Abstract
Depending on the source of the blastophore, there are various subtypes of laryngeal cancer, each with a unique metastatic risk and prognosis. The forecasting of their prognosis is a pressing issue that needs to be resolved. This study comprised 5953 patients with glottic carcinoma and 4465 individuals with non-glottic type (supraglottic and subglottic). Five clinicopathological characteristics of glottic and non-glottic carcinoma were screened using univariate and multivariate regression for CoxPH (Cox proportional hazards); for other models, 10 (glottic) and 11 (non-glottic) clinicopathological characteristics were selected using least absolute shrinkage and selection operator (LASSO) regression analysis, respectively; the corresponding survival models were established; and the best model was evaluated. We discovered that RSF (Random survival forest) was a superior model for both glottic and non-glottic carcinoma, with a projected concordance index (C-index) of 0.687 for glottic and 0.657 for non-glottic, respectively. The integrated Brier score (IBS) of their 1-year, 3-year, and 5-year time points is, respectively, 0.116, 0.182, 0.195 (glottic), and 0.130, 0.215, 0.220 (non-glottic), demonstrating the model's effective correction. We represented significant variables in a Shapley Additive Explanations (SHAP) plot. The two models are then combined to predict the prognosis for two distinct individuals, which has some effectiveness in predicting prognosis. For our investigation, we established separate models for glottic carcinoma and non-glottic carcinoma that were most effective at predicting survival. RSF is used to evaluate both glottic and non-glottic cancer, and it has a considerable impact on patient prognosis and risk factor prediction.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
Laryngeal carcinoma, which makes up 20% of all malignant tumors of the head and neck, is a prevalent type of these malignancies1,2. Laryngeal cancer is estimated to affect over 1,700,000 people annually, and 123,000 people died from it in 2019—86% of whom were men3,4. The majority of pathological kinds of laryngeal cancer, which include squamous cell carcinoma, are believed to be associated with smoking, alcohol use, human papillomavirus infection, and air pollution. Early detection and treatment are especially crucial for laryngeal cancer because it has an excellent prognosis and quality of life. Advanced laryngeal cancer can be treated with definitive Radiation therapy, combination chemotherapy, or adjuvant radiotherapy and total laryngectomy in addition to surgery, radiotherapy, chemotherapy, and other comprehensive treatments5,6. Different stages, forms, and treatments of laryngeal cancer have varying chances of survival, and the overall patient's 5-year relative survival rate is 64%2. Clinically, we use the conventional TNM staging and the 5-year survival rate to develop an overall understanding of the prognosis of patients, but it is challenging to quantify and depict it7. The conventional Cox proportional hazards (CoxPH) model is a prominent prediction model used to accomplish this goal, but it has limitations since it is based on the presumption that there is a linear relationship between survival outcomes and clinical variables8,9. At the same time, the standard survival analysis is unable to forecast an individual's survival prognosis with any degree of precision. The creation and prediction of tumor-related survival models now uses a large number of sophisticated Machine learning techniques (MLTs). When it comes to predicting the prognosis of tumor patients, MLTs—which include Random Survival Forest (RSF), Gradient Boosting Machine (GBM), eXtreme Gradient Boosting (XGBoost), and deep learning model Deepsurv—have been demonstrated to be more accurate than CoxPH model10,11,12,13,14. Studies on the equivalent machine learning of laryngeal cancer to create a survival model have been conducted concurrently15. Unfortunately, there are two flaws in the way that current research predicts the prognosis and survival of laryngeal cancer. First off, the performance of these models is constrained by the small sample size and homogeneity of the prediction algorithms. Furthermore, the lymph node metastatic rates of supraglottic and subglottic laryngeal carcinoma (non-glottic laryngeal carcinoma), which were 19.9% and 8.0% respectively, were higher. This contributed to the varied survival rates for glottic and non-glottic laryngeal cancer16,17. How to accurately predict the survival of patients with different types of laryngeal cancer has become a key problem. Therefore, in this study, we selected different types of laryngeal cancer patients using SEER database data, and developed survival models for glottic and non-glottic cancer, describing the main factors, to predict the survival of patients with laryngeal cancer more accurately. We develop the survival model using data from the SEER database, and compare the CoxPH model with four widely used machine learning techniques. Last but not least, we apply the model to forecast each person's prognosis, which aligns more with clinical application.
Methods
Data collection
The Surveillance, Epidemiology, and End Results Program (SEER) database was used to gather the study's data (Incidence-Seer Research Plus Data 17 Registries Nov 2021 Sub). Using the SEER*Stat program (version 8.4.2), we retrieved individuals who had been given a larynx carcinoma diagnosis by the third edition of the International Classification of Oncology Diseases (ICD-O-3). The period frame covers instances handled between 2000 and 2019. The following were the inclusion requirements: The behavior was identified as malignant and encoded by position and shape as "larynx".
Data clarity
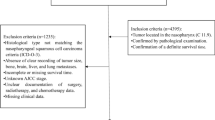
In total, 54,613 patients with primary laryngeal malignant tumors were included. The median follow-up duration of the sample in this study is 38 months. We used the following exclusion criteria to clean up the data: (1) Patients with limited follow-up information; (2) Patients without T stage (AJCC7), N stage (AJCC7), M stage (AJCC7), or AJCC stage grade information.
Feature selection
We selected variables that were directly related to the clinic, such as age, race, and gender, based on clinical experience. We chose the T stage, N stage, M stage, AJCC stage (AJCC stage 7), tumor size, and pathological categorization to assess the patient's health. Finally, to evaluate the patient's treatment plans, we also included radiation therapy, surgery, and chemotherapy.
Models for survival analysis
A classic model for survival analysis, the Cox proportional hazards (CoxPH) model has been the most commonly applied multifactor analysis technique in survival analysis to date18,19.
CoxPH is a statistical technique for survival analysis, which is mainly used to study the relationship between survival time and one or more predictors. The core of the model is the proportional risk hypothesis.
It is expressed as h(t|x) = h0 (t) exp (β|x), h(t|x) is the instantaneous risk function under the given covariable x, h0 (t) is the baseline risk function, on the other hand, exp (β x) represents the multiplicative effect of covariates on risk.
The random survival forest (RSF) model is an extremely efficient integrated learning model that can handle complex data linkages and is made up of numerous decision trees20.
RSF can improve the accuracy and robustness of the prediction, but it does not have a single expression because it is an integrated model consisting of multiple decision trees21. RSF constructs 1000 trees and calculates the importance of variables. To find the optimal model parameters, we adjust three key parameters: the maximum number of features of the tree (mtry), the minimum sample size of each node (nodesize), and the maximum depth of the tree (nodedepth). The values of these parameters are set to mtry from 1 to 10, nodesize from 3 to 30, and nodedepth from 3 to 6. We use a random search strategy (RandomSearch) to optimize the parameters. To evaluate the performance of the model under different parameter configurations, we use tenfold cross-validation and use C-index (ConcordanceIndex) as the evaluation index. The purpose of this process is to find the parameter configuration that can maximize the prediction accuracy of the model through many iterations.
One of the integrated learning methods called Boosting is the gradient boosting machine (GBM) model, which constructs a strong prediction model by combining several weak prediction models (usually decision trees). At each step, GBM adds a new weak learner by minimizing the loss function. The newly added model is trained to reduce the residual generated in the previous step, and the direction is determined by the gradient descent method. It can be expressed as Fm+1(x) = Fm(x) + αmhm(x). Where the Fm(x) is a weak model newly added, and the αm is the learning rate.
XGBoost is an efficient implementation of GBM, especially in optimizing computing speed and efficiency. To reuse the learner with the highest performance, it linearly combines the base learner with various weights22. eXtreme Gradient Boosting (XGBoost) is an optimization of the Gradient Boosting Decision Tree (GBDT), which boosts the algorithm's speed and effectiveness23. The neural network-based multi-task logic regression model developed by Deepsurv outperforms the conventional linear survival model in terms of performance24. DeepSurv uses a deep neural network to simulate the Cox proportional hazard model. Therefore, deepsurv can be expressed as h(t|x) = h0 (t) exp (g(x)), Where the g (x) is the output of the neural network, which represents the linear combination of the covariable x8.
Model training and validation
We categorize five models to adapt to various variable screening techniques used with various models. The RSF, GBM, and XGBoost models are screened using the least absolute shrinkage and selection operator (LASSO) regression analysis, while the CoxPH model is screened using the traditional Univariate and multivariate Cox regression analysis25,26,27.
In contrast, the Deepsurv model can automatically extract features and handle high-dimensional data and nonlinear relationships, so variable screening is not necessary28. We randomly split the data set into t and v datasets (training set and validation set) and test set in the ratio of 9:1 using spss (version 26) to further illustrate the model's dependability. Randomly selected 10% of the data as external verification. Once more, the ratio of 7:3 is used to divide the training set and validation set, and for both splits, the log-rank test is used to evaluate any differences between the two cohorts. The mlr3 package of R (version 4.2.2) uses the grid search approach to fine-tune the hyperparameters in the RSF, GBM, and XGBoost models in the validation set and chooses the most beneficial hyperparameters to build the survival model once the variables have been filtered following the aforementioned stages. Finally, the Deepsurv model is constructed using the Python (version 3.9) sksurv package, and the model is additionally optimized using grid search.
Model evaluation and interpretation
We used the integrated Brier score (IBS), which is appropriate for 1-year, 3-year, and 5-year time points, as the major assessment metric when evaluating the prediction performance of the model in the test set. In addition, the calibration curve is drawn and the conventional time-dependent receiver operating characteristic (ROC) curve as well as the area under the curve (AUC) (1 year, 3 years, and 5 years) are compared. By calculating the clinical net benefit to address the actual needs of clinical decisions, Decision Curve Analysis (DCA), a clinical evaluation prediction model, incorporates the preferences of patients or decision-makers into the analysis. Calculating the various clinicopathological characteristics is also required for the prognosis of contribution. We visualized the survival contribution of several clinicopathological characteristics for 1-year, 3-years, and 5-years using The Shapley Additive Explanations (SHAP) plot.
The particular prediction
Clinically speaking, various individuals require personalized care. Consequently, it is crucial to estimate the likelihood that a single patient will survive. The survival probability of a certain patient is predicted using the ggh4x package of R (version 4.2.2), along with the contribution of several clinicopathological characteristics to survival. This has major clinical work implications.
Results
Baseline characteristics
The information of 54,613 patients was included. After data cleaning, there were 5953 patients with glottic carcinoma and 4465 patients with non-glottic (supraglottic and subglottic) cancer as a result of the aforementioned exclusion criteria. Figure 1 shows specific cleaning procedures. Table 1 displays the clinical and clinicopathological characteristics of these patients as well as the relevant categorization ratio. In Fig. 2, the survival curve was displayed after patients with glottic and non-glottic cancer were divided into training and validation datasets and testing datasets, respectively.
Feature selection and model construction
Age, histology, tumor size, RN Eval (Regular Nodes Evaluation), AJCC T, AJCC N, AJCC M, AJCC Stage, surgery, and chemotherapy were the 10 significant mutations identified in the univariate Cox regression analysis for Glottic Carcinoma. Following multivariate Cox regression, age, AJCC T, AJCC N, AJCC Stage, and surgery were the final 5 effective variables to be included. Similarly, the effective variables of univariate Cox regression analysis and multivariate Cox regression analysis of non-glottic laryngeal carcinoma were age, sex, tumor size, RN Eval, AJCC T, AJCC N, AJCC M, AJCC Stage, surgery, radiotherapy and age, sex, AJCC Stage, surgery, radiotherapy. Table S1 displays the outcomes of the univariate and multivariate Cox regression. Machine learning characteristic variables (Fig. 3) were chosen using lasso regression analysis based on the lowest standard. A total of 10 efficient variables were chosen for glottic carcinoma: age, sex, histology, AJCC T, AJCC N, AJCC Stage, RN Eval, radiotherapy, surgery, and tumor size. The 11 effective variables were chosen for non-glottic carcinoma: age, sex, histology, AJCC T, AJCC N, AJCC M, AJCC Stage, chemotherapy, radiotherapy, surgery, and tumor size.
Constructing and evaluating survival analysis models
We built the CoxPH model, RSF model, GBM model, and XGBoost model for glottic and non-glottic cancer by the outcomes of multivariate Cox regression analysis and lasso regression analysis, respectively. The Deepsurv model does not require variable screening, hence all 12 variables are used in the model during model development. All survival models are trained to roughly estimate their performance and stability range using Ten-fold cross-validation C-index and IBS. Following the visual examination of the test set, we eventually obtained the following secondary outcomes: 1-year, 3-year, and 5-year ROC curve, calibration curve, C-index, and 1-year, 3-year, and 5-year IBS (Table 2 and Fig. 4). The indicators of the training set are shown in Table S2. All models have IBS that are less than 0.25, which suggests that they can be calibrated well. Figure 5 displays the 1-year, 3-year, and 5-year DCA decision curves for the best model RSF at the same period.
The chart "RSF Model for predicting 3-and 5-year Survival rates of Laryngeal Cancer patients: calibration Curve and time-dependent ROC Curve" shows the calibration curve and time-dependent ROC curve of RSF model for predicting 3-and 5-year survival rates of laryngeal cancer patients. The calibration curve shows the consistency between the predicted survival rate and the actual survival rate, while the ROC curve provides the performance of the model under different discriminant thresholds. Month is the unit of time. (a) glottic carcinoma, (b) non-glottic carcinoma.
The 3-year and 5-year decision-making curves are based on the RSF model. The decision curve shows the net benefit of the model prediction under different patient risk thresholds. By comparing the net income of model prediction with and without model prediction under a specific risk threshold, the application value of the model in clinical decision-making can be evaluated. In the figure, the horizontal axis represents the decision threshold and the vertical axis represents the net income. The points on the curve represent the relative net income that the model can predict under a given risk threshold. Ideally, the higher the curve, the greater the net income provided by the model within a wider threshold range, that is, the higher the value of the model in clinical application. (a) glottic carcinoma, (b) non-glottic carcinoma.
Visualization of the optimal model's evaluation indexes
The RSF model is the most effective one for both glottic and non-glottic carcinomas, and its C-index in the test set is 0.687 for glottic and 0.657 for non-glottic, respectively. Their 1-year, 3-year, and 5-year IBS were 0.116, 0.182, and 0.195 for glottic carcinomas, and 0.130, 0.215, and 0.220 for non-glottic carcinomas, respectively. Figure 6 depicts the impact of several clinicopathological characteristics on patient survival for the RSF model of two subtypes of laryngeal cancer. AJCC Stage, age, and AJCC T are the first three factors that have the greatest impact on glottic carcinoma. And it is AJCC Stage, age, surgery for non-glottic cancer.
The SHAP plot of the RSF model. The vertical axis lists many clinical characteristics, while the horizontal axis shows how the variable affected the outcomes. The likelihood of dying increases with a feature's SHAP value. The picture reflects the Shap values predicted by 1-year,3-year, and 5 years. (a) glottic carcinoma, (b) non-glottic carcinoma.
The particular forecast
Two patients with glottic carcinoma and two individuals with non-glottic carcinoma were chosen at random. Their clinicopathological data is listed below.
Glottic cancer: Patient 1 is a male, aged 70 to 79 years, with non-squamous cell carcinoma, T3N0, AJCC III, undergoing surgery and radiotherapy; tumor size is unknown; Patient 2 is a male, aged 60 to 69 years, with squamous cell carcinoma, T3N2c, AJCC IVA, not undergoing surgery and undergoing radiotherapy; tumor size is less than 1 cm.
Non-glottic laryngeal carcinoma: Patient 1 is a male, 50–59 years old, squamous cell cancer, T1N0M0, AJCC I, surgery, radiation, chemotherapy, tumor size less than 1 cm; Patient 2: Male, 50–59 years of age, squamous cell carcinoma, T2N3M0, AJCCIVB, surgery, radiation, no chemotherapy, tumor size less than 1 cm. Figure 7 depicts their unique forecasting chart.
Survival prediction of individual patients. The horizontal axis indicates various ages (1-year, 3-year, 5-year), the vertical axis shows the contribution to survival, while the lines of various hues represent various clinical parameters. Less than 0 indicates a detrimental contribution to survival, whereas more than 0 indicates a beneficial one. (a) glottic carcinoma, (b) non-glottic carcinoma.
Discussion
In otorhinolaryngology, head and neck surgery, laryngeal carcinoma is a common malignant tumor. Early laryngeal cancer has an occult quality. As examination and treatment techniques advance, the fibrolaryngoscope, for instance, is being used more frequently in clinics to play a significant role in early screening for laryngeal cancer. Nevertheless, since many patients ignore early symptoms such as hoarseness and throat discomfort, many people will mistake them for chronic diseases including chronic pharyngitis, leading to delayed diagnosis and treatment. More than 60% of patients are diagnosed with advanced cancer, based on studies, which significantly lowers the efficiency of laryngeal cancer treatment2.
Despite postoperative adjuvant radiotherapy and chemotherapy do not have a favorable prognosis, patients with advanced laryngeal cancer with lymph node metastases sometimes undergo partial laryngectomy or even total laryngectomy. Patients with early laryngeal cancer have a decent prognosis, and their quality of life will significantly diminish as a consequence of total laryngectomy29. The physical foundation and developmental base of the larynx are unique. Glottic type, supraglottic type, and subglottic type are three subtypes of laryngeal cancer. The glottic area and subglottic region's structure is derived from the storage trachea germ base, whereas the supraglottic region's structure is derived from the oropharynx germ. As a result, these two regions have different fibrous fascia and lymphatic drainage systems. Clinically, glottic and non-glottic cancer have significantly different risks of lymph nodes and distant metastases. The likelihood of survival varies significantly between various forms of laryngeal cancer, too.
There is some research on the likelihood of surviving laryngeal cancer, however, the majority of them use the outdated Kaplan–Meier estimator survival model (Kaplan and Meier, 1958), which is unable to incorporate the patient's variables. The obvious drawback of the KM survival model is that diverse clinicopathological variables influence how the tumor develops and evolves5,30.
The CoxPH model, which can handle censored and censored data as well as continuous and sub-type variables, is suggested as a solution to the problem of covariable fitting. The most used model for predicting survival, the CoxPH model, measures the effect of covariables on survival time using a partial regression coefficient and risk ratio. CoxPH model's assumptions that the risk rate is constant and that the logarithm of the risk rate is a linear function of the covariable are limiting. If the hypothesis is incorrect, the prediction will be biased8,31.
Machine learning-based survival analysis has been increasingly used in recent years to forecast the survival of tumor patients. Machine learning can handle complex, nonlinear data, extract relevant features and information, and enhance the model's generalizability and accuracy. Machine learning does not need to make as many assumptions about data distribution or risk functions as the CoxPH model requires. More significantly, we can estimate each patient's survival after the development of a machine-learning model, which has enormous clinical importance. Different machine learning algorithms can be used to handle various types of data and have varying properties. Based on the CoxPH model's survival analysis, RSF, XGBoost, GBM model, and deep learning model Deepsurv are added in this work.
Many academics have discovered that RSF is a survival analysis model with good performance in earlier investigations. It creates several decision trees by self-sampling and combines the outcomes of each tree's predictions by voting or averaging32,33.
Of course, similar machine learning studies on the survival analysis of head and neck malignancies, such as laryngeal cancer, hypopharyngeal carcinoma, oropharyngeal carcinoma, and nasopharyngeal carcinoma, have been conducted by various researchers34,35. However, as previously mentioned, different embryonic sources and fibrous fascia tissues cause laryngeal carcinoma to be divided into different subtypes. Because previous studies have not distinguished between different subtypes, there will inevitably be a discrepancy between the expected results and the actual results. Based on this, we created the five survival analysis models mentioned above, one for glottic carcinoma and the other for non-glottic carcinoma. RSF is an excellent model, to sum up. The C-index of two separate subtypes of laryngeal carcinoma RSF reached 0.687 and 0.657, respectively, in the final test set. The integrated Brier score (IBS) of their 1-year, 3-year, and 5-year time points is, respectively, 0.116, 0.182, 0.195 (glottic type), and 0.130, 0.215, 0.220 (non-glottic type). This demonstrates the RSF model's high degree of reliability and strengthens our conclusion. The SHAP plot can also more easily convey how risk factors affect specific survival outcomes when compared to the conventional CoxPH analysis nomogram plot. Furthermore, using the RSF machine learning model, we can build the individual survival probability curve for any patient and display their survival prognosis in a more precise manner. This raises the study's clinical relevance even further.
This study has some limitations. First of all, glottic carcinoma and supraglottic carcinoma account for the vast majority of subtypes of laryngeal carcinoma, and because there is a dearth of data on subglottic type, we are unable to develop a survival analysis model for subglottic carcinoma alone. It can only be split into glottic type and non-glottic type as a result. Theoretically, a more precise division results in a more precise forecast. Second, while not terrible, our model C-index still has to be refined by academics.
In conclusion, we compared the prognostic value of patients with various subtypes of laryngeal cancer using five survival prediction model algorithms, and we selected the best RSF algorithm based on which we established survival prognosis prediction for patients with two subtypes of laryngeal cancer, model it and depict it. To advance customized medicine, we also give professionals a tailored patient prognosis prediction model at the same time. Our research demonstrates that the RSF algorithm offers promising therapeutic potential for the prognostic prediction of laryngeal cancer.
Data availability
The data used in this study are available from the Surveillance, Epidemiology, and End Results Program (SEER) database (Incidence-Seer Research Plus Data 17 Registries Nov 2021 Sub).
Abbreviations
- CoxPH:
-
Cox proportional hazards
- RSF:
-
Random survival forest
- GBM:
-
Gradient boosting machine
- XGBoost:
-
EXtreme gradient boosting
- MLTs:
-
Machine learning techniques
- SEER:
-
The surveillance, epidemiology, and end results program
- IBS:
-
Integrated brier score
- LASSO:
-
Least absolute shrinkage and selection operator
- C-index:
-
The concordance index
References
Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. Cancer statistics, 2021. CA. Cancer J. Clin. 71, 7–33 (2021).
Santos, A., Santos, I. C., Dos Reis, P. F., Rodrigues, V. D. & Peres, W. A. F. Impact of nutritional status on survival in head and neck cancer patients after total laryngectomy. Nutr. Cancer 74, 1252–1260 (2022).
GBD Respiratory Tract Cancers Collaborators. Global, regional, and national burden of respiratory tract cancers and associated risk factors from 1990 to 2019: A systematic analysis for the Global Burden of Disease Study 2019. Lancet Respir. Med. 9, 1030–1049 (2021).
Zhao, Y., Qin, J., Qiu, Z., Guo, J. & Chang, W. Prognostic role of neutrophil-to-lymphocyte ratio to laryngeal squamous cell carcinoma: A meta-analysis. Braz. J. Otorhinolaryngol. 88, 717–724 (2022).
Forastiere, A. A. et al. Long-term results of RTOG 91–11: A comparison of three nonsurgical treatment strategies to preserve the larynx in patients with locally advanced larynx cancer. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 31, 845–852 (2013).
Kim, D. H. et al. The prognostic utilities of various risk factors for laryngeal squamous cell carcinoma: A systematic review and meta-analysis. Med. Kaunas Lith. 59, 497 (2023).
Ju, J. et al. Nomograms predicting long-term overall survival and cancer-specific survival in head and neck squamous cell carcinoma patients. Oncotarget 7, 51059–51068 (2016).
Katzman, J. L. et al. DeepSurv: Personalized treatment recommender system using a Cox proportional hazards deep neural network. BMC Med. Res. Methodol. 18, 24 (2018).
Kim, D. W. et al. Deep learning-based survival prediction of oral cancer patients. Sci. Rep. 9, 6994 (2019).
Lin, J. et al. The development of a prediction model based on random survival forest for the postoperative prognosis of pancreatic cancer: A SEER-based study. Cancers 14, 4667 (2022).
Hatano, Y., Ishihara, T., Hirokawa, S. & Onodera, O. Machine learning approach for the prediction of age-specific probability of SCA3 and DRPLA by survival curve analysis. Neurol. Genet. 9, e200075 (2023).
Thio, Q. C. B. S. et al. Can machine-learning techniques be used for 5-year survival prediction of patients with chondrosarcoma?. Clin. Orthop. 476, 2040–2048 (2018).
Ji, G.-W. et al. Development and validation of a gradient boosting machine to predict prognosis after liver resection for intrahepatic cholangiocarcinoma. BMC Cancer 22, 258 (2022).
Song, Y. et al. Multiple machine learnings revealed similar predictive accuracy for prognosis of PNETs from the surveillance, epidemiology, and end result database. J. Cancer 9, 3971–3978 (2018).
Howard, F. M., Kochanny, S., Koshy, M., Spiotto, M. & Pearson, A. T. Machine learning-guided adjuvant treatment of head and neck cancer. JAMA Netw. Open 3, e2025881 (2020).
Sanabria, A. et al. Incidence of occult lymph node metastasis in primary larynx squamous cell carcinoma, by subsite, T classification and neck level: A systematic review. Cancers 12, 1059 (2020).
Coskun, H. et al. Prognosis of subglottic carcinoma: Is it really worse?. Head Neck 41, 511–521 (2019).
Zhang, K. et al. Machine learning-based prediction of survival prognosis in esophageal squamous cell carcinoma. Sci. Rep. 13, 13532 (2023).
Cygu, S., Seow, H., Dushoff, J. & Bolker, B. M. Comparing machine learning approaches to incorporate time-varying covariates in predicting cancer survival time. Sci. Rep. 13, 1370 (2023).
Segev, N., Harel, M., Mannor, S., Crammer, K. & El-Yaniv, R. Learn on source, refine on target: A model transfer learning framework with random forests. IEEE Trans. Pattern Anal. Mach. Intell. 39, 1811–1824 (2017).
Strobl, C., Boulesteix, A.-L., Zeileis, A. & Hothorn, T. Bias in random forest variable importance measures: Illustrations, sources and a solution. BMC Bioinform. 8, 25 (2007).
Dash, T. K., Chakraborty, C., Mahapatra, S. & Panda, G. Gradient boosting machine and efficient combination of features for speech-based detection of COVID-19. IEEE J. Biomed. Health Inform. 26, 5364–5371 (2022).
Yuan, K.-C. et al. The development an artificial intelligence algorithm for early sepsis diagnosis in the intensive care unit. Int. J. Med. Inf. 141, 104176 (2020).
Kim, S. I., Kang, J. W., Eun, Y.-G. & Lee, Y. C. Prediction of survival in oropharyngeal squamous cell carcinoma using machine learning algorithms: A study based on the surveillance, epidemiology, and end results database. Front. Oncol. 12, 974678 (2022).
Chen, D. et al. Integrated machine learning and bioinformatic analyses constructed a novel stemness-related classifier to predict prognosis and immunotherapy responses for hepatocellular carcinoma patients. Int. J. Biol. Sci. 18, 360–373 (2022).
Ni, M. et al. Radiomics models for diagnosing microvascular invasion in hepatocellular carcinoma: Which model is the best model?. Cancer Imaging Off. Publ. Int. Cancer Imaging Soc. 19, 60 (2019).
Ding, T. et al. Assessment and quantification of ovarian reserve on the basis of machine learning models. Front. Endocrinol. 14, 1087429 (2023).
She, Y. et al. Development and validation of a deep learning model for non-small cell lung cancer survival. JAMA Netw. Open 3, e205842 (2020).
Department of Veterans Affairs Laryngeal Cancer Study Group. Induction chemotherapy plus radiation compared with surgery plus radiation in patients with advanced laryngeal cancer. N. Engl. J. Med. 324, 1685–1690 (1991).
Du, X. et al. Marital status and survival in laryngeal squamous cell carcinoma patients: A multinomial propensity scores matched study. Eur. Arch. Oto-Rhino-Laryngol. 279, 3005–3011 (2022).
Chansky, K. et al. The International Association for the Study of Lung Cancer Staging Project. Prognostic factors and pathologic TNM stage in surgically managed non-small cell lung cancer. Zhongguo Fei Ai Za Zhi Chin. J. Lung Cancer 13, 9–18 (2010).
Choi, Y. S. et al. Machine learning and radiomic phenotyping of lower grade gliomas: Improving survival prediction. Eur. Radiol. 30, 3834–3842 (2020).
Taylor, J. M. G. Random survival forests. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 6, 1974–1975 (2011).
Peng, J. et al. The prognostic value of machine learning techniques versus cox regression model for head and neck cancer. Methods San Diego Calif. 205, 123–132 (2022).
Kotevski, D. P., Smee, R. I., Vajdic, C. M. & Field, M. Machine learning and nomogram prognostic modeling for 2-year head and neck cancer-specific survival using electronic health record data: A multisite study. JCO Clin. Cancer Inform. 7, e2200128 (2023).
Funding
This work was funded by Natural Science Foundation of Fujian Science and Technology Department, 2020J02060 and Key Medical and Health Project of Xiamen Science and Technology Bureau, 3502Z20204009.
Author information
Authors and Affiliations
Contributions
Wei Wang conceived and designed the study. Wei Wang, Yongjun Hong analyzed the data and wrote the manuscript. Wei Wang and Wenhui Wang contributed equally to this work. Dongdong Zhang, Peiji Zeng, Yue Wang, Min Lei, and Wenhui Wang collected the data. Chengfu Cai, Dongdong Zhang, and Peiji Zeng modified the manuscript. Chengfu Cai made the final review and correction of this article. All authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, W., Wang, W., Zhang, D. et al. Creation of a machine learning-based prognostic prediction model for various subtypes of laryngeal cancer. Sci Rep 14, 6484 (2024). https://doi.org/10.1038/s41598-024-56687-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-56687-x
- Springer Nature Limited