Abstract
In silico interrogation of glioblastoma (GBM) in The Cancer Genome Atlas (TCGA) revealed upregulation of GNA12 (Gα12), encoding the alpha subunit of the heterotrimeric G-protein G12, concomitant with overexpression of multiple G-protein coupled receptors (GPCRs) that signal through Gα12. Glioma stem cell lines from patient-derived xenografts also showed elevated levels of Gα12. Knockdown (KD) of Gα12 was carried out in two different human GBM stem cell (GSC) lines. Tumors generated in vivo by orthotopic injection of Gα12KD GSC cells showed reduced invasiveness, without apparent changes in tumor size or survival relative to control GSC tumor-bearing mice. Transcriptional profiling of GSC-23 cell tumors revealed significant differences between WT and Gα12KD tumors including reduced expression of genes associated with the extracellular matrix, as well as decreased expression of stem cell genes and increased expression of several proneural genes. Thrombospondin-1 (THBS1), one of the genes most repressed by Gα12 knockdown, was shown to be required for Gα12-mediated cell migration in vitro and for in vivo tumor invasion. Chemogenetic activation of GSC-23 cells harboring a Gα12-coupled DREADD also increased THBS1 expression and in vitro invasion. Collectively, our findings implicate Gα12 signaling in regulation of transcriptional reprogramming that promotes invasiveness, highlighting this as a potential signaling node for therapeutic intervention.
Similar content being viewed by others
Introduction
G protein coupled receptors (GPCRs) transduce their signals through coupling to heterotrimeric G-proteins. Upon stimulation of GPCRs by binding of their cognate ligands, the G-protein alpha subunit (Gα) exchanges GDP for GTP and in this activated state can bind to and regulate its downstream effectors. The effectors of the earliest discovered G-proteins of the Gs, Gi, and Gq families are the enzymes adenylate cyclase and phospholipase C. The last discovered G-protein family (G12/13), encoded by GNA12 and GNA13, do not couple to these enzymes. Instead, the effectors for Gα12/Gα13 are GTP exchange factors (GEFs) for the low molecular weight GTPase RhoA1,2. Activation of Gα12 increases signaling to RhoA which elicits cytoskeletal responses3,4,5. More recently established mediators of RhoA signaling are the transcriptional co-activators YAP and MRTF-A6,7, which we previously linked to GSC and GBM tumor growth8.
The GPCRs that are most prominently coupled to G12/13 and RhoA signaling include those for thrombin and the lysophospholipids LPA and S1P9,10. Our previous work and that of others demonstrated that activated GPCRs that couple to Gα12 and RhoA are efficacious mitogens and transcriptional activators11,12,13. These receptors are highly expressed in the brain, activated by ligands that are generated in response to inflammation and have been implicated in tumor growth and progression14,15,16. Levels of mRNA for GPCRs, such as PAR1 (the receptor for thrombin) and S1P2 and S1P3 (receptors for sphingosine 1-phosphate), are elevated in GBM14,17,18. In addition to enhanced expression of a wide range of GPCRs and increased availability of their ligands, Gα12 mRNA is particularly elevated in higher grades of glioma8. While multiple distinct Gα12-coupled GPCRs could regulate GBM progression in vivo, these receptors all ultimately converge on Gα12 to transduce their signals. Accordingly, we reasoned that knocking down Gα12, the nodal mediator of signaling through these GPCRs, would demonstrate the importance of this pathway and provide clues as to what signals contribute to its role in aggressiveness and progression of GBM tumors.
Here we examine in vitro and in vivo the properties of GSCs in which Gα12 expression is knocked down with shRNA. Loss of the Gα12 pathway diminished invasion of GBM tumor cells in vivo, in association with altered extracellular matrix, proneural and stem cell gene expression. Amongst the ECM components, thrombospondin-1 (THBS1), previously associated with tumor progression, was one of the most significantly downregulated in cells and tumors lacking Gα12. Our study highlights the importance of Gα12 as an integrator of signals from GPCRs to transcriptional responses that alter glioma cell phenotype and tumor invasion.
Materials and methods
Glioblastoma stem Cells
GSC23 and HK281 were provided by Dr. Frederick Lang, MD Anderson and Dr. Harley Kornblum, UCLA, respectively. GSC neurospheres were cultured in suspension flasks containing DMEM/F12 medium supplemented with B27 (Gibco) in a 5% CO2 37 °C incubator. Lentivirus infection to knock down GNA12 or THBS1 or express Gα12 DREADD in GSC cells was followed by 48–96-h puromycin selection.
Extreme limiting dilution assay
Neurospheres dissociated into cell suspension by Accutase treatment were counted and plated in fresh media in pentaplicate into 96-wells at 1–100 cells/well. After 21 days spheres were counted to estimate stem cell frequency by ELDA (http://bioinf.wehi.edu.au/software/elda/) using χ2 to determine pair-wise differences.
Orthotopic GSC injections
1.5 or 5 × 105 control or knocked down GSC23 cells tagged with near infrared IRFP720 were intracranially injected into the mouse brain (6 mice per group), using a stereotactic system as previously described8. Survival experiments were performed twice. Tumor size was estimated by fluorescence emission detection by FMT 2500 Fluorescence Tomography (Perkin Elmer) at 720 nm. The onset of neurologic sequelae in the control group was used to determine time of euthanasia. Mice were euthanized by CO2 inhalation in accordance with our institutional guidelines for animal welfare and experimental conduct at University of California at San Diego. Brain samples were collected, and tissue samples were processed for histological examination by H&E and anti-human nuclei IHC at UCSD CALM and MCC Biorepository and Tissue Technology Core.
RNA analysis
Total RNA was isolated using Trizol reagent according to the manufacturer’s protocol, followed by RT-qPCR for relative quantification. RNA sequencing of tumors from mice injected with Gα12 KD or control GSC23 cells (4 mice tumors for biological replicates in each group) were submitted to RNA integrity analysis (Agilent Bioanalyzer, Tapestation results eRIN > 8.5), ribodepleted library preparation, and sequencing using Illumina NovaSeq 6000 (run set PE100 and 25 M reads). Gene-set enrichment analysis was performed using GSEA software.
Migration/invasion assay
Uncoated or Geltrex-coated membranes of Transwell 24-well plates (8 µm pore size, Corning, Cat#3422) were used to assess migration and invasion, respectively, as detailed in Supplementary Material.
Statistics
Statistical differences were analyzed using Graphpad Prism software version 8. Analysis of variance (ANOVA) followed by Tukey’s multiple test was applied for groups with several features. One-way ANOVA was used to analyze data from experiments with one independent variable, and two-way ANOVA for two independent factors. Data are presented as mean ± SEM and significances based on calculated probability values (*p < 0.05; **p < 0.01).
Ethical approval
In vivo experiments were executed under approval of animal protocol by University of California San Diego Institutional Animal Care and Use Committee (IACUC) Office #S00192M, which were performed following the protocols in complying with federal regulations by USDA, APHIS, CFR, Title 9, Parts 1, 2, and 3. The study was reported in accordance with the recommendations of the ARRIVE guidelines.
Results
GBMs overexpress GNA12 and G⍺12—coupled GPCRs
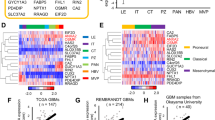
We interrogated the TCGA database generated from patient GBM surgical specimens and determined that GNA12 mRNA expression was elevated in 30% of 160 GBM patient samples profiled in TCGA, while its homolog, G⍺13, was far less frequently overexpressed (Fig. 1A). We also interrogated the TCGA for a series of GPCRs recently established to couple efficiently to G⍺12 (Supplemental Fig. 1). GPCRs that couple to G⍺12 and are altered in ≥ 5% of patients are shown in Fig. 1A. Notably, most GPCRs were overexpressed in fewer patients and in subsets of patients distinct from those with elevated GNA12. Overall expression of GNA12 mRNA in GBM based on TCGA analysis by Gliovis was 2.2-fold higher than in normal brain (Fig. 1B).
G⍺12 is upregulated in high grade glioblastoma tumor samples and GSC cell lines. (A) cBio oncoprint from TCGA PanCancer GBM study. Genomic alterations in GNA12 and predicted G⍺12-coupled GPCRs. Z-scores relative to diploid samples RNA-Seq V2 RSEM: ± 1.5 threshold. (B) G⍺12 upregulation in GBM. RNA-Seq sample counts from Gliovis, p < 0.001. (C) Heatmaps of RNA-seq, whole exome, and additional clinical phenotype data aggregated from TCGA. (D) G⍺12 mRNA expression relative to neural stem cells in GSC cell lines.
We examined expression of GNA12 in relation to the molecular classification and phenotypic characteristics of glioma samples, including consideration of their IDH1/p53/PTEN mutational status, tumor grade, patient age, and survival. GBMs have been also classified by transcriptional signatures into proneural, classical, and mesenchymal subtypes19. GNA12, but not GNA13, was highly expressed in the classical and mesenchymal GBM subtypes and enriched in elderly patients and those with worst performance status (Fig. 1C). Expression clustered with YAP1, a downstream transcriptional co-activator regulated through G⍺12-RhoA signaling7,20,21.
RNA-seq data from 40 patient-derived GSCs generated by the Rich laboratory and compiled and stored in a data base described previously22,23 was also analyzed; all GSCs were found to have levels of GNA12 expression at least one SD above that of neural stem cells (Fig. 1D). The GSC23 cell line, established from a patient-derived xenograft of a recurrent and aggressive tumor24, and also used in our previous work8, was intermediate in its expression of GNA12, providing a representative model to examine the role of G⍺12 signaling in GBM growth.
GSCs were transduced by lentiviral-directed short hairpin RNAs (shRNAs) encoding either a control sequence not found in the mammalian genome or one of two non-overlapping G⍺12 sequences. G⍺12 mRNA levels were reduced by over 75% relative to control cells without significant compensatory changes in G⍺13 (Supplemental Fig. 2). Westerns on whole cell lysates also demonstrated an approximately 50% decrease in G⍺12 protein in the knockdowns compared to control cells (Supplemental Fig. 2).
G⍺12 knockdown does not alter mouse survival or tumor size observed following orthotopic intracranial injection of GSC tumor cells. shRNA control or shG⍺12 knockdown GSC23 and HK281 cells labeled with IRFP720 were intracranially injected into syngeneic nu/nu mice. (A, B) Kaplan–Meier survival curves for GSC23 and HK281 control and shG⍺12 KD tumor-bearing mice (6 animals per group, in 2 independent experiments for GSC23, and 4 animals per group for HK281 in one experiment). Survival curves were not significantly different as determined using the Log rank test (C, D) Brain tumor growth of GSC23-engrafted mice as monitored using Fluorescence Molecular Tomography (FMT) emission at 720 nm. Representative FMT scan images at 17 days post intracranial injection and relative fluorescence quantification. (E) Additional set of animals injected with approximately 3 × 105 GSC23 cells and analyzed for tumor growth monitored by FMT did not show significant differences in tumor size (p = 0.1; n = 6).
Depletion of G⍺12 does not affect GBM lethality or tumor size
To determine whether G⍺12 protein signals are critical for tumorigenesis in the brain microenvironment we intracranially implanted mice with GSC23 control or G⍺12 shRNA transduced cells labeled with IRFP720. The experiment was repeated twice with 6 animals per group in each experiment. Inhibition of G⍺12 did not change overall survival of the tumor-bearing mice over a period of approximately 30 days (Fig. 2A). We used an additional GBM patient derived glioma stem cell, HK281, which our previous work established to show elevated G⍺12 mRNA (to an extent approximately double that of GSC23 cells)8. Knockdown of G⍺12 in HK281 was highly effective and occurred without concomitant changes in G⍺13 expression (Supplemental Fig. 2). Survival of mice implanted with G⍺12 KD HK 281 cells was not altered (Fig. 2B).
Although alterations in overall survival were not observed when G⍺12 expression was inhibited, we carried out additional analysis on GSC-23 cell implanted mice. Tumor size was assessed using fluorescent molecular tomography (FMT) of IRFP720-expressing WT and G⍺12 knockdown tumors as shown in Fig. 2C. Tumor size, assessed longitudinally, did not differ significantly between the mice bearing control and G⍺12 knockdown cells (Fig. 2D). In an additional series of orthotopic injections, we engrafted approximately half the number of GSC23 cells to minimize potential deleterious effects of massive tumor development and associated lethality. Here, again, there was no difference in the survival of the two groups of tumor-bearing mice followed until the time of sacrifice, and serial FMT imaging of tumor-bearing brains revealed insignificant fluorescence range distributions between groups (Fig. 2E). We also confirmed that G⍺12 mRNA levels remained downregulated in the KD tumors (Supplemental Fig. 2A,B).
GNA12 is essential for in vivo tumor invasion
Tumor-bearing brains were harvested, sectioned, and stained with hematoxylin and eosin (H&E) or analyzed by immunohistochemistry (IHC) (Fig. 3A–C). GSC23 control cells generated tumors with typical irregular invasive GBM borders (Fig. 3A). In contrast, the tumor mass in mice injected with G⍺12 KD GSC23 cells was largely confined to the injection site and the tumor border areas were clearly defined and compact. Analysis of tissue samples from two additional experiments confirmed that GSC23 control cells developed tumors with irregular borders and finger-like projections into the mice brain (Fig. 3C), while GSC23 cells with G⍺12 knockdown formed tumors with smoother and more defined borders. Quantitative analysis of the shape of the tumor border from sections shown in (Fig. 2A) was carried out using digital pathology QuPath software. While only two representative images were used to provide quantitative data, the multiforme tumor-stroma interfaces were significantly decreased in virtually all G⍺12 KD compared to the control tumors, indicative of diminished invasiveness (Fig. 3B). To further verify these histological observations, we visualized the GSC cells in the tumor by IHC, staining for a human nucleolar antigen protein; this further revealed micrometastasis along the tumor borders in control but not in KD tumors (Fig. 3C). We also demonstrated that G⍺12 knockdown in HK281 cells diminished the invasiveness of tumors formed in vivo (Fig. 3D). These data support the hypothesis that GPCR ligands in the tumor microenvironment utilize G⍺12 to trigger GSC invasion.
Orthotopic xenograft GSC tumors with G⍺12 knockdown exhibit diminished invasiveness. (A, B) Mouse brain cross sections at 10 days post GSC23 intracranial injection stained with H&E (hematoxylin and eosin) (2 per group). (B) Yellow lines define tumor border analyzed by digital pathology detection software QuPath for border area by square pixel of cross-sections (p < 0.05 vs. shControl, 2 per group) (C, D) H&E and corresponding IHC sections marking human cell nuclei (HPM, × 40) in tumors from GSC23 and HK281 tumor-bearing mice.
RNA-seq analysis of differentially expressed genes in G⍺12 knockdown and control GSC23 tumors
To explore molecular pathways downstream of GNA12 we performed a comprehensive analysis of gene expression profiles in tumors derived from control and G⍺12-depleted GSCs. We harvested tumors at 21 days and submitted 8 GSC23 tumor samples (4 WT and 4 G⍺12 knockdown) for RNA sequencing (RNA-seq). Data generated from the RNA-seq analysis identified 22,247 expressed genes with high confidence. Of these 272 genes were upregulated and 558 were downregulated in G⍺12-depleted tumors (p-adjusted < 0.05) as shown in the Heatmap and Volcano plot (Fig. 4A). A more stringent cut-off value of p-adjusted < 0.01 was used to rank the most significantly differentially regulated genes annotated in the Volcano plot. Gene oncology (GO) and Gene Set Enrichment Analysis (GSEA) were used to assess the pathways that were influenced by G⍺12 (Fig. 4B).
Differentially expressed genes in G⍺12 knockdown GSC23 tumor. (A) Heatmap obtained for DESeq2 analysis of GSC23 tumors (shG⍺12#3 or shControl) obtained at 17 days post injection (n = 4, p-adj = 0.05). (B) Volcano plot shows genes down or upregulated in shG⍺12 KD GSC23 tumor samples. (C) Gene Ontology (GO) annotation based on Gene-set enrichment analysis (GSEA) performed to assess biological function and related hallmark pathways for differentially expressed genes in control and G⍺12KD tumors. Those most relevant to tumor phenotype described here are highlighted.
Genes involved in the regulation of ECM components and organization, matrix adhesion and lamellipodia dynamics as well as stem cell properties and epithelial mesenchymal transition were differentially expressed as shown by RNA-Seq (Fig. 4B). The list includes several markers indicative of a decreased mesenchymal phenotype, for example expression of CHI3L1 (encoding YKL-40) was downregulated over fivefold in 3 of 4 G⍺12 KD tumor samples. THBS1, which encodes a matricellular protein and has been proposed as a robust clinical marker of the mesenchymal phenotype and GBM prognosis25,26, was downregulated by 80% in the G⍺12 KD tumors. Cadherin-11, a cell–cell adhesion molecule that is associated with EMT/PMT27, was also significantly downregulated in G⍺12 KD tumors; this gene is also highly expressed in GBM patient samples in TCGA data analysis (Supplemental Fig. 5). The categorization includes many genes that are in several GO categories since the biological process of EMT, stemness and cell migration are interrelated.
G⍺12 knockdown GSCs tumors have altered gene expression profiles compatible with phenotypic changes in GSC cell behavior observed in vitro. (A) mRNA expression of proneural-mesenchymal transition related genes in GSC23 tumor samples (n = 4, *p < 0.05, **p < 0.01 vs. shControl). (B) Cell plasticity of GSC23 grown on Matrigel-coated surfaces for 21 days visualized by light microscopy (20X). (C) mRNA expression of cancer-associated stem cell genes in GSC23 tumor samples (n = 8, *p < 0.05 vs. shControl). The total number of GSC23 spheres formed at 14 days in culture plotted by extreme limiting dilution analysis (ELDA, 0.95 confidence interval). Estimated stem cell frequency per group in the table.
A proneural-to-mesenchymal transition (PMT), similar to EMT, has been described for GBM24,28. To more specifically assess genes associated with the proneural/mesenchymal transition (PMT) characteristic of glioblastoma we looked for changes in known PMT associated genes in the GSC23 tumors by qPCR (Fig. 5A).
We observed increases in well-known proneural genes in G⍺12 KD tumors (Fig. 5A), specifically up-regulation of CD133, OLIG2, and TAZ along with a trend towards an increase in PATZ. The NF1 gene, disputably proneural29, was also increased as was YKL40 which was, however, significantly down-regulated by RNA-Seq. Overall, these data are consistent with deletion of G⍺12 reducing in vivo proneural to mesenchymal transition and leading to attenuated mesenchymal tumor cell properties. We also demonstrated in an in vitro analysis that the typical plasticity of cancer stem cells seen with prolonged culture (21 days) on Matrigel-coated plates was altered by G⍺12 knockdown (Fig. 5B). Control GSC23 cells formed large and stable spheres, with adherent cells migrating out of the spheres, while G⍺12 KD cells formed less stable spheres that yielded loose rounded-shape cells, consistent with an altered mesenchymal/proneural dynamic state.
We also analyzed mRNA levels of core cancer-associated stem cell genes in G⍺12 KD GSC23 (Fig. 5C) and in HK281 tumors (Supplemental Fig. 4B,C). We observed reduced mRNA levels for seven stem cell genes (CCND1, MYC, NANOG, NESTIN, OCT4, PAX6 and SOX2) in the G12 KD GSC23 tumors. To demonstrate that there were functional differences associated with these genetic changes we analyzed GSC self-renewal by in vitro sphere formation comparing control and two lentiviral constructs of G⍺12shRNA knockdown cells (Fig. 5D). Decreasing G⍺12 expression in GSC23 cells lead to diminished stem cell frequency assessed by extreme limiting dilution (ELDA) analysis. GSC23 control cells showed an average of one stem cell for every 35 cells, while the two shRNA knockdown cells averaged one for every ~ 50–80 cells, i.e., the knockdown of G⍺12 protein decreased the ability to generate new spheres by an average of 60% (Fig. 5D). Diminishing G⍺12 mRNA levels in HK281 cells, as in the GSC-23 cells, decreased stem cell properties as assessed by alterations in stem cell gene mRNA levels and growth in the extreme limiting dilution assay (Supplemental Fig. 4).
G⍺12 promotes a mesenchymal-like invasive phenotype through THBS1 signaling
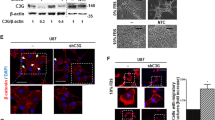
To further investigate the role of G⍺12 in GBM tumor invasion we examined the effect of GNA12 knockdown on GSC migration and invasion in vitro (Fig. 6A,B). We used sphingosine-1-phosphate (S1P) to activate GPCRs coupled to G⍺12 and effected a 2.5-fold increase in cell migration and a fivefold increase in invasion. Migration and invasion were significantly attenuated by G⍺12 KD supporting the role for G⍺12 signaling in GSC23 cell migration and invasion. There was no significant effect of S1P on proliferation of either control or G12 KD cells over this time period nor throughout 5 consecutive days, as assessed by Cyquant analysis (Supplemental Fig. 4D).
Cell migration and invasion of GSC23 cells are dependent on G⍺12 and Thrombospondin-1. Migration (A) or Invasion (B) of control or G⍺12KD GSC23 cells through microwells in response to 16 h treatment with 1.0 µM S1P is mediated through G⍺12 (n = 3, *p < 0.05). (C) Thrombospondin-1 gene expression (THBS1) in GSC23 cells is stimulated by S1P and mediated through G⍺12. (n = 3, *p < 0.05, **p < 0.01 vs. shControl). (D) Migration elicited by S1P is blocked by knockdown of THBS1. (n = 3, #p < 0.05 PBS vs S1P; **p < 0.01 KDs vs. shControl.). (E) G⍺12 DREADD expressing GSC23 cells stimulated by 10um of synthetic ligand CNO express canonical RhoA regulated genes CCN1 and CTGF. (F) THBS1 mRNA expression is induced by 10um CNO in G⍺12 DREADD expressing GSC23 cells (n = 3, *p < 0.05 vs. shControl). (G) Cell migration elicited by G⍺12 DREADD activation of 10um CNO treatment (n = 3, *p < 0.05 vs. PBS).
THBS1 was the one of the most highly down regulated genes in the RNA seq analysis and it has been functionally associated with cell migration, EMT and stemness. We confirmed the decrease in THBS1 expression demonstrated in the RNA-seq data of G⍺12 KD tumors by qPCR on GSC23 tumor samples (Supplemental Fig. 2E). Accordingly, we focused on THBS1 to interrogate the role that downstream transcriptionally regulated targets of G⍺12 play in GSC23 migration. S1P treatment of GSC23 cells increased THBS1 mRNA and this increase was fully abrogated by G⍺12 KD (Fig. 6C). Notably basal levels of THBS1 were also downregulated in the G⍺12-knockdown cells. To investigate the role of THBS1 in GSC23 cell migration we generated THBS1 KD GSC23 cells using lentiviral shRNA (Supplemental Fig. 2E). Migration stimulated by S1P was attenuated by nearly 80% in THBS1-depleted GSC23 cells (Fig. 6D). A gain of function approach was then used to further establish that pharmacological and specific activation of G⍺12 can regulate THBS1 expression and cell migration. GSC23 cells were engineered to express a G⍺12-coupled designer receptor (DREADD)9. Activation of the DREADD-expressing GSC23 cells with CNO (the synthetic ligand for the DREADD) was confirmed to be effective based on robust expression of two canonical G⍺12 and RhoA regulated targets genes, CYR61/CCN1 and CTGF/CCN27 (Fig. 6E). We further demonstrated that CNO treatment increased THBS1 mRNA in GSC23 cells expressing the G⍺12-coupled DREADD (Fig. 6F) and concomitantly increased cell migration (Fig. 6G).
Analysis of TCGA confirmed that THBS1 is highly upregulated in GBM (Fig. 7A). Notably THBS1 mRNA levels also correlated with those of G⍺12 for mesenchymal tumors in TCGA and the more extensive Chinese Glioma Gene Atlas (CGGA) (Fig. 7B). To evaluate involvement of THBS1 in GBM growth in vivo we implanted mice with GSC23 cells in which either of two shRNA constructs were used to knockdown THBS1. (shTHBS1#1 and #3; Supplemental Fig. 2). There were no significant differences in survival or onset of neurologic sequealae compared to controls after 3 weeks mirroring the lack of effect of G⍺12 knockdown on in vivo tumor growth of GSC23. However, brain sections of mice bearing GSC23 THBS1 KD cells revealed tumors that were less invasive (Fig. 7C). Taken together, these data indicate that THBS1 is a G⍺12-regulated gene critical for cell migration and GBM invasion.
Association of G⍺12 and thrombospondin-1 expression in mesenchymal GBM and requirement of THBS1 for GSC tumor invasion. (A) THBS1 is elevated in GBM patients in TCGA database assessed by GlioVis. (B) CCGA dataset indicates positive correlation between GNA12 and THBS1 in GBM patient tumor samples classified as mesenchymal. Pearson’s correlation, HSD p < 0.01. (C) Mouse brain cross sections showing the effect of shTHBS1#1 and shTHBS1#3 KD compared with shControl at 17 days post intracranial injection (H&E or IHC for human nuclei). (HPM, × 40).
Discussion
In this study, we demonstrate a unique and critical role for the heterotrimeric G-protein, Gα12, in GBM. Signaling through heterotrimeric G-proteins depends on their activation by GPCRs, thus our findings implicate endogenous GPCRs and their locally generated ligands in driving GBM tumor progression. G-protein coupled receptors are upregulated and implicated in growth and invasion of numerous cancer types15,18. Recently more than 30 GPCRs were demonstrated to couple to Gα12/139 and, based on their downstream signaling, are likely to regulate cancer progression. We established that many of these receptors had altered profiles in GBM samples included in the TCGA PanCancer dataset, including upregulation of S1PR2, LPAR4, EDNRA, FFAR4, HTR7, and OXGR1. Most strikingly, however, GNA12 was altered in almost one third of the profiled samples, a higher rate than that observed for any of the Gα12-coupled GPCRs. GNA12 was notably upregulated in tumor samples largely distinct from those showing overexpression of these GPCRs. Accordingly, even tumors in which GPCRs are not upregulated would be stimulated through Gα12 regulated pathways in a microenvironment in which their ligands (e.g., thrombin, S1P and LPA) are generated.
Knockdown of G⍺12 decreased G⍺12 mRNA without compensatory increases in G⍺13 mRNA, and with an associated decrease in G⍺12 protein. Tumors observed 2-to-3 weeks after implantation of G⍺12KD GSC23 or HK281 cells did not differ in size nor was there a difference in survival of tumor-bearing mice compared to WT controls. The lack of effect on tumor size likely results from the presence of multiple potential stimuli in the tumor microenvironment that could act independently of G⍺12-coupled receptors to stimulate tumor cell growth30, and indeed there are multiple pathways for YAP activation and YAP-mediated cell proliferation that would remain intact in the KD cells31,32. Consistent with these findings knockdown of G⍺12 did not alter in vitro GSC23 cell proliferation; in addition G⍺12 or G⍺13 deletion has also been reported to have no effect on in vivo growth of pancreatic, breast, or oral cancer cell-derived tumors21,33,34.
On the other hand, tumor cell invasion appears to be highly dependent on G⍺12, as it was significantly diminished in the G⍺12 KD tumors, demonstrated in three separate experiments. This was shown in experiments using both GSC23 and HK281 cells, suggesting that it is a generalizable feature of G⍺12 signaling. Our in vitro experiments confirmed this observation, demonstrating that GSC23 cell migration and invasion were enhanced by S1P in control cells, but not in cells in which G⍺12 was knocked down. Our G⍺12 knockdown studies were complemented by gain-of-function experiments using GSC23 cells expressing a DREADD coupled to G⍺12, in which we demonstrated ligand induced activation of cell migration. Defects in cell migration and invasion were also seen in pancreatic and breast cancer cell derived-tumors in which G⍺12 and or G⍺13 were deleted33,34, a finding extended by our in vivo orthotopic observations.
G⍺12 and G⍺13 couple to RhoGEFs thus their primary effect is the activation of RhoA. Actin cytoskeletal changes could acutely alter cell shape, motility and migration, well established responses induced through RhoA signaling3,35. It is now evident, however, that RhoA activation also leads to transcriptional responses.6,7,11,13,20. Our findings with Gα12 KD glioma stem cells suggest that signaling through Gα12 induces transcriptional responses including genes that characterize mesenchymal-like and stem cell-like states and which would contribute to their invasive phenotype. Of related interest, studies examining DNA copy number alterations in GBM using large-scale network modeling identified GNA12 as a major hub correlated with disease-relevant transcriptional effects36. Together these findings support the hypothesis that G⍺12 activity regulates tumor cell migration and invasion through chronic and sustained transcriptional alterations.
A proneural-to-mesenchymal transition (PMT), has been described for GBM24,28. This resembles the epithelial-to-mesenchymal transition (EMT) observed in other solid cancers, which has been associated with increased stemness and metastasis37. Suggestive evidence for a role of G12 signaling pathway in EMT has been reviewed38, but transcriptional responses mediated through G⍺12/13 signaling have not been previously linked to the proneural-mesenchymal transition (PMT) in glioma cells. Notably, however, glioma stem cells in early passage culture tend to revert to a less aggressive phenotype with a different molecular signature than that of their parental GBM39, and the addition of serum, which contains activators of GPCRs coupled to G⍺12/13 and to RhoA mediated gene expression6, stimulates their transition to a more mesenchymal phenotype24. In addition, expression of the GBM associated gene RPHN2 (rhophilin) activates RhoA and was also reported to lead to mesenchymal transition of GBM cells40. The possibility that transcriptional responses regulated by G⍺12 promote the process of PMT is further supported by our finding that knockdown of G⍺12 in GSCs leads to increases in several genes reflective of a more proneural signature, in particular OLIG2, PATZ1, and TAZ28,41,42. In addition, expression of YKL40 considered by most to be an early marker of a mesenchymal shift in recurrent GBM43,44 was reduced.
Our data demonstrate changes in stem cell frequency and expression of canonical stem cell genes, including key transcription factors like NANOG, SOX and NES, were decreased in the G12 knockdown cells. In addition, classical EMT protein families that modulate cell communication processes, e.g. cadherins (CDH11), collagens, and focal adhesion components (integrins) were highly differentially expressed in the tumor samples. Thus while our data do not conclusively support a PMT shift associated with G⍺12 deletion, as defined by GSEA using the Verhaak classification method19, or demonstrate all of the phenotypic changes associated with altered stemness, this is not unexpected since GBM presents a high degree of phenotypic variability due to its inter- and intra-tumor heterogeneity. Overall, however, the genomic changes we observed, along with the decrease in the tumor’s aggressive features, are compatible with transcriptional reprogramming through altered expression of proneural-mesenchymal and stem cell genes.
We identified thrombospondin-1 (THBS1), associated with the most aggressive and invasive GBM tumors, as one of the most highly downregulated G⍺12 dependent genes in the tumor samples analyzed by DESeq2, demonstrating that THBS1 expression was decreased by 90% in G⍺12 KD tumors. We extended our analysis using GSCs in vitro, demonstrating that THBS1 expression was induced through G⍺12 signaling by S1P, as well as by direct activation of G⍺12 through ligand stimulated DREADD activation. Previous work has linked TGFβ/STAT3 signaling to THBS1 expression25, but to our knowledge, the data we present are the first to implicate GPCR and G⍺12 signaling in expression of thrombospondin-1. Notably, our analysis of the promotor of the THBS1 gene sequence using prediction tools for transcription factors binding sites (e.g., TRANSFAC-based public tools PROMO or MATCH, and TFBSPred, https://www.michalopoulos.net/tfbspred/) revealed a variety of biding sites, including those for MRTFA/SRF and YAP/TAZ/TEADs, transcriptional effectors robustly regulated through G⍺12 signaling.
Our finding that THBS1 knockdown phenocopies that of G⍺12, with little effect on tumor size but a clear change in invasiveness at the tumor border, suggests that upregulation of THBS1 through G⍺12 signaling is one of the transcriptional targets that mediate GBM tumor invasiveness. We show here that its expression also correlated with that of G⍺12 in TCGA tumor samples. While the role of THBS-1 as regulator of cell invasion is less well established than its role in angiogenesis26, it has been shown to regulate the tumor microenvironment, bind to integrins, and activate several protein kinase pathways involved in cell migration including ERK, p38MAPK, and FAK25,45,46. Interestingly, our groups have shown that FAK is activated through integrin signaling downstream of RhoA and ligand induced G⍺12 activation47 and through RhoA signaling in uveal melanoma48. Thus, it will be of interest to determine if FAK serves as a downstream effector of THBS-1 to mediate invasiveness of GBM.
Taken together, our in vivo and in vitro data suggest that GBM tumor cells respond to endogenous GPCR agonists in the tumor environment to engage G⍺12 transcriptional signaling that alters molecular programming of GSCs. Our findings support the notion that activation of G⍺12 signaling contributes to a phenotypic shift towards a more invasive and mesenchymal tumor growth pattern. Thus, downregulation of this signaling pathway could be efficacious in treating GBM by decreasing therapeutic resistance and the tumor infiltration that contributes to recurrence.
Data availability
RNA-seq datasets have been submitted to the Gene Expression Omnibus (GEO) and the accession number provided is GSE229420.
References
Hart, M. J. et al. Direct stimulation of the guanine nucleotide exchange activity of p115 RhoGEF by Galpha13. Science 280, 2112–2114. https://doi.org/10.1126/science.280.5372.2112 (1998).
Suzuki, N., Hajicek, N. & Kozasa, T. Regulation and physiological functions of G12/13-mediated signaling pathways. Neurosignals 17, 55–70. https://doi.org/10.1159/000186690 (2009).
Ridley, A. J. Rho GTPases and actin dynamics in membrane protrusions and vesicle trafficking. Trends Cell Biol. 16, 522–529. https://doi.org/10.1016/j.tcb.2006.08.006 (2006).
Hall, A. Rho GTPases and the actin cytoskeleton. Science 279, 509–514. https://doi.org/10.1126/science.279.5350.509 (1998).
Ren, X. D., Kiosses, W. B. & Schwartz, M. A. Regulation of the small GTP-binding protein Rho by cell adhesion and the cytoskeleton. EMBO J. 18, 578–585. https://doi.org/10.1093/emboj/18.3.578 (1999).
Esnault, C. et al. Rho-actin signaling to the MRTF coactivators dominates the immediate transcriptional response to serum in fibroblasts. Genes Dev. 28, 943–958. https://doi.org/10.1101/gad.239327.114 (2014).
Yu, O. M., Miyamoto, S. & Brown, J. H. Myocardin-related transcription factor a and yes-associated protein exert dual control in g protein-coupled receptor- and RhoA-mediated transcriptional regulation and cell proliferation. Mol. Cell. Biol. 36, 39–49. https://doi.org/10.1128/MCB.00772-15 (2016).
Yu, O. M. et al. YAP and MRTF-A, transcriptional co-activators of RhoA-mediated gene expression, are critical for glioblastoma tumorigenicity. Oncogene 37, 5492–5507. https://doi.org/10.1038/s41388-018-0301-5 (2018).
Inoue, A. et al. Illuminating G-protein-coupling selectivity of GPCRs. Cell 177, 1933–1947. https://doi.org/10.1016/j.cell.2019.04.044 (2019).
Riobo, N. A. & Manning, D. R. Receptors coupled to heterotrimeric G proteins of the G12 family. Trends Pharmacol. Sci. 26, 146–154. https://doi.org/10.1016/j.tips.2005.01.007 (2005).
Yu, O. M. & Brown, J. H. G protein-coupled receptor and RhoA-stimulated transcriptional responses: Links to inflammation, differentiation, and cell proliferation. Mol. Pharmacol. 88, 171–180. https://doi.org/10.1124/mol.115.097857 (2015).
Aragay, A. M. et al. G12 requirement for thrombin-stimulated gene expression and DNA synthesis in 1321N1 astrocytoma cells. J. Biol. Chem. 270, 20073–20077 (1995).
Marinissen, M. J. et al. The small GTP-binding protein RhoA regulates c-jun by a ROCK-JNK signaling axis. Mol. Cell. 14, 29–41 (2004).
Arakaki, A. K. S., Pan, W. A. & Trejo, J. GPCRs in cancer: Protease-activated receptors, endocytic adaptors and signaling. Int. J. Mol. Sci. 19, 1886. https://doi.org/10.3390/ijms19071886 (2018).
Arang, N. & Gutkind, J. S. G Protein-Coupled receptors and heterotrimeric G proteins as cancer drivers. FEBS Lett. 594, 4201–4232. https://doi.org/10.1002/1873-3468.14017 (2020).
Mills, G. B. & Moolenaar, W. H. The emerging role of lysophosphatidic acid in cancer. Nat. Rev. Cancer 3, 582–591. https://doi.org/10.1038/nrc1143 (2003).
Feve, M. et al. Comparative expression study of the endo-G protein coupled receptor (GPCR) repertoire in human glioblastoma cancer stem-like cells, U87-MG cells and non malignant cells of neural origin unveils new potential therapeutic targets. PloS One 9, e91519. https://doi.org/10.1371/journal.pone.0091519 (2014).
Wu, V. et al. Illuminating the Onco-GPCRome: Novel G protein-coupled receptor-driven oncocrine networks and targets for cancer immunotherapy. J. Biol. Chem. 294, 11062–11086. https://doi.org/10.1074/jbc.REV119.005601 (2019).
Verhaak, R. G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110. https://doi.org/10.1016/j.ccr.2009.12.020 (2010).
Mo, J. S., Yu, F. X., Gong, R., Brown, J. H. & Guan, K. L. Regulation of the Hippo-YAP pathway by protease-activated receptors (PARs). Genes Dev. 26, 2138–2143. https://doi.org/10.1101/gad.197582.112 (2012).
Gan, C. P. et al. Heterotrimeric G-protein alpha-12 (Galpha12) subunit promotes oral cancer metastasis. Oncotarget 5, 9626–9640. https://doi.org/10.18632/oncotarget.2437 (2014).
Zhang, G. et al. Chromatin remodeler HELLS maintains glioma stem cells through E2F3 and MYC. JCI Insight https://doi.org/10.1172/jci.insight.126140 (2019).
Jung, J. et al. Nicotinamide metabolism regulates glioblastoma stem cell maintenance. JCI Insight https://doi.org/10.1172/jci.insight.90019 (2017).
Bhat, K. P. L. et al. Mesenchymal differentiation mediated by NF-kappaB promotes radiation resistance in glioblastoma. Cancer Cell 24, 331–346. https://doi.org/10.1016/j.ccr.2013.08.001 (2013).
Daubon, T. et al. Deciphering the complex role of thrombospondin-1 in glioblastoma development. Nat. Commun. 10, 1146. https://doi.org/10.1038/s41467-019-08480-y (2019).
Qi, C. et al. Thrombospondin-1 is a prognostic biomarker and is correlated with tumor immune microenvironment in glioblastoma. Oncol. Lett. 21, 22. https://doi.org/10.3892/ol.2020.12283 (2021).
Maeda, M., Johnson, K. R. & Wheelock, M. J. Cadherin switching: essential for behavioral but not morphological changes during an epithelium-to-mesenchyme transition. J. Cell Sci. 118, 873–887. https://doi.org/10.1242/jcs.01634 (2005).
Fedele, M., Cerchia, L., Pegoraro, S., Sgarra, R. & Manfioletti, G. Proneural-mesenchymal transition: phenotypic plasticity to acquire multitherapy resistance in glioblastoma. Int. J. Mol. Sci. 20, 2746. https://doi.org/10.3390/ijms20112746 (2019).
Marques, C. et al. NF1 regulates mesenchymal glioblastoma plasticity and aggressiveness through the AP-1 transcription factor FOSL1. Elife https://doi.org/10.7554/eLife.64846 (2021).
Bayik, D. & Lathia, J. D. Cancer stem cell-immune cell crosstalk in tumour progression. Nat. Rev. Cancer 21, 526–536. https://doi.org/10.1038/s41568-021-00366-w (2021).
Zhao, B., Li, L., Lei, Q. & Guan, K. L. The Hippo-YAP pathway in organ size control and tumorigenesis: an updated version. Genes Dev. 24, 862–874. https://doi.org/10.1101/gad.1909210 (2010).
Ando, T. et al. EGFR Regulates the Hippo pathway by promoting the tyrosine phosphorylation of MOB1. Commun. Biol. 4, 1237. https://doi.org/10.1038/s42003-021-02744-4 (2021).
Kelly, P. et al. The G12 family of heterotrimeric G proteins promotes breast cancer invasion and metastasis. Proc. Natl. Acad. Sci. U S A 103, 8173–8178. https://doi.org/10.1073/pnas.0510254103 (2006).
Kelly, P. et al. A role for the G12 family of heterotrimeric G proteins in prostate cancer invasion. J. Biol. Chem. 281, 26483–26490. https://doi.org/10.1074/jbc.M604376200 (2006).
Guilluy, C. et al. The Rho GEFs LARG and GEF-H1 regulate the mechanical response to force on integrins. Nat. Cell Biol. 13, 722–727. https://doi.org/10.1038/ncb2254 (2011).
Jornsten, R. et al. Network modeling of the transcriptional effects of copy number aberrations in glioblastoma. Mol. Syst. Biol. 7, 486. https://doi.org/10.1038/msb.2011.17 (2011).
Mani, S. A. et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133, 704–715. https://doi.org/10.1016/j.cell.2008.03.027 (2008).
Rasheed, S. A. K. et al. The emerging roles of Galpha12/13 proteins on the hallmarks of cancer in solid tumors. Oncogene 41, 147–158. https://doi.org/10.1038/s41388-021-02069-w (2022).
Nakod, P. S., Kim, Y. & Rao, S. S. Biomimetic models to examine microenvironmental regulation of glioblastoma stem cells. Cancer Lett. 429, 41–53. https://doi.org/10.1016/j.canlet.2018.05.007 (2018).
Danussi, C. et al. RHPN2 drives mesenchymal transformation in malignant glioma by triggering RhoA activation. Cancer Res. 73, 5140–5150. https://doi.org/10.1158/0008-5472.CAN-13-1168-T (2013).
Liang, Q. et al. Profiling pro-neural to mesenchymal transition identifies a lncRNA signature in glioma. J. Transl. Med. 18, 378. https://doi.org/10.1186/s12967-020-02552-0 (2020).
Bhat, K. P. et al. The transcriptional coactivator TAZ regulates mesenchymal differentiation in malignant glioma. Genes Dev. 25, 2594–2609. https://doi.org/10.1101/gad.176800.111 (2011).
Liu, Q. et al. Molecular properties of CD133+ glioblastoma stem cells derived from treatment-refractory recurrent brain tumors. J. Neurooncol. 94, 1–19. https://doi.org/10.1007/s11060-009-9919-z (2009).
Phillips, H. S. et al. Molecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesis. Cancer Cell 9, 157–173. https://doi.org/10.1016/j.ccr.2006.02.019 (2006).
Firlej, V. et al. Thrombospondin-1 triggers cell migration and development of advanced prostate tumors. Cancer Res. 71, 7649–7658. https://doi.org/10.1158/0008-5472.CAN-11-0833 (2011).
Huang, T., Sun, L., Yuan, X. & Qiu, H. Thrombospondin-1 is a multifaceted player in tumor progression. Oncotarget 8, 84546–84558. https://doi.org/10.18632/oncotarget.19165 (2017).
Del Re, D. P., Miyamoto, S. & Brown, J. H. Focal adhesion kinase as a RhoA-activable signaling scaffold mediating Akt activation and cardiomyocyte protection. J. Biol. Chem. 283, 35622–35629. https://doi.org/10.1074/jbc.M804036200 (2008).
Feng, X. et al. A platform of synthetic lethal gene interaction networks reveals that the GNAQ uveal melanoma oncogene controls the hippo pathway through FAK. Cancer Cell 35, 457–472. https://doi.org/10.1016/j.ccell.2019.01.009 (2019).
Acknowledgements
A special thanks to Dr. Donald Pizzo for the assistance for the histopathology preparations at the UC San Diego Human and Animal Tissue Technology Center at UCSD CALM. We also acknowledge data obtained from tissue processed by the UC San Diego Biorepository and Tissue Technology Shared Resources at Moores Cancer Center, directed by Dr. Valeria Estrada and Dr. Alfredo Molinolo. This project provided short research experience through UCSD Summer Program in Biomedical Sciences in 2019 and 2021, we acknowledge the participation of Taylor Gatesman and Emma Rudolph, both undergraduate trainees from Cellular and Molecular Biology Major Program, Westminster College, New Wilmington, PA, USA. This publication also includes data generated at the UC San Diego IGM Genomics Center utilizing an Illumina NovaSeq 6000 that was purchased with funding from a National Institutes of Health SIG grant (#S10 OD026929).
Funding
This work was supported by NIH grants R01CA218859 to JHB, R03NS131822 to OMC; R01HL145459, and AHA 15GRNTZ297009 to SMiy; R01CA238662, R35CA197718, and R01NS103434 to JNR; NS080939, NS116802, CA258248 to FBF; R01 CA 257505 to JSG.
Author information
Authors and Affiliations
Contributions
Conception and design: O.M.C., F.B.F., S.M., J.H.B. Development of methodology: O.M.C., S.M., J.M., J.L., A.Y.J., N.T., T.G., E.R., J.M.S., M.B. Acquisition of data (provided animals, provided facilities, etc.): O.M.C., J.M., S.M., J.S.G., F.B.F. Analysis and interpretation of data: O.M.C., S.M., B.C.P., J.H.B. Writing, review, and/or revision of the manuscript: O.M.C., S.M., B.C.P., J.S.G., S.M., J.N.R., F.B.F., J.H.B. Administrative, technical, or material support: J.H.B., J.S.G., J.N.R., F.B.F. Study supervision: J.H.B., F.B.F., O.M.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chaim, O.M., Miki, S., Prager, B.C. et al. Gα12 signaling regulates transcriptional and phenotypic responses that promote glioblastoma tumor invasion. Sci Rep 13, 22412 (2023). https://doi.org/10.1038/s41598-023-49164-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-49164-4
- Springer Nature Limited