cells48. NR3C2 and CSF2 are correlated with macrophage polarization and activation49,50. Some genes might be implicated in immunomodulatory activities, such as PLAUR51, PLXNB352, AMH53, KL54, and SEMA3G55. A previous study obtained data from the TCGA database and established a model based on 14 IRGs using the LASSO-COX method18. Unfortunately, the model suffers from platform bias and lacks validation data from other databases. Zhou et al. developed a prognostic model for immune-related genes pairs through bioinformatics analysis of papillary renal cell carcinoma; however, this model did not include clinical variables19.
To further assess the clinical effect of IRGPM, we studied its relationship with clinicopathological factors, IPS, and immune cell infiltration. First, we found that poorer pathologic state, higher histologic state, male sex, and higher TNM stage were associated with the high-risk group, suggesting that our prediction model is accurate in predicting the progression of ccRCC. Second, we assessed the proportion of immune cells in ccRCC samples. Studies have shown that the interaction between the microenvironment and the tumor is a determining factor in ccRCC progression. Therefore, we evaluated the potential of IRGPM to display the prognostic value of immune cells and immune cell infiltration. M0 macrophages and Tregs have been reported to promote tumor progression in various cancers 56,57, which are consistent with our study that high riskscore patients showed a higher enrichment of these cells. Interestingly, our study suggests that NK cells enhance the immune response and lead to a better prognosis based on TCGA-KIRC cohort (Fig. 5A). Conversely, analysis from TIMER in our study showed that high infiltration levels of activated NK cells is associated with poor prognosis (Fig. 5D). The potential reason of the inconsistent is that NK cells exert both oncogenic and tumor-suppressive effects in ccRCC. NK cells facilitate tumor growth and angiogenesis by releasing cytokines and growth factors in ccRCC. NK cells can directly recognize and killing ccRCC cells and activating other immune cells (T cells) to eliminate the ccRCC cells58. Another possible reason of the inconsistent is that the two results are based on different genomic data. Previously study demonstrated that enriched DCs were found to be closely associated with dysfunctional T cells, resulting in poor patient survival59. Similarly, our results indicated that the high riskscore group exhibited higher DC cell infiltration (Fig. 5A). Finally, we assessed immune checkpoint therapy in patients with ccRCC and found that the high riskscore group presented better IPS involving CTLA4 and PD1/PDL1/PDL2 combination blockades. This indicates that the high-risk group was more immunogenic to ICBs. This insight could guide the prediction of personalized cancer immunotherapy.
Yin et al. developed a gene signature to predict prognosis of ccRCC, while they only explore the response to anti-PD-1 therapy60. Zhang et al. constructed a gene-based model for prognosis prediction of ccRCC61. However, the study didn`t investigate relationship between the models and immune infiltration/immune checkpoints/ immunotherapy. Our study has several strengths compared with previously reported studies. First, the signature is confirmed and evaluated using multiple datasets, improving its reliability. Second, we performed several multifaceted studies that included discussions on the association of ICB and IRGPM with immune cells. Third, we established a nomogram for quantitative calculations, which is beneficial for clinical dissemination and application. Nonetheless, this study also had several limitations. First, our risk model was based on bioinformatics analyses of GEO and TCGA. To make this study more reliable, examining them experimentally in conjunction with clinical specimens is necessary. Furthermore, our study was retrospective in nature, and a prospective randomized trial should be designed to test and validate our hypotheses.
Conclusions
In conclusion, we established an IRGPM for ccRCC to provide a better picture of the effect of immunotherapy and accurately predict the survival of patients with ccRCC. The IRG model provides clinicians with an effective tool for more precise treatment of patients with ccRCC.
Data availability
The data that support the findings of this study are openly available found here: TCGA-KIRC (http://portal.gdc.cancer.gov/) and GEO-GSE29609 (https://www.ncbi. nlm.nih.gov/geo/).
References
Ferlay, J. et al. Cancer incidence and mortality patterns in Europe: Estimates for 40 countries and 25 major cancers in 2018. Eur. J. Cancer 103, 356–387. https://doi.org/10.1016/j.ejca.2018.07.005 (2018).
Hsieh, J. J. et al. Renal cell carcinoma. Nat. Rev. Dis. Primers 3, 17009. https://doi.org/10.1038/nrdp.2017.9 (2017).
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249. https://doi.org/10.3322/caac.21660 (2021).
Moch, H., Cubilla, A. L., Humphrey, P. A., Reuter, V. E. & Ulbright, T. M. The 2016 WHO classification of tumours of the urinary system and male genital organs-part A: Renal, penile, and testicular tumours. Eur. Urol. 70, 93–105. https://doi.org/10.1016/j.eururo.2016.02.029 (2016).
Ljungberg, B. et al. European association of urology guidelines on renal cell carcinoma: The 2019 update. Eur. Urol. 75, 799–810. https://doi.org/10.1016/j.eururo.2019.02.011 (2019).
Rini, B. I., Campbell, S. C. & Escudier, B. Renal cell carcinoma. Lancet 373, 1119–1132. https://doi.org/10.1016/S0140-6736(09)60229-4 (2009).
Hao, H. et al. Reduced GRAMD1C expression correlates to poor prognosis and immune infiltrates in kidney renal clear cell carcinoma. PeerJ 7, e8205. https://doi.org/10.7717/peerj.8205 (2019).
Frank, I. et al. An outcome prediction model for patients with clear cell renal cell carcinoma treated with radical nephrectomy based on tumor stage, size, grade and necrosis: the SSIGN score. J. Urol. 168, 2395–2400. https://doi.org/10.1097/01.ju.0000035885.91935.d5 (2002).
Li, W. et al. High expression of activated CD4(+) memory T cells and CD8(+) T cells and low expression of M0 macrophage are associated with better clinical prognosis in bladder cancer patients. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi 36, 97–103 (2020).
Raimondi, A. et al. Predictive biomarkers of response to immunotherapy in metastatic renal cell cancer. Front. Oncol. 10, 1644. https://doi.org/10.3389/fonc.2020.01644 (2020).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: The next generation. Cell 144, 646–674. https://doi.org/10.1016/j.cell.2011.02.013 (2011).
Sima, P., Vannucci, L. & Vetvicka, V. Immunity in cancer and atherosclerosis. Ann. Transl. Med. 7, 204. https://doi.org/10.21037/atm.2019.04.56 (2019).
Bai, D. et al. Integrated analysis of immune-related gene subtype and immune index for immunotherapy in clear cell renal cell carcinoma. Pathol. Res. Pract. 225, 153557. https://doi.org/10.1016/j.prp.2021.153557 (2021).
Management of hepatocellular carcinoma. European Association for the Study of the Liver. Electronic address, e. e. e. & European Association for the Study of the, L. EASL Clinical Practice Guidelines. J. Hepatol. 69, 182–236. https://doi.org/10.1016/j.jhep.2018.03.019 (2018).
Refolo, M. G., Lotesoriere, C., Messa, C., Caruso, M. G. & D’Alessandro, R. Integrated immune gene expression signature and molecular classification in gastric cancer: New insights. J. Leukoc. Biol. 108, 633–646. https://doi.org/10.1002/JLB.4MR0120-221R (2020).
Taube, J. M. et al. Implications of the tumor immune microenvironment for staging and therapeutics. Mod. Pathol. 31, 214–234. https://doi.org/10.1038/modpathol.2017.156 (2018).
Xu, W. H. et al. Prognostic value and immune infiltration of novel signatures in clear cell renal cell carcinoma microenvironment. Aging (Albany NY) 11, 6999–7020. https://doi.org/10.18632/aging.102233 (2019).
Zou, Y. & Hu, C. A 14 immune-related gene signature predicts clinical outcomes of kidney renal clear cell carcinoma. PeerJ 8, e10183. https://doi.org/10.7717/peerj.10183 (2020).
Zhou, X. et al. Development and validation of an individualized immune-related gene pairs prognostic signature in papillary renal cell carcinoma. Front. Genet. 11, 569884. https://doi.org/10.3389/fgene.2020.569884 (2020).
Bhattacharya, S. et al. ImmPort: Disseminating data to the public for the future of immunology. Immunol. Res. 58, 234–239. https://doi.org/10.1007/s12026-014-8516-1 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: A bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140. https://doi.org/10.1093/bioinformatics/btp616 (2010).
Villanueva, R. A. M. & Chen, Z. J. ggplot2: Elegant graphics for data analysis, 2nd edition. Meas-Interdiscip. Res. 17, 160–167 (2019).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16, 284–287. https://doi.org/10.1089/omi.2011.0118 (2012).
Harris, M. A. et al. The gene ontology (GO) database and informatics resource. Nucleic Acids Res. 32, D258-261. https://doi.org/10.1093/nar/gkh036 (2004).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51, D587–D592. https://doi.org/10.1093/nar/gkac963 (2023).
Ginestet, C. ggplot2: Elegant graphics for data analysis. J. R. Stat. Soc. A Stat. 174, 245–245. https://doi.org/10.1111/j.1467-985X.2010.00676_9.x (2011).
Subramanian, A. et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U. S. A. 102, 15545–15550. https://doi.org/10.1073/pnas.0506580102 (2005).
Liu, J. et al. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 173, 400–416. https://doi.org/10.1016/j.cell.2018.02.052 (2018).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Zheng, Y. & Heagerty, P. J. Semiparametric estimation of time-dependent ROC curves for longitudinal marker data. Biostatistics 5, 615–632. https://doi.org/10.1093/biostatistics/kxh013 (2004).
Foucher, Y. & Danger, R. Time dependent ROC curves for the estimation of true prognostic capacity of microarray data. Stat. Appl. Genet. Mol. Biol. https://doi.org/10.1515/1544-6115.1815 (2012).
Rhodes, D. R. et al. ONCOMINE: A cancer microarray database and integrated data-mining platform. Neoplasia 6, 1–6. https://doi.org/10.1016/s1476-5586(04)80047-2 (2004).
Bindea, G. et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39, 782–795. https://doi.org/10.1016/j.immuni.2013.10.003 (2013).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7. https://doi.org/10.1186/1471-2105-14-7 (2013).
Charoentong, P. et al. Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep. 18, 248–262. https://doi.org/10.1016/j.celrep.2016.12.019 (2017).
Ferris, R. L. et al. Nivolumab for recurrent squamous-cell carcinoma of the head and neck. N. Engl. J. Med. 375, 1856–1867. https://doi.org/10.1056/NEJMoa1602252 (2016).
Bai, Z. et al. Pyroptosis regulators exert crucial functions in prognosis, progression and immune microenvironment of pancreatic adenocarcinoma: A bioinformatic and in vitro research. Bioengineered 13, 1717–1735. https://doi.org/10.1080/21655979.2021.2019873 (2022).
Zhang, L. et al. The roles of programmed cell death ligand-1/ programmed cell death-1 (PD-L1/PD-1) in HPV-induced cervical cancer and potential for their use in blockade therapy. Curr. Med. Chem. 28, 893–909. https://doi.org/10.2174/0929867327666200128105459 (2021).
Li, F. et al. Identification of key biomarkers and potential molecular mechanisms in renal cell carcinoma by bioinformatics analysis. J. Comput. Biol. 26, 1278–1295. https://doi.org/10.1089/cmb.2019.0145 (2019).
Zhao, Q. J., Zhang, J., Xu, L. & Liu, F. F. Identification of a five-long non-coding RNA signature to improve the prognosis prediction for patients with hepatocellular carcinoma. World J. Gastroenterol. 24, 3426–3439. https://doi.org/10.3748/wjg.v24.i30.3426 (2018).
Bing, Z. et al. An integrative model of miRNA and mRNA expression signature for patients of breast invasive carcinoma with radiotherapy prognosis. Cancer Biother. Radiopharm. 31, 253–260. https://doi.org/10.1089/cbr.2016.2059 (2016).
Jumeau, C. et al. Expression of SAA1, SAA2 and SAA4 genes in human primary monocytes and monocyte-derived macrophages. PLoS ONE 14, e0217005. https://doi.org/10.1371/journal.pone.0217005 (2019).
Ho, I. C. & Miaw, S. C. Regulation of IL-4 expression in immunity and diseases. Adv. Exp. Med. Biol. 941, 31–77. https://doi.org/10.1007/978-94-024-0921-5_3 (2016).
Pinzon Grimaldos, A. et al. ANGPTL3 deficiency associates with the expansion of regulatory T cells with reduced lipid content. Atherosclerosis 362, 38–46. https://doi.org/10.1016/j.atherosclerosis.2022.09.014 (2022).
Sun, Y., Sedgwick, A. J., Palarasah, Y., Mangiola, S. & Barrow, A. D. A transcriptional signature of PDGF-DD activated natural killer cells predicts more favorable prognosis in low-grade glioma. Front Immunol 12, 668391. https://doi.org/10.3389/fimmu.2021.668391 (2021).
Li, W., Meng, X., Yuan, H., Xiao, W. & Zhang, X. A novel immune-related ceRNA network and relative potential therapeutic drug prediction in ccRCC. Front. Genet. 12, 755706. https://doi.org/10.3389/fgene.2021.755706 (2021).
Cai, H., Zhang, Y., Wang, J. & Gu, J. Defects in macrophage reprogramming in cancer therapy: The negative impact of PD-L1/PD-1. Front. Immunol. 12, 690869. https://doi.org/10.3389/fimmu.2021.690869 (2021).
Wang, Y. et al. Identification of PLAUR-related ceRNA and immune prognostic signature for kidney renal clear cell carcinoma. Front. Oncol. 12, 834524. https://doi.org/10.3389/fonc.2022.834524 (2022).
Plaza-Florido, A., Rodriguez-Ayllon, M., Altmae, S., Ortega, F. B. & Esteban-Cornejo, I. Cardiorespiratory fitness and targeted proteomics involved in brain and cardiovascular health in children with overweight/obesity. Eur. J. Sport Sci. https://doi.org/10.1080/17461391.2023.2167237 (2023).
Fei, H. & Chen, X. Development of a novel five-gene immune-related risk model for the prognosis evaluation of prostate adenocarcinoma patients. Am. J. Cancer Res. 12, 2337–2349 (2022).
Liang, J., Zhang, X., Wang, X., Yin, W. & Guo, Z. Pan-cancer analyses reveal the immunotherapeutic value of klotho. Heliyon 8, e11510. https://doi.org/10.1016/j.heliyon.2022.e11510 (2022).
Gao, X., Yang, J. & Chen, Y. Identification of a four immune-related genes signature based on an immunogenomic landscape analysis of clear cell renal cell carcinoma. J. Cell Physiol. 235, 9834–9850. https://doi.org/10.1002/jcp.29796 (2020).
Zhang, Y., Zou, J. & Chen, R. An M0 macrophage-related prognostic model for hepatocellular carcinoma. BMC Cancer 22, 791. https://doi.org/10.1186/s12885-022-09872-y (2022).
Finotello, F. & Trajanoski, Z. New strategies for cancer immunotherapy: Targeting regulatory T cells. Genome Med. 9, 10. https://doi.org/10.1186/s13073-017-0402-8 (2017).
Giraldo, N. A. et al. Orchestration and prognostic significance of immune checkpoints in the microenvironment of primary and metastatic renal cell cancer. Clin. Cancer Res. 21, 3031–3040. https://doi.org/10.1158/1078-0432.CCR-14-2926 (2015).
Brech, D. et al. Dendritic cells or macrophages? The microenvironment of human clear cell renal cell carcinoma imprints a mosaic myeloid subtype associated with patient survival. Cells https://doi.org/10.3390/cells11203289 (2022).
Yin, X. et al. Development of a novel gene signature to predict prognosis and response to PD-1 blockade in clear cell renal cell carcinoma. Oncoimmunology 10, 1933332. https://doi.org/10.1080/2162402X.2021.1933332 (2021).
Zhang, Z. et al. Construction of a novel gene-based model for prognosis prediction of clear cell renal cell carcinoma. Cancer Cell Int. 20, 27. https://doi.org/10.1186/s12935-020-1113-6 (2020).
Acknowledgements
We would like to thank the original data provided by the TCGA and GEO databases. J.G (CSC NO.201906370030) and Z.N.L(CSC NO.202108080086) are supported by a scholarship from the China Scholarship Council Program.
Author information
Authors and Affiliations
Contributions
J.G. and Z.N.L. designed the study. J.G., Z.N.L., X.B.Z., Z.Z.P., and Z.M.P. performed research and/or contributed to data analysis. J.G. and Z.N.L. wrote the original manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gu, J., Zhang, X., Peng, Z. et al. A novel immune-related gene signature for predicting immunotherapy outcomes and survival in clear cell renal cell carcinoma. Sci Rep 13, 18922 (2023). https://doi.org/10.1038/s41598-023-45966-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-45966-8
- Springer Nature Limited