Abstract
Mutations in Cullin-3 (Cul3), a conserved gene encoding a ubiquitin ligase, are strongly associated with autism spectrum disorder (ASD). Here, we characterize ASD-related pathologies caused by neuron-specific Cul3 knockdown in Drosophila. We confirmed that neuronal Cul3 knockdown causes short sleep, paralleling sleep disturbances in ASD. Because sleep defects and ASD are linked to metabolic dysregulation, we tested the starvation response of neuronal Cul3 knockdown flies; they starved faster and had lower triacylglyceride levels than controls, suggesting defects in metabolic homeostasis. ASD is also characterized by increased biomarkers of oxidative stress; we found that neuronal Cul3 knockdown increased sensitivity to hyperoxia, an exogenous oxidative stress. Additional hallmarks of ASD are deficits in social interactions and learning. Using a courtship suppression assay that measures social interactions and memory of prior courtship, we found that neuronal Cul3 knockdown reduced courtship and learning compared to controls. Finally, we found that neuronal Cul3 depletion alters the anatomy of the mushroom body, a brain region required for memory and sleep. Taken together, the ASD-related phenotypes of neuronal Cul3 knockdown flies establish these flies as a genetic model to study molecular and cellular mechanisms underlying ASD pathology, including metabolic and oxidative stress dysregulation and neurodevelopment.
Similar content being viewed by others
Introduction
Autism spectrum disorder (ASD) is a neurodevelopmental condition associated with dysfunctions in social communication, executive function and cognition, sleep, and metabolism1. Recent estimates find that the prevalence of ASD is one in 44 children2 and that ASD is approximately 83% heritable3. Large-scale genomic sequencing over the past decade has identified more than 100 risk genes and genomic regions associated with the development of ASD4,5,6,7,8,9, but the identification of these genetic risk factors has yet to have a significant impact on the clinical treatment of ASD10,11. As these genetic risk factors span a variety of potential mechanisms contributing to ASD pathology, it is crucial that various genetic models of ASD are generated and explored to advance our understanding of how ASD may arise from multiple mechanisms.
Genomic sequencing studies have implicated de novo mutations in Cullin-3 (Cul3) as causative factors for ASD9,12,13,14,15,16,17,18,19. Based on these studies, the Simons Foundation Autism Research Initiative (SFARI) gives Cul3 a SFARI Gene Score of one20, signifying it as a high confidence autism risk gene. Cul3 encodes a member of the Cullin protein family and acts as an E3 ubiquitin ligase, assembling with adaptor proteins that recruit specific protein substrates for ubiquitination21,22,23,24,25,26. Cul3 is evolutionarily conserved from fruit flies to humans, with 71.67% sequence identity. Drosophila shares approximately 75% of disease-causing genes with humans and offers a wide variety of relevant assays and powerful, publicly available genetic tools27,28,29. Thus, Drosophila melanogaster is a potentially advantageous model system for studying ASD pathophysiology.
Because of this close association with ASD, Cul3 mutants have previously been used to study protein degradation, brain development, and social behaviors in mice30,31,32,33,34,35,36,37. Cul3 has also surfaced in work regarding sleep, locomotion, and neurodevelopment in C. elegans38,39. In fruit flies, neuron-specific knockdown of Cul3 has been studied primarily in the context of neuronal development and sleep40,41,42,43. A conserved, putative Cul3 adaptor protein, Insomniac (Inc), was shown to be important for the proper development of sleep-regulatory neurons in the fruit fly brain44,45. Because sleep disturbances are common in autism46,47,48, these findings suggest a mechanism by which Cul3 lesions may cause a core symptom of ASD. It is not clear whether other symptoms and physiologies associated with ASD can also be observed in flies with neuronal Cul3 knockdown.
Here, we performed a broad characterization of RNAi-mediated Cul3 neuronal-specific knockdown as a potential genetic model for the study of ASD in fruit flies. We first validated the knockdown and the short-sleep phenotype. We then found that these flies were dysregulated in both metabolic and oxidative stress homeostasis, as indicated by increased sensitivity to starvation and hyperoxia. Neuronal Cul3 knockdown flies also had fewer triacylglyceride stores than controls. Through a courtship suppression assay, we found that neuronal Cul3 knockdown flies exhibited significantly less courtship behavior and a learning and memory deficit. Immunostaining revealed that Cul3 neuronal knockdown exhibited alterations in the structure of the mushroom body, a brain region associated with memory and sleep. Thus, neuronal knockdown of Cul3 in fruit flies recapitulates multiple phenotypes of ASD, including defects in sleep, metabolism, social interactions, cognition, and brain development, presenting a powerful model to dissect the underlying mechanisms by which this genetic risk factor might contribute to ASD.
Results
Validating the neuronal knockdown of Cul3 and sleep phenotype
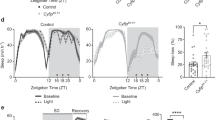
Cul3 encodes a core component of a multiprotein E3 ubiquitin ligase complex that targets specific proteins for ubiquitination (Fig. 1A). To knock down Cul3 expression, we performed RNAi depletion (UAS-Cul3-RNAi) using a neuron-specific Gal4 expression driver (elav-GAL4). We used 7- to 10-day old males for all assays, unless otherwise noted. To validate the neuronal knockdown of Cul3, we assessed Cul3 mRNA levels in male fly heads using quantitative real-time PCR (qRT-PCR). We found that neuronal Cul3 RNAi reduced Cul3 expression by 58% relative to controls containing the RNAi construct alone and 52% relative to controls containing the Gal4 driver alone (Fig. 1B). We then showed by Western blot analysis that neuronal Cul3 RNAi led to a 65% reduction in Cul3 protein compared to controls containing the RNAi construct alone and 64% relative to controls containing the Gal4 driver alone (Fig. 1C). We further confirmed that these flies exhibited a short-sleep phenotype, as previously described40,41 (Fig. 1D). Cul3 RNAi expressed with elav-GAL4 reduced sleep by 42% and 36% respectively relative to controls containing either the RNAi construct or the Gal4 driver.
Validation of Cul3 neuronal knockdown and sleep phenotype. (A) Schematic of the Cul3 protein, an E3 ligase, in the E3 ubiquitin ligase complex that targets proteins from degradation by the proteosome. (B) Neuronal Cul3 knockdown flies exhibited less Cul3 expression than controls by qRT-PCR (p = 0.0036 Gal4 control, p = 0.0010 UAS control, n = number of biological replicates, each containing 30 fly heads). (C) Western blot showing that neuronal Cul3-RNAi flies have reduced Cul3 protein compared to controls (p = 0.0112 Gal4 control, p = 0.0486 UAS control, n = number of biological replicates, each containing 10 fly heads). (D) Relative to controls (gray), flies with neuronal knockdown of Cul3 (teal) exhibited fewer average total minutes of sleep per day (p < 0.0001, each data point represents a single fly). P-values were obtained by Brown-Forsythe and Welch ANOVA test (B,D) and Kruskal–Wallis test (C). Averages are shown with error bars representing SEM.
Neuronal knockdown of Cul3 shortened survival under starvation
Because both sleep deprivation49,50,51 and ASD52,53,54,55 have been linked to metabolic dysregulation, we tested whether Cul3 neuronal knockdown flies exhibited metabolic differences. We performed a starvation survival assay and found that neuronal knockdown of Cul3 significantly shortened survival time under starvation compared to either control (Fig. 2A), suggesting that neuronal Cul3 reduction perturbs metabolic homeostasis. We next characterized the feeding behavior of these flies, using food labeled with a radioactive tracer (see Methods). We found that flies with neuronal knockdown of Cul3 consumed similar amounts of food relative to controls, although one parental control ate significantly more than the other (Fig. 2B). This result suggests that feeding behavior does not contribute to the metabolic differences caused by neuronal Cul3 knockdown. To further understand the starvation sensitivity phenotype, we measured levels of triacylglycerides (TAGs), the main fat storage molecules of both humans and Drosophila . Since neuronal Cul3 knockdown flies were more susceptible to starvation, we hypothesized that they would have lower levels of TAGs relative to controls. Indeed, we found that TAG levels in Cul3 knockdown flies were significantly lower than controls (Fig. 2C). Thus, insufficient TAG stores may contribute to the starvation sensitivity of neuronal Cul3 knockdown flies, suggesting that lipid metabolism is dysregulated in these flies.
Neuronal Cul3 knockdown flies exhibited starvation sensitivity and lower TAG levels but no difference in food consumption. (A) Flies with neuronal knockdown of Cul3 (teal) died faster on starvation media (1% agar) than controls (gray) (p < 0.0001, each line represents 32 flies). (B) Neuronal knockdown of Cul3 flies (teal) ate the same amount of food relative to control flies (gray) (Gal4 control, p = 0.3646; UAS control, p = 0.6578; each data point represents an average of 4–5 flies). (C) Neuronal knockdown of Cul3 flies (teal) had a significantly lower amount of triacylglycerides (TAGs) compared to control flies (Gal4 control, p = 0.0018; UAS control, p < 0.0001; each data point represents an average of 20 flies). p-values were obtained by log-rank test (A) and Brown-Forsythe and Welch ANOVA test (B,C). Averages are shown, with error bars representing SEM.
Neuronal knockdown of Cul3 conferred sensitivity to hyperoxia, an induced oxidative stress.
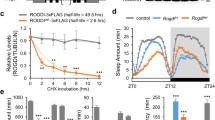
Biomarkers of oxidative stress are reported to be increased in ASD56,57,58. We previously found that short-sleeping flies, including those with neuronal knockdown of Cul3 or inc, are more sensitive to an induced oxidative stress, such as hydrogen peroxide feeding or paraquat injection42. Because these previous methods of inducing oxidative stress are laborious and potentially variable across individual flies, we sought to develop a more consistent and efficient method of applying oxidative stress. To assess sensitivity to oxidative stress, we applied 100% oxygen (hereafter, "hyperoxia")59 to flies in DAMs contained within airtight chambers and monitored their time until death, using terminal loss of activity as a proxy for death (Fig. 3A,B). Using this method, we found that neuronal knockdown of either Cul3 (Fig. 3C) or inc (Fig. 3D) significantly reduced survival time in hyperoxia with respect to controls. This result confirms our previous findings that neuronal knockdown of Cul3 and inc lead to increased sensitivity to oxidative stress, including acute oxidative stress via exogenous hyperoxia.
Flies with neuronal knockdown of Cul3 or inc have increased sensitivity to hyperoxia. (A) Schematic of Drosophila Activity Monitors (DAMs). (B) Schematic of the set-up to apply hyperoxia, or 100% oxygen, to flies in DAMs. (C) Flies with neuronal knockdown of Cul3 (teal) died faster in hyperoxia than controls (gray) (p < 0.0001). (D) Flies with neuronal knockdown of inc (pink) died faster in hyperoxia than controls (gray) (p < 0.0001). Each line represents 32 flies; p-values were obtained by log-rank test (C,D).
Neuronal knockdown of Cul3 caused a courtship defect, as well as a learning and memory defect
ASD is associated with motor difficulties and impairments, such as hypotonia, motor apraxia and motor delay60,61,62. To assess motor function in neuronal Cul3 knockdown flies, we assayed their climbing ability but found no difference in climbing ability relative to controls (Fig. S2).
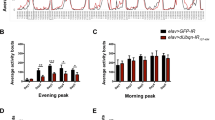
Individuals with ASD typically exhibit altered social interactions and often have cognitive impairments related to learning and memory63,64,65. To test the effects of neuronal knockdown of Cul3 on both social interactions and memory, we used the well-established courtship suppression assay (described in “Methods”, Fig. 4A)66. In brief, male Drosophila exhibit highly stereotyped behaviors to court female Drosophila; these behaviors can be videotaped and quantified. Here we used a combination of FlyTracker and JAABA software, together with a machine learning algorithm, for automated quantitation of male courtship behavior (see “Methods”)67,68. Courtship behavior by naïve males provides a measure of basal social interaction, whereas courtship in experienced (trained) males is also dependent on learning and memory. Males trained with sexually unreceptive females that have mated in the preceding 24 h learn to expect sexual rejection and are less likely to exhibit courtship behavior in their next encounter with a female66.
Flies with neuronal knockdown of Cul3 exhibited a learning and memory deficit and less courtship behavior. (A) Schematic depicting the timeline of our courtship suppression assay, used to assess both courtship behavior and learning and memory. (B) Trained rutabaga mutants (light purple) did not differ in courtship index compared to naïve rutabaga mutants (purple) (p = 0.8022). (C) Trained wCS flies (tan) had a significantly higher courtship index than naïve wCS flies (brown) (p < 0.0001). (D) Trained neuronal Cul3 knockdown flies (light teal) did not differ in courtship index as compared to naïve neuronal Cul3 knockdown flies (teal) (p > 0.9999), while trained controls had significantly lower courtship indices than naïve controls (Gal4 control, p = 0.0005; UAS control, p = 0.0152). Flies with neuronal knockdown of Cul3 (teal) exhibited a lower courtship index than controls (gray) (p < 0.0001). Each point represents an individual fly; p-values were obtained by Mann–Whitney test (B,C; neuronal Cul3 knockdown, D), Kruskal–Wallis test (D, all naïve groups compared), and Welch’s t test (controls, D). Averages are shown with error bars representing SEM.
To test learning and memory, 2- to 4-day-old virgin males of each genotype were split into two groups and housed for 6 h either alone (naïve condition), or with a sexually unreceptive female (trained condition). After 5 days, all males were introduced to a sexually unreceptive female and recorded for 10 min to track and quantify their courtship behavior, as described in the Methods. In the case of normal learning and memory, trained flies exhibit reduced courtship due to memory of their previously rejected courtship attempts. For mutants with learning and memory defects, such as rutabaga (rut)69, which lack function for the enzyme adenylyl cyclase required for synaptic plasticity, trained males will court at the same level as naïve males (Fig. 4B). In contrast, white Canton-S (wCS) males were able to learn and remember previous experiences of sexual rejection even though they have low courtship indices due to vision defects70,71 (Fig. 4C). Thus, the courtship suppression assay is a robust measure of both basal courtship behavior (seen with naïve males) and learning and memory (seen with trained males).
To assess basal courtship behavior, we compared courtship in naïve neuronal Cul3 knockdown males and naïve controls. We found that naïve males with neuronal knockdown of Cul3 had a significantly lower courtship index relative to controls (Fig. 4D). This result indicates that neuronal knockdown of Cul3 reduces courtship behavior, a critical social behavior in fruit flies. When we used this assay to compare courtship of naïve and trained neuronal Cul3 knockdown males, we found that there was no difference in courtship (Fig. 4D), in contrast to trained control males which courted significantly less than their naïve siblings. This result suggests that flies with neuronal knockdown of Cul3 have a learning and memory defect.
Neuronal Cul3 knockdown altered mushroom body anatomy
Autism is characterized by altered nervous system development72, and many ASD risk genes have developmental functions within the brain73. To assess whether deficits in memory and sleep caused by reduced neuronal Cul3 activity might reflect anatomical changes in relevant brain regions, we examined the mushroom body, a structure required for memory and sleep74,75,76. We stained adult brains with an antibody recognizing Fasciclin II (FasII), which prominently marks the axons of mushroom body αβ neurons77. Control brains bearing elav-Gal4 or UAS-Cul3-RNAi alone exhibited normal αβ projections that bifurcated into vertical and medial lobes in each hemisphere (Fig. 5A,B). In contrast, neuronal Cul3 RNAi caused severe abnormalities in αβ axons, with ~ 93% of brains missing at least one lobe and ~ 36% of brains lacking αβ projections entirely (Fig. 5A,B). The loss of αβ axons was associated with enlarged FasII+ clusters, suggesting the accumulation of axons that failed to bifurcate (Fig. 5A,B and Fig. S3). In some brains, αβ projections were present but attenuated, and other brains exhibited morphological irregularities in the ellipsoid body, also marked by FasII (Fig. S3). These results indicate that neuronal Cul3 activity is essential for the development and anatomy of the mushroom body, a structure important for memory and sleep.
Neuronal Cul3 knockdown alters mushroom body anatomy. (A) Adult brains stained with anti-FasII antibody are shown for neuronal driver control (elav-Gal4, left), RNAi construct control (UAS-Cul3-RNAi/ +, center), and neuronal knockdown of Cul3 (elav-Gal4; UAS-Cul3-RNAi/ +, right) genotypes. For Cul3 knockdown panel (right), note missing vertical lobe and attenuated medial lobe in left hemisphere and attenuated medial lobe in right hemisphere. Scale bars 100 μM. (B) Summary data for controls and Cul3 knockdown brains. Number of αβ lobes missing entirely is indicated.
Discussion
Cul3 mutations have been associated with autism spectrum disorder (ASD). In this work, we establish neuronal Cul3 knockdown flies as a model system for the study of ASD. These flies share a variety of behavioral and physiological phenotypes that are associated with ASD in humans. We confirmed that neuronal Cul3 knockdown flies are short sleeping, found that they exhibited increased sensitivity to both starvation and hyperoxia, and discovered that they exhibited decreased courtship behavior and a learning and memory defect. We also found that reduced Cul3 activity in neurons altered the structure of the mushroom body, a brain region vital for memory and sleep. Thus, neuronal Cul3 reduction in Drosophila may serve as a useful model to elucidate mechanisms underlying behavioral and metabolic differences in ASD.
As a ubiquitin ligase, Cul3 binds dozens of adaptors and is likely to ubiquitinate hundreds of substrates78,79,80. This may help to explain why a range of behavioral and physiological phenotypes including sleep, metabolism, oxidative stress response, learning and memory, and social behaviors like courtship are all impacted by the neuronal knockdown of Cul3 in flies and are implicated in ASD. While some specific targets of Cul3 have been identified81,82,83, a systematic investigation of proteins ubiquitinated by Cul3 may provide insight into how this variety of phenotypes in ASD and Cul3 mutants is connected.
While intellectual disabilities and social behaviors are aspects of ASD frequently studied in humans and model systems, our analysis of Cul3 neuronal knockdown flies also revealed two more unusual phenotypes: increased sensitivity to both starvation and hyperoxia, an induced oxidative stress. These findings suggest that reduced Cul3 activity in neurons alters metabolic and oxidative stress homeostasis, both of which are associated with ASD in humans52,53,54,55,56,57,58. Reactive oxygen species, the main propagators of oxidative stress in a cell, are generated by mitochondria during oxidative phosphorylation. Because oxidative phosphorylation is the main mechanism for cellular ATP production, changes in metabolism will directly impact cellular oxidative stress levels. This begs the question of whether neuronal Cul3 knockdown leads to perturbations in metabolism that subsequently cause an increase in oxidative stress, or whether these two phenotypes arise separately. We found that flies with neuronal knockdown of Cul3 had significantly lower triacylglyceride stores, which may explain their sensitivity to starvation and ultimately suggests lipid metabolic dysregulation in these flies. This finding is consistent with studies that have observed lipid dysregulation as well as elevated levels of lipid peroxidation, a form of lipid oxidative damage, that could contribute further to lipid dysregulation in ASD56,57,84. Further study may reveal other aspects of metabolism that are altered by neuronal Cul3 knockdown and whether such metabolic differences increase oxidative stress, establishing these as either related or separate pathologies in ASD.
Although many ASD risk genes are known to impact neuronal development, how specific ASD-associated mutations alter the development and function of discrete brain circuits is not well understood. We found that reduced Cul3 activity in neurons altered projections of the mushroom body, a brain region essential for sleep and memory. Because neuronal Cul3 knockdown also caused defects in sleep and memory, one simple hypothesis is that these phenotypes arise from developmental alterations of mushroom body neurons. Manipulations of Cul3 within the mushroom body and other neuronal subpopulations could test this hypothesis and might enable Cul3 phenotypes, including metabolic dysregulation and susceptibility to oxidative stress, to be mapped with greater resolution within the brain.
Our findings and earlier studies suggest that Cul3 impacts neuronal development by multiple mechanisms. Homozygous Cul3 mutant clones within the mushroom body were found to alter axonal and dendritic development of α′β′ and γ neurons, but have relatively few effects on αβ neurons43. The anatomical defects in αβ neurons caused by neuronal Cul3 RNAi suggest that Cul3 may impact neuronal development through additional, cell non-autonomous mechanisms. The latter may be particularly relevant to autism associated with heterozygous Cul3 mutations, which would be expected to broadly decrease Cul3 activity throughout the brain. Additional effects of Cul3 on neurogenesis, neuronal migration, and synaptic development have been observed in Cul3 mutant mice85,86,87,88. Whether these phenotypes are conserved in flies remains to be addressed.
While Cul3 likely contributes to ASD phenotypes through multiple adaptors and substrates, previous work on Inc, a putative Cul3 adaptor, has suggested their coordinated function during brain development45. Loss of inc function in developing neurons underlies sleep dysregulation in adult inc mutants, and inc mutants also exhibit alterations of mushroom body neurons similar to those caused by Cul3 knockdown. It is unknown whether Cul3 function during development or adulthood underlies the metabolic, oxidative stress, courtship, and learning and memory phenotypes caused by neuronal Cul3 knockdown. Future studies utilizing a conditional neuronal knockdown of Cul3, specific to development or adulthood, may reveal the temporal contributions of Cul3 in ASD-related behaviors and physiologies.
Our findings highlight neuronal knockdown of Cul3 in Drosophila as a useful model to further investigate the mechanisms linking ASD to sleep disturbances, metabolic dysregulation, oxidative stress response, learning and memory, and social deficits. We also establish exposure to 100% oxygen as a simple method to induce hyperoxia and study oxidative stress response in an ASD fly model. As the pathways underlying these various behaviors and physiologies are highly conserved between Drosophila and humans, we anticipate that further study of this Cul3 fly model will be valuable for understanding the pathology of ASD.
Methods
Fly strains and rearing conditions
All flies were raised on glucose food (Archon) in a temperature- (25 °C) and humidity- (65%) controlled incubator with a 12-h light–dark cycle. 7- to 10-day-old males were used for all experiments, unless otherwise noted.
The following flies were used to manipulate inc and Cul3 as described previously41,42: UAS-inc-RNAi (VDRC stock #18225), elavC155-Gal4, UAS-Dicer-2 (Bloomington stock #24651), UAS-Cul3-RNAi (NIG stock #11861R-2), along with the isogenic iso31 strain used for outcrossing. For neuronal Cul3 knockdown experiments, the UAS-Dicer-2 line (Bloomington stock #24651) was crossed into the elavC155-Gal4 line. Parental controls used for experiments were obtained by crossing expression driver and RNAi construct lines to the outcrossed wild-type line for heterozygous controls, accounting for differences in phenotypes affected by genetic background. rutabaga mutants (rut1) were acquired from David Schneider.
Sleep analysis
7- to 10-day-old male flies entrained on a 12:12 light–dark (LD) cycle were placed in individual 5 mm plastic tubes containing food. Tubes were placed in TriKinetics Drosophila Activity Monitors (DAMs) to record their locomotor activity for 2 days in 12:12 LD. Beam break data were grouped into 1 min bins using DAM File Scan, and pySolo (Python-based software) was used to analyze sleep architecture and waking activity. Sleep was defined as a period of at least 5 min of inactivity89.
Hyperoxia assay
7- to 10-day-old male flies entrained on a 12:12 light–dark cycle were placed in individual 5 mm plastic tubes containing food and loaded into Drosophila Activity Monitors. The monitors were placed in airtight chambers (Kent Scientific VetFlo™ Low Cost Induction Chamber) filled with 100% oxygen starting at ZT0 (“lights on”) until all flies were dead. Time of death was determined by complete loss of movement.
Starvation assay
7- to 10-day-old male flies entrained on a 12:12 light–dark cycle were placed in individual 5 mm plastic tubes containing 1% agar and loaded into Drosophila Activity Monitors. Time of death was determined by complete loss of movement.
Survival curves
Survival curves for starvation assays and hyperoxia assays are all plotted as Kaplan–Meier graphs. Log-rank analysis was performed using GraphPad Prism. All experiments were performed with a minimum of three independent trials and yielded statistically similar results, except where noted. Graphs and p-values in figures are from representative trials.
qRT-PCR
7 to 10-day-old male flies entrained to 12:12 LD were frozen at -80ºC and heads were separated by vortexing and collected on ice. RNA was extracted from 30 fly heads for each of 4 biological replicates per genotype with TRIzol (Invitrogen) following the manufacturer’s protocol. Samples were treated with DNaseI (Invitrogen), then heat inactivated. cDNA was synthesized by Revertaid First Strand cDNA Synthesis Kit (Thermo Scientific). PowerUp SYBR Mastermix (Applied Biosystems) was used to perform qRT-PCR using a CFXConnect thermal cycler (BioRad). Primer efficiency and relative quantification of transcripts were determined using a standard curve of serial diluted cDNA. Transcripts were normalized using ribosomal protein S3 (RPS3) as a reference gene.
Primer sequences:
Cul3-fwd-ATGCTACTTTTGTCGCCCATCGC
Cul3-rev-CTGGGTTATCCTTGGTTTATCCTGGCCT
RPS3-fwd-CGAACCTTCCGATTTCCAAGAAACGC
RPS3-rev-ACGACGGACGGCCAGTCCTCC
Feeding assay
As previously described90, approximately 7-day-old flies were fed 32P-labeled Archon food for 24 h. Scintillation counts of accumulated 32P were quantified for an average of 4–5 flies per sample to estimate the amount of food consumed.
Triacylglyceride measurement using thin-layer chromatography
Each biological replicate sample was comprised of 20 male flies. Samples were homogenized in a 2:1 chloroform:ethanol solution, then centrifuged for 10 min at 4 °C. 10 μL of each sample was loaded on a TLC plate (Sigma Aldrich Silica gel on TLC-PET foils 99577-25EA) and separated by polarity using a solvent solution (70 mL n-hexane, 30 mL diethyl ether, and 1 mL acetic acid). A reference sample of 20 wildtype female flies in 2:1 chloroform:ethanol solution was included on each plate. Plates were stained with 0.2% amido black (NAPHTHOL BLUE BLACK, Sigma Product N-3393-100G) in 1 M NaCl and imaged with an iBright™ FL1500 Imaging System. Images were analyzed using FIJI, subtracting individual lane background for each sample and normalizing to the reference sample.
Western blot analysis
Dissected head lysates of female flies (10 flies/sample) were separated by SDS-PAGE using standard procedures. Membranes were probed with antibodies raised against the conserved C-terminal region of human Cul3 at 1:1000 (Cell Signaling, 2759 s) and (HRP)-conjugated monoclonal mouse anti-actin antibody at 1:5000 (Sigma-Aldrich, A3854). Rabbit primary antibody was detected using HRP-conjugated anti-rabbit IgG antibodies at 1:2000 (Cell Signaling, 7074). ECL chemiluminescence reagent (Pierce) was used to visualize horseradish peroxidase activity and detected by CCD camera iBright 1500 (Thermo-Fisher). FIJI gel analysis tool was used for densitometric quantification. A minimum of 5 independent samples for each condition were used for statistical analysis.
Climbing assay
Assessment of climbing ability was performed as previously described91. Eight vials were transferred to an empty standard 23 mm × 95 mm plastic vial and then gently tapped to the bottom. The number of flies that reached the top quarter of the vial within 20 s were then scored as climbing.
Courtship suppression assay
Virgin males were collected within 6 h of eclosion and stored in individual vials. 3–4 days post-collection, individual males were transferred to training vials (15 mm × 65 mm). For the “trained” condition, a sexually unreceptive wCS female (mated within the last 24 h) was added to the training vial with the virgin male; for the “naïve” condition, no female was added to the training vial with the male. Six hours following conditioning, males were isolated and housed in individual vials for another 5 days (memory consolidation period). During testing, trained and naïve males were placed with an unreceptive wCS female in 16 mm diameter chambers without food and videotaped for 10 min to assess courtship memory.
The CalTech FlyTracker and JAABA softwares were used to automatically quantify the Courtship Index (CI), or percent of frames spent courting in a 10 min period, for each male. The Caltech FlyTracker automatically generates data for each video frame such as the location of the male and female, the angle of the wings, the velocity of the flies, etc. We visually inspected each video to ensure tracking accuracy; if a video had tracking errors in > 5% of frames, we discarded the video. We trained the machine learning algorithm (JAABA) to quantify the wing extension, chasing, and attempted mounting behaviors for courtship frame by frame based on the data from the Caltech FlyTracker. Since two or more behaviors can co-occur simultaneously (such as wing extension and chasing in courtship), it is typical in the field to use a hierarchical code that would not double count frames as “courtship” when computing total courtship levels. Our courtship code gives greatest priority to wing extension, followed by chasing and attempted mounting; that is, if wing extension and chasing co-occur in a video frame, JAABA annotates the frame for wing extension only.
Immunostaining and imaging
Brains from adult male progeny were dissected in ice-cold PBS and fixed in 4% paraformaldehyde in PBS for 20 min at room temperature on a nutator. All subsequent incubations and washes were performed at room temperature and on a nutator unless specified otherwise. Fixative was removed and brains were washed in PBS containing 0.3% Triton X-100 (PBST) three times for 15 min each. Samples were then incubated in blocking solution, containing 5% normal donkey serum in PBST, for 30 min, followed by incubation for 2 days at 4 °C in primary antibody cocktail, prepared in blocking solution and containing mouse anti-Fasciclin II (1:50, DSHB, 1D4) and rat anti-elav (1:50, DSHB, 7E8A10) antibodies. Primary antibody cocktail was removed and samples were washed in PBST three times for 15 min each. Samples were then incubated for 2 days at 4 °C in secondary antibody cocktail prepared in blocking solution and containing Alexa 488 donkey anti-mouse (1:1000, Life Technologies, A21202) or Alexa 647 donkey anti-mouse (1:1000, Life Technologies, A31571) and Rhodamine Red-X donkey anti-rat (1:1000, Jackson Immunoresearch, 712-295-153) antibodies; samples were wrapped in foil to protect them from light for this and subsequent steps. Samples were washed three times in PBST for 15 min each and subsequently incubated for 30 min in 0.8 μg/ml DAPI in PBST. Samples were rinsed three times in PBST and mounted in Vectashield (Vector labs) on slides and covered with #1.5 coverslips, using bridges prepared from #2 coverslips. Samples were imaged on Leica Stellaris or SP8 microscopes using a 20 × oil immersion objective to capture z-stacks at 512 × 512 or 1024 × 1024 resolution. The same gain settings were used for experimental and control samples. Maximal projections were generated in Fiji/Image J. The presence or absence of vertical and horizontal mushroom body lobes was scored in a non-blinded manner.
Statistical analysis
We assessed the normality of our data using the D’Agostino-Pearson omnibus normality test and the Shapiro–Wilk normality test, which have good power properties over a wide range of distributions92. For datasets that passed both normality tests, we used the unpaired Student’s t-test with Welch’s correction when comparing two groups and the one-way ANOVA (Dunn’s multiple comparisons) when comparing three or more groups. For data sets that failed either one of the normality tests, we used the Mann–Whitney U test when comparing two groups and the Kruskal–Wallis test with Dunn’s post hoc test when comparing three or more groups. Significance is expressed as pvalues (n.s., p > 0.05; *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001).
Data availability
The authors declare that all data supporting the findings of this study are available, including replicate experiments, and will be made available upon reasonable request to the corresponding author, Dr. Mimi Shirasu-Hiza.
References
Association, A. P. Diagnostic and Statistical Manual of Mental Disorders (DSM-5). (American Psychiatric Association Publishing, 2013).
Maenner, M. J., Bakian, A. V., Bilder, D. A., Durkin, M. S., Esler, A., Furnier, S. M., Hallas, L., Hall-Lande, J., Hudson, A., Hughes, M. M., Patrick, M., Pierce, K., Poynter, J. N., Salinas, A., Shenouda, J., Vehorn, A., Warren, Z., Constantino, J. N., DiRienzo, M., Fitzgerald, R. T., Grzybowski, A., Spivey, M. H., Pettygrove, S., Zahorodny, W., Ali, A., Andrews, J. G., Baroud, T., Gutierrez, J., Hewitt, A., Lee, L. C., Lopez, M., Mancilla, K. C., McArthur, D., Schwenk, Y. D., Washington, A., Williams, S., & Cogswell, M. E. Prevalence and characteristics of autism spectrum disorder among children aged 8 years—autism and developmental disabilities monitoring network. 1–16 (Centers for Disease Control and Prevention, United States, 2021).
Sandin, S. et al. The heritability of autism spectrum disorder. JAMA 318, 1182–1184. https://doi.org/10.1001/jama.2017.12141 (2017).
Sanders, S. J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241. https://doi.org/10.1038/nature10945 (2012).
Iossifov, I. et al. Low load for disruptive mutations in autism genes and their biased transmission. 112, E5600–E5607. https://doi.org/10.1073/pnas.1516376112 (2015).
Sanders, S. J. et al. Insights into autism spectrum disorder genomic architecture and biology from 71 risk loci. Neuron 87, 1215–1233. https://doi.org/10.1016/j.neuron.2015.09.016 (2015).
Satterstrom, F. K. et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell 180, 568-584.e523. https://doi.org/10.1016/j.cell.2019.12.036 (2020).
Searles Quick, V. B., Wang, B. & State, M. W. Leveraging large genomic datasets to illuminate the pathobiology of autism spectrum disorders. Neuropsychopharmacology 46, 55–69. https://doi.org/10.1038/s41386-020-0768-y (2021).
Zhou, X. et al. Integrating de novo and inherited variants in 42,607 autism cases identifies mutations in new moderate-risk genes. Nat. Genet. 54, 1305–1319. https://doi.org/10.1038/s41588-022-01148-2 (2022).
Manoli, D. S. & State, M. W. Autism spectrum disorder genetics and the search for pathological mechanisms. Am. J. Psychiatry 178, 30–38. https://doi.org/10.1176/appi.ajp.2020.20111608 (2021).
Patel, J., Lukkes, J. L. & Shekhar, A. Overview of genetic models of autism spectrum disorders. Prog. Brain Res. 241, 1–36. https://doi.org/10.1016/bs.pbr.2018.10.002 (2018).
O’Roak, B. J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485, 246–250. https://doi.org/10.1038/nature10989 (2012).
Kong, A. et al. Rate of de novo mutations and the importance of father’s age to disease risk. Nature 488, 471–475. https://doi.org/10.1038/nature11396 (2012).
Codina-Sola, M. et al. Integrated analysis of whole-exome sequencing and transcriptome profiling in males with autism spectrum disorders. Mol. Autism 6, 21. https://doi.org/10.1186/s13229-015-0017-0 (2015).
Wang, T. et al. De novo genic mutations among a Chinese autism spectrum disorder cohort. Nat. Commun. 7, 13316. https://doi.org/10.1038/ncomms13316 (2016).
Stessman, H. A. et al. Targeted sequencing identifies 91 neurodevelopmental-disorder risk genes with autism and developmental-disability biases. Nat. Genet. 49, 515–526. https://doi.org/10.1038/ng.3792 (2017).
Guo, H. et al. Inherited and multiple de novo mutations in autism/developmental delay risk genes suggest a multifactorial model. Mol. Autism 9, 64. https://doi.org/10.1186/s13229-018-0247-z (2018).
Nakashima, M. et al. De novo variants in CUL3 are associated with global developmental delays with or without infantile spasms. J. Hum. Genet. 65, 727–734. https://doi.org/10.1038/s10038-020-0758-2 (2020).
Ruzzo, E. K. et al. Inherited and de novo genetic risk for autism impacts shared networks. Cell 178, 850–866. https://doi.org/10.1016/j.cell.2019.07.015 (2019).
Cul3. SFARI Gene.
Jerabkova, K. & Sumara, I. Cullin 3, a cellular scripter of the non-proteolytic ubiquitin code. Semin. Cell Dev. Biol. 93, 100–110. https://doi.org/10.1016/j.semcdb.2018.12.007 (2019).
Harper, J. W. & Schulman, B. A. Cullin-RING ubiquitin ligase regulatory circuits: A quarter century beyond the F-Box hypothesis. Annu. Rev. Biochem. 90, 403–429. https://doi.org/10.1146/annurev-biochem-090120-013613 (2021).
Furukawa, M., He, Y. J., Borchers, C. & Xiong, Y. Targeting of protein ubiquitination by BTB-Cullin 3-Roc1 ubiquitin ligases. Nat. Cell Biol. 5, 1001–1007. https://doi.org/10.1038/ncb1056 (2003).
Geyer, R., Wee, S., Anderson, S., Yates, J. & Wolf, D. A. BTB/POZ domain proteins are putative substrate adaptors for cullin 3 ubiquitin ligases. Mol. Cell 12, 783–790. https://doi.org/10.1016/s1097-2765(03)00341-1 (2003).
Pintard, L. et al. The BTB protein MEL-26 is a substrate-specific adaptor of the CUL-3 ubiquitin-ligase. Nature 425, 311–316. https://doi.org/10.1038/nature01959 (2003).
Xu, L. et al. BTB proteins are substrate-specific adaptors in an SCF-like modular ubiquitin ligase containing CUL-3. Nature 425, 316–321. https://doi.org/10.1038/nature01985 (2003).
Ugur, B., Chen, K. & Bellen, H. J. Drosophila tools and assays for the study of human diseases. Dis. Model Mech. 9, 235–244. https://doi.org/10.1242/dmm.023762 (2016).
Bellosta, P. & Soldano, A. Dissecting the genetics of autism spectrum disorders: A drosophila perspective. Front. Physiol. 10, 987. https://doi.org/10.3389/fphys.2019.00987 (2019).
Coll-Tane, M., Krebbers, A., Castells-Nobau, A., Zweier, C. & Schenck, A. Intellectual disability and autism spectrum disorders 'on the fly': Insights from Drosophila. Dis. Model Mech. 12. https://doi.org/10.1242/dmm.039180 (2019).
McEvoy, J. D., Kossatz, U., Malek, N. & Singer, J. D. Constitutive turnover of cyclin E by Cul3 maintains quiescence. Mol. Cell Biol. 27, 3651–3666. https://doi.org/10.1128/MCB.00720-06 (2007).
Rapanelli, M. et al. Behavioral, circuitry, and molecular aberrations by region-specific deficiency of the high-risk autism gene Cul3. Mol. Psychiatr. 26, 1491–1504. https://doi.org/10.1038/s41380-019-0498-x (2021).
Dong, Z. Q. et al. CUL3 deficiency causes social deficits and anxiety-like behaviors by impairing excitation-inhibition balance through the promotion of cap-dependent translation. Neuron 105, 475. https://doi.org/10.1016/j.neuron.2019.10.035 (2020).
Amar, M. et al. Autism-linked Cullin3 germline haploinsufficiency impacts cytoskeletal dynamics and cortical neurogenesis through RhoA signaling. Mol. Psychiatr. 26, 3586–3613. https://doi.org/10.1038/s41380-021-01052-x (2021).
Morandell, J. et al. Cul3 regulates cytoskeleton protein homeostasis and cell migration during a critical window of brain development. Nat. Commun. 12. https://doi.org/10.1038/s41467-021-23123-x (2021).
Rapanelli, M. et al. Cholinergic neurons in the basal forebrain are involved in behavioral abnormalities associated with Cul3 deficiency: Role of prefrontal cortex projections in cognitive deficits. Transl. Psychiat. 13, 22. https://doi.org/10.1038/s41398-023-02306-8 (2023).
Xia, Q. Q. et al. Effects of heterozygous deletion of autism-related gene Cullin-3 in mice. PLoS One 18, e0283299. https://doi.org/10.1371/journal.pone.0283299 (2023).
Lin, P. et al. Current trends of high-risk gene Cul3 in neurodevelopmental disorders. Front. Psychiatry 14, 1215110. https://doi.org/10.3389/fpsyt.2023.1215110 (2023).
Wong, W. R. et al. Autism-associated missense genetic variants impact locomotion and neurodevelopment in Caenorhabditis elegans. Hum. Mol. Genet. 28, 2271–2281. https://doi.org/10.1093/hmg/ddz051 (2019).
Singh, K., Ju, J. Y., Walsh, M. B., DiIorio, M. A. & Hart, A. C. Deep conservation of genes required for both Drosophila melanogaster and Caenorhabditis elegans sleep includes a role for dopaminergic signaling. Sleep 37, 1439-U1256. https://doi.org/10.5665/sleep.3990 (2014).
Pfeiffenberger, C. & Allada, R. Cul3 and the BTB adaptor insomniac are key regulators of sleep homeostasis and a dopamine arousal pathway in Drosophila. PLoS Genet. 8, e1003003. https://doi.org/10.1371/journal.pgen.1003003 (2012).
Stavropoulos, N. & Young, M. W. insomniac and Cullin-3 regulate sleep and wakefulness in Drosophila. Neuron 72, 964–976. https://doi.org/10.1016/j.neuron.2011.12.003 (2011).
Hill, V. M. et al. A bidirectional relationship between sleep and oxidative stress in Drosophila. PLOS Biol. 16, e2005206. https://doi.org/10.1371/journal.pbio.2005206 (2018).
Zhu, S. J., Perez, R., Pan, M. & Lee, T. M. Requirement of Cul3 for axonal arborization and dendritic elaboration in Drosophila mushroom body neurons. J. Neurosci. 25, 4189–4197. https://doi.org/10.1523/Jneurosci.0149-05.2005 (2005).
Li, Q. et al. Conserved properties of Drosophila Insomniac link sleep regulation and synaptic function. PLoS Genet. 13, e1006815. https://doi.org/10.1371/journal.pgen.1006815 (2017).
Li, Q., Jang, H., Lim, K. Y., Lessing, A. & Stavropoulos, N. insomniac links the development and function of a sleep-regulatory circuit. Elife 10. https://doi.org/10.7554/eLife.65437 (2021).
Ballester, P., Richdale, A. L., Baker, E. K. & Peiro, A. M. Sleep in autism: A biomolecular approach to aetiology and treatment. Sleep Med. Rev. 54, 101357. https://doi.org/10.1016/j.smrv.2020.101357 (2020).
Carmassi, C. et al. Systematic review of sleep disturbances and circadian sleep desynchronization in autism spectrum disorder: Toward an integrative model of a self-reinforcing loop. Front. Psychiatry 10, 366. https://doi.org/10.3389/fpsyt.2019.00366 (2019).
Deliens, G., Leproult, R., Schmitz, R., Destrebecqz, A. & Peigneux, P. Sleep disturbances in autism spectrum disorders. Rev. J. Autism Dev. Disord. 2, 343–356. https://doi.org/10.1007/s40489-015-0057-6 (2015).
Knutson, K. L., Spiegel, K., Penev, P. & Van Cauter, E. The metabolic consequences of sleep deprivation. Sleep Med. Rev. 11, 163–178. https://doi.org/10.1016/j.smrv.2007.01.002 (2007).
Sharma, S. & Kavuru, M. Sleep and metabolism: An overview. Int. J. Endocrinol. 2010. https://doi.org/10.1155/2010/270832 (2010).
Mullington, J. M., Haack, M., Toth, M., Serrador, J. M. & Meier-Ewert, H. K. Cardiovascular, inflammatory, and metabolic consequences of sleep deprivation. Prog. Cardiovasc. Dis. 51, 294–302. https://doi.org/10.1016/j.pcad.2008.10.003 (2009).
De Angelis, M. et al. Fecal microbiota and metabolome of children with autism and pervasive developmental disorder not otherwise specified. PLoS One 8, e76993. https://doi.org/10.1371/journal.pone.0076993 (2013).
Lanz, T. A. et al. Transcriptomic analysis of genetically defined autism candidate genes reveals common mechanisms of action. Mol. Autism 4, 45. https://doi.org/10.1186/2040-2392-4-45 (2013).
Ming, X., Stein, T. P., Barnes, V., Rhodes, N. & Guo, L. Metabolic perturbance in autism spectrum disorders: A metabolomics study. J. Proteome Res. 11, 5856–5862. https://doi.org/10.1021/pr300910n (2012).
Orozco, J. S., Hertz-Picciotto, I., Abbeduto, L. & Slupsky, C. M. Metabolomics analysis of children with autism, idiopathic-developmental delays, and Down syndrome. Transl. Psychiatry 9, 243. https://doi.org/10.1038/s41398-019-0578-3 (2019).
Ming, X. et al. Increased excretion of a lipid peroxidation biomarker in autism. Prostaglandins Leukot Essent Fatty Acids 73, 379–384. https://doi.org/10.1016/j.plefa.2005.06.002 (2005).
Bjorklund, G. et al. Oxidative stress in autism spectrum disorder. Mol. Neurobiol. 57, 2314–2332. https://doi.org/10.1007/s12035-019-01742-2 (2020).
Manivasagam, T. et al. in Personalized food intervention and therapy for autism spectrum disorder management (eds M. Mohamed Essa & M. Walid Qoronfleh) 193–206 (Springer International Publishing, 2020).
Walker, D. W. & Benzer, S. Mitochondrial, “swirls” induced by oxygen stress and in the Drosophila mutant hyperswirl. Proc. Natl. Acad. Sci. USA 101, 10290–10295. https://doi.org/10.1073/pnas.0403767101 (2004).
Matson, M. L., Matson, J. L. & Beighley, J. S. Comorbidity of physical and motor problems in children with autism. Res. Dev. Disabil. 32, 2304–2308. https://doi.org/10.1016/j.ridd.2011.07.036 (2011).
Ming, X., Brimacombe, M. & Wagner, G. C. Prevalence of motor impairment in autism spectrum disorders. Brain Dev. 29, 565–570. https://doi.org/10.1016/j.braindev.2007.03.002 (2007).
Fournier, K. A., Hass, C. J., Naik, S. K., Lodha, N. & Cauraugh, J. H. Motor coordination in autism spectrum disorders: A synthesis and meta-analysis. J. Autism Dev. Disord. 40, 1227–1240. https://doi.org/10.1007/s10803-010-0981-3 (2010).
Goh, S. & Peterson, B. S. Imaging evidence for disturbances in multiple learning and memory systems in persons with autism spectrum disorders. Dev. Med. Child Neurol. 54, 208–213. https://doi.org/10.1111/j.1469-8749.2011.04153.x (2012).
Solomon, M., McCauley, J. B., Iosif, A. M., Carter, C. S. & Ragland, J. D. Cognitive control and episodic memory in adolescents with autism spectrum disorders. Neuropsychologia 89, 31–41. https://doi.org/10.1016/j.neuropsychologia.2016.05.013 (2016).
Williams, D. L., Goldstein, G. & Minshew, N. J. The profile of memory function in children with autism. Neuropsychology 20, 21–29. https://doi.org/10.1037/0894-4105.20.1.21 (2006).
Ejima, A. & Griffith, L. C. Assay for courtship suppression in Drosophila. Cold Spring Harb. Protoc. 2011, pdb prot5575. https://doi.org/10.1101/pdb.prot5575 (2011).
Eyjolfsdottir, E. et al. 772–787 (Springer International Publishing).
Kabra, M., Robie, A. A., Rivera-Alba, M., Branson, S. & Branson, K. JAABA: Interactive machine learning for automatic annotation of animal behavior. Nat. Methods 10, 64–67. https://doi.org/10.1038/nmeth.2281 (2013).
Livingstone, M. S., Sziber, P. P. & Quinn, W. G. Loss of calcium/calmodulin responsiveness in adenylate cyclase of rutabaga, a Drosophila learning mutant. Cell 37, 205–215. https://doi.org/10.1016/0092-8674(84)90316-7 (1984).
Krstic, D., Boll, W. & Noll, M. Influence of the White locus on the courtship behavior of Drosophila males. PLoS One 8, e77904. https://doi.org/10.1371/journal.pone.0077904 (2013).
Xiao, C., Qiu, S. & Robertson, R. M. The white gene controls copulation success in Drosophila melanogaster. Sci. Rep. 7, 7712. https://doi.org/10.1038/s41598-017-08155-y (2017).
Courchesne, E. et al. Mapping early brain development in autism. Neuron 56, 399–413. https://doi.org/10.1016/j.neuron.2007.10.016 (2007).
Hu, W. F., Chahrour, M. H. & Walsh, C. A. The diverse genetic landscape of neurodevelopmental disorders. Annu. Rev. Genom. Hum. Genet. 15, 195–213. https://doi.org/10.1146/annurev-genom-090413-025600 (2014).
de Belle, J. S. & Heisenberg, M. Associative odor learning in Drosophila abolished by chemical ablation of mushroom bodies. Science 263, 692–695. https://doi.org/10.1126/science.8303280 (1994).
Joiner, W. J., Crocker, A., White, B. H. & Sehgal, A. Sleep in Drosophila is regulated by adult mushroom bodies. Nature 441, 757–760. https://doi.org/10.1038/nature04811 (2006).
Pitman, J. L., McGill, J. J., Keegan, K. P. & Allada, R. A dynamic role for the mushroom bodies in promoting sleep in Drosophila. Nature 441, 753–756. https://doi.org/10.1038/nature04739 (2006).
Cheng, Y. et al. Drosophila fasciclinII is required for the formation of odor memories and for normal sensitivity to alcohol. Cell 105, 757–768. https://doi.org/10.1016/s0092-8674(01)00386-5 (2001).
Bennett, E. J., Rush, J., Gygi, S. P. & Harper, J. W. Dynamics of cullin-RING ubiquitin ligase network revealed by systematic quantitative proteomics. Cell 143, 951–965. https://doi.org/10.1016/j.cell.2010.11.017 (2010).
Stogios, P. J., Downs, G. S., Jauhal, J. J., Nandra, S. K. & Prive, G. G. Sequence and structural analysis of BTB domain proteins. Genome Biol. 6, R82. https://doi.org/10.1186/gb-2005-6-10-r82 (2005).
Emanuele, M. J. et al. Global identification of modular cullin-RING ligase substrates. Cell 147, 459–474. https://doi.org/10.1016/j.cell.2011.09.019 (2011).
Maerki, S. et al. The Cul3–KLHL21 E3 ubiquitin ligase targets Aurora B to midzone microtubules in anaphase and is required for cytokinesis. J. Cell Biol. 187, 791–800. https://doi.org/10.1083/jcb.200906117 (2009).
Rondou, P., Haegeman, G., Vanhoenacker, P. & Van Craenenbroeck, K. BTB protein KLHL12 targets the dopamine D4 receptor for ubiquitination by a Cul3-based E3 ligase. J. Biol. Chem. 283, 11083–11096. https://doi.org/10.1074/jbc.M708473200 (2008).
Zhang, D. D. et al. Ubiquitination of Keap1, a BTB-Kelch substrate adaptor protein for Cul3, targets Keap1 for degradation by a proteasome-independent pathway. J. Biol. Chem. 280, 30091–30099. https://doi.org/10.1074/jbc.M501279200 (2005).
Tamiji, J. & Crawford, D. A. The neurobiology of lipid metabolism in autism spectrum disorders. Neurosignals 18, 98–112. https://doi.org/10.1159/000323189 (2010).
Rapanelli, M. et al. Behavioral, circuitry, and molecular aberrations by region-specific deficiency of the high-risk autism gene Cul3. Mol. Psychiatry 26, 1491–1504. https://doi.org/10.1038/s41380-019-0498-x (2021).
Dong, Z. et al. CUL3 deficiency causes social deficits and anxiety-like behaviors by impairing excitation-inhibition balance through the promotion of cap-dependent translation. Neuron 105, 475–490. https://doi.org/10.1016/j.neuron.2019.10.035 (2020).
Amar, M. et al. Autism-linked Cullin3 germline haploinsufficiency impacts cytoskeletal dynamics and cortical neurogenesis through RhoA signaling. Mol. Psychiatry 26, 3586–3613. https://doi.org/10.1038/s41380-021-01052-x (2021).
Morandell, J. et al. Cul3 regulates cytoskeleton protein homeostasis and cell migration during a critical window of brain development. Nat. Commun. 12, 3058. https://doi.org/10.1038/s41467-021-23123-x (2021).
Huber, R. et al. Sleep homeostasis in Drosophila melanogaster. Sleep 27, 628–639. https://doi.org/10.1093/sleep/27.4.628 (2004).
Deshpande, S. A. et al. Quantifying Drosophila food intake: Comparative analysis of current methodology. Nat. Methods 11, 535–540. https://doi.org/10.1038/nmeth.2899 (2014).
Ulgherait, M. et al. Circadian autophagy drives iTRF-mediated longevity. Nature 598, 353–358. https://doi.org/10.1038/s41586-021-03934-0 (2021).
Yap, B. W. & Sim, C. H. Comparisons of various types of normality tests. J. Stat. Comput. Simul. 81, 2141–2155. https://doi.org/10.1080/00949655.2010.520163 (2011).
Acknowledgements
We thank all members of the Shirasu-Hiza and Canman labs for support, discussions, and feedback. We thank David Schneider for the rutabaga fly line. We also thank Shuhao Li for assistance with brain dissections. We thank the Waksman Institute Shared Imaging Facility, Rutgers, The State University of New Jersey, for use of microscopes. Work was supported by NIH F31AG074664 (SJT), Columbia University SURF (KO), AFAR Glenn Foundation Postdoctoral Fellowship for Aging Research (MU), T32GM141882 (AMM), NIH F31AG079601 (JAG), NIH R01NS112844 (NS), NIH R21NS111304 (NS), NIH R01DC020031 (WWJ), NIH R01GM117407 (JCC), NIH R01GM130764 (JCC), NIH R35GM127049 (MSH) and NIH R01AG045842 (MSH).
Author information
Authors and Affiliations
Contributions
S.J.T. and M.S.H. conceived all experiments except brain staining and imaging, which were conceived by E.V.B. and N.S. Experiments were performed and analyzed by S.J.T. (husbandry, sleep, hyperoxia survival, TAG measurement), Z.L. (sleep, starvation, courtship suppression assay), S.J.P. and W.W.J. (feeding assay), K.O. (sleep, hyperoxia survival), M.U. (qRT-PCR, western blot), E.V.B. and N.S. (brain staining and imaging), A.M.M. (qRT-PCR), M.P. (courtship suppression assay), J.A.G. (climbing assay), and J.V. (husbandry). S.J.T., Z.L., M.U., A.M.M., J.V., N.S., W.W.J., J.C.C., and M.S.H. made intellectual contributions. S.J.T., N.S., J.C.C., and M.S.H. designed the figures and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tener, S.J., Lin, Z., Park, S.J. et al. Neuronal knockdown of Cullin3 as a Drosophila model of autism spectrum disorder. Sci Rep 14, 1541 (2024). https://doi.org/10.1038/s41598-024-51657-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-51657-9
- Springer Nature Limited