Abstract
Abiotic stresses limit the quantity and quality of rice grain production, which is considered a strategic crop in many countries. In this study, a meta-analysis of different microarray data at seedling stage was performed to investigate the effects of multiple abiotic stresses (drought, salinity, cold situation, high temperature, alkali condition, iron, aluminum, and heavy metal toxicity, nitrogen, phosphorus, and potassium deficiency) on rice. Comparative analysis between multiple abiotic stress groups and their control groups indicated 561 differentially expressed genes (DEGs), among which 422 and 139 genes were up-regulated and down-regulated, respectively. Gene Ontology analysis showed that the process of responding to stresses and stimuli was significantly enriched. In addition, pathways such as metabolic process and biosynthesis of secondary metabolites were identified by KEGG pathway analysis. Weighted correlation network analysis (WGCNA) uncovered 17 distinct co-expression modules. Six modules were significantly associated with genes involved in response to abiotic stresses. Finally, to validate the results of the meta-analysis, five genes, including TIFY9 (JAZ5), RAB16B, ADF3, Os01g0124650, and Os05g0142900 selected for qRT-PCR analysis. Expression patterns of selected genes confirmed the results of the meta-analysis. The outcome of this study could help introduce candidate genes that may be beneficial for use in genetic engineering programs to produce more tolerant crops or as markers for selection.
Similar content being viewed by others
Introduction
Rice (Oryza sativa) is one of the world's most important cereals and a staple food for half of the world's population1. Since the global population is rapidly growing and is predicted to reach 9.9 billion by 20502, the production of rice must increase at least 1% annually to meet its demand3. On the other hand, environmental stresses have been primarily considered as the factors that reduce the quantity and quality of agricultural products. Abiotic stresses including drought, salinity, low and high temperatures, deficiency of essential nutrients, and accumulation of heavy metals can negatively affect the growth, development, and yield of the crops. Abiotic stresses annually reduce rice production by 32% (about three million tons) in the world4. Therefore, to improve sustainability, there is a need to increase the yield in breeding programs by introducing stress-tolerant varieties.
During the evolution of plants, different mechanisms including various physiological, cellular, and molecular modifications have been developed to cope with different stresses and to survive in adverse conditions5. Abiotic stresses significantly affect physiological processes such as flowering, grain filling, and maturation. It has been reported that abiotic stresses affect plant metabolisms including photosynthesis, enzyme activity, mineral nutrition intake, and respiration6. Plants respond to abiotic stresses by inducing a complex network of genes. They activate stress-related genes to adapt to new environmental conditions through the perception and transduction of stress signals7. Sensing, signaling, transcription, transcript processing, translation, and post-translational protein modifications are plant molecular mechanisms to respond to abiotic stresses8. The thickness of the cell wall, production of reactive oxygen species (ROS), and secretion of phytohormones including abscisic acid (ABA), jasmonic acid (JA), salicylic acid (SA), and ethylene (ET) are other defense responses of plant to abiotic stresses9.
To better understand the complex system of molecular processes and identify pathways and mechanisms involved in cell response to abiotic stresses, it is inevitable to use statistical and computational approaches. High-throughput technologies such as microarray and RNAseq, which are being used for gene expression analyses, have made it feasible to study a large number of genes simultaneously in different conditions. With the development of new technologies, a big step has been taken to decipher the gene regulatory networks in plants' stress-tolerance mechanisms.
Meta-analysis is a standard statistical procedure for combining datasets from multiple studies to systematically assess previously published data to derive more comprehensive conclusions about that research field10. This technique provides a broad perspective on specific biological questions and more reliable results than individual studies11,12.
Different studies have identified several responsive genes under abiotic stresses in rice through omics data analysis such as genomics and transcriptomics13,14,15,16,17. In a meta-analysis, de Abreu Neto et al. investigated the genes involved in redox homeostasis in rice under abiotic stresses and showed that only 4% of differentially expressed genes (DEGs) were in the ROS mechanism pathway directly13. ROS plays an important signaling role in plants, especially in response to biotic and abiotic stimuli14. Another meta-analysis reported by Cohen et al. investigated different abiotic stresses including drought, salinity, high and low temperature as well as biotic stresses such as Dwarf, Stripe, ZB13, and Guy11 viruses, and Xoc and Xoo bacteria in rice15. They showed that the number of DEGs varied from 1220 to 11,644 in different experiments, in which 5863 and 2154 genes were common in all abiotic and biotic stresses, respectively. Buti et al. investigated different responses of susceptible and tolerant genotypes of rice under osmotic, chilling, and salt stresses. They found 35 hub genes through gene network analysis, which 24 of them were located in at least one known QTL of rice such as qLRC-1, qGY-2b, qTGW-2a, rfw1b, rfw4a, qtl3.1, gpl11.1, gw11.1, yld11.1, rn3 and qSDW2 which are related to cold, drought, and salinity16. Recently, Ramkumar et al. identified 6657 multiple abiotic stress-responsive genes (salinity, drought, and heat stresses) in rice at the seedling stage. They found 10 modules containing 10 genes through gene network analysis that were common to all three studied stresses17.
In the present study, a large-scale meta-analysis was performed to integrate different microarray studies focused on abiotic stresses including drought, salinity, cold and high temperatures, alkali conditions, nutrients (nitrogen, phosphorus, and potassium) deficiency, toxicity of heavy metal, iron, and aluminum to find genes involved in different stresses. We hypothesized that there may be some common genes and pathways that are activated and expressed to alleviate stress and regulate plant metabolisms under different stress conditions. Therefore, it may lead to the introduction of genes and pathways in breeding programs to have resistant cultivars.
Materials and methods
Data collection and meta-analysis
Microarray data sets of studied abiotic stresses including drought, salinity, cold and high temperatures, alkalinity, deficiency of essential nutrients such as nitrogen, phosphorus and potassium as well as the toxicity of heavy metals, aluminum, and iron were extracted from Gene Expression Omnibus (http://www.ncbi.nlm.nih.gov/geo) (Table 1). Two technical considerations were applied to select microarray datasets: (1) The selected dataset must be in one of the two subspecies O. sativa japonica or O. sativa indica; (2) The RNA must have been extracted from the vegetative parts of plants including shoots, roots, and or whole seedlings (reproductive tissues such as seeds, panicles or flowers were excluded from the study).
The method used for meta-analysis of microarray data is based on Raw Data Integration. It integrates raw microarray data from multiple studies through the following steps:
-
1.
Data preprocessing: Extract raw microarray data (e.g., CEL files) from individual studies (raw microarray data from individual studies retrieved from NCBI).
-
2.
Quality control: Perform quality control checks for each dataset to identify and remove low-quality samples or datasets. The quality of each dataset was controlled by checking the boxplot of datasets.
-
3.
Normalization: The normality of each dataset was checked.
-
4.
Data integration: Merge normalized data into a unified dataset. All data series matrices are merged into one dataset. Treated samples (regardless of the type of stress) are categorized into the "stress" group, and all untreated samples are grouped into the "control" group.
-
5.
Batch effect correction: Address batch effects using the ComBat technique to ensure comparability between stress and control samples. The batch effect, as one of the major technical variations that make differences between different datasets, was removed by the SVA R package (v 3.38.0)18 according to the COMBAT method19.
-
6.
Statistical analysis: Conduct statistical analyses (e.g., Differential expression analysis) on the integrated dataset to identify genes that are differentially expressed across conditions or groups (stress samples vs control samples)20.
Meta-analysis of transcriptome data sets was carried out by merging expression data matrix in R software. The DEGs between stress and control samples were identified using the Limma R package (v 3.48.1)21. Genes with an adjusted P value (p-adj) < 0.05 and |log2 fold change| (|log2FC|) ≥ 0.5 were considered as DEGs. Probe IDs of DEGs were used as queries in the DAVID web-based tool (http://david.abcc.ncifcrf.gov/) to annotate them. A schematic workflow summarizing the main stages of the current study is presented in Fig. S1.
Enrichment analysis
The DEGs were subjected to singular enrichment analysis (SEA) in agriGO (http://bioinfo.cau.edu.cn/agriGO/) to identify enriched gene ontologies (GO). DEGs were classified into three biological process (BP), molecular function (MF), and cellular component (CC) with a significant threshold of p-value < 0.05. The Kyoto Encyclopedia of Genes and Genomes (KEGG) (https://www.genome.jp/kegg/) analysis was performed to identify enriched pathways, in which DEGs are significantly involved.
Protein–protein interaction network
To identify key genes responsible for abiotic stresses, hub gene determination analysis was performed for all DEGs. Protein–protein interaction (PPI) network was constructed using the STRING database (https://string-db.org/) by submitting DEGs as input. The output file was imported to Cytoscape (version 3.8.2) software for visualization and edition22. The CytoHubba plugin and Maximal Clique Centrality (MCC) algorithm were used to identify highly connected genes as hubs23.
Weighted correlation network analysis (WGCNA)
The WGCNA R package was used to identify the group of genes with similar expression patterns under stress situations24. The WGCNA is performed as a system biology approach for analyzing the correlation pattern between genes and spreading them into co-expression modules25. The co‐expression analysis was performed for paired genes using a Pearson correlation matrix. The weighted adjacency matrix was constructed using the power function (β), and then, transformed into a topological overlap measure (TOM) matrix to assess its connectivity in the network26. The clustering dendrogram of the TOM matrix was constructed using the average linkage hierarchical clustering. To obtain the correct module number, a restricted minimum gene number of 30 for each module was set and a threshold of 0.25 to merge similar modules was used. The network was visualized for the two most important modules using the Cytoscape software (version 3.8.2). Each module can lead to a real biological process, so to examine the significance of grouping, gene ontology analysis for each module was performed using the DAVID web-based tool.
Plant materials and experimental design
The rice seeds (Shiroodi variety; japonica subgroup) were obtained from the Iranian Rice Research Institute (Amol, Mazandaran, Iran). Plant studies comply with relevant institutional, national, and international guidelines and legislation. The surface of the seeds was sterilized using 70% ethanol for 2 min, followed by treatment with 1.5% NaClO4 for 1 min. The seeds were washed three times with distilled water to remove the detergents27. Seeds were placed on the moistened filter paper in the Petri dishes and incubated for the first 72 h in the dark to germinate. Then, they were transferred to pots filled with pearlite and kept in a 16 h light and 8 h dark photoperiod at 25 ± 2 °C and were irrigated every day. The Yoshida solution was used28 after the emergence of seedlings. Samples in the control treatment were kept under mentioned condition (16 h light and 8 h dark photoperiod at 25 ± 2 °C), and the Yoshida solution was renewed every 3 days. Seedlings were exposed to different stress treatments according to Table 2. The root and/or shoot samples of different stress treatments with their respective control treatments were collected, immediately immersed in the liquid nitrogen, and kept at − 80 °C until further analysis.
RNA extraction and cDNA synthesis
Total RNA was extracted using the DENAzist Column RNA Isolation Kit (DENAzist Inc., Mashhad, Iran). The quantity and quality of RNA samples were evaluated by a NanoDrop spectrophotometer and 1% agarose gel electrophoresis, respectively. The first-strand cDNA was synthesized using the EasyTM cDNA synthesis kit Pars Tous according to the manufacturer's instructions (Pars Tous Inc., Mashhad, Iran).
Validation of abiotic responses of candidate genes by real-time PCR
To validate the reliability of the meta-analysis approach, five genes were randomly selected for real-time PCR. Specific primers were designed by OLIGO Primer Analysis Software v.7.0 (National Bioscience Inc., Plymouth, USA). The optimize the amplification, 10 μl of Taq DNA Polymerase Master Mix RED (Ampliqon, Denmark), 1 μl of cDNA (~ 20 ng), 0.01 µM of each forward and reverse primer, and sterile distilled water (up to 20 μl) were used and PCR products were evaluated on 1% agarose gel for the presence of the desired band and the absence of non-specific amplicons and primers dimer. The optimized PCR program for each gene and oligonucleotide primers are presented in Table 3. Real-time PCR with three technical and five biological replicates was done by the Rotor-Gene Q (QIAGEN, Germany) and SYBR® Green Fluorescent DNA Stain-low ROX (Jena Bioscience, Germany) according to the optimized program for each candidate gene. The elongation factor 1-alpha (elF1α) gene of rice (LOC4331813) was selected as the reference gene. The gene expression was calculated using the Delta-Delta CT method40 in the REST2009 software according to the comparative threshold cycle, and the graphs were made using the GraphPad Prism9(GraphPad Software, United States).
Statistical analysis
Statistical analysis for all molecular data was performed using R version 3.5.321 and RStudio version 1.1.463. Data were analyzed by ANOVA for a completely randomized design with treatments as fixed effects and replicate as random effects. Mean values were compared using the Duncan test function provided in the agricolae package at 5% significance level of probability.
Results
Meta-analysis to identify DEGs of rice in response to multiple abiotic stresses
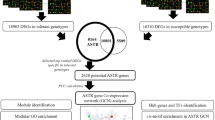
The meta-analysis included 11 studies containing 118 samples. Our result showed 561 DEGs among stressed and control samples, in which 421 and 139 were up- and downregulated, respectively (Table S1). The PCA plot illustrates that the presence of a batch effect leads to the segregation of each dataset based on their respective batches, as indicated by the distinct colors shown in Fig. 1a. However, following the implementation of the ComBat method, the impact of the batch effect is reduced, resulting in the mixing of datasets, regardless of their original batches, as demonstrated in Fig. 1b.
Gene ontology and KEGG analysis
The GO in the biological process associated with DEGs was grouped in 37 terms (p-value < 0.05) (Table S2). The top enriched biological processes were response to stress and stimulus (GO:0006950 GO:0050896), metabolic and catabolic processes of cell wall macromolecules (GO:0044036, GO:0016998), lipid transport (GO:0006869), and defense response (GO:0006952). GO enrichment analyses for functional annotation revealed that oxidoreductase (GO:0016491), hydrolase (GO:0004553, GO:0016798), and enzyme inhibitor (GO:0004857) activities were the top enriched molecular function. In this category, DEGs were grouped in 24 GO (p-value < 0.05) (Table S2) among them there are also regulation activities such as transcription factors (GO:0003700), transcription regulators (GO:0030528), DNA binding (GO:0003677), and enzyme regulator (GO:0030234) activities. Among GO terms in the cellular component, vesicles (GO:0031982), cytoplasmic membrane-bounded vesicles (GO:0016023), membrane-bounded vesicles (GO:0031988), cytoplasmic vesicles (GO:0031410), and extracellular region (GO:0005576) were significantly enriched (Fig. 2).
The KEGG analysis showed that DEGs were mostly enriched in the metabolic pathway, biosynthesis of secondary metabolites, plant hormone signal transduction, phenylpropanoid biosynthesis, amino sugar, and nucleotide sugar metabolism, and MAPK signaling pathways (Fig. 3)41.
Identification of DEGs encoding TF and PK in response to abiotic stresses
A total of 25 Transcription factor (TF) genes related to 7 TF families were identified among all DEGs (Table 4). The WRKY family with 8 genes and the Ethylene Response Factor (ERF) family with 6 genes represented the highest number of TFs. Moreover, HSF, MYB, 6HLH and NF-YB factors had 3, 3, 1 and 1 genes, respectively. Protein kinases were encoded by 19 genes that were classified as the receptor-like kinase (RLK) (19 genes) and calcium/calmodulin-dependent protein kinase (CAMK) families (1 gene). As shown in Table 4, the RLK family included 5 subgroups including leucine-rich repeat (8 genes), DLSV (4 genes), and S Domain 2b (4 genes) with the highest number of genes, respectively.
Protein–protein interaction network
The network of hub proteins is shown in Fig. 4. In this network, a total of 31 hub proteins interacted, which the stress response proteins such as JAZ, LEA, NAC, RAB, and WORKY families had the highest interaction in response to abiotic stresses. The top 12 hub proteins are represented in Table S3. The highest interaction scores were related to proteins LEA14, HSFA6B, RAB16B, OsJ_021637, RAB16C, and OS03T0305600-01with scores of 164, 142, 133, 133, 130 and 129, respectively (Table S3). The complete images of the gels are available in the Fig. S5.
WGCN analysis
A total of 17 WGCNA modules were identified based on the dynamic tree-cutting algorithm (Fig. S2). The number of genes in each module varied from 57 to 483. The turquoise (483 genes), blue (316 genes), brown (248 genes), and yellow (244 genes) were four major modules. As shown in Fig. 5, when the soft threshold power is defined as 6, the scale-free topological index is 0.9. Therefore, the network is closer to the real biological network state as it adheres to the power-law distribution. Results showed that the six modules (turquoise, blue, yellow, pink, magenta, and tan) were directly involved in abiotic stress processes. Turquoise and blue were the largest modules in the gene networks (Fig. 6). The most important genes in stress response pathways were the JAZ, WPKY, NAC, APR1, and GLU families in the networks.
Validation of DEGs using qRT-PCR
The specificity of the designed primers (Jasmonate-ZIM domain-containing protein 5 (JAZ5) also known as TIFY9, RAB16B, actin depolymerizing factor 3 (ADF3), Os01g0124650, and Os05g0142900) was evaluated by agarose gel electrophoresis 1% (Fig. S3), which no non-specific band and primer dimer were observed. To validate the result, the expression of these genes was evaluated by qRT-PCR in all of the studied stresses using elF1α as the reference gene to normalize CT values.
The ANOVA analysis of different DEGs in Rice that grew under abiotic stresses indicated that the single effects of stress treatments were highly significant (p < 0.01) in TIFY9, ADF3, Os01g0124650, RAB16B, and Os05g0142900 (Table 5).
The TYFY9 was significantly upregulated in all stresses except potassium deficiency for 6 h (K-6h), in which a non-significant decrease was shown. The highest expression of TYFY9 was observed under Iron toxicity, followed by Nitrogen starvation, cold (Exposure of plants to a temperature of 4 °C for 48 h), alkali situation, and potassium deficiency for 5 days (K-5d). The TYFY9 was highly upregulated in K-5d (log2FC = 5) but downregulated in K-6h (log2FC = − 0.88).
The expression of ADF3 increased in all stresses but was not significant under drought, cold-48h, and N starvation conditions. The highest expression of ADF3 was observed under alkali situations (log2FC = 9.8), K-5d (log2FC = 7.72), phosphorus deficiency ((log2FC = 6.65), salinity condition (log2FC = 6.47) and aluminum toxicity (log2FC = 5.84). The Os01g0124650 was significantly upregulated in six stresses including drought and alkali situation, aluminum and heavy metal toxicity as well as cold conditions for 6 h (C-6h) and potassium deficiency for 5 days (K-5d). Although this gene was upregulated in K-6h, high temperature (T), and salinity conditions, it was not statistically significant. The RAB16B was significantly upregulated in all the stresses except for N starvation, in which gene expression was equal to − 1.25 (significant downregulation). The highest expression of RAB16B was shown in the C-48h, alkali situation, salinity stress, phosphorus deficiency, and high temperature (T). The Os05g0142900 had expression levels of 9.87, 9.82, 7.17, 6.66, and 6.58 under phosphorus deficiency, C-48h, nitrogen starvation, Fe, and aluminum toxicity stresses, respectively. Significant downregulation of this gene was observed under K-5d. In general, the qRT-PCR results highly confirmed the outcome of the meta-analysis (Fig. 7) although some treatments had contradictory results.
Validation of selected genes using qRT-PCR in different stresses. dr: drought stress/st: salinity stress/Alkali: alkali condition/Al: aluminum toxicity/C-6h: cold exposed for 6 h/C-48h: cold exposed for 48 h/N: nitrogen deficiency/ Fe: Fe toxicity(Iron toxicity)/P: phosphor deficiency/K-5d: Potassium deficiency for 5 days/K-6h: Potassium deficiency for 6 h/T: high temperature/H.M: excess heavy metal (K2Cr2O7). Expressions with not significant changes are represented by n.s, other expressions are significantly up (positive data) or down (negative data) regulated.
Discussion
A meta-analysis was conducted to identify genes responsible for a wide range of abiotic stresses including drought, salinity, cold and high temperatures, alkali conditions, iron, aluminum, heavy metal toxicity, and nitrogen, phosphorus, and potassium deficiency. Plants change their transcriptome profile to endure unfavorable environmental conditions in response to different stresses42,43. Despite having a specific response to each stress in plants, this study indicates that there are a vast number of genes that are similarly expressed in response to different abiotic stresses, indicating that there is a core response system to all environmental stresses in rice. Our meta-analysis investigated 11 different abiotic stresses to find the central network of response in rice. Our study identified 561 DEGs, in which 422 genes were upregulated and 139 genes were downregulated (Table S1). Our mata-analysis approach was validated by real-time PCR, and the expression results mainly confirmed it. The TIYF9 protein has previously been identified that regulating defense response and signaling pathway mediated by jasmonic acid in response to wounding in rice44. The Os01g0124650 is a family of serine-type endopeptidase inhibitors, which are known as Bowman-Birk inhibitors (BBI). They modulate endogenous proteolytic activities in different developmental stages and prevent exogenous proteases as an element of defense mechanisms in plants45. The BBI may also be engaged in multiple abiotic stress responses apart from their important role in defense against pathogenesis45. Shan et al. showed that the expression of WRSI5 increased in a salt-tolerant wheat cultivar under salt, drought, or oxidative stresses46.
Actin-depolymerizing factor family (ADF3) has been recognized as a stress-responsive protein47. It has been reported that overexpression of OsADF3 in Arabidopsis increased the tolerance to drought/osmotic stress by regulating some downstream responsive genes to abiotic stress48. A proteomics analysis of rice in 2006 showed a high level of ADFs expression in drought-tolerant cultivars49. The Rab proteins belong to the small G protein family, which are involved in different activities including intracellular signaling events, vesicle trafficking, various physiological processes, and stress response50. It has been reported that Dehydrin Rab16B is involved in response to abscisic acid, water deprivation, and cold acclimation44. Rab16 has been known as an abscisic acid (ABA)-responsive gene, which can sense the ABA and induce downstream stress signaling responses51. The Os05g0142900 is an unknown gene, and its expression was determined significantly high under different stresses when it was compared with the control treatment, suggesting a novel gene in response to abiotic stresses. Therefore, a functional analysis is needed to elucidate the potential function of this gene in response to stress.
In our study, the most enriched GO terms in biological processes were "response to stress" and "response to stimulus". Enriched terms in molecular function were "catalytic activity", "oxidoreductase activity", "DNA binding", and "transcription regulator activity". Among the stress-responsive genes, transcription factors are very important because the expression of other stress-responsive genes is regulated by their products by attaching to regulatory elements52. Ethylene response factor (ERF) and WRKY family (WRKY23, WRKY30, WRKY50, WRKY56, WRKY70, and WRKY71) had the highest number of genes among the enriched transcription factors in our study. The ERFs are a large subfamily of APETALA2 (Ap2) and are known for their Ap2 domain53. Ethylene is essential for many developmental processes and responds to biotic and abiotic stresses53. Recent reports have shown that different ERFs attach to dehydration-responsive factors (DREs) and act as key regulatory factors in plant responses to abiotic stresses54. It has been reported that the expression of ERF increased under drought, salinity, light, cold, and high-temperature stresses53. Jasmonic acid and abscisic acid are also involved in regulating the expression of ERFs under abiotic stresses55. The ethylene signaling pathway is also interconnected with other phytohormone pathways, which are regulated by salicylic acids, gibberellins, and brassinosteroids when plants are adapted to abiotic stresses55. Previously it was reported that the application of exogenous phytohormones also increased the expression of ERF genes55.
The WRKY proteins have an important function in cellular metabolism including the biosynthesis of phytohormones, phytoalexins, and other chemicals engaged in cellular defense56. The WRKY transcription factors regulate plant growth57 and play an important role in plants' responses to biotic and abiotic stresses58,59,60. The WRKY may correlate biotic stress-responsive proteins and abiotic stress-responsive proteins58. In our study, vesicle groups and cytoplasmic and vesicular membranes had the highest number of DEGs among cellular components. In plants, endomembrane trafficking is an essential mechanism that responds to environmental stresses61,62. Plants as immobile organisms continuously monitor environmental changes to be capable of altering their metabolism and gene expression profile in response to shifted conditions63. It has been widely indicated that plants have an effective response system to deal with stresses including the primary and secondary perception of stress and signal transmission in cells64. Early perception of stress occurs with changes in membrane leucine-rich receptors65. In addition, secretory pathways regulated by vesicular walls act in the stress tolerance mechanism. The first stage of the secretory pathway is regulated by coat protein II from the vesicular and induces the transfer of cargo, which usually are stress-damaged proteins, from the endoplasmic reticulum to the Golgi apparatus66.
The KEGG result showed that DEGs were mostly involved in "metabolic pathways", "biosynthesis of secondary metabolites", "transport of plant hormones", "biosynthesis of phenylpropanoids, nucleotide, and amino acids glucose metabolites" and "MAPK signaling pathway." It has been previously reported that plant metabolites protect plants against high salinity and drought stresses67. Metabolites such as glutathione, ascorbic acid, anthocyanins, tocopherols, and carotenoids protect plants from oxidative damage associated with different stresses by inhibiting reactive oxygen species (ROS) production during oxidative stress. Jasmonic acid, methyl jasmonate, salicylic acid, methyl salicylate, and other small molecules produced during stress conditions can also be activated as signaling molecules to induce defense responses and lead to a systemic adaptation67. Mitogen-Activated Protein Kinase (MAPK) are highly conserved signal transmission modules and are involved in in many signal transmission processes through the MAPK cascade68. The messages were transmitted from the extracellular into the cell by activating downstream kinases, enzymes, and transcription factors69. Recently it was reported that elevated expression of genes associated with the MAPK pathway increased the resistance to stress in crops70.
Our findings have identified candidate genes associated with various abiotic stresses, which can be further investigated to understand the core mechanisms underlying the response of rice to multiple stress conditions. Many individual studies have been done to investigate the response of rice to stresses but our meta-analysis covers a wide range of abiotic stresses. The meta-analysis approach can be used to study different plants' mechanisms in different situations, particularly for plants with less information. The current study includes abiotic stresses, suggesting that further research should concentrate on environmental and biological stresses for a better understanding of the relationship between abiotic and biotic response systems in plants.
Data availability
The datasets analyzed during the current study are available in the [NCBI] repository, [GEO Datasets] with accession numbers of GSE14275, GSE93917, GSE3053, GSE107531, GSE45724, GSE37940, GSE109649, GSE37161, GSE60823, GSE131287, and GSE25206.
References
MaghboliBalasjin, N., Maki, J. S., Schläppi, M. R. & Marshall, C. W. Plant growth-promoting activity of bacteria isolated from Asian rice (Oryza sativa L.) depends on rice genotype. Microbiol. Spectr. 10, e02787-12 (2022).
Sosunova, I. & Porras, J. IoT-enabled smart waste management systems for smart cities: A systematic review. IEEE Access 10, 73326–73363. https://doi.org/10.1109/ACCESS.2022.3188308 (2022).
Normile, D. Reinventing rice to feed the world. Science 321(5887), 330–333 (2008).
Ray, D. K., Gerber, J. S., MacDonald, G. K. & West, P. C. Climate variation explains a third of global crop yield variability. Nat. Commun. 6(1), 5989. https://doi.org/10.1038/ncomms6989 (2015).
Ben Rejeb, I., Pastor, V. & Mauch-Mani, B. Plant responses to simultaneous biotic and abiotic stress: Molecular mechanisms. Plants 3(4), 458–475 (2014).
Maiti, R. K. & Satya, P. Research advances in major cereal crops for adaptation to abiotic stresses. GM Crops Food 5(4), 259–279 (2014).
Zhang, Y. et al. Sensing of abiotic stress and ionic stress responses in plants. Int. J. Mol. Sci. 19(11), 3298 (2018).
Zhang, H., Zhu, J., Gong, Z. & Zhu, J.-K. Abiotic stress responses in plants. Nat. Rev. Genet. 23(2), 104–119. https://doi.org/10.1038/s41576-021-00413-0 (2022).
Jones, J. D. G. & Dangl, J. L. The plant immune system. Nature 444(7117), 323–329. https://doi.org/10.1038/nature05286 (2006).
Amar, D. et al. Time trajectories in the transcriptomic response to exercise—A meta-analysis. Nat. Commun. 12(1), 1–12 (2021).
Haidich, A.-B. Meta-analysis in medical research. Hippokratia 14(Suppl 1), 29 (2010).
Yi, G., Shin, H., Min, K. & Lee, E. J. Expanded transcriptomic view of strawberry fruit ripening through meta-analysis. PLoS ONE 16(6), e0252685 (2021).
de Abreu Neto, J. B. & Frei, M. Microarray meta-analysis focused on the response of genes involved in redox homeostasis to diverse abiotic stresses in rice. Front. Plant Sci. 6, 1260 (2016).
Das, K. & Roychoudhury, A. Reactive oxygen species (ROS) and response of antioxidants as ROS-scavengers during environmental stress in plants. Front. Environ. Sci. 2, 53 (2014).
Cohen, S. P. & Leach, J. E. Abiotic and biotic stresses induce a core transcriptome response in rice. Sci. Rep. 9(1), 1–11 (2019).
Buti, M. et al. A meta-analysis of comparative transcriptomic data reveals a set of key genes involved in the tolerance to abiotic stresses in rice. Int. J. Mol. Sci. 20(22), 5662 (2019).
Ramkumar, M. K. et al. Identification of major candidate genes for multiple abiotic stress tolerance at seedling stage by network analysis and their validation by expression profiling in rice (Oryza sativa L.). 3 Biotech 12(6), 1–14 (2022).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28(6), 882–883 (2012).
Zhang, Y., Parmigiani, G. & Johnson, W. E. ComBat-seq: batch effect adjustment for RNA-seq count data. NAR genom. bioinform. 2(3), lqaa078 (2020).
Walsh, M. R., Cooley, F. IV., Biles, K. & Munch, S. B. Predator-induced phenotypic plasticity within-and across-generations: a challenge for theory?. Proceed. Royal Society B: Biol. Sci. 282(1798), 20142205 (2015).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic acids res. 43(7), e47–e47 (2015).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13(11), 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Chin, C.-H. et al. cytoHubba: Identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 8(4), 1–7 (2014).
Langfelder, P. & Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 9(1), 1–13 (2008).
Zhang, B. & Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol. https://doi.org/10.2202/1544-6115.1128 (2005).
Yip, A. M. & Horvath, S. Gene network interconnectedness and the generalized topological overlap measure. BMC Bioinform. 8(1), 1–14 (2007).
Stein, R. J. et al. Genotype variation in rice (Oryza sativa L.) tolerance to Fe toxicity might be linked to root cell wall lignification. Front. Plant Sci. 10, 746 (2019).
Yoshida, S., Forno, D. A. & Cock, J. H. Laboratory Manual for Physiological Studies of Rice (1971).
Hu, W., Hu, G. & Han, B. Genome-wide survey and expression profiling of heat shock proteins and heat shock factors revealed overlapped and stress specific response under abiotic stresses in rice. Plant Sci. 176(4), 583–590 (2009).
Wang, X. et al. The NAD kinase OsNADK1 affects the intracellular redox balance and enhances the tolerance of rice to drought. BMC plant biol. 20(1), 1–19 (2020).
Walia, H. et al. Comparative transcriptional profiling of two contrasting rice genotypes under salinity stress during the vegetative growth stage. Plant physiol. 139(2), 822–835 (2005).
Li, G. Z. et al. Transcription factor WRKY 22 promotes aluminum tolerance via activation of Os FRDL 4 expression and enhancement of citrate secretion in rice (Oryza sativa). New Phytol. 219(1), 149–162 (2018).
Zhang, Y. et al. Transcriptome alteration in a rice introgression line with enhanced alkali tolerance. Plant Physiol. Biochem. 68, 111–117 (2013).
Zhang, F. et al. Genome-wide gene expression profiling of introgressed indica rice alleles associated with seedling cold tolerance improvement in a japonica rice background. BMC Genom. 13(1), 1–15 (2012).
Hsieh, P. H., Kan, C. C., Wu, H. Y., Yang, H. C. & Hsieh, M. H. Early molecular events associated with nitrogen deficiency in rice seedling roots. Sci. Rep. 8(1), 12207 (2018).
Ma, T. L., Wu, W. H. & Wang, Y. Transcriptome analysis of rice root responses to potassium deficiency. BMC Plant Biol. 12, 1–13 (2012).
Wang, Z. et al. Rice SPX1 and SPX2 inhibit phosphate starvation responses through interacting with PHR2 in a phosphate-dependent manner. Proc. Natl. Acad. Sci. 111(41), 14953–14958 (2014).
Stein, R. J. et al. Genotype variation in rice (Oryza sativa L.) tolerance to Fe toxicity might be linked to root cell wall lignification. Front. Plant Sci. 10, 746 (2019).
Dubey, S. et al. Transcriptomic and metabolomic shifts in rice roots in response to Cr (VI) stress. BMC Genom. 11(1), 1–19 (2010).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 25(4), 402–408 (2001).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51(D1), D587–D592. https://doi.org/10.1093/nar/gkac963 (2023).
Ghanbari Moheb Seraj, R. et al. Metabolomics analysis of milk thistle lipids to identify drought-tolerant genes. Sci. Rep. 12(1), 12827 (2022).
Hasanpour, K. et al. Identification of drought-tolerant hub genes in Iranian KC-2226 genotype of Aegilops tauschii using transcriptomic analysis. Sci. Rep. 13(1), 9499 (2023).
Gaudet, P., Livstone, M. S., Lewis, S. E. & Thomas, P. D. Phylogenetic-based propagation of functional annotations within the Gene Ontology consortium. Brief. Bioinform. 12(5), 449–462 (2011).
Xie, Y., Ravet, K. & Pearce, S. Extensive structural variation in the Bowman-Birk inhibitor family in common wheat (Triticum aestivum L.). BMC Genomics 22(1), 1–21 (2021).
Shan, L., Li, C., Chen, F., Zhao, S. & Xia, G. A Bowman-Birk type protease inhibitor is involved in the tolerance to salt stress in wheat. Plant. Cell Environ. 31(8), 1128–1137 (2008).
Hussey, P. J., Ketelaar, T. & Deeks, M. J. Control of the actin cytoskeleton in plant cell growth. Annu. Rev. Plant Biol. 57, 109–125 (2006).
Huang, Y.-C., Huang, W.-L., Hong, C.-Y., Lur, H.-S. & Chang, M.-C. Comprehensive analysis of differentially expressed rice actin depolymerizing factor gene family and heterologous overexpression of OsADF3 confers Arabidopsis thaliana drought tolerance. Rice 5(1), 1–14 (2012).
Ali, G. M. & Komatsu, S. Proteomic analysis of rice leaf sheath during drought stress. J. Proteome Res. 5(2), 396–403 (2006).
Choudhary, M. & Padaria, J. C. Transcriptional profiling in pearl millet (Pennisetum glaucum LR Br.) for identification of differentially expressed drought responsive genes. Physiol. Mol. Biol. Plants 21(2), 187–196 (2015).
Liang, Y., Tabien, R. E., Tarpley, L., Mohammed, A. R. & Septiningsih, E. M. Transcriptome profiling of two rice genotypes under mild field drought stress during grain-filling stage. AoB Plants 13(4), plab043 (2021).
Singh, D. & Laxmi, A. Transcriptional regulation of drought response: a tortuous network of transcriptional factors. Front. plant sci. 6, 895 (2015).
Müller, M. & Munné-Bosch, S. Ethylene response factors: A key regulatory hub in hormone and stress signaling. Plant Physiol. 169(1), 32–41. https://doi.org/10.1104/pp.15.00677 (2015).
Cheng, M.-C., Liao, P.-M., Kuo, W.-W. & Lin, T.-P. The Arabidopsis ETHYLENE RESPONSE FACTOR1 regulates abiotic stress-responsive gene expression by binding to different cis-acting elements in response to different stress signals. Plant Physiol. 162(3), 1566–1582 (2013).
Ma, Y. et al. Expressing a citrus ortholog of Arabidopsis ERF1 enhanced cold-tolerance in tobacco. Sci. Hortic. 174, 65–76 (2014).
Liang, X., Chen, X., Li, C., Fan, J. & Guo, Z. Metabolic and transcriptional alternations for defense by interfering OsWRKY62 and OsWRKY76 transcriptions in rice. Sci. Rep. 7(1), 1–15 (2017).
Tian, X. et al. Transcription factor OsWRKY53 positively regulates brassinosteroid signaling and plant architecture. Plant Physiol. 175(3), 1337–1349 (2017).
Lee, H. et al. Rice WRKY11 plays a role in pathogen defense and drought tolerance. Rice 11(1), 1–12 (2018).
Wang, H. et al. Overexpression of rice WRKY89 enhances ultraviolet B tolerance and disease resistance in rice plants. Plant Mol. Biol. 65(6), 799–815 (2007).
Yokotani, N. et al. WRKY76 is a rice transcriptional repressor playing opposite roles in blast disease resistance and cold stress tolerance. J. Exp. Bot. 64(16), 5085–5097 (2013).
Rosquete, M. R. & Drakakaki, G. Plant TGN in the stress response: A compartmentalized overview. Curr. Opin. Plant Biol. 46, 122–129 (2018).
Yu, F. & Xie, Q. Non-26S proteasome endomembrane trafficking pathways in ABA signaling. Trends Plant Sci. 22(11), 976–985 (2017).
Ding, Y., Shi, Y. & Yang, S. Advances and challenges in uncovering cold tolerance regulatory mechanisms in plants. New Phytol. 222(4), 1690–1704 (2019).
Zhang, S. et al. Cellulose synthesis genes CESA6 and CSI1 are important for salt stress tolerance in Arabidopsis. J. Integr. Plant Biol. 58(7), 623–626 (2016).
Yang, Y. & Guo, Y. Elucidating the molecular mechanisms mediating plant salt-stress responses. New Phytol. 217(2), 523–539 (2018).
Chung, K. P., Zeng, Y. & Jiang, L. COPII paralogs in plants: Functional redundancy or diversity?. Trends Plant Sci. 21(9), 758–769 (2016).
Shulaev, V., Cortes, D., Miller, G. & Mittler, R. Metabolomics for plant stress response. Physiol. Plant. 132(2), 199–208 (2008).
He, X., Wang, C., Wang, H., Li, L. & Wang, C. The function of MAPK cascades in response to various stresses in horticultural plants. Front. Plant Sci. 11, 952 (2020).
Zhang, M., Su, J., Zhang, Y., Xu, J. & Zhang, S. Conveying endogenous and exogenous signals: MAPK cascades in plant growth and defense. Curr. Opin. Plant Biol. 45, 1–10 (2018).
Šamajová, O., Plíhal, O., Al-Yousif, M., Hirt, H. & Šamaj, J. Improvement of stress tolerance in plants by genetic manipulation of mitogen-activated protein kinases. Biotechnol. Adv. 31(1), 118–128 (2013).
Acknowledgements
The first author would like to thank Zahra Sadat Eslami, Masoumeh Fallah, and Sanaz Moghadam for their guidance and assistance.
Author information
Authors and Affiliations
Contributions
M.A. (Mahnaz Azad) collected and analyzed the data, carried out laboratory work, and wrote the manuscript. M.T. (Masoud Tohidfar) designed the project and reviewed the manuscript. R.G.M.S. (Rahele Ghanbari Moheb Seraj) advice on writing the paper, editing, and scientific corrections of the paper. M.M. (Mohammad Mehralian) edited and validated the manuscript. K.E. (Keyvan Esmaeilzadeh-Salestani) validated the data, reviewed and edited the manuscript, and proofread the language. All authors have read and agreed to the submitted version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Azad, M., Tohidfar, M., Ghanbari Moheb Seraj, R. et al. Identification of responsive genes to multiple abiotic stresses in rice (Oryza sativa): a meta-analysis of transcriptomics data. Sci Rep 14, 5463 (2024). https://doi.org/10.1038/s41598-024-54623-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-54623-7
- Springer Nature Limited