Abstract
The objective of this study was to analyze the antimicrobial resistance (AMR) characteristics produced by antibiotic resistance genes (ARGs), mobile genetic elements (MGEs) and gene cassettes in Escherichia coli isolated from the feces of captive black bears. Antimicrobial susceptibility testing was performed by using the disk diffusion method, and both MGEs and integron gene cassettes were detected by polymerase chain reaction. Our results showed that 43.7% (62/142) of the isolates were multidrug resistant strains and 97.9% (139/142) of the isolates were resistant to at least one antibiotic. The highest AMR phenotype was observed for tetracycline (79.6%, 113/142), followed by ampicillin (50.0%, 71/142), trimethoprim-sulfamethoxazole (43.7%, 62/142) and cefotaxime (35.9%, 51/142). However, all isolates were susceptible to tobramycin. tetA had the highest occurrence in 6 ARGs in 142 E. coli isolates (76.8%, 109/142). Ten mobile genetic elements were observed and IS26 was dominant (88.0%, 125/142). ISECP1 was positively associated with five β-lactam antibiotics. ISCR3/14, IS1133 and intI3 were not detected. Seventy-five E. coli isolates (65 intI1-positive isolates, 2 intI2-positive isolates and 8 intI1 + intI2-positive isolates) carried integrons. Five gene cassettes (dfrA1, aadA2, dfrA17-aadA5, aadA2-dfrA12 and dfrA1-aadA1) were identified in the intI1-positive isolates and 2 gene cassettes (dfrA1-catB2-sat2-aadA1 and dfrA1-catB2-sat1-aadA1) were observed in the intI2-positive isolates. Monitoring of ARGs, MGEs and gene cassettes is important to understand the prevalence of AMR, which may help to introduce measures to prevent and control of AMR in E. coli for captive black bears.
Similar content being viewed by others
Introduction
Antimicrobial resistance (AMR) is a rapidly growing global health problem in animals and humans1. Antimicrobial resistance genes (ARGs) have been considered as a major mechanism of bacterial resistance to antibiotics2. Furthermore, the presence of bacteria carrying ARGs has been widely reported in wildlife in previous studies, which are major factors in the emergence of global health challenges3. Mobile genetic elements (MGEs), such as integrons, transposons and plasmids, play an important role in the spread of ARGs in animals and the environment4. Integrons, a type of MGE, can easily transfer one or more gene cassettes between different bacteria. Different gene cassettes that are incorporated into integrons have a significant correlation with resistance to different antibiotics5,6.
Black bear (Ursus thibetanus) is one of the national first-class protected wild animals in China, and there are more than 97 facilities are maintained for black bears and other bears in China7,8. There have been more reports on the epidemiological investigation of antibiotic resistant bacteria in captive wild animals9,10,11,and study for E. coli from sloth bear in India showed high prevalence of antimicrobial resistance, indicating that the AMR E. coli from captive black bears should be concerned12.Overuse of antibiotics has led to a problem of antibiotic resistant in clinical practice13. Previous studies have shown that the feces of healthy wild animals may serve as a reservoir for antibiotic resistant Escherichia coli (E. coli), which could pose a threat to public health and environmental safety14,15. Captive black bears have close contact with humans, including animal keepers and veterinarians. Exposure to antibiotic resistant E. coli from black bear feces may pose a potential risk to other animals and public health. To our knowledge, there is little information on the prevalence of AGRs, MGEs, integron gene cassettes, and the association between AGRs and MGEs in E. coli from captive black bears. The aim of this study was to characterize the antimicrobial resistance, especially the MGEs, ARGs and integron gene cassettes, of 142 E. coli isolates collected from the feces of captive black bears.
Materials and methods
Sample collection and bacterial strain identification
This study collected 142 fecal samples from a black bear breeding farm in Dujiang Yan city, China. Each sample was collected from one individual. Fresh fecal specimens (approximately 10 g) from each black bear were collected immediately by feeders after defecation on the ground and then quickly transferred into individual 50-mL plastic containers. All isolates (per isolate correspond to per sample) were identified using Gram staining, MacConkey (Solarbio, Beijing), and then confirmed by eosin methylene blue agar growth (Chromagar, France), and biochemical identification by API 20E system (BioMerieux, France)16. The verified and confirmed E. coli isolates were resuspended in tryptic soya broth plus 20% glycerol and stored at -20 ℃. A total of 142 strains were identified as DJY1-DJY142. DJY indicates the source of isolates from Dujiang Yan city, which is used in our lab for E. coli isolates from captive black bears.
Analysis of antibiotic sensitivity and resistance patterns
Antimicrobial susceptibility was determined for all isolates against 13 antimicrobial agents via a standard disk diffusion test. The following antimicrobial disks (Oxoid) were used: aminoglycosides (gentamicin, GM, 10 μg; tobramycin, TOB, 10 μg), chloram phenicols (chloramphenicol, C, 30 μg), quinolones (ciprofloxacin, CIP, 5 μg), sulfonamide (sulfamethoxazole, RL, 25 μg), tetracyclines (tetracycline, TE, 30 μg), β-lactams (cephazolin, KZ, 30 μg; cefuroxime sodium, CXM, 30 μg; cefotaxime, CTX, 30 μg; cefepime, FEP, 30 μg; aztreonam, ATM, 30 μg; ampicillin, AMP, 10 μg; ampicillin/sulbactam 1:1, SAM, 10/10 μg). Results were interpreted based on the CLSI 2021 criteria. E. coli ATCC25922 was used as a control. The antibiotics used in this study were based on the information provided by the local farm veterinarians (CN, CIP, CTX, CXM and SAM were used for disease control); TOB, TE, C, KZ, FEP, CIP, SXT, ATM and AMP have been reported to be found antibiotic resistant in wildlife17,18.
Screening for ARGs, MGEs and gene cassettes
Total genomic DNA was extracted using a kit (Tiangen Biotech, China). All genomic DNA solutions were stored at -20℃. According to previous studies, 6 antibiotic resistance genes (ARGs), 13 mobile genetic elements (MGEs) and gene cassettes were selected and detected by polymerase chain reaction (PCR)19,20,21,22. PCR assays were carried out in 25 μL volumes containing 2 μL template DNA, 12.5 μL 2 × Taq PCR Master Mix (Tsingke, China), 8.5 μL ddH2O (Solarbio, China) and 1 μL each primer. The PCR products were separated by gel electrophoresis in a 1.5% agarose gel stained with GoldView™ (Sangon Biotech, China), visualized under ultraviolet light and photographed using a gel documentation system (BioRad, USA). The primers and amplification conditions used have been previously described in Table S123,24.
Statistical analysis
All positive PCR products were directly sequenced in both directions by BGI (Beijing, China). Sequences were analyzed online by BLAST (http://blast.ncbi.nlm.nih.gov). P-values < 0.05 were considered to be statistically significant. The association between AMR phenotypes and the MGEs was calculated, and was considered significant at a P-value of < 0.05 and was reported as an odds ratio (OR) with 95% confidence interval (CI). An OR > 1 was considered a positive association or an increasing likelihood of co-occurrence of the MGEs or AMR phenotype, while an OR < 1 was considered a negative association. The statistical analyses were performed using the SPSS 27 software (StataCorp Lp, College Station, TX, USA).
Ethics approval
This study was reviewed and approved by the Institutional Animal Care and Use Committee of Sichuan Agricultural University under permit number DYY-2020103018. Prior to the collection of fecal specimens from captive black bear, permission was obtained from the farm of black bear breeding, Dujiang Yan city, China.
Results
Antimicrobial susceptibility of 142 E. coli isolates
As shown in Table S2, a total of 142 E. coli isolates were obtained from the feces of 142 black bears in Sichuan, China. All E. coli strains showed resistance to the 13 antibiotics arranging from 0.0% (TOB) to 79.6% (TE) (Fig. 1). Of the142 strains, 139 strains (97.9%) were resistant to at least one of the 13 antibiotics tested. The most common resistances were to TE (79.6%, 113/142), followed by AMP (50.0%, 71/142) and SXT (43.7%, 62/142). Out of 142 strains, 62 strains (43.7%, 62/142) were found to be multidrug resistant (MDR), and 40 resistance patterns were observed in MDR strains (Fig. 2). Two strains (DJY14 and DJY43) were resistant to five classes of antibiotics, including: chloramphenicol, quinolone, sulfonamide, tetracycline and β-lactam. A total of 70 antibiotic resistance patterns were observed, with the three most prevalent patterns were TE (17.6%, 25 isolates), TE/SXT (7.0%, 10 isolates) and TE/C/AMP/SXT (4.2%, 6 isolates). In particular, the DJY127 strain was resistant to nine antibiotics (CN/TE/KZ/CXM/CTX/FEP/ATM/AMP/SXT).
The resistant phenotype patterns of E. coli isolates from black bears. (A) The 0 on the x-axis means the strains were sensitive to all antibiotics tested and 1–5 means the strains resistant to 1–5 categories of antibiotics, respectively. The bars indicating that 62 E. coli isolates (62/142, 43.7%) are MDR. (B) Color bars demonstrating the distribution of resistant phenotype patterns in E. coli isolates from black bears and only MDR isolates were analyzed (n = 62). By using disk diffusion assay, a total of 40 resistant patterns were observed in MDR E. coli strains.
Identification and characterization of ARGs from 142 E. coli isolates
As shown in Table S2, 5 ARGs were detected in 142 E. coli: tetA (70.8%, 109/142), qnrS (35.2%, 50/142), flor (23.9%, 34/142), sul1 (25.4%, 36/142) and blaCTX-M (12.7%, 18/142). The aminoglycosides resistant gene aphA3 was not detected in 142 E. coli. A total of 125/142 (88.0%) strains carried ARGs, with tetA being the most common pattern (36/142, 25.4%). Among the 142 E. coli isolates, 39 isolates (27.5%, 39/142) carried more than three types of ARGs. We further analyzed the concordance rate of ARGs and AMR detected in 142 E. coli strains. The results showed that 18.0% to 88.9% of ARGs corresponded to antibiotics resistance phenotypes (Table 1).
Mobile genetic elements prevalence in 142 E. coli
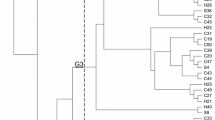
Ten out of the 13 MGEs were detected in the 142 isolates: IS26 (88.0%, 125/ 142), intI1 (52.8%, 75/142), trbC (22.5%, 32/142), tnpA/Tn21 (18.3%, 26/142), ISECP1 (9.2%, 13/142), tnsA (7.0%, 10/142), intI2 (6.3%, 9/142), merA (2.8%, 4/142), Isaba1 (2.1%, 4/142) and IS903 (1.4%, 2/142).However, IntI3, ISCR3/14 and IS1133 were not detected in any of the 142 E. coli. There were 23 patterns of combinations among 13 MGEs (Fig. 3), in which the IS26 was predominant (28.1%, 40/142), followed by IS26 + intI1 (21.8%, 31/142) and IS26 + trbC + intI1 (7.7%, 11/142). No mobile genetic element was detected in eleven isolates (DJY18, DJY36, DJY41, DJY44, DJY47, DJY56, DJY70, DJY80, DJY102, DJY103 and DJY119).
Characterization of integrons and gene cassettes in 142 E. coli isolates
As shown in Fig. 3, 52.8% (75/142) and 6.3% (9/142) of isolates were found to be positive for intI1 and intI2, respectively. intI1 and intI2 were both detected in six strains (DJY127, DJY130, DJY136, DJY140, DJY141 and DJY142). None of the 142 isolates tested positive for intI3 , and 45.1% (64/142) of isolates were negative for intI1 or intI2. Gene cassettes were also analyzed in intI1/2 positive isolates. As shown in Table 2, among 75 intI1-positive isolates, we detected 5 different gene cassette arrays:dfrA1-aadA1(1 isolate), dfrA1(2 isolates), dfrA17-aadA5,(4 isolates), aadA2(5 isolates), aadA2-dfrA12 (11 isolates) in 23 E. coli isolates. Among the 9 intI2-positive isolates, 6 E. coli isolates had 2 gene cassette arrays, dfrA1-catB2-sat1-aadA1(1 isolate), dfrA1-catB2-sat2-aadA1(5 isolates) arrays in 6 E. coli isolates.
Association among AMR phenotypes and MGEs
For the associations between AMR phenotypes and MGE, 14 pairs were positively associated (OR > 1) and 1 pair was negatively associated (ATM/IS26, OR, 0.308) (OR < 1) (Table 3). The strongest positive associations were observed between CN and tnpA (OR, 15.000; 95% CI 1.493–150.668), followed by CXM/ISECP1 (OR, 13.143; 95% CI 3.775–45.759) and FEP/ISECP1 (OR, 12.600; 95% CI 2.245–70.717). It is worth noting that ISECP1 was found be positively associated with five β-lactam antibiotics (CXM, CTX, FEP, ATM and AMP). No associations were observed for intI3, ISCR3/14 and IS1133 with 13 antibiotics.
Discussion
To enhance comprehension of the AMR of E. coli isolates from captive black bears, we assessed 142 E. coli strains from 142 captive black bears. Of these strains, 43.7% exhibited multidrug resistance (MDR). The MDR rate of E. coli in black bears is similar to that of sloth bear (51.1%) and giant panda (43.4%)24,25, but lower than that of domestic animals and poultry26. Antibiotics have been widely applied to promote growth and prevent diseases in China, which may lead to high MDR strains detected in domestic animals and captive wild animals27,28,29. Our study found a high rate of antibiotic resistance in E. coli isolates (97.9%) from black bears, which is higher than that observed in sloth bears in India (93.3%)25.The consumption and production of antibiotics are higher in China30, and the frequent exposure or misuse of antimicrobials in animals may contribute to the emergence and spread of resistance in E. coli from black bears. In our study, E. coli from black bear exhibited varying degrees of resistance to GM, CIP, CTX and CXM. GM, CIP, CTX and CXM have been used for disease control in black bears, which may contribute to antimicrobial resistance. Additionally, the resistance rate to TE (79.6%) in E. coli is the highest rate among 13 antibiotics31. The high resistance rate to TE (79.6%) has also been reported in sloth bears (51.1%)25. TE has been used to treat animal infections in the world32, however, the development of resistance has narrowed their utility and the use of TE is strictly regulated in China33. Our previous study was also found this phenomenon in giant panda, indicating that AMR persists for a longer time and is not easy to eliminate34. In general, ARGs were the primary cause of antibiotic resistance phenotypes, AGRs were expressed and translated into proteins, allowing bacteria to achieve resistance to antibiotics. Different ARGs mediate bacterial resistance to antibiotics in different ways35. In our study, 6 ARGs were detected and the most prevalent ARG was tetA (70.8%, 109/142). The high prevalence of tetA has also been observed in various wild animals (e.g., giant panda, monkey) and human studies23,24,36, suggesting that TE resistance will continue to exist for a period of time, and we should continue to monitor TE resistance. Moreover, we also found that some isolates (DJY62 and DJY129) showed antimicrobial susceptibility despite harboring ARGs. This phenomenon may be related to abnormal expression of ARGs, or the expression of ARGs may not have reached the level required to produce resistance37.
Previous studies have shown that MGEs play an important role in the dissemination of ARGs38,39. Our result showed that 10 MGEs and 23 MGEs combination patterns were detected in 142 E. coli strains from black bears. In comparison to our previous research, we found fewer MGEs (10) and MGEs combinations (23) in black bears than in giant pandas (11 MGEs, 35 MGEs combinations)24. We detected 10 blaCTX-M positive strains out of 13 strains carrying ISECP1 (Table S2). Studies have shown that ISECP1 and blaCTX-M are all located in the same plasmid, which may be the reason for their association with five β-lactams antibiotics40. Horizontal transfer of ARGs mediated by MGEs is one of the main mechanisms41,42, and it is necessary to regularly monitor MGEs in E. coli from black bears. To our knowledge, this study is the first to examine MGEs of E. coli isolates from captive black bears.
Integrons, as one of the MGEs, are known to capture gene cassettes, which could be transferable among bacteria and disseminate ARGs via transmissible plasmids and insertion sequences, posing a threat to public health16,43. In our results, class 1 integron (56.8%) was more prevalent compared to class 2 integron (6.3%), while class 3 integron was not detected, which is consistent with results from yaks, ducks and giant pandas16,24,44. Research has shown that due to the influence of human activities, the prevalence of Class I integron in natural environments is higher than that of Class II and III45. Gene cassettes harbored in integron-positive isolates are an important medium for the spread of ARGs46. In this study, seven types of gene cassettes were detected in class 1 or class 2 integron-positive isolates (Table 2). The gene cassette encoding the aminoglycoside adenyl transferase (aadA2) and dihydrofolate reductase (dfrA12) 34 is the most frequently detected among the isolates, with 11 instances34. Similar gene cassettes belonging to dfrA, sat, catB and aadA have also been detected in E. coli isolates from other animals, including giant pandas and rabbits24,47, indicating that the gene cassettes have spread between different species.
The above results indicate that different types of integrons and gene cassettes are carried by E. coli from captive black bears, posing a threat of spreading ARGs to other animals or environments. The high prevalence of AMR E. coli detected in black bears pose a potential risk to public health. Therefore, relevant people like veterinarians should pay attention during work. The origin of the AMR isolates analyzed in our present study is unclear, despite the high prevalence of AMR E. coli detected within the farm-raised black bear population.
Conclusion
This study found a high prevalence of antibiotic resistance and a diverse range of MGEs observed among the E. coli strains isolated from captive black bears. A positive association was observed between MGEs and AMR in E. coli strains. It is essential to monitor the distribution of ARGs, MGEs and gene cassettes, which may help to introduce interventions for the prevention and control of AMR in E. coli among captive black bears.
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Jang, J. et al. Environmental Escherichia coli: Ecology and public health implications-a review. J. Appl. Microbiol. 123, 570–581. https://doi.org/10.1111/jam.13468 (2017).
Tunyong, W. et al. Antibiotic resistance genes among carbapenem-resistant enterobacterales (CRE) isolates of Prapokklao Hospital, Chanthaburi Province, Thailand. Infect. Drug Resistance 14, 3485–3494. https://doi.org/10.2147/idr.S328521 (2021).
Meyers, B. C. & McLellan, S. L. Influence of nutrients and the native community on E. coli survival in the beach environment. Appl. Environ. Microbiol. 88, e0104322. https://doi.org/10.1128/aem.01043-22 (2022).
Shintani, M. The behavior of mobile genetic elements (MGEs) in different environments. Biosci. Biotechnol. Biochem. 81, 854–862. https://doi.org/10.1080/09168451.2016.1270743 (2017).
Baltazar, M. et al. Activation of class 1 integron integrase is promoted in the intestinal environment. PLoS Genet. 18, e1010177. https://doi.org/10.1371/journal.pgen.1010177 (2022).
Antelo, V. et al. Metagenomic strategies identify diverse integron-integrase and antibiotic resistance genes in the Antarctic environment. MicrobiologyOpen 10, e1219. https://doi.org/10.1002/mbo3.1219 (2021).
Zahoor, B. et al. Activity pattern study of Asiatic black bear (Ursus thibetanus) in the Qinling Mountains, China, by using infrared camera traps. Environ Sci Pollut Res Int 28, 25179–25186. https://doi.org/10.1007/s11356-020-12325-3 (2021).
Malcolm, K. D. et al. Analyses of fecal and hair glucocorticoids to evaluate short- and long-term stress and recovery of Asiatic black bears (Ursus thibetanus) removed from bile farms in China. Gen. Compar. Endocrinol. 185, 97–106. https://doi.org/10.1016/j.ygcen.2013.01.014 (2013).
Pontes, P. S. et al. Survey on pathogenic Escherichia coli and Salmonella spp. in captive cockatiels (Nymphicus hollandicus). Braz. J. Microbiol.: [Publication of the Brazilian Society for Microbiology] 49(Suppl 1), 76–82. https://doi.org/10.1016/j.bjm.2018.05.003 (2018).
Hu, T. et al. Geographic pattern of antibiotic resistance genes in the metagenomes of the giant panda. Microbial Biotechnol. 14, 186–197. https://doi.org/10.1111/1751-7915.13655 (2021).
Donkor, E. S., Newman, M. J. & Yeboah-Manu, D. Epidemiological aspects of non-human antibiotic usage and resistance: implications for the control of antibiotic resistance in Ghana. Trop. Med. Int. Health: TM & IH 17, 462–468. https://doi.org/10.1111/j.1365-3156.2012.02955.x (2012).
VinodhKumar, O. R. et al. Multi-drug resistant (MDR), extended spectrum beta-lactamase (ESBL) producing and carbapenem resistant Escherichia coli in rescued Sloth bears (Melursus ursinus), India. Vet. Res. Commun. 45, 163–170. https://doi.org/10.1007/s11259-021-09794-3 (2021).
Anjum, M. F., Schmitt, H., Börjesson, S. & Berendonk, T. U. The potential of using E. coli as an indicator for the surveillance of antimicrobial resistance (AMR) in the environment. Curr. Opin. Microbiol. 64, 152–158. https://doi.org/10.1016/j.mib.2021.09.011 (2021).
Smoglica, C. et al. Antibiotic-resistant bacteria dissemination in the wildlife, livestock, and water of Maiella National Park, Italy. Anim: Open Access J. MDPI https://doi.org/10.3390/ani13030432 (2023).
Homeier-Bachmann, T. et al. Genomic analysis of ESBL-producing E. coli in wildlife from North-Eastern Germany. Antibiotics (Basel, Switzerland) https://doi.org/10.3390/antibiotics11020123 (2022).
Zhang, S. et al. Class 1 integrons as predominant carriers in Escherichia coli isolates from waterfowls in Hainan, China. Ecotoxicol Environ Saf 183, 109514. https://doi.org/10.1016/j.ecoenv.2019.109514 (2019).
Nowaczek, A. et al. Antibiotic resistance and virulence profiles of Escherichia coli strains isolated from wild birds in Poland. Pathogens (Basel, Switzerland) https://doi.org/10.3390/pathogens10081059 (2021).
Asai, T., Usui, M., Sugiyama, M. & Andoh, M. A survey of antimicrobial-resistant Escherichia coli prevalence in wild mammals in Japan using antimicrobial-containing media. J Vet Med Sci 84, 1645–1652. https://doi.org/10.1292/jvms.22-0415 (2022).
Zhu, Z. et al. High prevalence of multi-drug resistances and diversity of mobile genetic elements in Escherichia coli isolates from captive giant pandas. Ecotoxicol. Environ. Saf. https://doi.org/10.1016/j.ecoenv.2020.110681 (2020).
Zhang, S. et al. High incidence of multi-drug resistance and heterogeneity of mobile genetic elements in Escherichia coli isolates from diseased ducks in Sichuan province of China. Ecotoxicol. Environ. Saf. https://doi.org/10.1016/j.ecoenv.2021.112475 (2021).
Weiss, D. et al. Antibiotic-resistant Escherichia coli and Class 1 integrons in humans, domestic animals, and wild primates in rural Uganda. Appl. Environ. Microbiol. 84, 55. https://doi.org/10.1128/aem.01632-18 (2018).
Velhner, M. et al. Fluoroquinolone-resistant and extended-spectrum beta-lactamase producing Escherichia coli isolates from free-living wild animals. Vet. Microbiol. 223, 168–172. https://doi.org/10.1016/j.vetmic.2018.08.011 (2018).
Zhu, Z. et al. Prevalence and characterization of antibiotic resistance genes and integrons in Escherichia coli isolates from captive non-human primates of 13 zoos in China. Sci. Total Environ. 798, 149268. https://doi.org/10.1016/j.scitotenv.2021.149268 (2021).
Zhu, Z. et al. High prevalence of multi-drug resistances and diversity of mobile genetic elements in Escherichia coli isolates from captive giant pandas. Ecotoxicol Environ Saf 198, 110681. https://doi.org/10.1016/j.ecoenv.2020.110681 (2020).
VinodhKumar, O. R. et al. Multi-drug resistant (MDR), extended spectrum beta-lactamase (ESBL) producing and carbapenem resistant Escherichia coli in rescued Sloth bears (Melursus ursinus), India. Vet Res Commun 45, 163–170. https://doi.org/10.1007/s11259-021-09794-3 (2021).
Ganta, R. R. et al. Antimicrobial resistance in clinical Escherichia coli isolates from poultry and livestock, China. Plos One https://doi.org/10.1371/journal.pone.0185326 (2017).
Liu, Z. et al. High prevalence and diversity characteristics of blaNDM, mcr, and blaESBLs harboring multidrug-resistant Escherichia coli from chicken, pig, and cattle in China. Front. Cell. Infect. Microbiol. https://doi.org/10.3389/fcimb.2021.755545 (2022).
Algammal, A. M. et al. Migratory wild birds carrying multidrug-resistant Escherichia coli as potential transmitters of antimicrobial resistance in China. Plos One https://doi.org/10.1371/journal.pone.0261444 (2021).
Yuan, J. et al. Extensive antimicrobial resistance and plasmid-carrying resistance genes in mcr-1-positive E. coli sampled in swine, in Guangxi, South China. BMC Vet. Res. 17, 55. https://doi.org/10.1186/s12917-021-02758-4 (2021).
Liu, X., Lu, S., Guo, W., Xi, B. & Wang, W. Antibiotics in the aquatic environments: A review of lakes, China. Sci. Total Environ. 627, 1195–1208. https://doi.org/10.1016/j.scitotenv.2018.01.271 (2018).
Roth, N. et al. The application of antibiotics in broiler production and the resulting antibiotic resistance in Escherichia coli: A global overview. Poultry Sci. 98, 1791–1804. https://doi.org/10.3382/ps/pey539 (2019).
Chopra, I. & Roberts, M. Tetracycline antibiotics: Mode of action, applications, molecular biology, and epidemiology of bacterial resistance. Microbiol. Mol. Biol. Rev.: MMBR 65, 232–260. https://doi.org/10.1128/mmbr.65.2.232-260.2001 (2001).
Roberts, M. C. & Schwarz, S. Tetracycline and phenicol resistance genes and mechanisms: Importance for agriculture, the environment, and humans. J. Environ. Quality 45, 576–592. https://doi.org/10.2134/jeq2015.04.0207 (2016).
Zhang, S. et al. High incidence of multi-drug resistance and heterogeneity of mobile genetic elements in Escherichia coli isolates from diseased ducks in Sichuan province of China. Ecotoxicol Environ Saf 222, 112475. https://doi.org/10.1016/j.ecoenv.2021.112475 (2021).
García, J. et al. A review of emerging organic contaminants (EOCs), antibiotic resistant bacteria (ARB), and antibiotic resistance genes (ARGs) in the environment: Increasing removal with wetlands and reducing environmental impacts. Bioresour. Technol. 307, 123228. https://doi.org/10.1016/j.biortech.2020.123228 (2020).
Bag, M. A. S. et al. Virulence determinants and antimicrobial resistance of E. coli isolated from bovine clinical mastitis in some selected dairy farms of Bangladesh. Saudi J. Biol. Sci. 28, 6317–6323. https://doi.org/10.1016/j.sjbs.2021.06.099 (2021).
Zhang, S. et al. Distribution and association of antimicrobial resistance and virulence traits in Escherichia coli isolates from healthy waterfowls in Hainan, China. Ecotoxicol. Environ. Saf. https://doi.org/10.1016/j.ecoenv.2021.112317 (2021).
Redhead, S. et al. Fate of antibiotic resistant E. coli and antibiotic resistance genes during full scale conventional and advanced anaerobic digestion of sewage sludge. PLoS One 15, e0237283. https://doi.org/10.1371/journal.pone.0237283 (2020).
Xiao, E. et al. Occurrence and dissemination of antibiotic resistance genes in mine soil ecosystems. Appl. Microbiol. Biotechnol. 106, 6289–6299. https://doi.org/10.1007/s00253-022-12129-0 (2022).
Zhao, Q. Y. et al. Transmission of plasmid-borne and chromosomal blaCTX-M-64 among Escherichia coli and Salmonella isolates from food-producing animals via ISEcp1-mediated transposition. J. Antimicrobial Chemother. 75, 1424–1427. https://doi.org/10.1093/jac/dkaa044 (2020).
Sultan, I., Siddiqui, M. T., Gogry, F. A. & Haq, Q. M. R. Molecular characterization of resistance determinants and mobile genetic elements of ESBL producing multidrug-resistant bacteria from freshwater lakes in Kashmir, India. Sci Total Environ 827, 154221. https://doi.org/10.1016/j.scitotenv.2022.154221 (2022).
Tansirichaiya, S. et al. Intracellular transposition and capture of mobile genetic elements following intercellular conjugation of multidrug resistance conjugative plasmids from clinical enterobacteriaceae isolates. Microbiol. Spectrum 10, e0214021. https://doi.org/10.1128/spectrum.02140-21 (2022).
Rehman, M. U. et al. Characteristics of integrons and associated gene cassettes in antibiotic-resistant Escherichia coli isolated from free-ranging food animals in China. J Food Sci 82, 1902–1907. https://doi.org/10.1111/1750-3841.13795 (2017).
Yang, X. et al. Prevalence of antimicrobial resistance and integron gene cassettes in Escherichia coli isolated from yaks (Poephagus grunniens) in Aba Tibetan Autonomous Prefecture, China. Microb Pathog 111, 274–279. https://doi.org/10.1016/j.micpath.2017.09.008 (2017).
Koczura, R., Mokracka, J., Taraszewska, A. & Łopacinska, N. Abundance of class 1 integron-integrase and sulfonamide resistance genes in river water and sediment is affected by anthropogenic pressure and environmental factors. Microbial Ecol. 72, 909–916. https://doi.org/10.1007/s00248-016-0843-4 (2016).
Algarni, S., Ricke, S. C., Foley, S. L. & Han, J. The dynamics of the antimicrobial resistance mobilome of Salmonella enterica and related enteric bacteria. Front. Microbiol. 13, 859854. https://doi.org/10.3389/fmicb.2022.859854 (2022).
Dotto, G. et al. High prevalence of oqxAB in Escherichia coli isolates from domestic and wild lagomorphs in Italy. Microbial Drug Resist. (Larchmont, NY) 20, 118–123. https://doi.org/10.1089/mdr.2013.0141 (2014).
Acknowledgements
This work was funded by the National Key Research and Development Program of China (2018YFD0500900, 2016YFD0501009), the Chengdu Giant Panda Breeding Research Foundation (CPF2017-05, CPF2015-4) and the Science and Technology Achievements Transfer Project in Sichuan Province (2022JDZH0026). We thank Dr Junai Gan for the English language revision.
Author information
Authors and Affiliations
Contributions
Z.Z., K.S. and X.Z.: Conceptualization, Methodology, Software. H.L., Y.W. and Z.Z.: Data curation, Writing-Original draft preparation. H.L., Y.C. and S.P.: Visualization, Investigation. H.L., W.Z., Y.Y. and L.Z.: Supervision. Z.Z., G.P. and Q.Y.: Software, Validation. Y.L., S.Z. and Z.Z.: Writing-Reviewing and Editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, H., Shi, K., Wang, Y. et al. Characterization of antibiotic resistance genes and mobile genetic elements in Escherichia coli isolated from captive black bears. Sci Rep 14, 2745 (2024). https://doi.org/10.1038/s41598-024-52622-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-52622-2
- Springer Nature Limited