Abstract
Bovine leukemia virus (BLV) is the causative agent of enzootic bovine leucosis (EBL), which has been reported worldwide. The expression of viral structural proteins: surface glycoprotein (gp51) and three core proteins - p15 (matrix), p24 (capsid), and p12 (nucleocapsid) induce a strong humoral and cellular immune response at first step of infection. CD4+ T-cell activation is generally induced by bovine leukocyte antigen (BoLA) region– positive antigen-presenting cells (APC) after processing of an exogenous viral antigen. Limited data are available on the BLV epitopes from the core proteins recognized by CD4+ T-cells. Thus, immunoinformatic analysis of Gag sequences obtained from 125 BLV isolates from Poland, Canada, Pakistan, Kazakhstan, Moldova and United States was performed to identify the presence of BoLA-DRB3 restricted CD4+ T-cell epitopes. The 379 15-mer overlapping peptides spanning the entire Gag sequence were run in BoLA-DRB3 allele-binding regions using a BoLA-DRB- peptide binding affinity prediction algorithm. The analysis identified 22 CD4+ T-cell peptide epitopes of variable length ranging from 17 to 22 amino acids. The predicted epitopes interacted with 73 different BoLA-DRB3 alleles found in BLV-infected cattle. Importantly, two epitopes were found to be linked with high proviral load in PBMC. A majority of dominant and subdominant epitopes showed high conservation across different viral strains, and therefore could be attractive targets for vaccine development.
Similar content being viewed by others
Introduction
Bovine leukemia virus (BLV) is the etiological agent of enzootic bovine leucosis (EBL), a chronic, lymphoproliferative disease associated with persistent lymphocytosis and B-cell lymphomas1. BLV, together with human T-cell leukemia viruses type 1 and 2 (HTLV-1, HTLV-2), belong to the genus Deltaretrovirus of the family Retroviridae. BLV infection has a worldwide distribution and causes substantial economic losses in the livestock industry2,3, infection with this virus result in a negative effect on dairy production and cow longevity, which is very likely based on the resulting impaired immune function following infection3,4,5.
The complex BLV genome encodes structural genes (env, gag, pol/pro) and nonstructural, regulatory genes (tax, rex). The env gene gives rise to two glycoproteins: extracellular surface subunit (SU, gp51) implicated in receptor recognition and virion attachment, and transmembrane subunit (TM, gp30) responsible for anchoring the SU-TM complex into lipid bilayers (reviewed in6).
The gag gene encodes the internal structural polyprotein- Gag (group-specific antigen), responsible for initiating the process of virion budding from the infected cell and RNA packaging in the viral particle formation process7,8. During viral maturation, the precursor Gag is processed into three separate proteins: matrix (MA, p15), capsid (CA, p24) and nucleocapsid (NC, p12), which undergo substantial conformational rearrangements—and confer infectivity to virions9. The tax gene encodes Tax protein, involved in activation of transcription of viral mRNA.
Both humoral and cell-mediated immune responses strongly limits BLV replicative cycle in cows naturally infected with the virus10,11,12,13,14. With respect to cellular immunity, CD4+ T-cell response to BLV, mediated through recognition of short viral peptides presented by bovine leukocyte antigen (BoLA) class II molecules on the surface of antigen-presenting cells (APC), is required for multiple anti-viral processes. These anti-viral processes confer protection from progression to persistent lymphocytosis and tumor development during BLV infection15,16. The CD4+ T-cell response can be directed toward any virion protein; yet, many studies into T cell immunity to BLV have focused on the external glycoprotein gp51. Indeed, existing molecular studies on CD4+ T-cells and BLV are limited to the gp51 CD4+ T-cell epitopes: peptide 98–117, peptide 169–188, and peptide 177–19217; peptide 51–70 and peptide 61–8018; peptide 98–11719. Recently, CD4+ T-cell epitopes on gp30 protein of the env gene were mapped (peptide gp30N5, peptide gp30N6 and peptide gp30N7)19. Furthermore, peptide 131–150 and peptide 111–13020; and peptides tax16/17, peptides tax19/20 and peptides tax 22–2419, were recognized as T-cell epitopes for Tax. To date, there are no structurally defined CD4+ T-cell epitopes from the internal gag proteins–MA (p15) and NC (p12). Experimental data regarding Gag polypeptide are limited to CA (p24). An in vitro study based on lymphocyte proliferation assays two non-universal CD4+ T-cell epitopes: peptide 31–55 and peptide 141–165 were defined13.
Resistance or susceptibility to progression of BLV infection in cattle was linked to the polymorphism in the BoLA class II gene21,22. Cattle express only two BoLA class II proteins, DR and DQ23. The only source of diversity in DR molecules is from the polymorphic DRB chain24. So far, 384 BoLA-DRB3 alleles have been identified according to the BoLA Nomenclature Committee of IPD-MHC Database [Available from: https://www.ebi.ac.uk/ipd/mhc/group/BoLA/species/]. Polymorphisms in the BoLA-DRB3 gene can influence immune response by peptide binding, antigen presentation, and T-cell receptor (TCR) gene sequences in a T-cell population and group of cytokines that mediate and regulate immunity25,26. Furthermore, the affinity of BLV epitopes to bind to certain BoLA-DRB3 molecules was suggested to correlate with the capacity to induce T-cell proliferation27,28. However, the BLV epitope peptides responsible for this binding remains unknown.
The aim of this study was to determine epitopes on Gag-derived proteins (MA, CA and NC) binding to different BoLA-DRB3 alleles and link them with blood proviral load (PVL) levels of BLV-infected cattle.
Results
BoLA-DRB3 allele genotyping
Out of 125 samples, 113 identified two alleles, 11 identified one allele, and genotyping failed in one sample. A total of 73 different BoLA-DRB3 alleles were identified (Table 1). Alleles with the highest frequencies were DRB3*01:01, *11:01, *10:01 and *15:01 with respective percentage values of 15.7%, 8.4%, 8.0% and 8.0%. Thirteen alleles (*12:01, *14:01:01, *27:03, *07:01, *18:01, *41:01, *105:02, *116:01, *160:01, *09:02, *24:33, *25:01:01 and *57:02) had frequencies ranging from 1.7% to 3.8%; 20 alleles (DRB3*09:01,*44:01, *107:01, *107:04, *130:01, *134:01,*139:01, *142:01, *02:01, *04:01, *05:02, *05:03, *08:01, *09:04, *13:01, *15:04, *17:01, *24:03, *31:01 and *20:01:01) had frequencies ranging from 0.8% to 1.3%; and the remaining 36 alleles had frequencies below 0.5% (Table 1).
Analysis of amino acid sequence variability of the Gag protein
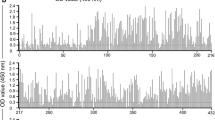
The Shannon entropy (Hx) plot exhibited 49 peaks with values ranging from 0.05 to 1.13 (Supplementary Fig. S1, Supplementary Table S1). Considering the three Gag domains, the highest total entropy (6.33) occurred in the matrix (MA). For the capsid (CA) and nucleocapsid (NC) domains, the total entropy was 4.09 and 1.38, respectively. A detailed multiple sequence alignment analysis on 395 amino acid sites indicated 345 as conserved and 49 with non-synonymous single nucleotide polymorphisms (nsSNPs). A substantial number of nsSNPs found in the MA and CA domains suggested the possibility of positive selection on variable sites of the protein (Supplementary Fig. S2). The dN/dS ratios were drawn over the midpoint window position (window length 9, step size 3) from the whole coding region. The following regions with putative positive selection sites were identified: 133–149 nt, 175–195 nt, 199–216 nt, 250–279 nt, 316–342 nt in MA, 829–843 nt, 952–972 nt in CA and 1087–1107 nt for NC domains, respectively. Thirteen codons located in these regions had dN/dS ratios > 1 that identified them as major sites for the occurrence of positive selection. These were codons 48, 61, 63, 69, 87, 88, 108, 109, 112 in MA domain; 278, 318, 323 in CA; and 365 in NC (Supplementary Table S2 and Supplementary Fig. S1).
Determination of epitope peptides in the Gag protein consensus sequence based on BoLA-DRB3
In order to search for common epitope peptides on Gag protein, we determined the consensus sequence from 125 sequences described in this study. To detect putative binding sites for BoLA-DRB3, we used the 379 15-mer overlapping peptides that spanned the entire Gag consensus sequence in 73 BoLA-DRB3 allele binding regions in NetBoLAIIpan (pan-specific predictor for BoLA-DRB3 Ag presentation). Analysis revealed 22 putative regions within Gag proteins with high binding affinity to BoLA-DRB3 alleles. The binding affinities of the epitopes and complete calculations are presented in Supplementary Table S3.
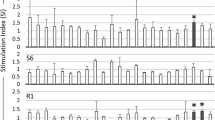
Detailed analysis indicated that the highest number of BoLA-DRB3 alleles, 44 out of 73 (60.3%), had significant binding affinity to peptide 320#QPAILVHTPGPKMPG and additionally to 5 overlapping peptides sharing the same ILVHTPGPK core sequence (as shown in Table 2). A combined sequence 317-KIKQPAILVHTPGPKMPGPR-336, formed by the group of these peptides, was designated epitope 1A (as shown in Fig. 1 and Supplementary Fig. S3). The same number of BoLA-DRB3 alleles—44 out of 73 (60.3%)—with high binding affinity were determined for peptide 257#VNRLQISLADNLPDG and to a lesser extent, for 6 overlapping peptides sharing the common core sequence LQISLADNL. Summary sequence 255-EFVNRLQISLADNLPDGVPKE-275 for a group of the peptides was designated epitope 1B. The third most frequent binding alleles, 42 out of 73 (57.5%), were peptide 296#GRGLVAAPVGQKLQA and 5 other contiguous peptides with the common core sequence LVAAPVGQK. The completed sequence 293-ILQGRGLVAAPVGQKLQACA-312 for a group of these peptides was designated epitope 2. The fourth peptide with significant binding affinity for multiple alleles, 38 out of 73 (52.1%), was peptide 249#PAESYVEFVNRLQIS and 6 other contiguous peptides. Deduced sequence 247-QGPAESYVEFVNRLQISLADN-267, based on those peptides with common core sequence VEFVNRLQI, was designated as epitope 3. Additionally, a large number of BoLA-DRB3 alleles, 32 out of 73 (43.8%), had significant binding affinity to the 40#LKNYIHWFHKTQKKP, 165#QLCQYIASPVDQTAH and 147/148#QTLRLAILQADPTPAD main peptides. Summary sequences 38-TDLKNYIHWFHKTQKKPW-55, 163-LEQLCQYIASPVDQTAH MTS-182 and 144-VWIQTLRLAILQADPTPADLE-164 were respectively designated epitopes 4A, 4B and 4C.
Distribution of the BoLA-DRB3-restricted CD4+ T-cell epitopes along the Gag polyprotein. The labeled blue bars in the upper part of figure refer to the identified 22 epitopes 1A–15B. The figure shows the localization of the epitopes for the most commonly detected BoLA-DRB3 (on the left side of the figure). The distribution of the epitopes for the all analysed alleles is shown in Supplementary Fig. S3.
The eighth peptide frequently bound by BoLA-DRB3 alleles, 27 out of 73 (37.0%), was sequence 72#RVPLVLATLNEVLSN plus 5 adjacent peptides with common core sequence LVLATLNEV. Completed sequence 69-KFGRVPLVLATLNEVLSNDE-88 for the group of peptides was designated epitope 5. The ninth peptide 42#NYIHWFHKTQKKPWT, for which 24 of 73 (32.9%) different alleles showed binding affinity together with 3 overlapping peptides with common core sequence FHKTQKKPW, created sequence 42-NYIHWFHKTQKKPWTFTS-59, which was designated epitope 6. The tenth peptide 225#LWLQAWKNLPTRPSV, to which 23 of 73 (31.5%) BoLA-DRB3 alleles had binding affinity together with the other 6 peptides with common core sequence WKNLPTRPS, represented the combined sequence 222-YQNLWLQAWKNLPTRPSVQPW-242 designated as epitope 7. The eleventh peptide 346#GPCYRCLKEGHWARD showing binding affinity for 16 of 73 alleles (21.9%), together with three adjacent peptides with YRCLKEGHW core sequence, presented summarized sequence 344-PPGPCYRCLKEGHWARDC-361 and was designated epitope 8 (Table 2, Fig. 1). The remaining eleven peptides (184#TAAIAAAEAANTLQG, 128#LQDIKKEIENKAPGS, 52#KKPWTFTSGGPASCP, 103#YDPPAVLPIISEGNR, 235/236#TRPSVQPWSTIVQGPA, 177#TAHMTSLTAAIAAAE, 141#GSQVWIQTLRLAILQ, 238/239#SVQPWSTIVQGPAESY, 116#NRNRHRAWALRELQD, 15/16#PSDWLNLLQSAQRLNP and 180#MTSLTAAIAAAEAAN) formed epitopes respectively named 9-10 (183-SLTAAIAAAEAANTLQG-199, 126-LRELQDIKKEIENKAPGS-143), 11A-11B (50-T QKKPWTFTSGGPASCP-66, 102-PPYDPPAVLPIISEGNRNR-120), 12A-12B (235-PTRPSVQPWSTIVQGPA-251, 176-DQTAHMTSLTAAIAAAE-192), 13A-13B (142-GSQVW IQTLRLAILQAD-158, 238-PSVQPWSTIVQGPAESYVE-256), 14 (117-NRNRHRAWALRELQDI-132) and 15A-15B (15-PSDWLNLLQSAQRLNP-30, 181-MTSLTAAIAAAEAAN-195) were subdominant. These epitopes characterized a high BoLA-DRB3 binding specificity and were consequently dedicated to a small group of alleles (not exceeding 18% of all alleles in this study) (Table 2, Supplementary Fig. S3).
A total of 22 CD4+ T-cell epitopes were identified. Out of 22, five epitopes (15A, 4A, 6, 11A and 5) were located in the matrix; one epitope (11B) within the matrix-capsid; 14 epitopes (14, 10, 13C, 4C, 4B, 12B, 15B, 9, 7, 12A, 13B, 3, 1B and 2) in the capsid; one epitope (1A) at the capsid-nucleocapsid; and one epitope in the nucleocapsid domain (8) (Supplementary Fig. S2). The majority of these epitopes could be considered vaccine candidate antigens as they are exposed on the surface (a topological approach was presented in Fig. 2A–D).
(a–d) Representation of the dominant and subdominant CD4+ T cell epitopes on 3D protein structure model of BLV Gag. The model is shown as space-filled images of opposite sides arbitrarily named side A and side B. (A–B) Gag structure contain 11 dominant epitopes: 1A (red), 1B (forest green), 2 (blue), 3 (yellow), 4A (cyan), 4B (orange), 4C (green), 5 (cornflower blue), 6 (magenta), 7 (goldenrod), 8 (sky blue), 1B + 3 [255–267] (brown), 4A + 6 [42–55] (purple), 4B + 4C (163–164): silver. (C–D) Gag structure contain 11 subdominant epitopes: 9 (red), 10 (forest green), 11A (blue), 11B (yellow), 12A (cyan), 12B (orange), 13A (green), 13B (cornflower blue), 14 (magenta), 15A (goldenrod), 15B (all residues overlap with one or more other sequences), 9 + 15B [192–194] (brown), 12B + 15B [180–181] (purple), 10 + 13A [141–142] (silver), 11B + 14 [116–119] (navy blue), 12A + 13B [237–250] (coral), 10 + 14 [125–131] (dark red), 9 + 12B + 15B [182–191] (black).
Relation between the incidence of BoLA-DRB3 alleles and number of CD4+ T-cell epitopes
Predictions of BoLA-DRB3 peptide’s binding affinity were performed for 73 alleles determined in BLV-infected cattle tested in this study. The results shown in Supplementary Table S4 indicate that the examined alleles had significant binding specificity to the epitope peptides. BoLA-DRB3 molecules were found to interact with binding core sequences of 3–11 CD4+ T-cell epitopes of Gag protein. Notably, the first ten BoLA-BRB3 alleles distinguishable by binding the lowest number of epitopes were distributed as follows: *01:01, *11:01, *11:02, *20:01:01 and *112:02 had affinity to three epitopes; *116:01, *134:01, *31:01, *114:01 and *14:04 had affinity to four epitopes (Supplementary Table S4). The combined incidence of these alleles was 72 out of 237 possible pairs (30.4%). Conversely, the first ten alleles distinguishable by binding affinity for the highest number of Gag epitopes were *006:01 (eleven epitopes); *03:01:01 (ten epitopes); *24:32, *24:33, *57:02, *005:08 and *80:01 (nine epitopes); and *09:01, *02:01 and *24:03 (eight epitopes) (Supplementary Table S4). In total, the incidence of these alleles was 21 out of 237 (8.9%). These results indicate that it needs to be determined if there is a relation with the number of binding sites versus susceptibility for BLV or progression to clinical disease.
Next, the numbers of epitopes were analyzed with respect to proviral copy numbers in BLV-infected cattle. Figure 3 depicts results of such analysis and demonstrates that the number of CD4+ T-cell epitopes on Gag protein identified for different BoLA-DRB3 alleles is not significantly correlated with BLV proviral load (R2 = 0.0231, P value = 0.0793, n = 113).
Association between BoLA-DRB3 alleles, BoLA-DRB3-restricted CD4+ T-cell epitopes and BLV proviral load
We tested 73 alleles for their affinity to particular epitopes and BLV proviral load. Supplementary Fig. S4 shows the distribution of all detected epitopes within Gag protein and their binding affinity for 73 BoLA-DRB3 alleles identified in the current study. Based on literature data, BoLA-DRB3*015:01 and *012:01 alleles are known susceptibility-associated markers related to high PVL in blood cells (> 10,000 per 105), and cattle with susceptible alleles may be at a high risk of BLV transmission via direct contact with healthy cows. In contrast, BoLA-DRB3*09:02, *02:01 and *014:01:01 alleles comprise resistant markers associated with the development of low PVL in blood cells (< 10,000 per 105), and cattle with resistant alleles may be low-risk spreaders for BLV transmission29,30,31,32,33. To determine possible associations of BoLA-DRB3 alleles related to susceptibility or resistance to BLV proviral load with BoLA-DRB3-restricted CD4 + T-Cell epitopes, the five (5/73) BoLA-DRB3 alleles like *012:01, *012:03, *015:01, *015:05, *016:01 were placed in a single group (Group A) and marked in red. Group A contained those BoLA-DRB3 alleles associated with a high BLV copy number, as previously described. The five BoLA-DRB3 alleles like *09:02, *09:01, *09:04, *02:01 and *14:01:01 were compiled as a second group (Group B) marked in gray and consisted of the alleles previously correlated in BLV-infected cattle with low proviral load.
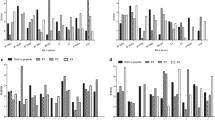
As a result of the analysis, none of Group A alleles had affinity for epitopes 1A and 2 (Fig. 4), in contrast to group B alleles’ significant affinity for epitopes 1A and 2. For other epitopes, allele binding from the two groups did not differ and no other patterns were observed between the two groups as far as binding affinity of alleles to these epitopes (Supplementary Fig. S4). Subsequently, the affinity or lack of affinity for epitopes 1A and 2 was determined for the remaining 63 alleles (63/73). As a result, 17 alleles (17/63) were assigned to Group A, (*005:02, *005:04, *005:07, *010:01, *010:03, *010:04, *011:01, *011:02, *017:01, *020:01:01, *024:15, *041:01, *044:01, *086:03, *105:02, *139:01 and *142:01), and remaining 46 alleles (46/63) were classified to group B (*001:01, *003:01, *004:01, *005:03, *005:08,*006:01, *007:01, *008:01, *13:01, *14:03, *14:04, *15:04, *18:01, *19:02, *24:03, *24:32, *24:33, *25:01, *27:03, *27:13, *27:18, *28:01, *28:05, *31:01, *32:01, *35:01, *38:01, *43:03, *45:01, *57:02, *75:03, *80:01, *81:01, *86:02, *89:01, *107:01, *107:04, *112:02, *112:03, *114:01, *116:01, *130:01, *134:01, *141:01, *157:01 and *160:01. Thus, one might infer that Group A alleles’ lack of affinity for epitopes 1A and 2 is related to the number of BLV proviral copies in BLV-infected cattle. Therefore, we conducted a BLV copy number comparison of cattle carrying at least one Group A allele and those with only Group B alleles. Statistical analysis using the student t-test showed that cattle carrying one or both Group A alleles (with no affinity for 1A and 2 epitopes) had a significantly increased number of BLV proviral copies per 1000 cells, as opposed to Group B alleles (t-value = 2.06255, P value = 0.040269) (Fig. 5 and Supplementary Table S5).
Association between 73 BoLA-DRB3 alleles and two BoLA-DRB3- restricted CD4 + T cell epitopes (1A and 2). Log2 Rank predicted binding score for Gag peptides observed for the BoLA-DRB3 alleles distinguished in the two groups: Group A (n = 22 alleles) marked in red line on the graph; Group B (n = 51 alleles) marked in grey line.
Changes in amino acid sequence of Gag vary BoLA-DRB3-peptide binding affinity
The 13 codons, which were identified as the major sites with a process of positive selection were evaluated for epitope affiliation. Out of these 13, 11 codons (84.6%) were located in the following epitopes: codon 48 in epitopes 4A and 6; codons 61 and 63 in epitope 11A; codons 69, 87 and 88 in epitope 5; codons 108, 109 and 112 in epitope 11B; and codons 318 and 323 in epitope 1A. Therefore, to assess the observed mutations’ impact on the binding affinity of BoLA-DRB3, the individual sequences of 125 isolates with corresponding BoLA-DRB3 genotypes were submitted for analysis using NetBoLAIIpan. As result, 58 out of the 125 BLV strains had different amino acid changes (n = 25) in the epitope sequences that was predicted to alter the binding affinity of BoLA-DRB3 to the epitopes. Out of 25, 13 changes were located in positive selection sites (H48R, H48Y, G61S, A63T, A63V, K69R, D87E, E88G, V108I, L109M, I112V, I318M, I323V), and an additional 9 other changes (V76I , A78T, E82D, E82K, D104N, A189T, A193T, A250T, V254I) have predicted a significant effect on the binding affinity of BoLA-DRB3 in combination with different BoLA-DRB3 alleles (Supplementary Table S6). Detailed descriptions of the changes on epitope peptides, and the BoLA-DRB3 epitope binding level are shown in Table 3 and Fig. 6.
Gag protein sequence alignment for selected BLV isolates, containing amino acid changes within CD4+ T cell epitopes that change the degree of binding affinity of BoLA-DRB3. The names of the isolates and their corresponding BoLA-DRB3 are listed on the left side of the alignment. Amino acid changes, which generate new BoLA-DRB3 binding affinity site are marked with red arrows; amino acid changes that impair BoLA-DRB3 binding affinity sites are marked with green arrows; changes that enhance BoLA-DRB3 affinity are marked with orange arrows, the changes that generate lack of peptides interactions with BoLA-DRB3 are marked with grey arrows. BoLA-DRB3-restricted CD4 + T-cell epitopes along the Gag polyprotein are labeled in the upper part of the figure as blue bars.
Analysis of sequence conservation in the binding core of the CD4 + T-cell epitopes
The binding core is the anchoring region of the epitope and is defined as the central nine amino acid (aa) sequence of the predicted 15-mer epitope that is flanked by three aa residues on the N- and C-terminal ends. Therefore, the conservation of the 22 identified putative CD4+ T-cell epitopes was evaluated based on the number of changed amino acid residues in their binding-core sequences (Table 4). The most conserved epitope cores were present in 1B (CA), 4B (CA), 4C (CA), 7 (CA), 8 (NC), 12A (CA), 13A (CA), 14 (CA) and 15A (MA) epitopes, with complete sequence conservation in 125 sequences (100%). Core sequences of epitopes 15B (CA), 2 (CA), 10 (CA) and 12B (CA) showed the second highest level of conservation, in the range of 96.8% (in 121 of 125 sequences) to 99.2% (in 124 of 125 sequences), respectively. Importantly, the mutations found in these core sequences were predicted to no affect the epitope binding affinity to the BoLA-DRB3 molecules (Table 3). Epitopes 1A (CA/NC), 5 (MA), 9 (CA), 3 (CA) and 13B (CA) showed the conservation in the range of 87.2% (in 109 of 125 sequences) to 92.8% (in 116 of 125 sequences), respectively, and contained the mutations I323V, V254I, A78T, E82D, V76I, A193T, and A25T, which may affect epitope binding affinity to certain BoLA-DRB3 molecules. The variable epitopes were 11A (in 59 of 125 MA sequences, 47.2%), 4A and 6 (in 94 of 125 MA sequences, 75.2%) and 11B (in 98 of 125 MA/CA) sequences, 78.4%), containing amino acid changes H48R/Y, G61S, A63V/T, L109M, I112V and V108I, which were predicted to affect binding to BoLA-DRB3.
Discussion
The utilisation of bioinformatics to identify T-cell responses to retroviral infections has increased over the last few years34,35,36,37,38. The in silico prediction methods NetMHCIIpan and NetBoLAIIpan developed to predict HLA and BoLA class II restricted peptide binding, respectively, have proven to be among the best methods currently available35,39,40. Here, we used NetBoLAIIpan prediction method to determine BoLA-DRB3- restricted BLV peptides on p15, p24 and p12 of the Gag polyprotein, with broad BoLA allelic coverage. Of all tested epitope candidates, 11 top-scoring epitopes (1A, 1B, 2, 3, 4A, 4B, 4C, 5 -8) were selected to identify broadly reactive BLV-specific CD4+ T-cell responses, by up to 60% of the analysed BoLA-DRB3 alleles. Another 11 subdominant epitopes (9, 10, 11A, 11B, 12A, 12B, 13A, 13B, 14, 15A, 15B) were predicted to be more restricted to one or more of the donor’s BoLA class II alleles, up to 17.8% of all examined BoLA-DRB3 alleles.
These identified epitopes often overlapped, thus creating long regions within the Gag protein that had BoLA-DRB3 binding affinity. In MA, dominant epitopes occupied 40%, in CA 57% and in NC 47% of the protein sequences. Importantly, the Gag peptide-binding motifs were detected for all bovine BoLA-DRB3 proteins (n = 73) recognized for the 125 cows used in the current studies. Each of the alleles had at least three of the identified target epitopes. The presented data demonstrates a high promiscuity of Gag protein to BoLA-DRB3. BoLA-DRB3-restricted CD4+ T-Cell epitopes 1A and 1B were considered the most promiscuous binders as they contained binding cores of epitopes predicted for 44 different BoLA-DRB3 types. They were located within a conserved part of the p24 region, C-terminal domain, which is required for capsid dimerization, Gag oligomerization and viral formation41,42,43. Moreover, the binding core sequences of the other 12 epitopes (60% of all identified epitopes) had a high conservancy—from 99.2 to 100%. These results show that the majority of the detected epitopes provide a solid guide for vaccine development.
The two previously defined regions of p24 protein, being recognized by specific T-lymphocytes, represented by amino acids 31-PGSQVWIQTLRLAILQADPTPADLE-55 and 141-AESYVEFVNRLQISLADNLPDGVPK-16513 were compared with the newly identified epitopes in this study. The first epitope (residues 31–55) region corresponded to T-cell epitopes 4C (144-VWIQTLRLAILQADPTPADLE-164), 13A (142-GSQVWIQTLRLAILQAD-158) and 10 (126-LRELQDIKKEIENKAPGS-143) (specific residues are underlined). The second epitope (residues 141–165) region corresponded to T-cell epitopes 1B (255-EFVNRLQISLADNLPDGVPKE-275), 3 (247-QGPAESYVEFVNRLQISLADN-267), and part of 13B (238-PSVQPWSTIVQGPAESYVE-256). We noted that the previously defined epitopes were nested within newly identified epitopes. Therefore, we strongly suggest that these regions may be alternatively extended by the amino acids contained in the epitopes defined by BoLA-DRB3 II peptide-binding prediction method. Interestingly, in contrast to the second epitope (residues 141–165) region, epitopes 1B, 3 and 13B cover the full length of the major homology region (MHR) 244-IVQGPAESYVEFVNRLQISL-263, which was found to be essential for the stability and folding of the monomer, and hence for viral assembly, maturation and infectivity. This region is conserved throughout the whole retrovirus group and thus offers a novel and stable target for viral vaccines.
In the current study, we detected a greater number of BoLA-DRB3-restricted epitopes than Mager and coworkers13. This is likely related to the fact that the previously defined epitope regions are based on PBMCs from only four animals, which represented a particular allele of the BoLA-DRB3 genes interacting with the two specific p24 regions. The type of BoLA-DRB3 alleles in the tested PBMCs in prior published experiments are not defined. Nevertheless, based on our data, it appears that there indeed exist BoLA-DRB3 alleles which have affinity only for one or the other identified epitope regions; there are inter alia: BoLA-DRB3 *007:01, *009:04, *020:01:01 for residues 31–55 and BoLA-DRB3 *002:01, *004:01, *008:01, *010:03, *015:01, *018:01, *025:01:01, *043:03, *081:01, *116:01 and *160:01 for residues 141–165.
We observed that a higher number of Gag protein epitopes recognized by certain BoLA-DRB3 alleles accompanied the alleles associated with BLV resistance in cattle. Interestingly, relatively few of them were observed in the population of BLV-infected animals. Likewise, fewer epitopes recognized by particular BoLA-DRB3 alleles were associated with the BLV susceptible alleles. Noteworthy, the percentage of these alleles in the examined population of virus-infected cows was relatively high. Indeed, the affinity for the interaction of certain BoLA–DRB3 alleles with a longer region of the Gag protein (where epitopes overlap) or more Gag regions may elicit a stronger cellular response. Thus, our results confirm the hypothesis that disease-susceptible cattle may have fewer epitopes than resistant cattle, resulting in weaker immune responses. Moreover, these results indicate a significant role of bovine MHC II polymorphisms in the mapping of BLV epitopes recognized by CD4+ T-cells on viral proteins.
Bai and coworkers studied gp51, gp30 and Tax protein epitopes related to the BoLA–DRB3 genotype and found that fewer CD4+ T-cell epitopes were observed in susceptible cattle than in resistant cattle19. Takeshima and colleagues suggested that the BoLA-DRB3 gene may regulate both antigen epitope recognition and the magnitude of the antigen-specific T-cell response that is processed after exposure to infection23. Accordingly, our studies confirm that BLV antigens are restricted according to BoLA-DRB3, and that genotyping of cattle is important for determining antigenic epitopes recognized by the bovine immune system.
In this work, we also analyzed whether the number of BoLA-DRB3-restricted epitopes in Gag protein is related to the number of BLV proviral copies in PBMCs in the analyzed DNA samples; however, we did not find a significant correlation. Additionally, Bai and colleagues observed that the number of CD4+ T-cell epitopes was positively related to proviral load, which depended on the BoLA class II genotype19. This discrepancy may be due to the fact that the current study used a 25-fold larger group of cattle for the analysis. It is well known that the BLV proviral load varies greatly as it is the result of many different factors such us the time of exposure to the virus, biochemical and hematological factors of the cow, or the age of the cow, to name a few, which can generate erroneous results when experiments are conducted on a small number of animals44,45,46. In comparison, another retrovirus, HIV, specifically targets the HIV Gag peptides by CD4+ T-cells has been associated with lower viremia in both adults and children47,48,49. Ranasinghe and coworkers demonstrated an inverse correlation between viral load and the number of Gag peptides targeted by CD4+ T-cells50. Buggert and coworkers confirmed this finding, suggesting that broadly reactive Gag-specific CD4+ T-cell responses could have an impact on HIV disease progression51. However, whether the frequent targeting of Gag peptides is the cause or the consequence of the reduced viremia remains to be clarified.
Nevertheless, an association polymorphisms of the BoLA-DRB3 gene with BLV PVL is described in the literature23,31,45,52,53. Published data indicates some BoLA-DRB3 alleles such as *15:01, *12:01 and *16:01 are associated with high PVL in BLV-infected cattle but BoLA-DRB3 alleles like *09:02, *02:01 and *14:01:01 are associated with low PVL29,30,31,32. Of the 22 epitopes, we found two epitopes—1A (317-KIKQPAILVHTPGPKMPGPR-336) and 2 (293-ILQGRGLVAAPVGQKLQACA-312) that were significantly related to cattle resistant to developing high BLV proviral load. These epitopes were located in CTD-CA and between CA/NC proteins, respectively, highly conserved regions for retroviruses (Supplementary Fig. S6). In addition, these epitopes were broadly recognized for most of the BoLA-DRB3 alleles (70%). Interestingly, epitopes 1A and 2 were not recognizable by the types of BoLA-DRB3 alleles, which were previously reported in the literature as being associated with the development of subclinical infection and high BLV PVL29,30,31,54,55. Some of these have never been investigated for PVL dependence therefore, additional functional studies are required to further confirm these findings. Nevertheless, in the case of HIV, there are certain epitopes that determine resistance to infection56. Our results suggest that the 1A and 2 epitopes may have a key and powerful effect in inducing a strong cellular response and fighting BLV within the host. It is noteworthy that epitopes 1A and 2 were the strongest epitopes to which the most alleles bound. Therefore, they seem to be an indispensable element that would be instructive in the design of synthetic peptide vaccine.
Antigenic variation within T-cell epitopes has been demonstrated for HIV-1, and this 'antigenic escape' may be responsible for viral persistence. Generally, although external proteins are highly immunogenic, antigenic shift limits their capacity to provide cross-protective immunity to novel viral strains. In contrast, the internal proteins are more conserved and may better mediate cross-protective T-cell responses57,58.
BLV exhibits less genetic variation among strains as compared with most other retroviruses, and the genomes of viruses isolated from multiple countries around the world share approximately between 94.5 and 99.5% of their nucleotide sequences. However, variation within the sequences encoding the Gag protein is poorly characterized. In our study the pairwise identity for 125 gag nucleotide sequences was 97.3%. Despite the internal proteins MA, CA and NC that exhibit higher levels of conservation relative to SU (gp51), sequence variation was still present, in which most sequence variation can be attributed to a single mutation. Based on the resulting proviral mutation profile, we revealed that the mutations are driven by immune selection pressure, suggesting mechanisms of positive selection and mutation hotspots.
Diversified positions were preferentially located within bovine CD4+ T-cell epitopes. Of 13 hotspots, 7 were located in the 9-mer core epitopes and had predicted a significant effect on the binding affinity of BoLA-DRB3 molecules. This is consistent with what is known about the peptide-binding core of epitopes that primarily interact with the BoLA-DR antigen binding groove. The peptide–BoLA-DR binding affinity is primarily determined by the amino acid sequence of the peptide binding core59. However, it has been shown that peptide flanking regions (PFRs) on either side of the binding core affect peptide–BoLA-DR binding and thereby ultimately also influence the peptide immunogenicity40. Indeed, 4 hotspots were located in PFRs and have input on binding affinity. These mutations may upend the presentation of virus-derived peptides via BoLA-DR. Based on the obtained results, certain mutations reduced while other mutations increased the affinity to bovine MHCII. Additionally, some mutations exhibited a neutral affinity. On the basis of our analysis of mutations, we selected 12 mutant peptides with predicted decreased BoLA-DR-binding strength 19 and mutant peptides with predicted increased BoLA-DR-binding strength for further biophysical and functional analyses. Our study provides evidence that single nonsynonymous mutations in BLV can subvert the immune response to CD4+ T-cell epitopes.
Our hypothesis was that the substitution of single amino acids in CD4+ T-cell epitope may influence the BoLA-DRB3 binding affinity and that nsSNP might be associated with variations in individual immune responses to antigens and susceptibility or resistance to disease. There are therefore many factors that make it difficult to predict peptide binding affinities to BoLA-DRB3 molecules, including the polymorphic sites of Gag epitopes. However, without functional analysis, the impact of single anchor residue substitutions on the response of CD4+ T-cells is still unclear. This study does not allow direct conclusions to be drawn concerning potential selection pressures, which shape the mutational landscape of CD4+ T-cell epitopes. This would invariably involve accounting for the BoLA-DRB3 genotype of all individuals from whom BLV genomes were sequenced. Moreover, how T-cell escape mutations within BLV are maintained during virus transmission between individuals with differing BoLA types and how viruses carrying epitope mutations affect disease severity requires further investigation.
Many CD4+ T-cell epitopes for BLV have been described in this study. The CD4+ T-cell response against BLV was associated with broad epitope recognition of, on average, 6 CD4+ T-cell epitopes per antigen per BoLA-DRB3 allele, which raises the question whether and how mutations in single epitopes affect virus control60. This may be of particular importance for BLV subunit vaccines to induce responses against an unlimited number of CD4 epitopes. These results highlight the capacity of BLV to evade cellular immune responses through sporadically emerging mutations in BoLA-BRB3 epitopes.
Taking into account the very conservative and wide range of identified epitopes and, on the other hand, the lack of progress in obtaining an effective vaccine, the new discovery has a high chance of success. The new vaccine could be an important element in protecting herds against BLV infections, especially in dairy cattle, where this category of cattle is especially susceptible to BLV infection61. Moreover, preventive vaccinations based on selected peptide immunogens could become an integral part of BLV eradication programs62,63. Finally, the importance of immunopeptidomics should be emphasized in subsequent studies taking into account other exotic and local cattle breeds as well as the circulation of endemic BLV variants.
Conclusions
In the present study, BLV Gag protein was characterized by immunoinformatic techniques to identify potential T-cell epitopes. Twenty-two BoLA-DRB3 class II epitopes were available across the entire BLV Gag polyprotein, however the p24 protein was identified as the main target for recognition by antigen-specific CD4+ T-lymphocytes. The thirteen broadly conserved BoLA-DRB3-restricted CD4+ T-cell epitopes shared between BLV isolates from different countries and 9 epitopes with changes in the binding core were identified. Among them two promiscuous conserved pBoLA-(gag)peptides, 1A and 2, related to hosts that mounted a successful host–pathogen immune response (animals with low proviral load) were discovered. We believe the newly-identified pBoLA-(gag) peptides, together with additional peptides that have been shown within gp51, gp30 and Tax proteins, will be important for inclusion in a multivalent antigen peptide vaccine for BLV that can provide protection against BLV infection caused by geographically distant viral strains in cattle that express different BoLA class II DRB3 haplotypes.
Methods
Ethics declaration
The study was approved by the Veterinary Sciences Animal Care Committee No. AC21-0210, Canada; the Institutional Animal Care and Use Committee No. PROTO202000096 from 4/13/2020 to 4/14/2023, Michigan State University, United States; the Ethics Review Board, COMSATS Institute of Information Technology, Islamabad, Pakistan, no. CIIT/Bio/ERB/17/26 and the Bioethics Commission No. 06-18 on 30 January 2018, Almaty, Kazakhstan. Blood samples from Polish and Moldovan cattle, naturally infected with BLV, were selected from collections at local diagnostic laboratories as part of the Enzootic bovine leukosis (EBL) monitoring program between 2012 and 2018 and sent to the National Veterinary Research Institute (NVRI) in Pulawy for confirmation study. The approval for collection of these samples from ethics committee was not required according to Polish regulation (“Act on the Protection of Animals Used for Scientific or Educational Purposes”, Journal of Laws of 2015).
Sample collection and preparation
A total of 125 DNA samples obtained from blood of naturally BLV-infected cattle from Canada, United States, Poland, Moldova, Pakistan and Kazakhstan were used for this study. Seventy-six of them were archival DNA samples obtained between 2013 and 2018 as described in our previous studies on samples from Poland (n = 22)64,65, Moldova (n = 14)66, Pakistan (n = 20)67 and Kazakhstan (n = 21)68. Between 2020 and 2021 48 peripheral blood and serum samples from naturally BLV-infected cattle were obtained from three dairy farms of Alberta, Canada and two dairy farms of Michigan, US (see Table 1). All cattle were positive for anti-BLV antibodies, as determined by commercially available ELISA kit (IDEXX Leukosis Serum X2 Ab Test, IDEXX). Genomic DNA were isolated using a Quick DNA Miniprep Plus kit (Zymo Research) and a DNeasy Blood & Tissue Kit (Qiagen) for Canadian (n = 24) and US (n = 24) whole blood samples, respectively, following the manufacturer’s protocol.
PCR amplification of BoLA-DRB3 exon 2 and sequencing
A 247 bp fragment of BoLA-DRB3 exon 2 containing the hypervariable domain was amplified on all DNA samples (n = 125) by a PCR as described in a previous study69. Reactions were carried out in 30 µl final volume containing 1 × Pol Buffer B, 2.5 mM MgCl2, 115 µM of each dNTP, 0.3 μM of each primer, 0.4 U of OptiTaq DNA Polymerase and 12 ng genomic DNA. The thermal cycling protocol was initial denaturation at 94°C for 3 min; 35 cycles at 94°C for 20 s, 60°C for 20 s, and at 72°C for 1 min, followed by final extension at 72°C for 5 min. Reactions were carried out in a TAdvanced Twin PCR Thermal Cycler (Biometra). PCR products were clean-ed up using ExoSAP-IT (Applied Biosystems) by incubation at 37°C for 15 min, followed by ExoSAP-IT inactivation by heating to 80°C for 15 min. The DNA was directly sequenced (Genomed SA Company) with BoLA-DRB3 exon 2 forward and reverse primers HL030 (5′-AGATCTATCCTCTCTCTGCAGCACATTTCC-3′) and HL031 (5′-CGCGCTCACCTCGCCGCT-3′) respectively69.
Sequence-based typing of BoLA-DRB3 alleles
The raw sequences of the 247 bp fragment of BoLA-DRB3 gene were visualized and aligned in Geneious Prime software. Consensus sequences were generated, in which the heterozygous positions were assigned ambiguity codes according to the IUPAC coding system (Supplementary file 1). The consensus sequences were initially compared to the 389 DRB3 allele sequences deposited in the IPD-MHC database (available via web https://www.ebi.ac.uk/ipd/) using Haplofinder script (http://bioinformatics.roslin.ed.ac.uk/haplofinder/haplofinder.py) and Python 2.7.18 software (https://www.python.org)70. To confirm that the assigned alleles did indeed match the pairwise sequence combination, a custom BLAST database was created in Geneious Prime and the data from IPD-MHC were implemented and the query centric alignment was generated using Megablast configuration for highly similar sequences and Low Complexity Filter with Scoring: 1–2 and max E-value of < 0.05.
Amplification and sequencing of gag gene
The full-length 1353 bp BLV gag gene was amplified from DNA samples (n = 125) by nested PCR using oligonucleotide primers as previously reported71 (Supplementary Fig. S5). Both rounds of amplification was performed using PrimeSTAR GXL DNA Polymerase (Takara Bio). Thermal cycling parameters were as follows: initial denaturation at 98 °C for 2 min followed by 38 cycles (36 cycles for the second round) of denaturation at 98 °C for 15 s, annealing at 60 °C for 15 s, extension at 68 °C for 1 min 50 s (1 min 30 s for the second round) and final extension at 72 °C for 5 min. PCR products were separated by electrophoresis on 1.5% agarose gel containing SimplySafe (EURx) and purified using a NucleoSpin Extract II Kit (Marcherey Nagel GmbH & Co KG). Sequencing was performed by Genomed SA Company (Warsaw, Poland). Each sequencing reaction was carried out using: 3 µl BigDye™ Terminator v3.1 Ready Reaction Mix, 1 µl BigDye™ Terminator v1.1 & v3.1 5× Sequencing Buffer, 5 pmol of the primer and 150 ng of DNA were mixed in a final 10 µl volume.
Cycle sequencing was performed in 100 µl PCR tubes as follows: incubation at 96 °C for 1 min as the initial denaturation step followed by 25 cycles of 96 °C for 10 s, 54 °C for 5 s, and 60 °C for 4 min according to standard protocol routinely used by the Genomed SA Company. Prior to purification, the reaction mix was incubated for 10 min at 4 °C. Purified reaction products were separated by electrophoresis on the 3730 xl DNA Analyzer (Thermo Fisher) according to the manufacturer’s instructions. The following pairs of primers were used to direct sequencing as shown in Supplementary Fig. S5.
Analysis of genetic variation among the gag gene sequences
The raw sequence reads in both directions were proofread and analyzed in Geneious Prime 2021.0.1 (Biomatters Ltd). The consensus sequences were determined and deposited in the GenBank database under accession numbers OP146492-OP146601 (Table 1 and Supplementary Fig. S2). The amino acid sequences were translated according to the IUPAC amino acid code and aligned using the Clustal Omega 1.2.2 algorithm. The substitution analysis and pairwise genetic distance assessment were performed in Geneious Prime. The Shannon’s entropy (a quantitative measure of diversity in the alignment, where H = 0 indicates complete conservation) was estimated using BioEdit v. 7.2.5 software72 for each position in the amino-acid alignment, and the entropy values were summed for each Gag domain. The DnaSP v. 6.12.03 program was used to quantify strength of selection by comparing synonymous substitution rates (dS) with nonsynonymous substitution rates (dN)73. Ratio of dN/dS < 1 was interpreted as negative or purifying selection and > 1 was interpreted as positive selection pressure.
Prediction of BoLA-DRB3-presented epitopes
A BoLA-DRB- peptide binding affinity prediction method, NetBoLAIIpan—1.0 was used to predict the Gag protein peptides presented by the BoLA-DRB3 molecule. The consensus Gag protein sequence calculated for the 125 sequences alignment was submitted to the server in FASTA format. For this analysis, a comprehensive list of BoLA molecules available in the server for prediction was updated with new additional sequences deposited in the IPD-MHC database. The prediction values represented by likelihood for BoLA antigen presentation and %Rank score were calculated. In detail, the percentile rank for each Gag peptide was generated by comparing its score against the scores of 100,000 random natural peptides74. For example, if a peptide was assigned a rank of 1%, it meant that its predicted affinity was among the top 1% scores for the specified molecule. The %Rank score of < 1.0, ≥ 1 to < 5.0 and ≥ 5.0 were interpreted as strong binders, weak binders, and non-binders, respectively. All %Rank score predictions ≤ 5.0 were considered as epitope peptides.
Proviral load quantification
The qPCR assays for the BLV pol gene and H3F3A gene were performed according to previously published methods75,76. Briefly, genomic DNA was amplified using primers and probes for pol gene and H3F3A gene, and QuantiTect Multiplex PCR NoROX master mix (Qiagen AG GmbH) according to the protocol: 95 °C for 15 min, followed by 45 cycles each of 94 °C for 60 s and 60 °C for 60 s. Ten-fold dilutions of the pBLV1 and pH3F3A plasmids from 1 × 106 copies/μl to 100 copies/μl were used as the standard to estimate BLV copy numbers. The BLV proviral load (copies/1000 cells) was calculated as [copies of BLV pol gene/(copies of H3F3A gene/2)] × 1000 cells75.
Statistical analysis
The correlation between number of CD4 + T-cell epitopes and proviral load in BLV-infected cattle was calculated using the Spearman non-parametric test (with P value < 0.05). The difference in proviral DNA copy number between two groups of samples with different BoLA-DRB3 genotypes was calculated using the Student’s t-test, where a P value of < 0.05 was considered to be significant. The statistical analysis was performed using STATISTICA ver. 10 (StatSoft).
Modeling of peptides
A PDB file of a putative BLV Gag structure77 was used to identify the location of the epitopes. The structure was opened in UCSF Chimera78 and the surface representation was used to display epitopes by color.
Data availability
The gag gene sequences generated and analyzed during the current study are available in the GenBank repository under accession numbers OP146492-OP146601. The raw sequences of the fragment of BoLA-DRB3 gene with assigned ambiguity codes, according to the IUPAC coding system, in the heterozygous positions generated and analyzed in this study are available in Supplementary file 1.
References
Schwartz, I. & Levy, D. Pathobiology of bovine leukemia virus. Vet. Res. 25, 521–536 (1994).
Bartlett, P. C. et al. Current developments in the epidemiology and control of enzootic bovine leukosis as caused by bovine leukemia virus. Pathogens 9, 1058 (2020).
Bartlett, P. C. et al. Bovine leukemia virus and cow longevity in Michigan dairy herds. J. Dairy Sci. 96, 1591–1597 (2013).
Nekouei, O., VanLeeuwen, J., Stryhn, H., Kelton, D. & Keefe, G. Lifetime effects of infection with bovine leukemia virus on longevity and milk production of dairy cows. Prev. Vet. Med. 133, 1–9 (2016).
Blagitz, M. G. et al. Immunological implications of bovine leukemia virus infection. Res. Vet. Sci. 114, 109–116 (2017).
Gillet, N. et al. Mechanisms of leukemogenesis induced by bovine leukemia virus: Prospects for novel anti-retroviral therapies in human. Retrovirology 4, 18 (2007).
Kakker, N. K., Mikhailov, M. V., Nermut, M. V., Burny, A. & Roy, P. Bovine leukemia virus Gag particle assembly in insect cells: Formation of chimeric particles by domain-switched leukemia/lentivirus Gag polyprotein. Virology 265, 308–318 (1999).
Franke, E. K., Yuan, H. E., Bossolt, K. L., Goff, S. P. & Luban, J. Specificity and sequence requirements for interactions between various retroviral Gag proteins. J. Virol. 68, 5300–5305 (1994).
Olivero-Deibe, N. et al. Expression, purification, and characterization of bovine leukemia virus-like particles produced in drosophila S2 cells. Front. Virol. 1, 151 (2021).
Frie, M. C. & Coussens, P. M. Bovine leukemia virus: A major silent threat to proper immune responses in cattle. Vet. Immunol. Immunopathol. 163, 103–114 (2015).
Florins, A. et al. Viral expression directs the fate of B cells in bovine leukemia virus-infected sheep. J. Virol. 86, 621–624 (2012).
Florins, A. et al. Cell dynamics and immune response to BLV infection: A unifying model. Front. Biosci. 12, 1520–1531 (2007).
Mager, A., Masengo, R., Mammerickx, M. & Letesson, J. J. T cell proliferative response to bovine leukaemia virus (BLV): Identification of T cell epitopes on the major core protein (p24) in BLV-infected cattle with normal haematological values. J. Gen. Virol. 75(Pt 9), 2223–2231 (1994).
Bai, L., Takeshima, S. N., Isogai, E., Kohara, J. & Aida, Y. Novel CD8(+) cytotoxic T cell epitopes in bovine leukemia virus with cattle. Vaccine 33, 7194–7202 (2015).
Orlik, O. & Splitter, G. A. Progression to persistent lymphocytosis and tumor development in bovine leukemia virus (BLV)-infected cattle correlates with impaired proliferation of CD4+ T cells in response to gag- and env-encoded BLV proteins. J. Virol. 70, 7584–7593 (1996).
Stone, D. M., Norton, L. K., Chambers, J. C. & Meek, W. J. CD4 T lymphocyte activation in BLV-induced persistent B lymphocytosis in cattle. Clin. Immunol. 96, 280–288 (2000).
Callebaut, I. et al. Mapping of B-neutralizing and T-helper cell epitopes on the bovine leukemia virus external glycoprotein gp51. J. Virol. 67, 5321–5327 (1993).
Gatei, M. H., Good, M. F., Daniel, R. C. & Lavin, M. F. T-cell responses to highly conserved CD4 and CD8 epitopes on the outer membrane protein of bovine leukemia virus: Relevance to vaccine development. J. Virol. 67, 1796–1802 (1993).
Bai, L. et al. Mapping of CD4(+) T-cell epitopes in bovine leukemia virus from five cattle with differential susceptibilities to bovine leukemia virus disease progression. Virol. J. 16, 157 (2019).
Sakakibara, N., Kabeya, H., Ohashi, K., Sugimoto, C. & Onuma, M. Epitope mapping of bovine leukemia virus transactivator protein Tax. J. Vet. Med. Sci. 60, 599–605 (1998).
Nagaoka, Y. et al. Ovine MHC class II DRB1 alleles associated with resistance or susceptibility to development of bovine leukemia virus-induced ovine lymphoma. Cancer Res. 59, 975–981 (1999).
Xu, A., van Eijk, M. J., Park, C. & Lewin, H. A. Polymorphism in BoLA-DRB3 exon 2 correlates with resistance to persistent lymphocytosis caused by bovine leukemia virus. J. Immunol. 151, 6977–6985 (1993).
Takeshima, S. N. & Aida, Y. Structure, function and disease susceptibility of the bovine major histocompatibility complex. Anim. Sci. J. 77, 138–150 (2006).
Norimine, J. & Brown, W. C. Intrahaplotype and interhaplotype pairing of bovine leukocyte antigen DQA and DQB molecules generate functional DQ molecules important for priming CD4+ T-lymphocyte responses. Immunogenetics 57, 750–762 (2005).
Baumgartner, C. K., Ferrante, A., Nagaoka, M., Gorski, J. & Malherbe, L. P. Peptide-MHC class II complex stability governs CD4 T cell clonal selection. J. Immunol. 184, 573–581 (2010).
Kedzierska, K. & Koutsakos, M. The ABC of major histocompatibility complexes and T cell receptors in health and disease. Viral Immunol. 33, 160–178 (2020).
Aida, Y. et al. A novel bovine leukemia virus peptide vaccine targeting susceptible cattle-Production by 3-D modelling and nanotechnology. Retrovirology 12, P48 (2015).
Connelley, T. et al. Immunopeptidomic analysis of BoLA-I and BoLA-DR Presented peptides from theileria parva infected cells. Vaccines Basel 10, 1907 (2022).
Nakatsuchi, A. et al. BoLA-DRB3 Polymorphism Controls Proviral Load and Infectivity of Bovine Leukemia Virus (BLV) in Milk. Pathogens 11, 210 (2022).
Takeshima, S. N., Ohno, A. & Aida, Y. Bovine leukemia virus proviral load is more strongly associated with bovine major histocompatibility complex class II DRB3 polymorphism than with DQA1 polymorphism in Holstein cow in Japan. Retrovirology 16, 14 (2019).
Lo, C. W. et al. BoLA-DRB3 polymorphism is associated with differential susceptibility to bovine leukemia virus-induced lymphoma and proviral load. Viruses 12, 352 (2020).
Notsu, K. et al. Identifying pathogen and allele type simultaneously in a single well using droplet digital PCR. mSphere 8, e00493-e1422 (2023).
Nakatsuchi, A., Matsumoto, Y. & Aida, Y. Influence of BoLA-DRB3 polymorphism and bovine leukemia virus (BLV) infection on dairy cattle productivity. Vet. Sci. 10, 250 (2023).
Samad, A., Meghla, N. S., Nain, Z., Karpinski, T. M. & Rahman, M. S. Immune epitopes identification and designing of a multi-epitope vaccine against bovine leukemia virus: A molecular dynamics and immune simulation approaches. Cancer Immunol. Immunother. 71, 2535–2548 (2022).
Lin, H. H., Zhang, G. L., Tongchusak, S., Reinherz, E. L. & Brusic, V. Evaluation of MHC-II peptide binding prediction servers: Applications for vaccine research. BMC Bioinf. 9, S22 (2008).
Rolland, M. et al. Genetic impact of vaccination on breakthrough HIV-1 sequences from the STEP trial. Nat. Med. 17, 366–371 (2011).
Fischer, W. et al. Polyvalent vaccines for optimal coverage of potential T-cell epitopes in global HIV-1 variants. Nat. Med. 13, 100–106 (2007).
Hooshmand, N., Fayazi, J., Tabatabaei, S. & Ghaleh-Golab-Behbahan, N. Prediction of B cell and T-helper cell epitopes candidates of bovine leukaemia virus (BLV) by in silico approach. Vet. Med. Sci. 6, 730–739 (2020).
Fisch, A. et al. Integral use of immunopeptidomics and immunoinformatics for the characterization of antigen presentation and rational identification of BoLA-DR-presented peptides and epitopes. BioRxiv 45, 47 (2020).
Reynisson, B. et al. Improved prediction of MHC II antigen presentation through integration and motif deconvolution of mass spectrometry MHC eluted ligand data. BioRxiv 166, 799882 (2020).
Sun, M., Grigsby, I. F., Gorelick, R. J., Mansky, L. M. & Musier-Forsyth, K. Retrovirus-specific differences in matrix and nucleocapsid protein-nucleic acid interactions: Implications for genomic RNA packaging. J. Virol. 88, 1271–1280 (2014).
Rayne, F., Kajava, A. V., Lalanne, J. & Mamoun, R. Z. In vivo homodimerisation of HTLV-1 Gag and MA gives clues to the retroviral capsid and TM envelope protein arrangement. J. Mol. Biol. 343, 903–916 (2004).
Gamble, T. R. et al. Structure of the carboxyl-terminal dimerization domain of the HIV-1 capsid protein. Science 278, 849–853 (1997).
Ohno, A., Takeshima, S. N., Matsumoto, Y. & Aida, Y. Risk factors associated with increased bovine leukemia virus proviral load in infected cattle in Japan from 2012 to 2014. Virus Res. 210, 283–290 (2015).
Lohr, C. E. et al. Phenotypic selection of dairy cattle infected with bovine leukemia virus demonstrates immunogenetic resilience through NGS-based genotyping of BoLA MHC Class II Genes. Pathogens 11, 104 (2022).
Juliarena, M. A., Barrios, C. N., Lützelschwab, C. M., Esteban, E. N. & Gutierrez, S. E. Bovine leukemia virus: Current perspectives. Virus Adapt. Treatment 13, 26 (2017).
Ramduth, D. et al. Immunodominant HIV-1 Cd4+ T cell epitopes in chronic untreated clade C HIV-1 infection. PLoS One 4, e5013 (2009).
Huang, S. et al. Deficiency of HIV-Gag-specific T cells in early childhood correlates with poor viral containment. J. Immunol. 181, 8103–8111 (2008).
Prendergast, A. et al. Gag-specific CD4+ T-cell responses are associated with virological control of paediatric HIV-1 infection. AIDS 25, 1329–1331 (2011).
Ranasinghe, S. et al. HIV-specific CD4 T cell responses to different viral proteins have discordant associations with viral load and clinical outcome. J. Virol. 86, 277–283 (2012).
Buggert, M. et al. Characterization of HIV-specific CD4+ T cell responses against peptides selected with broad population and pathogen coverage. PLoS One 7, e39874 (2012).
Hayashi, T. et al. Cattle with the BoLA class II DRB3*0902 allele have significantly lower bovine leukemia proviral loads. J. Vet. Med. Sci. 79, 1552–1555 (2017).
LaHuis, C. H. et al. Identification of BoLA alleles associated with BLV proviral load in US beef cows. Pathogens 11, 1093 (2022).
Juliarena, M. A. et al. Association of BLV infection profiles with alleles of the BoLA-DRB32 gene. Anim. Genet. 39, 432–438 (2008).
Le, T. D. et al. Association between BoLA-DRB3 polymorphism and bovine leukemia virus proviral load in Vietnamese Holstein Friesian cattle. Hla 99, 105–112 (2022).
Sahay, B., Nguyen, C. Q. & Yamamoto, J. K. Conserved HIV epitopes for an effective HIV vaccine. J. Clin. Cell Immunol. 8, 4 (2017).
Wang, Y. E. et al. Protective HLA class I alleles that restrict acute-phase CD8+ T-cell responses are associated with viral escape mutations located in highly conserved regions of human immunodeficiency virus type 1. J. Virol. 83, 1845–1855 (2009).
Li, G. et al. An integrated map of HIV genome-wide variation from a population perspective. Retrovirology 12, 18 (2015).
Barra, C. et al. Footprints of antigen processing boost MHC class II natural ligand predictions. Genome Med. 10, 84 (2018).
Pyasi, S., Sharma, V., Dipti, K., Jonniya, N. A. & Nayak, D. Immunoinformatics approach to design multi-epitope- subunit vaccine against bovine ephemeral fever disease. Vaccines Basel 9, 925 (2021).
Shrestha, S. et al. Effects of bovine leukemia virus seropositivity and proviral load on milk, fat, and protein production of dairy cows. J. Dairy Sci. 2023, 859 (2023).
Borjigin, L. et al. Effectiveness of integrated bovine leukemia virus eradication strategies utilizing cattle carrying resistant and susceptible histocompatibility complex class II DRB3 alleles. J. Dairy Sci. 2023, 14 (2023).
Nakatsuchi, A. et al. Anti-BLV antibodies in whey correlate with bovine leukemia virus disease progression and BoLA-DRB3 polymorphism. Front. Vet. Sci. 9, 1038101 (2022).
Pluta, A., Rola-Łuszczak, M., Douville, R. N. & Kuźmak, J. Bovine leukemia virus long terminal repeat variability: Identification of single nucleotide polymorphisms in regulatory sequences. Virol. J. 15, 165 (2018).
Pluta, A., Willems, L., Douville, R. N. & Kuźmak, J. Effects of naturally occurring mutations in bovine leukemia virus 5’-LTR and tax gene on viral transcriptional activity. Pathogens 9, 836 (2020).
Pluta, A. et al. Molecular characterization of bovine leukemia virus from Moldovan dairy cattle. Arch. Virol. 162, 1563–1576 (2017).
Rola-Łuszczak, M. et al. Molecular characterization of the env gene of bovine leukemia virus in cattle from Pakistan with NGS-based evidence of virus heterogeneity. Pathogens 10, 910 (2021).
Sultanov, A. et al. Molecular characterization of bovine leukemia virus with the evidence of a new genotype circulating in cattle from Kazakhstan. Pathogens 11, 180 (2022).
Miltiadou, D., Law, A. S. & Russell, G. C. Establishment of a sequence-based typing system for BoLA-DRB3 exon 2. Tissue Antigens 62, 55–65 (2003).
Baxter, R., Hastings, N., Law, A. & Glass, E. J. A rapid and robust sequence-based genotyping method for BoLA-DRB3 alleles in large numbers of heterozygous cattle. Anim. Genet. 39, 561–563 (2008).
Suzuki, A. et al. Phylogenetic analysis of South African bovine leukaemia virus (BLV) isolates. Viruses 12, 14 (2020).
Hall, T. A. BioEdit A a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98 (1999).
Rozas, J. et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 34, 3299–3302 (2017).
Fisch, A. et al. Integral use of immunopeptidomics and immunoinformatics for the characterization of antigen presentation and rational identification of BoLA-DR-presented peptides and epitopes. J. Immunol. 206, 2489–2497 (2021).
Pluta, A. et al. Analysis of nucleotide sequence of Tax, miRNA and LTR of bovine leukemia virus in cattle with different levels of persistent lymphocytosis in Russia. Pathogens 10, 246 (2021).
Rola-Łuszczak, M., Finnegan, C., Olech, M., Choudhury, B. & Kuźmak, J. Development of an improved real time PCR for the detection of bovine leukaemia provirus nucleic acid and its use in the clarification of inconclusive serological test results. J. Virol. Methods 189, 258–264 (2013).
Qualley, D. F. et al. Solution conformation of bovine leukemia virus gag suggests an elongated structure. J. Mol. Biol. 431, 1203–1216 (2019).
Pettersen, E. F. et al. UCSF Chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
T.K. and A.P. thank Lauren Eichberger, Madison Wieland, Lauren St. Andrew and Nancy Raney for DNA extractions (Michigan State University, USA). Acknowledgement of farms includes Black Locust Farms, LLC and Michigan State University Dairy Cattle Teaching and Research Center (Michigan State University, USA). A.P. thanks Morten Nielsen (Technical University of Denmark, Denmark) for the NetBoLAIIpan—1.0 server lessons.
Author information
Authors and Affiliations
Contributions
Proposed the conception and design of the study, A.P.; data curation, A.P., T.M.T., S.S, M.R.-Ł, A.R., A.S. and S.M.; investigation, A.P., T.M.T., S.S., M.R.-Ł and A.R., formal analysis, A.P.; database analysis, A.P., visualization of the results, A.P.; visualization – modeling, D.Q.; resources, A.S., S.M. and J.K; writing—original draft preparation, A.P.; writing—review and editing, A.P., T.M.T., F.M, D.Q., S.S, P.C. and J.K; project administration, A.P. All authors read and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pluta, A., Taxis, T.M., van der Meer, F. et al. An immunoinformatics study reveals a new BoLA-DR-restricted CD4+ T cell epitopes on the Gag protein of bovine leukemia virus. Sci Rep 13, 22356 (2023). https://doi.org/10.1038/s41598-023-48899-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-48899-4
- Springer Nature Limited